Modular Engineering of Biomass Degradation Pathways
Abstract
1. Introduction
2. Lignocellulose Composition
2.1. Cellulose Composition
2.2. Hemicellulose Composition
2.3. Lignin Composition
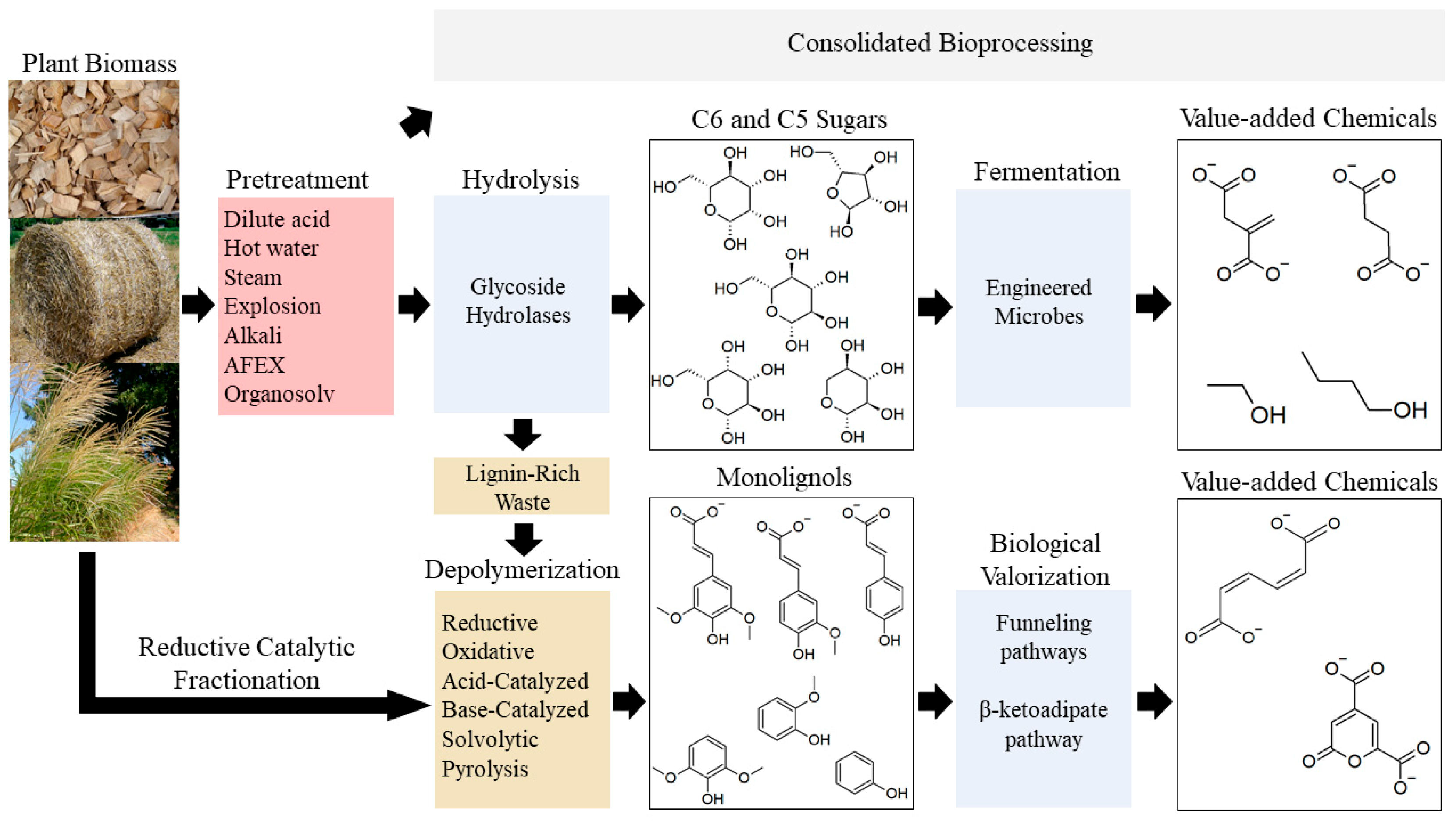
3. Pathways for Microbial Catabolism of Lignocellulose
3.1. Catabolism of Cellulose & Hemicellulose-Derived Compounds
3.1.1. Glucose
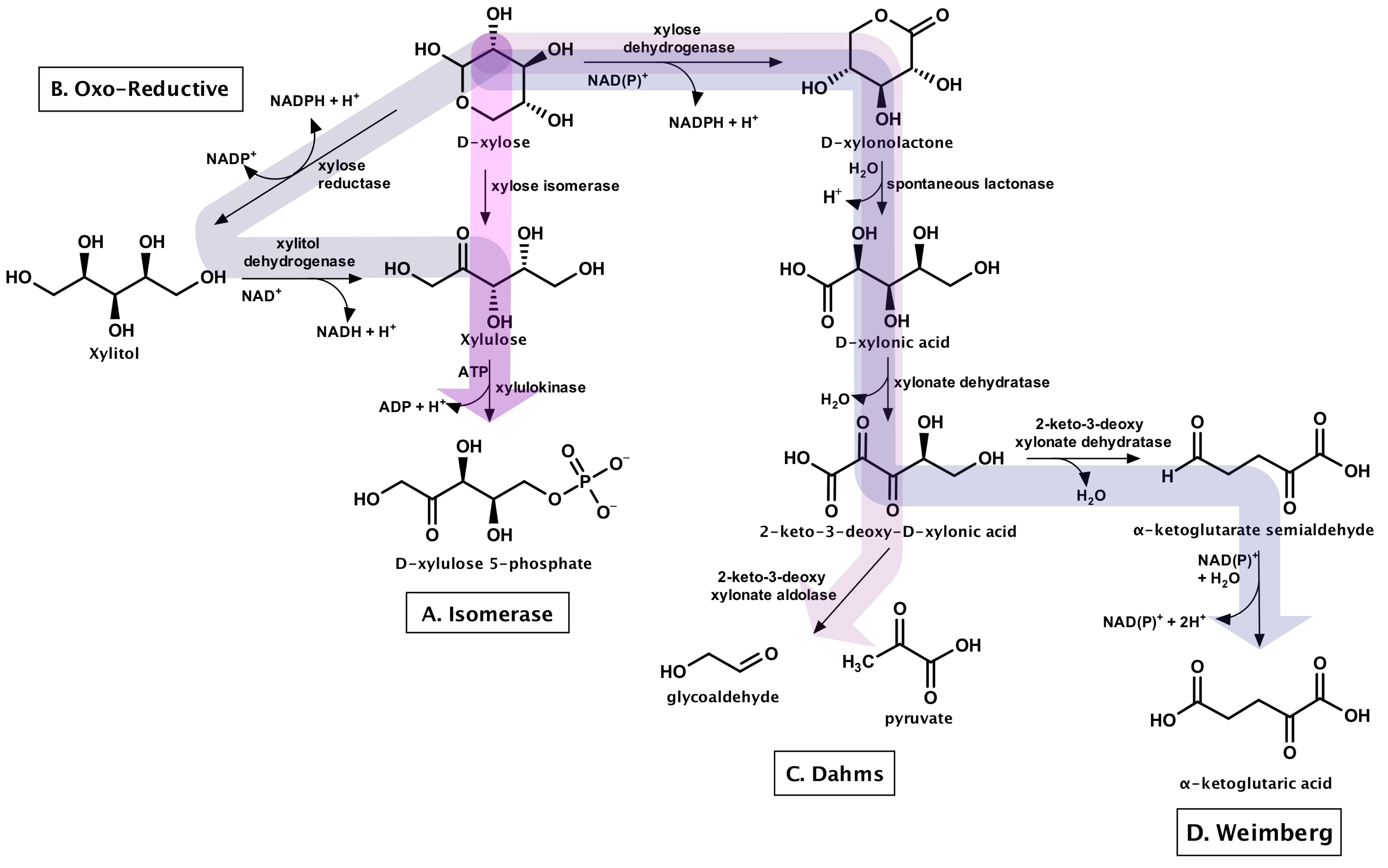
3.1.2. Xylose
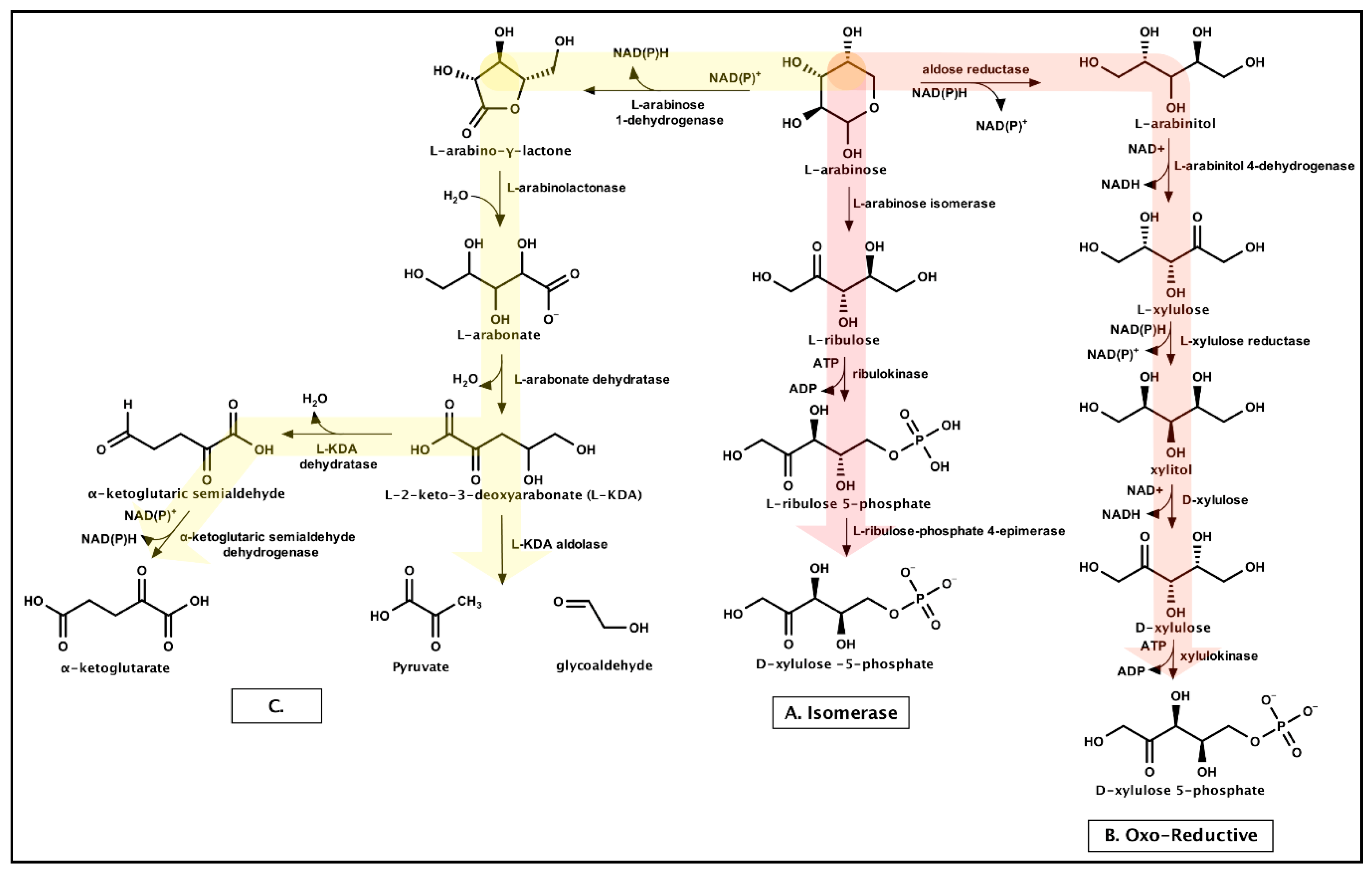
3.1.3. Arabinose
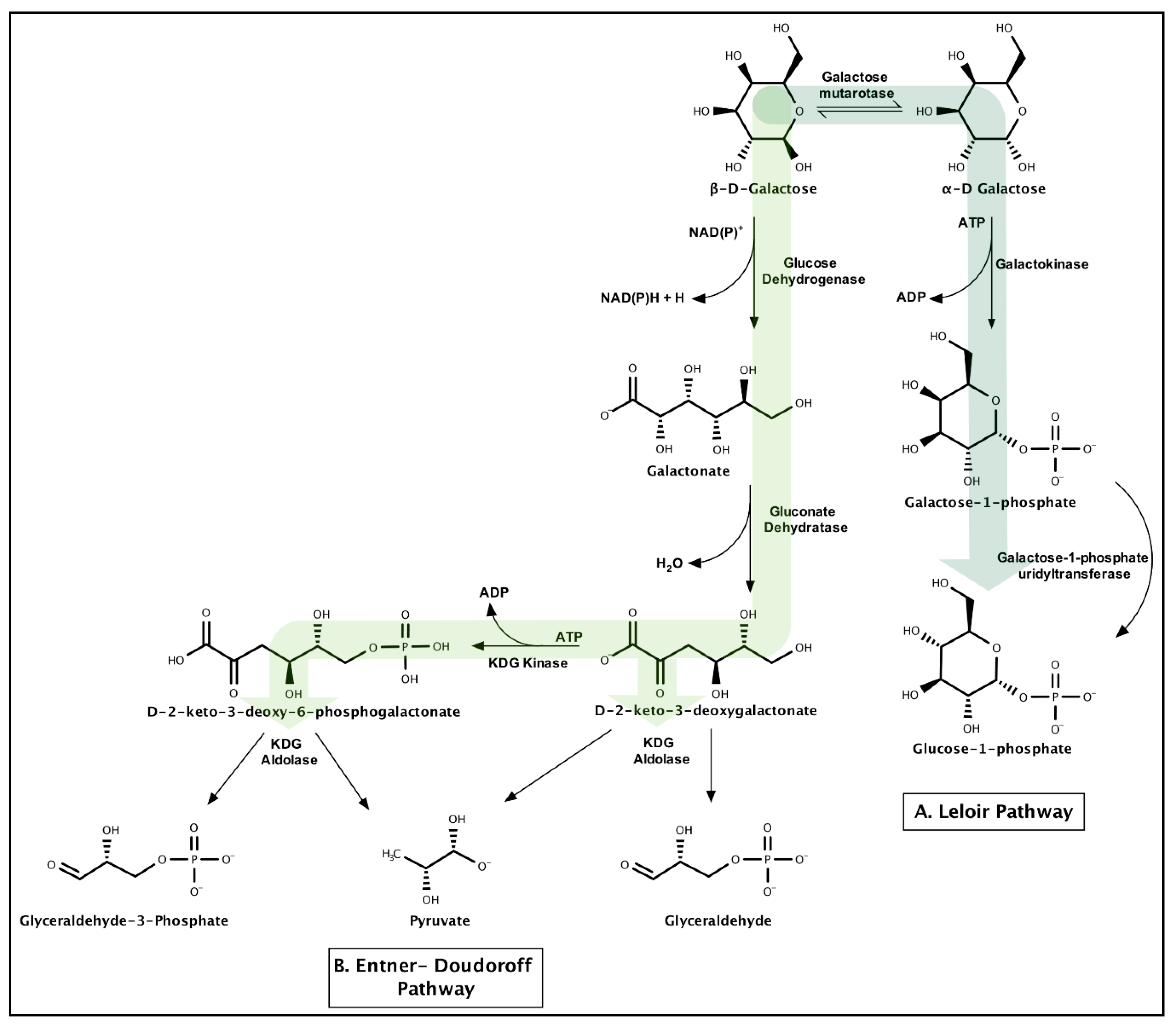
3.1.4. Galactose
3.1.5. Other Hemicellulosic Compounds
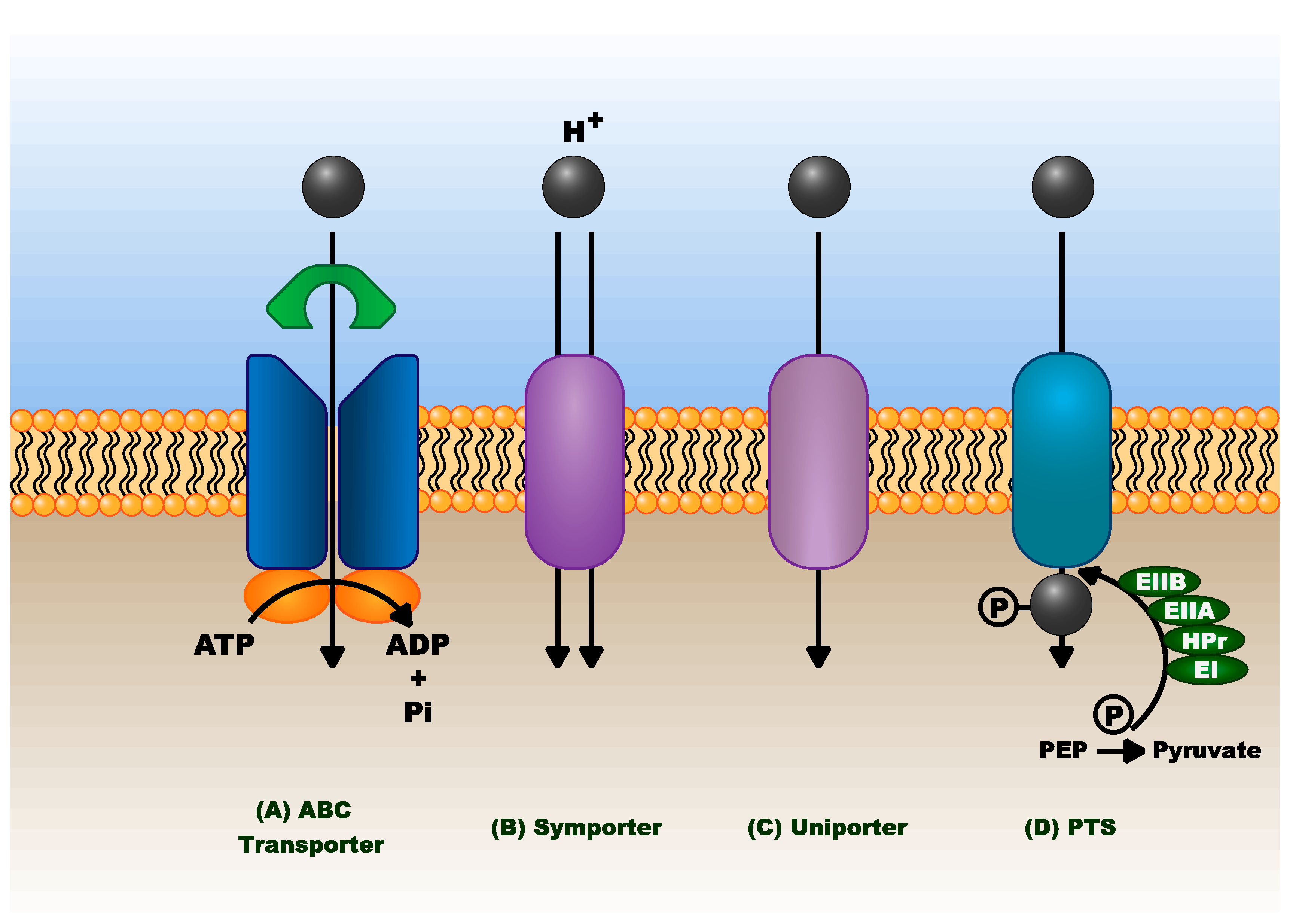
3.1.6. Sugar Transport
3.1.7. Regulation of Sugar Catabolism
3.2. Catabolism of Lignin-Derived Compounds
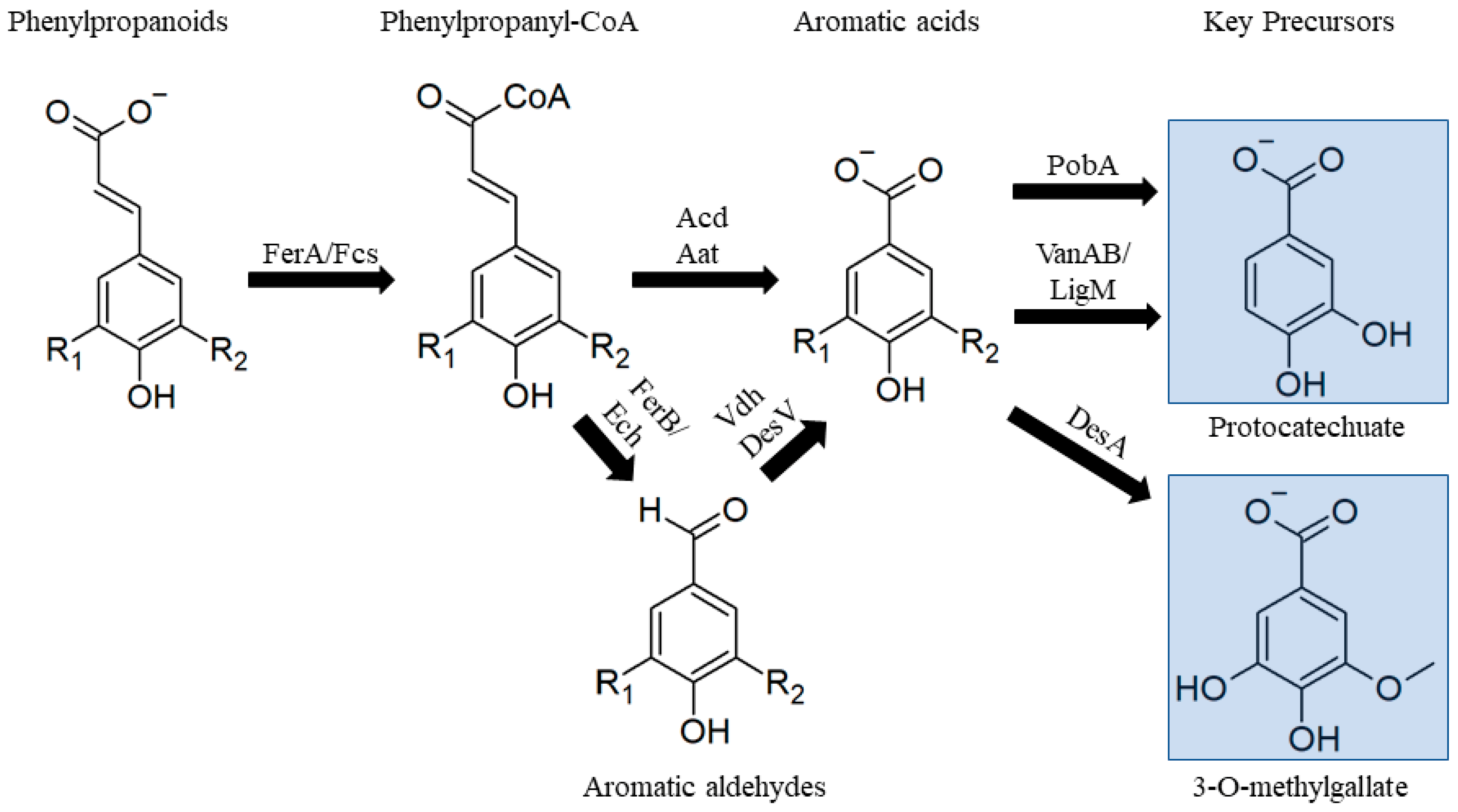
3.2.1. Phenylpropanoids
3.2.2. Guaiacol
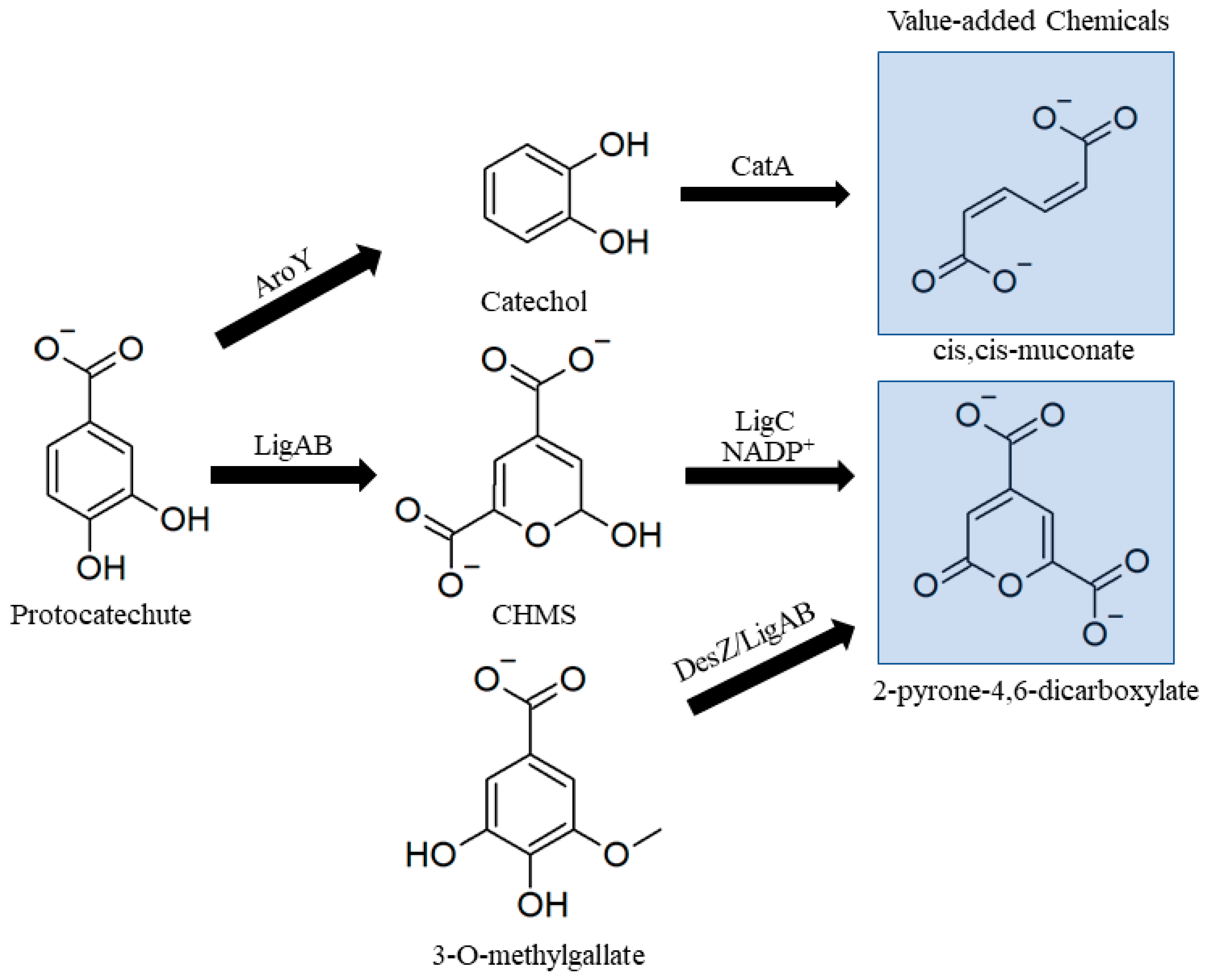
3.2.3. Lower Catabolic Pathways for Biological Valorization
3.2.4. Transport of Aromatic Compounds
4. Challenges to Engineering Catabolic Pathways
4.1. Transport
4.2. Redox & Energy Balance/Co-Factor Preference
4.3. Regulation
4.4. Expression and Activity of Metabolic Pathway Enzymes
4.5. Interactions with Native Metabolic Pathways
5. Application of Modular Design for Carbon Catabolic Pathway Engineering
5.1. Characterization of Engineered Pathways in Diverse Hosts and Combinations
5.2. Minimizing Interactions with the Host or Other Pathways
5.3. Tolerating Host and Pathway Variability
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Langholtz, M.H.; Stokes, B.J.; Eaton, L.M. 2016 Billion-Ton Report: Advancing Domestic Resources for a Thriving Bioeconomy, Volume 1: Economic Availability of Feedstocks; U.S. Department of Energy’s Office of Energy Efficiency and Renewable Energy: Oak Ridge, TN, USA, 2016.
- Dutta, A. Forecasting ethanol market volatility: New evidence from the corn implied volatility index. Biofuels Bioprod. Biorefining 2019, 13, 48–54. [Google Scholar] [CrossRef]
- Bajpai, P. Pretreatment of Lignocellulosic Biomass for Biofuel Production; Springer: Singapore, 2016. [Google Scholar] [CrossRef]
- Keegstra, K. Plant cell walls. Plant Physiol. 2010, 154, 483–486. [Google Scholar] [CrossRef]
- McCann, M.; Rose, J.; Bauer, W.; Albersheim, P. Blueprints for Building Plant Cell Walls. Plant Physiol. 2010, 153, 365. [Google Scholar] [CrossRef]
- Vorwerk, S.; Somerville, S.; Somerville, C. The role of plant cell wall polysaccharide composition in disease resistance. Trends Plant Sci. 2004, 9, 203–209. [Google Scholar] [CrossRef] [PubMed]
- Barakat, A.; Mayer-Laigle, C.; Solhy, A.; Arancon, R.A.D.; de Vries, H.; Luque, R. Mechanical pretreatments of lignocellulosic biomass: Towards facile and environmentally sound technologies for biofuels production. RSC Adv. 2014, 4, 48109–48127. [Google Scholar] [CrossRef]
- Garrote, G.; Domínguez, H.; Parajó, J.C. Hydrothermal processing of lignocellulosic materials. Holz als Roh- und Werkst. 1999, 57, 191–202. [Google Scholar] [CrossRef]
- Vom Stein, T.; Grande, P.M.; Kayser, H.; Sibilla, F.; Leitner, W.; Domínguez de María, P. From biomass to feedstock: One-step fractionation of lignocellulose components by the selective organic acid-catalyzed depolymerization of hemicellulose in a biphasic system. Green Chem. 2011, 13, 1772. [Google Scholar] [CrossRef]
- Rashid, B.; Leman, Z.; Jawaid, M.; Ghazali, M.J.; Ishak, M.R. Physicochemical and thermal properties of lignocellulosic fiber from sugar palm fibers: Effect of treatment. Cellulose 2016, 23, 2905–2916. [Google Scholar] [CrossRef]
- Vasco-Correa, J.; Ge, X.; Li, Y. Biological Pretreatment of Lignocellulosic Biomass. Biomass Fractionation Technol. a Lignocellul. Feed. Based Biorefinery 2016, 561–585. [Google Scholar] [CrossRef]
- Olson, D.G.; McBride, J.E.; Joe Shaw, A.; Lynd, L.R. Recent progress in consolidated bioprocessing. Curr. Opin. Biotechnol. 2012, 23, 396–405. [Google Scholar] [CrossRef]
- Salvachua, D.; Karp, E.M.; Nimlos, C.T.; Vardon, D.R.; Beckham, G.T. Towards lignin consolidated bioprocessing: Simultaneous lignin depolymerization and product generation by bacteria. Green Chem. 2015, 17, 4951–4967. [Google Scholar] [CrossRef]
- Yaguchi, A.; Spagnuolo, M.; Blenner, M. Engineering yeast for utilization of alternative feedstocks. Curr. Opin. Biotechnol. 2018, 53, 122–129. [Google Scholar] [CrossRef]
- Sun Yim, S.; Woong Choi, J.; Hwa Lee, S.; Jun Jeong, K. Modular Optimization of a Hemicellulose-Utilizing Pathway in Corynebacterium glutamicum for Consolidated Bioprocessing of Hemicellulosic Biomass. ACS Synth. Biol. 2016. [Google Scholar] [CrossRef]
- Kim, S.M.; Choi, B.Y.; Ryu, Y.S.; Jung, S.H.; Park, J.M.; Kim, G.-H.; Lee, S.K. Simultaneous utilization of glucose and xylose via novel mechanisms in engineered Escherichia coli. Metab. Eng. 2015, 30, 141–148. [Google Scholar] [CrossRef]
- Xia, J.; Yang, Y.; Liu, C.-G.; Yang, S.; Bai, F.-W. Engineering Zymomonas mobilis for Robust Cellulosic Ethanol Production. Trends Biotechnol. 2019. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Li, N.; Wang, Z.; Tang, Y.-J.; Chen, T.; Zhao, X. Inverse metabolic engineering of Bacillus subtilis for xylose utilization based on adaptive evolution and whole-genome sequencing. Appl. Microbiol. Biotechnol. 2015, 99, 885–896. [Google Scholar] [CrossRef]
- Dvořák, P.; de Lorenzo, V. Refactoring the upper sugar metabolism of Pseudomonas putida for co-utilization of cellobiose, xylose, and glucose. Metab. Eng. 2018, 48, 94–108. [Google Scholar] [CrossRef] [PubMed]
- Alper, H.; Stephanopoulos, G. Engineering for biofuels: Exploiting innate microbial capacity or importing biosynthetic potential? Nat. Rev. Microbiol. 2009, 7, 715–723. [Google Scholar] [CrossRef] [PubMed]
- Jimenez, J.I.; Minambres, B.; Garcia, J.L.; Diaz, E. Genomic analysis of the aromatic catabolic pathways from Pseudomonas putida KT2440. Environ. Microbiol. 2002, 4, 824–841. [Google Scholar] [CrossRef]
- Ballesteros, M.; Oliva, J.M.; Negro, M.J.; Manzanares, P.; Ballesteros, I. Ethanol from lignocellulosic materials by a simultaneous saccharification and fermentation process (SFS) with Kluyveromyces marxianus CECT 10875. Process Biochem. 2004, 39, 1843–1848. [Google Scholar] [CrossRef]
- Jeffries, T.W.; Grigoriev, I.V.; Grimwood, J.; Laplaza, J.M.; Aerts, A.; Salamov, A.; Schmutz, J.; Lindquist, E.; Dehal, P.; Shapiro, H.; et al. Genome sequence of the lignocellulose-bioconverting and xylose-fermenting yeast Pichia stipitis. Nat. Biotechnol. 2007, 25, 319–326. [Google Scholar] [CrossRef] [PubMed]
- Schäfer, A.; Tauch, A.; Jäger, W.; Kalinowski, J.; Thierbach, G.; Pühler, A. Small mobilizable multi-purpose cloning vectors derived from the Escherichia coli plasmids pK18 and pK19: Selection of defined deletions in the chromosome of Corynebacterium glutamicum. Gene 1994, 145, 69–73. [Google Scholar] [CrossRef]
- Kushwaha, M.; Salis, H.M. A portable expression resource for engineering cross-species genetic circuits and pathways. Nat. Commun. 2015, 6, 7832. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Copley, S.D. Inhibitory cross-talk upon introduction of a new metabolic pathway into an existing metabolic network. Proc. Natl. Acad. Sci. USA 2012, 109, E2856–E2864. [Google Scholar] [CrossRef]
- Close, D.; Cooper, S.; Wang, X.; Giannone, R.J.; Engle, N.L.; Tschaplinski, T.J.; Hedstrom, L.; Parks, J.M.; Michener, J.K. Horizontal transfer of a pathway for coumarate catabolism unexpectedly inhibits purine nucleotide biosynthesis. bioRxiv 2018, 315036. [Google Scholar] [CrossRef]
- Michener, J.K.; Camargo Neves, A.A.; Vuilleumier, S.; Bringel, F.; Marx, C.J. Effective use of a horizontally-transferred pathway for dichloromethane catabolism requires post-transfer refinement. Elife 2014, 3. [Google Scholar] [CrossRef]
- Abdel-Rahman, M.A.; Tashiro, Y.; Zendo, T.; Shibata, K.; Sonomoto, K. Isolation and characterisation of lactic acid bacterium for effective fermentation of cellobiose into optically pure homo l-(+)-lactic acid. Appl. Microbiol. Biotechnol. 2011, 89, 1039–1049. [Google Scholar] [CrossRef] [PubMed]
- Adsul, M.; Khire, J.; Bastawde, K.; Gokhale, D. Production of lactic acid from cellobiose and cellotriose by Lactobacillus delbrueckii mutant Uc-3. Appl. Environ. Microbiol. 2007, 73, 5055–5057. [Google Scholar] [CrossRef] [PubMed]
- Carr, F.J.; Chill, D.; Maida, N. The Lactic Acid Bacteria: A Literature Survey. Crit. Rev. Microbiol. 2002, 28, 281–370. [Google Scholar] [CrossRef]
- Ha, S.-J.; Galazka, J.M.; Kim, S.R.; Choi, J.-H.; Yang, X.; Seo, J.-H.; Glass, N.L.; Cate, J.H.D.; Jin, Y.-S. Engineered Saccharomyces cerevisiae capable of simultaneous cellobiose and xylose fermentation. Proc. Natl. Acad. Sci. USA 2011, 108, 504–509. [Google Scholar] [CrossRef] [PubMed]
- Shin, H.-D.; Wu, J.; Chen, R. Comparative engineering of Escherichia coli for cellobiose utilization: Hydrolysis versus phosphorolysis. Metab. Eng. 2014, 24, 9–17. [Google Scholar] [CrossRef]
- Barsett, H.; Ebringerová, A.; Harding, S.; Heinze, T. Polysaccharides I: Structure, Characterisation and Use; Springer Science & Business Media: Berlin/Heidelberg, Germany, 2005. [Google Scholar]
- Viikari, L.; Kantelinen, A.; Buchert, J.; Puls, J. Enzymatic accessibility of xylans in lignocellulosic materials. Appl. Microbiol. Biotechnol. 1994, 41, 124–129. [Google Scholar] [CrossRef]
- Hu, Z.; Sykes, R.; Davis, M.F.; Charles Brummer, E.; Ragauskas, A.J. Chemical profiles of switchgrass. Bioresour. Technol. 2010, 101, 3253–3257. [Google Scholar] [CrossRef]
- Zhang, M.; Su, R.; Qi, W.; He, Z. Enhanced Enzymatic Hydrolysis of Lignocellulose by Optimizing Enzyme Complexes. Appl. Biochem. Biotechnol. 2010. [Google Scholar] [CrossRef] [PubMed]
- Moxley, G.; Zhang, Y.-H.P. More Accurate Determination of Acid-Labile Carbohydrates in Lignocellulose by Modified Quantitative Saccharification. Energy Fuels 2007, 21, 3684–3688. [Google Scholar] [CrossRef]
- Sorek, N.; Yeats, T.H.; Szemenyei, H.; Youngs, H.; Somerville, C.R. The Implications of Lignocellulosic Biomass Chemical Composition for the Production of Advanced Biofuels. Bioscience 2014, 64, 192–201. [Google Scholar] [CrossRef]
- Chen, H. Chemical Composition and Structure of Natural Lignocellulose. In Biotechnology of Lignocellulose; Springer: Dordrecht, The Netherlands, 2014; pp. 25–71. [Google Scholar] [CrossRef]
- Pawar, P.M.-A.; Koutaniemi, S.; Tenkanen, M.; Mellerowicz, E.J. Acetylation of woody lignocellulose: Significance and regulation. Front. Plant Sci. 2013, 4, 118. [Google Scholar] [CrossRef] [PubMed]
- Zabotina, O.A. Xyloglucan and its biosynthesis. Front. Plant Sci. 2012, 3, 134. [Google Scholar] [CrossRef]
- Luterbacher, J.S.; Alonso, D.M.; Dumesic, J.A. Targeted chemical upgrading of lignocellulosic biomass to platform molecules. Green Chem. 2014, 16, 4816–4838. [Google Scholar] [CrossRef]
- Lenihan, P.; Orozco, A.; O’Neill, E.; Ahmad, M.N.M.; Rooney, D.W.; Walker, G.M. Dilute acid hydrolysis of lignocellulosic biomass. Chem. Eng. J. 2010, 156, 395–403. [Google Scholar] [CrossRef]
- Li, C.; Zhao, Z.K. Efficient Acid-Catalyzed Hydrolysis of Cellulose in Ionic Liquid. Adv. Synth. Catal. 2007, 349, 1847–1850. [Google Scholar] [CrossRef]
- Li, C.; Wang, Q.; Zhao, Z.K. Acid in ionic liquid: An efficient system for hydrolysis of lignocellulose. Green Chem. 2008, 10, 177–182. [Google Scholar] [CrossRef]
- Alonso, D.M.; Hakim, S.H.; Zhou, S.; Won, W.; Hosseinaei, O.; Tao, J.; Garcia-Negron, V.; Motagamwala, A.H.; Mellmer, M.A.; Huang, K.; et al. Increasing the revenue from lignocellulosic biomass: Maximizing feedstock utilization. Sci. Adv. 2017, 3, e1603301. [Google Scholar] [CrossRef] [PubMed]
- Schutyser, W.; Renders, T.; Van den Bosch, S.; Koelewijn, S.F.; Beckham, G.T.; Sels, B.F. Chemicals from lignin: An interplay of lignocellulose fractionation, depolymerisation, and upgrading. Chem. Soc. Rev. 2018, 47, 852–908. [Google Scholar] [CrossRef] [PubMed]
- Ragauskas, A.J.; Beckham, G.T.; Biddy, M.J.; Chandra, R.; Chen, F.; Davis, M.F.; Davison, B.H.; Dixon, R.A.; Gilna, P.; Keller, M.; et al. Lignin Valorization: Improving Lignin Processing in the Biorefinery. Science 2014, 344, 709. [Google Scholar] [CrossRef] [PubMed]
- Abdelaziz, O.Y.; Brink, D.P.; Prothmann, J.; Ravi, K.; Sun, M.Z.; Garcia-Hidalgo, J.; Sandahl, M.; Hulteberg, C.P.; Turner, C.; Liden, G.; et al. Biological valorization of low molecular weight lignin. Biotechnol. Adv. 2016, 34, 1318–1346. [Google Scholar] [CrossRef]
- Higuchi, T. Lignin biochemistry—Biosynthesis and biodegradation. Wood Sci. Technol. 1990, 24, 23–63. [Google Scholar] [CrossRef]
- Ralph, J. Hydroxycinnamates in lignification. Phytochem. Rev. 2010, 9, 65–83. [Google Scholar] [CrossRef]
- Lan, W.; Rencoret, J.; Lu, F.C.; Karlen, S.D.; Smith, B.G.; Harris, P.J.; del Rio, J.C.; Ralph, J. Tricin-lignins: Occurrence and quantitation of tricin in relation to phylogeny. Plant J. 2016, 88, 1046–1057. [Google Scholar] [CrossRef]
- El Hage, R.; Brosse, N.; Chrusciel, L.; Sanchez, C.; Sannigrahi, P.; Ragauskas, A. Characterization of milled wood lignin and ethanol organosolv lignin from miscanthus. Polym. Degrad. Stab. 2009, 94, 1632–1638. [Google Scholar] [CrossRef]
- Villaverde, J.J.; Li, J.; Ek, M.; Ligero, P.; de Vega, A. Native Lignin Structure of Miscanthus x giganteus and Its Changes during Acetic and Formic Acid Fractionation. J. Agric. Food Chem. 2009, 57, 6262–6270. [Google Scholar] [CrossRef] [PubMed]
- Auxenfans, T.; Crônier, D.; Chabbert, B.; Paës, G. Understanding the structural and chemical changes of plant biomass following steam explosion pretreatment. Biotechnol. Biofuels 2017, 10, 36. [Google Scholar] [CrossRef] [PubMed]
- David, K.; Ragauskas, A.J. Switchgrass as an energy crop for biofuel production: A review of its ligno-cellulosic chemical properties. Energy Environ. Sci. 2010, 3, 1182. [Google Scholar] [CrossRef]
- Yan, J.; Hu, Z.; Pu, Y.; Charles Brummer, E.; Ragauskas, A.J. Chemical compositions of four switchgrass populations. Biomass Bioenergy 2010, 34, 48–53. [Google Scholar] [CrossRef]
- Samuel, R.; Foston, M.; Jiang, N.; Allison, L.; Ragauskas, A.J. Structural changes in switchgrass lignin and hemicelluloses during pretreatments by NMR analysis. Polym. Degrad. Stab. 2011, 96, 2002–2009. [Google Scholar] [CrossRef]
- Dutta, T.; Papa, G.; Wang, E.; Sun, J.; Isern, N.G.; Cort, J.R.; Simmons, B.A.; Singh, S. Characterization of Lignin Streams during Bionic Liquid-Based Pretreatment from Grass, Hardwood, and Softwood. ACS Sustain. Chem. Eng. 2018, 6, 3079–3090. [Google Scholar] [CrossRef]
- Zeng, J.; Helms, G.L.; Gao, X.; Chen, S. Quantification of Wheat Straw Lignin Structure by Comprehensive NMR Analysis. J. Agric. Food Chem. 2013, 61, 10848–10857. [Google Scholar] [CrossRef]
- Min, D.; Wang, S.; Chang, H.; Jameel, H.; Lucia, L. Molecular Changes in Corn Stover Lignin Resulting from Pretreatment Chemistry. BioResources 2017, 12, 6262–6275. [Google Scholar] [CrossRef]
- Stewart, J.J.; Akiyama, T.; Chapple, C.; Ralph, J.; Mansfield, S.D. The Effects on Lignin Structure of Overexpression of Ferulate 5-Hydroxylase in Hybrid Poplar. Plant Physiol. 2009, 150, 621–635. [Google Scholar] [CrossRef]
- Sannigrahi, P.; Ragauskas, A.J.; Tuskan, G.A. Poplar as a feedstock for biofuels: A review of compositional characteristics. Biofuels Bioprod. Biorefining 2010, 4, 209–226. [Google Scholar] [CrossRef]
- Wagner, A.; Tobimatsu, Y.; Phillips, L.; Flint, H.; Geddes, B.; Lu, F.; Ralph, J. Syringyl lignin production in conifers: Proof of concept in a Pine tracheary element system. Proc. Natl. Acad. Sci. USA 2015, 112, 6218–6223. [Google Scholar] [CrossRef]
- Capanema, E.A.; Balakshin, M.Y.; Kadla, J.F. A comprehensive approach for quantitative lignin characterization by NMR spectroscopy. J. Agric. Food Chem. 2004, 52, 1850–1860. [Google Scholar] [CrossRef]
- Constant, S.; Wienk, H.L.J.; Frissen, A.E.; de Peinder, P.; Boelens, R.; van Es, D.S.; Grisel, R.J.H.; Weckhuysen, B.M.; Huijgen, W.J.J.; Gosselink, R.J.A.; et al. New insights into the structure and composition of technical lignins: A comparative characterisation study. Green Chem. 2016, 18, 2651–2665. [Google Scholar] [CrossRef]
- Wadenbäck, J.; Clapham, D.; Gellerstedt, G.; von Arnold, S. Variation in content and composition of lignin in young wood of Norway spruce. Holzforschung 2004, 58, 107–115. [Google Scholar] [CrossRef]
- Ralph, J.; Lundquist, K.; Brunow, G.; Lu, F.; Kim, H.; Schatz, P.F.; Marita, J.M.; Hatfield, R.D.; Ralph, S.A.; Christensen, J.H.; et al. Lignins: Natural polymers from oxidative coupling of 4-hydroxyphenyl- propanoids. Phytochem. Rev. 2004, 3, 29–60. [Google Scholar] [CrossRef]
- Li, M.; Pu, Y.Q.; Ragauskas, A.J. Current Understanding of the Correlation of Lignin Structure with Biomass Recalcitrance. Front. Chem. 2016, 4. [Google Scholar] [CrossRef] [PubMed]
- Bugg, T.D.H.; Ahmad, M.; Hardiman, E.M.; Rahmanpour, R. Pathways for degradation of lignin in bacteria and fungi. Nat. Prod. Rep. 2011, 28, 1883–1896. [Google Scholar] [CrossRef]
- Gupta, A.; Verma, J.P. Sustainable bio-ethanol production from agro-residues: A review. Renew. Sustain. Energy Rev. 2015, 41, 550–567. [Google Scholar] [CrossRef]
- Van den Bosch, S.; Koelewijn, S.F.; Renders, T.; Van den Bossche, G.; Vangeel, T.; Schutyser, W.; Sels, B.F. Catalytic Strategies Towards Lignin-Derived Chemicals. Top. Curr. Chem. 2018, 376. [Google Scholar] [CrossRef]
- Lancefield, C.S.; Rashid, G.M.M.; Bouxin, F.; Wasak, A.; Tu, W.C.; Hallett, J.; Zein, S.; Rodriguez, J.; Jackson, S.D.; Westwood, N.J.; et al. Investigation of the Chemocatalytic and Biocatalytic Valorization of a Range of Different Lignin Preparations: The Importance of beta-O-4 Content. ACS Sustain. Chem. Eng. 2016, 4, 6921–6930. [Google Scholar] [CrossRef]
- Bouxin, F.P.; McVeigh, A.; Tran, F.; Westwood, N.J.; Jarvis, M.C.; Jackson, S.D. Catalytic depolymerisation of isolated lignins to fine chemicals using a Pt/alumina catalyst: Part 1-impact of the lignin structure. Green Chem. 2015, 17, 1235–1242. [Google Scholar] [CrossRef]
- Studer, M.H.; DeMartini, J.D.; Davis, M.F.; Sykes, R.W.; Davison, B.; Keller, M.; Tuskan, G.A.; Wyman, C.E. Lignin content in natural Populus variants affects sugar release. Proc. Natl. Acad. Sci. USA 2011, 108, 6300–6305. [Google Scholar] [CrossRef]
- Rinaldi, R.; Jastrzebski, R.; Clough, M.T.; Ralph, J.; Kennema, M.; Bruijnincx, P.C.A.; Weckhuysen, B.M. Paving the Way for Lignin Valorisation: Recent Advances in Bioengineering, Biorefining and Catalysis. Angew. Chem. Int. Ed. 2016, 55, 8164–8215. [Google Scholar] [CrossRef] [PubMed]
- Barta, K.; Warner, G.R.; Beach, E.S.; Anastas, P.T. Depolymerization of organosolv lignin to aromatic compounds over Cu-doped porous metal oxides. Green Chem. 2014, 16, 191–196. [Google Scholar] [CrossRef]
- Katahira, R.; Mittal, A.; McKinney, K.; Chen, X.W.; Tucker, M.P.; Johnson, D.K.; Beckham, G.T. Base-Catalyzed Depolymerization of Biorefinery Lignins. ACS Sustain. Chem. Eng. 2016, 4, 1474–1486. [Google Scholar] [CrossRef]
- Luo, H.; Klein, I.M.; Jiang, Y.; Zhu, H.Y.; Liu, B.Y.; Kenttamaa, H.I.; Abu-Omar, M.M. Total Utilization of Miscanthus Biomass, Lignin and Carbohydrates, Using Earth Abundant Nickel Catalyst. ACS Sustain. Chem. Eng. 2016, 4, 2316–2322. [Google Scholar] [CrossRef]
- Renders, T.; Schutyser, W.; Van den Bosch, S.; Koelewijn, S.F.; Vangeel, T.; Courtin, C.M.; Sels, B.F. Influence of Acidic (H3PO4) and Alkaline (NaOH) Additives on the Catalytic Reductive Fractionation of Lignocellulose. ACS Catal. 2016, 6, 2055–2066. [Google Scholar] [CrossRef]
- Renders, T.; Van den Bosch, S.; Koelewijn, S.F.; Schutyser, W.; Sels, B.F. Lignin-first biomass fractionation: The advent of active stabilisation strategies. Energy Environ. Sci. 2017, 10, 1551–1557. [Google Scholar] [CrossRef]
- Papagianni, M. Recent advances in engineering the central carbon metabolism of industrially important bacteria. Microb. Cell Fact. 2012, 11, 50. [Google Scholar] [CrossRef] [PubMed]
- Linger, J.G.; Vardon, D.R.; Guarnieri, M.T.; Karp, E.M.; Hunsinger, G.B.; Franden, M.A.; Johnson, C.W.; Chupka, G.; Strathmann, T.J.; Pienkos, P.T.; et al. Lignin valorization through integrated biological funneling and chemical catalysis. Proc. Natl. Acad. Sci. USA 2014, 111, 12013–12018. [Google Scholar] [CrossRef] [PubMed]
- Bar-Even, A.; Flamholz, A.; Noor, E.; Milo, R. Rethinking glycolysis: On the biochemical logic of metabolic pathways. Nat. Chem. Biol. 2012, 8, 509–517. [Google Scholar] [CrossRef] [PubMed]
- Gottschalk, G. Bacterial Metabolism; Springer: New York, NY, USA, 2012. [Google Scholar]
- Conway, T. The Entner-Doudoroff pathway: History, physiology and molecular biology. FEMS Microbiol. Lett. 1992, 103, 1–28. [Google Scholar] [CrossRef] [PubMed]
- Kurn, N.; Contreras, I.; Shapiro, L. Galactose Catabolism in Caulobacter crescentus. J. Bacteriol. 1978, 135, 517–520. [Google Scholar] [PubMed]
- Bender, R.; Andreesen, J.R.; Gottschalk, G. 2-keto-3-deoxygluconate, an intermediate in the fermentation of gluconate by clostridia. J. Bacteriol. 1971, 107, 570–573. [Google Scholar]
- Ahmed, H.; Ettema, T.J.G.; Tjaden, B.; Geerling, A.C.M.; van der Oost, J.; Siebers, B. The semi-phosphorylative Entner-Doudoroff pathway in hyperthermophilic archaea: A re-evaluation. Biochem. J. 2005, 390, 529–540. [Google Scholar] [CrossRef]
- Sprenger, G.A. Genetics of pentose-phosphate pathway enzymes ofEscherichia coli K-12. Arch. Microbiol. 1995, 164, 324–330. [Google Scholar] [CrossRef] [PubMed]
- Sudarsan, S.; Dethlefsen, S.; Blank, L.M.; Siemann-Herzberg, M.; Schmid, A. The functional structure of central carbon metabolism in Pseudomonas putida KT2440. Appl. Environ. Microbiol. 2014, 80, 5292–5303. [Google Scholar] [CrossRef] [PubMed]
- Valdehuesa, K.N.G.; Ramos, K.R.M.; Nisola, G.M.; Bañares, A.B.; Cabulong, R.B.; Lee, W.-K.; Liu, H.; Chung, W.-J. Everyone loves an underdog: Metabolic engineering of the xylose oxidative pathway in recombinant microorganisms. Appl. Microbiol. Biotechnol. 2018, 102, 7703–7716. [Google Scholar] [CrossRef]
- Watanabe, S.; Kodaki, T.; Kodak, T.; Makino, K. Cloning, expression, and characterization of bacterial L-arabinose 1-dehydrogenase involved in an alternative pathway of L-arabinose metabolism. J. Biol. Chem. 2006, 281, 2612–2623. [Google Scholar] [CrossRef] [PubMed]
- Deanda, K.; Zhang, M.; Eddy, C.; Picataggio, S. Development of an arabinose-fermenting Zymomonas mobilis strain by metabolic pathway engineering. Appl. Environ. Microbiol. 1996, 62, 4465–4470. [Google Scholar]
- Kawaguchi, H.; Sasaki, M.; Vertès, A.A.; Inui, M.; Yukawa, H. Engineering of an l-arabinose metabolic pathway in Corynebacterium glutamicum. Appl. Microbiol. Biotechnol. 2008, 77, 1053–1062. [Google Scholar] [CrossRef] [PubMed]
- Xiong, X.; Wang, X.; Chen, S. Engineering of an L-arabinose metabolic pathway in Rhodococcus jostii RHA1 for biofuel production. J. Ind. Microbiol. Biotechnol. 2016, 43, 1017–1025. [Google Scholar] [CrossRef] [PubMed]
- Seiboth, B.; Metz, B. Fungal arabinan and l-arabinose metabolism. Appl. Microbiol. Biotechnol. 2011, 89, 1665–1673. [Google Scholar] [CrossRef]
- McClintock, M.K.; Wang, J.; Zhang, K. Application of Nonphosphorylative Metabolism as an Alternative for Utilization of Lignocellulosic Biomass. Front. Microbiol. 2017, 8, 2310. [Google Scholar] [CrossRef] [PubMed]
- Holden, H.M.; Rayment, I.; Thoden, J.B. Structure and function of enzymes of the Leloir pathway for galactose metabolism. J. Biol. Chem. 2003, 278, 43885–43888. [Google Scholar] [CrossRef]
- Berka, T.R.; Lessie, T.G. Enzymes related to galactose utilization inPseudomonas cepacia. Curr. Microbiol. 1984, 11, 43–48. [Google Scholar] [CrossRef]
- Neves, A.R.; Pool, W.A.; Solopova, A.; Kok, J.; Santos, H.; Kuipers, O.P. Towards enhanced galactose utilization by Lactococcus lactis. Appl. Environ. Microbiol. 2010, 76, 7048–7060. [Google Scholar] [CrossRef]
- Voragen, A.G.J.; Coenen, G.-J.; Verhoef, R.P.; Schols, H.A. Pectin, a versatile polysaccharide present in plant cell walls. Struct. Chem. 2009, 20, 263–275. [Google Scholar] [CrossRef]
- Protzko, R.J.; Latimer, L.N.; Martinho, Z.; de Reus, E.; Seibert, T.; Benz, J.P.; Dueber, J.E. Engineering Saccharomyces cerevisiae for co-utilization of d-galacturonic acid and d-glucose from citrus peel waste. Nat. Commun. 2018, 9, 5059. [Google Scholar] [CrossRef]
- Jojima, T.; Omumasaba, C.A.; Inui, M.; Yukawa, H. Sugar transporters in efficient utilization of mixed sugar substrates: Current knowledge and outlook. Appl. Microbiol. Biotechnol. 2010, 85, 471–480. [Google Scholar] [CrossRef]
- Görke, B.; Stülke, J. Carbon catabolite repression in bacteria: Many ways to make the most out of nutrients. Nat. Rev. Microbiol. 2008, 6, 613–624. [Google Scholar] [CrossRef]
- Stülke, J.; Hillen, W. Carbon catabolite repression in bacteria. Curr. Opin. Microbiol. 1999, 2, 195–201. [Google Scholar] [CrossRef]
- Masai, E.; Harada, K.; Peng, X.; Kitayama, H.; Katayama, Y.; Fukuda, M.; Harada, Y.; Peng, X. Cloning and characterization of the ferulic acid catabolic genes of Sphingomonas paucimobilis SYK-6. Appl. Environ. Microbiol. 2002, 68, 4416–4424. [Google Scholar] [CrossRef] [PubMed]
- Overhage, J.; Priefert, H.; Steinbuchel, A. Biochemical and genetic analyses of ferulic acid catabolism in Pseudomonas sp strain HR199. Appl. Environ. Microbiol. 1999, 65, 4837–4847. [Google Scholar]
- Kamimura, N.; Goto, T.; Takahashi, K.; Kasai, D.; Otsuka, Y.; Nakamura, M.; Katayama, Y.; Fukuda, M.; Masai, E. A bacterial aromatic aldehyde dehydrogenase critical for the efficient catabolism of syringaldehyde. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef]
- Priefert, H.; Rabenhorst, J.; Steinbuchel, A. Biotechnological production of vanillin. Appl. Microbiol. Biotechnol. 2001, 56, 296–314. [Google Scholar] [CrossRef]
- Ravi, K.; Garcia-Hidalgo, J.; Gorwa-Grauslund, M.F.; Liden, G. Conversion of lignin model compounds by Pseudomonas putida KT2440 and isolates from compost. Appl. Microbiol. Biotechnol. 2017, 101, 5059–5070. [Google Scholar] [CrossRef] [PubMed]
- Seibold, B.; Matthes, M.; Eppink, M.H.M.; Lingens, F.; Berkel, W.J.H.; Muller, R.; vanBerkel, W.J.H.; Muller, R. 4-hydroxybenzoate hydroxylase from Pseudomonas sp CBS3—Purification, characterization, gene cloning, sequence analysis and assignment of structural features determining the coenzyme specificity. Eur. J. Biochem. 1996, 239, 469–478. [Google Scholar] [CrossRef]
- Brunel, F.; Davison, J. Cloning and sequencing of pseudomonas genes encoding vanillate demethylase. J. Bacteriol. 1988, 170, 4924–4930. [Google Scholar] [CrossRef] [PubMed]
- Morawski, B.; Segura, A.; Ornston, L.N. Substrate range and genetic analysis of Acinetobacter vanillate demethylase. J. Bacteriol. 2000, 182, 1383–1389. [Google Scholar] [CrossRef] [PubMed]
- Mohan, K.; Phale, P.S. Carbon Source-Dependent Inducible Metabolism of Veratryl Alcohol and Ferulic Acid in Pseudomonas putida CSV86. Appl. Environ. Microbiol. 2017, 83. [Google Scholar] [CrossRef]
- Harada, A.; Kamimura, N.; Takeuchi, K.; Yu, H.Y.; Masai, E.; Senda, T. The crystal structure of a new O-demethylase from Sphingobium sp strain SYK-6. FEBS J. 2017, 284, 1855–1867. [Google Scholar] [CrossRef] [PubMed]
- Abe, T.; Masai, E.; Miyauchi, K.; Katayama, Y.; Fukuda, M. A tetrahydrofolate-dependent O-demethylase, LigM, is crucial for catabolism of vanillate and syringate in Sphingomonas paucimobilis SYK-6. J. Bacteriol. 2005, 187, 2030–2037. [Google Scholar] [CrossRef] [PubMed]
- Cecil, J.H.; Garcia, D.C.; Giannone, R.J.; Michener, J.K. Rapid, Parallel Identification of Catabolism Pathways of Lignin-Derived Aromatic Compounds in Novosphingobium aromaticivorans. Appl. Environ. Microbiol. 2018, 84. [Google Scholar] [CrossRef] [PubMed]
- Nishimura, M.; Nishimura, Y.; Abe, C.; Kohhata, M. Expression and Substrate Range of Streptomyces Vanillate Demethylase. Biol. Pharm. Bull. 2014, 37, 1564–1568. [Google Scholar] [CrossRef]
- Kamimura, N.; Takahashi, K.; Mori, K.; Araki, T.; Fujita, M.; Higuchi, Y.; Masai, E. Bacterial catabolism of lignin-derived aromatics: New findings in a recent decade: Update on bacterial lignin catabolism. Environ. Microbiol. Rep. 2017, 9, 679–705. [Google Scholar] [CrossRef]
- Masai, E.; Sasaki, M.; Minakawa, Y.; Abe, T.; Sonoki, T.; Miyauchi, K.; Katayama, Y.; Fukuda, M. A novel tetrahydrofolate-dependent O-demethylase gene is essential for growth of Sphingomonas paucimobilis SYK-6 with syringate. J. Bacteriol. 2004, 186, 2757–2765. [Google Scholar] [CrossRef] [PubMed]
- Mallinson, S.J.B.; Machovina, M.M.; Silveira, R.L.; Garcia-Borras, M.; Gallup, N.; Johnson, C.W.; Allen, M.D.; Skaf, M.S.; Crowley, M.F.; Neidle, E.L.; et al. A promiscuous cytochrome P450 aromatic O-demethylase for lignin bioconversion. Nat. Commun. 2018, 9. [Google Scholar] [CrossRef]
- Tumen-Velasquez, M.; Johnson, C.W.; Ahmed, A.; Dominick, G.; Fulk, E.M.; Khanna, P.; Lee, S.A.; Schmidt, A.L.; Linger, J.G.; Eiteman, M.A.; et al. Accelerating pathway evolution by increasing the gene dosage of chromosomal segments. Proc. Natl. Acad. Sci. USA 2018, 115, 7105–7110. [Google Scholar] [CrossRef]
- Harwood, C.S.; Parales, R.E. The β-ketoadipate pathway and the biology of self-identity. Annu. Rev. Microbiol. 1996, 50, 553–590. [Google Scholar] [CrossRef] [PubMed]
- Kasai, D.; Masai, E.; Miyauchi, K.; Katayama, Y.; Fukuda, M. Characterization of the 3-O-Methylgallate Dioxygenase Gene and Evidence of Multiple 3-O-Methylgallate Catabolic Pathways in Sphingomonas paucimobilis SYK-6. J. Bacteriol. 2004, 186, 4951–4959. [Google Scholar] [CrossRef] [PubMed]
- Beckham, G.T.; Johnson, C.W.; Karp, E.M.; Salvachua, D.; Vardon, D.R.; Salvachúa, D.; Vardon, D.R. Opportunities and challenges in biological lignin valorization. Curr. Opin. Biotechnol. 2016, 42, 40–53. [Google Scholar] [CrossRef]
- Yoshida, T.; Inami, Y.; Matsui, T.; Nagasawa, T. Regioselective carboxylation of catechol by 3,4-dihydroxybenzoate decarboxylase of Enterobacter cloacae P. Biotechnol. Lett. 2010, 32, 701–705. [Google Scholar] [CrossRef] [PubMed]
- Suastegui, M.; Matthiesen, J.E.; Carraher, J.M.; Hernandez, N.; Quiroz, N.R.; Okerlund, A.; Cochran, E.W.; Shao, Z.; Tessonnier, J.-P. Combining Metabolic Engineering and Electrocatalysis: Application to the Production of Polyamides from Sugar. Angew. Chemie Int. Ed. 2016, 55, 2368–2373. [Google Scholar] [CrossRef] [PubMed]
- Walmsley, A.R.; Barrett, M.P.; Bringaud, F.; Gould, G.W. Sugar transporters from bacteria, parasites and mammals: Structure–activity relationships. Trends Biochem. Sci. 1998, 23, 476–481. [Google Scholar] [CrossRef]
- Bernaudat, F.; Frelet-Barrand, A.; Pochon, N.; Dementin, S.; Hivin, P.; Boutigny, S.; Rioux, J.-B.; Salvi, D.; Seigneurin-Berny, D.; Richaud, P.; et al. Heterologous Expression of Membrane Proteins: Choosing the Appropriate Host. PLoS ONE 2011, 6, e29191. [Google Scholar] [CrossRef]
- Claassens, N.J.; Siliakus, M.F.; Spaans, S.K.; Creutzburg, S.C.A.; Nijsse, B.; Schaap, P.J.; Quax, T.E.F.; van der Oost, J. Improving heterologous membrane protein production in Escherichia coli by combining transcriptional tuning and codon usage algorithms. PLoS ONE 2017, 12, e0184355. [Google Scholar] [CrossRef]
- Van Dijken, J.P.; Scheffers, W.A. Redox balances in the metabolism of sugars by yeasts (NAD(H); NADP(H); glucose metabolism; xylose fermentation; ethanol; Crabtree effect; Custers effect). FEMS Microbiol. Rev. 1986, 1, 199–224. [Google Scholar]
- Chavarría, M.; Nikel, P.I.; Pérez-Pantoja, D.; de Lorenzo, V. The Entner-Doudoroff pathway empowers Pseudomonas putida KT2440 with a high tolerance to oxidative stress. Environ. Microbiol. 2013, 15, 1772–1785. [Google Scholar] [CrossRef]
- Chen, X.; Li, S.; Liu, L. Engineering redox balance through cofactor systems. Trends Biotechnol. 2014, 32, 337–343. [Google Scholar] [CrossRef]
- Matsushika, A.; Inoue, H.; Kodaki, T.; Sawayama, S. Ethanol production from xylose in engineered Saccharomyces cerevisiae strains: Current state and perspectives. Appl. Microbiol. Biotechnol. 2009, 84, 37–53. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.-C.; Liu, J.-J.; Kong, I.I.; Kwak, S.; Jin, Y.-S. Combining C6 and C5 sugar metabolism for enhancing microbial bioconversion. Curr. Opin. Chem. Biol. 2015, 29, 49–57. [Google Scholar] [CrossRef] [PubMed]
- Meijnen, J.-P.; de Winde, J.H.; Ruijssenaars, H.J. Engineering Pseudomonas putida S12 for efficient utilization of D-xylose and L-arabinose. Appl. Environ. Microbiol. 2008, 74, 5031–5037. [Google Scholar] [CrossRef] [PubMed]
- Johnson, C.W.; Salvachúa, D.; Khanna, P.; Smith, H.; Peterson, D.J.; Beckham, G.T. Enhancing muconic acid production from glucose and lignin-derived aromatic compounds via increased protocatechuate decarboxylase activity. Metab. Eng. Commun. 2016, 3, 111–119. [Google Scholar] [CrossRef] [PubMed]
- Johnson, C.W.; Abraham, P.E.; Linger, J.G.; Khanna, P.; Hettich, R.L.; Beckham, G.T. Eliminating a global regulator of carbon catabolite repression enhances the conversion of aromatic lignin monomers to muconate in Pseudomonas putida KT2440. Metab. Eng. Commun. 2017, 5, 19–25. [Google Scholar] [CrossRef] [PubMed]
- Mitraki, A.; King, J. Protein Folding Intermediates and Inclusion Body Formation. Nat. Biotechnol. 1989, 7, 690–697. [Google Scholar] [CrossRef]
- Michener, J.K.; Nielsen, J.; Smolke, C.D. Identification and treatment of heme depletion attributed to overexpression of a lineage of evolved P450 monooxygenases. Proc. Natl. Acad. Sci. USA 2012, 109, 19504–19509. [Google Scholar] [CrossRef]
- Keasling, J.D. Manufacturing Molecules Through Metabolic Engineering. Science 2010, 330, 1355–1358. [Google Scholar] [CrossRef] [PubMed]
- Stephanopoulos, G. Metabolic Fluxes and Metabolic Engineering. Metab. Eng. 1999, 1, 1–11. [Google Scholar] [CrossRef]
- Brockman, I.M.; Prather, K.L.J. Dynamic knockdown of E. coli central metabolism for redirecting fluxes of primary metabolites. Metab. Eng. 2015, 28, 104–113. [Google Scholar] [CrossRef]
- George, K.W.; Thompson, M.G.; Kim, J.; Baidoo, E.E.K.; Wang, G.; Benites, V.T.; Petzold, C.J.; Chan, L.J.G.; Yilmaz, S.; Turhanen, P.; et al. Integrated analysis of isopentenyl pyrophosphate (IPP) toxicity in isoprenoid-producing Escherichia coli. Metab. Eng. 2018, 47, 60–72. [Google Scholar] [CrossRef] [PubMed]
- Bentley, F.K.; Zurbriggen, A.; Melis, A. Heterologous Expression of the Mevalonic Acid Pathway in Cyanobacteria Enhances Endogenous Carbon Partitioning to Isoprene. Mol. Plant 2014, 7, 71–86. [Google Scholar] [CrossRef] [PubMed]
- Pitera, D.J.; Paddon, C.J.; Newman, J.D.; Keasling, J.D. Balancing a heterologous mevalonate pathway for improved isoprenoid production in Escherichia coli. Metab. Eng. 2007, 9, 193–207. [Google Scholar] [CrossRef] [PubMed]
- Akkaya, Ö.; Pérez-Pantoja, D.R.; Calles, B.; Nikel, P.I.; de Lorenzo, V. The Metabolic Redox Regime of Pseudomonas putida Tunes Its Evolvability toward Novel Xenobiotic Substrates. MBio 2018, 9, e01512-18. [Google Scholar] [CrossRef]
- Notebaart, R.A.; Kintses, B.; Feist, A.M.; Papp, B. Underground metabolism: Network-level perspective and biotechnological potential. Curr. Opin. Biotechnol. 2018, 49, 108–114. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.-H.; Block, D.E.; Mills, D.A. Simultaneous consumption of pentose and hexose sugars: An optimal microbial phenotype for efficient fermentation of lignocellulosic biomass. Appl. Microbiol. Biotechnol. 2010, 88, 1077–1085. [Google Scholar] [CrossRef]
- Barry, K.P.; Taylor, E.A. Characterizing the promiscuity of LigAB, a lignin catabolite degrading extradiol dioxygenase from Sphingomonas paucimobilis SYK-6. Biochemistry 2013, 52, 6724–6736. [Google Scholar] [CrossRef]
- Yun, E.J.; Oh, E.J.; Liu, J.-J.; Yu, S.; Kim, D.H.; Kwak, S.; Kim, K.H.; Jin, Y.-S. Promiscuous activities of heterologous enzymes lead to unintended metabolic rerouting in Saccharomyces cerevisiae engineered to assimilate various sugars from renewable biomass. Biotechnol. Biofuels 2018, 11, 140. [Google Scholar] [CrossRef]
- Qian, Y.; Otsuka, Y.; Sonoki, T.; Mukhopadhyay, B.; Nakamura, M.; Jellison, J.; Goodell, B. Engineered Microbial Production of 2-Pyrone-4,6-Dicarboxylic Acid from Lignin Residues for Use as an Industrial Platform Chemical. Bioresources 2016, 11, 6097–6109. [Google Scholar] [CrossRef]
- Johnson, C.W.; Beckham, G.T. Aromatic catabolic pathway selection for optimal production of pyruvate and lactate from lignin. Metab. Eng. 2015, 28, 240–247. [Google Scholar] [CrossRef]
- Michener, J.K.; Vuilleumier, S.; Bringel, F.; Marx, C.J. Phylogeny Poorly Predicts the Utility of a Challenging Horizontally Transferred Gene in Methylobacterium Strains. J. Bacteriol. 2014, 196, 2101–2107. [Google Scholar] [CrossRef]
- Porse, A.; Schou, T.S.; Munck, C.; Ellabaan, M.M.H.; Sommer, M.O.A. Biochemical mechanisms determine the functional compatibility of heterologous genes. Nat. Commun. 2018, 9, 522. [Google Scholar] [CrossRef]
- Amin, S.A.; Endalur Gopinarayanan, V.; Nair, N.U.; Hassoun, S. Establishing synthesis pathway-host compatibility via enzyme solubility. Biotechnol. Bioeng. 2019, 26959. [Google Scholar] [CrossRef] [PubMed]
- Agostini, F.; Cirillo, D.; Livi, C.M.; Delli Ponti, R.; Tartaglia, G.G. ccSOL omics: A webserver for solubility prediction of endogenous and heterologous expression in Escherichia coli. Bioinformatics 2014, 30, 2975–2977. [Google Scholar] [CrossRef] [PubMed]
- Knight, T. Draft Standard for Biobrick Biological Parts. 2007. Available online: http://dspace.mit.edu/handle/1721.1/45138 (accessed on 22 April 2019).
- Smolke, C.D. Building outside of the box: iGEM and the BioBricks Foundation. Nat. Biotechnol. 2009, 27, 1099–1102. [Google Scholar] [CrossRef]
- Canton, B.; Labno, A.; Endy, D. Refinement and standardization of synthetic biological parts and devices. Nat. Biotechnol. 2008, 26, 787–793. [Google Scholar] [CrossRef]
- Exley, K.; Reynolds, C.R.; Suckling, L.; Chee, S.M.; Tsipa, A.; Freemont, P.S.; McClymont, D.; Kitney, R.I. Utilising datasheets for the informed automated design and build of a synthetic metabolic pathway. J. Biol. Eng. 2019, 13, 8. [Google Scholar] [CrossRef] [PubMed]
- Lee, T.; Krupa, R.A.; Zhang, F.; Hajimorad, M.; Holtz, W.J.; Prasad, N.; Lee, S.; Keasling, J.D. BglBrick vectors and datasheets: A synthetic biology platform for gene expression. J. Biol. Eng. 2011, 5, 12. [Google Scholar] [CrossRef] [PubMed]
- Radeck, J.; Kraft, K.; Bartels, J.; Cikovic, T.; Dürr, F.; Emenegger, J.; Kelterborn, S.; Sauer, C.; Fritz, G.; Gebhard, S.; et al. The Bacillus BioBrick Box: Generation and evaluation of essential genetic building blocks for standardized work with Bacillus subtilis. J. Biol. Eng. 2013, 7, 29. [Google Scholar] [CrossRef] [PubMed]
- Xu, P.; Vansiri, A.; Bhan, N.; Koffas, M.A.G. ePathBrick: A Synthetic Biology Platform for Engineering Metabolic Pathways in E. coli. ACS Synth. Biol. 2012, 1, 256–266. [Google Scholar] [CrossRef] [PubMed]
- Holkenbrink, C.; Dam, M.I.; Kildegaard, K.R.; Beder, J.; Dahlin, J.; Doménech Belda, D.; Borodina, I. EasyCloneYALI: CRISPR/Cas9-Based Synthetic Toolbox for Engineering of the Yeast Yarrowia lipolytica. Biotechnol. J. 2018, 13, 1700543. [Google Scholar] [CrossRef] [PubMed]
- Ellinger, J.; Schmidt-Dannert, C. Construction of a BioBrickTM compatible vector system for Rhodococcus. Plasmid 2017, 90, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Pascuala, A.; de Lorenzo, V.; Nikel, P.I. Refactoring the Embden–Meyerhof–Parnas Pathway as a Whole of Portable GlucoBricks for Implantation of Glycolytic Modules in Gram-Negative Bacteria. ACS Synth. Biol. 2017, 6, 793–805. [Google Scholar] [CrossRef]
- Silva-Rocha, R.; Martínez-García, E.; Calles, B.; Chavarría, M.; Arce-Rodríguez, A.; de las Heras, A.; Páez-Espino, A.D.; Durante-Rodríguez, G.; Kim, J.; Nikel, P.I.; et al. The Standard European Vector Architecture (SEVA): A coherent platform for the analysis and deployment of complex prokaryotic phenotypes. Nucleic Acids Res. 2013, 41, D666–D675. [Google Scholar] [CrossRef]
- Latimer, L.N.; Lee, M.E.; Medina-Cleghorn, D.; Kohnz, R.A.; Nomura, D.K.; Dueber, J.E. Employing a combinatorial expression approach to characterize xylose utilization in Saccharomyces cerevisiae. Metab. Eng. 2014, 25, 20–29. [Google Scholar] [CrossRef]
- Sonoki, T.; Morooka, M.; Sakamoto, K.; Otsuka, Y.; Nakamura, M.; Jellison, J.; Goodell, B. Enhancement of protocatechuate decarboxylase activity for the effective production of muconate from lignin-related aromatic compounds. J. Biotechnol. 2014, 192, 71–77. [Google Scholar] [CrossRef] [PubMed]
- Payne, K.A.P.; White, M.D.; Fisher, K.; Khara, B.; Bailey, S.S.; Parker, D.; Rattray, N.J.W.; Trivedi, D.K.; Goodacre, R.; Beveridge, R.; et al. New cofactor supports α,β-unsaturated acid decarboxylation via 1,3-dipolar cycloaddition. Nature 2015, 522, 497–501. [Google Scholar] [CrossRef] [PubMed]
- Kohlstedt, M.; Starck, S.; Barton, N.; Stolzenberger, J.; Selzer, M.; Mehlmann, K.; Schneider, R.; Pleissner, D.; Rinkel, J.; Dickschat, J.S.; et al. From lignin to nylon: Cascaded chemical and biochemical conversion using metabolically engineered Pseudomonas putida. Metab. Eng. 2018, 47, 279–293. [Google Scholar] [CrossRef]
- Xia, P.-F.; Zhang, G.-C.; Liu, J.-J.; Kwak, S.; Tsai, C.-S.; Kong, I.I.; Sung, B.H.; Sohn, J.-H.; Wang, S.-G.; Jin, Y.-S. GroE chaperonins assisted functional expression of bacterial enzymes in Saccharomyces cerevisiae. Biotechnol. Bioeng. 2016, 113, 2149–2155. [Google Scholar] [CrossRef]
- Bengtsson, O.; Hahn-Hägerdal, B.; Gorwa-Grauslund, M.F. Xylose reductase from Pichia stipitis with altered coenzyme preference improves ethanolic xylose fermentation by recombinant Saccharomyces cerevisiae. Biotechnol. Biofuels 2009, 2, 9. [Google Scholar] [CrossRef]
- Myers, K.S.; Riley, N.M.; MacGilvray, M.E.; Sato, T.K.; McGee, M.; Heilberger, J.; Coon, J.J.; Gasch, A.P. Rewired cellular signaling coordinates sugar and hypoxic responses for anaerobic xylose fermentation in yeast. PLOS Genet. 2019, 15, e1008037. [Google Scholar] [CrossRef] [PubMed]
- Endalur Gopinarayanan, V.; Nair, N.U. A semi-synthetic regulon enables rapid growth of yeast on xylose. Nat. Commun. 2018, 9, 1233. [Google Scholar] [CrossRef] [PubMed]
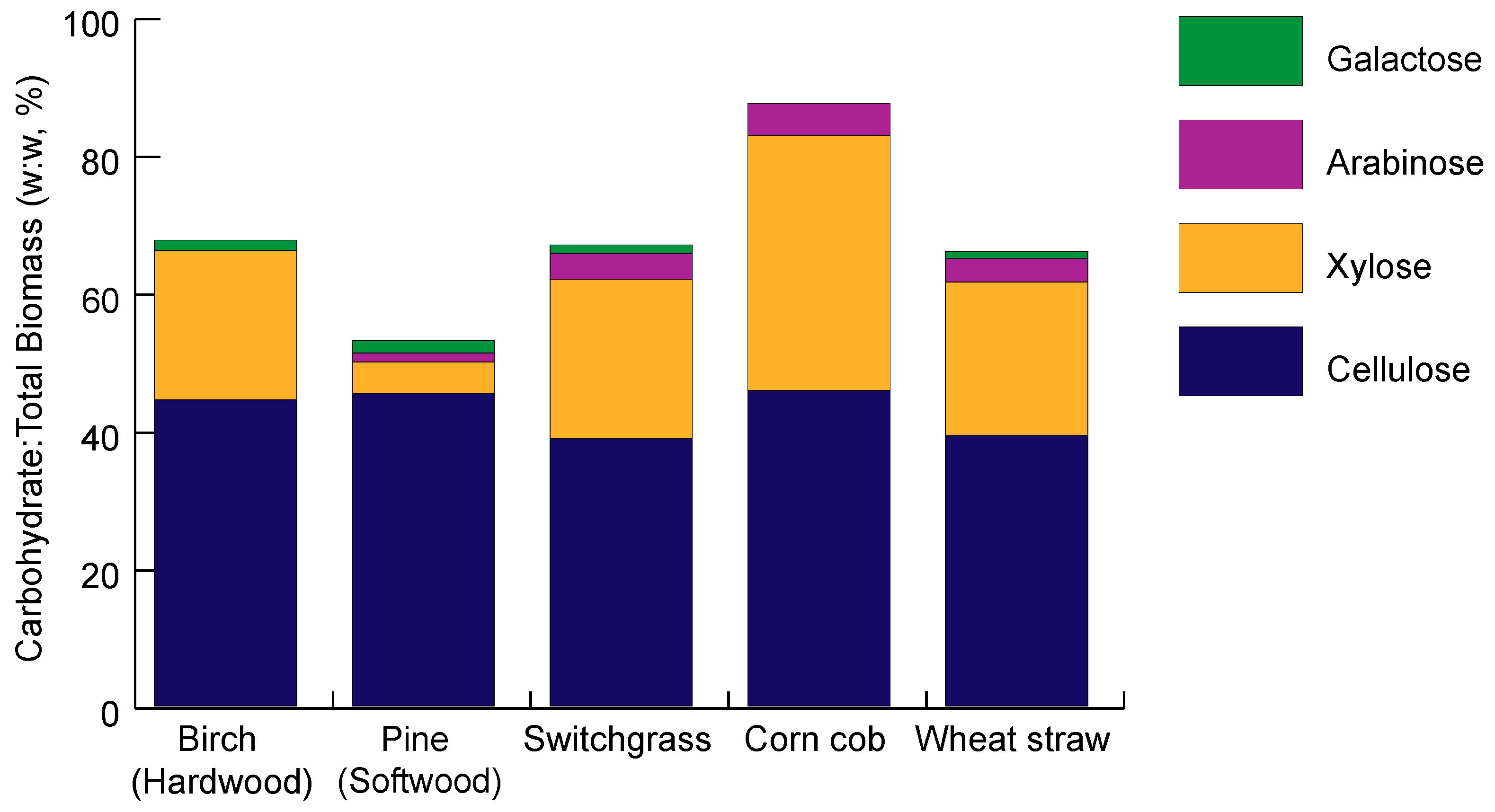
| Miscanthus | Switchgrass | Wheat Straw | Corn Stover | Poplar | Eucalyptus | Pine | Spruce | |
|---|---|---|---|---|---|---|---|---|
| % lignin | 9–27 | 17–25 | 19–27 | 18–23 | 21–31 | 29–35 | 28–35 | 27–31 |
| S/G ratio | 0.7–1.3 | 0.46–0.8 | 0.63–1.1 | 1.1 | 1.3–2.6 | 1.15 | 0 | 0 |
| Bond types (per aromatic ring) | ||||||||
| β-O-4 (aryl ether) | 0.41–0.42 | 0.42–0.56 | 0.29–0.45 | 0.457 | 0.43–0.68 | 0.62 | 0.48 | 0.45–0.53 |
| β-5 (phenylcoumaran) | 0.09 | 0.09 | 0.05–0.08 | 0.021 | 0.05–0.12 | 0.06 | 0.17 | 0.09–0.12 |
| β-β (resinol) | 0.04 | 0.1 | 0.01–0.06 | 0.015 | 0.06–0.11 | 0.17 | 0.05 | 0.02–0.04 |
| β-1 (Spirodienone) | trace | N/A | 0.004 | 0.005 | 0.09–0.12 | N/A | N/A | 0.01–0.02 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chaves, J.E.; Presley, G.N.; Michener, J.K. Modular Engineering of Biomass Degradation Pathways. Processes 2019, 7, 230. https://doi.org/10.3390/pr7040230
Chaves JE, Presley GN, Michener JK. Modular Engineering of Biomass Degradation Pathways. Processes. 2019; 7(4):230. https://doi.org/10.3390/pr7040230
Chicago/Turabian StyleChaves, Julie E., Gerald N. Presley, and Joshua K. Michener. 2019. "Modular Engineering of Biomass Degradation Pathways" Processes 7, no. 4: 230. https://doi.org/10.3390/pr7040230
APA StyleChaves, J. E., Presley, G. N., & Michener, J. K. (2019). Modular Engineering of Biomass Degradation Pathways. Processes, 7(4), 230. https://doi.org/10.3390/pr7040230





