1. Introduction
Lactic acid, a naturally occurring organic acid also known as 2-hydroxypropanoic acid with two optical isomers: L-lactic acid and D-lactic acid [
1]. It and its derivatives have extensive applications in the food, feed, pharmaceutical, cosmetic, and textile industries, serving roles as preservatives, acidulants, antimicrobial agents, moisturizers, and cleansers [
2]. Furthermore, when lactic acid meets specific chemical and optical purity standards, it is classified as polymer-grade. This high-purity grade is the primary feedstock for polylactic acid (PLA), a biodegradable polymer that offers a promising alternative to conventional plastics [
3]. Due to its potential to address energy and environmental challenges, PLA has broad application prospects [
4].
Lactic acid is primarily produced through fermentation and chemical synthesis, with fermentation being the preferred method due to its safety and cost-effectiveness [
5]. As a result, microbial fermentation is gradually becoming the industry standard, prompting increased research on identifying and developing superior microbial strains for this purpose. Currently, the most commonly used strains for fermentative lactic acid production are
Rhizopus and
Lactobacillus species [
6,
7]. Among
Rhizopus species,
Rhizopus oryzae,
Rhizopus nigricane,
Rhizopus chinensis, and
Rhizopus stolonifer are all capable of fermenting lactic acid.
Rhizopus strains, particularly
R. oryzae, are notably protein-rich, and their mycelial or granular structure facilitates easy separation from the fermentation broth. It is the key characteristic that effectively reduces separation costs. However, a significant limitation of
Rhizopus species is the heterofermentative metabolism. This process yields various byproducts such as ethanol, fumaric acid, and succinic acid except lactic acid, which complicates purification [
8]. Consequently, homofermentative lactic acid bacteria (LAB) are increasingly used for industrial production.
LAB are Gram-positive, non-spore-forming bacteria known for fermenting sugars into lactic acid [
9,
10]. Research indicates that these bacteria are favored for their high lactic acid production capacity, low byproduct formation, tolerance to higher temperatures, and relatively high acid production rates [
11]. However, these advantages are partly counterbalanced by their concomitant demand for more complex nutritional supplementation. The primary bacteria currently used for lactic acid production primarily include lactic acid bacteria (LAB),
Bacillus strains,
Escherichia coli, and
Corynebacterium glutamicum [
12]. In recent years,
Lactobacillus rhamnosus has been the potential specie for industrial lactic acid production [
13]. Preliminary laboratory work observed a rapid pH decline in cultures of
L. sakei, indicating it may also possess strong acid-producing capabilities. Although this is relatively apparent common in lactic acid bacteria, it is still an important phenomenon for evaluating the potential of strain for lactic acid production.
L. sakei belongs to the genus
Lactobacillus within the phylum firmicutes. It is a common microorganism in food fermentation, frequently applied in meat, dairy [
14], and vegetable products [
15]. This bacterium can alter food flavor and texture and produce bacteriocins [
16,
17]. Despite its well-documented applications, its potential for lactic acid fermentation remains largely unexplored. The extent to which the basal medium composition influences the apparent fermentation of this bacterial strain remains inadequately characterized.
This study focused on L. sakei L-7, a strain with high potential for lactic acid production. Using a single-factor method, we identified basic fermentation conditions for preliminary optimal production. Subsequently, response surface methodology (RSM) was employed to refine these conditions and maximize lactic acid production. The aforementioned research preliminarily established the optimal fermentation conditions for L. sakei L-7 lactic acid production and clarified the interactions among various factors, providing a foundation for investigation into its industrial-scale potential for lactic acid production.
2. Materials and Methods
2.1. Materials
The de Man, Rogosa and Sharpe (MRS) medium is a weakly selective medium commonly used in food microbiology and probiotic research for isolating, identifying, and cultivating lactic acid bacteria. It is typically composed of: glucose (20 g/L), yeast extract (5 g/L), trypticase (10 g/L), beef extract (10 g/L), CH3COONa (5 g/L), K2HPO4 (2 g/L), ammonium citrate (2 g/L), MgSO4·7H2O (0.58 g/L), and MnSO4·H2O (0.25 g/L). Additionally, Tween 80 (polyoxyethylene (20) sorbitan monolaurate) was added at 1 mL/L, with a pH of the medium was adjusted to 6.0.
The solidification of plate media was achieved by supplementing MRS medium with 15 g/L agar and 20 g/L calcium carbonate.
Reagents were sourced from various suppliers: glucose, ammonium citrate, CH3COONa, K2HPO4, MgSO4·7H2O, MnSO4 · H2O, and Tween 80 were purchased from Tianjin Jiangtian Chemical Technology Co., Ltd., Tianjin, China, while soybean meal extract, soya peptone, corn flour, and ammonium sulfate came from Shanghai Shengong Bioengineering Technology Service Co., Ltd., Shanghai, China. Yeast extract, trypticase, and beef extract were provided by Beijing Dingguo Changsheng Biotechnology Co., Ltd., Beijing, China.
2.2. Isolation and Identification of Bacterial Strains Used in Experiments
The strain
Latilactobacillus sakei L-7, isolated from homemade sausages, was used in this study. In the preliminary work, the probiotic effects of
L. sakei L-7 and characterized the relevant properties of their exopolysaccharide metabolites were investigated [
18,
19]. As an unmodified lactic acid bacterium, its metabolism pathway has been characterized as heterolactic fermentation in shaking flask culture. The 16S rRNA gene sequence has been submitted to the NCBI GenBank database under accession numbers OP648256–OP648257 [
18]. The isolation and identification of this strain were both carried out by Feng Fang [
19]. The brief process of the isolation and identification was as follows.
Homemade sausage was aseptically sectioned and homogenized in a mortar with MRS medium. Serial dilutions of the homogenate were prepared, yielding concentrations from 10−6 to 10−9. Aliquots (200 µL) from each dilution were spread onto the selective agar plates and incubated at 30 °C. After colony formation, isolates demonstrating CaCO3 hydrolysis through distinct clear zones were selected. These isolates were cultivated in MRS broth at 30 °C for 24 h. Strains associated with high lactic acid production, based on the size of the clear zones, were further purified through successive streak plating. The resulting pure cultures were cataloged and stored.
Total genomic DNA was extracted from well-grown fermentation broths using the commercial bacterial genomic DNA extraction kit (Tiangen Biochemical Technology Co., Ltd., Beijing, China). The 16S rDNA region was subsequently amplified by polymerase chain reaction (PCR) with the universal primers 16SF (5′–AGGTTTGATCCTGCTCAG–3′) and 16SR (5′–AGGTTACCTTGTTACGACTT–3′), which were synthesized by the company (Genewiz Biotechnology Co., Ltd., Beijing, China). The PCR amplicons were verified by agarose gel electrophoresis, purified using the MiniBEST Agarose Gel DNA Extraction Kit (Version 4.0) (Takara Biomedical Technology Co., Ltd., Dalian, China), and finally sent to the company (Genewiz Biotechnology Co., Ltd., Beijing, China) for sequencing.
2.3. Single-Factor Experimental Methods
2.3.1. Effect of Carbon Source Concentration on Strain Growth and Lactic Acid Production
The bacterial strain was cultivated in a defined medium with 10 g/L trypticase, 10 g/L beef extract, 5 g/L yeast extract, 2 g/L K
2HPO
4 (The effective buffering range of K
2HPO
4 was pH 5.8–8.0, while the target pH of the MRS medium used in this experiment was 6.0), 5 g/L CH
3COONa, 2 g/L ammonium citrate, 0.58 g/L MgSO
4·7H
2O, 0.25 g/L MnSO
4·H
2O, and 1 mL/L Tween 80 (as an emulsifier and stabilizer for cell membranes). This basal medium was supplemented with varying concentrations of glucose (20–140 g/L) as the sole carbon source. Each culture was inoculated with a 2 mL/L activated bacterial suspension and incubated at 30 °C with shaking at 150 rpm for 24 h. Following incubation, bacterial growth was measured by the optical density at 600 nm (OD
600) using the UV spectrophotometer (UV-1601, Mapada Instruments, Shanghai, China), with uninoculated medium as a blank. Finally, the lactic acid content in the culture broth was quantified via liquid chromatography. More information about liquid chromatography conditions is explained in
Section 2.5.
2.3.2. Effects of Nitrogen Source Type and Concentration on Lactic Acid Fermentation
Nitrogen source optimization was conducted using the previously optimized medium, which contained 40 g/L glucose, 2 g/L K
2HPO
4, 5 g/L CH
3COONa, 0.58 g/L MgSO
4·7H
2O, 0.25 g/L MnSO
4·H
2O, and 1 mL/L Tween 80. First, several nitrogen sources were evaluated including ammonium citrate, soybean meal extract, soya peptone, tryptone, yeast extract, and beef extract, which were as the sole nitrogen source at a 10 g/L concentration. Cultures were inoculated with a 2 mL/L activated bacterial suspension and incubated under the conditions specified in
Section 2.3.1 to assess their impact on OD
600 and lactic acid production. Following the identification of yeast extract as the preliminary optimal nitrogen source, its concentration was further investigated. Gradient concentrations of yeast extract (10–45 g/L) were added to the medium as the sole source of nitrogen. Fermentation was then carried out with a 2 mL/L inoculum under the same conditions as before, and the resulting OD
600 and lactic acid production were measured.
2.3.3. Effect of Salt Concentration on Lactic Acid Fermentation
Building on the optimized medium, next experiments were conducted to examine the effects of individual organic and inorganic and organic salts. The concentrations of CH3COONa (4–14 g/L), K2HPO4 (0–8 g/L), MgSO4·7H2O (0–1 g/L), and MnSO4·H2O (0–4 g/L) were varied while all other culture conditions remained constant. The OD600 and lactic acid production were monitored to determine the optimal concentration for each salt.
2.3.4. Effect of Temperature on Lactic Acid Fermentation
To determine the optimal growth temperature, the seed culture was inoculated at 2 mL/L into the optimized medium (40 g/L glucose, 35 g/L yeast extract, 6 g/L K2HPO4, 10 g/L CH3COONa, 4 g/L MgSO4·7H2O, 2 g/L MnSO4·H2O, and 1 mL/L Tween 80) at an initial, unadjusted pH of approximately 6.5. The cultures were incubated at temperatures ranging from 28 to 42 °C at 150 rpm. The optimal temperature was identified by measuring the OD600 and lactic acid production.
2.3.5. Effect of Initial pH on Lactic Acid Fermentation
The activated bacterial suspension was inoculated into the optimized medium adjusted to different initial pH values (5.5–9.5), including an unadjusted control (approximately pH 6.5). All cultures were incubated at 30 °C and 150 rpm for 24 h, after which the effects on fermentation production were determined by measuring lactic acid production and OD600.
2.4. RSM Experimental Methods
2.4.1. RSM for Optimizing Lactic Acid Fermentation Conditions
RSM is a statistical technique used to optimize multivariate systems. It quantifies the effects of factor interactions on response values and identifies optimal conditions by constructing second-order polynomial models [
20]. These models are visualized using three-dimensional surface and two-dimensional contour plots. In the surface plots, the steepness of the slope indicates the significance of a factor’s effect. Correspondingly, in contour plots, the density of the lines reflects the intensity of the factor’s influence, while an elliptical shape suggests a strong interaction between factors. This approach provides an intuitive and reliable basis for experimental optimization.
Leveraging the insights from the single-factor experiments, this study employed RSM to further enhance lactic acid yield. Three key variables were selected for optimization: nitrogen source addition, initial pH, and cultivation temperature. A three-factor, three-level Box–Behnken design was adopted, with the independent variables defined as: (A) nitrogen source concentration, (B) initial pH, and (C) temperature. Each factor was tested at three coded levels: −1 (lower limit), 0 (reference level), and +1 (upper limit). All experiments used the previously optimized base medium, with lactic acid production as the response variable. Each experimental group was conducted in triplicate to ensure reliability, and the specific factor-level combinations are detailed in
Table 1. The experimental design and data analysis were performed using Design Expert 13 software to model the results and determine the optimal fermentation conditions.
2.4.2. Response Surface Model Validation Experiments
This study utilized Design-Expert software to perform a RSM analysis. A quadratic polynomial model was fitted to the experimental data to describe the relationships between the independent variables and the response. The significance of this model was then evaluated through regression analysis and analysis of variance (ANOVA). This analysis identified the equation’s extrema and the optimal combination of variables. Finally, validation experiments for lactic acid fermentation were conducted based on the model’s predictions. The close agreement between the predicted and experimental results confirmed the model’s accuracy and established the optimized process conditions. To verify the accuracy of the model, independent replicate experiments should be performed at the predicted optimal conditions at least three times.
2.5. Analytical Method
The optical density (OD) of bacterial growth was measured at 600 nm using a UV spectrophotometer (UV-1601, Mapada Instruments, Shanghai, China).
Following filtration and centrifugation of the fermentation broth, the supernatant was collected for lactic acid production quantification via high-performance liquid chromatography (HPLC). The analysis was performed on a Waters E2695 HPLC system equipped with a diode array UV–visible detector and an Agilent C18 column (4.6 mm × 250 mm, 5 μm). The mobile phase consisted of 0.02 mol/L KH2PO4 aqueous solution (pH 2.3) and acetonitrile in a ratio of 90:10 (v/v), delivered at a flow rate of 1 mL/min. The injection volume was 10 μL, the column temperature was maintained at 35 °C, and detection was carried out at 215 nm.
All experiments were repeated three times. For each experimental factor, the sample size was at least three. The data in the bar chart are presented as the mean ± standard deviation. Statistical analysis was performed using ANOVA in Design-Expert software (version 13.0). Differences between means were further analyzed by the least significant difference (LSD) method with SPSS 25.0 software, considering p-values less than 0.05 as statistically significant. Data groups labeled with the same letter are not significantly different from each other (p > 0.05), whereas data groups marked with different letters indicate statistically significant differences (p < 0.05). Groups sharing a common letter do not differ statistically (p > 0.05).
3. Results and Discussion
3.1. Effects of Carbon Source Concentration on Lactic Acid Fermentation
In microbial fermentation, the selection of carbon source is critical for maximizing lactic acid production. A suitable carbon source not only supplies energy but also serves as a building block for cellular components, aiding nutrient transport, boosting metabolism, and promoting bacterial growth [
21].
L. sakei L-7 efficiently converts glucose into lactic acid through glycolysis, making glucose an ideal choice for optimizing fermentation. Therefore, optimizing glucose concentration is essential for achieving high yields cost-effectively. Based on this, glucose was selected to investigate the effect of its initial concentration (20–140 g/L) on lactic acid production.
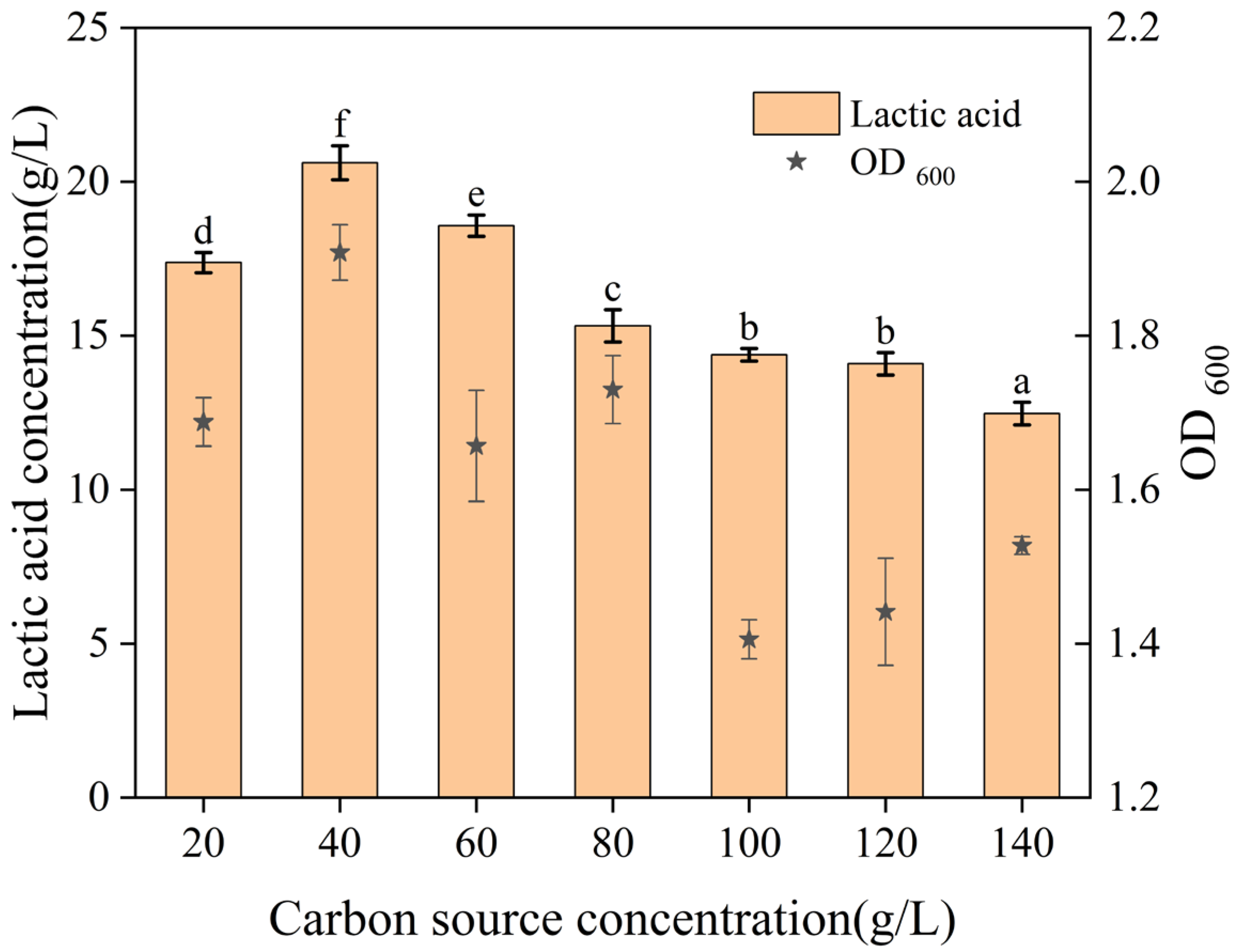
As shown in
Figure 1, lactic acid production initially increased and then decreased with rising glucose concentration, peaking at 20.61 ± 0.55 g/L with 40 g/L of glucose. This trend suggested that high glucose concentrations increase osmotic pressure, thereby inhibiting bacterial growth and metabolic activity. The subsequent decline in biomass beyond 40 g/L confirmed that excessively high carbon source concentrations inhibit
L. sakei L-7.
3.2. Effects of Nitrogen Source Type and Concentration on Lactic Acid Fermentation
Nitrogen sources are essential nutrients for microbial growth and metabolism, as they are primary components of biomolecules like proteins, nucleic acids, and coenzymes [
22]. However, different microbial strains vary significantly in their nitrogen utilization. This study aimed to assess the effects of various nitrogen sources on lactic acid fermentation of
L. sakei L-7. In addition to conventional organic nitrogen sources (yeast extract, beef extract, soya peptone and tryptone) [
22,
23,
24,
25], the investigation encompassed cost-effective alternatives such as soybean meal extract and corn flour, as well as inorganic sources including ammonium citrate and ammonium sulfate. The medium contained 40 g/L glucose, with all nitrogen sources set at 10 g/L.
As shown in
Figure 2, fermentation with organic nitrogen sources, such as beef extract (15.76 ± 0.16 g/L), soya peptone (17.32 ± 0.59 g/L), and tryptone (15.35 ± 0.22 g/L), were relatively more effective than with inexpensive nitrogen sources like corn flour (5.52 ± 0.30 g/L) and inorganic sources ammonium citrate (14.95 ± 0.57 g/L) and ammonium sulfate (14.64 ± 0.37 g/L). Yeast extract yielded the highest lactic acid production (18.93 ± 0.61 g/L) and supported robust cell growth. This outcome aligns with previous reports [
26,
27,
28], likely due to yeast extract not only contained abundant proteins, peptides, and free amino acids but also various vitamins and growth factors, providing comprehensive nutritional support for lactic acid bacteria growth, reproduction, and metabolic activities [
29].
Interestingly, while soybean peptone also supported high lactic acid production (17.32 ± 0.59 g/L), the corresponding microbial growth remained at a unremarkable level. This indicated that L. sakei L-7 may preferentially convert nutrients from soybean peptone into lactic acid rather than biomass. Furthermore, the inexpensive soybean meal extract yielded a lactic acid production (17.78 ± 0.38 g/L) only slightly lower than that achieved with yeast extract. However, since this study aimed to determine the maximum lactic acid production theoretically achievable by L. sakei L-7 under single-factor optimization, yeast extract powder was retained as the nitrogen source for subsequent experiments. Although yeast extract is optimal, cost-effective alternatives like soybean meal extract warrant further investigation for industrial applications where balancing yield and cost is crucial for market competitiveness.
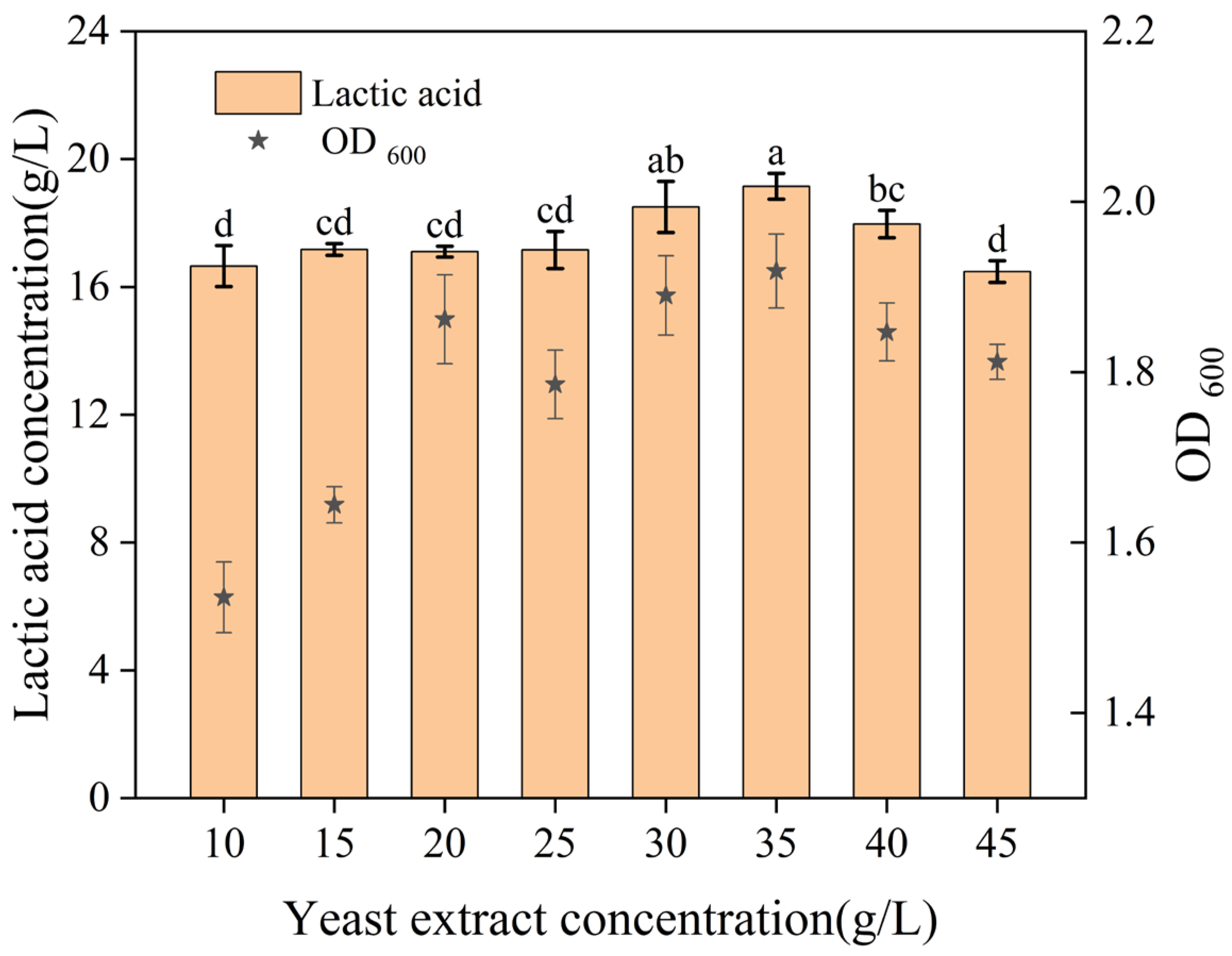
After the identification of yeast extract as the most effective nitrogen source, its concentration was varied to assess its impact.
Figure 3 showed that lactic acid production increased with yeast extract concentration up to a point. However, when the yeast extract concentration exceeded 35 g/L, it significantly decreased lactic acid production and probably inhibited bacterial growth. This phenomenon may be related to the inhibitory effects of high nitrogen concentrations on lactic acid bacteria. Consequently, the preliminary optimal yeast extract concentration was determined to be 35 g/L.
3.3. Effect of Salt Concentrations on Lactic Acid Fermentation
During lactic acid fermentation, salts served as essential nutrients that not only stabilized cell structures and regulated metabolic processes but also provided microorganisms with necessary micronutrients [
30]. While low concentrations of these ions significantly promoted microbial growth and metabolism, excessively high levels became inhibitory. This study investigated the effects of one organic salt and three inorganic salts—CH
3COONa, K
2HPO
4, MgSO
4, and MnSO
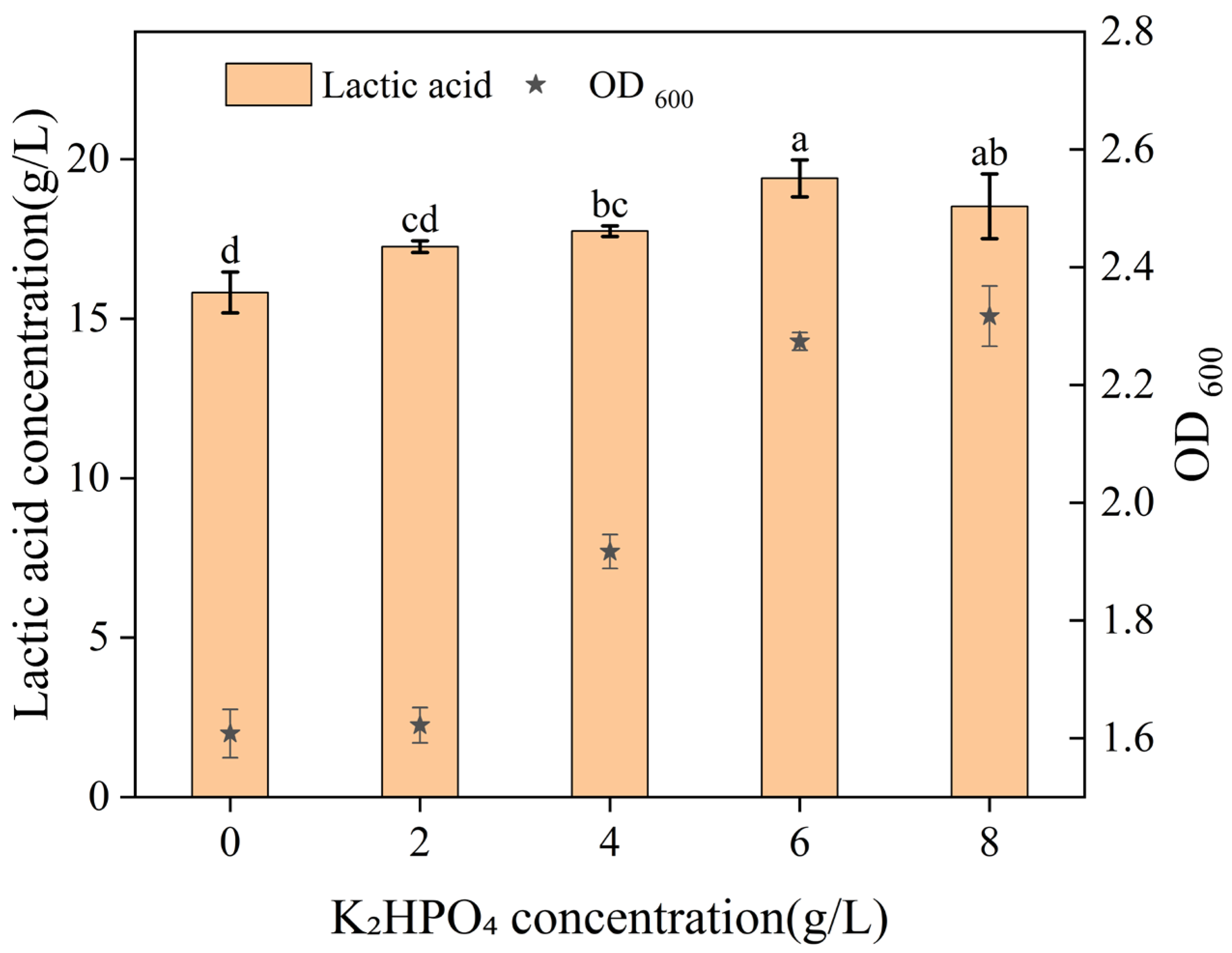
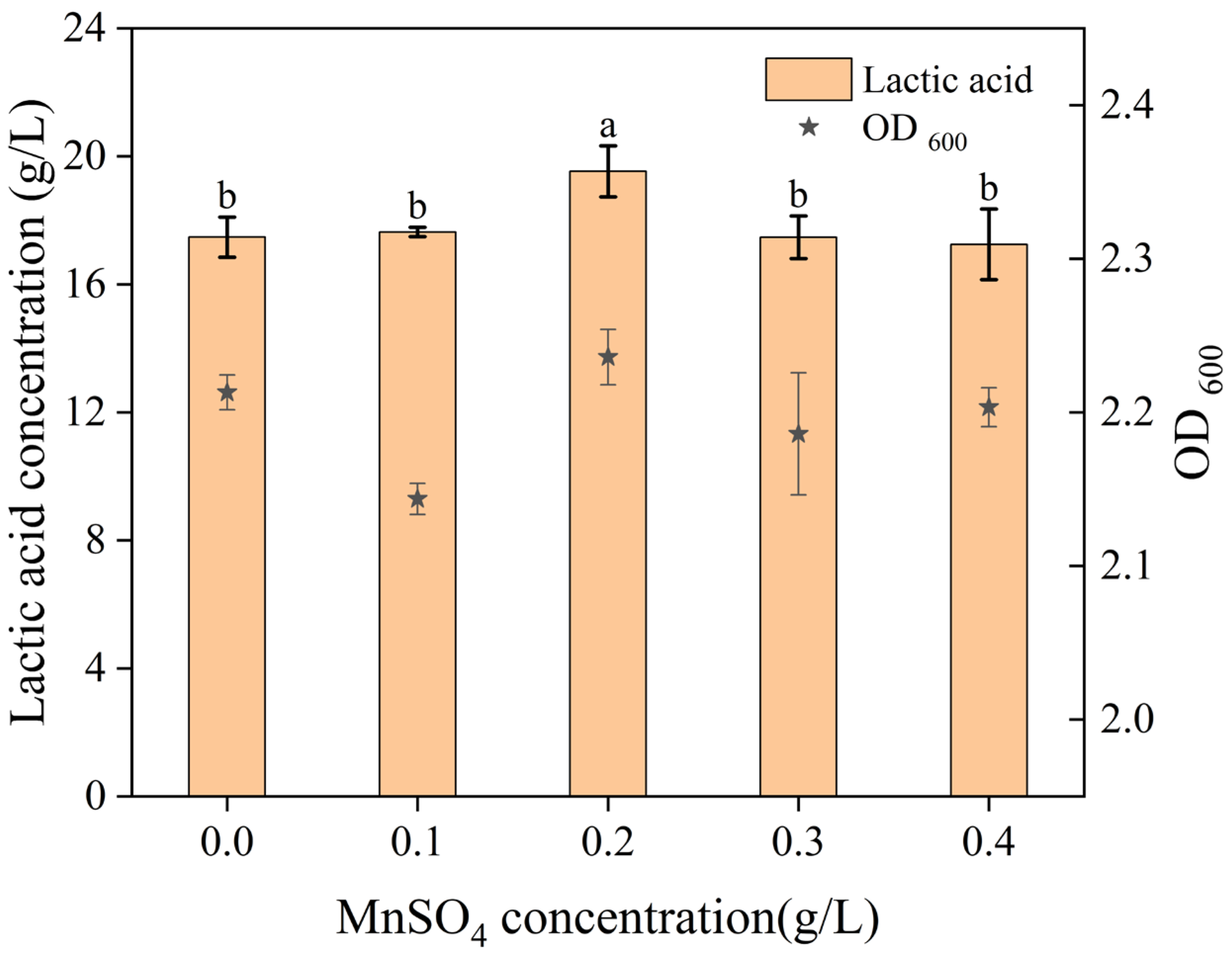
4—on the fermentation process under varying concentration conditions. As shown in
Figure 4,
Figure 5,
Figure 6 and
Figure 7, the optimal concentrations were determined to be 6 g/L for K
2HPO
4, 10 g/L for CH
3COONa, 0.2 g/L for MnSO
4, and 0.4 g/L for MgSO
4.
3.4. Effect of pH on Lactic Acid Fermentation
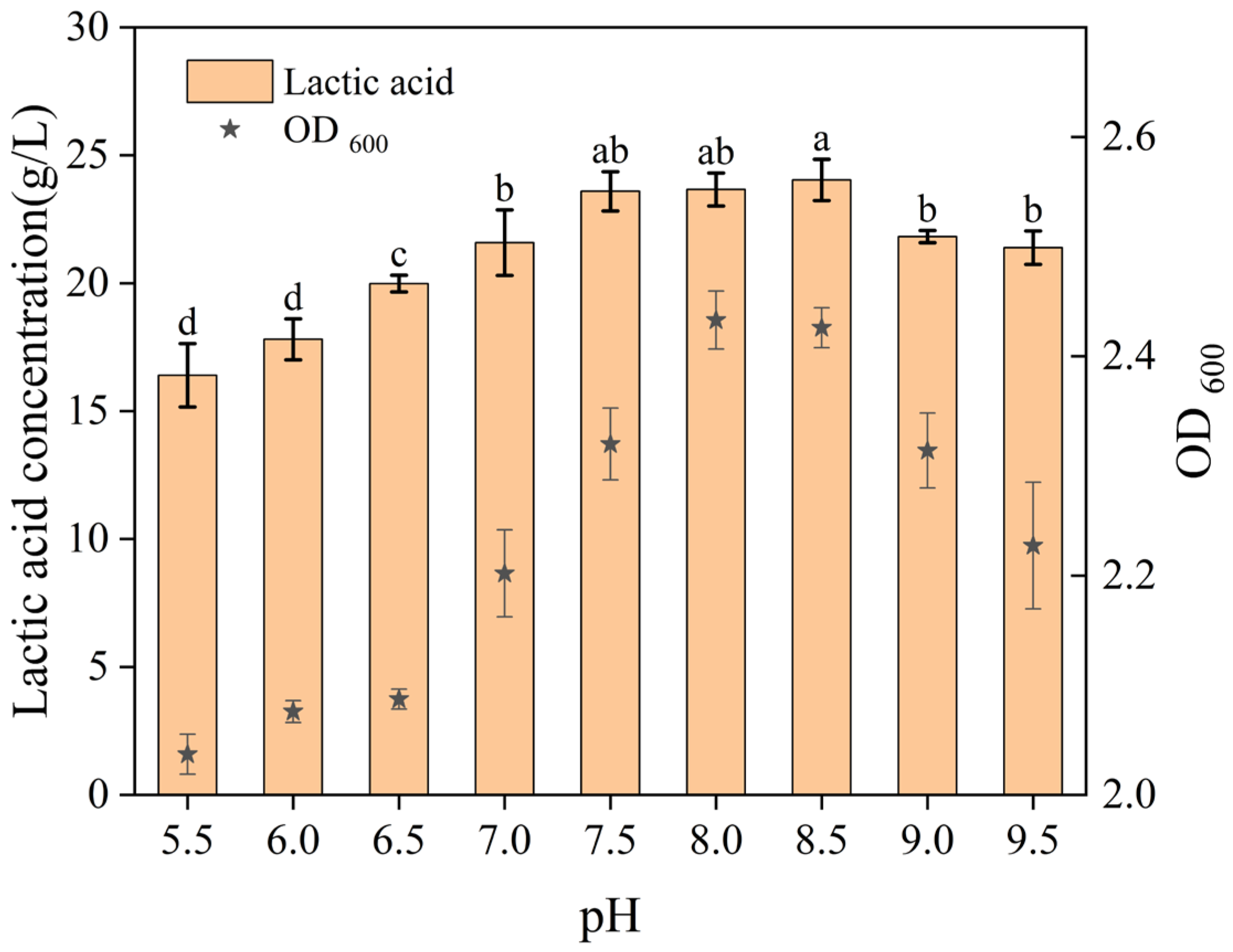
This study examined the effect of initial pH on bacterial growth and lactic acid production using the optimized MRS medium, with pH values ranging from 5.5 to 9.5. As shown in
Figure 8, the highest lactic acid production occurred at pH 8.0 and 8.5, which also supported robust strain growth. The corresponding lactic acid production reached 23.66 ± 0.65 g/L and 24.03 ± 0.81 g/L, respectively. According to the significance analysis, the labeling of significance indicates no statistically significant differences in lactic acid production among pH 7.5, 8.0, and 8.5 (sharing “a” or “ab”). Therefore, based on the highest mean value within this non-significant range, pH 8.5 was selected for subsequent experiments. The thriving bacterial growth and enhanced lactic acid yield at these moderately alkaline pH levels might be attributed to several factors. An initial pH above the pKa of lactic acid reduces its undissociated, inhibitory form, thereby minimizing acid stress on the cells [
31]. Furthermore, the buffering capacity of the medium at these starting pH values likely helped to maintain a more favorable microenvironment for a longer duration, supporting sustained metabolic activity [
32].
3.5. Effect of Temperature on Lactic Acid Fermentation
Temperature was a momentous factor in lactic acid fermentation, as it directly influenced the growth, metabolism, and product synthesis of lactic acid bacteria. Specifically, temperature affected the catalytic activity of intracellular enzymes, which in turn regulated the entire fermentation process. Within a specific range, increasing the temperature enhanced enzymatic efficiency, accelerating bacterial growth and metabolism, and ultimately promoting lactic acid accumulation. However, temperature exceeding a critical threshold can cause enzyme denaturation and inactivation, which accelerated cellular aging and death, thereby reducing lactic acid yield. Therefore, precise temperature control was essential for efficient production.
This study explored the effects of eight cultivation temperatures (28, 30, 32, 34, 36, 38, 40, and 42 °C) on lactic acid fermentation. As shown in
Figure 9, the lactic acid production followed a pattern similar to bacterial growth, with both parameters peaking at 30 °C. Below this optimum, the production increased with temperature; above it, the yield declined. This decrease is likely due to the inhibition of enzymatic activity and a shortened bacterial life cycle at higher temperatures. Based on these results, 30 °C was identified as the optimal temperature for lactic acid fermentation by
L. sakei L-7.
3.6. Optimization of Fermentation Conditions Using RSM
Building on the results of the single-factor experiments, this study utilized Design-Expert 13.0 software to further optimize the fermentation parameters. The RSM design was employed, focusing on the three key factors previously identified: nitrogen source concentration, pH, and temperature.
According to the results of preliminary single-factor experiments, this study utilized a three-factor, three-level Box–Behnken design. The specific experimental results were presented in
Table 2. A multiple regression analysis of this data, performed using Design-Expert software, produced the following mathematical model:
In this equation, Y represented lactic acid production, while A, B, and C were defined in
Section 2.4.1. Furthermore, regression analysis revealed that the absolute values of the regression coefficients for each variable in the equation positively correlated with their respective influence on the response value. A comparative analysis of the coefficient magnitudes reveals that pH exerted the most substantial influence on the experimental outcomes, followed by nitrogen source concentration, while temperature demonstrated the least pronounced effect.
As shown in
Table 3, the analysis of variance confirmed that the regression model is highly significant (
p < 0.0001). Meanwhile, the lack-of-fit test is not significant (
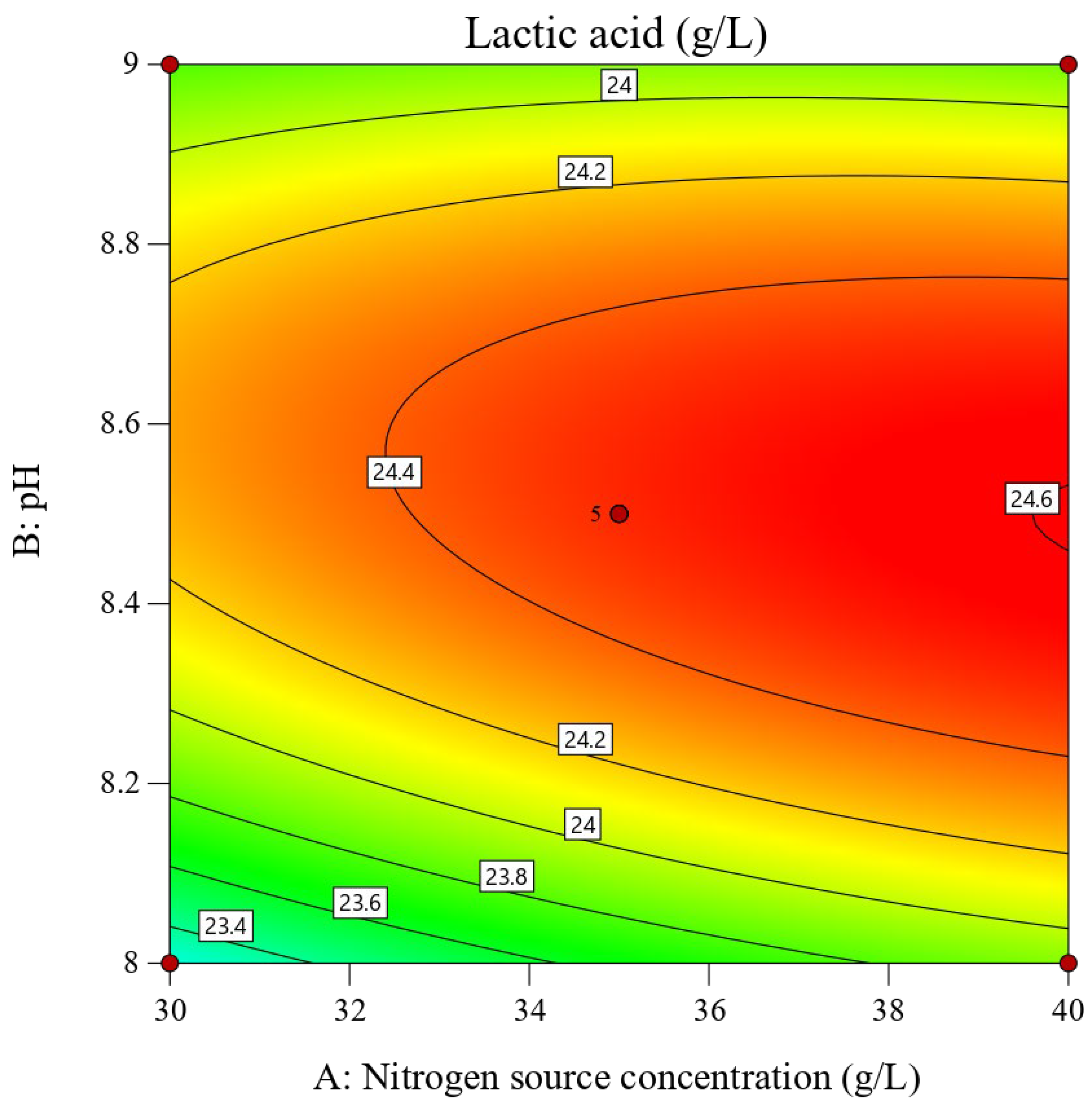
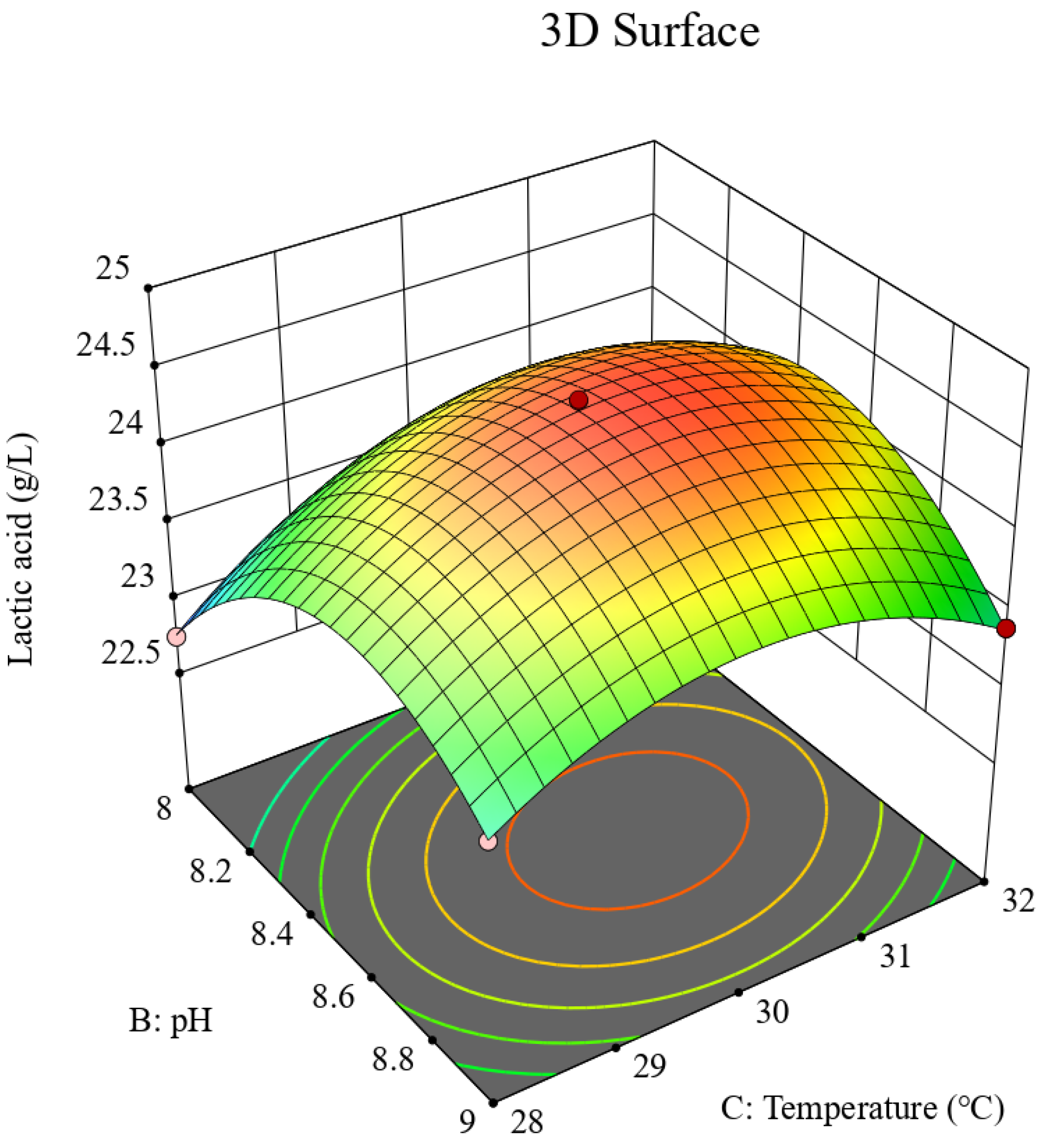
p > 0.05), indicating the model fitted the data well and had high predictive accuracy. To visualize the influence of the three parameters on the lactic acid fermentation process, interaction contour plots and three-dimensional response surface diagrams (
Figure 10,
Figure 11,
Figure 12,
Figure 13,
Figure 14 and
Figure 15) were generated. Finally, the fitted quadratic regression equation was used to determine the optimal values: a nitrogen source concentration of 39.54 g/L, a pH of 8.46, and a temperature of 30.27 °C, yielding a predicted lactic acid concentration of 24.62 g/L.
In this optimization experiment, lactic acid production was selected as the key indicator for determining the optimal conditions. In the experimental design, yeast extract was chosen as the nitrogen source, which may lead to an economically unfavorable balance between actual profit and incurred costs. The results obtained here primarily reflect the maximum theoretical lactic acid yield. However, if a low-cost nitrogen source is substituted in subsequent practical production, the cost-effectiveness of the process could be significantly enhanced.
In the development and optimization of fermentation processes, studying the concentrations of individual components independently serves as a necessary but insufficient starting point. The widespread presence of interactions (synergistic or antagonistic)—among nutritional components implies that simply superimposing results from single-factor experiments often fails to predict the actual fermentation performance in complex medium systems. While RSM has been employed to some extent to address this issue, it may not provide a complete picture. In subsequent phases of this study, multifactorial experimental designs beyond RSM are planned to be incorporated. This approach will allow for a systematic investigation of the interactions among key nutritional components, thereby enabling the construction of a composite medium system that not only contains appropriate component concentrations but also achieves internal balance to maximize the synthesis of the target product.
3.7. Response Surface Model Validation
To validate the model, a fermentation experiment was conducted using the optimized medium (glucose 40 g/L, yeast extract 39.54 g/L, CH
3COONa 10 g/L, K
2HPO
4 6 g/L, MnSO
4 0.2 g/L, MgSO
4 0.4 g/L, and 1 mL/L of Tween 80). The fermentation parameters were set as follows: a 100 mL liquid volume, a 14 h seed culture age, and a 2 mL/L inoculum. The culture was incubated at 30.27 °C and pH 8.46 with shaking at 150 rpm for 24 h. This process yielded 26.18 ± 0.44 g/L of lactic acid, a result that closely aligned with the model’s prediction of 24.62 g/L. The production obtained from the RSM validation experiment increased by a total of 50.6% compared to the yield obtained in
Section 3.1 (the lactic acid production of the 20 g/L glucose group was 17.38 ± 0.33 g/L). Consequently, these experimental findings confirmed that the RSM effectively simulated the lactic acid fermentation process for
L. sakei L-7, thereby verifying the model’s high reliability.
The results achieved through medium optimization are comparable to data published by other research groups. Kostov G et al. reported a lactic acid yield of 20.7 g/L from the cultivation of
L. casei ssp.
rhamnosus NBIMCC 1013 in a laboratory bioreactor under standard conditions for lactic acid bacteria without pH control after optimizing the experiment [
33]. In a study by Bhatt et al., the utilization of sugarcane molasses as a carbon source, coupled with
Lactobacillus delbrueckii NCIM 2025 as the microbial strain, resulted in a 6% enhancement in lactic acid production following a statistically designed multifactorial optimization experiment [
34].
It is worth noting that the OD600 value in this article is only used as a qualitative indicator of the growth status and non toxicity of the culture, rather than a quantitative standard for precise comparison of biomass. Moreover, it is not appropriate to evaluate bacterial growth solely by measuring the optical density at 600 nm. Because optical density cannot distinguish between live and dead cells, OD600 may remain at a high level even if the cell viability significantly decreases. For growth and meaningful quantitative assessment, it requires calibration by determining CFU with plate counting. Of course, the process of determining the number of viable bacteria in the culture medium through plate counting is very cumbersome. It is very convenient to evaluate bacterial growth by measuring the optical density at 600 nm. This is also the advantage of this method.
In the findings of this study, it was frequently observed that the conditions yielding the highest lactate production did not necessarily correspond to the highest OD
600 within the same experimental group. While lactate yield is the target product, OD
600 serves as an indicator of cellular growth within the fermenter. Different medium components can redirect the metabolic flux of the cells. Although the same microbial strain (
L. sakei) was investigated, the optimal medium formulation for maximizing its biomass, as reported by Tashakor A et al. [
35], differs considerably from the experimental results presented here. Consequently, it is postulated that an appropriate nutritional composition can channel more resources toward efficient product synthesis rather than toward biomass accumulation.
Previous research on
L. sakei primarily focuses on three areas: food, probiotics, and metabolomics [
36,
37,
38], while optimization experiments typically target bacteriocins and bacterial biomass [
35,
39]. Transcriptomic studies reveal that
L. sakei has a broad range of energetic substance utilization, suggesting its potential for industrial lactic acid production [
40]. However, only a limited number of studies have explored the use of
L. sakei in acid production experiments with agricultural and sideline products as substrates [
41]. There remains a significant deficiency in optimizing fermentation mediums using single-factor method and RSM, particularly in understanding how medium components influence lactic acid production by
L. sakei L-7. Additionally, research indicates that adding 1 mL/L ethanol, isopropanol, or other small molecules to the culture medium of
Escherichia coli induces stress and enhances protein production [
42,
43]. In light of these findings, our subsequent experiments will investigate the addition of growth factors or stress agents to the culture medium, while examining the strain’s performance in a more complex fermentation environment.
4. Conclusions
Through single-factor experiments, the preliminary optimal fermentation medium for L. sakei L-7 was determined to contain: 40 g/L glucose, 35 g/L yeast extract, 10 g/L CH3COONa, 6 g/L K2HPO4, 0.2 g/L MnSO4, 0.4 g/L MgSO4, and 1 mL/L Tween 80. The preliminary optimal fermentation conditions were established as a temperature of 30 °C and an initial pH of 8.5, without the use of neutralizing agents.
Subsequently, RSM was employed to further investigate the influence of key parameters. Temperature, nitrogen source concentration, and initial pH were selected as the critical factors. Using the established mathematical model, the optimal parameter combination was identified as a temperature of 30.27 °C, a nitrogen source concentration of 39.54 g/L, and a pH of 8.46. Noteworthily, the optimal values derived from response surface methodology should take precedence over those identified in single-factor experiments. Validation experiments under these conditions yielded 26.18 ± 0.44 g/L of lactic acid, a result consistent with the model’s prediction. The corresponding production rate was 1.091 g/L/h.
These experimental results partially address the existing knowledge gap regarding the influence of culture medium composition and concentration on lactic acid production by L.sakei. Furthermore, they provide preliminary references for material usage in practical production.
This productivity was relatively high, especially in the absence of prior, extensive fermentation process optimization. Future work should therefore focus on further process refinement to facilitate the industrial-scale production of lactic acid. And it is also possible to consider combining L.sakei with inexpensive raw materials for fermentation experiments, fully utilizing the metabolic characteristics of L.sakei.