Helical Foldamers and Stapled Peptides as New Modalities in Drug Discovery: Modulators of Protein-Protein Interactions
Abstract
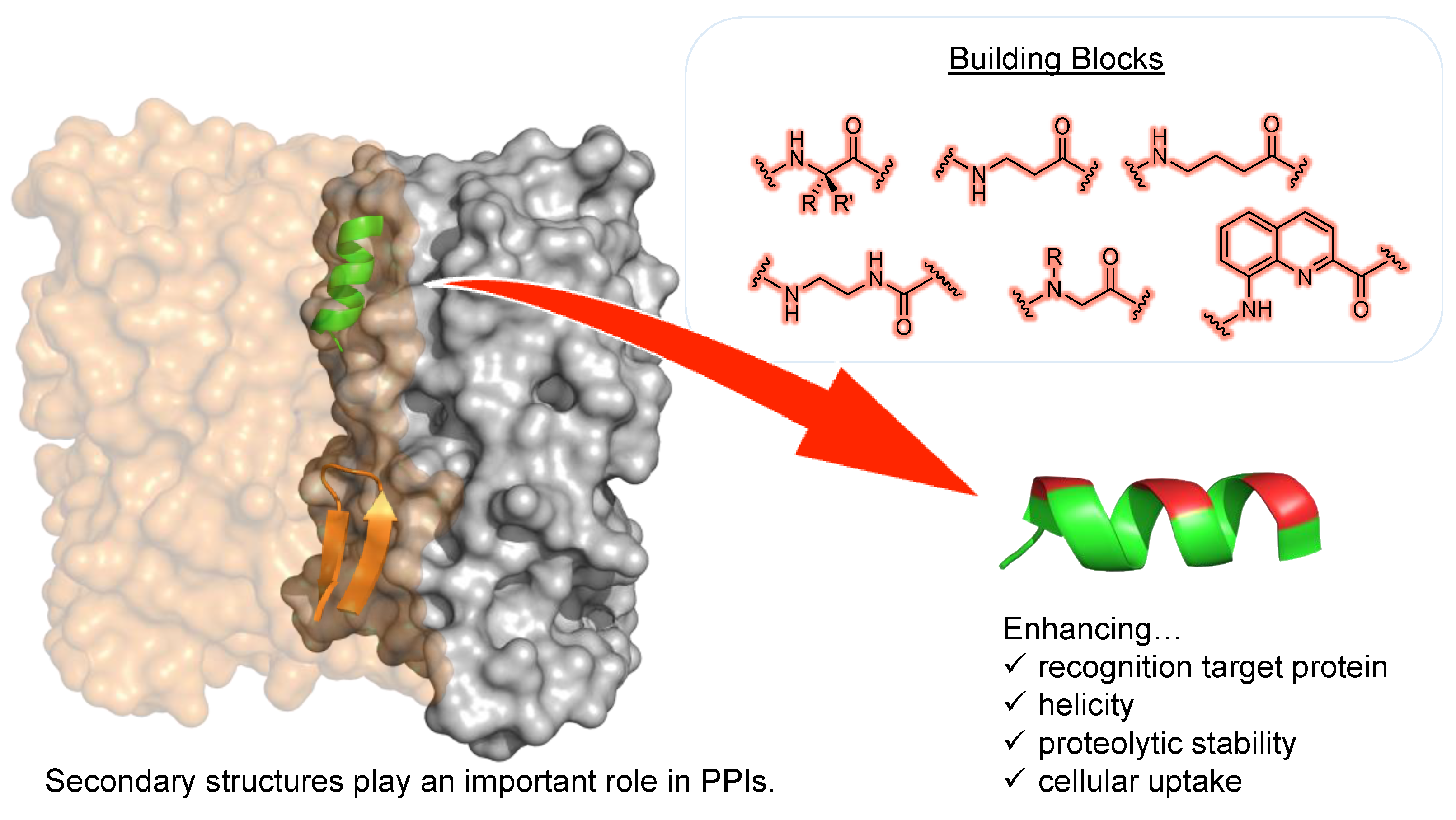
1. Introduction
2. α-Peptides
3. β-Peptides
4. γ-Peptides
5. Peptoids
6. Urea-Type Foldamers
7. Aromatic Foldamers and the Terphenyl Scaffold
8. Stapled Peptides
| Peptide Sequence and Structure | Target Protein | Ref. |
|---|---|---|
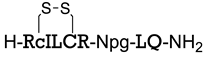  Lowercase letters: D-amino acid | ERα | [143] |
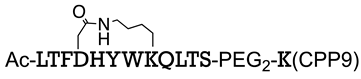 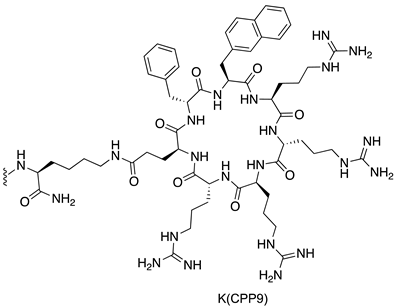 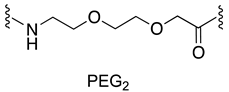 | MDM2/p53 | [145] |
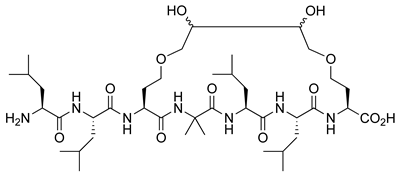 | VDR | [57] |
 | β-catenin/TCF | [150] |
 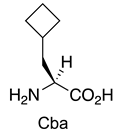 | MDM2/p53 MDMX/p53 | [151] |
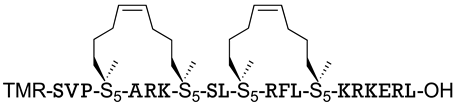 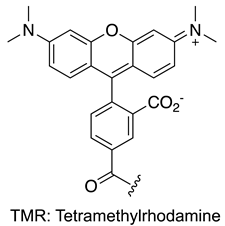 | MYC | [153] |
 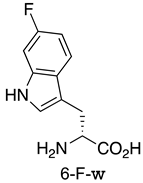 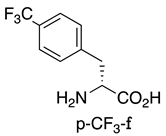 | MDM2/p53 MDM4/p53 | [155] |
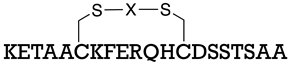  | S-protein | [157] |
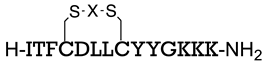 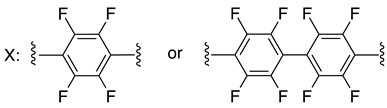 | C-terminal domain of an HIV-1 capsid assembly polyprotein (C-CA) | [158] |
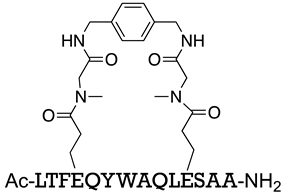 | MDM2/p53 MDMX/p53 | [160] |
X-NI-†-SLLRVQAHIRKKMV-NH2 | myoA tail interacting protein (MTIP) | [162] |
  | ERα | [168] |
9. Outlook
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Gellman, S.H. Foldamers: A Manifesto. Acc. Chem. Res. 1998, 31, 173–180. [Google Scholar] [CrossRef]
- Tyrikos-Ergas, T.; Fittolani, G.; Seerberger, P.H.; Delianco, M. Structural studies using unnatural oligosaccharides; toward sugar foldamers. Biomacromolecules 2020, 21, 18–29. [Google Scholar] [CrossRef] [PubMed]
- Hill, D.J.; Mio, M.J.; Prince, R.B.; Hughes, T.S.; Moore, J.S. A field guide to foldamers. Chem. Rev. 2001, 101, 3893–4012. [Google Scholar] [CrossRef]
- Girvine, Z.C.; Gellman, S.H. Foldamer catalysis. J. Am. Chem. Soc. 2020, 142, 17211–17223. [Google Scholar] [CrossRef]
- Legrand, B.; Aguesseau-kondrotas, J.; Simon, M.; Maillard, L. Catalytic foldamers: When the structure guides the functions. Catalysts 2020, 10, 700. [Google Scholar] [CrossRef]
- Kulkarni, K.; Habila, N.; Del Borgo, M.P.; Aguilar, M.I. Novel materials from the supramolecular self-assembly of short helical β3-peptide foldamers. Front. Chem. 2019, 7, 70. [Google Scholar] [CrossRef] [PubMed]
- Dey, S.; Misra, R.; Saseendran, A.; Pahan, S.; Gopi, H.N. Metal-coordinated supramolecular polymers from the minimalistic hybrid peptide foldamers. Angew. Chem. Int. Ed. Engl. 2021, 60, 9863–9868. [Google Scholar] [CrossRef] [PubMed]
- Vezenkov, L.L.; Martin, V.; Bettache, N.; Simon, M.; Messerschmitt, A.; Legrand, B.; Bantignies, J.L.; Subra, G.; Maynadier, M.; Bellet, V.; et al. Ribbon-like foldamers for cellular uptake and drug delivery. ChemBioChem 2017, 18, 2110–2114. [Google Scholar] [CrossRef]
- Bornerie, M.; Brion, A.; Guichard, G.; Kichler, A.; Douat, C. Delivery of siRNA by tailored cell-penetrating urea-based foldamers. Chem. Commun. 2021, 57, 1458–1461. [Google Scholar] [CrossRef]
- Bhaumik, K.N.; Hetényi, A.; Olajos, G.; Martins, A.; Spohn, R.; Németh, L.; Jojart, B.; Szili, P.; Dunai, A.; Jangir, P.K.; et al. Rationally designed foldameric adjucants enhance antibiotics efficacy via promoting membrane hyperpolarization. Mol. Syst. Des. Eng. 2021, 7, 21–33. [Google Scholar] [CrossRef]
- Bonnel, C.; Legrand, B.; Simon, M.; Clavié, M.; Masnou, A.; Jumas-Bilak, E.; Kang, Y.K.; Licznar-Fajardo, P.; Maillard, L.T.; Masurier, N. Tailoring the physicochemical properties of antimicrobial peptides onto a thiazole-based γ-peptide foldamers. J. Med. Chem. 2020, 63, 9168–9180. [Google Scholar] [CrossRef] [PubMed]
- Ferrand, Y.; Huc, I. Designing Helical Molecular Capsules Based on Folded Aromatic Amide Oligomers. Acc. Chem. Res. 2018, 51, 1970–1977. [Google Scholar] [CrossRef] [PubMed]
- Checco, J.W.; Gellman, S.H. Targeting recognition surfaces on natural proteins with peptidic foldamers. Curr. Opin. Struct. Biol. 2016, 39, 96–105. [Google Scholar] [CrossRef] [PubMed]
- Rinaldi, S. The Diverse World of Foldamers: Endless Possibilities of Self-Assembly. Molecules 2020, 25, 3276. [Google Scholar] [CrossRef]
- Gopalakrishnan, R.; Frolov, A.I.; Knerr, L.; Drury, W.J., III; Valeur, E. Therapeutic Potential of Foldamers: From Chemical Biology Tools to Drug Candidates? J. Med. Chem. 2016, 59, 9599–9621. [Google Scholar] [CrossRef]
- Haggag, Y.A.; Donia, A.A.; Osman, M.A.; El-Gizawy, S.A. Peptides as Drug Candidates: Limitations and Recent Development Perspectives. Biomed. J. Sci. Technol. Res. 2018, 8, 6659–6662. [Google Scholar] [CrossRef]
- Gentilucci, L.; De Marco, R.; Cerisoli, L. Chemical modifications designed to improve peptide stability: Incorporation of non-natural amino acids, pseudo-peptide bonds, and cyclization. Curr. Pharm. Des. 2010, 16, 3185–3203. [Google Scholar] [CrossRef]
- Antosova, Z.; Mackova, M.; Kral, V.; Macek, T. Therapeutic application of peptides and proteins: Parenteral forever? Trends Biotechnol. 2009, 27, 628–635. [Google Scholar] [CrossRef]
- Yokoo, H.; Misawa, T.; Demizu, Y. De Novo Design of Cell-Penetrating Foldamers. Chem. Rec. 2020, 20, 912–921. [Google Scholar] [CrossRef]
- Yokoo, H.; Hirano, M.; Misawa, T.; Demizu, Y. Helical Antimicrobial Peptide Foldamers Containing Non-proteinogenic Amino Acids. ChemMedChem 2021, 16, 1226–1233. [Google Scholar] [CrossRef]
- Oba, M. Cell-Penetrating Peptide Foldamers: Drug-Delivery Tools. ChemBioChem 2019, 20, 2041–2045. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.; Zhou, Q.; He, J.; Jiang, Z.; Peng, C.; Tong, R.; Shi, J. Recent advances in the development of protein-protein interactions modulators: Mechanisms and clinical trials. Signal Transduct. Target. Ther. 2020, 5, 213. [Google Scholar] [CrossRef] [PubMed]
- Qiu, Y.; Li, X.; He, X.; Pu, J.; Zhang, J.; Lu, S. Computational methods-guided design of modulators targeting protein-protein interactions (PPIs). Eur. J. Med. Chem. 2020, 207, 112764. [Google Scholar] [CrossRef]
- Edwards, T.A.; Wilson, A.J. Helix-mediated protein-protein interactions as targets for intervention using foldamers. Amino Acids 2011, 41, 743–754. [Google Scholar] [CrossRef] [PubMed]
- Watkins, A.M.; Arora, P.S. Anatomy of the β-strands at protein-protein interfaces. ACS Chem. Biol. 2014, 9, 1747–1754. [Google Scholar] [CrossRef] [PubMed]
- Sharma, G.V.; Babu, B.S.; Ramakrishna, K.V.; Nagendar, P.; Kunwar, A.C.; Schramm, P.; Baldauf, C.; Hofmann, H.J. Synthesis and structure of alpha/delta-hybrid peptides—Access to novel helix patterns in foldamers. Chemistry 2009, 15, 5552–5566. [Google Scholar] [CrossRef]
- Sharma, G.V.; Babu, B.S.; Chatterjee, D.; Ramakrishna, K.V.; Kunwar, A.C.; Schramm, P.; Hofmann, H.J. Theoretical and experimental studies on alpha/epsilon-hybrid peptides: Design of a 14/12-helix from peptides with alternating (S)-C-linked carbo-epsilon-amino acid [(S)-epsilon-Caa((x))] and L-ala. J. Org. Chem. 2009, 74, 6703–6713. [Google Scholar] [CrossRef]
- Laurencin, M.; Simon, M.; Fleury, Y.; Baudy-Floc’h, M.; Bondon, A.; Legrand, B. Selectivity Modulation and Structure of α/aza-β3 Cyclic Antimicrobial Peptides. Chemistry 2018, 24, 6191–6201. [Google Scholar] [CrossRef]
- Raghuraman, A.; Ko, E.; Perez, L.M.; Ioerger, T.R.; Burgess, K. Pyrrolonone-pyrrolidine oligomers as universal peptidemimetics. J. Am. Chem. Soc. 2011, 133, 12350–12353. [Google Scholar] [CrossRef]
- Sinatra, L.; Kolano, L.; Icker, M.; Fritzsche, S.R.; Volke, D.; Gockel, I.; Thieme, R.; Hoffmann, R.; Hansen, F.K. Hybrid Peptides Based on α-Aminoxy Acids as Antimicrobial and Anticancer Foldamers. ChemPlusChem 2021, 86, 827–835. [Google Scholar] [CrossRef]
- Guarracino, D.A.; Riordan, J.A.; Barreto, G.M.; Oldfield, A.L.; Kouba, C.M.; Agrinsoni, D. Macrocyclic control in helix mimetics. Chem. Rev. 2019, 119, 9915–9949. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, M. Design and synthesis of chiral α,α-disubstituted amino acids and conformational study of their oligopeptides. Chem. Pharm. Bull. 2007, 55, 349–358. [Google Scholar] [CrossRef] [PubMed]
- Crisma, M.; Toniolo, C. Helical screw-sense preferences of peptides based on chiral, Cα-tetrasubstituted α-amino acids. Biopolymers 2015, 104, 46–64. [Google Scholar] [CrossRef] [PubMed]
- Crisma, M.; De Zotti, M.; Formaggio, F.; Peggion, C.; Moretto, A.; Toniolo, C. Handedness preference and switching of peptide helices. Part II: Helices based on noncoded α-amino acids. J. Pept. Sci. 2015, 21, 148–177. [Google Scholar] [CrossRef]
- Demizu, Y.; Doi, M.; Kurihara, M.; Okuda, H.; Nagano, M.; Suemune, H.; Tanaka, M. Conformational studies on peptides containing α,α-disubstituted α-amino acids: Chiral cyclic α,α-disubstituted α-amino acid as an α-helical inducer. Org. Biomol. Chem. 2011, 9, 3303–3312. [Google Scholar] [CrossRef]
- Demizu, Y.; Okitsu, K.; Yamashita, H.; Doi, M.; Misawa, T.; Oba, M.; Tanaka, M.; Kurihara, M. α-Helical structures of oligopeptides with an alternating L-Leu-Aib segment. Eur. J. Org. Chem. 2016, 2815–2820. [Google Scholar] [CrossRef]
- Kobayashi, H.; Misawa, T.; Matsuno, K.; Demizu, Y. Preorganized cyclic α,α-disubstituted α-amino acids bearing functionalized side chains that act as peptide-helix inducers. J. Org. Chem. 2017, 82, 10722–10726. [Google Scholar] [CrossRef]
- Akagawa, K.; Higuchi, J.; Yoshikawa, I.; Kudo, K. Kinetic resolution of ansa cyclophanes by peptide-catalyzed aldol/retro-aldol reactions. Eur. J. Org. Chem. 2018, 38, 5278–5281. [Google Scholar] [CrossRef]
- Sato, K.; Umeno, T.; Ueda, A.; Kato, T.; Doi, M.; Tanaka, M. Asymmetric 1,4-addition reactions catalyzed by N-terminal thiourea-modified helical L-Leu peptide with cyclic amino acids. Chem. Eur. J. 2021, 27, 11216–11220. [Google Scholar] [CrossRef]
- Umeno, T.; Ueda, A.; Doi, M.; Kato, T.; Oba, M.; Tanaka, M. Helical foldamer-catalyzed enantioselective 1,4-addition reaction of dialkyl malonates to cyclic enones. Tetrahedron Lett. 2019, 60, 151301. [Google Scholar] [CrossRef]
- Yamashita, H.; Demizu, Y.; Shoda, T.; Sato, Y.; Oba, M.; Tanaka, M.; Kurihara, M. Amphipathic short helix-stabilized peptides with cell-membrane penetrating ability. Bioorg. Med. Chem. 2014, 22, 2403–2408. [Google Scholar] [CrossRef] [PubMed]
- Yamashita, H.; Oba, M.; Misawa, T.; Tanaka, M.; Hattori, T.; Naito, M.; Kurihara, M.; Demizu, Y. A helix-stabilized cell-penetrating peptide as an intracellular delivery tool. ChemBioChem 2016, 17, 137–140. [Google Scholar] [CrossRef] [PubMed]
- Yamashita, H.; Misawa, T.; Oba, M.; Tanaka, M.; Naito, M.; Kurihara, M.; Demizu, Y. Development of helix-stabilized cell-penetrating peptides containing cationic α,α-disubstituted amino acids as helical promoters. Bioorg. Med. Chem. 2017, 25, 1846–1851. [Google Scholar] [CrossRef]
- Misawa, T.; Ohoka, N.; Oba, M.; Yamashita, H.; Tanaka, M.; Naito, M.; Demizu, Y. Development of 2-aminoisobutyric acid (Aib)-rich cell-penetrating foldamers for efficient siRNA delivery. Chem. Commun. 2019, 55, 7792–7795. [Google Scholar] [CrossRef] [PubMed]
- Oba, M.; Ito, Y.; Umeno, T.; Kato, T.; Tanaka, M. Plasmid DNA delivery using cell-Penetrating peptide foldamers composed of Arg-Arg-Aib repeating sequences. ACS Biomater. Sci. Eng. 2019, 5, 5660–5668. [Google Scholar] [CrossRef] [PubMed]
- Uchida, S.; Yamaberi, Y.; Tanaka, M.; Oba, M. A helix foldamer oligopeptide improves intracellular stability and prolongs protein expression of the delivered mRNA. Nanoscale 2021, 13, 18941–18946. [Google Scholar] [CrossRef] [PubMed]
- Yokum, T.S.; Elzer, P.H.; McLaughlin, M.L. Antimicrobial α,α-dialkylated amino acid rich peptides with in-vivo activity against an intracellular pathogen. J. Med. Chem. 1996, 39, 3603–3605. [Google Scholar] [CrossRef]
- Misawa, T.; Imamura, M.; Ozawa, Y.; Haishima, K.; Kurihara, M.; Kikuchi, Y.; Demizu, Y. Development of helix-stabilized antimicrobial peptides composed of lysine and hydrophobic α,α-disubstituted α-amino acid residues. Bioorg. Med. Chem. Lett. 2017, 27, 3950–3953. [Google Scholar] [CrossRef] [PubMed]
- Goto, C.; Hirano, M.; Hayashi, K.; Kikuchi, Y.; Kudo-Hara, Y.; Misawa, T.; Demizu, Y. Development of Amphipathic Antimicrobial Peptide Foldamers Based on Magainin 2 Sequence. ChemMedChem 2019, 14, 1911–1916. [Google Scholar] [CrossRef]
- Hirano, M.; Saito, C.; Goto, C.; Yokoo, H.; Kawano, R.; Misawa, T.; Demizu, Y. Rational design of helix-stabilized antimicrobial peptide foldamers containing α,α-disubstituted amino acids or side-chain stapling. ChemPlusChem 2020, 85, 2731–2736. [Google Scholar] [CrossRef]
- Paterson, Y.; Rumsey, S.M.; Benedetti, E.; Neḿethy, G.; Scheraga, H. Sensitivity of polypeptide conformation to geometry. Theoretical conformational analysis of oligomers of alpha-aminoisobutyric acid. J. Am. Chem. Soc. 1981, 103, 2947–2955. [Google Scholar] [CrossRef]
- Benedetti, E. X-ray crystallography of peptides: The contribution of the Italian laboratories. Biopolymers 1996, 40, 3–44. [Google Scholar] [CrossRef]
- Karle, I.L. Controls exerted by the Aib residue: Helix formation and helix reversal. Biopolymers 2001, 60, 351–365. [Google Scholar] [CrossRef]
- Banerjee, R.; Basu, G.; Chène, P.; Roy, S. Aib-based peptide backbone as scaffolds for helical peptide mimics. J. Pept. Res. 2002, 60, 88–94. [Google Scholar] [CrossRef] [PubMed]
- Morimoto, J.; Hosono, Y.; Sando, S. Isolation of a peptide containing D-amino acid residues that inhibits the α-helix-mediated p53–MDM2 interaction from a one-bead one-compound library. Bioorg. Med. Chem. Lett. 2018, 28, 231–234. [Google Scholar] [CrossRef]
- Dhar, A.; Mallick, S.; Ghosh, P.; Maiti, A.; Ahmed, I.; Bhattacharya, S.; Mandal, T.; Manna, A.; Roy, K.; Singh, S.; et al. Simultaneous inhibition of key growth pathways in melanoma cells and tumor regression by a designed bidentate constrained helical peptide. Biopolymers 2014, 101, 344–358. [Google Scholar] [CrossRef]
- Demizu, Y.; Nagoya, S.; Shirakawa, M.; Kawamura, M.; Yamagata, N.; Sato, Y.; Doi, M.; Kurihara, M. Development of stapled short helical peptides capable of inhibiting vitamin D receptor (VDR)-coactivator interactions. Bioorg. Med. Chem. Lett. 2013, 23, 4292–4296. [Google Scholar] [CrossRef]
- Misawa, T.; Demizu, Y.; Kawamura, M.; Yamagata, N.; Kurihara, M. Structural development of stapled short helical peptides as vitamin D receptor (VDR)-coactivator interaction inhibitors. Bioorg. Med. Chem. 2015, 23, 1055–1061. [Google Scholar] [CrossRef]
- Wang, X.; Espinosa, J.F.; Gellman, S.H. 12-Helix formation in aqueous solution with short β-peptides containing purrolidine-based residues. J. Am. Chem. Soc. 2000, 122, 4821–4822. [Google Scholar] [CrossRef]
- Hart, S.A.; Bahadoor, A.B.F.; Matthews, E.E.; Qiu, X.J.; Schepartz, A. Helix macrodipole control of β3-peptide 14-helix stability in water. J. Am. Chem. Soc. 2003, 125, 4022–4023. [Google Scholar] [CrossRef]
- Seebach, D.; Hook, F.F.; Glättli, A. Helices and other secondary structures of β- and γ-peptides. Pep. Sci. 2006, 84, 23–37. [Google Scholar] [CrossRef] [PubMed]
- Reguse, T.R.; Lai, J.R.; Gellman, S.H. Evidence that the β-peptide 14-helix is stabilized by β3-residues with sidechain branching adjacent to the β-carbon. Helv. Chim. Acta 2002, 85, 4154. [Google Scholar] [CrossRef]
- Rueping, M.; Shreiber, J.V.; Lelais, G.; Jaun, B.; Seebach, D. Mixed β2/β3-hexapeptides and β2/β3-nonapeptides folding to (P)-helices with alternating twelve- and ten- membered hydrogen bonded rings. Helv. Chim. Acta 2002, 85, 2577–2593. [Google Scholar] [CrossRef]
- Appella, D.H.; Christiason, L.A.; Karle, I.L.; Powell, D.P.; Gellman, S.H. β-peptide foldamers: Robust helix formation in a new family of β-amino acid oligomers. J. Am. Chem. Soc. 1996, 118, 13071–13072. [Google Scholar] [CrossRef]
- Zhang, D.; Chen, Q.; Zhang, W.; Liu, H.; Wan, J.; Qian, Y.; Li, B.; Tang, S.; Liu, Y.; Chen, S.; et al. Silk-inspired β-peptide materials resist fouling and the foreign body response. Angew. Chem. Int. Ed. 2020, 59, 9586–9593. [Google Scholar]
- Girvin, Z.C.; Andrews, M.K.; Liu, X.; Gellman, S.H. Foldamer-templated catalysis of macrocyclic formation. Science 2019, 366, 1528–1531. [Google Scholar] [CrossRef] [PubMed]
- Andrews, M.K.; Liu, X.; Gellman, S.H. Tailoring reaction selectivity by modulating a catalytic diad on a foldamer scaffold. J. Am. Chem. Soc. 2022, 144, 2225–2232. [Google Scholar] [CrossRef]
- Checco, J.W.; Lee, E.F.; Evangelista, M.; Sleebs, N.; Rogers, K.; Pettikiriarachchi, A.; Kershaw, N.J.; Eddinger, G.A.; Belair, D.G.; Wilson, J.L.; et al. α/β-Peptide foldamers targeting intracellular protein-protein interaction with activity in living cells. J. Am. Chem. Soc. 2016, 137, 11365–11375. [Google Scholar] [CrossRef]
- Horne, W.S.; Johnson, L.M.; Ketas, T.J.; Klasse, P.J.; Lu, M.; Moore, J.P.; Gellman, S.H. Structural and biological mimicry of protein surface recognition by α/β-peptide foldamers. Proc. Natl. Acad. Sci. USA 2009, 106, 14751–14756. [Google Scholar] [CrossRef]
- Johnson, L.M.; Gellman, S.H. α-Helix mimicry with α/β-peptides. Methods Enzymol. 2013, 523, 407–429. [Google Scholar]
- Nagy, A.; Goldschmidt Gőz, V.; Pintér, I.; Farkas, V.; Perczel, A. α/β-Chimera peptide synthesis with cyclic β-sugar amino acids: The efficient coupling protocol. Amino Acids 2019, 51, 669–678. [Google Scholar] [CrossRef] [PubMed]
- Milbeo, P.; Martinez, J.; Amblard, M.; Calmès, M.; Legrand, B. 1-Aminobicyclic [2.2.2]octane-2-carboxylic acid and derivatives as chiral constrained bridged scaffolds for foldamers and chiral catalysts. Acc. Chem. Res. 2021, 54, 685–696. [Google Scholar] [CrossRef] [PubMed]
- Demizu, Y.; Oba, M.; Okitsu, K.; Yamashita, H.; Misawa, T.; Tanaka, M.; Kurihara, M.; Gellman, S.H. A preorganized β-amino acid bearing a guanidinium side chain and its use in cell-penetrating peptides. Org. Biomol. Chem. 2015, 13, 5617–5620. [Google Scholar] [CrossRef] [PubMed]
- Hamuro, Y.; Schneider, J.P.; Degrado, W.F. De novo design of antibacterial β-peptide. J. Am. Chem. Soc. 1999, 121, 12200–12201. [Google Scholar] [CrossRef]
- Porter, E.A.; Wang, X.; Lee, H.S.; Weisblum, B.; Gellman, S.H. Non-haemolytic β-amino acid oligomers. Nature 2000, 404, 565. [Google Scholar] [CrossRef] [PubMed]
- Kritzer, J.A.; Hodsdon, M.E.; Schepartz, A. Solution structure of a β-peptide ligand for hDM2. J. Am. Chem. Soc. 2005, 127, 4118–4119. [Google Scholar] [CrossRef]
- Stephens, O.M.; Kim, S.; Welch, B.D.; Hodsdon, M.E.; Kay, M.S.; Schepartz, A. Inhibiting HIV fusion with a β-peptide foldamer. J. Am. Chem. Soc. 2005, 127, 13126–13127. [Google Scholar] [CrossRef]
- Kritzer, J.A.; Stephens, O.M.; Guarracino, D.A.; Reznik, S.K.; Schepartz, A. β-Peptides as inhibitors of protein-protein interactions. Bioorg. Med. Chem. 2005, 13, 11–16. [Google Scholar] [CrossRef]
- Johnson, L.M.; Mortenson, D.E.; Yun, H.G.; Horne, W.S.; Ketas, T.J.; Lu, M.; Moore, J.P.; Gellman, S.H. Enhancement of α-helix mimicry by an α/β-peptide foldamer via incorporation of dense ionic side-chain array. J. Am. Chem. Soc. 2012, 134, 7317–7320. [Google Scholar] [CrossRef]
- Checco, J.W.; Kreitler, D.F.; Thomas, N.C.; Belair, D.G.; Rettko, N.J.; Murphy, W.L.; Forest, K.T.; Gellman, S.H. Targeting diverse protein-protein interaction interfaces with α/β-peptides derived from the Z-domain scaffold. Proc. Natl. Acad. Sci. USA 2015, 112, 4552–4557. [Google Scholar] [CrossRef]
- Checco, J.W.; Gellman, S.H. Iterative nonproteinogenic residue incorporation yields α/β-peptides with Helix-loop-helix tertiary structure and high affinity for VEGF. ChemBioChem 2017, 18, 291–299. [Google Scholar] [CrossRef] [PubMed]
- Hanessian, S.; Luo, X.; Shaum, R.; Michnick, S. Design of secondary structure in unnatural peptide: Stable helical γ-tetra-, hexa-, and and octapeptides and consequences of α-substitution. J. Am. Chem. Soc. 1998, 120, 8569–8570. [Google Scholar] [CrossRef]
- Ferrrera-Sinfreu, J.; Zaccaro, L.; Vidal, D.; Salvatella, X.; Giralt, E.; Pons, M.; Albericio, F.; Royo, M. A new class of foldamers based on cis-γ-amino-L-proline. J. Am. Chem. Soc. 2004, 126, 6048–6057. [Google Scholar] [CrossRef]
- Ferrrera-Sinfreu, J.; Giralt, E.; Castel, S.; Albericio, F.; Royo, M. Cell-penetrating cis-γ-amino-L-proline-derived peptides. J. Am. Chem. Soc. 2005, 127, 9459–9468. [Google Scholar] [CrossRef]
- Baldauf, C.; Günther, R.; Hofman, H.-J. Control of helix formation in vinylogous γ-peptide by (E)- and (Z)-double bonds: A way to ion channels and monomolecular nanotubes. J. Org. Chem. 2005, 70, 5351–5361. [Google Scholar] [CrossRef]
- Legrand, B.; Maillard, L.T. α,β-unsaturated γ-peptide foldamers. ChemPlusChem 2021, 86, 629–645. [Google Scholar] [CrossRef] [PubMed]
- Fisher, B.F.; Gellman, S.H. Impact of γ-amino acid residue preorganization on α/γ-peptide foldamer helicity in aqueous solution. J. Am. Chem. Soc. 2016, 138, 10766–10769. [Google Scholar] [CrossRef]
- Bouillère, F.; Thétiot-Laurent, S.; Kouklovsky, C.; Alezra, V. Foldamers containing γ-amino acid residues or their analogues: Structural features and application. Amino Acids 2011, 41, 687–707. [Google Scholar] [CrossRef]
- Bandyopadhyay, A.; Jadhav, S.V.; Gopi, H.N. α/γ4-Hybrid peptide helices: Synthesis, conformations and analogy with the α-helix. Chem. Commun. 2012, 57, 7170–7172. [Google Scholar] [CrossRef]
- Sawada, T.; Gellman, S.H. Structural mimicry of the α-helix in aqueous solution with an isoatomic α,β,γ-backbone. J. Am. Chem. Soc. 2011, 133, 7336–7339. [Google Scholar] [CrossRef]
- Grison, C.M.; Miles, J.A.; Robin, S.; Wilson, A.J.; Aitken, D.J. An α-helix mimicking 12,13-helix: Designed α/β/γ-foldamers as selective inhibitors of protein-protein interactions. Angew. Chem. Int. Ed. 2016, 55, 11096–11100. [Google Scholar] [CrossRef] [PubMed]
- Grison, C.M.; Robin, S.; Aitken, D.J. 13-Helix folding of α,β,γ-peptide manifold designed from a minimal-constraint blueprint. Chem. Commun. 2016, 52, 7802–7805. [Google Scholar] [CrossRef] [PubMed]
- She, F.; Teng, P.; Peguero-Tejada, A.; Wang, M.; Ma, N.; Odom, T.; Zhou, M.; Gjonaj, E.; Wojtas, L.; Vaart, A.V.D.; et al. De novo left-handed synthetic peptidemimetic foldamers. Angew. Chem. Int. Ed. 2018, 57, 9916–9920. [Google Scholar] [CrossRef] [PubMed]
- Sang, P.; Shi, Y.; Xue, S.; Odom, T.; Cai, J. Sulfono-γ-AApeptides as helical mimetics: Crystal structures and applications. Acc. Chem. Res. 2020, 53, 2425–2442. [Google Scholar] [CrossRef]
- Sang, P.; Zhang, M.; Shi, Y.; Li, C.; Abdulladir, S.; Li, Q.; Ji, H.; Cai, J. Inhibition of β-catenine/B cell lymphoma 9 protein-protein interaction using α-helix mimicking sulfono-γ-AApeptide inhibitors. Proc. Natl. Acad. Sci. USA 2019, 116, 10757–10762. [Google Scholar] [CrossRef]
- Sang, P.; Shi, Y.; Lu, J.; Chen, L.; Yang, L.; Borcherds, W.; Abdulladir, S.; Li, Q.; Daughdrill, G.; Chen, J.; et al. α-Helix-mimicking sulfono-γ-AApeptide inhibitors for p53-MDM2/MDMX protein-protein interactions. J. Med. Chem. 2020, 63, 975–986. [Google Scholar] [CrossRef]
- Mathieu, L.; Legrand, B.; Deng, C.; Vezenkov, L.; Wenger, E.; Didierjean, C.; Amblard, M.; Averlant-Petit, M.C.; Masurier, N.; Lisowski, V.; et al. Helical foldamers of thiazole-based γ-amino acids: Synthesis and structural studies. Angew. Chem. Int. Ed. 2013, 52, 6006–6610. [Google Scholar] [CrossRef]
- Kaffy, J.; Berardet, C.; Mathieu, L.; Legrand, B.; Taverna, M.; Halgand, F.; Van Der Rest, G.; Maillard, L.T.; Ongeri, S. Helical γ-peptide foldamers as dual inhibitors of amyloid-β peptide and islet amyloid polypeptide oligomerization and fibrilization. Chemistry 2020, 26, 14612–14622. [Google Scholar] [CrossRef]
- Mándity, I.M.; Fülöp, F. An overview of peptide and peptoid foldamers in medicinal chemistry. Expert Opin. Drug Discov. 2015, 10, 1163–1177. [Google Scholar] [CrossRef]
- Kirshenbaum, K.; Barron, A.E.; Goldsmit, R.A.; Armand, P.; Bradley, E.K.; Truong, K.T.; Dill, K.A.; Cohen, F.E.; Zuckermann, R.N. Sequence-specific polypeptoids: A diverse family of heteropolymers with stable secondary structure. Proc. Natl. Acad. Sci. USA 1998, 95, 4303–4308. [Google Scholar] [CrossRef]
- Gorske, B.C.; Bastian, B.L.; Geske, G.D.; Blackwell, H.E. Local and tunable n→π* interactions regulate amide isomerism in the peptoid backbone. J. Am. Chem. Soc. 2007, 129, 8928–8929. [Google Scholar] [CrossRef] [PubMed]
- Szekely, T.; Roy, O.; Dériaud, E.; Job, A.; Lo-Man, R.; Leclerc, C.; Taillefumier, C. Design, Synthesis, and Immunological Evaluation of a Multicomponent Construct Based on a Glycotripeptoid Core Comprising B and T Cell Epitopes and a Toll-like Receptor 7 Agonist That Elicits Potent Immune Responses. J. Med. Chem. 2018, 61, 9568–9582. [Google Scholar] [CrossRef] [PubMed]
- Shyam, R.; Charbonnel, N.; Job, A.; Blavignac, C.; Forestier, C.; Taillefumier, C.; Faure, S. 1,2,3-triazolium-based cationic amphipathic peptoid oligomers mimicking antimicrobial peptides. ChemMedChem 2018, 13, 1513–1516. [Google Scholar] [CrossRef] [PubMed]
- Morimoto, J.; Fukuda, Y.; Kuroda, D.; Watanabe, T.; Yoshida, F.; Asada, M.; Nakamura, T.; Senoo, A.; Nagatoishi, S.; Tsumoto, K.; et al. A peptoid with extended shape in water. J. Am. Chem. Soc. 2019, 141, 14612–14623. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Kodadek, T. Synthesis and screening of stereochemically diverse combinatorial libraries of peptide tertiary amides. Chem. Biol. 2013, 20, 360–369. [Google Scholar] [CrossRef]
- Udugamasooriya, D.G.; Dineen, S.P.; Brekken, R.A.; Kodadek, R. A peptoid antibody surrogate that antagonizes VEGF receptor 2 activity. J. Am. Chem. Soc. 2008, 130, 5744–5752. [Google Scholar] [CrossRef]
- Udugamasooriya, D.G.; Dunham, G.; Ritchie, C.; Brekken, R.A.; Kodadek, T. The pharmacophore of a peptoid VEGF receptor 2 antagonist includes both side chain and main chain residues. Bioorg. Med. Chem. Lett. 2008, 18, 5892–5894. [Google Scholar] [CrossRef]
- Udugamasooriya, D.G.; Ritchie, C.; Brekken, R.A.; Kodadek, T. A peptoid antagonist of VEGF receptor 2 recognizes a ‘hotspot’ in the extracellular domain distinct from the hormone-binding site. Bioorg. Med. Chem. 2008, 16, 6338–6343. [Google Scholar] [CrossRef][Green Version]
- Lynn, K.D.; Udugamasooriya, D.G.; Roland, C.L.; Castrillon, D.H.; Kodadek, T.; Brekken, R.A. GU81, a VEGFR2 antagonist peptoid, enhances the anti-tumor activity of doxorubicin in the murine MMTV-PyMT transgenic model of breast cancer. BMC Cancer 2010, 10, 397. [Google Scholar] [CrossRef]
- Schneider, J.A.; Craven, T.W.; Kasper, A.C.; Yun, C.; Haugbro, M.; Briggs, E.M.; Svetlov, V.; Nudler, E.; Knaut, H.; Bonneau, R.; et al. Design of peptoid-peptide macrocycles to inhibit the β-catenin TCF interaction in prostate cancer. Nat. Commun. 2018, 9, 4396. [Google Scholar] [CrossRef]
- Trader, D.J.; Simanski, S.; Kodadek, T. A reversible and highly selective inhibitor of the proteasomal ubiquitin receptor Rpn13 is toxic to multiple myeloma cells. J. Am. Chem. Soc. 2015, 137, 6312–6319. [Google Scholar] [CrossRef] [PubMed]
- Oh, M.; Lee, J.H.; Moon, H.; Hyun, Y.J.; Lim, H.S. A chemical inhibitor of the Skp2/p300 interaction that promotes p53-mediated apoptosis. Angew. Chem. Int. Ed. 2016, 55, 602–606. [Google Scholar] [CrossRef] [PubMed]
- Fukuda, Y.; Yokomine, M.; Kuroda, D.; Tsumoto, K.; Morimoto, J.; Sando, S. Peptoid-based reprogrammable template for cell-permeable inhibitors of protein–protein interactions. Chem. Sci. 2021, 12, 13292–13300. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, J.T.; Turck, C.W.; Cohen, F.E.; Zuckermann, R.N.; Lim, W.A. Exploiting the basis of proline recognition by SH3 and WW domains: Design of N-substituted inhibitors. Science 1998, 282, 2088–2092. [Google Scholar] [CrossRef]
- Demmer, O.; Frank, A.O.; Hagn, F.; Schottelius, M.; Marinelli, L.; Cosconati, S.; Brack-Werner, R.; Kremb, S.; Wester, H.-J.; Kessler, H. A conformationally frozen peptoid boosts CXCR4 affinity and anti-HIV activity. Angew. Chem. Int. Ed. 2012, 51, 8110–8113. [Google Scholar] [CrossRef]
- Yoo, S.H.; Li, B.; Dolain, C.; Pasco, M.; Guichard, G. Urea Based Foldamers. Methods Enzymol. 2021, 656, 59–92. [Google Scholar]
- Pasco, M.; Dolain, C.; Guichard, G. Foldamers in Medicinal Chemistry. Compr. Supramol. Chem. II 2017, 89–125. [Google Scholar]
- Burgess, K.; Shin, H.; Linthicum, D.S. Solid-Phase Syntheses of Unnatural Biopolymers Containing Repeating Urea Units. Angew. Chem. Int. Ed. Engl. 1995, 34, 907–909. [Google Scholar] [CrossRef]
- Burgess, K.; Ibarzo, J.; Linthicum, D.S.; Russell, D.H.; Shin, H.; Shitangkoon, A.; Totani, R.; Zhang, A.J. Solid Phase Syntheses of Oligoureas. J. Am. Chem. Soc. 1997, 119, 1556–1564. [Google Scholar] [CrossRef]
- Guichard, G.; Semetey, V.; Rodriguez, M.; Briand, J.-P. Solid Phase Synthesis of Oligoureas Using O-Succinimidyl-(9H-Fluoren-9-ylmethoxycarbonylamino)Ethylcarbamate Derivatives as Activated Monomers. Tetrahedron Lett. 2000, 41, 1553–1557. [Google Scholar] [CrossRef]
- Violette, A.; Averlant-Petit, M.C.; Semetey, V.; Hemmerlin, C.; Casimir, R.; Graft, R.; Marraud, M.; Briand, J.P.; Rognan, D.; Guichard, G. N,N′-Linked Oligoureas as Foldamers: Chain Length Requirements for Helix Formation in Protic Solvent Investigated by Circular Dichroism, NMR Spectroscopy, and Molecular Dynamics. J. Am. Chem. Soc. 2005, 127, 2156–2164. [Google Scholar] [CrossRef]
- Fischer, L.; Guichard, G. Folding and Self-Assembly of Aromatic and Aliphatic Urea Oligomers: Towards Connecting Structure and Function. Org. Biomol. Chem. 2010, 8, 3102–3117. [Google Scholar] [CrossRef] [PubMed]
- Fremaux, J.; Venin, C.; Mauran, L.; Zimmer, R.H.; Guichard, G.; Goudreau, S.R. Peptide-Oligourea Hybrids Analogue of GLP-1 with Improved Action in Vivo. Nat. Commun. 2019, 10, 924. [Google Scholar] [CrossRef] [PubMed]
- Cussol, L.; Mauran-Ambrosino, L.; Buratto, J.; Belorusova, A.Y.; Neuville, M.; Osz, J.; Fribourg, S.; Fremaux, J.; Dolain, C.; Goudreau, S.R.; et al. Structural Basis for α-Helix Mimicry and Inhibition of Protein–Protein Interactions with Oligourea Foldamers. Angew. Chem. Int. Ed. 2021, 60, 2296–2303. [Google Scholar] [CrossRef] [PubMed]
- Hoe, K.K.; Verma, C.S.; Lane, D.P. Drugging the P53 Pathway: Understanding the Route to Clinical Efficacy. Nat. Rev. Drug Discov. 2014, 13, 217–236. [Google Scholar]
- Roodman, G.D.; Windle, J.J. Paget Disease of Bone. J. Clin. Investig. 2005, 115, 200–208. [Google Scholar] [CrossRef]
- Mbianda, J.; Bakail, M.; André, C.; Moal, G.; Perrin, M.E.; Pinna, G.; Guerois, R.; Becher, F.; Legrand, P.; Traoré, S.; et al. Optimal Anchoring of a Foldamer Inhibitor of ASF1 Histone Chaperone through Backbone Plasticity. Sci. Adv. 2021, 7, eabd9153. [Google Scholar] [CrossRef]
- Im, J.S.; Keaton, M.; Lee, K.Y.; Kumar, P.; Park, J.; Dutta, A. ATR Checkpoint Kinase and CRL1βTRCP Collaborate to Degrade ASF1a and Thus Repress Genes Overlapping with Clusters of Stalled Replication Forks. Genes Dev. 2014, 28, 875–887. [Google Scholar] [CrossRef]
- Huc, I. Aromatic Oligoamide Foldamers. Eur. J. Org. Chem. 2004, 1, 17–29. [Google Scholar] [CrossRef]
- Hamuro, Y.; Geib, S.J.; Hamilton, A.D. Novel Molecular Scaffolds: Formation of Helical Secondary Structure in a Family of Oligoanthranilamides. Angew. Chem. Int. Ed. 1994, 33, 446–448. [Google Scholar] [CrossRef]
- Hamuro, Y.; Geib, S.J.; Hamilton, A.D. Oligoanthranilamides. Non-Peptide Subunits That Show Formation of Specific Secondary Structure. J. Am. Chem. Soc. 1996, 118, 7529–7541. [Google Scholar] [CrossRef]
- Jiang, H.; Léger, J.M.; Dolain, C.; Guionneau, P.; Huc, I. Aromatic δ-Peptides: Design, Synthesis and Structural Studies of Helical, Quinoline-Derived Oligoamide Foldamers. Tetrahedron 2003, 59, 8365–8374. [Google Scholar] [CrossRef]
- Jiang, H.; Léger, J.M.; Huc, I. Aromatic δ-Peptides. J. Am. Chem. Soc. 2003, 125, 3448–3449. [Google Scholar] [CrossRef] [PubMed]
- Kleman, A.F.; Dufek, D.L.; Fobe, T.L.; McCaslin, D.R.; Cary, B.P.; Shirts, M.R.; Gellman, S.H. Potential Foldamers Based on an Ortho-Terphenyl Amino Acid. Org. Lett. 2021, 23, 4855–4859. [Google Scholar] [CrossRef]
- Vallade, M.; Jewginski, M.; Fischer, L.; Buratto, J.; Bathany, K.; Schmitter, J.M.; Stupfel, M.; Godde, F.; Mackereth, C.D.; Huc, I. Assessing Interactions between Helical Aromatic Oligoamide Foldamers and Protein Surfaces: A Tethering Approach. Bioconjug. Chem. 2019, 30, 54–62. [Google Scholar] [CrossRef]
- Ziach, K.; Chollet, C.; Parissi, V.; Prabhakaran, P.; Marchivie, M.; Corvaglia, V.; Bose, P.P.; Laxmi-Reddy, K.; Godde, F.; Schmitter, J.M.; et al. Single Helically Folded Aromatic Oligoamides That Mimic the Charge Surface of Double-Stranded B-DNA. Nat. Chem. 2018, 10, 511–518. [Google Scholar] [CrossRef]
- Corvaglia, V.; Amar, I.A.M.; Garambois, V.; Letast, S.; Garcin, A.; Gongora, C.; Del Rio, M.; Denevault-Sabourin, C.; Joubert, N.; Huc, I.; et al. Internalization of Foldamer-Based DNA Mimics through a Site-Specific Antibody Conjugate to Target Her2-Positive Cancer Cells. Pharmaceuticals 2021, 14, 624. [Google Scholar] [CrossRef]
- Marqusee, S.; Baldwin, R.L. Helix stabilization by Glu-…Lys+ salt bridges in short peptides of de novo design. Proc. Natl. Acad. Sci. USA 1987, 84, 8898–8902. [Google Scholar] [CrossRef]
- Ruan, F.; Chen, Y.; Hopkins, P.B. Metal Ion Enhanced Helicity in Synthetic Peptides Containing Unnatural, Metal-Ligating Residues. J. Am. Chem. Soc. 1990, 112, 9403–9404. [Google Scholar] [CrossRef]
- Ghadiri, M.R.; Fernholz, A.K. Peptide Architecture. Design of Stable α-Helical Metallopeptides via a Novel Exchange-Inert RuIII Complex. J. Am. Chem. Soc. 1990, 112, 9633–9635. [Google Scholar] [CrossRef]
- Smith, S.J.; Du, K.; Radford, R.J.; Tezcan, F.A. Functional, metal-based crosslinkers for α-helix induction in short peptides. Chem. Sci. 2013, 4, 3740–3747. [Google Scholar] [CrossRef] [PubMed]
- Leduc, A.M.; Trent, J.O.; Wittliff, J.L.; Bramlett, K.S.; Briggs, S.L.; Chirgadze, N.Y.; Wang, Y.; Burris, T.P.; Spatola, A.F. Helix-stabilized cyclic peptides as selective inhibitors of steroid receptor–coactivator interactions. Proc. Natl. Acad. Sci. USA 2003, 100, 11273–11278. [Google Scholar] [CrossRef] [PubMed]
- Galande, A.K.; Bramlett, K.S.; Trent, J.O.; Burris, T.P.; Wittliff, J.L.; Spatola, A.F. Potent Inhibitors of LXXLL-Based Protein– Protein Interactions. ChemBioChem 2005, 6, 1991–1998. [Google Scholar] [CrossRef]
- Nagakubo, T.; Demizu, Y.; Kanda, Y.; Misawa, T.; Shoda, T.; Okuhira, K.; Sekino, Y.; Naito, M.; Kurihara, M. Development of Cell-Penetrating R7 Fragment-Conjugated Helical Peptides as Inhibitors of Estrogen Receptor-Mediated Transcription. Bioconjug. Chem. 2014, 25, 1921–1924. [Google Scholar] [CrossRef] [PubMed]
- Dougherty, P.G.; Wen, J.; Pan, X.; Koley, A.; Ren, J.G.; Sahni, A.; Basu, R.; Salim, H.; Kubi, G.A.; Qian, Z.; et al. Enhancing the Cell Permeability of Stapled Peptides with a Cyclic Cell-Penetrating Peptide. J. Med. Chem. 2019, 62, 10098–10107. [Google Scholar] [CrossRef] [PubMed]
- Walensky, L.D.; Kung, A.L.; Escher, I.; Malia, T.J.; Barbuto, S.; Wright, R.D.; Wagner, G.; Verdine, G.L.; Korsmeyer, S.J. Activation of Apoptosis in Vivo by a Hydrocarbon-Stapled BH3 Helix. Science 2004, 305, 1466–1470. [Google Scholar] [CrossRef]
- Blackwell, H.E.; Grubbs, R.H. Highly Efficient Synthesis of Covalently Cross-Linked Peptide Helices by Ring-Closing Metathesis. Angew. Chem. Int. Ed. 1998, 37, 3281–3284. [Google Scholar] [CrossRef]
- Schafmeister, C.E.; Po, J.; Verdine, G.L. An All-Hydrocarbon Cross-Linking System for Enhancing the Helicity and Metabolic Stability of Peptides. J. Am. Chem. Soc. 2000, 122, 5891–5892. [Google Scholar] [CrossRef]
- Verdine, G.L.; Hilinski, G.J. All-hydrocarbon stapled peptides as Synthetic Cell-Accessible Mini-Proteins. Drug Discov. Today 2012, 9, e41–e47. [Google Scholar] [CrossRef]
- Tom, N.; Grossmann, T.N.; Yeha, J.T.-H.; Bowmana, B.R.; Chu, Q.; Moellering, R.E.; Verdine, G.L. Inhibition of oncogenic Wnt signaling through direct targeting of β-catenin. Proc. Natl. Acad. Sci. USA 2012, 109, 17942–17947. [Google Scholar]
- Chang, Y.S.; Graves, B.; Guerlavais, V.; Tovar, C.; Packman, K.; To, K.H.; Olson, K.A.; Kesavan, K.; Gangurde, P.; Mukherjee, A.; et al. Stapled α−helical peptide drug development: A potent dual inhibitor of MDM2 and MDMX for p53-dependent cancer therapy. Proc. Natl. Acad. Sci. USA 2013, 110, E3445–E3454. [Google Scholar] [CrossRef]
- Saleh, M.N.; Patel, M.R.; Bauer, T.M.; Goel, S.; Falchook, G.S.; Shapiro, G.I.; Chung, K.Y.; Infante, J.R.; Conry, R.M.; Rabinowits, G.; et al. Phase 1 Trial of ALRN-6924, a Dual Inhibitor of MDMX and MDM2, in Patients with Solid Tumors and Lymphomas Bearing Wild-Type TP53. Clin. Cancer Res. 2021, 27, 5236–5247. [Google Scholar] [CrossRef]
- Suzuki, K.; Takaoka, Y.; Ueda, M. Rational design of a stapled JAZ9 peptide inhibiting protein-protein interaction of a plant transcription factor. RSC Chem. Biol. 2021, 2, 499–502. [Google Scholar] [CrossRef] [PubMed]
- Hilinski, G.J.; Kim, Y.W.; Hong, J.; Kutchukian, P.S.; Crenshaw, C.M.; Berkovitch, S.S.; Chang, A.; Ham, S.; Verdine, G.L. Stitched α-helical peptides via bis ring-closing metathesis. J. Am. Chem. Soc. 2014, 136, 12314–12322. [Google Scholar] [CrossRef] [PubMed]
- Kannan, S.; Aronica, P.G.A.; Ng, S.; Lian, D.T.G.; Frosi, Y.; Chee, S.; Shimin, J.; Yuen, T.Y.; Sadruddin, A.; Kaan, H.Y.K.; et al. Macrocyclization of an all-D linear α-helical peptide imparts cellular permeability. Chem. Sci. 2020, 11, 5577–5591. [Google Scholar] [CrossRef]
- Flint, D.G.; Kumita, J.R.; Smart, O.S.; Woolley, G.A. Using an azobenzene cross-linker to either increase or decrease peptide helix content upon trans-to-cis photoisomerization. Chem. Biol. 2002, 9, 391–397. [Google Scholar] [CrossRef]
- Jankovic, B.; Gulzar, A.; Zanobini, C.; Bozovic, O.; Wolf, S.; Stock, G.; Hamm, P. Photocontrolling Protein-Peptide Interactions: From Minimal Perturbation to Complete Unbinding. J. Am. Chem. Soc. 2019, 141, 10702–10710. [Google Scholar] [CrossRef]
- Spokoyny, A.M.; Zou, Y.; Ling, J.J.; Yu, H.; Lin, Y.S.; Pentelute, B.L. A perfluoroaryl-cysteine SNAr chemistry approach to unprotected peptide stapling. J. Am. Chem. Soc. 2013, 35, 5946–5949. [Google Scholar] [CrossRef]
- Vasco, A.V.; Méndez, Y.; Porzel, A.; Balbach, J.; Wessjohann, L.A.; Rivera, D.G. A Multicomponent Stapling Approach to Exocyclic Functionalized Helical Peptides: Adding Lipids, Sugars, PEGs, Labels, and Handles to the Lactam Bridge. Bioconjug. Chem. 2019, 30, 253–259. [Google Scholar] [CrossRef]
- Ricardo, M.G.; Ali, A.M.; Plewka, J.; Surmiak, E.; Labuzek, B.; Neochoritis, C.G.; Atmaj, J.; Skalniak, L.; Zhang, R.; Holak, T.A.; et al. Multicomponent Peptide Stapling as a Diversity-Driven Tool for the Development of Inhibitors of Protein-Protein Interactions. Angew. Chem. Int. Ed. 2020, 59, 5235–5241. [Google Scholar] [CrossRef] [PubMed]
- Patgiri, A.; Jochim, A.L.; Arora, P.S. A hydrogen bond surrogate approach for stabilization of short peptide sequences in alpha-helical conformation. Acc. Chem. Res. 2008, 41, 1289–1300. [Google Scholar] [CrossRef] [PubMed]
- Douse, C.H.; Maas, S.J.; Thomas, J.C.; Garnett, J.A.; Sun, Y.; Cota, E.; Tate, E.W. Crystal structures of stapled and hydrogen bond surrogate peptides targeting a fully buried protein-helix interaction. ACS Chem. Biol. 2014, 9, 2204–2209. [Google Scholar] [CrossRef] [PubMed]
- Burslem, G.M.; Crews, C.M. Proteolysis-Targeting Chimeras as Therapeutics and Tools for Biological Discovery. Cell 2020, 181, 102–114. [Google Scholar] [CrossRef]
- Békés, M.; Langley, D.R.; Crews, C.M. PROTAC targeted protein degraders: The past is prologue. Nat. Rev. Drug Discov. 2022, 21, 181–200. [Google Scholar] [CrossRef]
- Liao, H.; Li, X.; Zhao, L.; Wang, Y.; Wang, X.; Wu, Y.; Zhou, X.; Fu, W.; Liu, L.; Hu, H.G.; et al. A PROTAC peptide induces durable β-catenin degradation and suppresses Wnt-dependent intestinal cancer. Cell Discov. 2020, 6, 35. [Google Scholar] [CrossRef]
- Moellering, R.E.; Cornejo, M.; Davis, T.N.; Del Bianco, C.; Aster, J.C.; Blacklow, S.C.; Kung, A.L.; Gilliland, D.G.; Verdine, G.L.; Bradner, J.E. Direct inhibition of the NOTCH transcription factor complex. Nature 2009, 462, 182–188. [Google Scholar] [CrossRef]
- Ohoka, N.; Misawa, T.; Kurihara, M.; Demizu, Y.; Naito, M. Development of a peptide-based inducer of protein degradation targeting NOTCH1. Bioorg. Med. Chem. Lett. 2017, 27, 4985–4988. [Google Scholar] [CrossRef]
- Yokoo, H.; Ohoka, N.; Takyo, M.; Ito, T.; Tsuchiya, K.; Kurohara, T.; Fukuhara, K.; Inoue, T.; Naito, M.; Demizu, Y. Peptide Stapling Improves the Sustainability of a Peptide-Based Chimeric Molecule That Induces Targeted Protein Degradation. Int. J. Mol. Sci. 2021, 22, 8772. [Google Scholar] [CrossRef]


| Peptide Sequence | Target Protein | Ref. |
|---|---|---|
ETFUDUWKULUE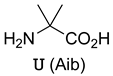 | hDM2 | [54] |
dtwUelvedlnGGGSBR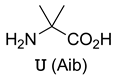  Lowercase letters: D-amino acids | MDM2 | [55] |
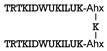 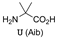  | S100B | [56] |
 | VDR | [57] |
 | VDR | [58] |
| Peptide Sequence and Structure | Target PPI | Ref. |
|---|---|---|
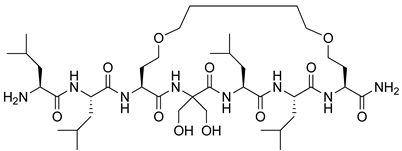
H-β3O-β3V-β3W-β3E-β3V-β3W-β3O-β3V-β3I-β3E-OH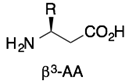 | HIV gp41 | [77] |
| H-β3K-β3V-β3L-β3E-β3V-β3W-β3K-β3V-β3F-β3E-OH | p53-hDM2 | [78] |
Ac-(β3R)-TWE-(β3E)-WD-(β3R)-AIA-(β3E)-YA-(β3R)-RIE-(β3E)-LIZAAQ-(β3E)-QQZKNE-(β3E)-ALZEL-NH2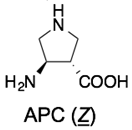 | HIV gp41 | [79] |
H-V-(β3D)-NK-(β3F)-NKEXCNZRAIEUALDPNLNDUQFHUKIWZIKXDC-NH2 | VEGF | [80] |
| Peptide Sequence and Structure | Target PPI | Ref. |
|---|---|---|
Boc-F-(γ4A)-(tACBC)-(γ4W)-(tACBC)-(β3L)-OMe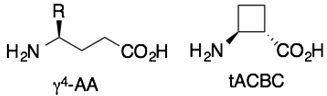 | p53-hDM2 | [91] |
Ac-γ4E(EtNH2)-γ4R(Me)-γ4R(Me)-γ4E(Me)-γ4L(EtNH2)-γ4T(iBu)-γ4L(EtNH2)-γ4R(iBu)-γ4A(Me)-γ4L(iBu)-γ4L(tol)-NH2 | β-catenin/BCL9 | [95] |
| Ac-γ4L(p-ClPh)-γ4F(Me)-γ4E(Me)-γ4W(Me)-γ4K(Me)-γ4cBA(iBu)-γ4A(Me)-NH2 | P53-MDM2/MDMX | [96] |
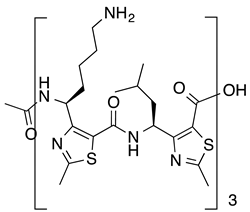 | Amyloid-β | [98] |
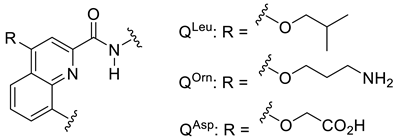
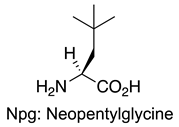
| Sequence or Structure | Target Protein | Ref. |
|---|---|---|
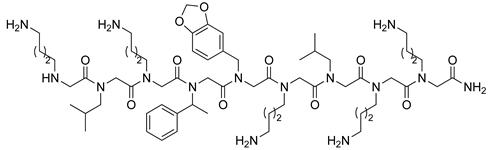 | VEGFR2 | [106] |
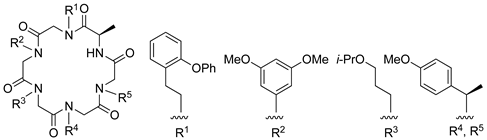 | β-catenin | [110] |
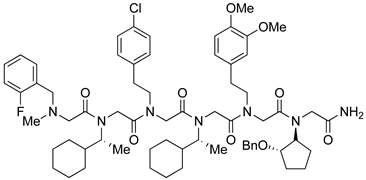 | Rpn13 | [111] |
 | Skp2 | [112] |
 | MDM2 | [113] |
YEVPPPVXPRRR | Grb2 SH3 | [114] |
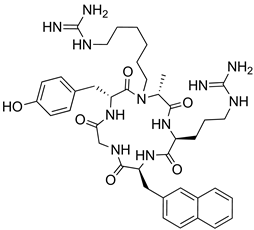 | CXCR4 | [115] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tsuchiya, K.; Kurohara, T.; Fukuhara, K.; Misawa, T.; Demizu, Y. Helical Foldamers and Stapled Peptides as New Modalities in Drug Discovery: Modulators of Protein-Protein Interactions. Processes 2022, 10, 924. https://doi.org/10.3390/pr10050924
Tsuchiya K, Kurohara T, Fukuhara K, Misawa T, Demizu Y. Helical Foldamers and Stapled Peptides as New Modalities in Drug Discovery: Modulators of Protein-Protein Interactions. Processes. 2022; 10(5):924. https://doi.org/10.3390/pr10050924
Chicago/Turabian StyleTsuchiya, Keisuke, Takashi Kurohara, Kiyoshi Fukuhara, Takashi Misawa, and Yosuke Demizu. 2022. "Helical Foldamers and Stapled Peptides as New Modalities in Drug Discovery: Modulators of Protein-Protein Interactions" Processes 10, no. 5: 924. https://doi.org/10.3390/pr10050924
APA StyleTsuchiya, K., Kurohara, T., Fukuhara, K., Misawa, T., & Demizu, Y. (2022). Helical Foldamers and Stapled Peptides as New Modalities in Drug Discovery: Modulators of Protein-Protein Interactions. Processes, 10(5), 924. https://doi.org/10.3390/pr10050924