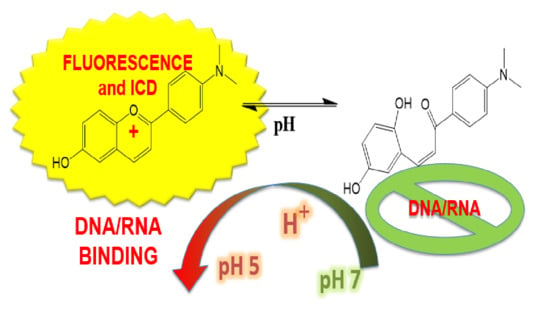
Flavylium Dye as pH-Tunable Fluorescent and CD Probe for Double-Stranded DNA and RNA
Abstract
1. Introduction
2. Materials and Methods
3. Results
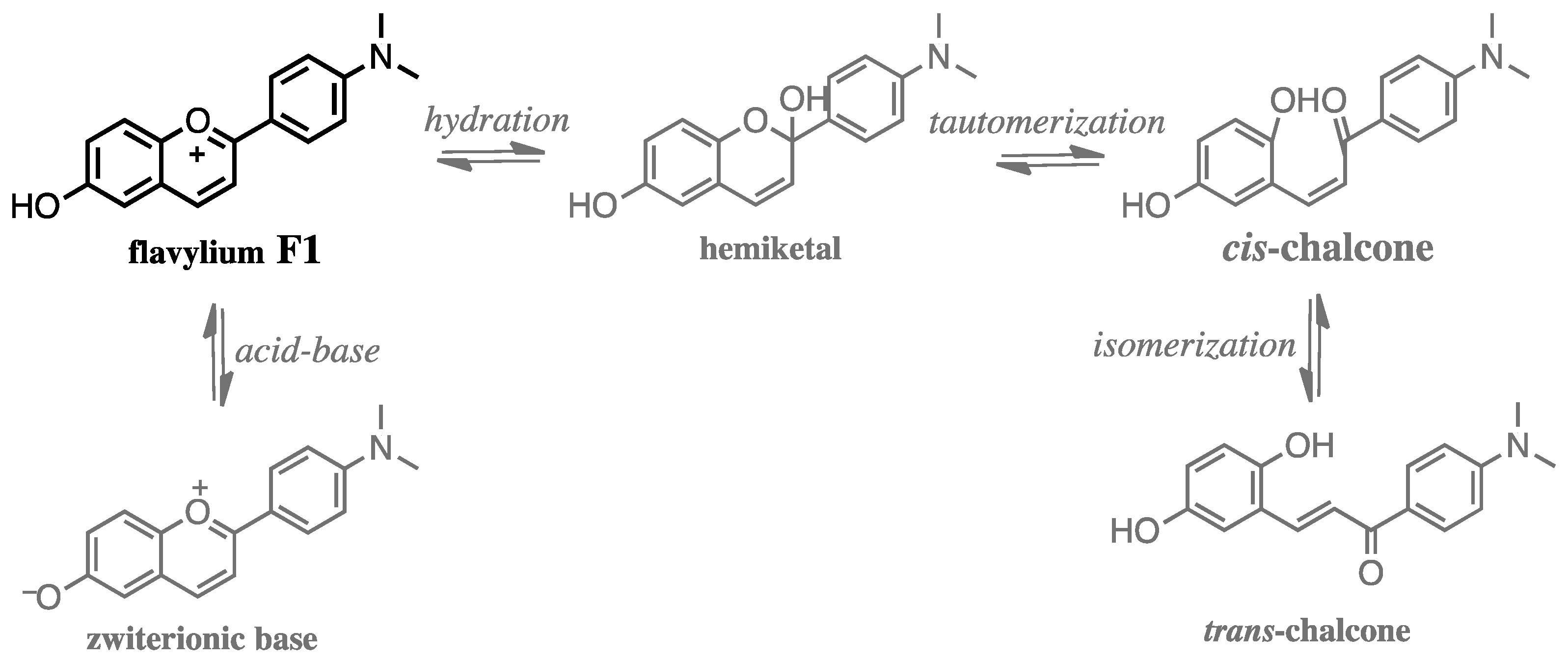
3.1. Physicochemical Properties of Aqueous Solutions of Studied Compound F1
3.2. Study of Interactions of F1 with ds-DNA and ds-RNA in Aqueous Media
3.2.1. UV/Vis Titrations
3.2.2. Fluorimetric Titrations
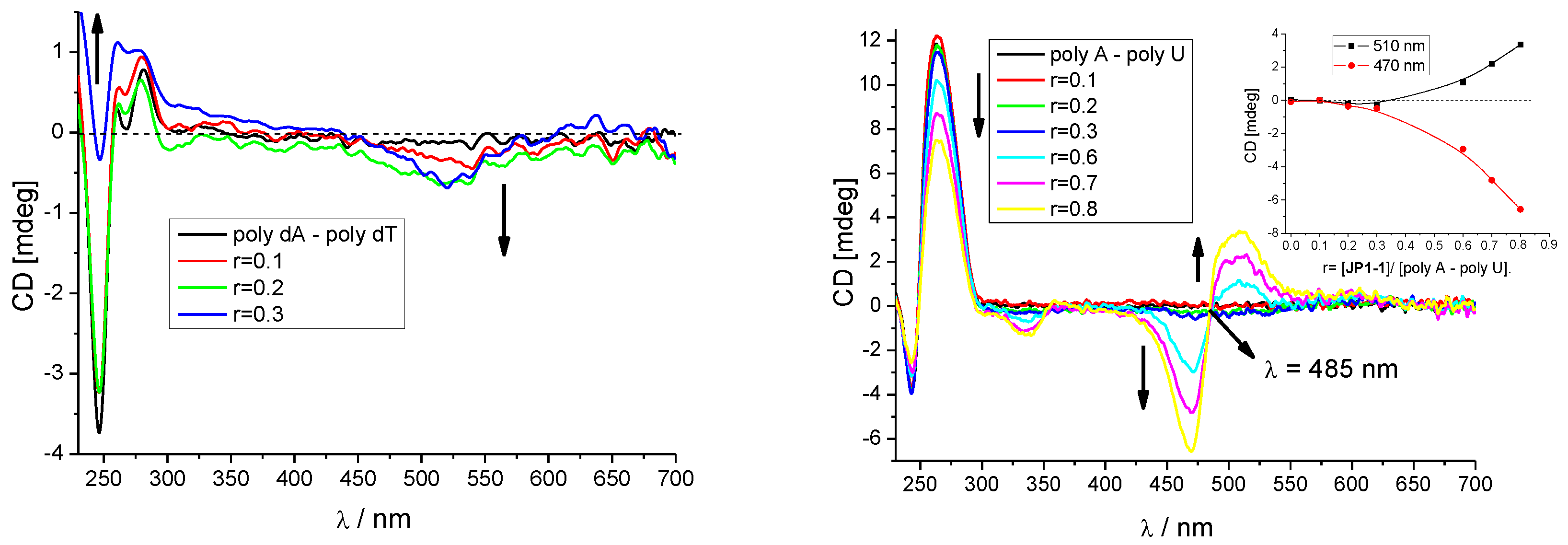
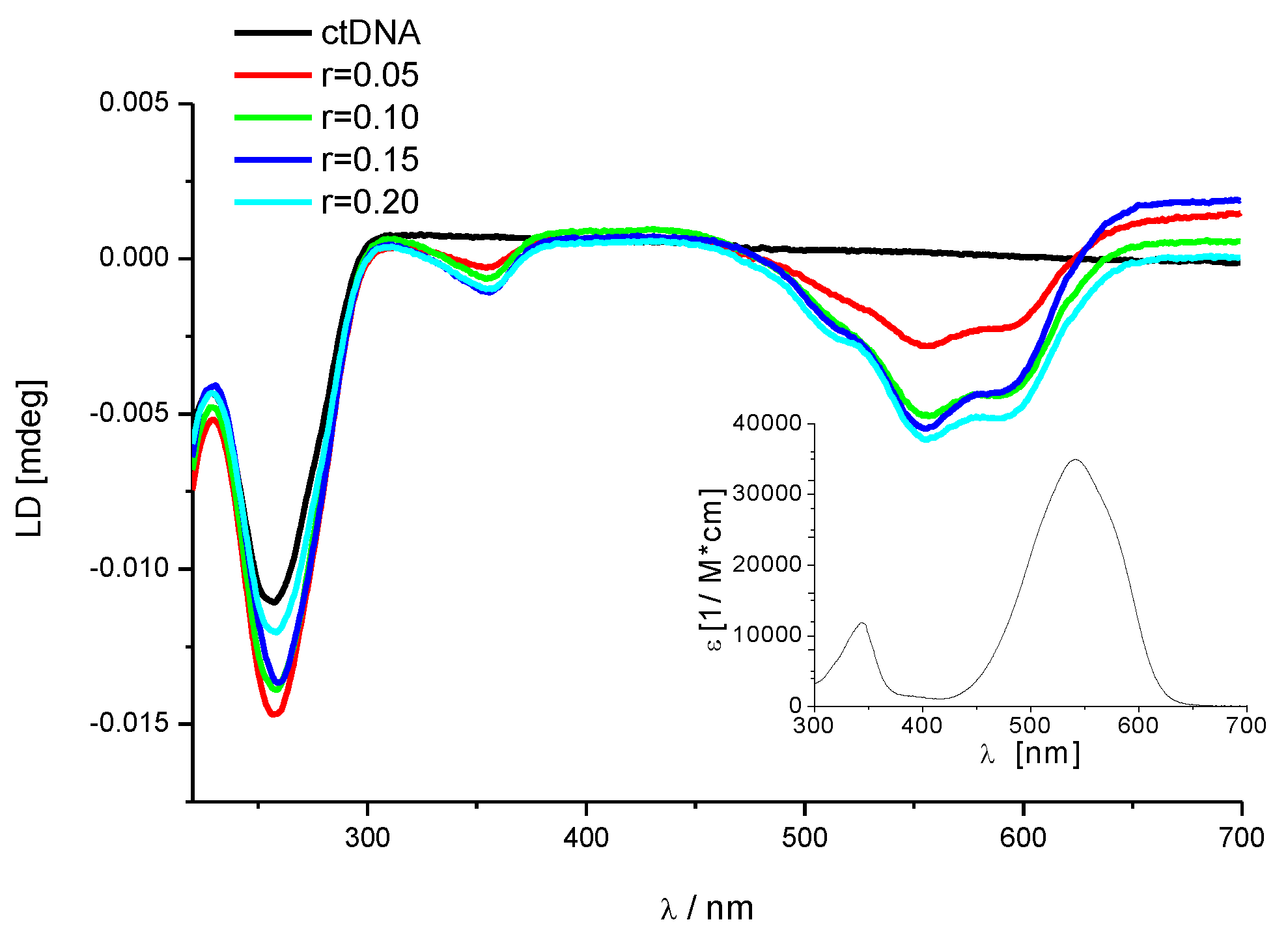
3.2.3. Circular Dichroism (CD) Titrations
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Melo, M.J. Missal Blue: Anthocyanins in Nature and Art. In Dyes in History and Archaeology; Kirby, J., Ed.; Archetype Publications: London, UK, 2008; Volume 21, pp. 65–74. [Google Scholar]
- Pina, F. Chemical applications of anthocyanins and related compounds A source of bio-inspiration. J. Agric. Food Chem. 2014, 62, 6885–6897. [Google Scholar] [CrossRef] [PubMed]
- Pina, F.; Parola, A.J.; Melo, M.J.; Lima, J.C.; De Freitas, V. Chemistry of Anthocyanins. In Anthocyanins from Natural Sources: Exploiting Targeted Delivery for Improved Health; Brook, M.S.-L., Celli, G.B., Eds.; The Royal Society of Chemistry: Cambridge, UK, 2019; Volume 12, Chapter 2; pp. 34–76. [Google Scholar]
- Cheynier, V.; Tomas-Barberan, F.A.; Yoshida, K. Polyphenols, From Plants to a Variety of Food and Nonfood Uses. J. Agric. Food Chem. 2015, 63, 7589–7594. [Google Scholar] [CrossRef] [PubMed]
- Dangles, O.; Fenger, J.-A. The Chemical Reactivity of Anthocyanins and Its Consequences in Food Science and Nutrition. Molecules 2018, 23, 1970. [Google Scholar] [CrossRef] [PubMed]
- Basílio, N.; Pina, F. Chemistry and Photochemistry of Anthocyanins and Related Compounds: A Thermodynamic and Kinetic Approach. Molecules 2016, 21, 1502. [Google Scholar] [CrossRef]
- Pina, F.; Melo, M.J.; Parola, A.J.; Maestri, M.; Balzani, V. pH-Controlled Photochromism of Hydroxyflavylium ions. Chem. Eur. J. 1998, 4, 2001–2007. [Google Scholar] [CrossRef]
- Moncada, M.C.; Parola, A.J.; Lodeiro, C.; Pina, F.; Maestri, M.; Balzani, V. Multistate/Multifunctional Behaviour of 6-nitro-4’-hydroxyflavylium A Write-lock/Read/Unlock/Enable-erase/Erase Cycle Driven by Light and pH Stimulation. Chem. Eur. J. 2004, 10, 1519–1526. [Google Scholar] [CrossRef]
- Pina, F.; Petrov, V.; Laia, C.A.T. Photochromism of flavylium systems: An overview of a versatile multistate system. Dyes Pigment. 2012, 92, 877–889. [Google Scholar] [CrossRef]
- Giestas, L.; Folgosa, F.; Lima, J.C.; Parola, A.J.; Pina, F. Bio-Inspired Multistate Networks Responsive to Light, pH and Thermal Inputs: An Example of a Multistate System Operating Through Different Algorithms. Eur. J. Org. Chem. 2005, 4187–4200. [Google Scholar] [CrossRef]
- Jimenez, A.; Pinheiro, C.; Parola, A.J.; Maestri, M.; Pina, F. The Chemistry of 6-Hydroxyflavylium, Zwitterionic base, and p-Quinoidal Chalcones A Multiswitchable System Operated by Proton, Electron and Photon Inputs. Photochem. Photobiol. Sci. 2007, 6, 372–380. [Google Scholar] [CrossRef]
- Pina, F.; Melo, M.J.; Laia, C.A.T.; Parola, A.J.; Lima, J.C. Chemistry and Applications of Flavylium Compounds, a Handful of Colours. Chem. Soc. Rev. 2012, 41, 869–908. [Google Scholar] [CrossRef]
- Yoshida, K.; Mori, M.; Kondo, T. Blue flower color development by anthocyanins, from chemical structure to cell physiology. Nat. Prod. Rep. 2009, 26, 884–915. [Google Scholar] [CrossRef] [PubMed]
- Gomes, R.; Parola, A.J.; Bastkowski, F.; Polkowska, J.; Klärner, F.-G. Host-guest interactions between molecular clips and multistate systems based on flavylium salts. J. Am. Chem. Soc. 2009, 131, 8922–8938. [Google Scholar] [CrossRef] [PubMed]
- Basílio, N.; Pina, F. Flavylium network of chemical reactions in confined media, modulation of 3′,4′,7-trihydroxyflavilium reactions by host-guest interactions with cucurbit[7]uril. ChemPhysChem 2014, 15, 2295–2302. [Google Scholar] [CrossRef] [PubMed]
- Zubillaga, A.; Ferreira, P.; Parola, A.J.; Gago, S.; Basílio, N. pH-Gated Photoresponsive Shuttling in a Water-Soluble Pseudorotaxane. Chem. Commun. 2018, 54, 2743–2746. [Google Scholar] [CrossRef] [PubMed]
- Seco, A.; Diniz, A.M.; Sarrato, J.; Mourão, H.; Cruz, H.; Parola, A.J.; Basílio, N. A pseudorotaxane formed from a cucurbit[7]uril wheel and a bioinspired molecular axle with pH, light and redox-responsive properties. Pure Appl. Chem. 2020, 92, 301–314. [Google Scholar] [CrossRef]
- Mistry, T.V.; Cai, Y.; Lilley, T.H.; Haslam, E. Polyphenol interactions. Part 5. Anthocyanin co-pigmentation. J. Chem. Soc. Perkin Trans. 1991, 8, 1287–1296. [Google Scholar] [CrossRef]
- Sarma, A.D.; Sharma, R. Anthocyanin-DNA copigmentation complex: Mutual protection against oxidative damage. Phytochemistry 1999, 52, 1313–1318. [Google Scholar] [CrossRef]
- Mas, T.; Susperregui, J.; Berke, B.; Chèze, C.; Moreau, S.; Nuhrich, A.; Vercauteren, J. DNA triplex stabilization prop-erty of natural anthocyanins. Phytochemistry 2000, 53, 679–687. [Google Scholar] [CrossRef]
- Habermeyer, M.; Fritz, J.; Barthelmes, H.U.; Christensen, M.O.; Larsen, M.K.; Boege, F.; Marko, D. Anthocyanidins modulate the activity of human DNA topoisomerases I and II and affect cellular DNA integrity. Chem. Res. Toxicol. 2005, 18, 1395–1404. [Google Scholar] [CrossRef]
- Webb, M.R.; Min, K.; Ebeler, S.E. Anthocyanin interactions with DNA: Intercalation, topoisomerase I inhibition and oxidative reactions. J. Food Biochem. 2008, 32, 576–596. [Google Scholar] [CrossRef]
- Katritzky, A.R.; Czerney, P.; Levell, J.R.; Du, W. Molecular Engineering of Benzo[b]pyrylium Salts by Indirect Electrophilic Substitution. Eur. J. Org. Chem. 1998, 2623–2629. [Google Scholar] [CrossRef]
- Haucke, G.; Czerney, P.; Steen, D.; Rettig, W.; Hartmann, H. Radiationless Transitions in Substituted Benzopyrylium Dyes, Competing Adiabatic Photoreactions Towards Nonradiative Funnels. Ber Bunsenges Phys. Chem. 1993, 97, 561–570. [Google Scholar] [CrossRef]
- Laia, C.A.T.; Parola, A.J.; Folgosa, F.; Pina, F. Multistate reaction kinetics of 6-hydroxy-4’-dimethylaminoflavylium driven by pH A stopped-flow study. Org Biomol. Chem. 2007, 5, 69–77. [Google Scholar] [CrossRef]
- Tumir, L.M.; Piantanida, I.; Cindric, I.J.; Hrenar, T.; Meic, Z.; Zinic, M. New permanently charged phenanthridinium-nucleobase conjugates. Interactions with nucleotides and polynucleotides and recognition of ds-polyAH(+). J. Phys. Org. Chem. 2003, 16, 891–899. [Google Scholar] [CrossRef]
- Zhao, H.Y.; Schuck, P. Global Multi-Method Analysis of Affinities and Cooperativity in Complex Systems of Macromolecular Interactions. Anal. Chem. 2012, 84, 9513–9519. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.Y.; Piszczek, G.; Schuck, P. SEDPHAT—A platform for global ITC analysis and global multi-method analysis of molecular interactions. Methods 2015, 76, 137–148. [Google Scholar] [CrossRef] [PubMed]
- Scatchard, G. The attractions of proteins for small molecules and ions. Ann. N. Y. Acad. Sci. 1949, 51, 660–672. [Google Scholar] [CrossRef]
- McGhee, J.D.; von Hippel, P.H. Theoretical aspects of DNA-protein interactions, Co-operative and non-co-operative binding of large ligands to a one-dimensional homogeneous lattice. J. Mol. Biol. 1976, 103, 679–684. [Google Scholar] [CrossRef]
- Steenken, S.; Jovanovic, S.V. How Easily Oxidizable Is DNA? One-Electron Reduction Potentials of Adenosine and Guanosine Radicals in Aqueous Solution. J. Am. Chem. Soc. 1997, 119, 617–618. [Google Scholar] [CrossRef]
- da Silva, P.F.; Lima, J.C.; Quina, F.H.; Maçanita, A.L. Excited-State Electron Transfer in Anthocyanins and Related Flavylium Salts. J. Phys. Chem. A 2004, 108, 10133–10140. [Google Scholar] [CrossRef]
- Piantanida, I.; Mašić, L.; Rusak, G. Structure-spectrophotometric selectivity relationship in interactions of quercetin related flavonoids with double stranded and single stranded RNA. J. Mol. Struct. 2009, 924, 138–143. [Google Scholar] [CrossRef]
- Marinić, M.; Piantanida, I.; Rusak, G.; Žinić, M. Interactions of quercetin and its lanthane complex with double stranded DNA/RNA and single stranded RNA, Spectrophotometric sensing of poly G. J. Inorg. Biochem. 2006, 100, 288–298. [Google Scholar] [CrossRef] [PubMed]
- Demeunynck, M.; Bailly, C.; Wilson, W.D. Small Molecule DNA and RNA Binders: From Synthesis to Nucleic Acid Complexes; Wiley-VCH: Weinheim, Germany, 2004. [Google Scholar]
- Long, E.C.; Barton, J.K. On demonstrating DNA intercalation. Acc. Chem. Res. 1990, 23, 271–273. [Google Scholar] [CrossRef]
- Šmidlehner, T.; Piantanida, I.; Pescitelli, G. Polarization spectroscopy methods in the determination of interactions of small molecules with nucleic acids-tutorial. Beilstein J. Org. Chem. 2018, 14, 84–105. [Google Scholar] [CrossRef] [PubMed]
- Eriksson, M.; Nordén, B. Drug–Nucleic Acid Interactions. In Methods in Enzymology; Academic Press: San Diego, CA, USA, 2001; Volume 340, pp. 68–98. [Google Scholar]
- Hernandez-Folgado, L.; Schmuck, C.; Tomić, S.; Piantanida, I. A novel pyrene-guanidiniocarbonyl-pyrrole cation efficiently differentiates between ds-DNA and ds-RNA by two independent, sensitive spectroscopic methods. Bioorg. Med. Chem. Lett. 2008, 18, 2977–2981. [Google Scholar] [CrossRef] [PubMed]
- Armitage, B.A. DNA Binders and Related Subjects: Cyanine dye-DNA interactions: Intercalation, groove binding, and aggregation. Top. Curr. Chem. 2005, 253, 55–76. [Google Scholar]
- Mergny, J.L.; Lacroix, L. Analysis of thermal melting curves. Oligonucleotides 2003, 13, 515–537. [Google Scholar] [CrossRef]
- Piantanida, I.; Palm, B.S.; Žinić, M.; Schneider, H.J. A new 4,9-diazapyrenium intercalator for single- and double-stranded nucleic acids: Distinct differences from related diazapyrenium compounds and ethidium bromide. Perkin Trans. 2001, 1808–1816. [Google Scholar] [CrossRef]
- Wong, P.; Lee, C.; Tannock, I.F. Reduction of intracellular pH as a strategy to enhance the pH-dependent cytotoxic effects of melphalan for human breast cancer cells. Clin. Cancer Res. 2005, 11, 3553–3557. [Google Scholar] [CrossRef][Green Version]
- Gillies, R.J.; Robey, I.; Gatenby, R.A. Causes and consequences of increased glucose metabolism of cancers. J. Nucl. Med. 2008, 49, 24s–42s. [Google Scholar] [CrossRef]
- Raghunand, N.; Gillies, R.J. pH and drug resistance in tumors. Drug Resist. Update 2000, 3, 39–47. [Google Scholar] [CrossRef] [PubMed]
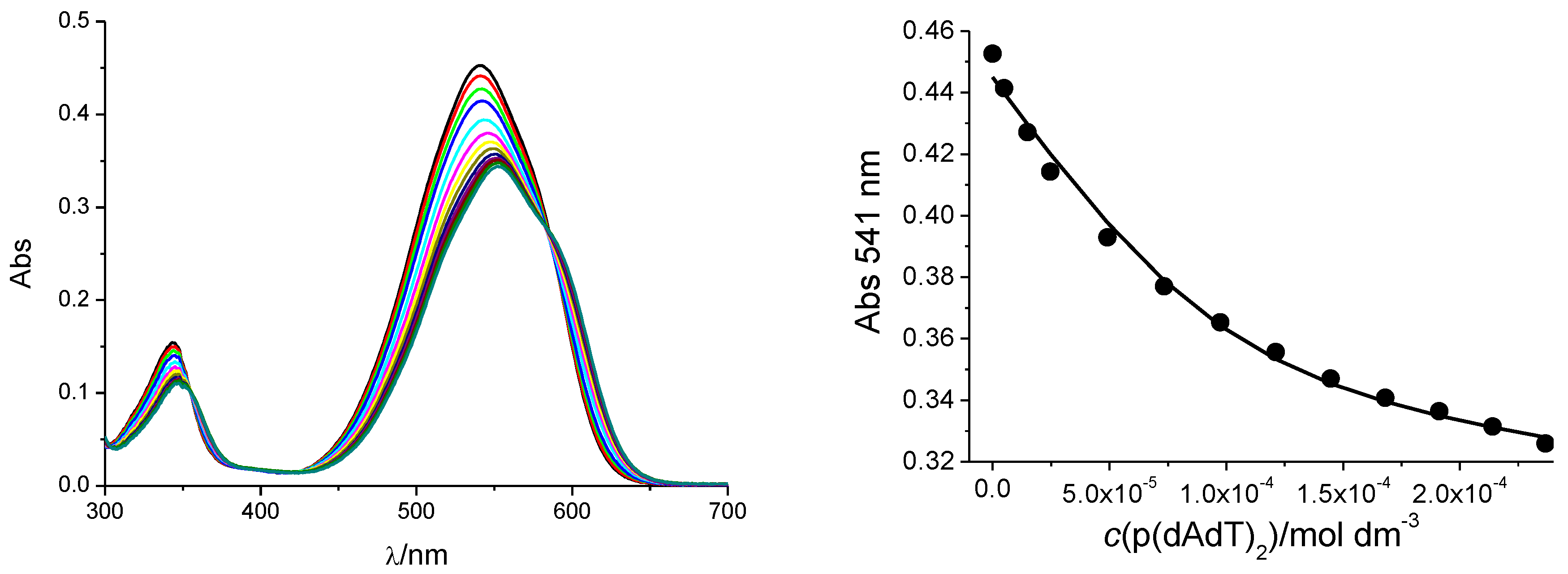
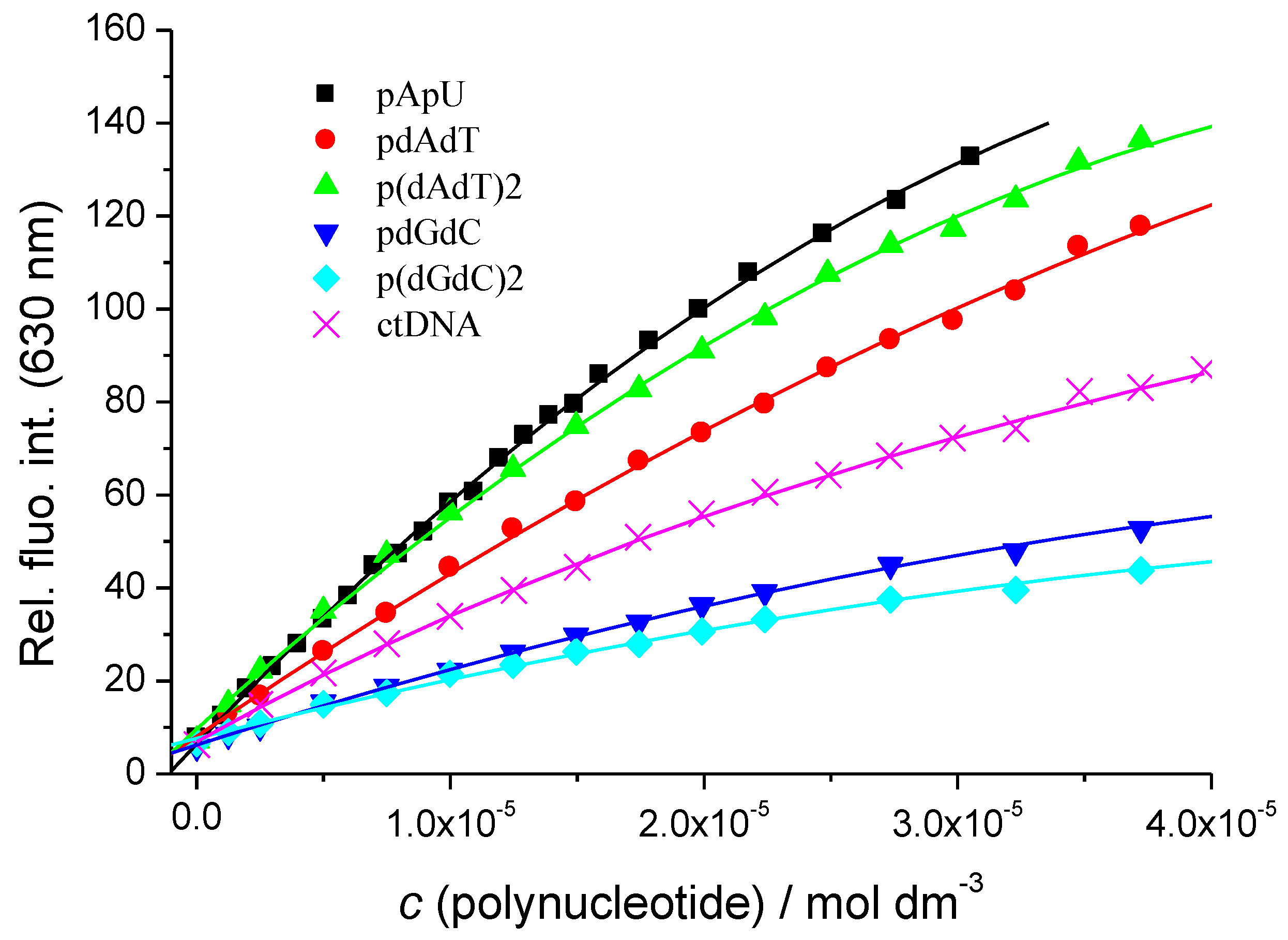
| Sample | τ/ps |
|---|---|
| F1 | 47 ± 7 |
| F1 + pApU | 284 ± 10 |
| F1 + pdAdT | 232 ± 6 |
| F1 + p(dAdT)2 | 220 ± 12 |
| F1 + p(dGdC)2 | 117 ± 8 |
| F1 + pdGdC | 118 ± 16 |
| Polynucleotides | logKs | n | a Δλ/nm | b ΔIcalc | c Stokes Shift/nm |
|---|---|---|---|---|---|
| ctDNA | 5.2 | 0.12 | +12 | 227 | 88 |
| poly A–poly U | 4.8 | 0.18 | +17 | 375 | 88 |
| poly dA–poly dT | 6.1 | 0.05 | +10 | 419 | 90 |
| poly dG–poly dC | 5.5 | 0.07 | +18 | 122 | 97 |
| poly (dA–dT)2 | 5.2 | 0.14 | +11 | 296 | 90 |
| poly (dG–dC)2 | 4.9 | 0.23 | +15 | 89 | 92 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Crnolatac, I.; Giestas, L.; Horvat, G.; Parola, A.J.; Piantanida, I. Flavylium Dye as pH-Tunable Fluorescent and CD Probe for Double-Stranded DNA and RNA. Chemosensors 2020, 8, 129. https://doi.org/10.3390/chemosensors8040129
Crnolatac I, Giestas L, Horvat G, Parola AJ, Piantanida I. Flavylium Dye as pH-Tunable Fluorescent and CD Probe for Double-Stranded DNA and RNA. Chemosensors. 2020; 8(4):129. https://doi.org/10.3390/chemosensors8040129
Chicago/Turabian StyleCrnolatac, Ivo, Letícia Giestas, Gordan Horvat, António Jorge Parola, and Ivo Piantanida. 2020. "Flavylium Dye as pH-Tunable Fluorescent and CD Probe for Double-Stranded DNA and RNA" Chemosensors 8, no. 4: 129. https://doi.org/10.3390/chemosensors8040129
APA StyleCrnolatac, I., Giestas, L., Horvat, G., Parola, A. J., & Piantanida, I. (2020). Flavylium Dye as pH-Tunable Fluorescent and CD Probe for Double-Stranded DNA and RNA. Chemosensors, 8(4), 129. https://doi.org/10.3390/chemosensors8040129