Dysarthria Speech Detection Using Convolutional Neural Networks with Gated Recurrent Unit
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Collection
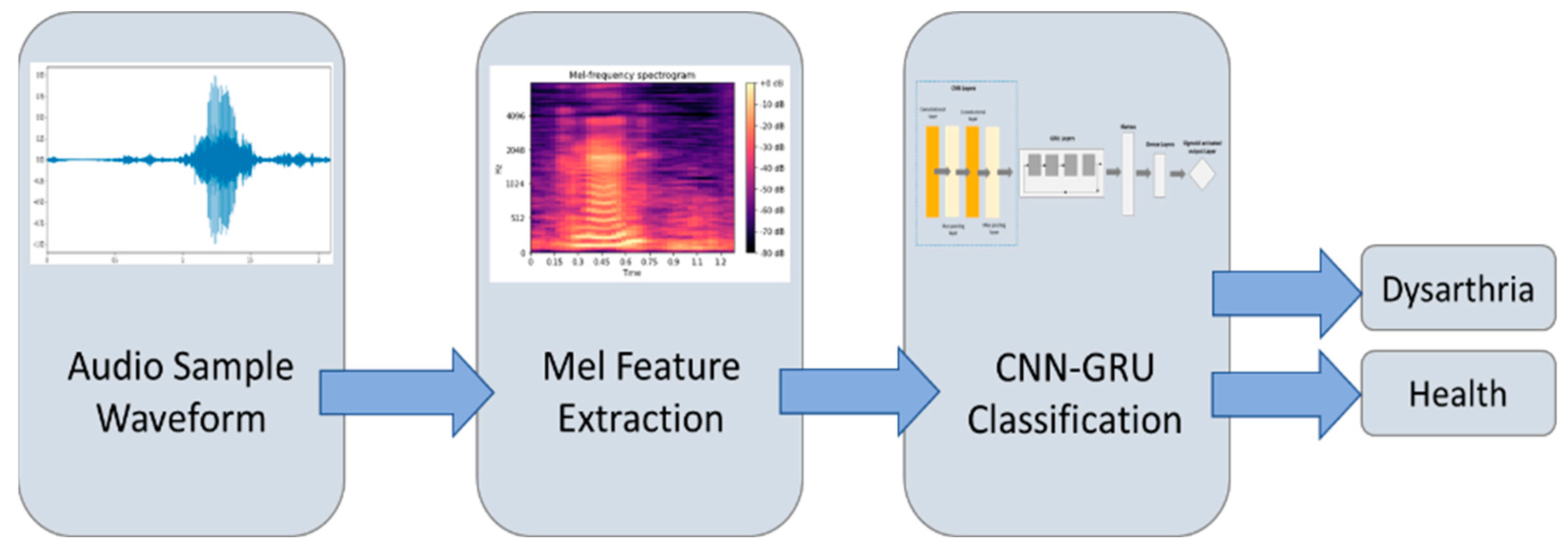
2.2. Method
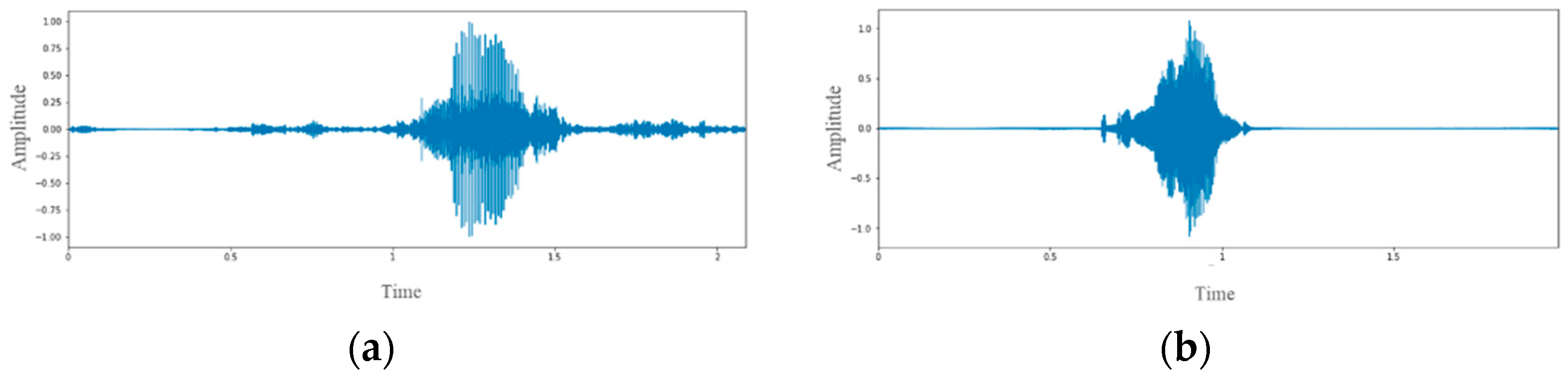
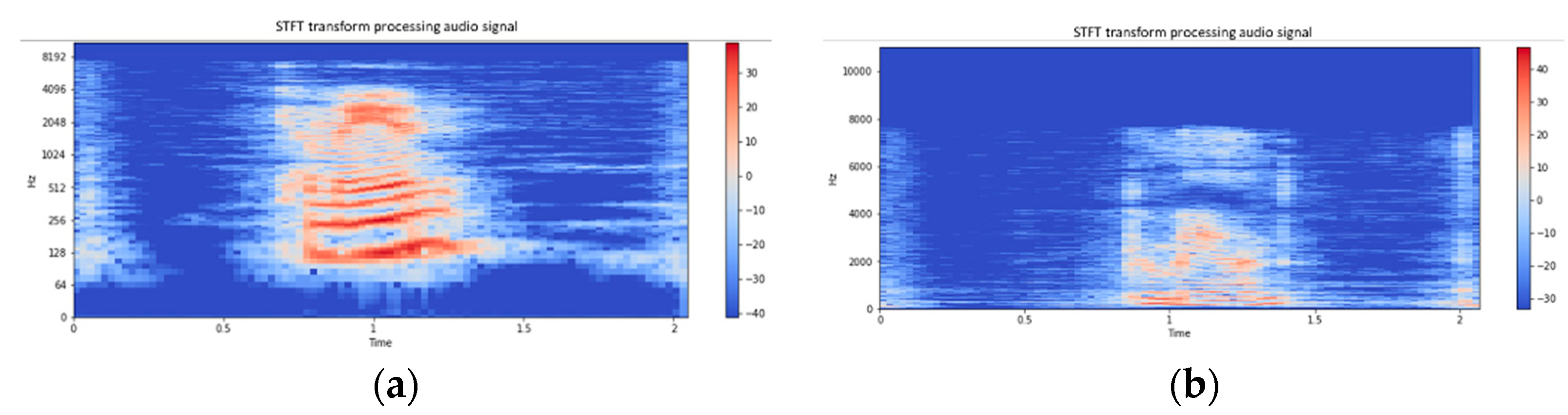
2.3. Data Preprocessing
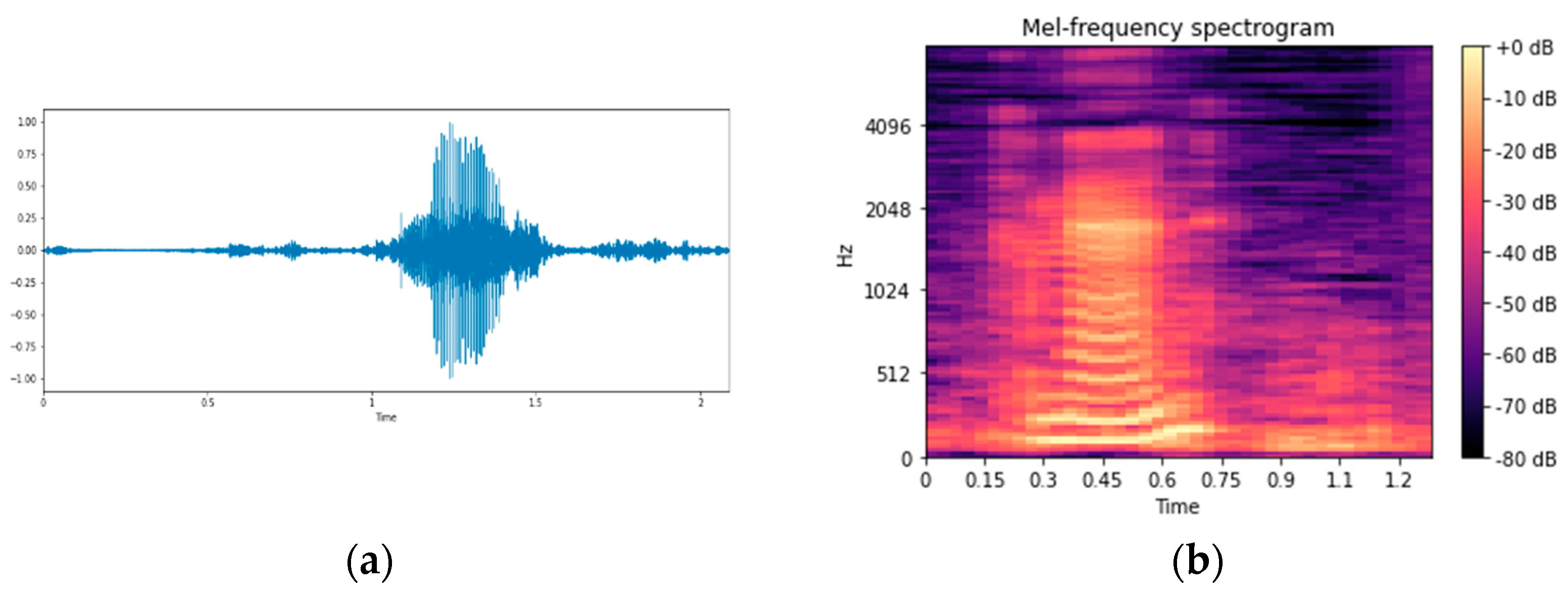
2.4. Feature Selection
2.5. Deep Learning Algorithms
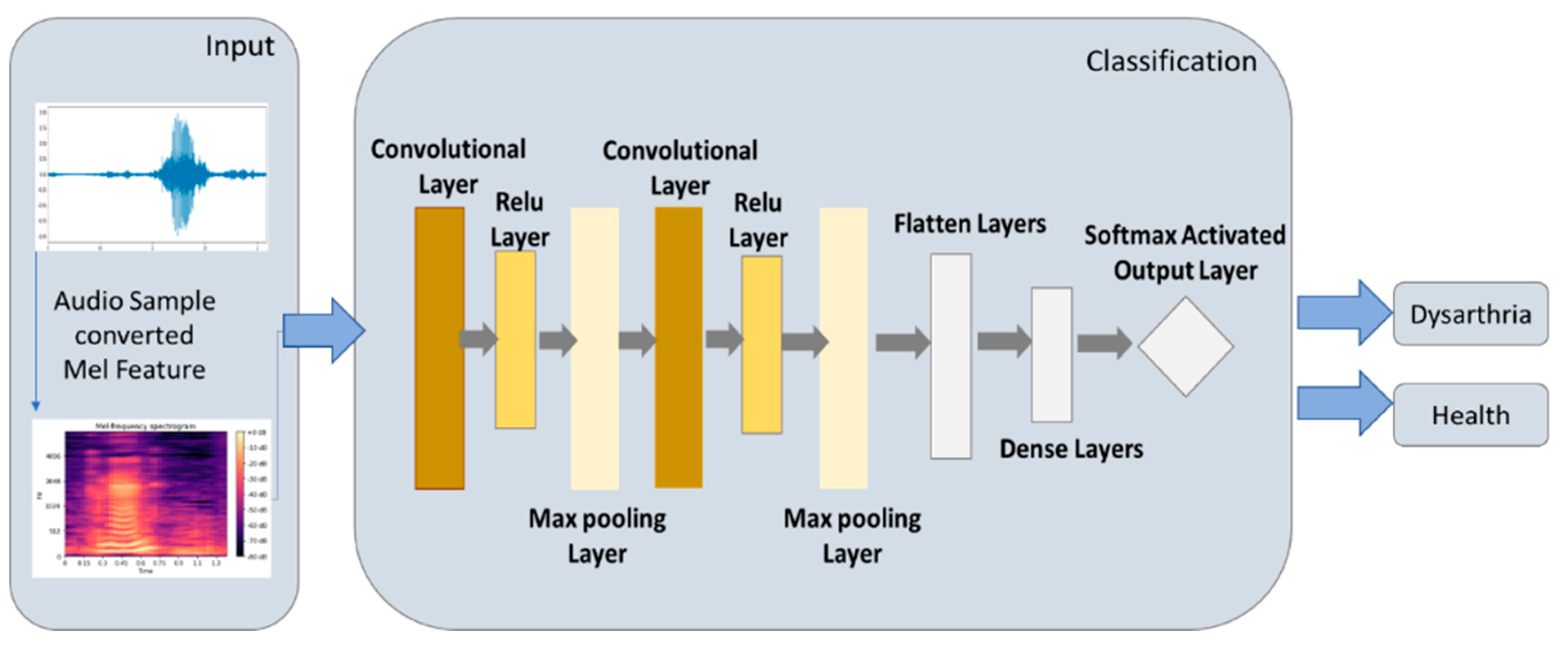
2.5.1. CNN Model
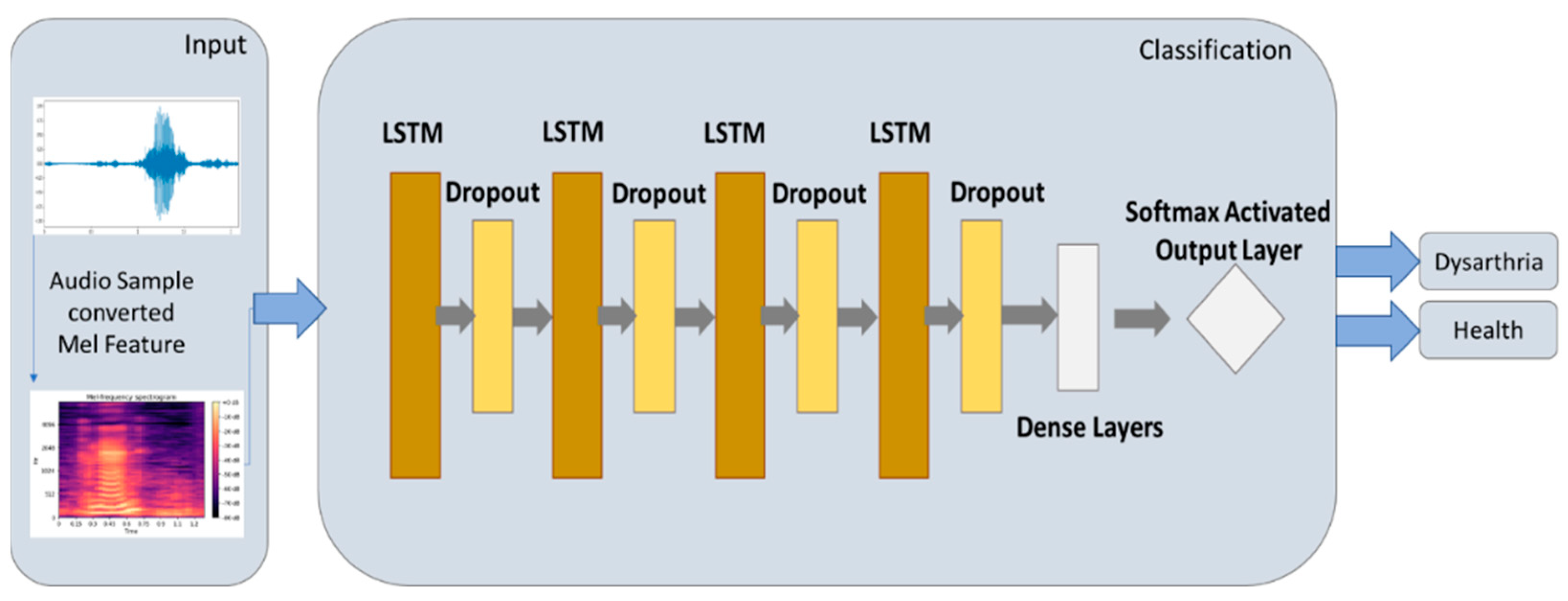
2.5.2. LSTM Model
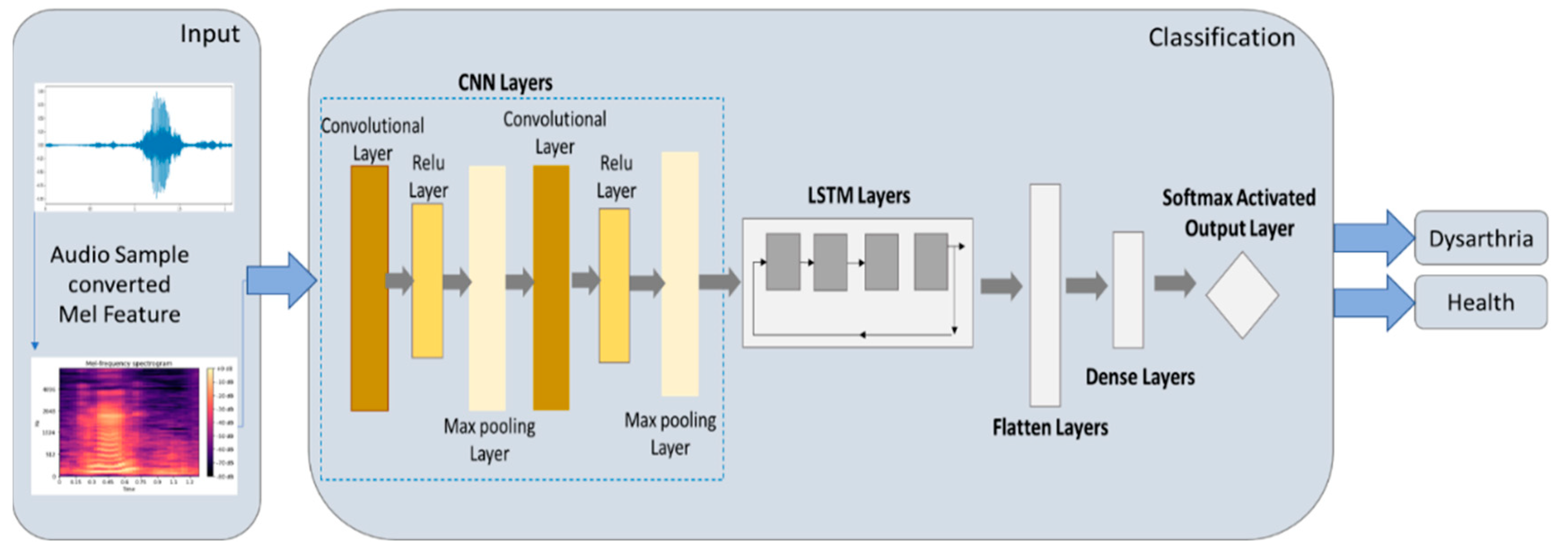
2.5.3. CNN-LSTM Model
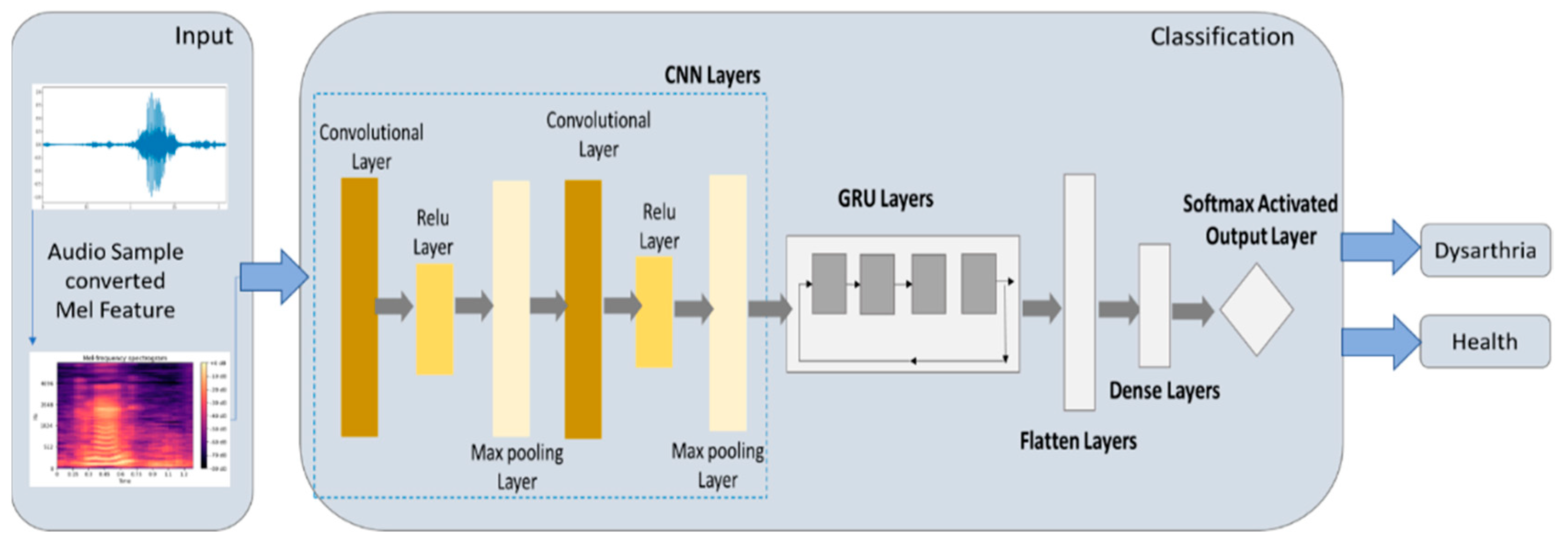
2.5.4. CNN-GRU Model
2.6. Experimental Design
2.7. Model Evaluation
3. Experimental Results
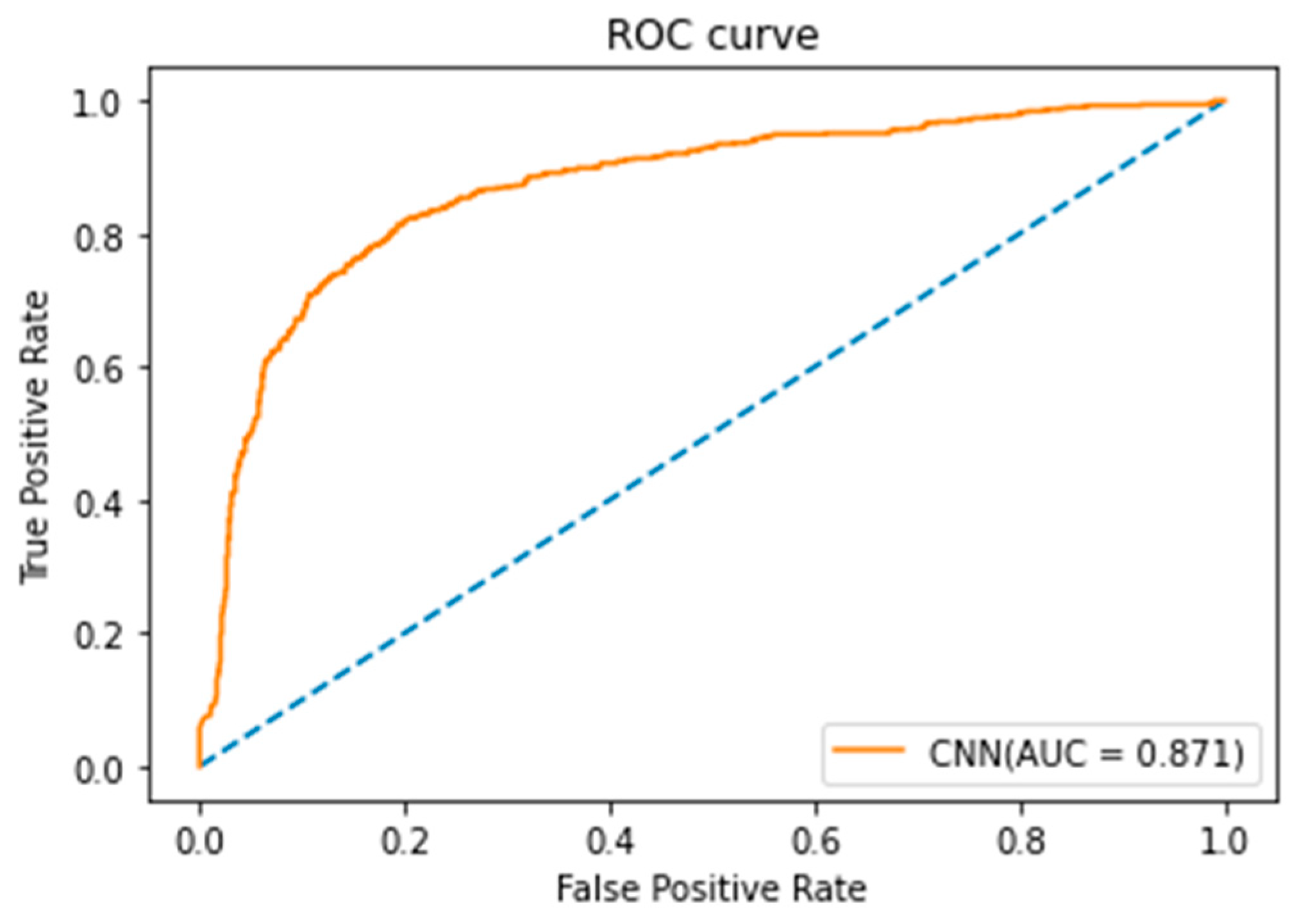
3.1. Experimental Results of CNN Model
3.2. Experimental Results of LSTM Model
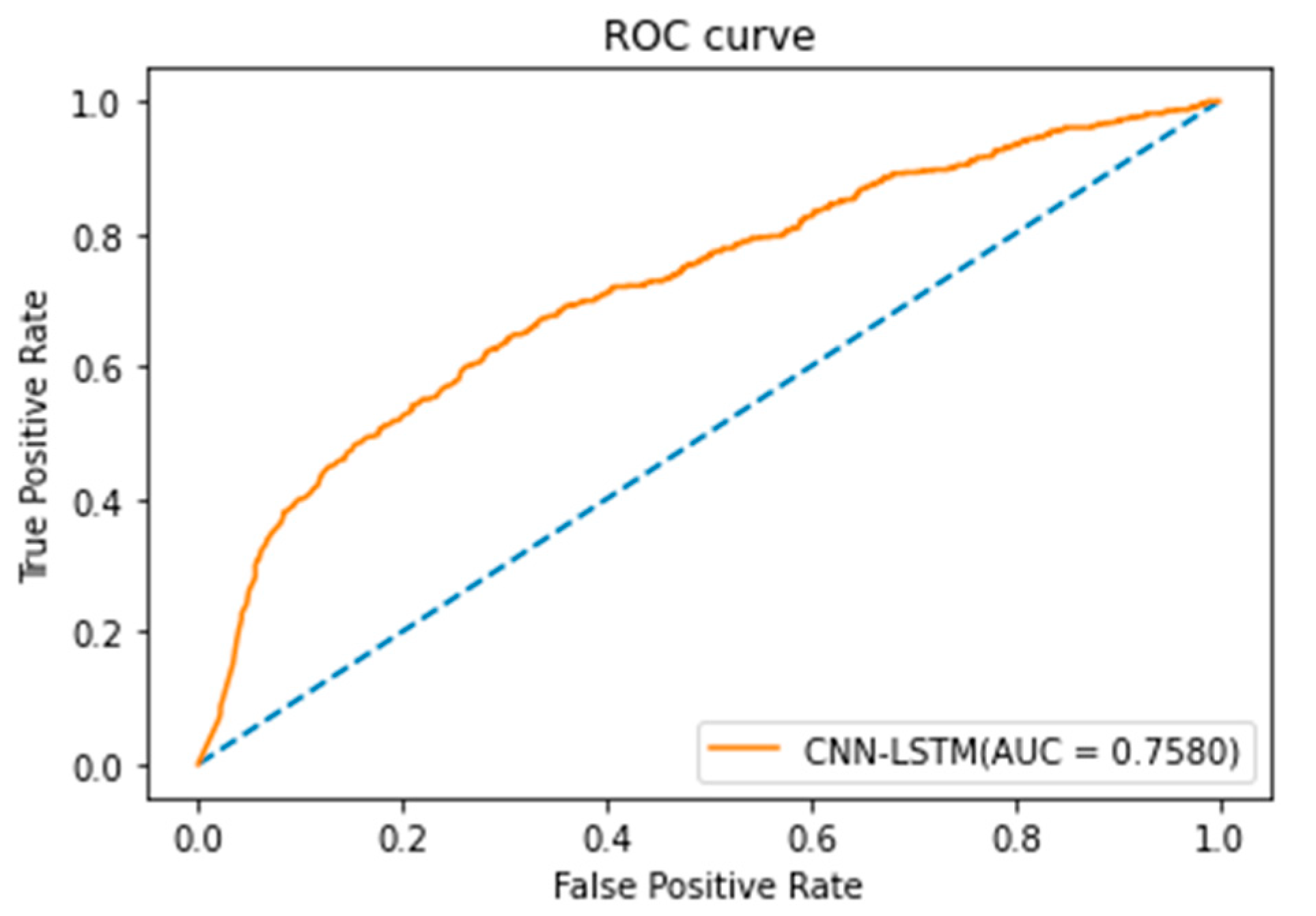
3.3. Experimental Results of CNN-LSTM
3.4. Experimental Results of CNN-GRU Model
4. Discussion of Results
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Gentil, M.; Pollak, P.; Perret, J. Parkinsonian dysarthria. Rev. Neurol. 1995, 151, 105–112. [Google Scholar] [PubMed]
- Rampello, L.; Rampello, L.; Patti, F.; Zappia, M. When the word doesn’t come out: A synthetic overview of dysarthria. J. Neurol. Sci. 2016, 369, 354–360. [Google Scholar] [CrossRef] [PubMed]
- Marmor, S.; Horvath, K.J.; Lim, K.O.; Misono, S. Voice problems and depression among adults in the U nited S tates. Laryngoscope 2016, 126, 1859–1864. [Google Scholar] [CrossRef] [PubMed]
- Van Nuffelen, G.; Middag, C.; De Bodt, M.; Martens, J. Speech technology-based assessment of phoneme intelligibility in dysarthria. Int. J. Lang. Commun. Disord. 2009, 44, 716–730. [Google Scholar] [CrossRef] [PubMed]
- Vashkevich, M.; Rushkevich, Y. Classification of ALS patients based on acoustic analysis of sustained vowel phonations. Biomed. Signal Process. Control 2020, 65, 102350. [Google Scholar] [CrossRef]
- Muhammad, G.; Alsulaiman, M.; Ali, Z.; Mesallam, T.A.; Farahat, M.; Malki, K.H.; Al-Nasheri, A.; Bencherif, M.A. Voice pathology detection using interlaced derivative pattern on glottal source excitation. Biomed. Signal Process. Control 2017, 31, 156–164. [Google Scholar] [CrossRef]
- Karan, B.; Sahu, S.S.; Mahto, K. Parkinson disease prediction using intrinsic mode function based features from speech signal. Biocybern. Biomed. Eng. 2019, 40, 249–264. [Google Scholar] [CrossRef]
- Moro-Velazquez, L.; Gómez-García, J.A.; Godino-Llorente, J.I.; Villalba, J.; Orozco-Arroyave, J.R.; Dehak, N. Analysis of speaker recognition methodologies and the influence of kinetic changes to automatically detect Parkinson’s Disease. Appl. Soft Comput. 2018, 62, 649–666. [Google Scholar] [CrossRef]
- Albaqshi, H.; Sagheer, A. Dysarthric Speech Recognition using Convolutional Recurrent Neural Networks. Int. J. Intell. Eng. Syst. 2020, 13, 384–392. [Google Scholar] [CrossRef]
- Narendra, N.P.; Alku, P. Glottal source information for pathological voice detection. IEEE Access 2020, 8, 67745–67755. [Google Scholar] [CrossRef]
- Schlauch, R.S.; Anderson, E.S.; Micheyl, C. A demonstration of improved precision of word recognition scores. J. Speech, Lang. Heart Res. 2014, 57, 543–555. [Google Scholar] [CrossRef]
- Kim, H.; Hasegawa-Johnson, M.; Perlman, A.; Gunderson, J.; Huang, T.S.; Watkin, K.; Frame, S. Dysarthric speech database for universal access research. Interspeech 2008, 2008, 480. [Google Scholar] [CrossRef]
- Dumane, P.; Hungund, B.; Chavan, S. Dysarthria Detection Using Convolutional Neural Network. Techno-Soc. 2021, 2020, 449–457. [Google Scholar] [CrossRef]
- Gers, F.; Schmidhuber, E. LSTM recurrent networks learn simple context-free and context-sensitive languages. IEEE Trans. Neural. Netw. 2001, 12, 1333–1340. [Google Scholar] [CrossRef] [PubMed]
- Chaiani, M.; Selouani, S.A.; Boudraa, M.; Yakoub, M.S. Voice disorder classification using speech enhancement and deep learning models. Biocybern. Biomed. Eng. 2022, 42, 463–480. [Google Scholar] [CrossRef]
- Hasannezhad, M.; Ouyang, Z.; Zhu, W.P.; Champagne, B. An integrated CNN-GRU framework for complex ratio mask estimation in speech enhancement. In Proceedings of the 2020 Asia-Pacific Signal and Information Processing Association Annual Summit and Conference (APSIPA ASC), Auckland, New Zealand, 7–10 December 2020; pp. 764–768. [Google Scholar]
- Yerima, S.; Alzaylaee, M.; Shajan, A.; Vinod, P. Deep learning techniques for android botnet detection. Electronics 2021, 10, 519. [Google Scholar] [CrossRef]
- Chung, J.; Gulcehre, C.; Cho, K.; Bengio, Y. Empirical evaluation of gated recurrent neural networks on sequence modeling. arXiv preprint 2014, arXiv:1412.3555. [Google Scholar]
- Goutte, C.; Gaussier, E. A probabilistic interpretation of precision, recall and F-score, with implication for evaluation. In European Conference on Information Retrieval; Springer: Berlin/Heidelberg, Germany, 2005; pp. 345–359. [Google Scholar]
- Fawcett, T. An Introduction to ROC analysis. Pattern Recogn. Lett. 2006, 27, 861–874. [Google Scholar] [CrossRef]
- Vasilev, I.; Slater, D.; Spacagna, G.; Roelants, P.; Zocca, V. Python Deep Learning: Exploring Deep Learning Techniques and Neural Network Architectures with Pytorch, Keras, and TensorFlow; Packt Publishing Ltd.: Birmingham, UK, 2019. [Google Scholar]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Duchesnay, E. Scikit-learn: Machine learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Hernandez, A.; Chung, M. Dysarthria classification using acoustic properties of fricatives. In Proceedings of Seoul International Conference on Speech Sciences (SICSS) 2019, Seoul, Korea, 15–16 November 2019. [Google Scholar]
- Narendra, N.; Alku, P. Dysarthric speech classification from coded telephone speech using glottal features. Speech Commun. 2019, 110, 47–55. [Google Scholar] [CrossRef]
- Rajeswari, R.; Devi, T.; Shalini, S. Dysarthric Speech Recognition Using Variational Mode Decomposition and Convolutional Neural Networks. Wirel. Pers. Commun. 2021, 122, 293–307. [Google Scholar] [CrossRef]
- Priyanka, A.; Ganesan, K. Radiomic features based severity prediction in dementia MR images using hybrid SSA-PSO optimizer and multi-class SVM classifier. IRBM, 2022; in press. [Google Scholar]


| Network | Layer | No. of Activations | No. of Parameters |
|---|---|---|---|
| CNN | Cov1 | (27,27,32) | 160 |
| Maxpooling1 | (13,13,32) | 0 | |
| Cov2 | (12,12,64) | 8256 | |
| Maxpooling2 | (6,6,64) | 0 | |
| Flatten | (2304) | 0 | |
| Dense | (3) | 387 |
| Network | Layer | No. of Activations | No. of Parameters |
|---|---|---|---|
| LSTM | LSTM | (26,10) | 480 |
| Dropout | (26,10) | 0 | |
| LSTM | (26,10) | 840 | |
| Dropout | (26,10) | 0 | |
| LSTM | (26,10) | 840 | |
| Dropout | (26,10) | 0 | |
| LSTM | (26,10) | 840 | |
| Dropout | (26,10) | 0 | |
| Dense | (26,2) | 22 |
| Network | Layer | No. of Activations | No. of Parameters |
|---|---|---|---|
| CNN-LSTM | Cov1 | (23,32) | 320 |
| Maxpooling1 | (11,32) | 0 | |
| Cov2 | (11,64) | 14,400 | |
| Maxpooling2 | (5,64) | 0 | |
| LSTM | (2,128) | 20,608 | |
| Flatten | (64) | 0 | |
| Dense | (44) | 2860 |
| Network | Layer | No. of Activations | No. of Parameters |
|---|---|---|---|
| CNN-GRU | Cov1 | (23,32) | 320 |
| Maxpooling1 | (11,32) | 0 | |
| Cov2 | (11,64) | 14,400 | |
| Maxpooling2 | (5,64) | 0 | |
| GRU | (2,32) | 15,552 | |
| Flatten | (64) | 0 | |
| Dense | (44) | 2860 |
| Model | Batch Size | Learning Rate | Accuracy% | Precision% | Recall | F1-Score |
|---|---|---|---|---|---|---|
| CNN | 32 | 0.1 | 70.00 | 70.01 | 0.7002 | 0.7010 |
| 0.01 | 88.89 | 69.44 | 0.8334 | 0.7575 | ||
| 0.001 | 75.00 | 75.50 | 0.7500 | 0.7506 | ||
| 64 | 0.1 | 88.88 | 86.66 | 0.8333 | 0.8148 | |
| 0.01 | 89.89 | 87.50 | 0.8333 | 0.8285 | ||
| 0.001 | 86.67 | 79.99 | 0.666 | 0.6249 | ||
| 128 | 0.1 | 93.33 | 87.50 | 0.8333 | 0.8285 | |
| 0.01 | 94.36 | 90.39 | 0.8913 | 0.8896 | ||
| 0.001 | 94.35 | 86.66 | 0.8333 | 0.8148 |
| Epoch | Execution Time (ms) | Accuracy (Training) (%) | Loss Function | Accuracy (Validation) (%) | Accuracy (Testing) (%) |
|---|---|---|---|---|---|
| 1 | 3 | 82.23 | 0.4972 | 45.40 | 79.20 |
| 2 | 3 | 91.26 | 0.2129 | 83.41 | 90.27 |
| 3 | 3 | 93.42 | 0.1674 | 90.35 | 91.27 |
| 4 | 3 | 94.18 | 0.1444 | 92.64 | 92.30 |
| 5 | 3 | 94.86 | 0.1314 | 93.90 | 93.24 |
| 6 | 3 | 95.77 | 0.1134 | 94.87 | 93.50 |
| 7 | 3 | 96.60 | 0.0931 | 95.48 | 93.52 |
| 8 | 3 | 96.52 | 0.0894 | 96.37 | 94.27 |
| 9 | 3 | 97.21 | 0.0757 | 97.13 | 94.20 |
| 10 | 3 | 97.88 | 0.0638 | 97.53 | 94.36 |
| Model | Batch Size | Learning Rate | Accuracy% | Precision% | Recall | F1-Score |
|---|---|---|---|---|---|---|
| LSTM | 32 | 0.1 | 50.32 | 50.10 | 0.5001 | 0.5002 |
| 0.01 | 54.29 | 53.21 | 0.5321 | 0.5321 | ||
| 0.001 | 54.67 | 54.60 | 0.5460 | 0.5430 | ||
| 64 | 0.1 | 55.60 | 44.21 | 0.4421 | 0.4421 | |
| 0.01 | 54.30 | 54.12 | 0.5420 | 0.5411 | ||
| 0.001 | 56.61 | 53.43 | 0.5435 | 0.5324 | ||
| 128 | 0.1 | 55.21 | 44.25 | 0.6550 | 0.5220 | |
| 0.01 | 56.60 | 43.21 | 0.5521 | 0.4201 | ||
| 0.001 | 55.37 | 50.20 | 0.5020 | 0.5020 |
| Epoch | Execution Time (ms) | Accuracy (Training) (%) | Loss Function | Accuracy (Validation) (%) | Accuracy (Testing) (%) |
|---|---|---|---|---|---|
| 1 | 2 | 53.32 | 0.7346 | 54.89 | 53.20 |
| 2 | 2 | 53.63 | 0.7360 | 55.32 | 53.39 |
| 3 | 2 | 54.29 | 0.7375 | 55.36 | 53.65 |
| 4 | 2 | 54.42 | 0.7394 | 54.22 | 54.30 |
| 5 | 2 | 54.68 | 0.7153 | 55.89 | 54.39 |
| 6 | 2 | 54.56 | 0.7163 | 55.90 | 54.37 |
| 7 | 2 | 54.04 | 0.7309 | 55.91 | 54.49 |
| 8 | 2 | 55.60 | 0.7316 | 56.01 | 55.60 |
| 9 | 2 | 56.01 | 0.7557 | 56.43 | 55.97 |
| 10 | 2 | 56.60 | 0.7562 | 56.42 | 56.61 |
| Model | Batch Size | Learning Rate | Accuracy% | Precision% | Recall | F1-Score |
|---|---|---|---|---|---|---|
| CNN-LSTM | 32 | 0.1 | 62.49 | 65.99 | 0.6666 | 0.6549 |
| 0.01 | 66.66 | 64.44 | 0.6656 | 0.6333 | ||
| 0.001 | 73.20 | 68.54 | 0.6756 | 0.6723 | ||
| 64 | 0.1 | 70.21 | 69.45 | 0.6230 | 0.7165 | |
| 0.01 | 69.20 | 69.44 | 0.7333 | 0.7175 | ||
| 0.001 | 73.21 | 67.54 | 0.6740 | 0.6740 | ||
| 128 | 0.1 | 75.30 | 70.15 | 0.6563 | 0.7490 | |
| 0.01 | 78.57 | 70.33 | 0.6660 | 0.7500 | ||
| 0.001 | 77.33 | 69.44 | 0.7475 | 0.7375 |
| Epoch | Execution Time (ms) | Accuracy (Training) (%) | Loss Function | Accuracy (Validation) (%) | Accuracy (Testing) (%) |
|---|---|---|---|---|---|
| 1 | 5 | 42.11 | 0.9493 | 50.00 | 43.50 |
| 2 | 5 | 57.89 | 0.8010 | 66.67 | 50.65 |
| 3 | 5 | 63.16 | 0.6720 | 66.67 | 51.27 |
| 4 | 8 | 73.68 | 0.5617 | 66.67 | 65.90 |
| 5 | 4 | 84.21 | 0.3367 | 83.33 | 66.37 |
| 6 | 8 | 84.21 | 0.3256 | 83.33 | 67.47 |
| 7 | 5 | 84.21 | 0.3102 | 83.33 | 70.30 |
| 8 | 6 | 84.21 | 0.3060 | 83.33 | 75.98 |
| 9 | 5 | 84.21 | 0.2665 | 83.33 | 76.35 |
| 10 | 5 | 84.21 | 0.2745 | 83.33 | 78.57 |
| Model | Batch Size | Learning Rate | Accuracy% | Precision% | Recall | F1-Score |
|---|---|---|---|---|---|---|
| CNN-GRU | 32 | 0.1 | 92.27 | 93.21 | 0.9121 | 0.9220 |
| 0.01 | 94.52 | 94.23 | 0.9422 | 0.9420 | ||
| 0.001 | 95.21 | 93.20 | 0.9220 | 0.9231 | ||
| 64 | 0.1 | 96.41 | 95.51 | 0.9421 | 0.9412 | |
| 0.01 | 96.70 | 90.24 | 0.9026 | 0.9633 | ||
| 0.001 | 96.38 | 96.31 | 0.9427 | 0.9532 | ||
| 128 | 0.1 | 97.71 | 96.21 | 0.9621 | 0.9621 | |
| 0.01 | 98.02 | 90.47 | 0.9030 | 0.9021 | ||
| 0.001 | 98.88 | 91.47 | 0.9147 | 0.9147 |
| Epoch | Execution Time (ms) | Accuracy (Training) (%) | Loss Function | Accuracy (Validation) (%) | Accuracy (Testing) (%) |
|---|---|---|---|---|---|
| 1 | 2 | 79.20 | 0.157 | 90.77 | 89.21 |
| 2 | 2 | 92.27 | 0.1267 | 91.08 | 90.20 |
| 3 | 2 | 94.52 | 0.3353 | 90.97 | 93.45 |
| 4 | 2 | 95.21 | 0.2937 | 91.16 | 94.60 |
| 5 | 2 | 96.41 | 0.1553 | 90.77 | 95.88 |
| 6 | 2 | 96.70 | 0.1274 | 91.36 | 97.56 |
| 7 | 2 | 96.83 | 0.1029 | 91.20 | 96.30 |
| 8 | 2 | 97.71 | 0.2396 | 91.40 | 97.13 |
| 9 | 2 | 98.02 | 0.2084 | 91.63 | 97.79 |
| 10 | 2 | 98.14 | 0.1621 | 91.52 | 98.38 |
| Author | Classification Method | Dataset | Accuracy (%) | Execution Time |
|---|---|---|---|---|
| Hernandez et al. (2019) [23] | SVM | UA-Speech | 72% | - |
| Narendra et al. (2019) [24] | SVM | UA-Speech | 96.38% | - |
| Narendra et al. (2020) [10] | CNN-MLP | UA-Speech | 87.93% | - |
| CNN-LSTM | 77.57% | - | ||
| Rajeswari et al. (2022) [25] | CNN | UA-Speech | 95.95% | - |
| Our Approach | CNN-GRU | UA-Speech | 98.38% | 2 ms |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shih, D.-H.; Liao, C.-H.; Wu, T.-W.; Xu, X.-Y.; Shih, M.-H. Dysarthria Speech Detection Using Convolutional Neural Networks with Gated Recurrent Unit. Healthcare 2022, 10, 1956. https://doi.org/10.3390/healthcare10101956
Shih D-H, Liao C-H, Wu T-W, Xu X-Y, Shih M-H. Dysarthria Speech Detection Using Convolutional Neural Networks with Gated Recurrent Unit. Healthcare. 2022; 10(10):1956. https://doi.org/10.3390/healthcare10101956
Chicago/Turabian StyleShih, Dong-Her, Ching-Hsien Liao, Ting-Wei Wu, Xiao-Yin Xu, and Ming-Hung Shih. 2022. "Dysarthria Speech Detection Using Convolutional Neural Networks with Gated Recurrent Unit" Healthcare 10, no. 10: 1956. https://doi.org/10.3390/healthcare10101956
APA StyleShih, D.-H., Liao, C.-H., Wu, T.-W., Xu, X.-Y., & Shih, M.-H. (2022). Dysarthria Speech Detection Using Convolutional Neural Networks with Gated Recurrent Unit. Healthcare, 10(10), 1956. https://doi.org/10.3390/healthcare10101956








