An Exhaustive Power Comparison of Normality Tests
Abstract
1. Introduction
- it can provide insights about the observed process;
- parameters of model can be inferred from the characteristics of data distributions; and
- it can help in choosing more specific and computationally efficient methods.
2. Statistical Methods
2.1. Chi-Square Test (CHI2)
2.2. Kolmogorov–Smirnov (KS)
2.3. Anderson–Darling (AD)
2.4. Cramer–Von Mises (CVM)
2.5. Shapiro–Wilk (SW)
2.6. Lilliefors (LF)
2.7. D’Agostino (DA)
2.8. Shapiro–Francia (SF)
2.9. D’Agostino–Pearson (DAP)
2.10. Filliben (Filli)
2.11. Martinez–Iglewicz (MI)
2.12. Epps–Pulley (EP)
2.13. Jarque–Bera (JB)
2.14. Hosking (
2.15. Cabaña–Cabaña (CC1-CC2)
2.16. The Chen–Shapiro Test (ChenS)
2.17. Modified Shapiro-Wilk (SWRG)
2.18. Doornik–Hansen (DH)
2.19. Zhang (ZQ), (ZQstar), (ZQQstar)
2.20. Barrio–Cuesta-Albertos–Matran–Rodriguez-Rodriguez (BCMR)
2.21. Glen–Leemis–Barr (GLB)
2.22. Bonett–Seier (BS)
2.23. Bontemps–Meddahi (BM1–, BM2–)
2.24. Zhang–Wu (ZW1–, ZW2–)
2.25. Gel–Miao–Gastwirth (GMG)
2.26. Robust Jarque–Bera (RJB)
2.27. Coin
2.28. Brys–Hubert–Struyf (BHS)
2.29. Brys–Hubert–Struyf–Bonett–Seier (BHSBS)
2.30. Desgagné–Lafaye de Micheaux–Leblanc (DLDMLRn), (DLDMXAPD), (DLDMZEPD)
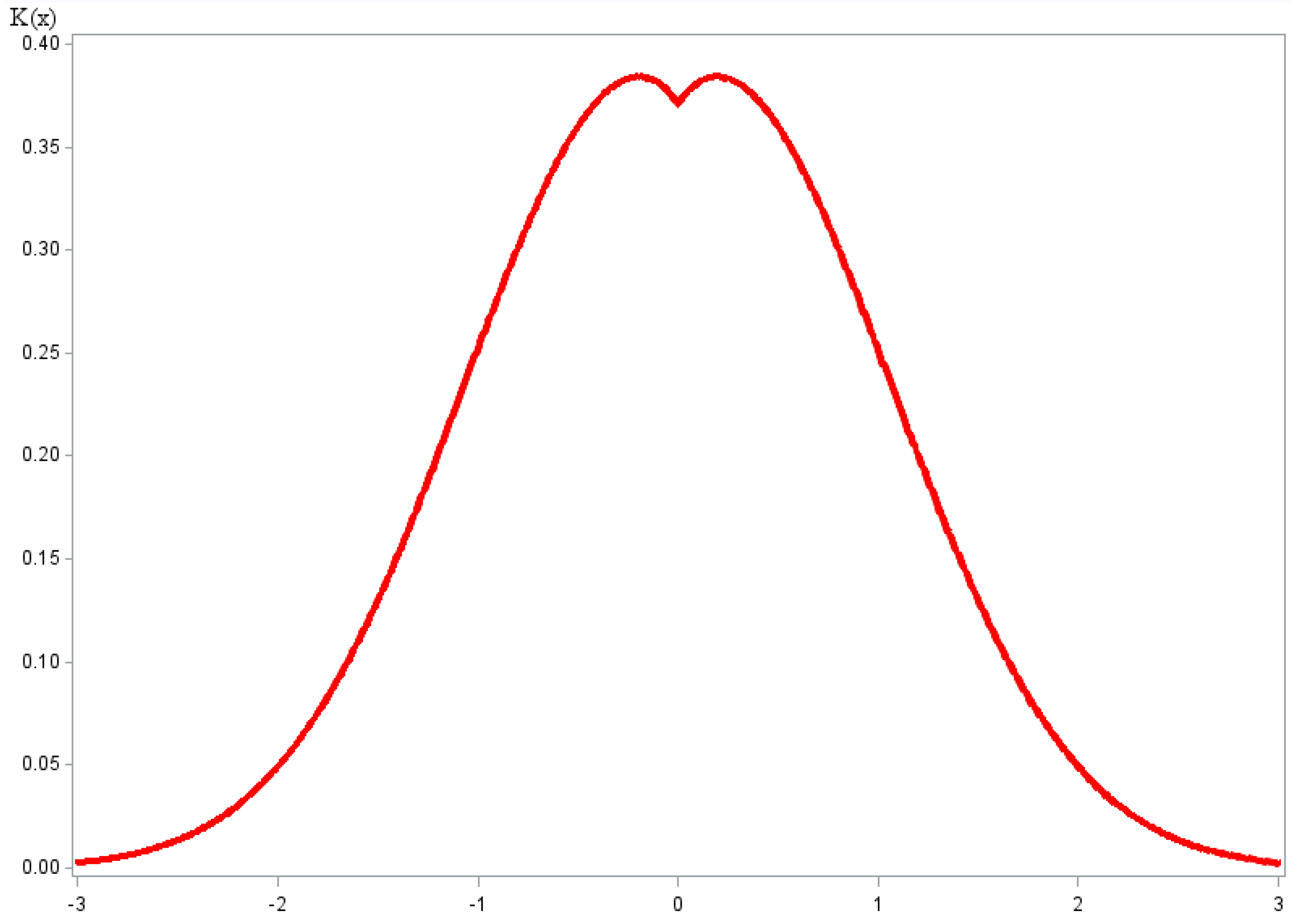
2.31. N-Metric
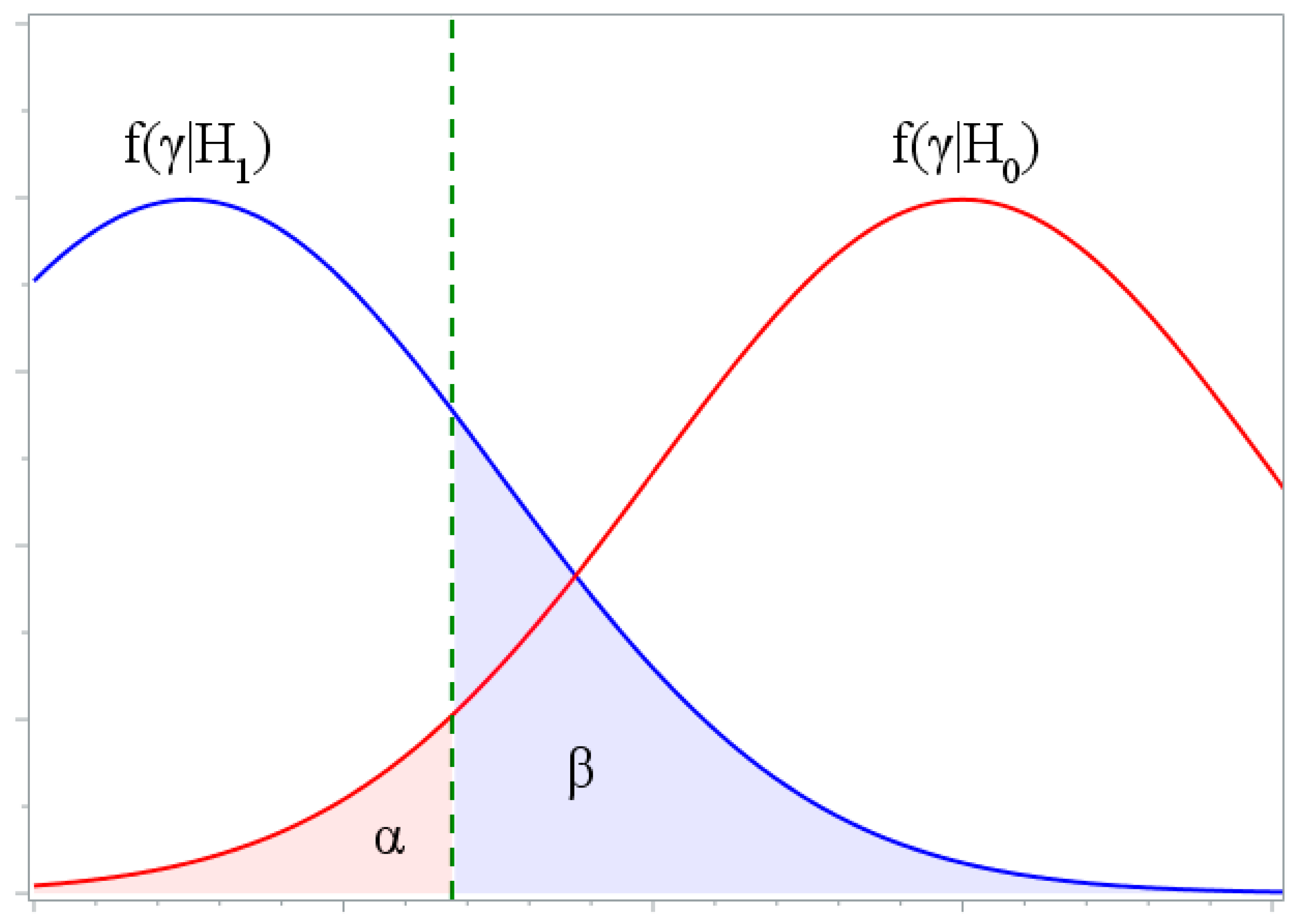
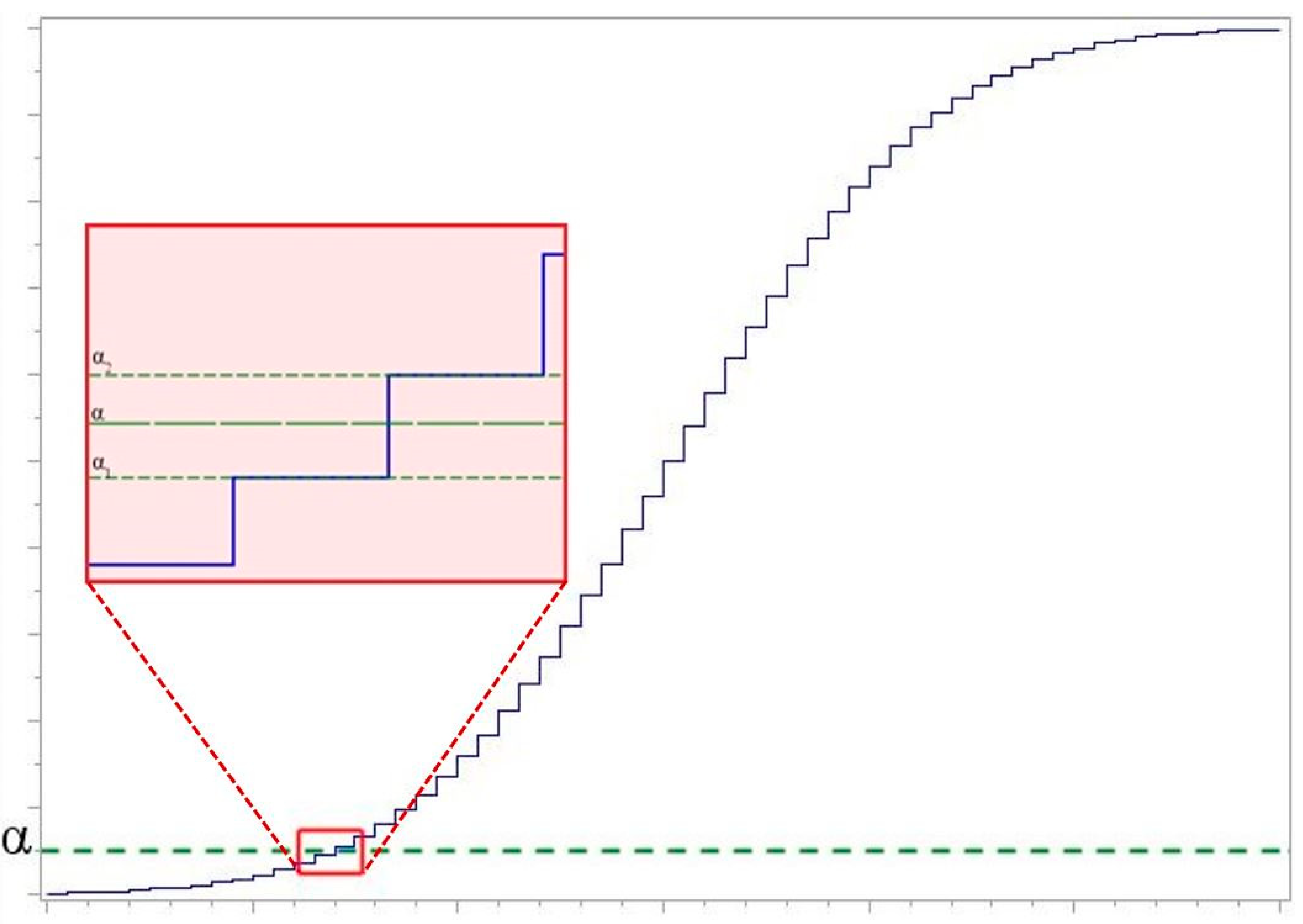
3. The Power of Test
- The distribution of the analyzed data is formed.
- Statistics of the compatibility hypothesis test criteria are calculated. If the obtain value of statistic is greater than the corresponding critical value ( is used), then hypothesis is rejected.
- Steps 1 and 2 are repeated for (in our experiments, ) times.
- The power of a test is calculated as , where is the number of false hypotheses rejections.
4. Statistical Distributions
4.1. Symmetric Distributions
- three cases of the distributionand , where and are the shape parameters;
- three cases of the distribution—, and , where and are the location and scale parameters;
- one case of the distribution, where and are the location and scale parameters;
- one case of the distribution, where and are the location and scale parameters;
- four cases of the distribution and , where is the number of degrees of freedom;
- five cases of the distribution and , where is the shape parameter; and
- one case of the standard normal distribution.
4.2. Asymmetric Distributions
- four cases of the distribution and ;
- four cases of the - distribution, and, where is the number of degrees of freedom;
- six cases of the distribution—, and , where and are the shape and scale parameters;
- one case of the distribution, where and are the location and scale parameters;
- one case of the distribution, where and are the location and scale parameters; and
- four cases of the distribution and , where and are the shape and scale parameters.
4.3. Modified Normal Distributions
- six cases of the standard normal distribution truncated at and and which are referred to as NORMAL1;
- nine cases of a location-contaminated standard normal distribution, hereon termed and , which are referred to as NORMAL2;
- nine cases of a scale-contaminated standard normal distribution, hereon termed and , which are referred to as NORMAL3; and
- twelve cases of a mixture of normal distributions, hereon termed and , which are referred to as NORMAL4.
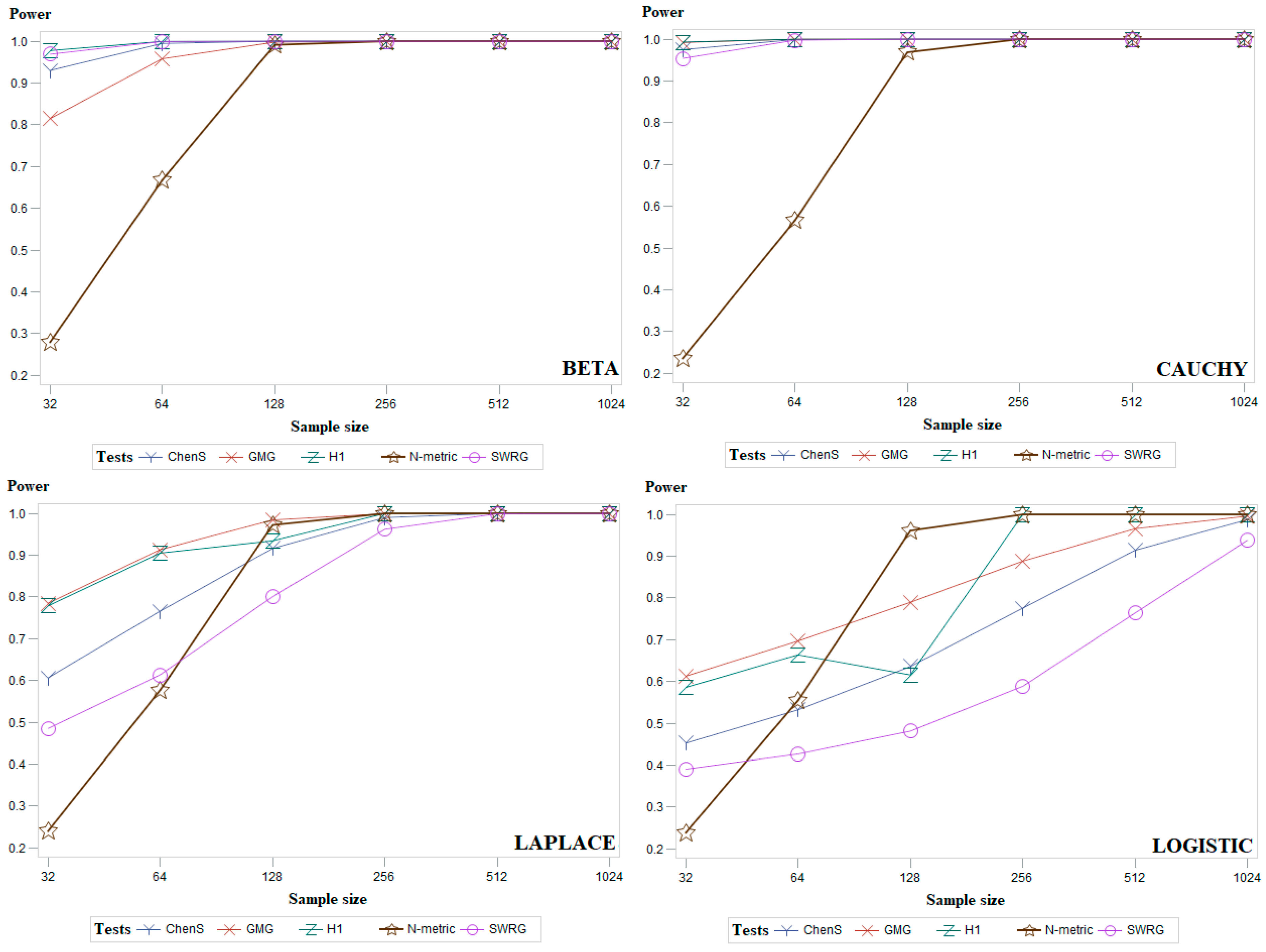
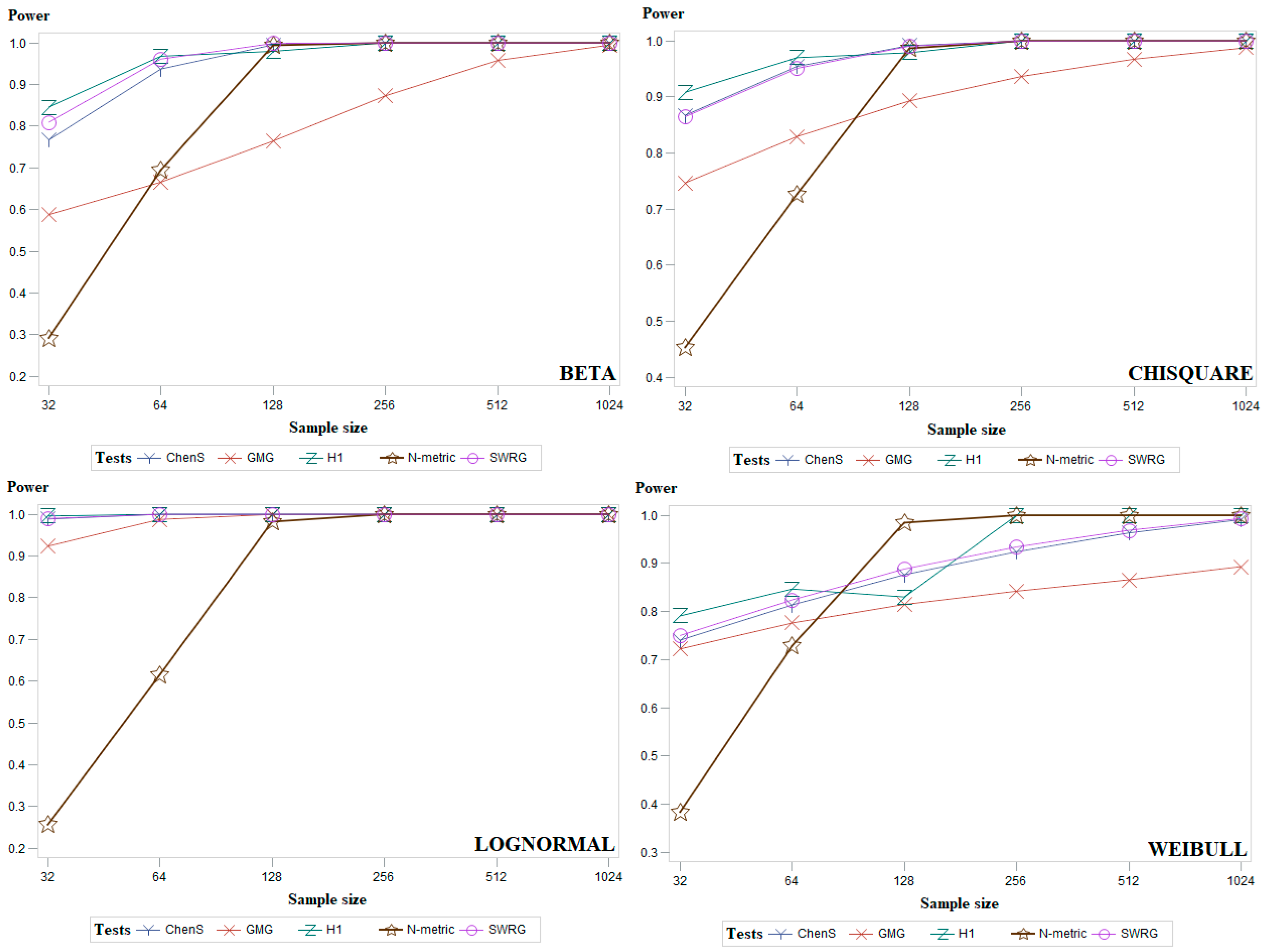
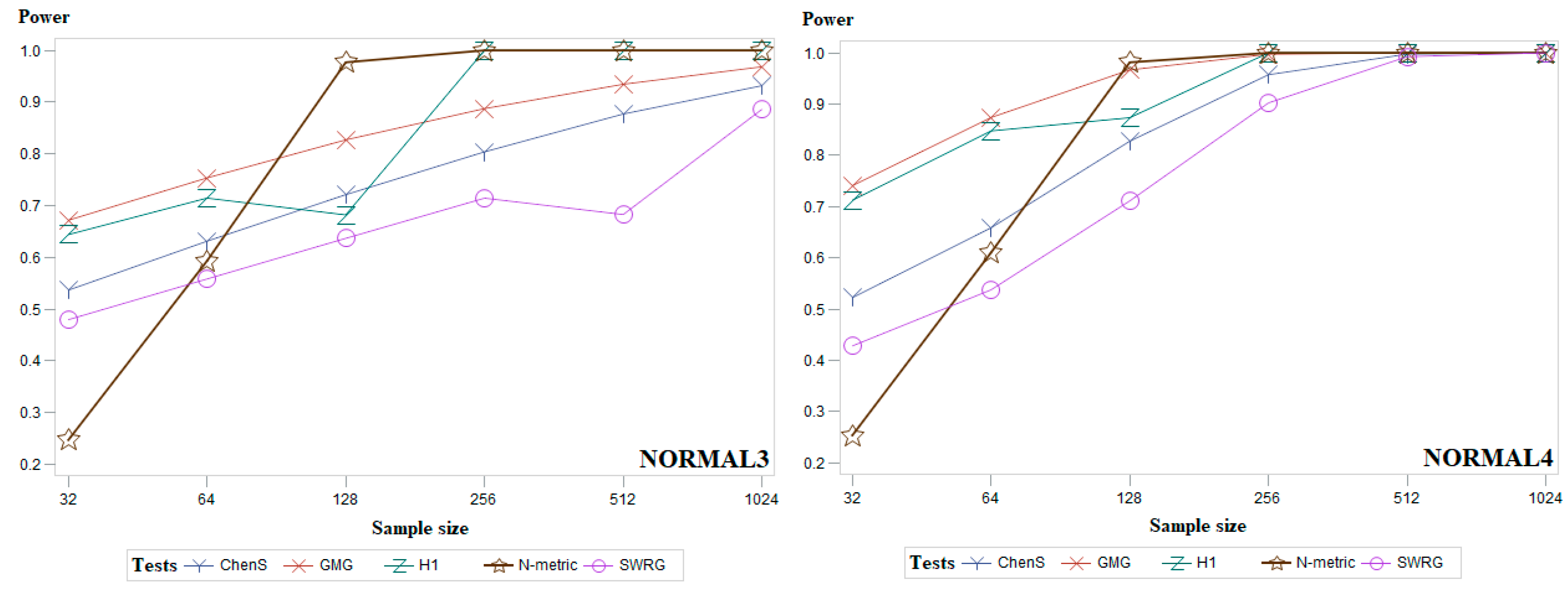
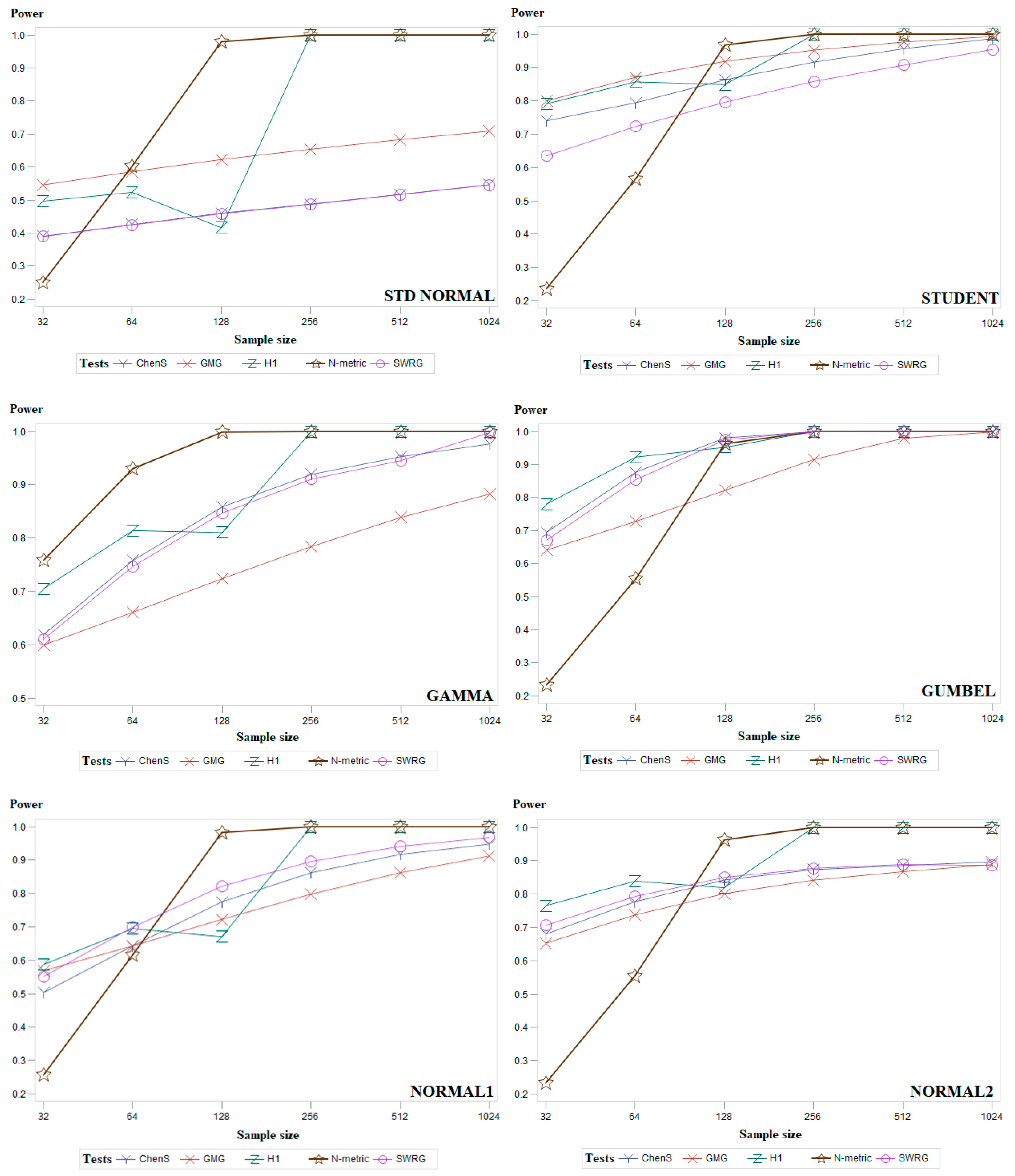
5. Simulation Study and Discussion
6. Conclusions and Future Work
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Appendix A
References
- Barnard, G.A.; Barnard, G.A. Introduction to Pearson (1900) on the Criterion That a Given System of Deviations from the Probable in the Case of a Correlated System of Variables is Such That it Can be Reasonably Supposed to Have Arisen from Random Sampling; Springer Series in Statistics Breakthroughs in Statistics; Springer: Cham, Switzerland, 1992; pp. 1–10. [Google Scholar]
- Kolmogorov, A. Sulla determinazione empirica di una legge di distribuzione. Inst. Ital. Attuari Giorn. 1933, 4, 83–91. [Google Scholar]
- Adefisoye, J.; Golam Kibria, B.; George, F. Performances of several univariate tests of normality: An empirical study. J. Biom. Biostat. 2016, 7, 1–8. [Google Scholar]
- Anderson, T.W.; Darling, D.A. Asymptotic theory of certain “goodness-of-fit” criteria based on stochastic processes. Ann. Math. Stat. 1952, 23, 193–212. [Google Scholar] [CrossRef]
- Hosking, J.R.M.; Wallis, J.R. Some statistics useful in regional frequency analysis. Water Resour. Res. 1993, 29, 271–281. [Google Scholar] [CrossRef]
- Cabana, A.; Cabana, E.M. Goodness-of-Fit and Comparison Tests of the Kolmogorov-Smirnov Type for Bivariate Populations. Ann. Stat. 1994, 22, 1447–1459. [Google Scholar] [CrossRef]
- Chen, L.; Shapiro, S.S. An Alternative Test for Normality Based on Normalized Spacings. J. Stat. Comput. Simul. 1995, 53, 269–288. [Google Scholar] [CrossRef]
- Rahman, M.M.; Govindarajulu, Z. A modification of the test of Shapiro and Wilk for normality. J. Appl. Stat. 1997, 24, 219–236. [Google Scholar] [CrossRef]
- Ray, W.D.; Shenton, L.R.; Bowman, K.O. Maximum Likelihood Estimation in Small Samples. J. R. Stat. Soc. Ser. A 1978, 141, 268. [Google Scholar] [CrossRef]
- Zhang, P. Omnibus test of normality using the Q statistic. J. Appl. Stat. 1999, 26, 519–528. [Google Scholar] [CrossRef]
- Barrio, E.; Cuesta-Albertos, J.A.; Matrán, C.; Rodríguez-Rodríguez, J.M. Tests of goodness of fit based on the L2-Wasserstein distance. Ann. Stat. 1999, 27, 1230–1239. [Google Scholar]
- Glen, A.G.; Leemis, L.M.; Barr, D.R. Order statistics in goodness-of-fit testing. IEEE Trans. Reliab. 2001, 50, 209–213. [Google Scholar] [CrossRef]
- Bonett, D.G.; Seier, E. A test of normality with high uniform power. Comput. Stat. Data Anal. 2002, 40, 435–445. [Google Scholar] [CrossRef]
- Psaradakis, Z.; Vávra, M. Normality tests for dependent data: Large-sample and bootstrap approaches. Commun. Stat.-Simul. Comput. 2018, 49, 283–304. [Google Scholar] [CrossRef]
- Zhang, J.; Wu, Y. Likelihood-ratio tests for normality. Comput. Stat. Data Anal. 2005, 49, 709–721. [Google Scholar] [CrossRef]
- Gel, Y.R.; Miao, W.; Gastwirth, J.L. Robust directed tests of normality against heavy-tailed alternatives. Comput. Stat. Data Anal. 2007, 51, 2734–2746. [Google Scholar] [CrossRef]
- Coin, D. A goodness-of-fit test for normality based on polynomial regression. Comput. Stat. Data Anal. 2008, 52, 2185–2198. [Google Scholar] [CrossRef]
- Desgagné, A.; Lafaye de Micheaux, P. A powerful and interpretable alternative to the Jarque–Bera test of normality based on 2nd-power skewness and kurtosis, using the Rao’s score test on the APD family. J. Appl. Stat. 2017, 45, 2307–2327. [Google Scholar] [CrossRef]
- Steele, C.M. The Power of Categorical Goodness-Of-Fit Statistics. Ph.D. Thesis, Australian School of Environmental Studies, Warrandyte, Victoria, Australia, 2003. [Google Scholar]
- Romão, X.; Delgado, R.; Costa, A. An empirical power comparison of univariate goodness-of-fit tests for normality. J. Stat. Comput. Simul. 2010, 80, 545–591. [Google Scholar] [CrossRef]
- Choulakian, V.; Lockhart, R.; Stephens, M. Cramérvon Mises statistics for discrete distributions. Can. J. Stat. 1994, 22, 125–137. [Google Scholar] [CrossRef]
- Shapiro, S.S.; Wilk, M.B. An analysis of variance test for normality (complete samples). Biometrika 1965, 52, 591–611. [Google Scholar] [CrossRef]
- Lilliefors, H.W. On the Kolmogorov-Smirnov test for normality with mean and variance unknown. J. Am. Stat. Assoc. 1967, 62, 399–402. [Google Scholar] [CrossRef]
- Ahmad, F.; Khan, R.A. A power comparison of various normality tests. Pak. J. Stat. Oper. Res. 2015, 11, 331. [Google Scholar] [CrossRef]
- D’Agostino, R.B.; Pearson, E.S. Testing for departures from normality. I. Fuller empirical results for the distribution of b2 and √b1. Biometrika 1973, 60, 613–622. [Google Scholar]
- Filliben, J.J. The Probability Plot Correlation Coefficient Test for Normality. Technometrics 1975, 17, 111–117. [Google Scholar] [CrossRef]
- Martinez, J.; Iglewicz, B. A test for departure from normality based on a biweight estimator of scale. Biometrika 1981, 68, 331–333. [Google Scholar] [CrossRef]
- Epps, T.W.; Pulley, L.B. A test for normality based on the empirical characteristic function. Biometrika 1983, 70, 723–726. [Google Scholar] [CrossRef]
- Jarque, C.; Bera, A. Efficient tests for normality, homoscedasticity andserial independence of regression residuals. Econ. Lett. 1980, 6, 255–259. [Google Scholar] [CrossRef]
- Bakshaev, A. Goodness of fit and homogeneity tests on the basis of N-distances. J. Stat. Plan. Inference 2009, 139, 3750–3758. [Google Scholar] [CrossRef]
- Hill, T.; Lewicki, P. Statistics Methods and Applications; StatSoft: Tulsa, OK, USA, 2007. [Google Scholar]
- Kasiulevičius, V.; Denapienė, G. Statistikos taikymas mokslinių tyrimų analizėje. Gerontologija 2008, 9, 176–180. [Google Scholar]
- Damianou, C.; Kemp, A.W. New goodness of statistics for discrete and continuous data. Am. J. Math. Manag. Sci. 1990, 10, 275–307. [Google Scholar]
| Sample Size | |||||||
|---|---|---|---|---|---|---|---|
| 32 | 64 | 128 | 256 | 512 | 1024 | ||
| Tests | AD | 0.714 | 0.799 | 0.863 | 0.909 | 0.939 | 0.955 |
| BCMR | 0.718 | 0.809 | 0.875 | 0.920 | 0.947 | 0.947 | |
| BHS | 0.431 | 0.551 | 0.663 | 0.752 | 0.818 | 0.868 | |
| BHSBS | 0.680 | 0.778 | 0.783 | 0.903 | 0.938 | 0.959 | |
| BM2 | 0.726 | 0.835 | 0.905 | 0.945 | 0.965 | 0.974 | |
| BS | 0.717 | 0.810 | 0.877 | 0.920 | 0.947 | 0.961 | |
| CC2 | 0.712 | 0.805 | 0.873 | 0.920 | 0.949 | 0.936 | |
| CHI2 | 0.663 | 0.778 | 0.842 | 0.884 | 0.941 | 0.945 | |
| CVM | 0.591 | 0.733 | 0.805 | 0.855 | 0.919 | 0.949 | |
| ChenS | 0.729 | 0.806 | 0.871 | 0.915 | 0.943 | 0.960 | |
| Coin | 0.735 | 0.830 | 0.891 | 0.930 | 0.952 | 0.963 | |
| DA | 0.266 | 0.295 | 0.314 | 0.319 | 0.315 | 0.311 | |
| DAP | 0.723 | 0.820 | 0.883 | 0.924 | 0.948 | 0.962 | |
| DH | 0.709 | 0.805 | 0.877 | 0.925 | 0.950 | 0.963 | |
| DLDMZEPD | 0.730 | 0.826 | 0.889 | 0.929 | 0.952 | 0.963 | |
| EP | 0.706 | 0.828 | 0.974 | 0.910 | 0.946 | 0.959 | |
| Filli | 0.712 | 0.805 | 0.875 | 0.922 | 0.949 | 0.962 | |
| GG | 0.658 | 0.760 | 0.850 | 0.915 | 0.949 | 0.962 | |
| GLB | 0.712 | 0.798 | 0.863 | 0.909 | 0.943 | 0.918 | |
| GMG | 0.787 | 0.862 | 0.914 | 0.946 | 0.965 | 0.975 | |
| H1 | 0.799 | 0.862 | 0.852 | 0.999 | 0.999 | 0.999 | |
| JB | 0.643 | 0.762 | 0.856 | 0.918 | 0.949 | 0.963 | |
| KS | 0.585 | 0.723 | 0.789 | 0.836 | 0.905 | 0.939 | |
| Lillie | 0.669 | 0.758 | 0.828 | 0.883 | 0.921 | 0.947 | |
| MI | 0.632 | 0.676 | 0.705 | 0.724 | 0.736 | 0.745 | |
| N-metric | 0.245 | 0.585 | 0.971 | 0.999 | 0.999 | 0.999 | |
| SF | 0.715 | 0.807 | 0.876 | 0.923 | 0.949 | 0.962 | |
| SW | 0.718 | 0.808 | 0.874 | 0.919 | 0.946 | 0.962 | |
| SWRG | 0.694 | 0.775 | 0.834 | 0.882 | 0.916 | 0.946 | |
| ZQstar | 0.513 | 0.576 | 0.630 | 0.669 | 0.697 | 0.718 | |
| ZW2 | 0.715 | 0.806 | 0.869 | 0.912 | 0.939 | 0.957 | |
| Sample Size | |||||||
|---|---|---|---|---|---|---|---|
| 32 | 64 | 128 | 256 | 512 | 1024 | ||
| Tests | AD | 0.729 | 0.835 | 0.908 | 0.949 | 0.969 | 0.984 |
| BCMR | 0.749 | 0.856 | 0.924 | 0.971 | 0.995 | 0.991 | |
| BHS | 0.529 | 0.664 | 0.769 | 0.855 | 0.915 | 0.950 | |
| BHSBS | 0.538 | 0.652 | 0.747 | 0.914 | 0.902 | 0.944 | |
| BM2 | 0.737 | 0.859 | 0.931 | 0.965 | 0.981 | 0.993 | |
| BS | 0.506 | 0.588 | 0.665 | 0.738 | 0.805 | 0.859 | |
| CC2 | 0.579 | 0.682 | 0.777 | 0.853 | 0.938 | 0.956 | |
| CHI2 | 0.645 | 0.799 | 0.881 | 0.934 | 0.965 | 0.980 | |
| CVM | 0.594 | 0.755 | 0.836 | 0.887 | 0.935 | 0.957 | |
| ChenS | 0.756 | 0.862 | 0.928 | 0.961 | 0.978 | 0.991 | |
| Coin | 0.480 | 0.556 | 0.630 | 0.700 | 0.769 | 0.916 | |
| DA | 0.237 | 0.223 | 0.209 | 0.198 | 0.191 | 0.192 | |
| DAP | 0.705 | 0.826 | 0.910 | 0.955 | 0.977 | 0.990 | |
| DH | 0.724 | 0.845 | 0.921 | 0.957 | 0.977 | 0.991 | |
| DLDMXAPD | 0.726 | 0.843 | 0.918 | 0.955 | 0.975 | 0.989 | |
| EP | 0.753 | 0.846 | 0.913 | 0.967 | 0.975 | 0.993 | |
| Filli | 0.732 | 0.842 | 0.915 | 0.953 | 0.974 | 0.991 | |
| GG | 0.672 | 0.805 | 0.898 | 0.949 | 0.973 | 0.988 | |
| GLB | 0.725 | 0.831 | 0.905 | 0.987 | 0.970 | 0.984 | |
| GMG | 0.683 | 0.751 | 0.809 | 0.859 | 0.901 | 0.932 | |
| H1 | 0.816 | 0.896 | 0.896 | 0.999 | 0.999 | 0.999 | |
| JB | 0.662 | 0.808 | 0.904 | 0.953 | 0.975 | 0.989 | |
| KS | 0.582 | 0.736 | 0.810 | 0.863 | 0.921 | 0.945 | |
| Lillie | 0.671 | 0.786 | 0.872 | 0.929 | 0.959 | 0.976 | |
| MI | 0.644 | 0.731 | 0.798 | 0.843 | 0.872 | 0.913 | |
| N-metric | 0.464 | 0.761 | 0.990 | 0.999 | 0.999 | 0.999 | |
| SF | 0.736 | 0.846 | 0.918 | 0.955 | 0.975 | 0.989 | |
| SW | 0.753 | 0.859 | 0.925 | 0.959 | 0.977 | 0.991 | |
| SWRG | 0.758 | 0.861 | 0.927 | 0.960 | 0.977 | 0.999 | |
| ZQstar | 0.570 | 0.639 | 0.693 | 0.732 | 0.761 | 0.748 | |
| ZW2 | 0.764 | 0.870 | 0.932 | 0.962 | 0.980 | 0.997 | |
| Sample Size | |||||||
|---|---|---|---|---|---|---|---|
| 32 | 64 | 128 | 256 | 512 | 1024 | ||
| Tests | AD | 0.662 | 0.756 | 0.825 | 0.872 | 0.905 | 0.931 |
| BCMR | 0.652 | 0.756 | 0.831 | 0.880 | 0.913 | 0.935 | |
| BHS | 0.463 | 0.585 | 0.676 | 0.744 | 0.796 | 0.834 | |
| BHSBS | 0.568 | 0.701 | 0.787 | 0.847 | 0.890 | 0.918 | |
| BM2 | 0.641 | 0.770 | 0.854 | 0.904 | 0.934 | 0.953 | |
| BS | 0.587 | 0.688 | 0.770 | 0.833 | 0.881 | 0.916 | |
| CC2 | 0.576 | 0.675 | 0.763 | 0.833 | 0.887 | 0.923 | |
| CHI2 | 0.566 | 0.728 | 0.808 | 0.866 | 0.914 | 0.939 | |
| CVM | 0.557 | 0.708 | 0.779 | 0.833 | 0.897 | 0.930 | |
| ChenS | 0.656 | 0.759 | 0.833 | 0.882 | 0.915 | 0.937 | |
| Coin | 0.579 | 0.691 | 0.781 | 0.846 | 0.889 | 0.918 | |
| DA | 0.314 | 0.342 | 0.367 | 0.388 | 0.405 | 0.418 | |
| DAP | 0.617 | 0.733 | 0.818 | 0.872 | 0.906 | 0.930 | |
| DH | 0.617 | 0.727 | 0.815 | 0.872 | 0.907 | 0.930 | |
| DLDMXAPD | 0.651 | 0.754 | 0.831 | 0.879 | 0.912 | 0.935 | |
| EP | 0.640 | 0.748 | 0.819 | 0.865 | 0.906 | 0.931 | |
| Filli | 0.637 | 0.743 | 0.823 | 0.877 | 0.911 | 0.933 | |
| GG | 0.529 | 0.657 | 0.775 | 0.860 | 0.906 | 0.932 | |
| GLB | 0.659 | 0.755 | 0.823 | 0.870 | 0.903 | 0.930 | |
| GMG | 0.688 | 0.771 | 0.836 | 0.883 | 0.917 | 0.942 | |
| H1 | 0.743 | 0.816 | 0.799 | 0.999 | 0.999 | 0.999 | |
| JB | 0.515 | 0.662 | 0.783 | 0.861 | 0.904 | 0.930 | |
| KS | 0.564 | 0.710 | 0.772 | 0.825 | 0.893 | 0.924 | |
| Lillie | 0.626 | 0.724 | 0.796 | 0.850 | 0.889 | 0.917 | |
| MI | 0.494 | 0.536 | 0.563 | 0.578 | 0.585 | 0.590 | |
| N-metric | 0.243 | 0.582 | 0.972 | 0.999 | 0.999 | 0.999 | |
| SF | 0.642 | 0.747 | 0.826 | 0.879 | 0.912 | 0.934 | |
| SW | 0.654 | 0.758 | 0.832 | 0.882 | 0.915 | 0.937 | |
| SWRG | 0.643 | 0.746 | 0.818 | 0.864 | 0.901 | 0.931 | |
| ZQstar | 0.394 | 0.423 | 0.450 | 0.472 | 0.487 | 0.498 | |
| ZW2 | 0.640 | 0.749 | 0.826 | 0.876 | 0.907 | 0.931 | |
| Nr. | Distribution | Groups of Distributions | |
|---|---|---|---|
| 1. | Standard normal | Symmetric | 46 |
| 2. | Beta | Symmetric | 88 |
| 3. | Cauchy | Symmetric | 257 |
| 4. | Laplace | Symmetric | 117 |
| 5. | Logistic | Symmetric | 71 |
| 6. | Student | Symmetric | 96 |
| 7. | Beta | Asymmetric | 108 |
| 8. | Chi-square | Asymmetric | 123 |
| 9. | Gamma | Asymmetric | <32 |
| 10. | Gumbel | Asymmetric | 125 |
| 11. | Lognormal | Asymmetric | 255 |
| 12. | Weibull | Asymmetric | 65 |
| 13. | Normal1 | Modified normal | 70 |
| 14. | Normal2 | Modified normal | 93 |
| 15. | Normal3 | Modified normal | 72 |
| 16. | Normal4 | Modified normal | 117 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Arnastauskaitė, J.; Ruzgas, T.; Bražėnas, M. An Exhaustive Power Comparison of Normality Tests. Mathematics 2021, 9, 788. https://doi.org/10.3390/math9070788
Arnastauskaitė J, Ruzgas T, Bražėnas M. An Exhaustive Power Comparison of Normality Tests. Mathematics. 2021; 9(7):788. https://doi.org/10.3390/math9070788
Chicago/Turabian StyleArnastauskaitė, Jurgita, Tomas Ruzgas, and Mindaugas Bražėnas. 2021. "An Exhaustive Power Comparison of Normality Tests" Mathematics 9, no. 7: 788. https://doi.org/10.3390/math9070788
APA StyleArnastauskaitė, J., Ruzgas, T., & Bražėnas, M. (2021). An Exhaustive Power Comparison of Normality Tests. Mathematics, 9(7), 788. https://doi.org/10.3390/math9070788







