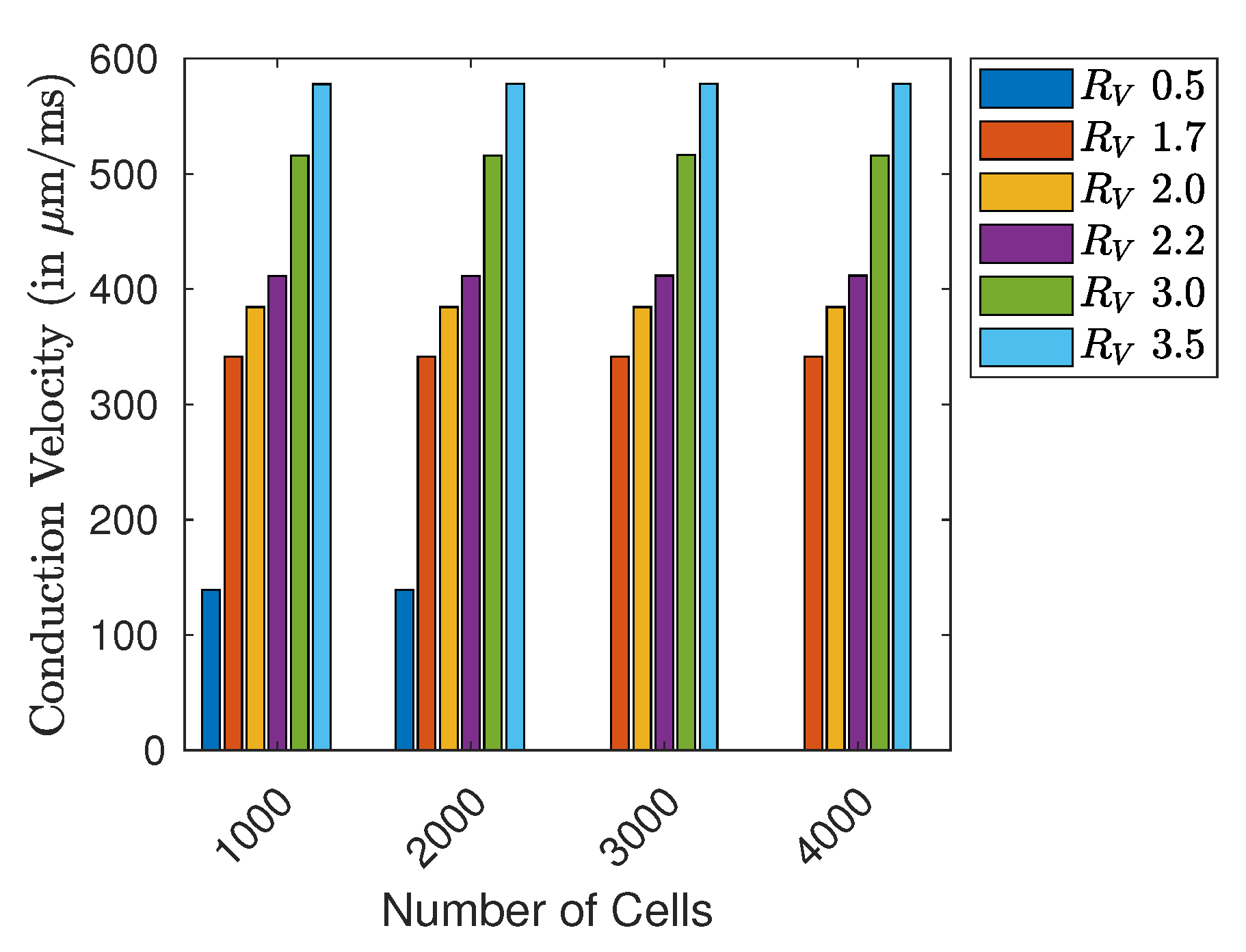1. Introduction
Computational cardiology provides a safe, non-invasive, and ethical method to investigate the heart and its dysfunctionalities. Early mathematical models [
1,
2,
3,
4] extend the pioneering work of Hodgkin-Huxley [
5] to describe and explain, with a set of ordinary differential equations (ODEs), the ionic mechanisms responsible for the initiation and the electrical propagation of action potentials traversing excitable cells such as cardiac myocytes and neurons [
6,
7]. These early studies enabled innovative in-silico research and clinically oriented applications [
8]. For example, they allow us to understand a variety of nonlinear phenomena affecting cardiac dynamics [
9,
10,
11,
12,
13] and unexpected, chaotic properties of the heart [
14,
15]. The mathematical complexity of such models has stimulated several works [
16,
17,
18] trying to improve the analysis by simplifying the model representation.
Cellular Automata (CA) are a well-known class of discrete simplified dynamical systems widely used to reproduce pattern formation and growth [
19,
20]. One of the first approaches using CA and its theory was proposed by von Neumann et al. [
21]. A cellular automaton consists of a regular grid of cells where a discrete state is associated to each cell. The state of each cell can evolve in time according to a set of rules operating over the states of the neighboring cells. This class of models provides a simple mathematical framework to reproduce several emergent behaviors that can be observed in excitable media such as the cardiac tissue [
22] and the nervous system [
23]. Other extensions of CA allow one to customize the model describing how each cell reacts to the external stimuli from the neighboring cells. For example, the electrophysiological process in a myocyte is driven by a set of voltage-dependent thresholds that control the opening/closing of ionic channels regulating the flux of ions across the cellular membrane. These thresholds generally identify specific phases of the myocyte’s behavior, such as the rapid depolarization, the initial repolarization, the plateau, and the late repolarization/resting phase, characterized by different dynamics.
A well-established approach to describe such processes is via Hybrid Automata [
24,
25,
26], a modeling formalism that extends finite automata with a set of ODEs describing the continuous behavior of the action potential at each phase (a phase corresponds to a discrete state in the finite automata). A set of guard conditions over voltage-dependent thresholds determine how each cell switches the continuous dynamics that modeling the inward/outward flux of ionic currents at each phase. Previous works [
24,
25,
27] have shown that HA can accurately reproduce the action potential occurring in myocytes. HA can be simulated using numerical integration, but they are also amenable to formal analysis techniques such as model checking [
18,
28]. HA and CA can be then combined in a Hybrid Cellular Automata (HCA) that can model both the
reaction of each cell to external stimuli and the
diffusion of the ionic currents among neighboring cells.
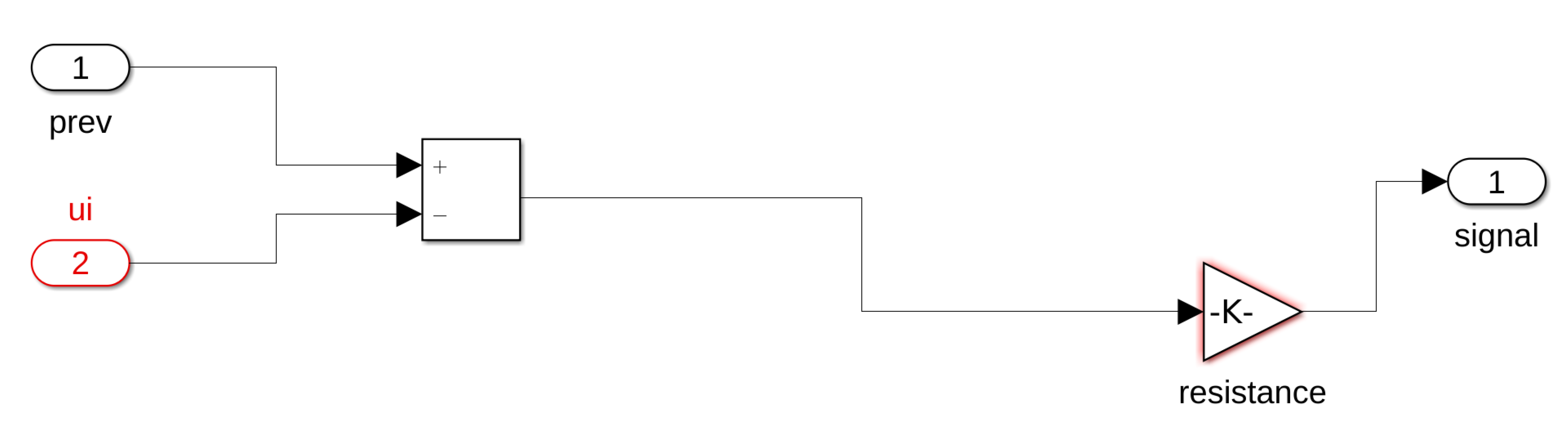
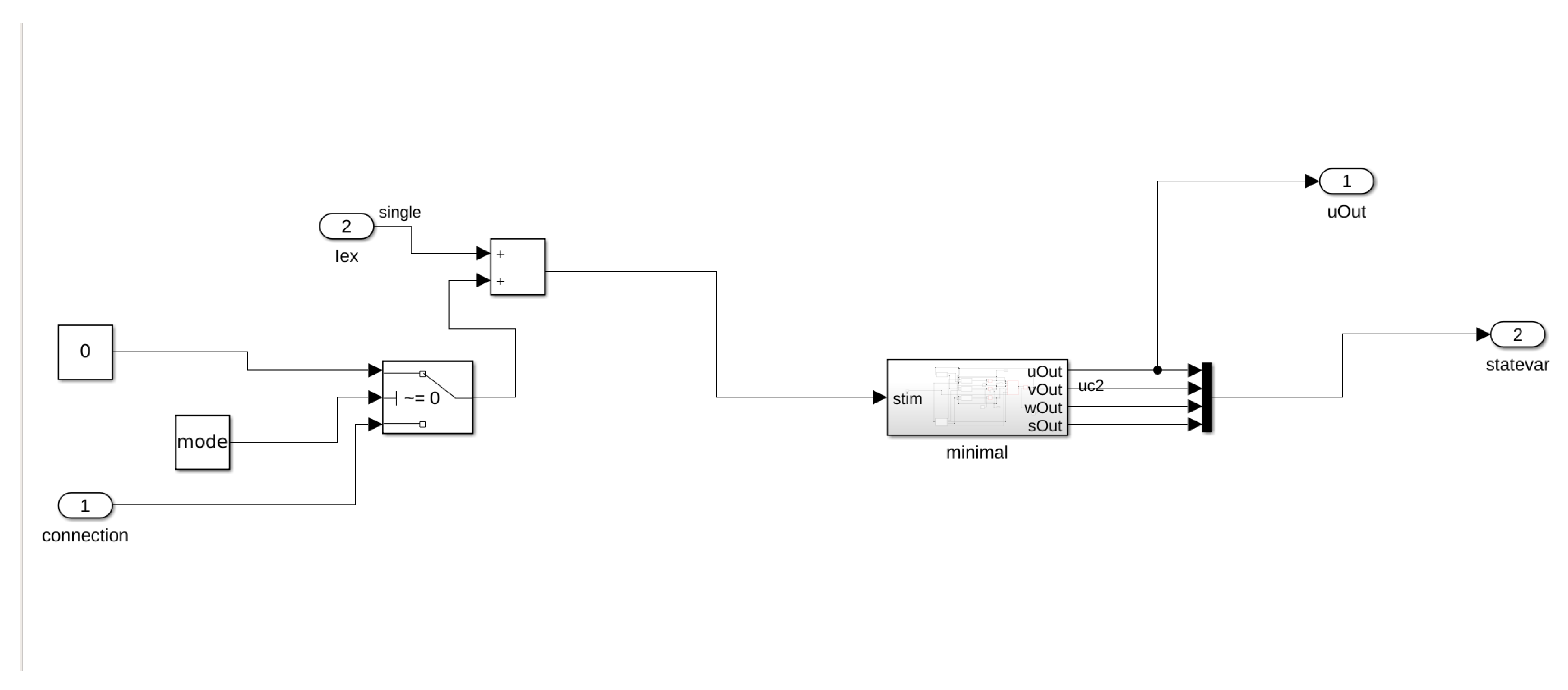
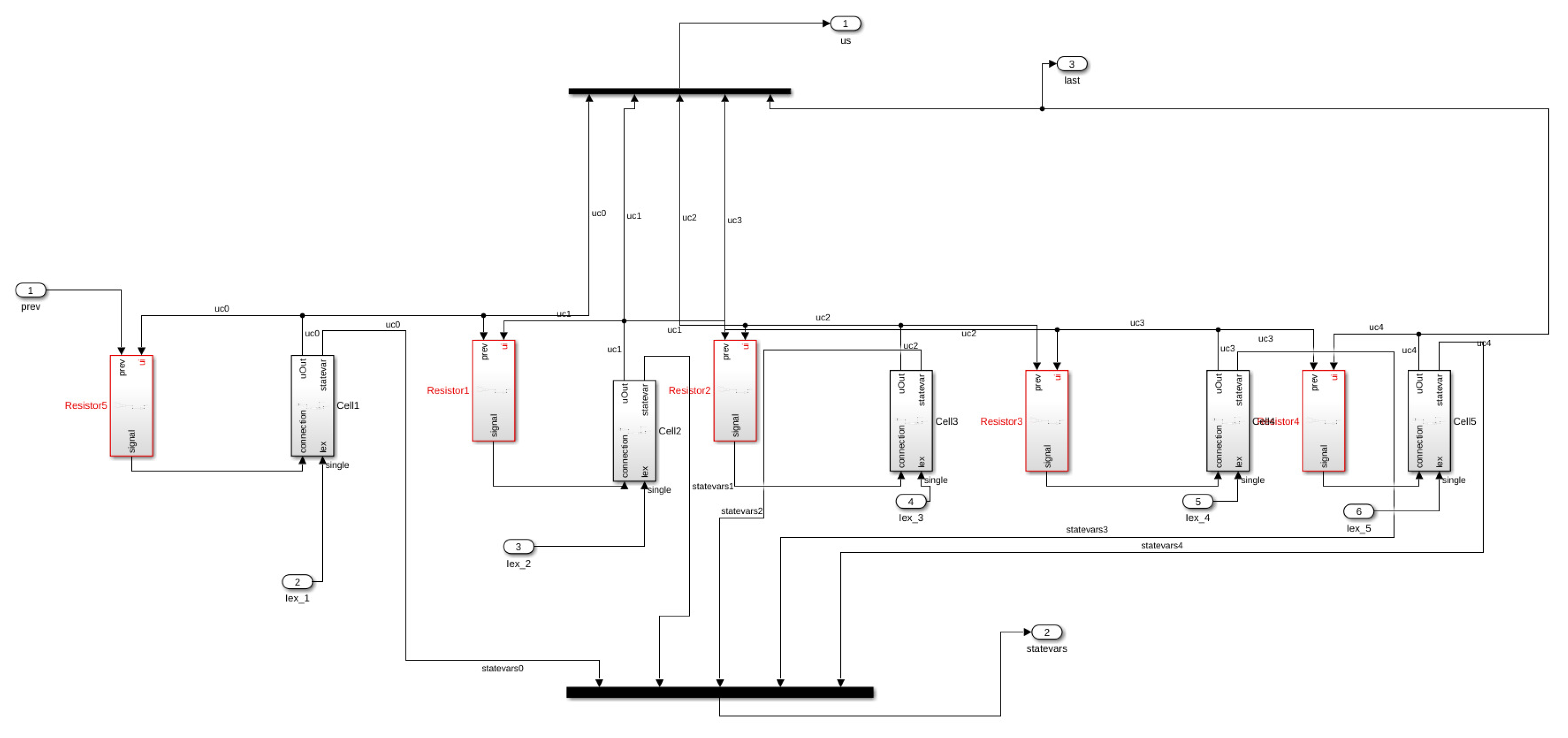
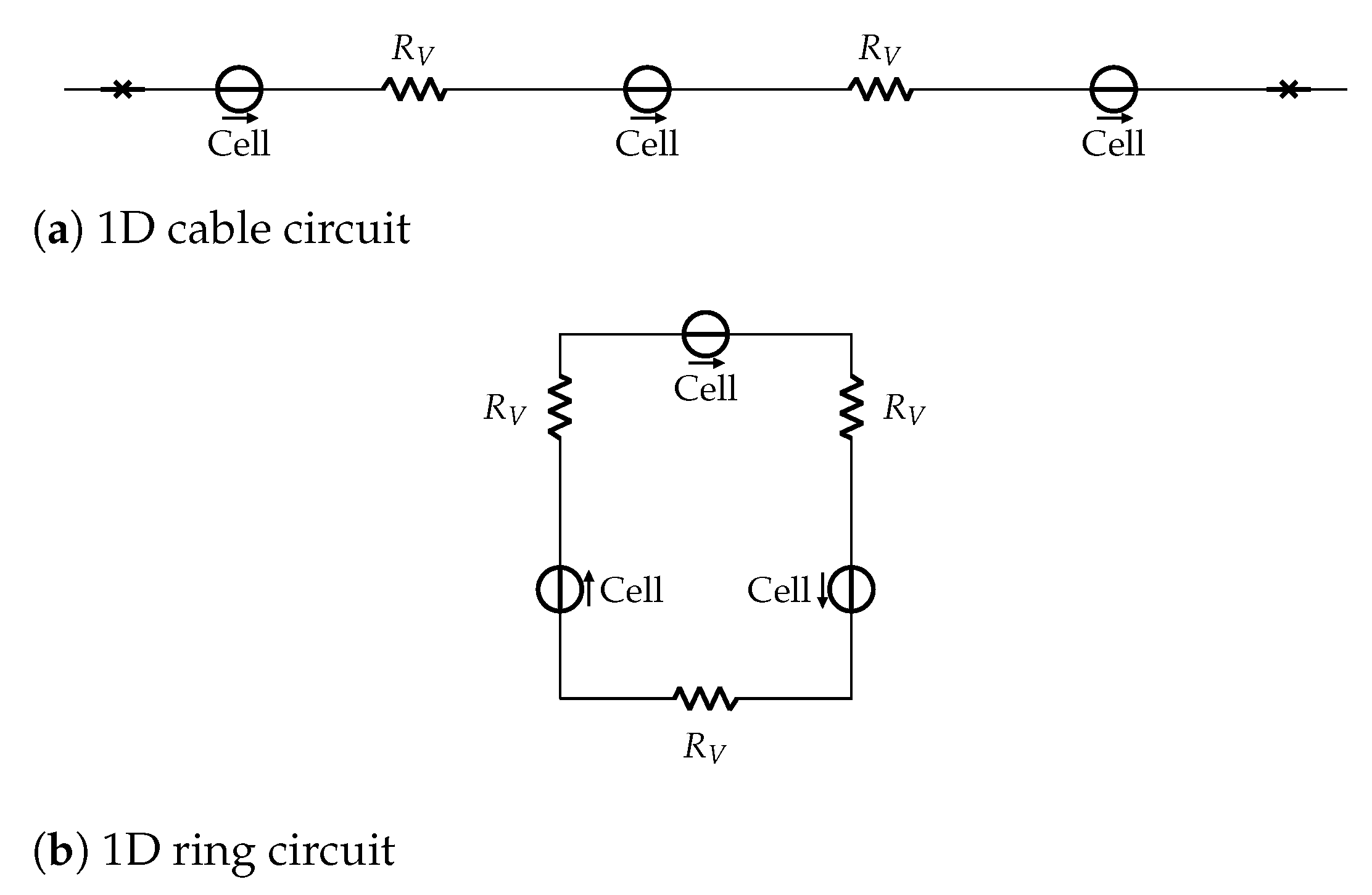
In this work, we consider an HCA model where the single cell behavior is described by a simplified but very accurate four-variables hybrid model proposed in [
16] and the cells are coupled with a simple link (resistance) to propagate signals as a unidirectional cable or as a ring of cardiac cells. The considered phenomenological modeling approach stems from experimental works on one-dimensional rings of cardiac cells and whole heart models studying the onset of alternans and irregular rhythms [
29,
30,
31].
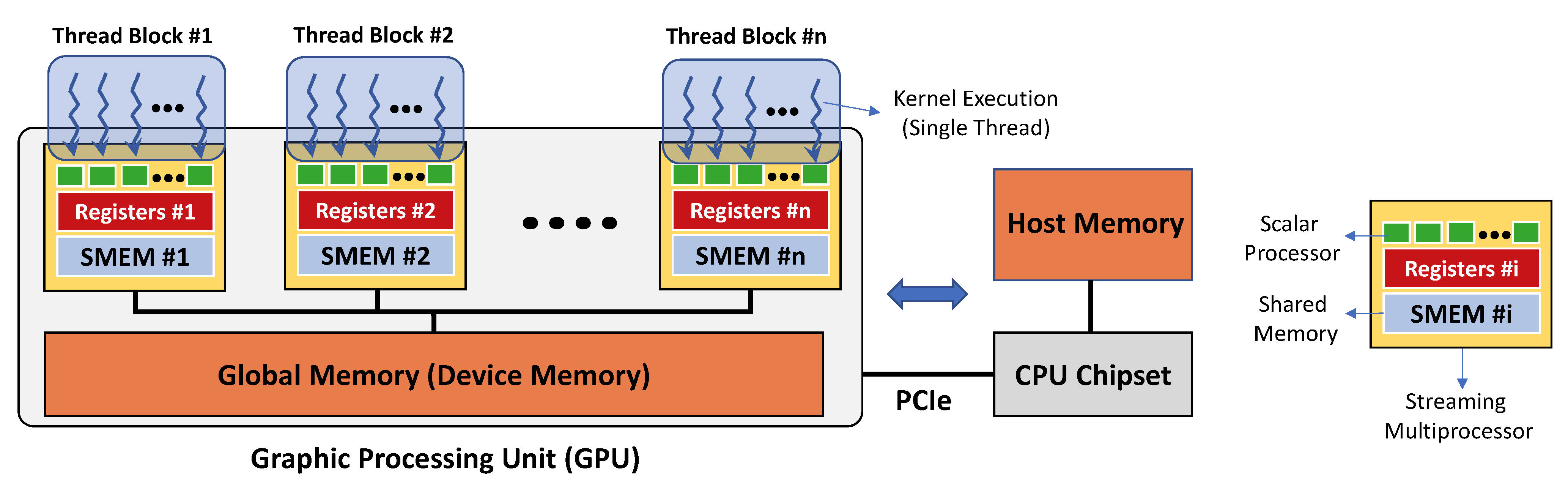
In addition to modeling, one can reduce simulation time and associated computational costs using special hardware accelerators such as Graphical Processing Units (GPU) [
32,
33,
34] and Field Programmable Gate Arrays (FPGA) [
35,
36]. To optimally simulate our cardiac cell model, we compare common numerical methods in combination with different coding approaches: Matlab
®Simulink, a sequential C implementation running on a CPU and an NVIDIA
®CUDA-based implementation running on a modern GPU. Such a comparison is a required step to evaluate the limitations and trade-offs of the different technologies as well as the reliability of our modeling approach.
The literature is populated by a multitude of physiological and phenomenological mathematical models of cardiac cell electrophysiology [
37]. These dynamical systems are usually analyzed in terms of local restitution features (e.g., changes of the action potential duration versus decreasing stimulation pacing periods) as well as bifurcation diagrams (when period-doubling occurs) and space-time visualization [
38,
39,
40,
41,
42]. Like the biological original, models depend on a variety of additional parameters, i.e., including temperature, membrane resistance, and nonlinearities within the tissue [
11,
43,
44,
45].
One-dimensional cables and rings of cardiac cells fulfill standard physiology studies on reentry circuits in cardiology [
46,
47]. Monolayers of cardiomyocytes [
29], as well as multiorgan structures [
30], introduced a fine control of system’s dynamics improving our understanding of complex neurocardiac diseases. Due to certain pathological conditions, such as ischemia, current sinks might develop. This behavior can be simulated by changing the amount of heterogeneity within the tissue and the coupling between cells [
48]. In this perspective, we will conduct an extended analysis investigating emergent phenomena associated with alternans and conduction blocks by using bifurcation diagrams and modeling the cell-cell coupling with a random variable. First, we investigate the propagation and benchmarks of our model, with respect to conduction velocity (CV) and action potential duration (APD). Then we analyze the resiliency of the proposed HCA by decreasing the length (cell count) within the ring structure after a freely chosen amount of time-steps. Doing so, we achieve shortening and spatiotemporal adaptation of the excitation wave, without the need of electrical pacing, also observing the onset of electrical alternans.
The final object of the paper is to assess the proposed HCA approach under different implementations. We do so by investigating the results of a Monte Carlo simulation analysis. As we are using a free resistance variable, we test the robustness of the approach using random values for this variable. We then use a numerical analysis of the bifurcation over APD in a decreasing number of cells. This study allows us to draw conclusions on usability, deficiencies, and further improvement of HCA as a cardiac modeling technique.
The paper is structured as follows: we first present the considered computational models and analysis techniques in
Section 2. In
Section 3, we summarize the main findings of our computational experiments, and we compare the performance of different model implementations. We discuss our results in
Section 4. We conclude in
Section 5 by discussing limitations and perspectives of our work.
4. Discussion
We presented a simplified phenomenological cardiac model based on a Hybrid Cellular Automata formulation by introducing the cell-cell coupling membrane resistance as a free variable. This approach enables us to thoroughly investigate the CV and APD features in the presence of distributed conductivity thus linked to excitation wave spatiotemporal organization and alternans onset [
55,
56,
57,
58].
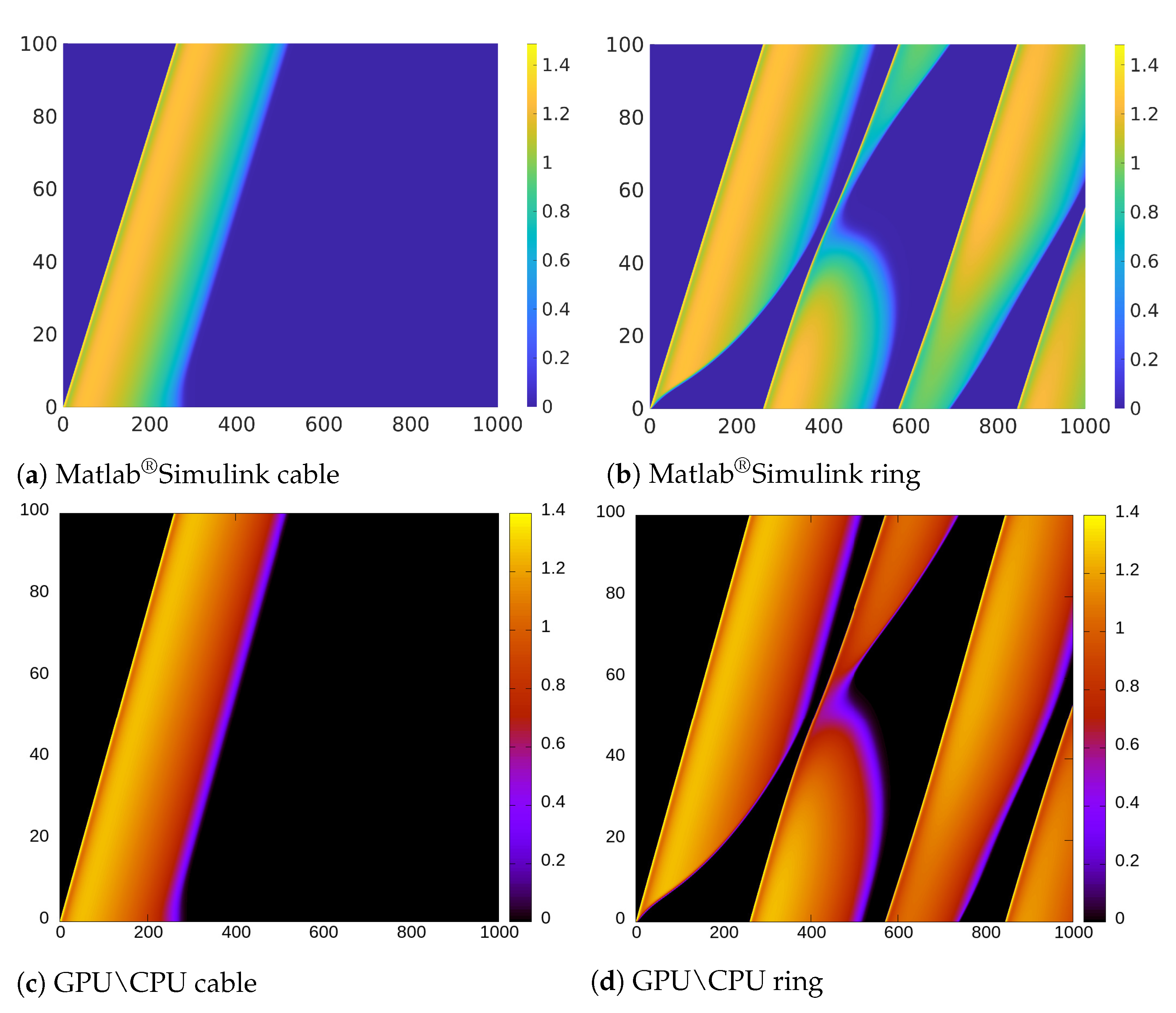
From the implementation point of view, we found Matlab
®Simulink very suitable for the development of rapid model prototypes. However, it is limited when it is necessary to handle large scale simulations. Furthermore, it was not possible to change the length of the cable/ring at runtime. Thus, we were not able to test the self-adapting properties described in the previous sections. In addition, the values of the variables at a certain time step cannot be updated using just the values of the previous step but rather all the values already computed in the current step. Furthermore, we found that the computational resources required to compile large HCA models with several cells were prohibitive (we could consider only models with a length of max. 3000 cells). Due to these restrictions, we could not implement the decreasing ring and the related Monte Carlo analysis in this environment. Nevertheless, as illustrated in the space-time plot shown in
Figure 11b, we were able to induce electrical alternans with a trapped wave in a ring with a small number of cells by setting the value of the resistance to
.
We performed the same numerical experiments by using CPU-based and GPU-based implementations (see
Figure 11d). Due to the negligible error between the results obtained using GPU and CPU, in
Figure 11 we only provide the result obtained using CPU as representative. While
Figure 11b,d look very similar, there is actually a small difference in the APD alternans profile. Specifically, the APD in
Figure 11d, is longer than in
Figure 11b.
If we observe the color representing the value of the variable
u, we notice that the transition in
Figure 11a is softer and shorter than in
Figure 11c. This is due to the different mechanisms for updating the variables in Matlab
®Simulink previously discussed and that are not suitable to implement CA updating rules that depend on the values of the previous step.
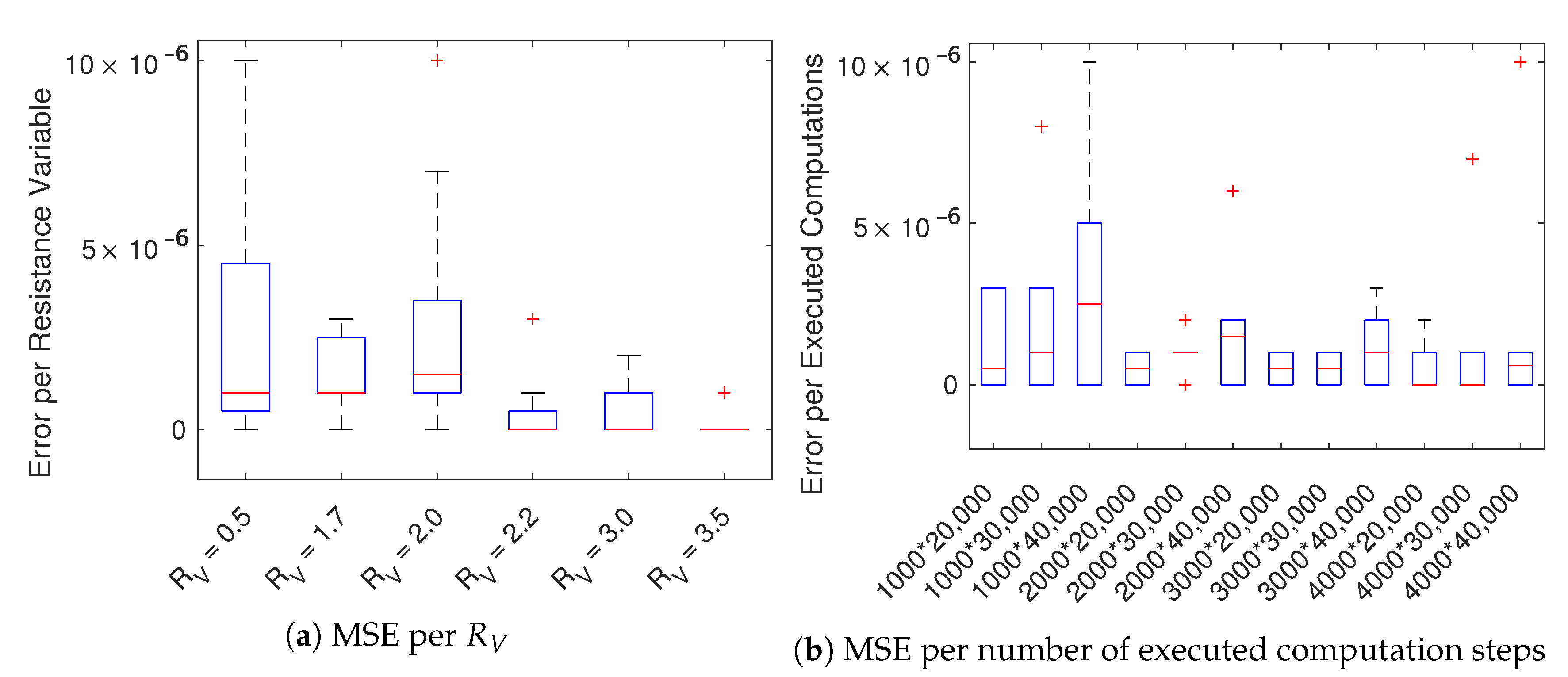
We, therefore, focused on the implementations using CPU (sequential) and GPU (parallel), producing instead results with a negligible difference (the average mean squared error is
). As expected, the parallel GPU-based implementation benefits of the use of many-cores accelerating considerable the simulation. In
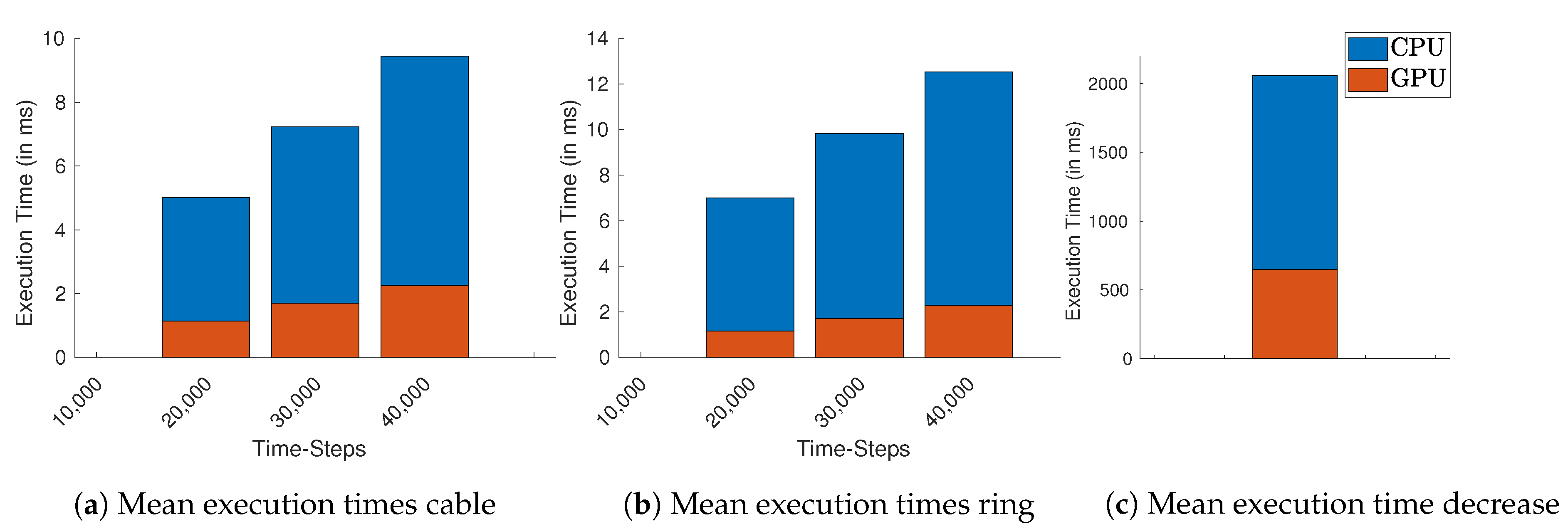
Figure 5 we compute the average execution times for all used lengths and
. For simulations with a fixed number of time-steps, only the number of cells and time-steps increase the execution times.
The longest execution time measured was the decreasing ring simulation in both implementations, as we stop when no AP can be recovered. Still, the average execution time overall lengths and
for this scenario using GPU is
compared to the measured times using CPU, as shown in
Figure 5c. Since this kind of analysis does not depend on a fixed number of time-steps, but it relies on the AP dynamics, the execution time for this scenario depends highly on the number of cells and the chosen value for the resistance
, shown in
Table A4 and
Table A5.
If we consider simulations with a fixed number of time-steps, the GPU implementation started to perform approximately 10× faster on average than the CPU-based implementation for simulations with more than 40,000 time-steps. If we consider the simulations with the highest number of required time-steps, this becomes more evident: the average measured execution time for simulating the ring with 40,000 time-steps and 2750 cells on GPU is ~2.60 ms while on the CPU required on average 21.60 ms. For the cable using these lengths and number of time-steps, the GPU implementation finishes consistently with an average of ~2.60 ms, while the CPU implementation required (less time than in the ring setting) within ~14.86 ms. In all implementations, we were able to control the CV with the free resistance variable.
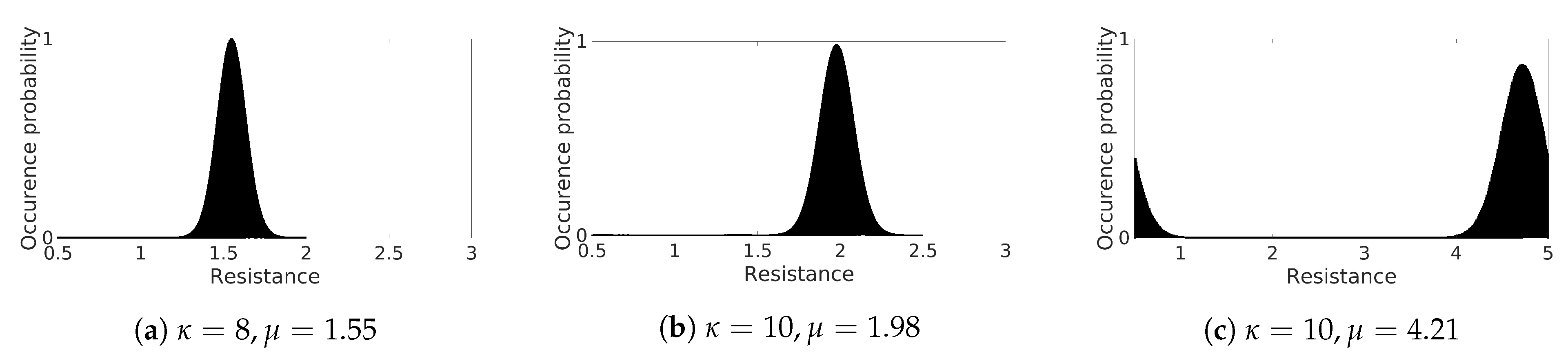
Although it is possible to observe APD fluctuations at the beginning of the Monte Carlo simulations of shorter rings, the critical limit in length for the onset of the alternans is consistent in all ring sizes simulated using both CPU and GPU implementations. The occurrence of alternans depends only on the distribution of the resistance and thus on the resulting CV.
The observed decrease in the CV and the occurrence of APD fluctuations in inhomogeneous settings can be compared to the behavior observed in studies of ischemic cardiac tissue [
59,
60]. Even though we have slight inhomogeneities in the cell’s resistance, our analysis shows that the APD is stable until the length falls below a certain critical point when too many cells undergo large oscillations, thus a continuous propagation. Such a condition depends on the chosen initial number of cells and CV, as we could observe an early onset with both von Mises distributions that we have considered. Considering a CV of
, the obtained APD is similar to the results in [
11] attained considering a regular body temperature of 37 °C. It is also similar to the original measurement data for epicardial cells referenced in [
16].
5. Conclusions
We have presented an HCA modeling the cardiac cell-cell membrane resistance with a free variable.
We provide three different implementations comparing pros and cons: one using the Matlab®Simulink graphical environment, a second implementation running on a CPU sequentially, and a third implementation running on a GPU in parallel. While Matlab®Simulink provides a suitable environment for fast prototyping complex models is less efficient in handling simulation instances of large HCA models. The GPU-based implementation could handle instead larger instances accelerating considerably the simulation (in our experiments, we could observe 10× speed-up with respect to the CPU-based implementation) and indeed the overall analysis of the model.
We then study the cardiac behavior within a unidimensional domain considering inhomogeneous resistance by performing a Monte Carlo simulation. We show that our modeling approach can reproduce important and complex spatiotemporal properties such as self-adaptive properties and the onset of alternans, paving the ground for promising future applications.
In future work, we plan to extend our analysis from the one-dimensional cases to two and three spatial dimensions, including the possibility to use different cell types. We foresee to generalize the present study towards an efficient computation of ECG signals [
61] simulating pathological conditions adapting critical parameters such as the free resistance variable and the ionic time constants. Additionally, we aim to apply our HCA computational approach in a more physiological context by implementing species-specific models [
62,
63,
64,
65] and investigating the underlying genetics [
66] linked to cell-to-cell communications problems [
67,
68,
69].
As mentioned in [
58,
70,
71], the estimation of cardiac conductivity is still under investigation; therefore, we need to consider approaches to determine a more precise value for this parameter for future work. One strategy worth investigating is to use machine learning approaches or optimization algorithms such as the one described in [
58] that map the free resistance variable to a specific conduction velocity. We also plan to improve our GPU-based implementation by combining our current approach with other optimizations described in other papers such as [
34,
72,
73].
The use of other hardware accelerators, such as FPGAs that are becoming more and more employed to handle computational extensive problems in simulation [
74,
75] and graphic processing [
76,
77] represents a promising perspective.