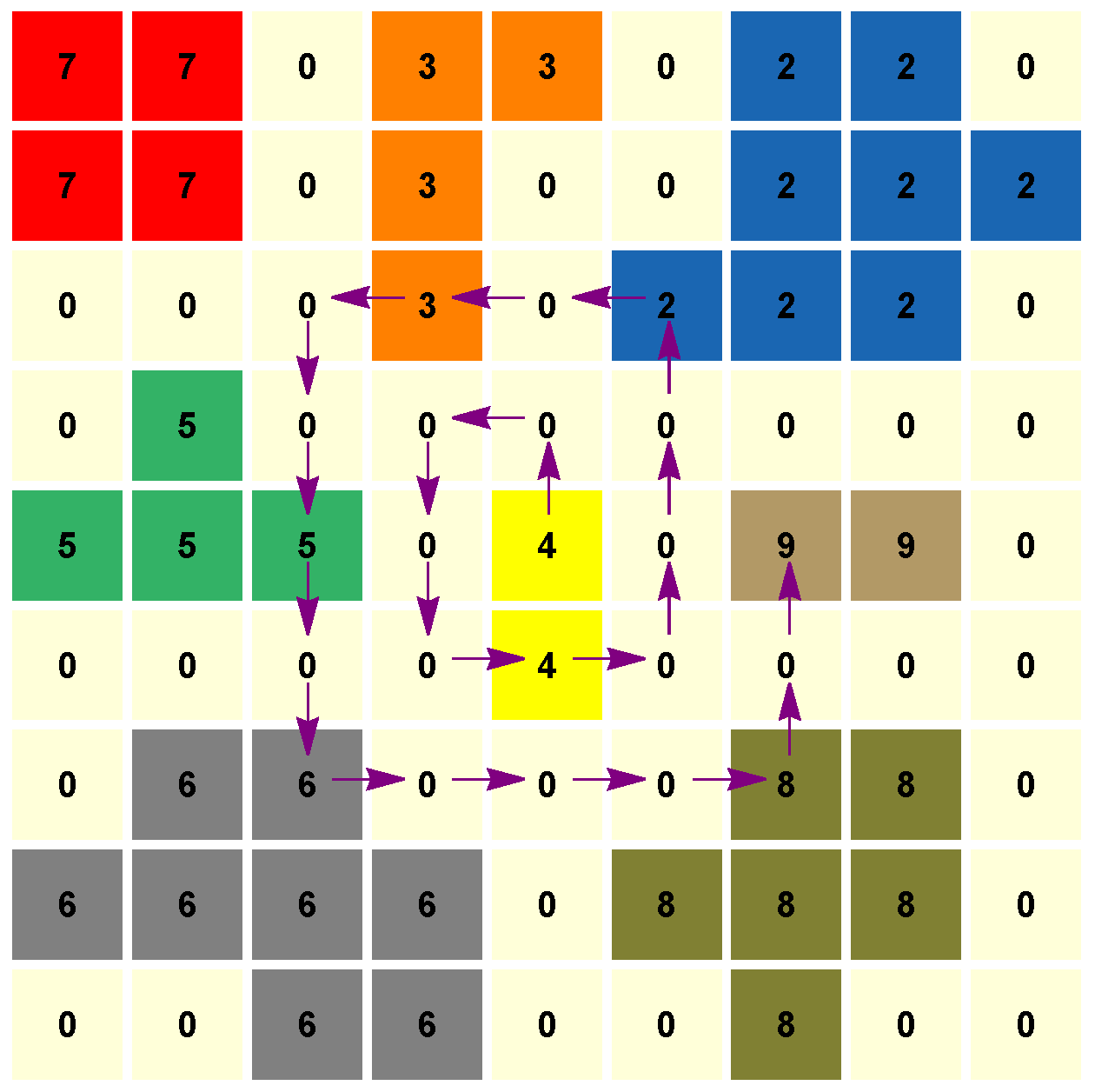
Figure 1.
Algorithm 1 runs over the pixels following a spiral until the desired number of labels (of cells) is reached. In this toy example, taking as input parameter , the output is the set of labels . Boundary pixels are labeled by 0.
Figure 1.
Algorithm 1 runs over the pixels following a spiral until the desired number of labels (of cells) is reached. In this toy example, taking as input parameter , the output is the set of labels . Boundary pixels are labeled by 0.
Figure 2.
An example of the sub filtration for . The top row corresponds to dNP and the bottom to cEE.
Figure 2.
An example of the sub filtration for . The top row corresponds to dNP and the bottom to cEE.
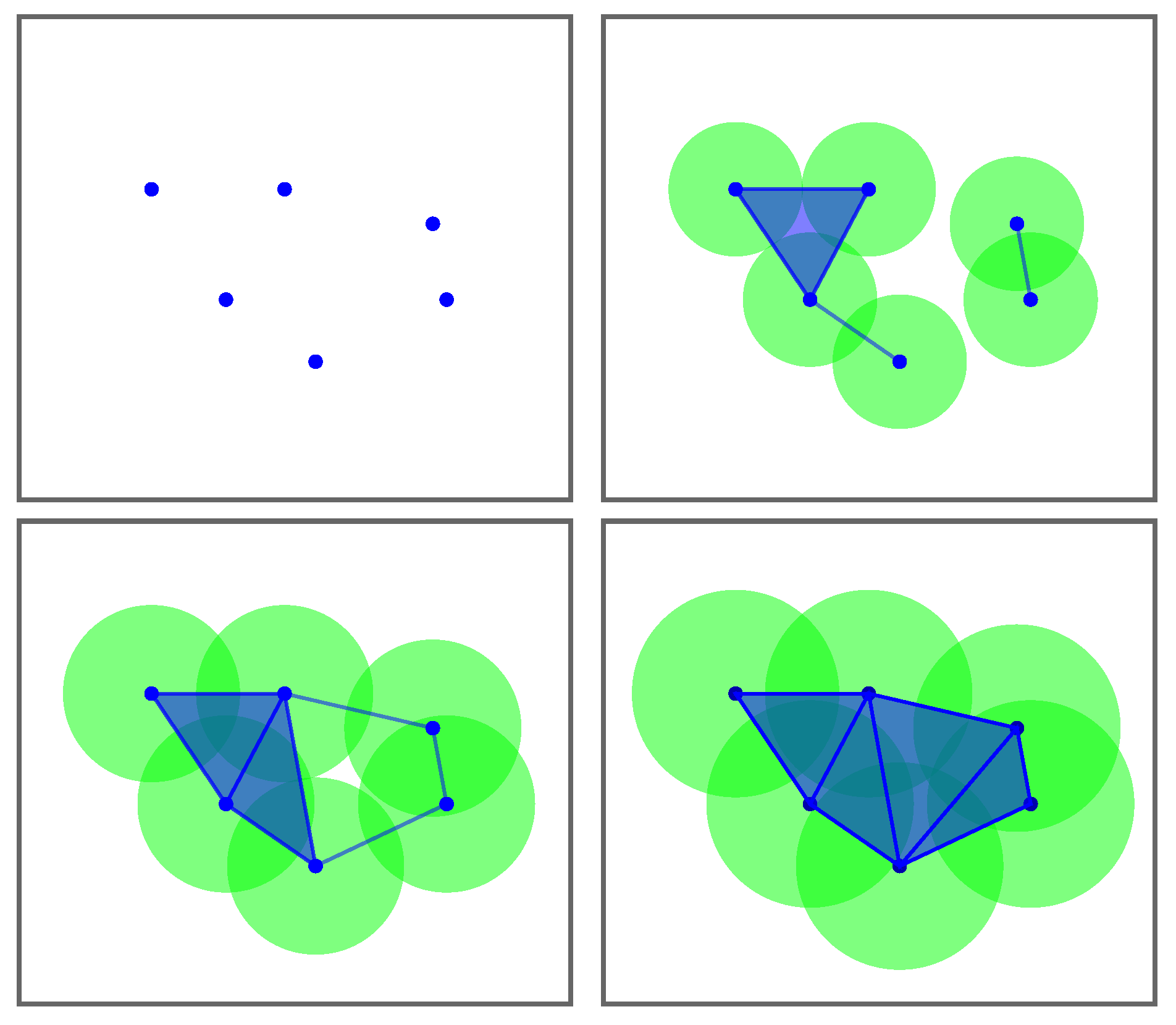
Figure 3.
An example of a rips filtration with 6 points in the Euclidean plane. Note that a simplex arises when the distance between the corresponding centroids is smaller than or equal to twice the radius.
Figure 3.
An example of a rips filtration with 6 points in the Euclidean plane. Note that a simplex arises when the distance between the corresponding centroids is smaller than or equal to twice the radius.
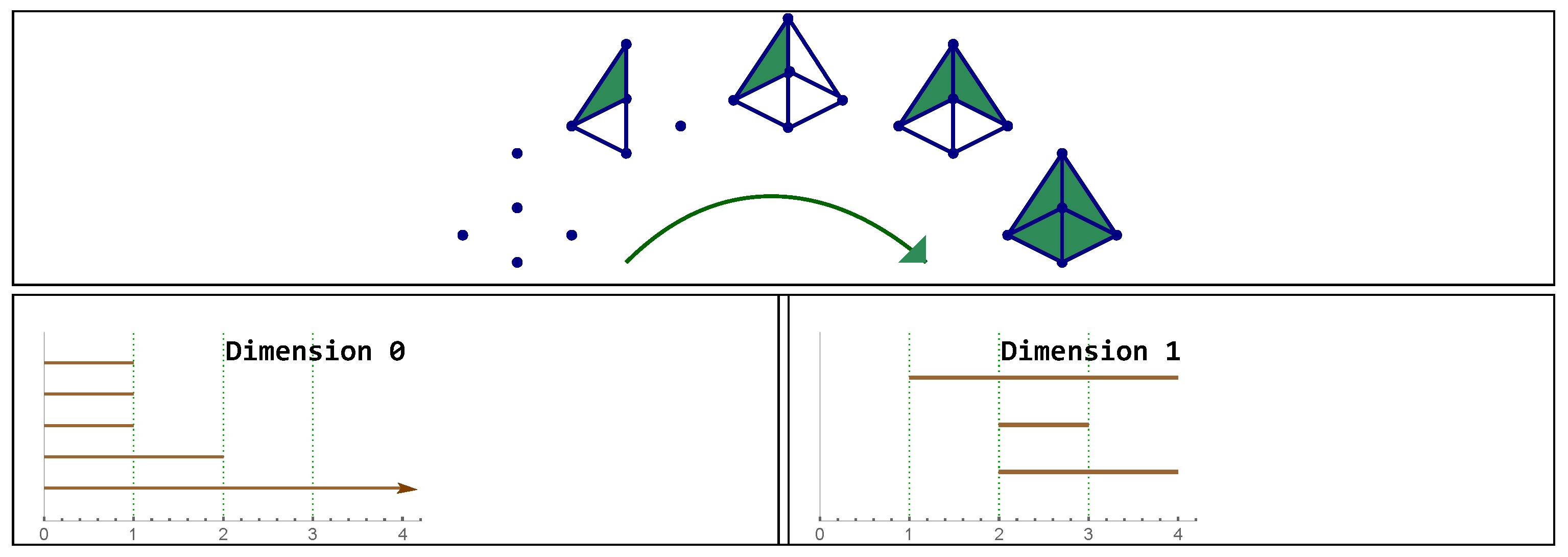
Figure 4.
Top: example of a filtration . Bottom: barcodes representing connected components and cycles. Note there is a bar in the 0 dimensional persistent homology. In this example: and then appears once in the 1-dimensional barcode.
Figure 4.
Top: example of a filtration . Bottom: barcodes representing connected components and cycles. Note there is a bar in the 0 dimensional persistent homology. In this example: and then appears once in the 1-dimensional barcode.
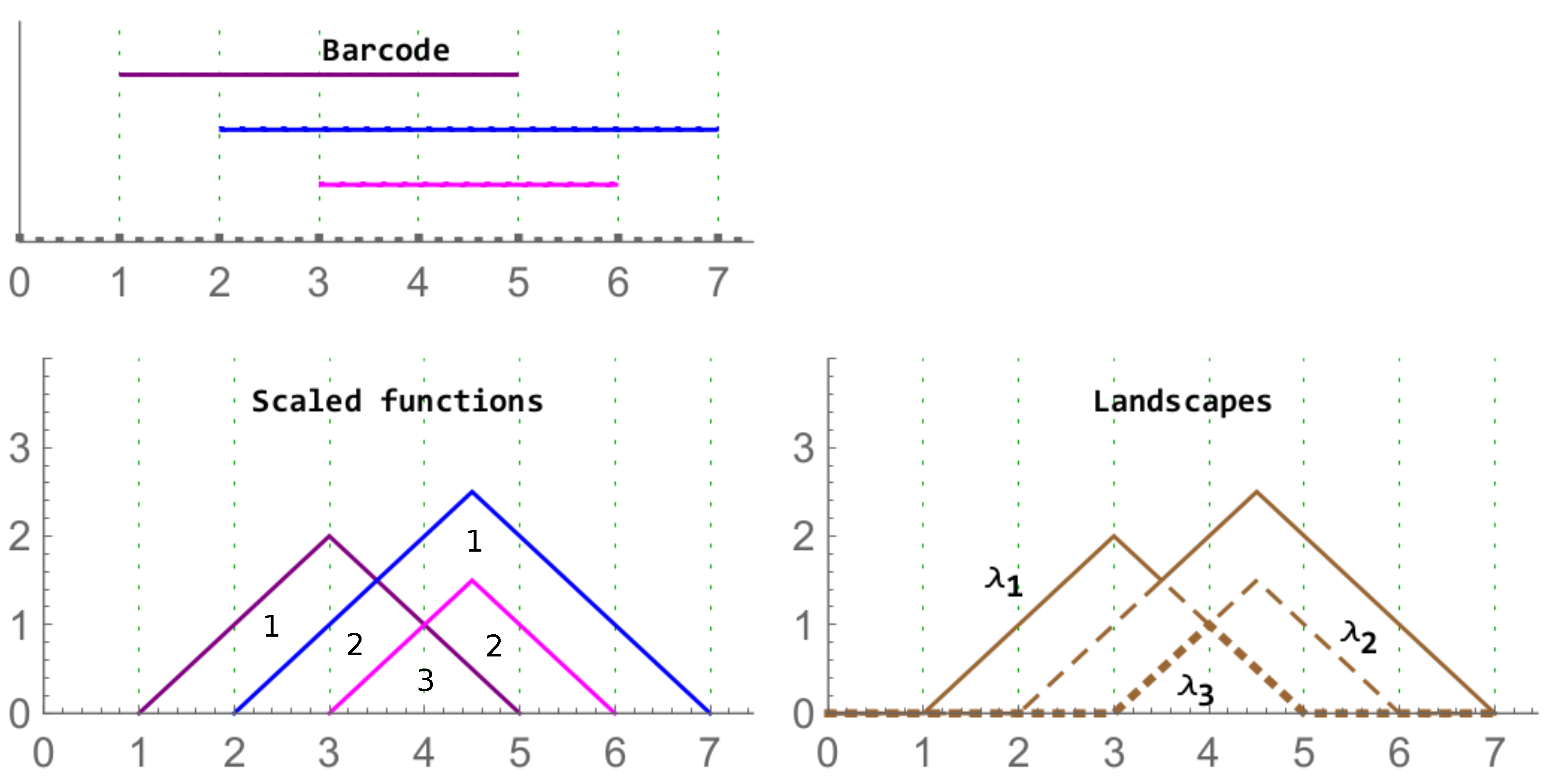
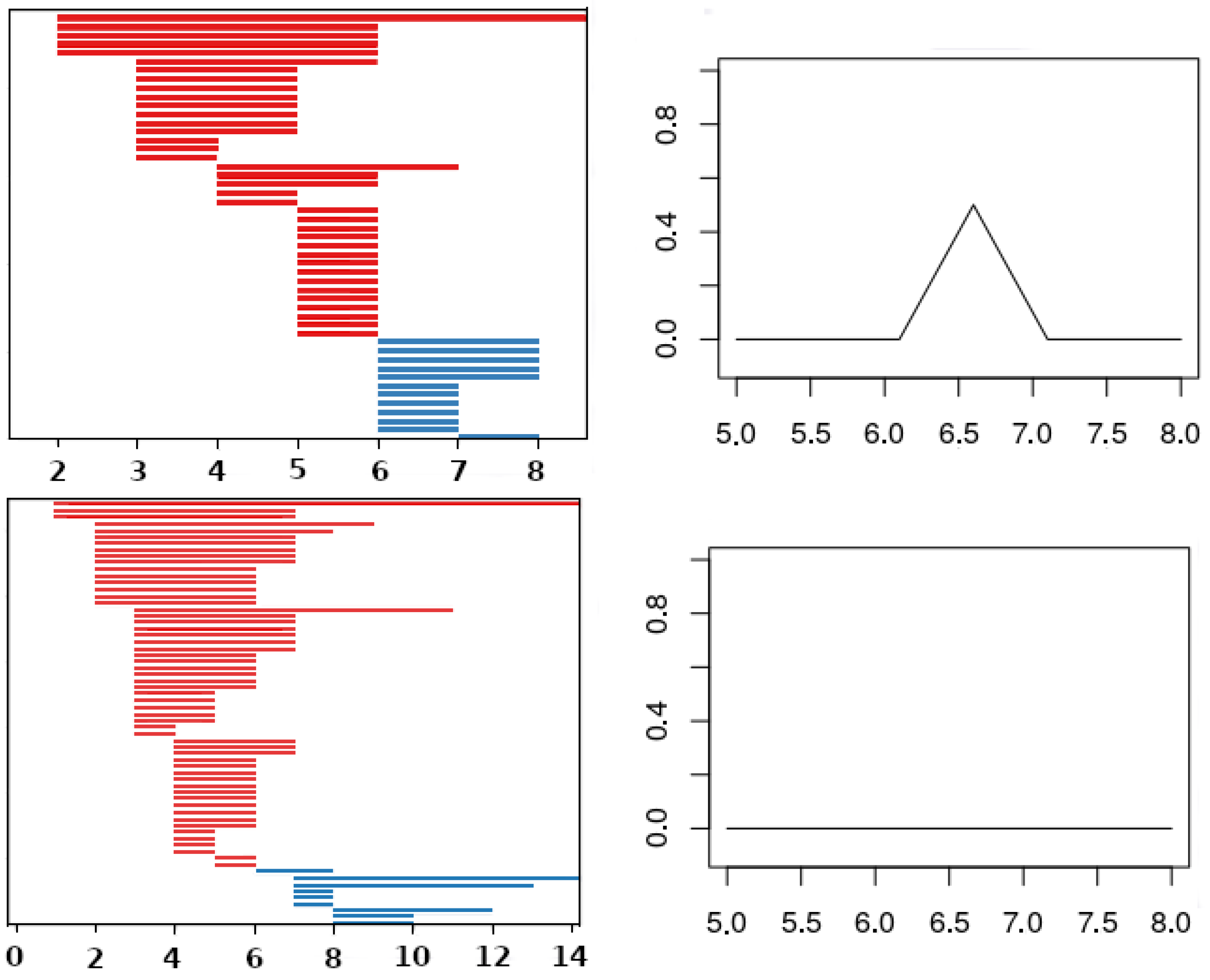
Figure 5.
Left column: A barcode (top) and its corresponding rescaled rank functions (bottom). The values of the functions in the corresponding region are provided. On the right, the associated landscape with the functions , , and are displayed with different layouts.
Figure 5.
Left column: A barcode (top) and its corresponding rescaled rank functions (bottom). The values of the functions in the corresponding region are provided. On the right, the associated landscape with the functions , , and are displayed with different layouts.
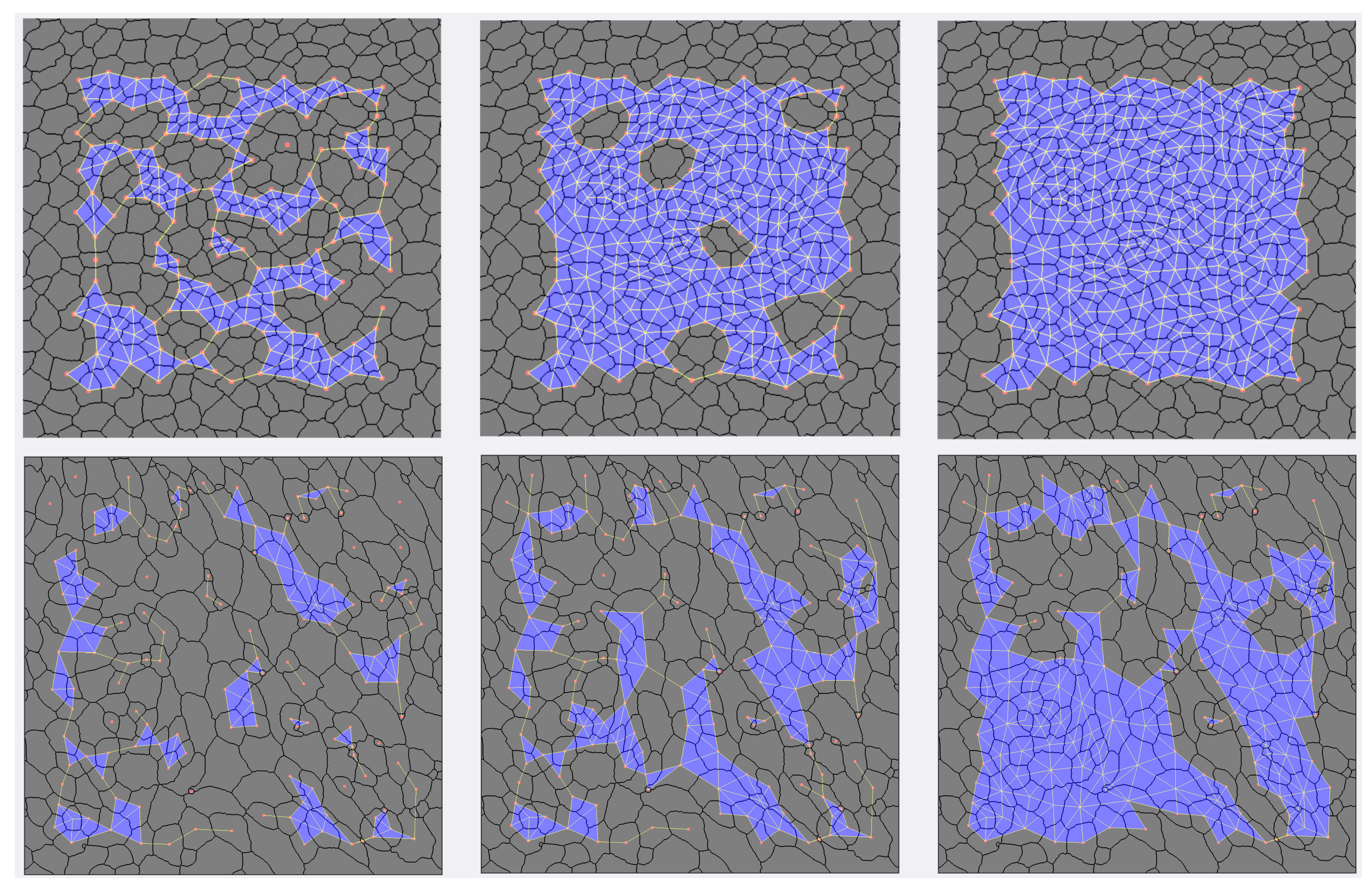
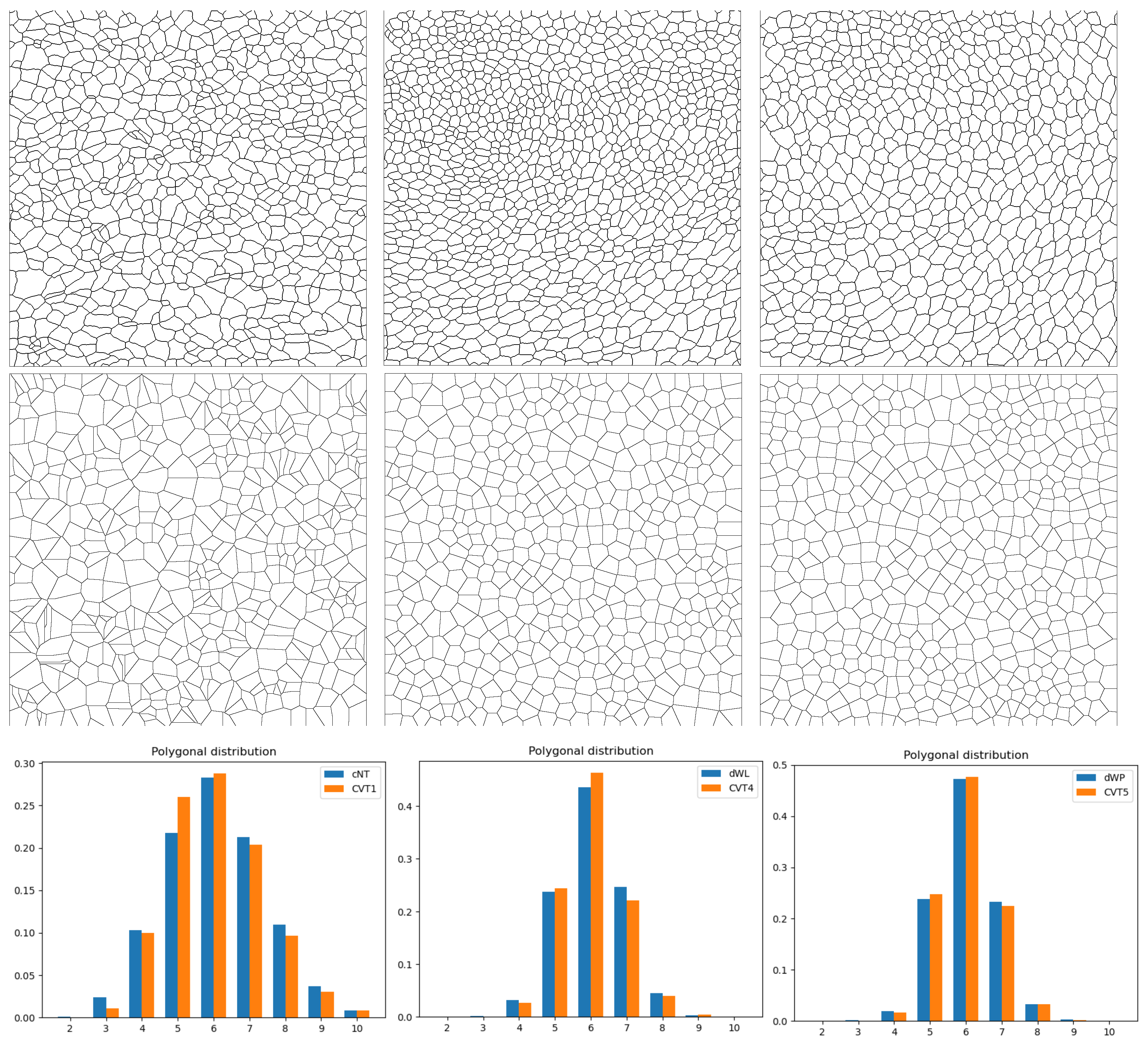
Figure 6.
An example of images of epithelial tissues (top row), their CVT-path counterpart (middle row), and a histogram with their polygonal distribution (bottom row). In the first column, we show cNT and ; in the second, dWL and , and in the third, and .
Figure 6.
An example of images of epithelial tissues (top row), their CVT-path counterpart (middle row), and a histogram with their polygonal distribution (bottom row). In the first column, we show cNT and ; in the second, dWL and , and in the third, and .
Figure 7.
An example of barcodes (0-bars in red and 1-bars in blue) with their landscape . The top corresponds to dNP and the bottom to cEE. Note that in the cEE case, there are not 9 1-dimensional holes simultaneously alive, so its landscape is zero.
Figure 7.
An example of barcodes (0-bars in red and 1-bars in blue) with their landscape . The top corresponds to dNP and the bottom to cEE. Note that in the cEE case, there are not 9 1-dimensional holes simultaneously alive, so its landscape is zero.
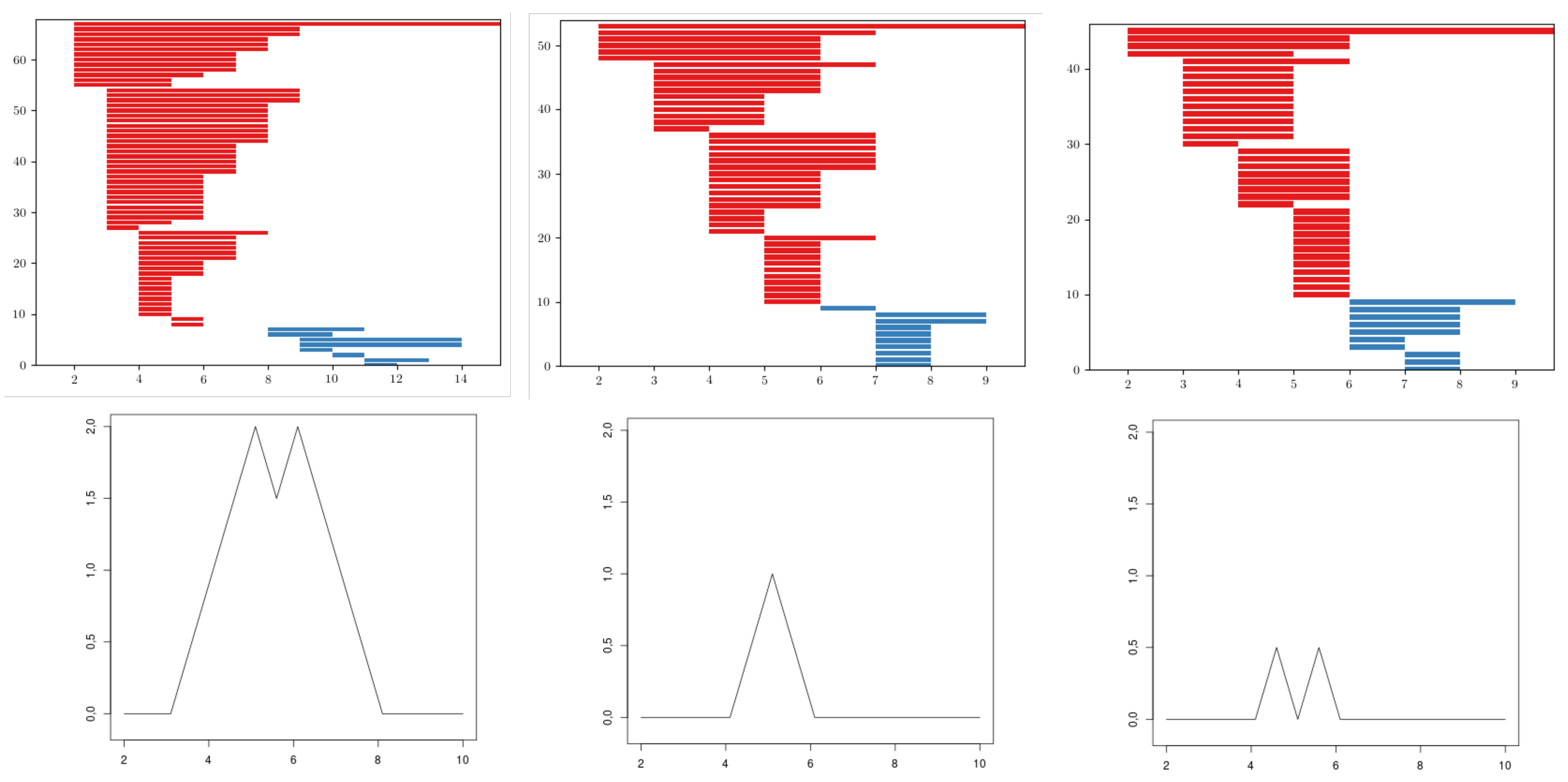
Figure 8.
An example of barcodes and its landscapes from cEE, cNT, and dWL, respectively. Note that the domain of the landscapes of cNT and dWL are the same, but the area of cNT is greater, since in that domain the 18 bars are the same, while for dWL the bars vary.
Figure 8.
An example of barcodes and its landscapes from cEE, cNT, and dWL, respectively. Note that the domain of the landscapes of cNT and dWL are the same, but the area of cNT is greater, since in that domain the 18 bars are the same, while for dWL the bars vary.
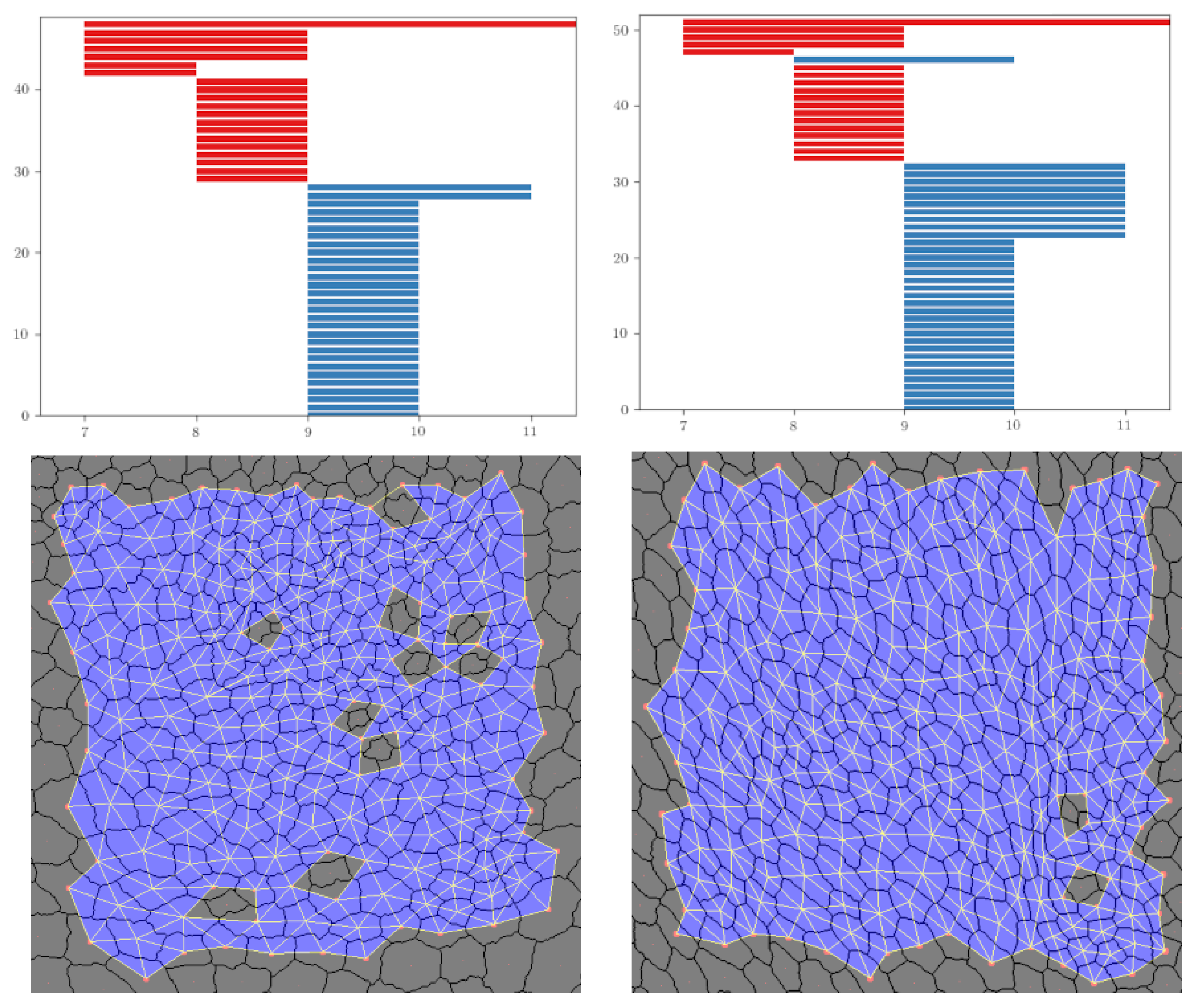
Figure 9.
An example with the persistence barcodes of dWL and dWP, which provide the median for and their corresponding sup filtration when i = 10.
Figure 9.
An example with the persistence barcodes of dWL and dWP, which provide the median for and their corresponding sup filtration when i = 10.
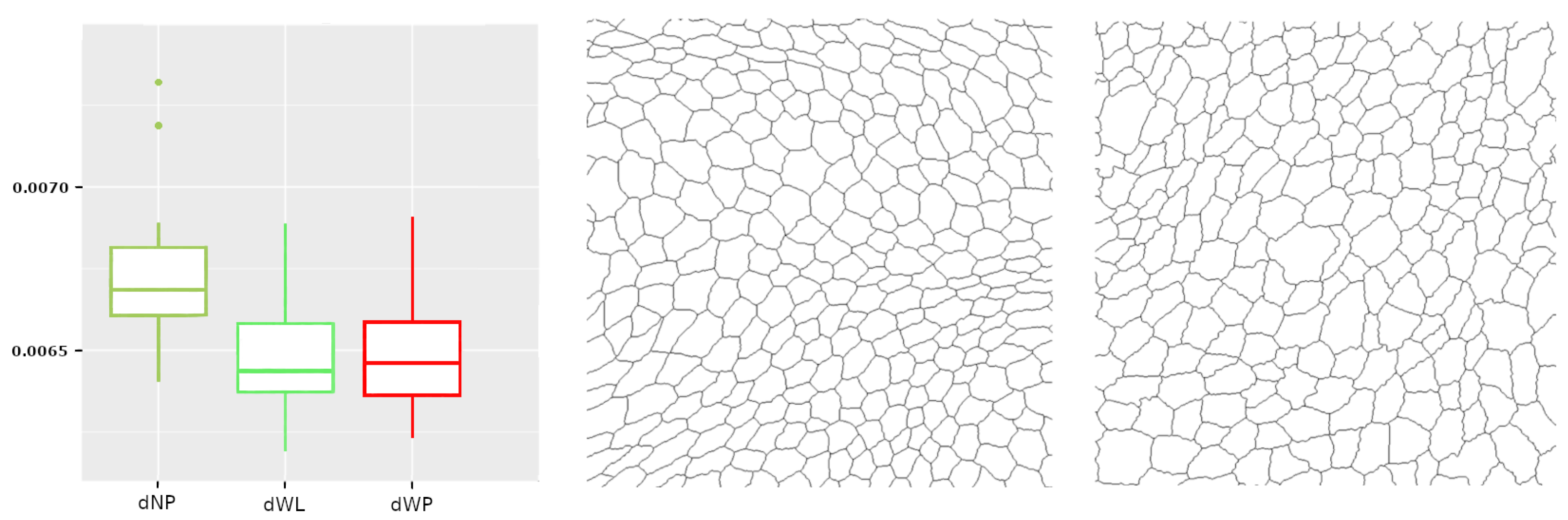
Figure 10.
On the left, the boxplot corresponds to . On the right are the images of dNP and dWL, which provide the median for each of these sets of images.
Figure 10.
On the left, the boxplot corresponds to . On the right are the images of dNP and dWL, which provide the median for each of these sets of images.
Table 1.
The number of valid cells in each image of the epithelial tissues.
Table 1.
The number of valid cells in each image of the epithelial tissues.
| | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 |
|---|
| cEE | 140 | 206 | 229 | 241 | 385 | 380 | 261 | 187 | 405 | 246 | 327 | 204 | 348 | 270 | | |
| cNT | 666 | 661 | 566 | 574 | 669 | 532 | 420 | 592 | 744 | 527 | 594 | 473 | 704 | 748 | 469 | 834 |
| dNP | 513 | 723 | 588 | 525 | 439 | 823 | 1102 | 533 | 309 | 575 | 302 | 375 | | | | |
| dWL | 432 | 556 | 485 | 525 | 501 | 936 | 890 | 790 | 977 | 913 | 606 | 835 | 785 | 748 | 622 | |
| dWP | 748 | 806 | 566 | 415 | 454 | 654 | 752 | 713 | 504 | 430 | 387 | 516 | 419 | 455 | 277 | 257 |
Table 2.
Differences between the tissues for 187 cells. A check mark implies that the p-value of that variable is smaller than in the Dunn test and a cross mark that we could not find significant differences using that variable.
Table 2.
Differences between the tissues for 187 cells. A check mark implies that the p-value of that variable is smaller than in the Dunn test and a cross mark that we could not find significant differences using that variable.
| 187 Cells | | | | | |
|---|
| cEE vs. cNT | √ | × | × | × | × |
| cEE vs. dNP | √ | × | √ | × | √ |
| cNT vs. dNP | √ | × | √ | × | × |
| cEE vs. dWL | √ | × | √ | √ | √ |
| cNT vs. dWL | × | × | √ | √ | √ |
| dNP vs. dWL | × | × | × | √ | × |
| cEE vs. dWP | √ | √ | √ | √ | √ |
| cNT vs. dWP | × | √ | √ | √ | √ |
| dNP vs. dWP | × | × | × | √ | √ |
| dWL vs. dWP | × | √ | × | × | × |
Table 3.
The p-values of some variables for the Mann–Whitney U test between dWL and dWP. The number of cells is set at 187 and 257.
Table 3.
The p-values of some variables for the Mann–Whitney U test between dWL and dWP. The number of cells is set at 187 and 257.
| dWL vs. dWP | | | |
|---|
| N = 187 | 0.013 | 0.01 | 0.019 |
| N = 257 | 0.012 | 0.006 | 0.005 |
Table 4.
Differences between some tissues and their CVT-path counterpart. A check mark implies that the p-value of that variable is smaller than in the Mann–Whitney U test and a cross mark that we could not find significant differences using that variable.
Table 4.
Differences between some tissues and their CVT-path counterpart. A check mark implies that the p-value of that variable is smaller than in the Mann–Whitney U test and a cross mark that we could not find significant differences using that variable.
| 257 Cells | | | | |
|---|
| cNT vs. | √ | √ | √ | √ |
| dWL vs. | × | √ | √ | × |
| dWP vs. | × | √ | × | √ |
Table 5.
Classification of tissues using all TDA variables, all network variables, all variables together, mean and variance of the degree, and a combination of these two with . The accuracy for the training/validation sets are displayed. Best results are in bold.
Table 5.
Classification of tissues using all TDA variables, all network variables, all variables together, mean and variance of the degree, and a combination of these two with . The accuracy for the training/validation sets are displayed. Best results are in bold.
| 187 Cells | TDA | Network | Mixed | m & v | m & v &
|
|---|
| cEE | 85.5/86.7 | 86.2/87.8 | 86.1/87.8 | 89.7/94.3 | 97.1/98.7 |
| cNT | 82.6/83 | 93.4/93.7 | 92.2/92.9 | 89.2/89.7 | 96.2/97.7 |
| Drosophila | 99.7/99.8 | 100/100 | 100/100 | 100/100 | 100/100 |
| overall | 92.9/93.4 | 95.8/96.1 | 95.5/96 | 95.5/96.2 | 98.6/99 |
Table 6.
Classification using Random Forests of the epithelial tissues. On the left, results using the number of neigbors and mean and variance of the degree. Best results are in bold. It can be seen that performs better.
Table 6.
Classification using Random Forests of the epithelial tissues. On the left, results using the number of neigbors and mean and variance of the degree. Best results are in bold. It can be seen that performs better.
| 257 Cells | Network | |
|---|
| dWL | 63.8/67.4 | 59.9/59.9 |
| dWP | 64.9/66.5 | 82.6/82.8 |
| global | 63.1/65.5 | 70/71.8 |