1. Introduction
Finding the maximum clique in a graph is a well-studied NP-complete problem [
1]. Recently developed algorithms significantly reduce the time required to search for a maximum clique, which is of great practical importance in many fields such as bioinformatics, social network analysis, and computational chemistry [
2,
3].
There have been many advances in the search for faster algorithms for maximum cliques, many of which focus on specific domains of graphs [
4,
5,
6,
7]. To make the algorithm work fast on general graphs, some good heuristics have been proposed to speed up the branch-and-bound search [
1,
4,
8,
9,
10,
11,
12,
13,
14,
15]. One such algorithm is MCQD, on which we have built [
4]. It has been shown that the MCQD algorithm is faster than many other similar branch-and-bound algorithms in finding maximum cliques [
1]. In the MCQD algorithm, there is a single parameter that can be set before the algorithm is executed. This parameter, called Tlimit, controls the fraction of a graph on which tighter upper bounds apply to the size of a maximal clique. These upper bounds require that (O(N
2)) be computed. The fraction of a graph on which looser upper bounds are used (O(NlogN)) is empirically estimated to be 0.025 for random graphs. Even though MCQD seems to progress quickly with a default value of Tlimit in many graphs, there are some graphs where Tlimit performs poorly [
4]. In particular, the Tlimit parameter is suboptimal in some dense and synthetic graphs of the DIMACS benchmark [
16]. Here, we present an improvement to the original MCQD algorithm that automatically determines the value of the Tlimit parameter for the MCQD algorithm. We predict that the Tlimit parameter uses machine learning for the input graph. The code used to perform the experiments is freely available at
http://insilab.org/mcqd-ml (accessed on 9 November 2021).
1.1. Problem Description and Notation
Let G = (V, E) be an undirected graph, where V = 1, …, n is a set of vertices and E⊂V × V is a set of edges. A clique C in the graph G is a set of nodes defined such that there exists an edge between every two nodes in C. We say that C is a maximum clique if its cardinality |C| is the largest among all cliques in the graph G. The maximum clique problem (MCP) is an optimization problem that seeks the maximum clique in a given graph. The clique number w(G) of graph G is the number of nodes in the maximum clique of graph G. The maximum clique problem is strictly equivalent to a maximum independent set (MIS) as well as the minimum vertex cover problem (MVC). Finding the maximum clique is an NP-complete problem. We do not know if there is an algorithm for this group of problems that can find the solution in polynomial time. It is likely that no such algorithm exists.
1.2. Maximum Clique Dynamic (MCQD) Algorithm
The MCQD algorithm is based on a branch and bound principle [
4]. It uses approximate graph coloring to estimate the upper bound of the maximum clique size and is shown in Algorithm 1.
| Algorithm 1. Dynamic algorithm for maximum clique search. |
| 1: procedure MaxCliqueDyn(R, C, level) |
| 2: S[level] ← S[level] + S[level - 1] – Sold[level] |
| 3: Sold[level] ← S[level - 1] |
| 4: while R ≠ Ø do |
| 5: choose a vertex p with maximum C(p) (last vertex) from R |
| 6: R ← R \ {p} |
| 7: if |Q| + C[index_of_p_in_R] > |Qmax| then |
| 8: Q ← Q ∪ {p} |
| 9: if R ∩ Γ(p) ≠ Ø then |
| 10: if S[level] / ALL_STEPS < Tlimit then |
| 11: calculate the degrees of vertices in G(R ∩ Γ(p)) |
| 12: sort vertices in R ∩ Γ(p) in descending order with respect to their degrees |
| 13: ColorSort(R ∩ Γ(p), C’) |
| 14: S[level] ← S[level] + 1 |
| 15: ALL_STEPS ← ALL_STEPS + 1 |
| 16: MaxCliqueDyn(R ∩ Γ(p), C’, level + 1) |
| 17: else if |Q| > |Qmax| then |
| 18: Qmax ← Q |
| 19: Q ← Q \ {p} |
The algorithm stores the current clique in the variable Q and keeps track of the current maximum clique size in the variable Qmax. As an input, it accepts an ordered set of nodes based on their color, a set of colors, and the level variable which provides the current depth of the recursive function. The algorithm also uses two global variables, S[level] and Sold [level], which store the sum of steps up to the current level of algorithm progression and the previous level Sold [level] = S[level − 1]. With the Tlimit parameter, we can limit the use of the graph coloring of vertices R sorted by their degree. When the proportion of steps up to a certain level of recursion is less than Tlimit, we perform additional operations of recalculating the vertex degrees for the remainder of the graph and of resorting these vertices according to their descending degrees. This additional work increases the tendency of the ColorSort function to estimate a tighter upper bound for the size of a maximum clique, generally reducing the number of steps and time necessary for the algorithm to find a maximum clique. The Tlimit value used in the original paper [
4] was empirically determined on a sample of random graphs and was set to a value of 0.025.
1.3. Protein Product Graphs and Use of Molecular Docking Graphs in Drug Discovery
To move drugs from the research phase to the trial phase, the most promising molecules must be identified from a set of potential candidates. This requires a detailed knowledge of the functions of drug target proteins, which is often lacking. Protein functions can be determined by comparing the structure of unknown proteins to proteins with known functions [
2]. To compare proteins with each other, we can represent them as protein graphs, such as we did with the ProBiS (Protein Binding Sites) algorithm [
17]. Two protein graphs can be compared by constructing a protein product graph, which is a Cartesian product of the two protein graphs and captures all possible overlaps of one protein with the other. Finding a maximum clique in this protein product graph is directly equivalent to finding the alignment that overlaps most of the vertices of the protein graphs. The quality of the overlap is an indication of the similarity of the proteins.
Another application for maximum clique search is molecular docking, which is often performed as a high-throughput screening approach whose goal is to predict the binding position and binding affinity of potential ligands of a target protein [
18]. In a particular class of molecular docking called fragment docking, which was explored in our ProBiS-Dock docking algorithm, a maximum clique algorithm is used to reconstruct a docking graph of the small molecule in a protein-bound conformation from fragments of the previously docked molecule. The calculated binding affinities of the docked fragments can be included in this graph as node weights, resulting in a weighted docking graph. A clique with maximum weight in such a graph represents the docked conformation of a small molecule with the highest binding affinity among all possible conformations of that small molecule. This allows the algorithm to discover potential new ligands of a protein that could become drugs in the future.
2. Overview of Graph Theory and Neural Networks Approaches
We describe the novel developed MCQD-ML (Maximum Clique Dynamic–Machine Learning) algorithm that was tested with different types of graphs and incorporates different machine learning models.
2.1. Graphs Used for Training and Testing
To train the machine learning algorithm, we first create a variety of graphs. In order to capture the largest possible variety of target graphs in our training set, we include 10,000 sparse and dense random graphs, as well as 15 complete protein graphs and 200 molecular docking graphs. The random graphs are generated such that each edge exists with probability d, where d is greater than 0.99 in dense graphs. The types of graphs are presented in the following sections.
2.2. Molecular Docking Graphs
To identify energetically preferred docking conformations of potential ligands, we performed a maximum clique search in molecular docking graphs. A molecular docking graph is a graph whose nodes are docked molecular fragments and in which two nodes are connected if the docked fragments can be connected with linker atoms to reconstruct the original docked molecule. Each node is assigned a weight representing the binding energy (or binding affinity) of a docked fragment. By performing a maximum weight clique search on docking graphs, we can find the combination of docked fragments that yields the conformation with the lowest energy of the docked small molecule with a given protein. We use the ProBiS-Dock algorithm to build molecular docking graphs. The algorithm is used to find the ligands with the highest potential when screening multiple ligands on a target protein [
18,
19].
2.3. Protein Product Graphs
In the ProBiS algorithm [
17], proteins are represented as protein graphs. Each node in a protein graph represents the spatial coordinates of the surface amino acid functional groups. If the distance between nodes u and v is less than 15 Å, there is an edge between two nodes in a protein graph. We can formulate the comparison of two proteins as a maximum clique search by using the notion of a protein product graph. A maximum clique in a protein product graph is a superposition of protein graphs in which the majority of the nodes of two graphs are aligned. The protein product graph of two protein graphs G1 and G2 is defined by a set of nodes, V (G1, G2) = V (G1) × V (G2). Each node in a product graph consists of a node u from graph G1 and a node v from graph G2, both of which represent a similar functional group in the original proteins. In general, a protein product graph can have |V1| × |V2| nodes, but this number is reduced by keeping only the nodes from the original protein graphs G1 and G2 that have similar neighbourhoods in a 6 Å sphere.
2.4. Small Protein Product Graphs
The problem with protein product graphs is the large size of the adjacency matrix, which can exceed the available memory depending on the size of the proteins being compared. It is possible to split a large protein product graph into smaller product graphs that are much denser and contain only a subset of the nodes of the original product graph. The advantage of smaller and denser graphs is the speed at which they can be processed. A disadvantage of smaller protein graphs is the loss of information. If we look for a maximum clique in a small product graph, there is no guarantee that the same clique will be the maximum clique in the entire protein product graph.
2.5. Protocol for Machine Learning on Graphs
To gather as much information as possible about the graph, it is necessary to perform machine learning directly on the graph. To this end, we tested several different graph neural network models and a support vector regression algorithm with the Weisfeiler–Lehman kernel function [
20,
21,
22,
23,
24,
25,
26,
27,
28,
29,
30], which are listed in
Table 1. We tested three different graph neural network models that can model data of different complexity with inductive biases. They are (i) Graph Convolutional Networks (GCN) [
28], (ii) Graph Attention Networks (GAT) [
29,
30], and (iii) Graph Isomorphism Networks (GIN) [
15,
25]. We trained the models on a given training set and then used them to predict Tlimit values for graphs on the test set. The test set contained 15 dense random graphs, 10 small product graphs, 3 product graphs, and 10 docking graphs. We evaluated the performance of the algorithms and calculated the average speed of the standard MCQD algorithm for each set of test graphs. We also calculated the combined speed for the entire test set by summing the runtimes of the algorithms for many different types of graphs and dividing the sum by the runtime required for the MCQD algorithm.
3. Materials and Methods
3.1. Preparation of a Labeled Training Set
Before attempting to use machine learning to improve the selection of the Tlimit parameter value for specific input graphs, we prepared a labeled training set in which different Tlimit values were identified for each graph with the time required to detect the maximum clique. So, we performed the maximum clique search with different Tlimit values on a set of graphs and recorded the time taken by the MCQD algorithm to find the maximum clique. For each generated graph, we ran the MCQD algorithm multiple times for different values of the Tlimit parameter to record the Tlimit values approximately uniformly on a logarithmic scale from 0 to 1. When running MCQD for many graphs and many Tlimit values for each graph, this step becomes computationally intensive. After collecting all Tlimit pairs and their corresponding computation time, we selected the Tlimit value with the lowest time as the best Tlimit value for a graph. This value was then used as the label value for training the machine learning models. The training set consists of graphs as input and the optimal Tlimit value for each graph as the target variable.
3.2. Maximum Clique Dynamic Algorithm with Machine Learning (MCQD-ML)
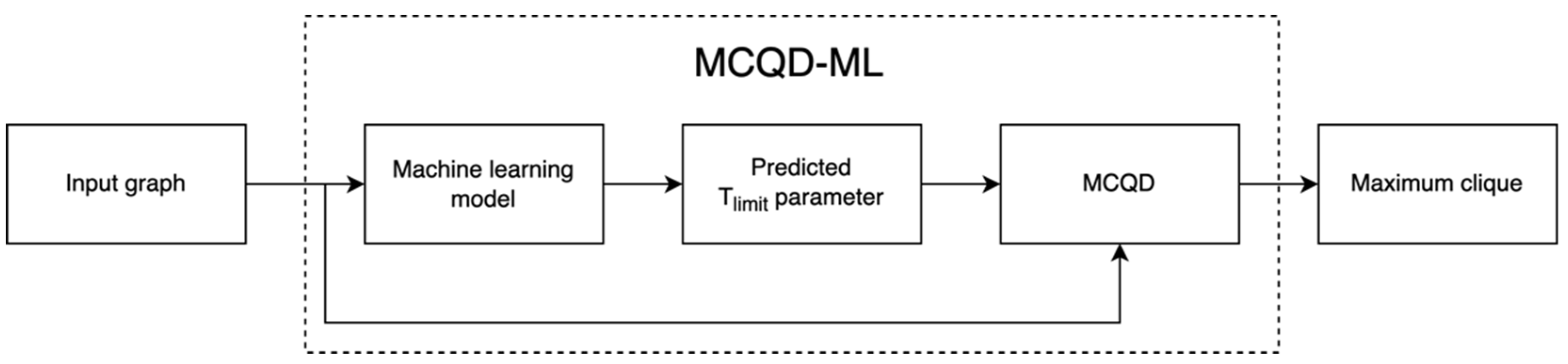
The idea behind the MCQD-ML algorithm is shown in
Figure 1. The algorithm performs inference on the graph to determine a Tlimit parameter before the MCQD algorithm starts, and the MCQD algorithm then uses this parameter instead of the hard-coded parameter. In this way, we obtain the best Tlimit parameter for a given graph and use it to make the MCQD algorithm run faster.
We used an implementation of the MCQD algorithm that can search for a maximum clique as well as a maximum weighted clique. This algorithm is available as source code at
https://gitlab.com/janezkonc/insidrug/-/blob/master/lib/glib/mcqd.cpp (accessed on 9 November 2021). For experimental purposes, we created two training sets and two test sets for molecular docking graphs. One set contains the docking graphs with weights, and in the other set we omit the weights from the docking graphs and assume that all nodes have the same weight. All other graphs are unweighted.
3.3. Evaluation of Possible Acceleration of the MCQD Algorithm
To determine if any speed-ups are possible by tuning the parameter Tlimit, we plot the time needed for MCQD to find the maximum clique at different values of the Tlimit parameter. In
Figure 2, it can be observed that on a random 150 node graph, the default value of parameter is well suited and the maximum clique can be found relatively quickly compared to other values of Tlimit.
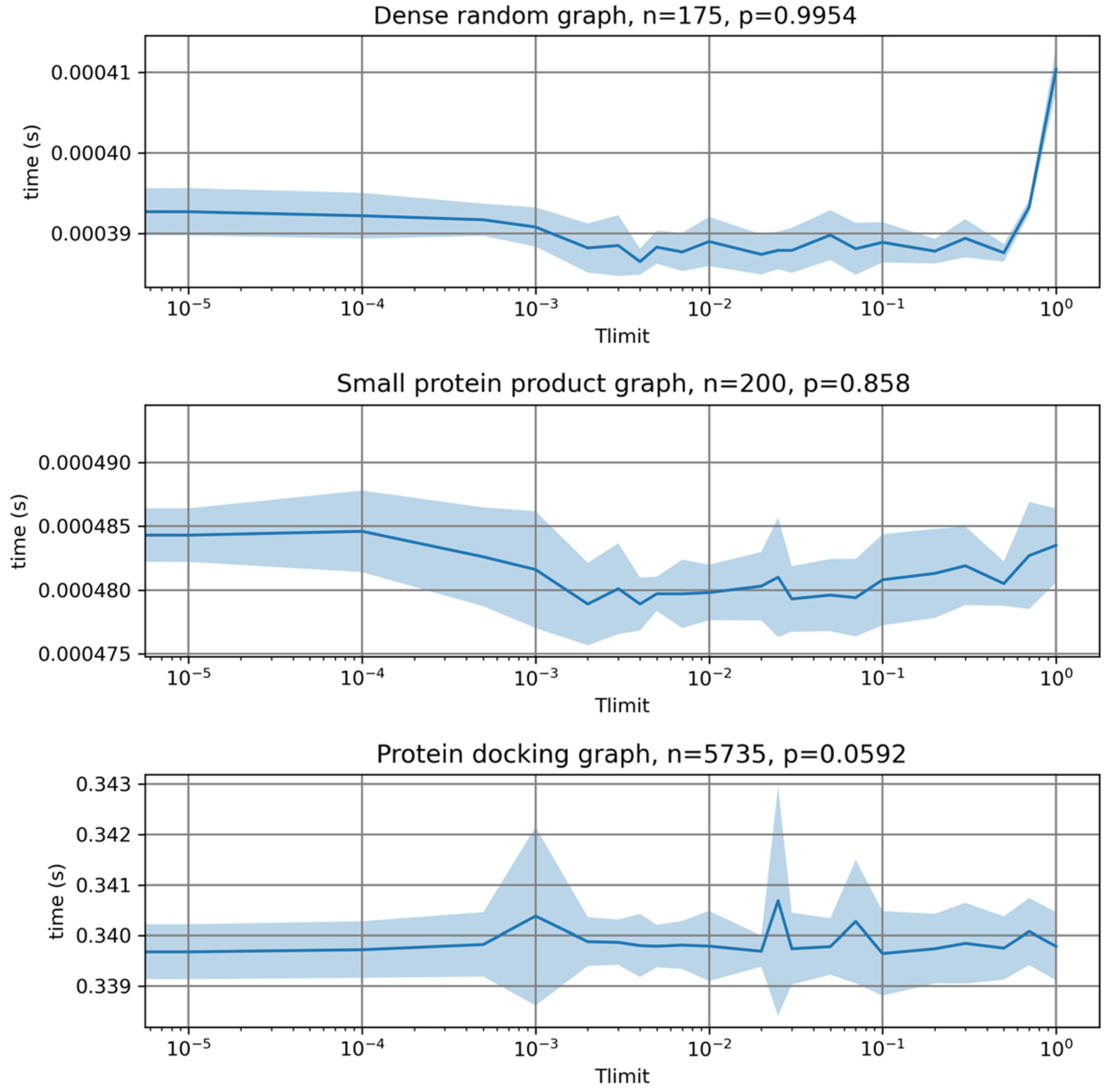
In
Figure 3 we evaluate the impact of the initial sorting of vertices on the time required for MCQD to finish searching. We found that initial sorting of vertices has no significant impact on the time needed by MCQD to find the maximum clique.
3.4. Evaluation of the Effect of Machine Learning Models on Validation Sets
We perform an evaluation of the trained machine learning models we presented. The models are evaluated using the R
2 score on the validation set, which contains graphs from different domains. This value (also called coefficient of determination) is used in statistics to evaluate statistical models. Values of R
2 typically range from 0 to 1, with 1 being the best possible value. If the model predicts the mean of the data (constant value), the R
2 value is 0. The value can also be negative if the model does not perform as well as the mean of the data. The results of our evaluation are shown in
Table 2.
We find that the model GAT achieves the highest R2 value, with any machine learning model performing better than the standard MCQD parameter choice, which is nearly equal to 0. Thus, we expect the GAT model to perform the best, while the other models in the test set are not as fast. In the next section, we evaluate the models based on the time they take to find the maximum clique.
4. Results
Our Maximum Clique Dynamic–Machine Learning (MCQD-ML) algorithm was implemented in Python (ML part) and C++ (MCQD part) and uses only 1 CPU core. Here we evaluated the MCQD-ML algorithm on several previously described sets and compared the results with the standard MCQD algorithm. The MCQD algorithm was extensively compared and benchmarked [
1,
2,
4]. The computational experiments were performed on an AMD Ryzen 9 3900X 12-core with a CPU frequency of 2 GHz. The MCQD-ML maximum clique algorithm was compared with the original MCQD algorithm on random graphs, protein product graphs, and molecular docking graphs. We limited the time available for the algorithms to 2000 s. To compare the performance of the algorithms, we use two metrics: (i) the speed-up on a test set, i.e., the time taken by the MCQD algorithm to find the maximum clique for each graph in a test set divided by the time taken by the MCQD-ML algorithm to find the maximum clique on a given set of graphs and (ii) the average speed-up on a test set is calculated by taking the speed-up of the MCQD-ML algorithm for each graph and averaging it over all graphs.
We used various machine learning models to predict the value of the Tlimit parameter, and then used this value in the MCQD-ML algorithm to evaluate its performance on several test sets, including random graphs, protein product graphs, and molecular docking graphs. We compared it with the basic MCQD algorithm with default value Tlimit = 0.025. MCQD-ML is implemented with the following machine learning models: XGBoost (XGB), Graph Convolutional Neural Network (GCN), Graph Attention Neural Network (GAT), Graph Isomorphism Network (GIN), and Support Vector Regressor with the Weisfeiler–Lehman Kernel (SVR-WL). For each model, we record the time it takes MCQD to find the maximum clique with a predicted value of the parameter Tlimit.
4.1. Dense Random Graphs
In a series of tests with dense random graphs, we found that GAT outperforms other models, including the original MCQD algorithm. The faster speed of GAT compared to MCQD is not great, as GAT is about 18% faster on average and only 4% faster on the entire test set of dense random graphs.
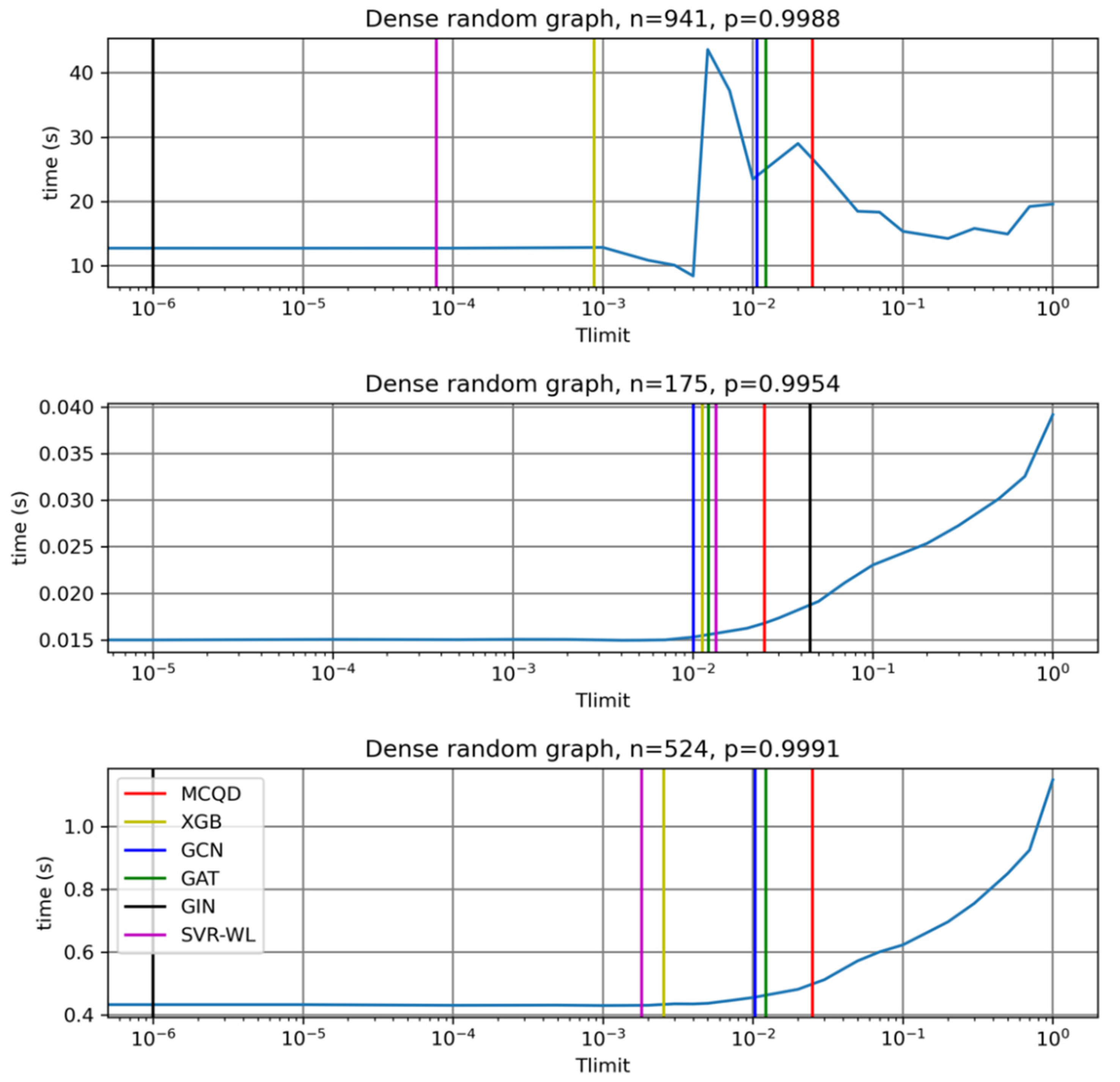
From
Table 3 and
Figure 4 we can see that the default MCQD algorithm is nearly optimal for some graphs and almost two times slower compared to tests with a better choice of the value of the parameter. There exists no Tlimit for which MCQD will find the maximum clique substantially faster.
4.2. Small Protein Product Graphs
From
Table 4 it can be observed that most ML models fail to reach the performance of the default MCQD algorithm.
4.3. Protein Product Graphs
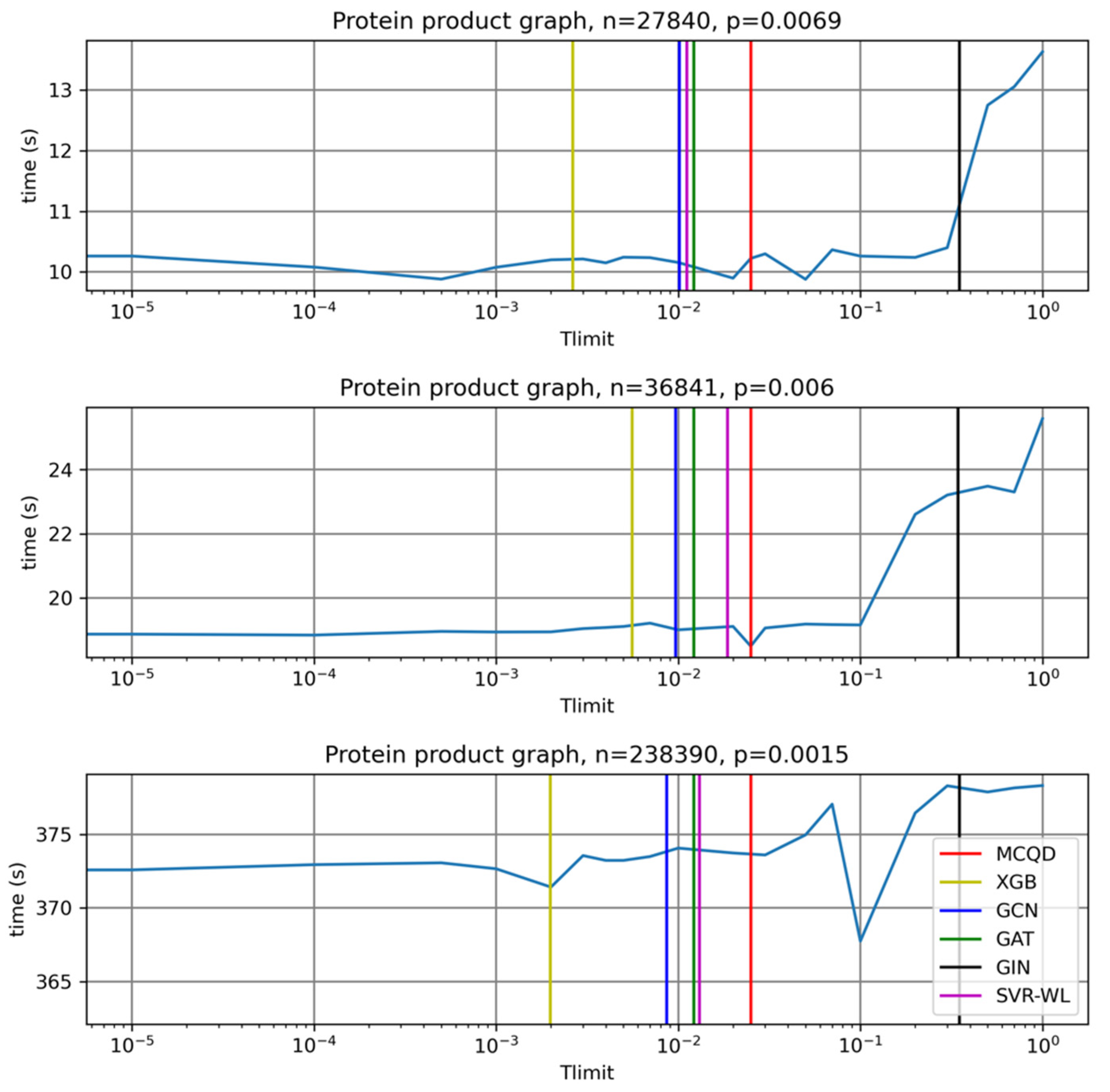
In
Table 5 and
Figure 5 we observe that any substantial speed-ups on product graphs are not achievable because the default value of parameter Tlimit is almost optimal for all product graphs in the test set.
4.4. Molecular Docking Graphs
On the test set of molecular docking graphs, we observe in
Table 6 that the GAT model and SVR-WL outperform every other model, including the MCQD algorithm. The performance of GAT and SVR-WL is almost two times faster with the whole test set, and 34% faster on average. On
Figure 6 we observe that the molecular docking graphs vary in the optimal parameter value. While on a graph with 1779 nodes the default value of the parameter is nearly optimal, it is not suitable for the graph with 5309 nodes where it is more than three times slower than with the optimal parameter value.
From these experiments we see that the prediction of Tlimit is not an easy task and differs between graphs from the same general domain. For the XGB model, we conclude that it does not have sufficient information about the graph to be able to predict a good Tlimit value. For models GCN and GIN, we hypothesize that due to their expressive power (GIN, for example, can distinguish between isomorphic graphs), they are harder to train with relatively small sets and thus perform more poorly than, for example, the GAT model.
4.5. Weighted Molecular Docking Graphs
On a test set of weighted molecular docking graphs, we observed that unlike with the set of unweighted docking graphs, there are only minor speed-ups with the GAT model (
Table 7).
From these experiments, above we can see that we can speed up the maximum clique search with MCQD by augmenting it with the GAT model. The speed-ups were achieved on random graphs and docking graphs, while on other graph domains we saw very little improvement.
5. Conclusions
We have developed a new approach to find the maximum clique on a protein graph using both neural networks and artificial intelligence approaches. It is a new approach that has not been developed before, and its results show a remarkable speed-up in determining the correct maximum clique on the product graph. Therefore, we expect that this approach will be widely applicable in various scientific fields, such as computer science.
Having fast algorithms that solve maximum clique problem is of great importance in the discovery of new drugs and of protein behavior. We applied a couple of machine learning methods on a regression problem in order to speed up a dynamic algorithm for maximum clique search and obtained several variants of the new MCQD-ML algorithm, which we applied to graph topologies that are particularly important in bioinformatics.
We concluded that improvements using deep learning methods are possible. The most well-suited model that we tested is the graph attention network (GAT), which can speed up the maximum clique search on average by 18% on random graphs and by 34% on docking graphs. The computational cost introduced with the machine learning model is negligible compared to the maximum clique search.
From experiments on protein product graphs, we can assume that further improvements using the same MCQD algorithm are unlikely to be achievable. In further work, we could improve the quality of the set with more samples from different graph topologies such as social network graphs. It would be interesting to test possible speed-ups on other algorithms that operate on a domain of graphs and use empirically determined parameters that determine the progress of the algorithm.
Author Contributions
Conceptualization, M.G., D.J. and J.K.; methodology, K.R. (Kristjan Reba), M.G., K.R. (Kati Rozman), D.J. and J.K.; software, K.R. (Kristjan Reba), M.G., K.R. (Kati Rozman), D.J. and J.K.; validation, K.R. (Kristjan Reba), M.G. and K.R. (Kati Rozman); formal analysis, M.G., K.R. (Kati Rozman), D.J. and J.K.; investigation, K.R. (Kristjan Reba), M.G., K.R. (Kati Rozman), D.J. and J.K.; resources, M.G., D.J. and J.K.; data curation, K.R. (Kristjan Reba), M.G., K.R. (Kati Rozman), D.J. and J.K.; writing—original draft preparation, K.R. (Kristjan Reba), M.G., D.J. and J.K.; writing—review and editing, K.R. (Kristjan Reba), M.G., K.R. (Kati Rozman), D.J. and J.K.; visualization, K.R. (Kristjan Reba); supervision, J.K., M.G. and D.J.; project administration, J.K., M.G. and D.J.; funding acquisition, J.K. and D.J. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the Slovenian Research Agency Project, grants N1-0142, L7-8269, N1-0209, J1-1715, and J1-9186.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Conflicts of Interest
The authors declare no conflict of interest.
References
- Wu, Q.; Hao, J.-K. A review on algorithms for maximum clique problems. Eur. J. Oper. Res. 2015, 242, 693–709. [Google Scholar] [CrossRef]
- Depolli, M.; J Konc, K.R.; Trobec, R.; Janezic, D. Exact parallel maximum clique algorithm for general and protein graphs. J. Chem. Inf. Model. 2013, 53, 2217–2228. [Google Scholar] [CrossRef]
- Butenko, S.; Wilhelm, W.E. Clique-detection models in computational biochemistry and genomics. Eur. J. Oper. Res. 2006, 173, 1–17. [Google Scholar] [CrossRef]
- Konc, J.; Janezic, D. An improved branch and bound algorithm for the maximum clique problem. Match Commun. Math. Comput. Chem. 2007, 58, 569–590. [Google Scholar]
- Prates, M.; Avelar, P.H.; Lemos, H.; Lamb, L.C.; Vardi, M.Y. Learning to solve np-complete problems: A graph neural network for decision tsp. Proc. AAAI Conf. Artif. Intell. 2019, 33, 4731–4738. [Google Scholar] [CrossRef]
- Walteros, J.L.; Buchanan, A. Why is maximum clique often easy in practice? Oper. Res. 2020, 68, 1625–1931. [Google Scholar] [CrossRef]
- Li, C.-M.; Jiang, H.; Manyà, F. On minimization of the number of branches in branch-and-bound algorithms for the maximum clique problem. Comput. Oper. Res. 2017, 84, 1–15. [Google Scholar] [CrossRef]
- Tomita, E.; Seki, T. An efficient branch-and-bound algorithm for finding a maxi-mum clique. In Proceedings of the International Conference on Discrete Mathematics and Theoretical Computer Science, Dijon, France, 7–12 July 2003; Springer: Berlin/Heidelberg, Germany, 2003; pp. 278–289. [Google Scholar]
- Tomita, E.; Matsuzaki, S.; Nagao, A.; Ito, H.; Wakatsuki, M. A much faster algorithm for finding a maximum clique with computational experiments. J. Inf. Process. 2017, 25, 667–677. [Google Scholar] [CrossRef] [Green Version]
- Carraghan, R.; Pardalos, P.M. An exact algorithm for the maximum clique problem. Oper. Res. Lett. 1990, 9, 375–382. [Google Scholar] [CrossRef]
- Li, C.-M.; Quan, Z. An efficient branch-and-bound algorithm based on maxsat for the maximum clique problem. Proc. AAAI Conf. Artif. Intell. 2010, 24, 128–133. [Google Scholar]
- Segundo, P.S.; Lopez, A.; Pardalos, P.M. A new exact maximum clique algorithm for large and massive sparse graphs. Comput. Oper. Res. 2016, 66, 81–94. [Google Scholar] [CrossRef]
- Bengio, Y.; Lodi, A.; Prouvost, A. Machine learning for combinatorial optimization: A methodological tour d’horizon. Eur. J. Oper. Res. 2021, 290, 405–421. [Google Scholar] [CrossRef]
- Abe, K.; Xu, Z.; Sato, I.; Sugiyama, M. Solving np-hard problems on graphs with extended AlphaGO Zero. arXiv 2019, arXiv:1905.11623. [Google Scholar]
- Zhou, J.; Cui, G.; Zhang, Z.; Yang, C.; Liu, Z.; Wang, L.; Li, C.; Sun, M. Graph neural networks: A review of methods and applications. arXiv 2018, arXiv:1812.08434. [Google Scholar] [CrossRef]
- Johnson, D.S.; Trick, M.A. Cliques, Coloring, and Satisfiability: Second DIMACS Implementation Challenge, 11–13 October 1993; American Mathematical Society: Boston, MA, USA, 1996. [Google Scholar]
- Konc, J.; Janežič, D. ProBiS algorithm for detection of structurally similar protein binding sites by local structural alignment. Bioinformatics 2010, 26, 1160–1168. [Google Scholar] [CrossRef] [PubMed]
- Fine, J.; Konc, J.; Samudrala, R.; Chopra, G. CANDOCK: Chemical atomic network-based hierarchical flexible docking algorithm using generalized statistical potentials. J. Chem. Inf. Model. 2020, 60, 1509–1527. [Google Scholar] [CrossRef]
- Lešnik, S.; Konc, J. In silico laboratory: Tools for similarity-based drug discovery. In Targeting Enzymes for Pharmaceutical Development; Springer: Berlin/Heidelberg, Germany, 2020; pp. 1–28. [Google Scholar]
- Cortes, C.; Vapnik, V. Support-vector networks. Mach. Learn. 1995, 20, 273–297. [Google Scholar] [CrossRef]
- Friedman, J.H. Greedy function approximation: A gradient boosting machine. Ann. Stat. 2001, 29, 1189–1232. [Google Scholar] [CrossRef]
- Chen, T.; Guestrin, C. Xgboost: A scalable tree boosting system. In Proceedings of the 22nd ACM Sigkdd International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13 August 2016; pp. 785–794. [Google Scholar]
- Shervashidze, N.; Vishwanathan, S.; Petri, T.; Mehlhorn, K.; Borgwardt, K. Efficient graphlet kernels for large graph comparison. Artif. Intell. Stat. PMLR 2009, 5, 488–495. [Google Scholar]
- Shervashidze, N.; Schweitzer, P.; van Leeuwen, E.J.; Mehlhorn, K.; Borgwardt, K.M. Weisfeiler-lehman graph kernels. J. Mach. Learn. Res. 2011, 12, 2539–2561. [Google Scholar]
- Leskovec, J. Stanford cs224w: Machine Learning with Graphs. Traditional Methods for Machine Learning in Graphs 2019. Available online: http://web.stanford.edu/class/cs224w/ (accessed on 1 May 2021).
- Xu, K.; Hu, W.; Leskovec, J.; Jegelka, S. How powerful are graph neural networks? arXiv 2018, arXiv:1810.00826. [Google Scholar]
- Khalil, E.; Dai, H.; Zhang, Y.; Dilkina, B.; Song, L. Learning combinatorial optimization algorithms over graphs. Adv. Neural Inf. Process. Syst. 2017, 1704, 6348–6358. [Google Scholar]
- Kipf, T.N.; Welling, M. Semi-supervised classification with graph convolutional networks. arXiv 2016, arXiv:1609.02907. [Google Scholar]
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, L.; Polosukhin, I. Attention is all you need. arXiv 2017, arXiv:1706.03762. [Google Scholar]
- Veličković, P.; Cucurull, G.; Casanova, A.; Romero, A.; Lio, P.; Bengio, Y. Graph attention networks. arXiv 2016, arXiv:1609.02907. [Google Scholar]
| Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).