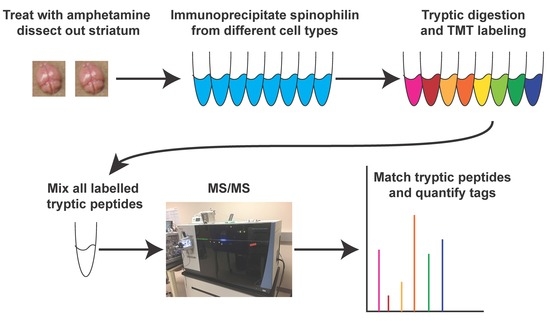
Proteomic Analysis of the Spinophilin Interactome in Rodent Striatum Following Psychostimulant Sensitization
Abstract
1. Introduction
2. Materials and Methods
2.1. Animals—HA Spinophilin Mice Generation
2.2. Animals—Proteomics Studies
2.3. Animals—Immunoblotting Studies
2.4. d-Amphetamine Sensitization
2.5. Brain Tissue Lysis
2.6. Transfections
2.7. Immunoprecipitations
2.8. TMT Labeling and Mass Spectrometry
2.9. Immunoblotting
2.10. Pathway Analysis
2.11. Statistics
3. Results
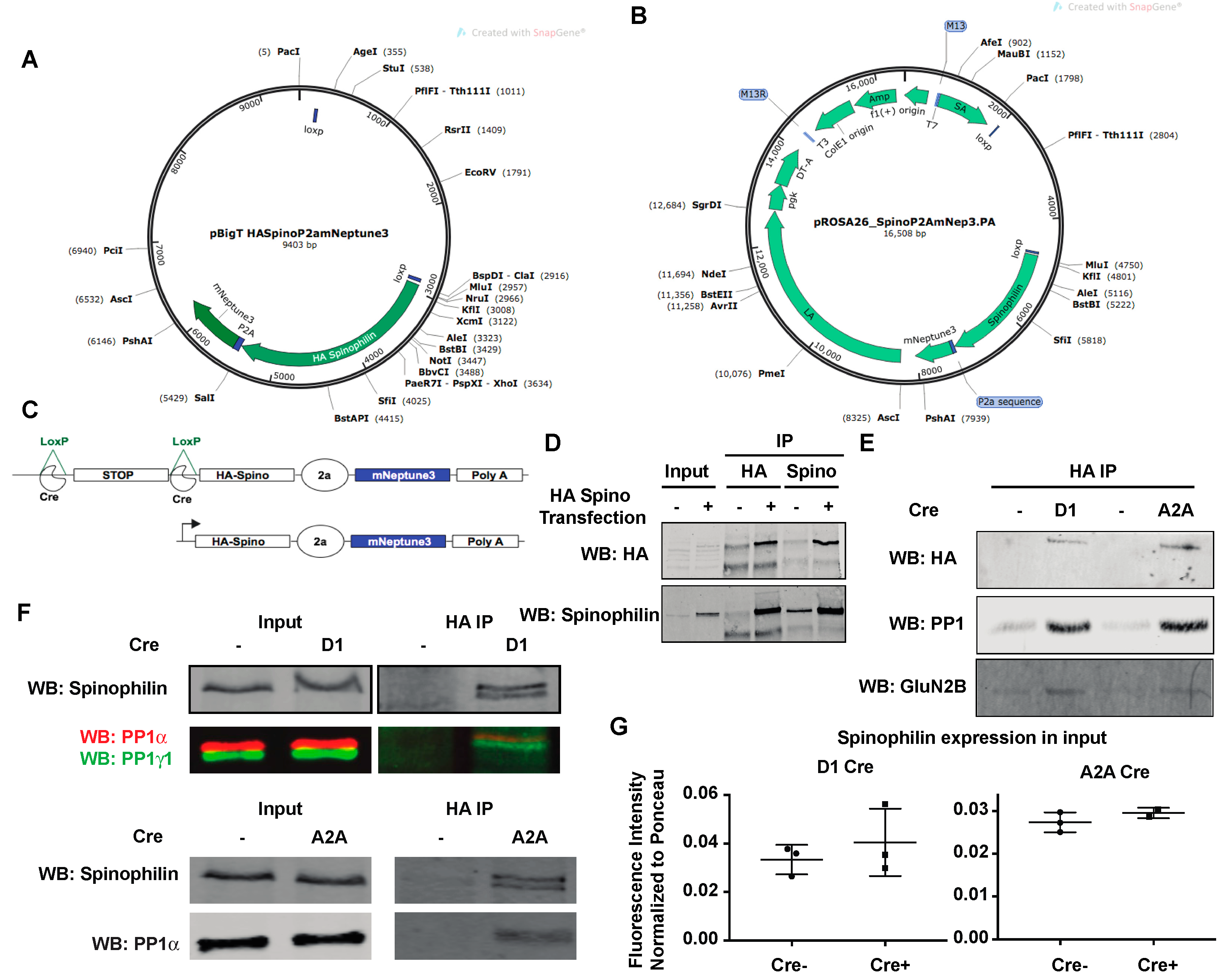
3.1. Generation of HA-Tagged Spinophilin Mice
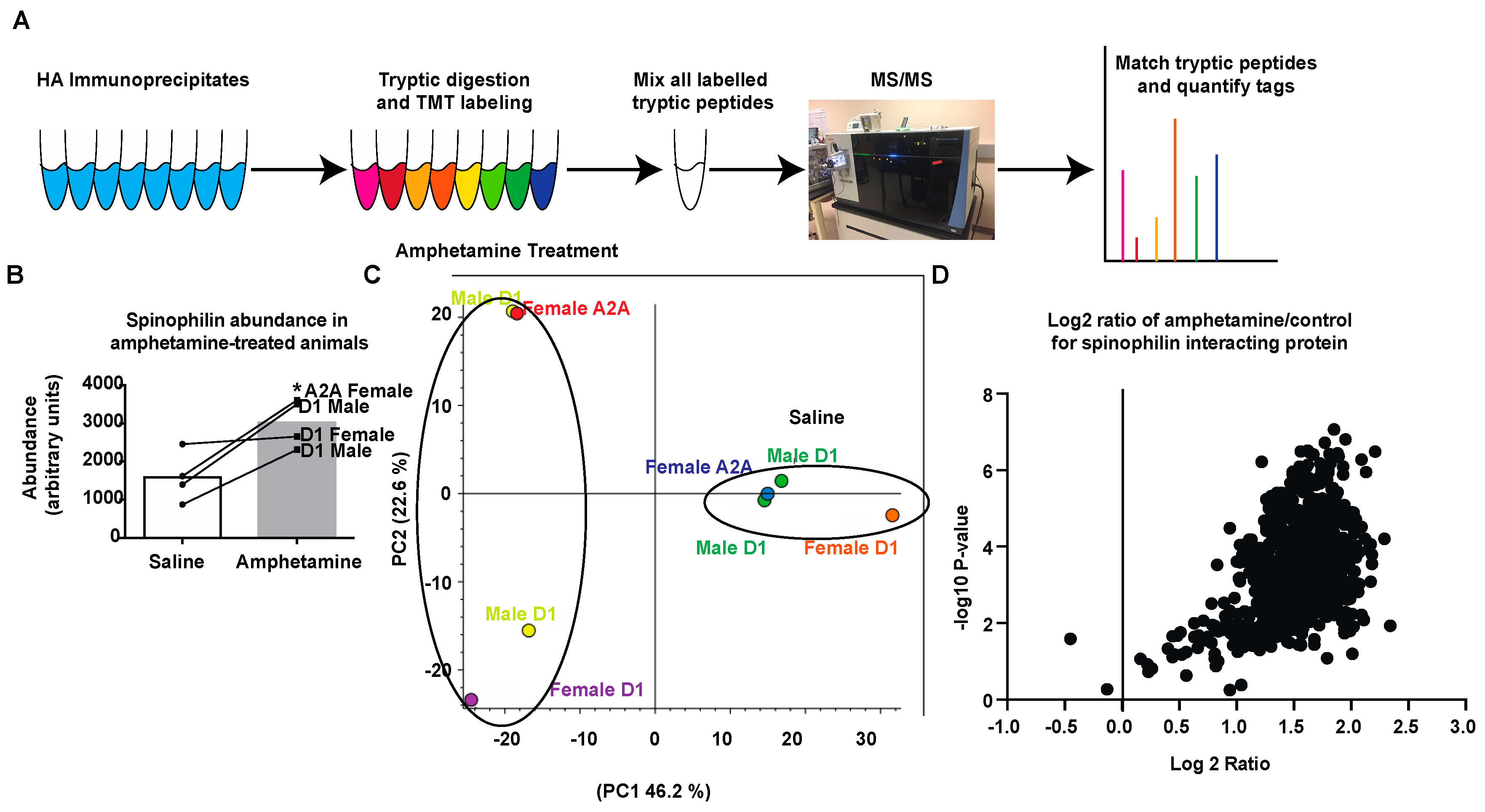
3.2. Amphetamine Modulates Spinophilin Expression and Interactions
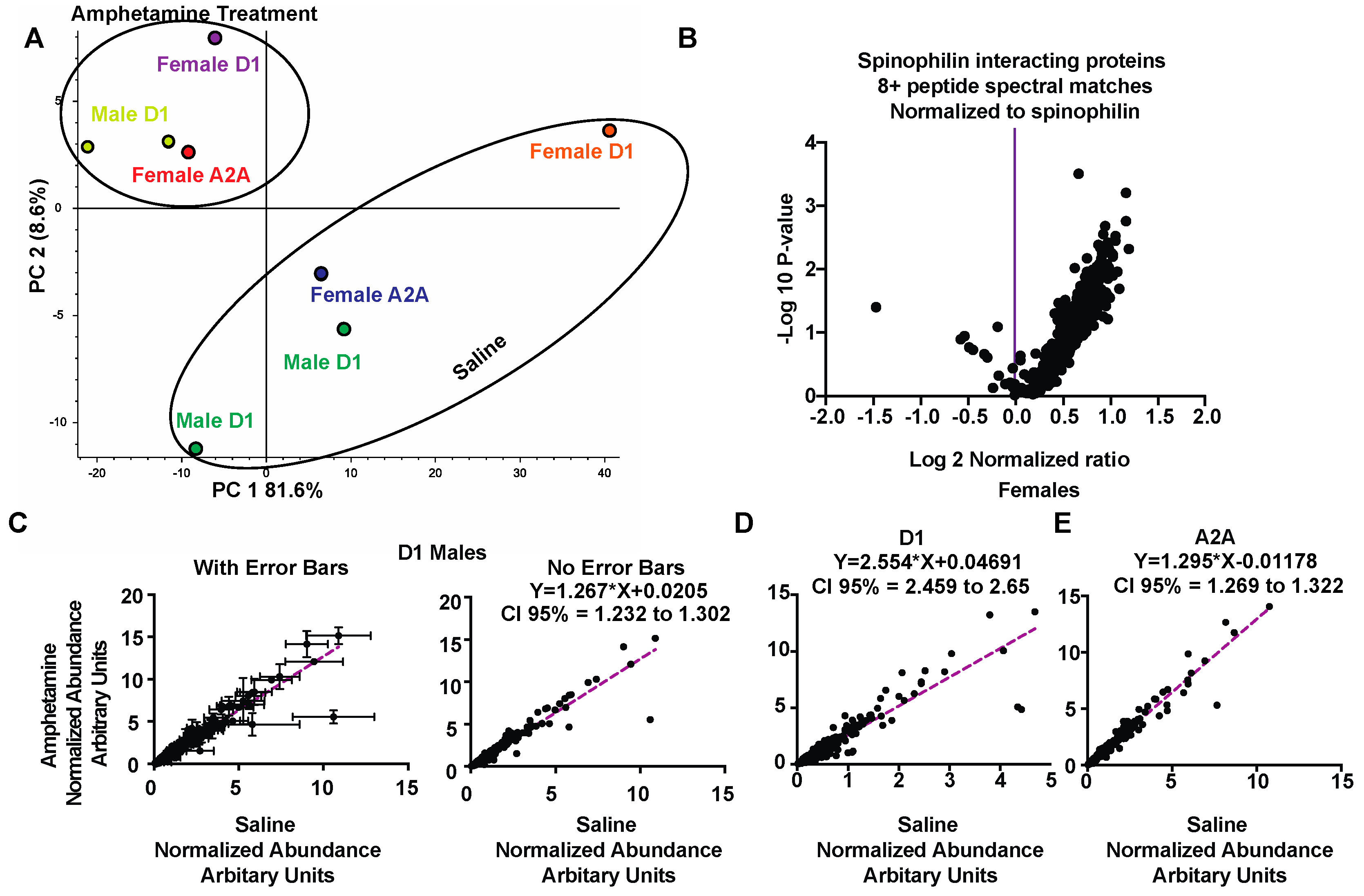
3.3. Regulation of Spinophilin Interactions by Amphetamine
3.4. Pathway and GO Analysis of Spinophilin Interacting Proteins Enhanced by Amphetamine
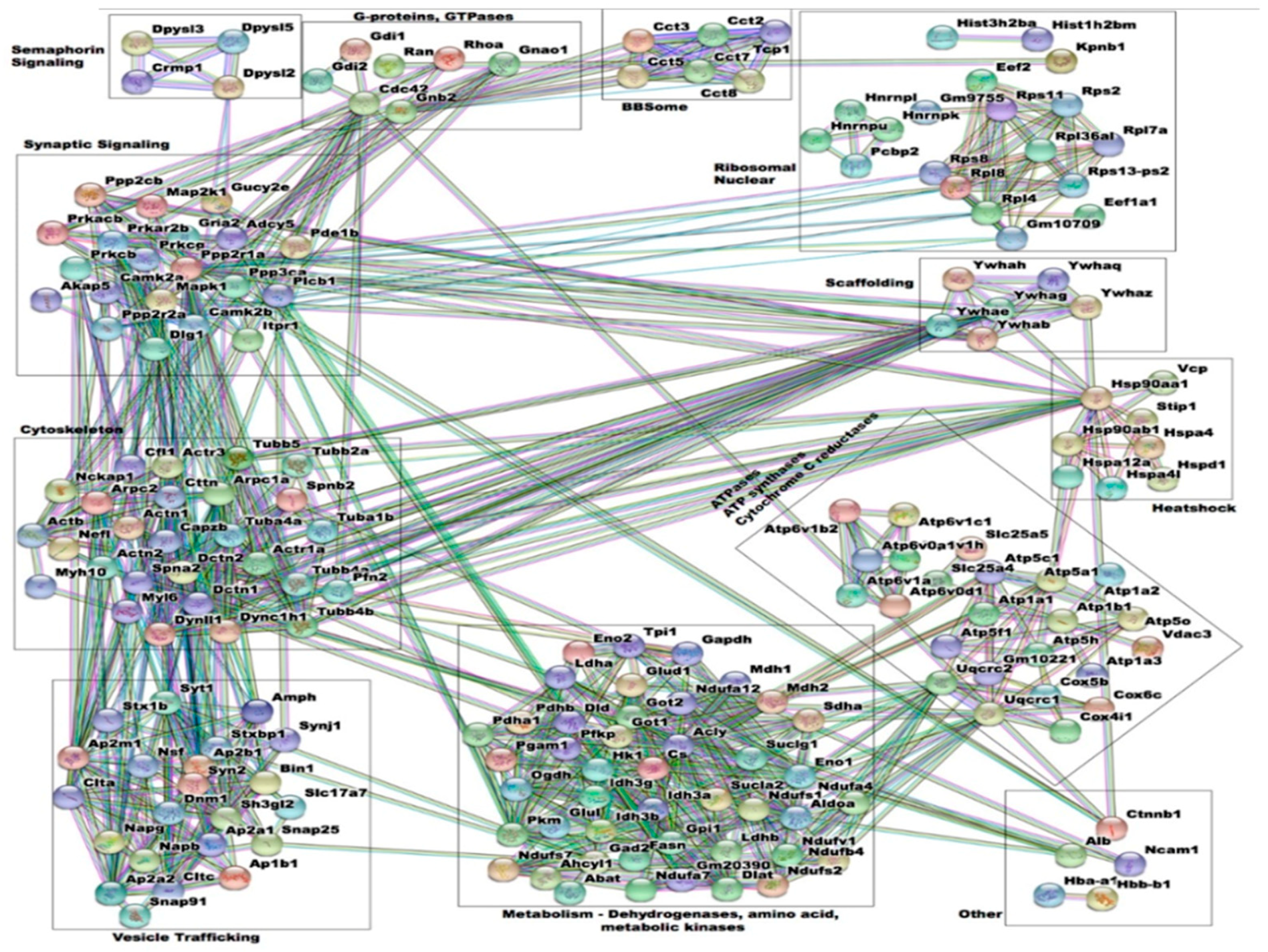
3.5. Interactome Analysis of Spinophilin Interacting Proteins Enhanced by Amphetamine
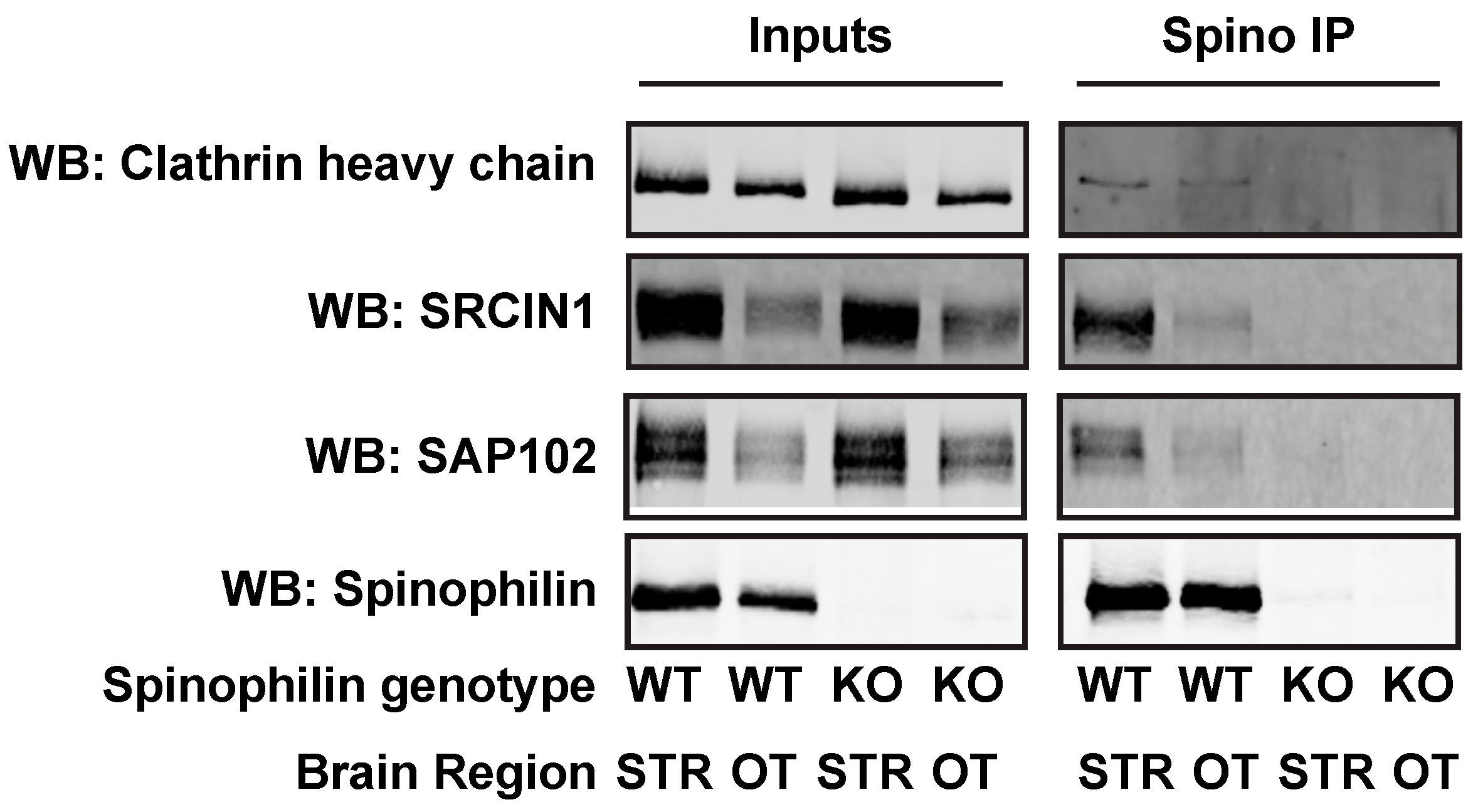
3.6. Alteration and Validation of Novel Spinophilin Interacting Proteins
4. Discussion
4.1. Spinophilin Functional Localization
4.2. Regulation of the Spinophilin Interactome Following Amphetamine Treatment
4.3. Classes and Specific Spinophilin Protein Interactions that are Modulated by Amphetamine
4.4. Direct and Indirect Pathway Striatal MSNs and Spinophilin Interactions
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Degenhardt, L.; Hall, W. Extent of illicit drug use and dependence, and their contribution to the global burden of disease. Lancet 2012, 379, 55–70. [Google Scholar] [CrossRef]
- Ashok, A.H.; Mizuno, Y.; Volkow, N.D.; Howes, O.D. Association of Stimulants With Dopaminergic Alterations in Users of Cocaine, Amphetamine, and Methamphetamine: A Systematic Review and Meta-analysis. JAMA Psychiatry 2017, 74, 511–519. [Google Scholar] [CrossRef] [PubMed]
- Burke, A.R.; Miczek, K.A. Stress in adolescence and drugs of abuse in rodent models: Role of dopamine, CRF, and HPA axis. Psychopharmacology 2014, 231, 1557–1580. [Google Scholar] [CrossRef] [PubMed]
- Gerdeman, G.L.; Partridge, J.G.; Lupica, C.R.; Lovinger, D.M. It could be habit forming: Drugs of abuse and striatal synaptic plasticity. Trends Neurosci. 2003, 26, 184–192. [Google Scholar] [CrossRef]
- Sanchez-Ramos, J. Chapter Seven–Neurologic Complications of Psychomotor Stimulant Abuse. In International Review of Neurobiology; Taba, P., Lees, A., Sikk, K., Eds.; Academic Press: Amsterdam, The Netherlands, 2015; Volume 120, pp. 131–160. [Google Scholar]
- Chiodi, V.; Mallozzi, C.; Ferrante, A.; Chen, J.F.; Lombroso, P.J.; Di Stasi, A.M.M.; Popoli, P.; Domenici, M.R. Cocaine-Induced Changes of Synaptic Transmission in the Striatum are Modulated by Adenosine A(2A) Receptors and Involve the Tyrosine Phosphatase STEP. Neuropsychopharmacology 2014, 39, 569–578. [Google Scholar] [CrossRef] [PubMed]
- Centonze, D.; Picconi, B.; Baunez, C.; Borrelli, E.; Pisani, A.; Bernardi, G.; Calabresi, P. Cocaine and Amphetamine Depress Striatal GABAergic Synaptic Transmission through D2 Dopamine Receptors. Neuropsychopharmacology 2002, 26, 164. [Google Scholar] [CrossRef]
- Ungless, M.A.; Whistler, J.L.; Malenka, R.C.; Bonci, A. Single cocaine exposure in vivo induces long-term potentiation in dopamine neurons. Nature 2001, 411, 583. [Google Scholar] [CrossRef]
- Di Chiara, G.; Imperato, A. Drugs abused by humans preferentially increase synaptic dopamine concentrations in the mesolimbic system of freely moving rats. Proc. Natl. Acad. Sci. USA 1988, 85, 5274–5278. [Google Scholar] [CrossRef]
- Volkow, N.D.; Wang, G.-J.; Fowler, J.S.; Logan, J.; Gatley, S.J.; Wong, C.; Hitzemann, R.; Pappas, N.R. Reinforcing Effects of Psychostimulants in Humans Are Associated with Increases in Brain Dopamine and Occupancy of D2 Receptors. J. Pharmacol. Exp. Ther. 1999, 291, 409–415. [Google Scholar]
- Siviy, S.M.; McDowell, L.S.; Eck, S.R.; Turano, A.; Akopian, G.; Walsh, J.P. Effects of amphetamine on striatal dopamine release, open-field activity, and play in Fischer 344 and Sprague–Dawley rats. Behav. Pharmacol. 2015, 26, 720–732. [Google Scholar] [CrossRef]
- dela Peña, I.; Gevorkiana, R.; Shi, W.-X. Psychostimulants affect dopamine transmission through both dopamine transporter-dependent and independent mechanisms. Eur. J. Pharmacol. 2015, 764, 562–570. [Google Scholar] [CrossRef]
- Cass, W.A.; Gerhardt, G.A.; Mayfield, R.D.; Curella, P.; Zahniser, N.R. Differences in Dopamine Clearance and Diffusion in Rat Striatum and Nucleus Accumbens Following Systemic Cocaine Administration. J. Neurochem. 1992, 59, 259–266. [Google Scholar] [CrossRef] [PubMed]
- Bertran-Gonzalez, J.; Bosch, C.; Maroteaux, M.; Matamales, M.; Hervé, D.; Valjent, E.; Girault, J.-A. Opposing patterns of signaling activation in dopamine D1 and D2 receptor-expressing striatal neurons in response to cocaine and haloperidol. J. Neurosci. 2008, 28, 5671. [Google Scholar] [CrossRef] [PubMed]
- Kalivas, P.W.; Stewart, J. Dopamine transmission in the initiation and expression of drug- and stress-induced sensitization of motor activity. Brain Res. Rev. 1991, 16, 223–244. [Google Scholar] [CrossRef]
- Robinson, T.E.; Becker, J.B. Enduring changes in brain and behavior produced by chronic amphetamine administration: A review and evaluation of animal models of amphetamine psychosis. Brain Res. Rev. 1986, 11, 157–198. [Google Scholar] [CrossRef]
- Joshua, M.; Adler, A.; Bergman, H. The dynamics of dopamine in control of motor behavior. Curr. Opin. Neurobiol. 2009, 19, 615–620. [Google Scholar] [CrossRef] [PubMed]
- Ishiguro, A.; Inagaki, M.; Kaga, M. Stereotypic circling behavior in mice with vestibular dysfunction: Asymmetrical effects of intrastriatal microinjection of a dopamine agonist. Int. J. Neurosci. 2007, 117, 1049–1064. [Google Scholar] [CrossRef] [PubMed]
- Ferro, M.M.; Bellissimo, M.I.; Anselmo-Franci, J.A.; Angellucci, M.E.M.; Canteras, N.S.; Da Cunha, C. Comparison of bilaterally 6-OHDA- and MPTP-lesioned rats as models of the early phase of Parkinson’s disease: Histological, neurochemical, motor and memory alterations. J. Neurosci. Method. 2005, 148, 78–87. [Google Scholar] [CrossRef]
- Archer, T.; Danysz, W.; Fredriksson, A.; Jonsson, G.; Luthman, J.; Sundström, E.; Teiling, A. Neonatal 6-hydroxydopamine-induced dopamine depletions: Motor activity and performance in maze learning. Pharmacol. Biochem. Behav. 1988, 31, 357–364. [Google Scholar] [CrossRef]
- Herbin, M.; Simonis, C.; Revéret, L.; Hackert, R.; Libourel, P.-A.; Eugène, D.; Diaz, J.; de Waele, C.; Vidal, P.-P. Dopamine Modulates Motor Control in a Specific Plane Related to Support. PLoS ONE 2016, 11, e0155058. [Google Scholar] [CrossRef]
- Wong, L.S.; Eshel, G.; Dreher, J.; Ong, J.; Jackson, D.M. Role of dopamine and GABA in the control of motor activity elicited from the rat nucleus accumbens. Pharmacol. Biochem. Behav. 1991, 38, 829–835. [Google Scholar] [CrossRef]
- Rikani, A.A.; Choudhry, Z.; Choudhry, A.M.; Rizvi, N.; Ikram, H.; Mobassarah, N.J.; Tuli, S. The mechanism of degeneration of striatal neuronal subtypes in Huntington disease. Annal. Neurosci. 2014, 21, 112–114. [Google Scholar] [CrossRef] [PubMed]
- Belluscio, M.A.; Escande, M.V.; Keifman, E.; Riquelme, L.A.; Murer, M.G.; Zold, C.L. Oscillations in the basal ganglia in Parkinson’s disease: Role of the striatum. Basal Ganglia 2014, 3, 203–212. [Google Scholar] [CrossRef]
- Greenberg, B.D.; Gabriels, L.A.; Malone, D.A., Jr.; Rezai, A.R.; Friehs, G.M.; Okun, M.S.; Shapira, N.A.; Foote, K.D.; Cosyns, P.R.; Kubu, C.S.; et al. Deep brain stimulation of the ventral internal capsule/ventral striatum for obsessive-compulsive disorder: Worldwide experience. Mol. Psychiatry 2008, 15, 64. [Google Scholar] [CrossRef] [PubMed]
- Yip, S.W.; Worhunsky, P.D.; Rogers, R.D.; Goodwin, G.M. Hypoactivation of the Ventral and Dorsal Striatum During Reward and Loss Anticipation in Antipsychotic and Mood Stabilizer-Naive Bipolar Disorder. Neuropsychopharmacology 2014, 40, 658. [Google Scholar] [CrossRef] [PubMed]
- Yager, L.M.; Garcia, A.F.; Wunsch, A.M.; Ferguson, S.M. The ins and outs of the striatum: Role in drug addiction. Neuroscience 2015, 301, 529–541. [Google Scholar] [CrossRef] [PubMed]
- Burguière, E.; Monteiro, P.; Mallet, L.; Feng, G.; Graybiel, A.M. Striatal circuits, habits, and implications for obsessive-compulsive disorder. Curr. Opin. Neurobiol. 2015, 0, 59–65. [Google Scholar] [CrossRef] [PubMed]
- Graybiel, A.M.; Grafton, S.T. The Striatum: Where Skills and Habits Meet. CSH Perspect. Biol. 2015, 7, a021691. [Google Scholar] [CrossRef] [PubMed]
- Balleine, B.W.; O’Doherty, J.P. Human and Rodent Homologies in Action Control: Corticostriatal Determinants of Goal-Directed and Habitual Action. Neuropsychopharmacology 2010, 35, 48–69. [Google Scholar] [CrossRef] [PubMed]
- Durieux, P.F.; Bearzatto, B.; Guiducci, S.; Buch, T.; Waisman, A.; Zoli, M.; Schiffmann, S.N.; de Kerchove d’Exaerde, A. D2R striatopallidal neurons inhibit both locomotor and drug reward processes. Nat. Neurosci. 2009, 12, 393. [Google Scholar] [CrossRef] [PubMed]
- Flanigan, M.; LeClair, K. Shared Motivational Functions of Ventral Striatum D1 and D2 Medium Spiny Neurons. J. Neurosci. 2017, 37, 6177. [Google Scholar] [CrossRef] [PubMed]
- Ikemoto, S. Dopamine reward circuitry: Two projection systems from the ventral midbrain to the nucleus accumbens-olfactory tubercle complex. Brain Res. Rev. 2007, 56, 27–78. [Google Scholar] [CrossRef] [PubMed]
- Belin, D.; Everitt, B.J. Cocaine Seeking Habits Depend upon Dopamine-Dependent Serial Connectivity Linking the Ventral with the Dorsal Striatum. Neuron 2008, 57, 432–441. [Google Scholar] [CrossRef] [PubMed]
- Kravitz, A.V.; Kreitzer, A.C. Striatal Mechanisms Underlying Movement, Reinforcement, and Punishment. Physiology 2012, 27. [Google Scholar] [CrossRef] [PubMed]
- Gerfen, C.R.; Engber, T.M.; Mahan, L.C.; Susel, Z.; Chase, T.N.; Monsma, F.J.; Sibley, D.R. D1 and D2 dopamine receptor-regulated gene expression of striatonigral and striatopallidal neurons. Science 1990, 250, 1429. [Google Scholar] [CrossRef] [PubMed]
- Gerfen, C.R.; Scott Young, W. Distribution of striatonigral and striatopallidal peptidergic neurons in both patch and matrix compartments: An in situ hybridization histochemistry and fluorescent retrograde tracing study. Brain Res. 1988, 460, 161–167. [Google Scholar] [CrossRef]
- Lanciego, J.L.; Luquin, N.; Obeso, J.A. Functional Neuroanatomy of the Basal Ganglia. CSH Perspect. Med. 2012, 2, a009621. [Google Scholar] [CrossRef] [PubMed]
- Burns, R.S.; LeWitt, P.A.; Ebert, M.H.; Pakkenberg, H.; Kopin, I.J. The Clinical Syndrome of Striatal Dopamine Deficiency. N. Eng. J. Med. 1985, 312, 1418–1421. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.Y.; Wang, E.A.; Cepeda, C.; Levine, M.S. Dopamine imbalance in Huntington’s disease: A mechanism for the lack of behavioral flexibility. Front. Neurosci. 2013, 7, 114. [Google Scholar] [CrossRef]
- Surmeier, D.J.; Graves, S.M.; Shen, W. Dopaminergic modulation of striatal networks in health and Parkinson’s disease. Curr. Opin. Neurobiol. 2014, 29, 109–117. [Google Scholar] [CrossRef] [PubMed]
- Goto, A.; Nakahara, I.; Yamaguchi, T.; Kamioka, Y.; Sumiyama, K.; Matsuda, M.; Nakanishi, S.; Funabiki, K. Circuit-dependent striatal PKA and ERK signaling underlies rapid behavioral shift in mating reaction of male mice. Proc. Natl. Acad. Sci. USA 2015, 112, 6718–6723. [Google Scholar] [CrossRef] [PubMed]
- Flores-Hernández, J.; Cepeda, C.; Hernández-Echeagaray, E.; Calvert, C.R.; Jokel, E.S.; Fienberg, A.A.; Greengard, P.; Levine, M.S. Dopamine Enhancement of NMDA Currents in Dissociated Medium-Sized Striatal Neurons: Role of D1 Receptors and DARPP-32. J. Neurophysiol. 2002, 88, 3010–3020. [Google Scholar] [CrossRef]
- Surmeier, D.J.; Ding, J.; Day, M.; Wang, Z.; Shen, W. D1 and D2 dopamine-receptor modulation of striatal glutamatergic signaling in striatal medium spiny neurons. Trends Neurosci. 2007, 30, 228–235. [Google Scholar] [CrossRef]
- Bahuguna, J.; Weidel, P.; Morrison, A. Exploring the role of striatal D1 and D2 medium spiny neurons in action selection using a virtual robotic framework. Eur. J. Neurosci. 2018. [Google Scholar] [CrossRef] [PubMed]
- Peti, W.; Nairn, A.C.; Page, R. Structural Basis for Protein Phosphatase 1 Regulation and Specificity. FEBS J. 2013, 280, 596–611. [Google Scholar] [CrossRef] [PubMed]
- Woolfrey, K.M.; Dell’Acqua, M.L. Coordination of Protein Phosphorylation and Dephosphorylation in Synaptic Plasticity. J. Biol. Chem. 2015, 290, 28604–28612. [Google Scholar] [CrossRef] [PubMed]
- Allen, P.B.; Ouimet, C.C.; Greengard, P. Spinophilin, a novel protein phosphatase 1 binding protein localized to dendritic spines. Proc. Natl. Acad. Sci. USA 1997, 94, 9956–9961. [Google Scholar] [CrossRef]
- Hsieh-Wilson, L.C.; Allen, P.B.; Watanabe, T.; Nairn, A.C.; Greengard, P. Characterization of the Neuronal Targeting Protein Spinophilin and Its Interactions with Protein Phosphatase-1. Biochemistry 1999, 38, 4365–4373. [Google Scholar] [CrossRef]
- Colbran, R.J.; Bass, M.A.; McNeill, R.B.; Bollen, M.; Zhao, S.; Wadzinski, B.E.; Strack, S. Association of brain protein phosphatase 1 with cytoskeletal targeting/regulatory subunits. J. Neurochem. 1997, 69, 920–929. [Google Scholar] [CrossRef]
- Morishita, W.; Connor, J.H.; Xia, H.; Quinlan, E.M.; Shenolikar, S.; Malenka, R.C. Regulation of Synaptic Strength by Protein Phosphatase 1. Neuron 2001, 32, 1133–1148. [Google Scholar] [CrossRef]
- Ragusa, M.J.; Dancheck, B.; Critton, D.A.; Nairn, A.C.; Page, R.; Peti, W. Spinophilin directs Protein Phosphatase 1 specificity by blocking substrate binding sites. Nat. Struct. Mol. Biol. 2010, 17, 459–464. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.D.; Huang, Q.; Roadcap, D.W.; Shenolikar, S.S.; Xia, H. Actin-associated neurabin–protein phosphatase-1 complex regulates hippocampal plasticity. J. Neurochem. 2006, 98, 1841–1851. [Google Scholar] [CrossRef] [PubMed]
- Muly, E.C.; Smith, Y.; Allen, P.; Greengard, P. Subcellular distribution of spinophilin immunolabeling in primate prefrontal cortex: Localization to and within dendritic spines. J. Comp. Neurol. 2004, 469, 185–197. [Google Scholar] [CrossRef] [PubMed]
- Muly, E.C.; Allen, P.; Mazloom, M.; Aranbayeva, Z.; Greenfield, A.T.; Greengard, P. Subcellular Distribution of Neurabin Immunolabeling in Primate Prefrontal Cortex: Comparison with Spinophilin. Cereb. Cortex 2004, 14, 1398–1407. [Google Scholar] [CrossRef] [PubMed]
- Feng, J.; Yan, Z.; Ferreira, A.; Tomizawa, K.; Liauw, J.A.; Zhuo, M.; Allen, P.B.; Ouimet, C.C.; Greengard, P. Spinophilin regulates the formation and function of dendritic spines. Proc. Natl. Acad. Sci. USA 2000, 97, 9287–9292. [Google Scholar] [CrossRef]
- Roze, E.; Cahill, E.; Martin, E.; Bonnet, C.; Vanhoutte, P.; Betuing, S.; Caboche, J. Huntington’s Disease and Striatal Signaling. Front. Neuroanat. 2011, 5, 55. [Google Scholar] [CrossRef] [PubMed]
- Boikess, S.R.; O’Dell, S.J.; Marshall, J.F. A sensitizing d-amphetamine dose regimen induces long-lasting spinophilin and VGLUT1 protein upregulation in the rat diencephalon. Neurosci. Lett. 2010, 469, 49–54. [Google Scholar] [CrossRef]
- Boikess, S.R.; Marshall, J.F. A sensitizing d-amphetamine regimen induces long-lasting spinophilin protein upregulation in the rat striatum and limbic forebrain. Eur. J. Neurosci. 2008, 28, 2099–2107. [Google Scholar] [CrossRef]
- Xu, J.; Chen, Y.; Lu, R.; Cottingham, C.; Jiao, K.; Wang, Q. Protein Kinase A Phosphorylation of Spinophilin Modulates Its Interaction with the α2A-Adrenergic Receptor (AR) and Alters Temporal Properties of α2AAR Internalization. J. Biol. Chem. 2008, 283, 14516–14523. [Google Scholar] [CrossRef]
- Brown, A.M.; Baucum, A.J.; Bass, M.A.; Colbran, R.J. Association of protein phosphatase 1 gamma 1 with spinophilin suppresses phosphatase activity in a Parkinson disease model. J. Biol. Chem. 2008, 283, 14286–14294. [Google Scholar] [CrossRef]
- Hiday, A.C.; Edler, M.C.; Salek, A.B.; Morris, C.W.; Thang, M.; Rentz, T.J.; Rose, K.L.; Jones, L.M.; Baucum, A.J. Mechanisms and Consequences of Dopamine Depletion-Induced Attenuation of the Spinophilin/Neurofilament Medium Interaction. Neural Plast. 2017. [Google Scholar] [CrossRef] [PubMed]
- Morris, C.W.; Watkins, D.S.; Salek, A.B.; Edler, M.C.; Baucum, A.J., II. The association of spinophilin with disks large-associated protein 3 (SAPAP3) is regulated by metabotropic glutamate receptor (mGluR) 5. Mol. Cell. Neurosci. 2018, 90, 60–69. [Google Scholar] [CrossRef] [PubMed]
- Gerfen, C.R.; Paletzki, R.; Heintz, N. GENSAT BAC cre-recombinase driver lines to study the functional organization of cerebral cortical and basal ganglia circuits. Neuron 2013, 80, 1368–1383. [Google Scholar] [CrossRef] [PubMed]
- Gong, S.; Doughty, M.; Harbaugh, C.R.; Cummins, A.; Hatten, M.E.; Heintz, N.; Gerfen, C.R. Targeting Cre recombinase to specific neuron populations with bacterial artificial chromosome constructs. J. Neurosci. 2007, 27, 9817–9823. [Google Scholar] [CrossRef] [PubMed]
- Edler, M.C.; Salek, A.B.; Watkins, D.S.; Kaur, H.; Morris, C.W.; Yamamoto, B.K.; Baucum, A.J. Mechanisms Regulating the Association of Protein Phosphatase 1 with Spinophilin and Neurabin. ACS Chem. Neurosci. 2018. [Google Scholar] [CrossRef] [PubMed]
- Smith-Kinnaman, W.R.; Berna, M.J.; Hunter, G.O.; True, J.D.; Hsu, P.; Cabello, G.I.; Fox, M.J.; Varani, G.; Mosley, A.L. The interactome of the atypical phosphatase Rtr1 in Saccharomyces cerevisiae. Mol. Biosyst. 2014, 10, 1730–1741. [Google Scholar] [CrossRef] [PubMed]
- Mosley, A.L.; Sardiu, M.E.; Pattenden, S.G.; Workman, J.L.; Florens, L.; Washburn, M.P. Highly reproducible label free quantitative proteomic analysis of RNA polymerase complexes. Mol. Cell. Proteom. 2011, 10, M110000687. [Google Scholar] [CrossRef]
- Huang da, W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef]
- Huang da, W.; Sherman, B.T.; Lempicki, R.A. Bioinformatics enrichment tools: Paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009, 37, 1–13. [Google Scholar] [CrossRef]
- Chu, J.; Haynes, R.D.; Corbel, S.Y.; Li, P.; Gonzalez-Gonzalez, E.; Burg, J.S.; Ataie, N.J.; Lam, A.J.; Cranfill, P.J.; Baird, M.A.; et al. Non-invasive intravital imaging of cellular differentiation with a bright red-excitable fluorescent protein. Nat. Methods 2014, 11, 572–578. [Google Scholar] [CrossRef]
- Baucum, A.J.; Brown, A.M.; Colbran, R.J. Differential association of postsynaptic signaling protein complexes in striatum and hippocampus. J. Neurochem. 2013, 124, 490–501. [Google Scholar] [CrossRef]
- Baucum, A.J.; Jalan-Sakrikar, N.; Jiao, Y.; Gustin, R.M.; Carmody, L.C.; Tabb, D.L.; Ham, A.J.; Colbran, R.J. Identification and validation of novel spinophilin-associated proteins in rodent striatum using an enhanced ex vivo shotgun proteomics approach. Mol. Cell. Proteom. 2010, 9, 1243–1259. [Google Scholar] [CrossRef]
- Gerfen, C.R.; Surmeier, D.J. Modulation of striatal projection systems by dopamine. Annu. Rev. Neurosci. 2011, 34, 441–466. [Google Scholar] [CrossRef]
- Hu, X.T.; Wang, R.Y. Comparison of effects of D-1 and D-2 dopamine receptor agonists on neurons in the rat caudate putamen: An electrophysiological study. J. Neurosci. 1988, 8, 4340. [Google Scholar] [CrossRef]
- Muhammad, K.; Reddy-Alla, S.; Driller, J.H.; Schreiner, D.; Rey, U.; Bohme, M.A.; Hollmann, C.; Ramesh, N.; Depner, H.; Lutzkendorf, J.; et al. Presynaptic spinophilin tunes neurexin signalling to control active zone architecture and function. Nat. Commun. 2015, 6, 8362. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Franceschini, A.; Wyder, S.; Forslund, K.; Heller, D.; Huerta-Cepas, J.; Simonovic, M.; Roth, A.; Santos, A.; Tsafou, K.P.; et al. STRING v10: Protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 2015, 43, D447–D452. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Morris, J.H.; Cook, H.; Kuhn, M.; Wyder, S.; Simonovic, M.; Santos, A.; Doncheva, N.T.; Roth, A.; Bork, P.; et al. The STRING database in 2017: Quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res. 2017, 45, D362–D368. [Google Scholar] [CrossRef]
- Allen, P.B.; Zachariou, V.; Svenningsson, P.; Lepore, A.C.; Centonze, D.; Costa, C.; Rossi, S.; Bender, G.; Chen, G.; Feng, J.; et al. Distinct roles for spinophilin and neurabin in dopamine-mediated plasticity. Neuroscience 2006, 140, 897–911. [Google Scholar] [CrossRef]
- Lu, R.; Chen, Y.; Cottingham, C.; Peng, N.; Jiao, K.; Limbird, L.E.; Wyss, J.M.; Wang, Q. Enhanced hypotensive, bradycardic, and hypnotic responses to alpha2-adrenergic agonists in spinophilin-null mice are accompanied by increased G protein coupling to the alpha2A-adrenergic receptor. Mol. Pharmacol. 2010, 78, 279–286. [Google Scholar] [CrossRef]
- Ingham, C.A.; Hood, S.H.; Arbuthnott, G.W. Spine density on neostriatal neurones changes with 6-hydroxydopamine lesions and with age. Brain Res. 1989, 503, 334–338. [Google Scholar] [CrossRef]
- Ingham, C.A.; Hood, S.H.; van Maldegem, B.; Weenink, A.; Arbuthnott, G.W. Morphological changes in the rat neostriatum after unilateral 6-hydroxydopamine injections into the nigrostriatal pathway. Exp. Brain Res. 1993, 93, 17–27. [Google Scholar] [CrossRef]
- Day, M.; Wang, Z.; Ding, J.; An, X.; Ingham, C.A.; Shering, A.F.; Wokosin, D.; Ilijic, E.; Sun, Z.; Sampson, A.R.; et al. Selective elimination of glutamatergic synapses on striatopallidal neurons in Parkinson disease models. Nat. Neurosci. 2006, 9, 251–259. [Google Scholar] [CrossRef]
- Lee, K.W.; Kim, Y.; Kim, A.M.; Helmin, K.; Nairn, A.C.; Greengard, P. Cocaine-induced dendritic spine formation in D1 and D2 dopamine receptor-containing medium spiny neurons in nucleus accumbens. Proc. Natl. Acad. Sci. USA 2006, 103, 3399–3404. [Google Scholar] [CrossRef]
- Li, Y.; Kolb, B.; Robinson, T.E. The location of persistent amphetamine-induced changes in the density of dendritic spines on medium spiny neurons in the nucleus accumbens and caudate-putamen. Neuropsychopharmacology 2003, 28, 1082–1085. [Google Scholar] [CrossRef]
- Evans, J.C.; Robinson, C.M.; Shi, M.; Webb, D.J. The Guanine Nucleotide Exchange Factor (GEF) Asef2 Promotes Dendritic Spine Formation via Rac Activation and Spinophilin-dependent Targeting. J. Biol. Chem. 2015, 290, 10295–10308. [Google Scholar] [CrossRef]
- Hsieh-Wilson, L.C.; Benfenati, F.; Snyder, G.L.; Allen, P.B.; Nairn, A.C.; Greengard, P. Phosphorylation of spinophilin modulates its interaction with actin filaments. J. Biol. Chem. 2003, 278, 1186–1194. [Google Scholar] [CrossRef]
- Satoh, A.; Nakanishi, H.; Obaishi, H.; Wada, M.; Takahashi, K.; Satoh, K.; Hirao, K.; Nishioka, H.; Hata, Y.; Mizoguchi, A.; et al. Neurabin-II/spinophilin. An actin filament-binding protein with one pdz domain localized at cadherin-based cell-cell adhesion sites. J. Biol. Chem. 1998, 273, 3470–3475. [Google Scholar] [CrossRef]
- Mellacheruvu, D.; Wright, Z.; Couzens, A.L.; Lambert, J.P.; St-Denis, N.A.; Li, T.; Miteva, Y.V.; Hauri, S.; Sardiu, M.E.; Low, T.Y.; et al. The CRAPome: A contaminant repository for affinity purification-mass spectrometry data. Nat. Methods 2013, 10, 730–736. [Google Scholar] [CrossRef]
- Berman, S.; O’Neill, J.; Fears, S.; Bartzokis, G.; London, E.D. Abuse of amphetamines and structural abnormalities in the brain. Ann. N. Y. Acad. Sci. 2008, 1141, 195–220. [Google Scholar] [CrossRef]
- Kimura, T.; Allen, P.B.; Nairn, A.C.; Caplan, M.J. Arrestins and spinophilin competitively regulate Na+,K+-ATPase trafficking through association with a large cytoplasmic loop of the Na+,K+-ATPase. Mol. Biol. Cell 2007, 18, 4508–4518. [Google Scholar] [CrossRef]
- King, L.J.; Carl, J.L.; Lao, L. Cocaine and amphetamine modification of cerebral energy metabolism in vivo. Psychopharmacologia 1975, 44, 43–45. [Google Scholar] [CrossRef]
- Valvassori, S.S.; Calixto, K.V.; Budni, J.; Resende, W.R.; Varela, R.B.; de Freitas, K.V.; Goncalves, C.L.; Streck, E.L.; Quevedo, J. Sodium butyrate reverses the inhibition of Krebs cycle enzymes induced by amphetamine in the rat brain. J. Neural Transm. 2013, 120, 1737–1742. [Google Scholar] [CrossRef]
- Klongpanichapak, S.; Phansuwan-Pujito, P.; Ebadi, M.; Govitrapong, P. Melatonin protects SK-N-SH neuroblastoma cells from amphetamine-induced neurotoxicity. J. Pineal Res. 2007, 43, 65–73. [Google Scholar] [CrossRef]
- Straiko, M.M.; Coolen, L.M.; Zemlan, F.P.; Gudelsky, G.A. The effect of amphetamine analogs on cleaved microtubule-associated protein-tau formation in the rat brain. Neuroscience 2007, 144, 223–231. [Google Scholar] [CrossRef]
| Eartag | Sex | Condition | Genotype | Cre | Initial Weight | Final Weight | Birth Date | Sacrifice Date |
|---|---|---|---|---|---|---|---|---|
| 2450 | M | Saline | Het/Cre+ | D1 | 23.8 | 25.0 | 31 January 2018 | 30 March 2018 |
| 2452 | M | Treated | Het/Cre+ | D1 | 22.4 | 23 | 31 January 2018 | 30 March 2018 |
| 2453 | M | Treated | tdHet/HA-Het/Cre+ | D1 | 23.0 | 23.3 | 31 January 2018 | 30 March 2018 |
| 2454 | M | Saline | Het/Cre+ | D1 | 22.3 | 22.8 | 31 January 2018 | 30 March 2018 |
| 2390 | F | Saline | Het/Cre+ | A2A | 22.9 | 22.8 | 3 January 2018 | 30 March 2018 |
| 2393 | F | Treated | Het/Cre+ | A2A | 20.9 | 20.8 | 3 January 2018 | 30 March 2018 |
| 2443 | F | Saline | Het/Cre+ | D1 | 21.5 | 21.7 | 29 January 2018 | 30 March 2018 |
| 2444 | F | Treated | Het/Cre+ | D1 | 19.6 | 19.8 | 29 January 2018 | 30 March 2018 |
| Description | # PSMs | Normalized Abundance Ratio (Treatment)/(Control) | Normalized Abundance Ratio (log2): (Treatment)/(Control) |
|---|---|---|---|
| Decreased Interactions | |||
| E3 ubiquitin-protein ligase XIAP | 22 | 0.69 | −0.54 |
| Disks large homolog 3 | 143 | 0.67 | −0.58 |
| Granulins | 10 | 0.36 | −1.47 |
| Increased Interactions | |||
| Myelin proteolipid protein | 38 | 2.28 | 1.19 |
| Hemoglobin subunit alpha | 17 | 2.24 | 1.16 |
| ADP/ATP translocase 2 | 36 | 2.23 | 1.16 |
| Clathrin light chain A | 17 | 2.13 | 1.09 |
| Adenylyl cyclase-associated protein 2 | 8 | 2.10 | 1.07 |
| MCG10343, isoform CRA_b | 35 | 2.08 | 1.05 |
| Tubulin alpha-1B chain | 54 | 2.07 | 1.05 |
| Tubulin alpha chain (Fragment) | 54 | 2.03 | 1.02 |
| Cytochrome c oxidase subunit NDUFA4 | 13 | 2.03 | 1.02 |
| Profilin-2 | 14 | 2.02 | 1.02 |
| Reticulon (Fragment) | 20 | 2.02 | 1.01 |
| Myelin-oligodendrocyte glycoprotein | 18 | 2.01 | 1.01 |
| Serine/threonine-protein phosphatase 2A 65 kDa regulatory subunit A alpha isoform | 28 | 2.00 | 1.00 |
| Description | PSMs | Female A2A Ratios | Female D1 Ratios | Male D1 Avg Ratios | Avg D1/A2A Ratios |
|---|---|---|---|---|---|
| Endophilin-A2 | 12 | 0.50 | 1.38 | 1.28 | 2.63 |
| Cofilin-1 | 14 | 1.13 | 4.14 | 1.57 | 2.52 |
| Malate dehydrogenase, mitochondrial | 27 | 0.92 | 3.22 | 1.38 | 2.51 |
| Calreticulin | 8 | 0.87 | 3.15 | 1.11 | 2.46 |
| Alpha-synuclein | 21 | 1.06 | 3.56 | 1.61 | 2.43 |
| Malate dehydrogenase, cytoplasmic | 41 | 1.05 | 3.59 | 1.38 | 2.36 |
| Glutamate dehydrogenase 1, mitochondrial | 23 | 1.05 | 3.58 | 1.39 | 2.36 |
| Fructose-bisphosphate aldolase | 87 | 1.14 | 3.93 | 1.41 | 2.35 |
| Protein disulfide-isomerase A3 | 9 | 0.93 | 3.19 | 1.13 | 2.31 |
| Citrate synthase, mitochondrial | 32 | 1.37 | 4.58 | 1.55 | 2.24 |
| Synapsin-1 | 42 | 1.11 | 3.51 | 1.45 | 2.23 |
| Tubulin polymerization-promoting protein | 9 | 1.08 | 3.24 | 1.56 | 2.23 |
| Nucleoside diphosphate kinase | 12 | 1.20 | 3.92 | 1.39 | 2.21 |
| Fascin | 26 | 0.89 | 2.78 | 1.13 | 2.20 |
| Cytochrome b-c1 complex subunit 1, mitochondrial | 19 | 0.99 | 3.09 | 1.26 | 2.19 |
| Carbonic anhydrase 2 | 14 | 1.22 | 3.75 | 1.54 | 2.18 |
| Rab GDP dissociation inhibitor alpha | 22 | 1.12 | 3.58 | 1.26 | 2.16 |
| Protein kinase C and casein kinase substrate in neurons protein 1 | 18 | 1.10 | 3.36 | 1.30 | 2.12 |
| Endophilin-A1 | 23 | 1.11 | 3.37 | 1.33 | 2.12 |
| Cytochrome c oxidase subunit 5B, mitochondrial | 13 | 1.17 | 3.52 | 1.43 | 2.12 |
| Adenylyl cyclase-associated protein 2 | 8 | 1.43 | 4.36 | 1.63 | 2.09 |
| Cytochrome b-c1 complex subunit 2, mitochondrial | 40 | 1.04 | 3.08 | 1.22 | 2.06 |
| Pyruvate kinase PKM | 36 | 1.10 | 3.22 | 1.30 | 2.06 |
| Profilin-2 | 14 | 1.33 | 3.85 | 1.58 | 2.04 |
| Myelin proteolipid protein | 38 | 1.58 | 4.56 | 1.89 | 2.04 |
| 60S ribosomal protein L17 | 8 | 1.06 | 3.20 | 1.12 | 2.03 |
| 40S ribosomal protein S23 | 11 | 0.89 | 2.40 | 1.23 | 2.03 |
| Beta-synuclein | 16 | 1.18 | 3.31 | 1.47 | 2.03 |
| Heat shock 70 kDa protein 4 | 23 | 1.07 | 3.01 | 1.27 | 2.01 |
| L-lactate dehydrogenase B chain | 28 | 1.23 | 3.58 | 1.35 | 2.00 |
| Term | Count | % | p Value | Bonferroni |
|---|---|---|---|---|
| mmu01200:Carbon metabolism | 27 | 9.54 | 3.52 × 10−18 | 7.04 × 10−16 |
| mmu05012:Parkinson’s disease | 29 | 10.25 | 2.34 × 10−17 | 4.69 × 10−15 |
| mmu05016:Huntington’s disease | 32 | 11.31 | 9.23 × 10−17 | 2.22 × 10−14 |
| mmu00190:Oxidative phosphorylation | 26 | 9.19 | 4.42 × 10−15 | 8.88 × 10−13 |
| mmu04721:Synaptic vesicle cycle | 19 | 6.71 | 5.04 × 10−15 | 9.99 × 10−13 |
| mmu00020:Citrate cycle (TCA cycle) | 15 | 5.30 | 7.94 × 10−15 | 1.60 × 10−12 |
| mmu05010:Alzheimer’s disease | 27 | 9.54 | 1.82 × 10−13 | 3.65 × 10−11 |
| mmu01130:Biosynthesis of antibiotics | 29 | 10.25 | 3.85 × 10−13 | 7.70 × 10−11 |
| mmu04961:Endocrine and other factor-regulated calcium reabsorption | 15 | 5.30 | 1.92 × 10−11 | 3.84 × 10−9 |
| mmu00010:Glycolysis/Gluconeogenesis | 16 | 5.65 | 5.11 × 10−11 | 1.02 × 10−8 |
| Term | Count | % | PValue | Bonferroni |
|---|---|---|---|---|
| Biological Process | ||||
| GO:0006099~tricarboxylic acid cycle | 13 | 4.59 | 3.85 × 10−15 | 6.78 × 10−12 |
| GO:0006810~transport | 68 | 24.03 | 1.55 × 10−12 | 2.70 × 10−9 |
| GO:0006096~glycolytic process | 11 | 3.89 | 8.12 × 10−11 | 1.42 × 10−7 |
| GO:0006734~NADH metabolic process | 8 | 2.83 | 1.11 × 10−10 | 1.93 × 10−7 |
| GO:0046034~ATP metabolic process | 11 | 3.89 | 2.57 × 10−10 | 4.48 × 10−7 |
| GO:0015992~proton transport | 12 | 4.24 | 7.90 × 10−10 | 1.38 × 10−6 |
| GO:0015991~ATP hydrolysis coupled proton transport | 10 | 3.53 | 8.81 × 10−10 | 1.54 × 10−6 |
| GO:0050821~protein stabilization | 15 | 5.30 | 7.11 × 10−9 | 1.24 × 10−5 |
| GO:0015986~ATP synthesis coupled proton transport | 8 | 2.83 | 2.98 × 10−8 | 5.20 × 10−5 |
| GO:1904871~positive regulation of protein localization to Cajal body | 6 | 2.12 | 3.79 × 10−8 | 6.61 × 10−5 |
| Cellular Compartment | ||||
| GO:0043209~myelin sheath | 92 | 32.51 | 3.70 × 10−120 | 1.53 × 10−117 |
| GO:0070062~extracellular exosome | 159 | 56.18 | 2.16 × 10−65 | 8.91 × 10−63 |
| GO:0005739~mitochondrion | 88 | 31.10 | 3.66 × 10−27 | 1.51 × 10−24 |
| GO:0005829~cytosol | 88 | 31.10 | 4.60 × 10−26 | 1.90 × 10−23 |
| GO:0005737~cytoplasm | 173 | 61.13 | 3.24 × 10−22 | 1.34 × 10−19 |
| GO:0016020~membrane | 178 | 62.90 | 6.47 × 10−22 | 2.66 × 10−19 |
| GO:0014069~postsynaptic density | 32 | 11.31 | 5.02 × 10−21 | 2.07 × 10−18 |
| GO:0043005~neuron projection | 40 | 14.13 | 6.70 × 10−21 | 2.76 × 10−18 |
| GO:0043234~protein complex | 46 | 16.25 | 1.63 × 10−19 | 6.71 × 10−17 |
| GO:0005743~mitochondrial inner membrane | 37 | 13.07 | 2.23 × 10−19 | 9.20 × 10−17 |
| Molecular Function | ||||
| GO:0005515~protein binding | 138 | 48.8 | 1.12 × 10−22 | 6.18 × 10−20 |
| GO:0032403~protein complex binding | 36 | 12.7 | 2.54 × 10−18 | 1.40 × 10−15 |
| GO:0019901~protein kinase binding | 36 | 12.7 | 1.24 × 10−15 | 6.74 × 10−13 |
| GO:0044822~poly(A) RNA binding | 55 | 19.4 | 4.52 × 10−14 | 2.49 × 10−11 |
| GO:0000166~nucleotide binding | 74 | 26.1 | 2.72 × 10−13 | 1.50 × 10−10 |
| GO:0019904~protein domain specific binding | 27 | 9.5 | 5.67 × 10−13 | 3.13 × 10−10 |
| GO:0098641~cadherin binding involved in cell-cell adhesion | 26 | 9.2 | 1.77 × 10−12 | 9.79 × 10−10 |
| GO:0008022~protein C-terminus binding | 22 | 7.8 | 1.30 × 10−11 | 7.15 × 10−9 |
| GO:0005516~calmodulin binding | 19 | 6.7 | 5.09 × 10−10 | 2.81 × 10−7 |
| GO:0003779~actin binding | 25 | 8.8 | 6.35 × 10−10 | 3.51 × 10−7 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Watkins, D.S.; True, J.D.; Mosley, A.L.; Baucum, A.J., II. Proteomic Analysis of the Spinophilin Interactome in Rodent Striatum Following Psychostimulant Sensitization. Proteomes 2018, 6, 53. https://doi.org/10.3390/proteomes6040053
Watkins DS, True JD, Mosley AL, Baucum AJ II. Proteomic Analysis of the Spinophilin Interactome in Rodent Striatum Following Psychostimulant Sensitization. Proteomes. 2018; 6(4):53. https://doi.org/10.3390/proteomes6040053
Chicago/Turabian StyleWatkins, Darryl S., Jason D. True, Amber L. Mosley, and Anthony J. Baucum, II. 2018. "Proteomic Analysis of the Spinophilin Interactome in Rodent Striatum Following Psychostimulant Sensitization" Proteomes 6, no. 4: 53. https://doi.org/10.3390/proteomes6040053
APA StyleWatkins, D. S., True, J. D., Mosley, A. L., & Baucum, A. J., II. (2018). Proteomic Analysis of the Spinophilin Interactome in Rodent Striatum Following Psychostimulant Sensitization. Proteomes, 6(4), 53. https://doi.org/10.3390/proteomes6040053