Performance of a Multiplex Serological Helicobacter pylori Assay on a Novel Microfluidic Assay Platform
Abstract
:1. Introduction
2. Materials and Methods
2.1. Technical Platform: Evalution™
2.2. Antigens and Human Sera
2.3. Antigen Immobilization
2.4. Generation of Quality Control Samples
2.5. Multiplex Assay Procedure
2.6. Intra- and Inter-Assay Precision
2.7. Cutoff Definition and Statistical Analysis
3. Results
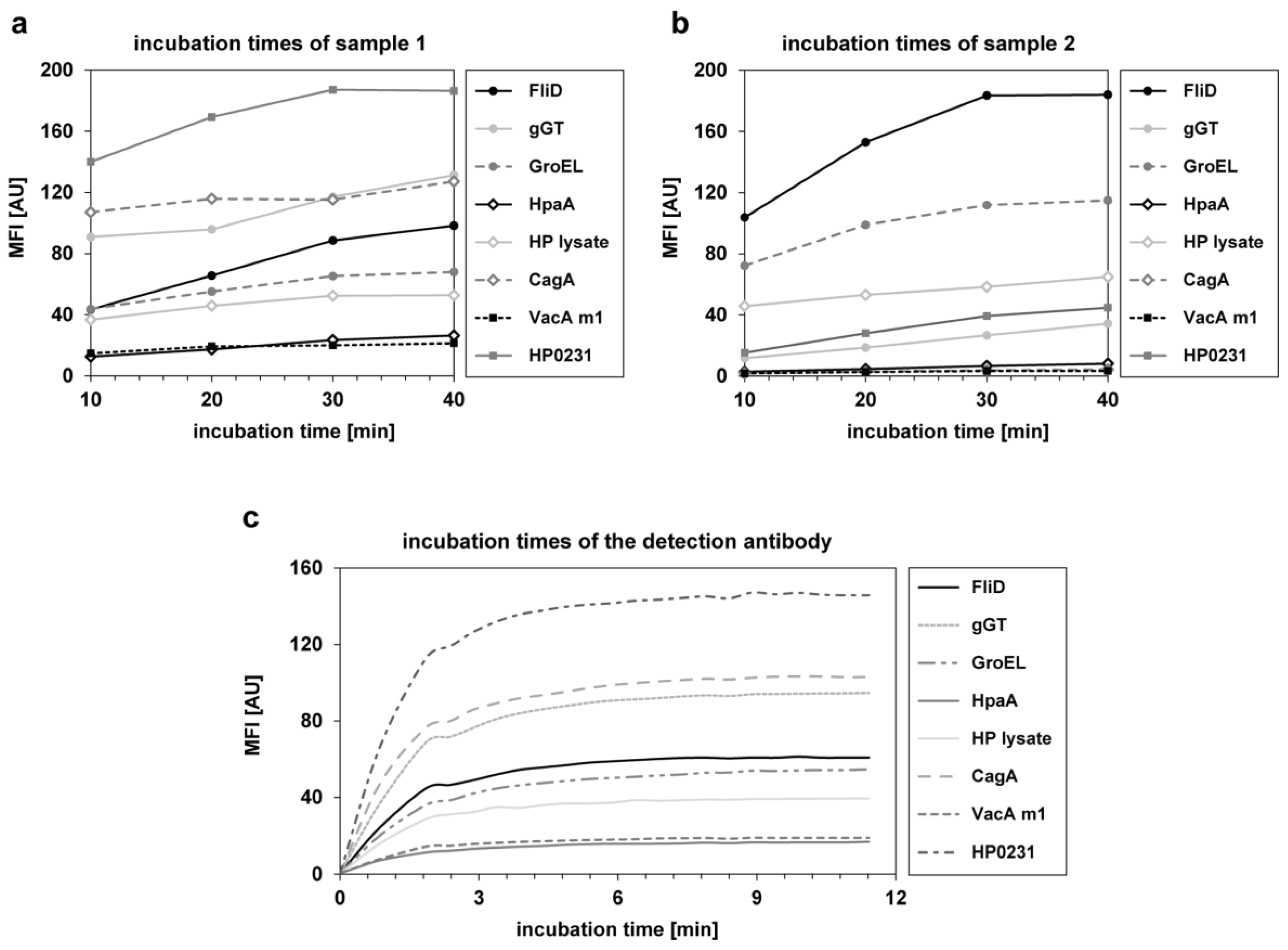
3.1. Assay Development and Technical Assay Validation
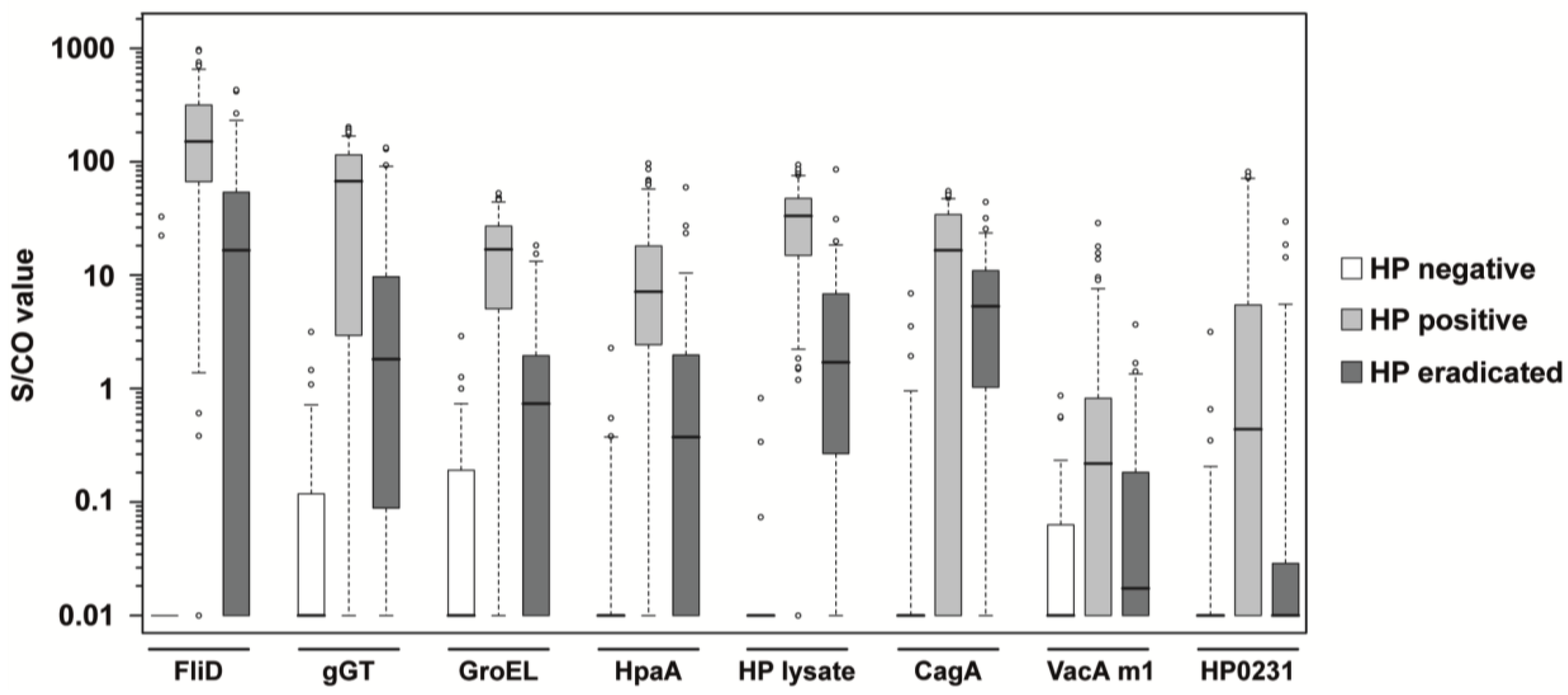
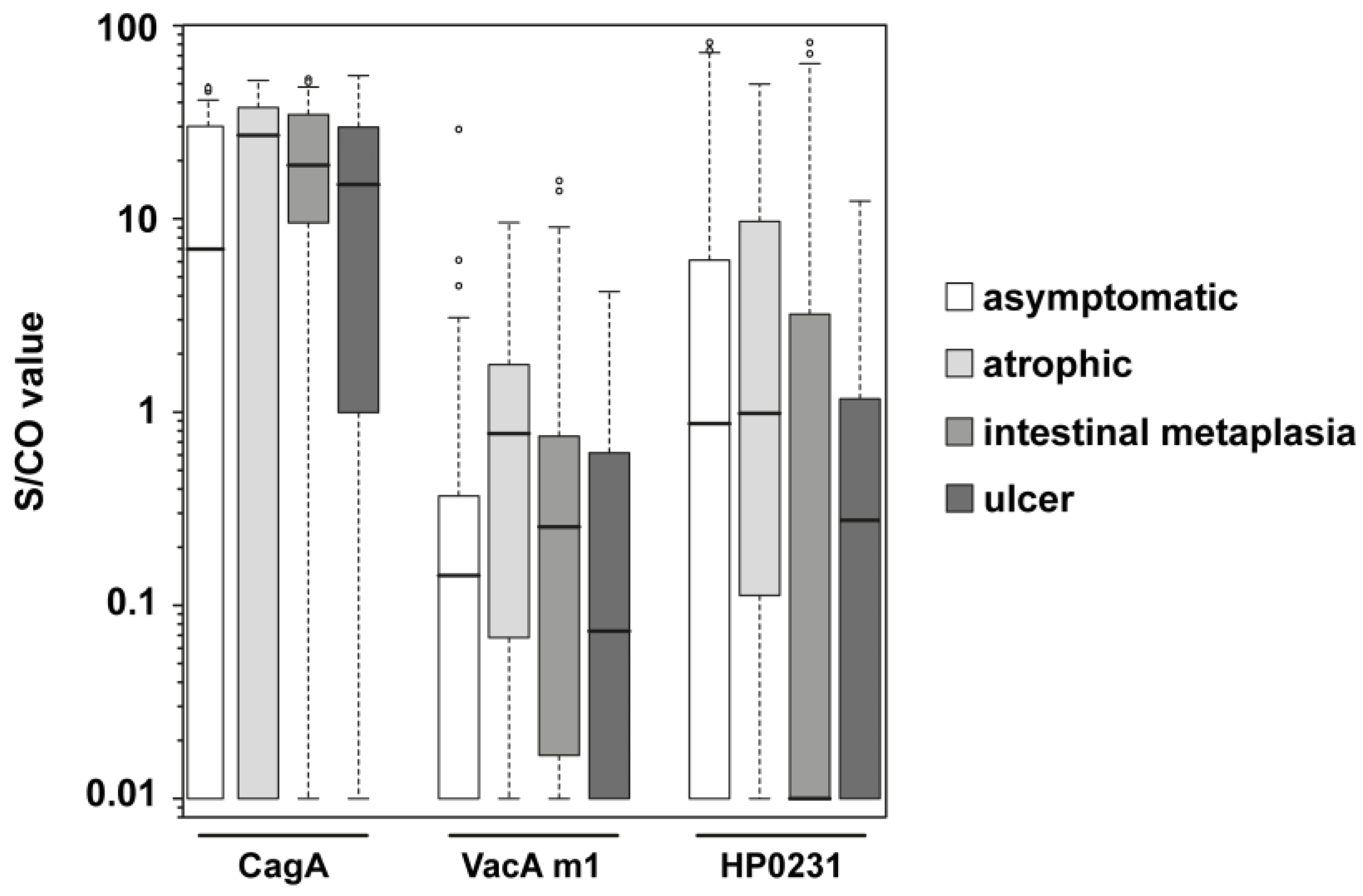
3.2. Analysis of Clinical Samples
4. Discussion
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Falconnet, D.; She, J.; Tornay, R.; Leimgruber, E.; Bernasconi, D.; Lagopoulos, L.; Renaud, P.; Demierre, N.; van den Bogaard, P. Rapid, sensitive and real-time multiplexing platform for the analysis of protein and nucleic-acid biomarkers. Anal. Chem. 2015, 87, 1582–1589. [Google Scholar] [CrossRef] [PubMed]
- Dunn, B.E.; Cohen, H.; Blaser, M.J. Helicobacter pylori. Clin. Microbiol. Rev. 1997, 10, 720–741. [Google Scholar] [PubMed]
- Blaser, M.J. Helicobacter pylori and the pathogenesis of gastroduodenal inflammation. J. Infect. Dis. 1990, 161, 626–633. [Google Scholar] [CrossRef] [PubMed]
- Iarc Working Group on the Evaluation of Carcinogenic Risks to Humans. Schistosomes, Liver Flukes and Helicobacter Pylori. IARC Monographs on the Evaluation of Carcinogenic Risks to Humans; World Health Organization, International Agency for Research on Cancer: Lyon, France, 1994; Volume 61, pp. 1–241. [Google Scholar]
- Atherton, J.C.; Cao, P.; Peek, R.M., Jr.; Tummuru, M.K.; Blaser, M.J.; Cover, T.L. Mosaicism in vacuolating cytotoxin alleles of helicobacter pylori. Association of specific vaca types with cytotoxin production and peptic ulceration. J. Biol. Chem. 1995, 270, 17771–17777. [Google Scholar] [CrossRef] [PubMed]
- Xiang, Z.; Censini, S.; Bayeli, P.F.; Telford, J.L.; Figura, N.; Rappuoli, R.; Covacci, A. Analysis of expression of caga and vaca virulence factors in 43 strains of helicobacter pylori reveals that clinical isolates can be divided into two major types and that caga is not necessary for expression of the vacuolating cytotoxin. Infect. Immun. 1995, 63, 94–98. [Google Scholar] [PubMed]
- Blaser, M.J.; Perez-Perez, G.I.; Kleanthous, H.; Cover, T.L.; Peek, R.M.; Chyou, P.H.; Stemmermann, G.N.; Nomura, A. Infection with helicobacter pylori strains possessing caga is associated with an increased risk of developing adenocarcinoma of the stomach. Cancer Res. 1995, 55, 2111–2115. [Google Scholar] [PubMed]
- Shakeri, R.; Malekzadeh, R.; Nasrollahzadeh, D.; Pawlita, M.; Murphy, G.; Islami, F.; Sotoudeh, M.; Michel, A.; Etemadi, A.; Waterboer, T.; et al. Multiplex H. pylori serology and risk of gastric cardia and noncardia adenocarcinomas. Cancer Res. 2015, 75, 4876–4883. [Google Scholar] [CrossRef] [PubMed]
- Gao, L.; Michel, A.; Weck, M.N.; Arndt, V.; Pawlita, M.; Brenner, H. Helicobacter pylori infection and gastric cancer risk: Evaluation of 15 H. pylori proteins determined by novel multiplex serology. Cancer Res. 2009, 69, 6164–6170. [Google Scholar] [CrossRef] [PubMed]
- Khalifeh Gholi, M.; Kalali, B.; Formichella, L.; Gottner, G.; Shamsipour, F.; Zarnani, A.H.; Hosseini, M.; Busch, D.H.; Shirazi, M.H.; Gerhard, M. Helicobacter pylori flid protein is a highly sensitive and specific marker for serologic diagnosis of H. pylori infection. Int. J. Med. Microbiol. 2013, 303, 618–623. [Google Scholar] [CrossRef] [PubMed]
- Schmees, C.; Prinz, C.; Treptau, T.; Rad, R.; Hengst, L.; Voland, P.; Bauer, S.; Brenner, L.; Schmid, R.M.; Gerhard, M. Inhibition of t-cell proliferation by H. pylori gamma-glutamyl transpeptidase. Gastroenterology 2007, 132, 1820–1833. [Google Scholar] [CrossRef] [PubMed]
- Cai, H.; Ye, F.; Michel, A.; Murphy, G.; Sasazuki, S.; Taylor, P.R.; Qiao, Y.L.; Park, S.K.; Yoo, K.Y.; Jee, S.H.; et al. H. pylori blood biomarker for gastric cancer risk in east asia. Int. J. Epidemiol. 2016, 45, 774–781. [Google Scholar] [CrossRef] [PubMed]
- Carlsohn, E.; Nystrom, J.; Bolin, I.; Nilsson, C.L.; Svennerholm, A.M. Hpaa is essential for helicobacter pylori colonization in mice. Infect. Immun. 2006, 74, 920–926. [Google Scholar] [CrossRef] [PubMed]
- Zhong, Y.; Anderl, F.; Kruse, T.; Schindele, F.; Jagusztyn-Krynicka, E.K.; Fischer, W.; Gerhard, M.; Mejias-Luque, R. Helicobacter pylori hp0231 influences bacterial virulence and is essential for gastric colonization. PLoS ONE 2016, 11, e0154643. [Google Scholar] [CrossRef] [PubMed]
- Fischer, W.; Winhager, L.; Rohrer, S.; Zeiller, M.; Karnholz, A.; Hoffmann, R.; Zimmer, R.; Haas, R. Strain-specific genes of Helicobacter pylori: Genome evolution driven by a novel type IV secretion system and genomic island transfer. Nucleic Acids Res. 2010, 38, 6089–6101. [Google Scholar] [CrossRef] [PubMed]
- Haas, R.; Meyer, T.F.; van Putten, J.P.M. Aflagellated mutants of Helicobacter pylori generated by genetic transformation of naturally competent strains using transposon shuttle mutagenesis. Mol. Microbiol. 1993, 8, 753–760. [Google Scholar] [CrossRef] [PubMed]
- Formichella, L.; Romberg, L.; Bolz, C.; Vieth, M.; Geppert, M.; Gottner, G.; Nolting, C.; Walter, D.; Schepp, W.; Schneider, A.; et al. A novel line immunoassay based on recombinant virulence factors enables highly specific and sensitive serologic diagnosis of helicobacter pylori infection. Clin. Vaccine Immunol. 2013, 20, 1703–1710. [Google Scholar] [CrossRef] [PubMed]
- Planatscher, H.; Rimmele, S.; Michel, G.; Potz, O.; Joos, T.; Schneiderhan-Marra, N. Systematic reference sample generation for multiplexed serological assays. Sci. Rep. 2013, 3, 3259. [Google Scholar] [CrossRef] [PubMed]
- Rudi, J.; Kolb, C.; Maiwald, M.; Zuna, I.; von Herbay, A.; Galle, P.R.; Stremmel, W. Serum antibodies against helicobacter pylori proteins vaca and caga are associated with increased risk for gastric adenocarcinoma. Dig. Dis. Sci. 1997, 42, 1652–1659. [Google Scholar] [CrossRef] [PubMed]
- Yamaoka, Y.; Kodama, T.; Kashima, K.; Graham, D.Y. Antibody against helicobacter pylori caga and vaca and the risk for gastric cancer. J. Clin. Pathol. 1999, 52, 215–218. [Google Scholar] [CrossRef] [PubMed]
- Sugano, K. Premalignant conditions of gastric cancer. J. Gastroenterol. Hepatol. 2013, 28, 906–911. [Google Scholar] [CrossRef] [PubMed]
- Michel, A.; Waterboer, T.; Kist, M.; Pawlita, M. Helicobacter pylori multiplex serology. Helicobacter 2009, 14, 525–535. [Google Scholar] [CrossRef] [PubMed]
- Atherton, J.C. H. pylori virulence factors. Br. Med. Bull. 1998, 54, 105–120. [Google Scholar] [CrossRef] [PubMed]
- Kato, S.; Furuyama, N.; Ozawa, K.; Ohnuma, K.; Iinuma, K. Long-term follow-up study of serum immunoglobulin g and immunoglobulin a antibodies after helicobacter pylori eradication. Pediatrics 1999, 104, e22. [Google Scholar] [CrossRef] [PubMed]
- Marchildon, P.; Balaban, D.H.; Sue, M.; Charles, C.; Doobay, R.; Passaretti, N.; Peacock, J.; Marshall, B.J.; Peura, D.A. Usefulness of serological igg antibody determinations for confirming eradication of helicobacter pylori infection. Am. J. Gastroenterol. 1999, 94, 2105–2108. [Google Scholar] [CrossRef] [PubMed]
- Sorberg, M.; Engstrand, L.; Strom, M.; Jonsson, K.A.; Jorbeck, H.; Granstrom, M. The diagnostic value of enzyme immunoassay and immunoblot in monitoring eradication of helicobacter pylori. Scand. J. Infect. Dis. 1997, 29, 147–151. [Google Scholar] [CrossRef] [PubMed]
- Veijola, L.; Oksanen, A.; Sipponen, P.; Rautelin, H. Evaluation of a commercial immunoblot, helicoblot 2.1, for diagnosis of helicobacter pylori infection. Clin. Vaccine Immunol. 2008, 15, 1705–1710. [Google Scholar] [CrossRef] [PubMed]
- Pan, K.F.; Formichella, L.; Zhang, L.; Zhang, Y.; Ma, J.L.; Li, Z.X.; Liu, C.; Wang, Y.M.; Goettner, G.; Ulm, K.; et al. Helicobacter pylori antibody responses and evolution of precancerous gastric lesions in a chinese population. Int. J. Cancer 2014, 134, 2118–2125. [Google Scholar] [CrossRef] [PubMed]
- Gwack, J.; Shin, A.; Kim, C.S.; Ko, K.P.; Kim, Y.; Jun, J.K.; Bae, J.; Park, S.K.; Hong, Y.C.; Kang, D.; et al. Caga-producing helicobacter pylori and increased risk of gastric cancer: A nested case-control study in korea. Br. J. Cancer 2006, 95, 639–641. [Google Scholar] [CrossRef] [PubMed]
- Gao, L.; Weck, M.N.; Michel, A.; Pawlita, M.; Brenner, H. Association between chronic atrophic gastritis and serum antibodies to 15 helicobacter pylori proteins measured by multiplex serology. Cancer Res. 2009, 69, 2973–2980. [Google Scholar] [CrossRef] [PubMed]
- Nomura, A.M.; Perez-Perez, G.I.; Lee, J.; Stemmermann, G.; Blaser, M.J. Relation between helicobacter pylori caga status and risk of peptic ulcer disease. Am. J. Epidemiol. 2002, 155, 1054–1059. [Google Scholar] [CrossRef] [PubMed]
- Saber, T.; Ghonaim, M.M.; Yousef, A.R.; Khalifa, A.; Al Qurashi, H.; Shaqhan, M.; Samaha, M. Association of helicobacter pylori caga gene with gastric cancer and peptic ulcer in saudi patients. J. Microbiol. Biotechnol. 2015, 25, 1146–1153. [Google Scholar] [CrossRef] [PubMed]
| Gender | Age | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| HP* Status | n (Total) | Female | Male | Female + Male | Female | Male | |||
| n | % | n | % | ||||||
| negative | 63 | 33 | 52.4 | 30 | 47.6 | 53.6 ± 13.3 | 53.8 ± 14.2 | 53.3 ± 12.2 | |
| positive (total) | 139 | 73 | 52.5 | 66 | 47.5 | 55.1 ± 14.9 | 56.8 ± 15.4 | 53.2 ± 14.0 | |
| asymptomatic | 66 | 34 | 51.5 | 32 | 48.5 | 55.2 ± 15.1 | 58.9 ± 14.6 | 53.0 ± 14.0 | |
| atrophy | 18 | 11 | 61.1 | 7 | 38.9 | 55.1 ± 13.6 | 54.1 ± 13.0 | 56.7 ± 14.4 | |
| intestinal metaplasia | 41 | 20 | 48.8 | 21 | 51.2 | 54.7 ± 13.7 | 57.6 ± 12.1 | 51.9 ± 14.5 | |
| ulcer | 14 | 8 | 57.1 | 6 | 42.9 | 55.6 ± 18.3 | 56.4 ± 22.4 | 54.5 ± 10.7 | |
| eradicated | 63 | 32 | 50.8 | 31 | 49.2 | 56.3 ± 13.5 | 57.4 ± 14.5 | 55.2 ± 12.4 | |
| p Values | |||||||
|---|---|---|---|---|---|---|---|
| Antigen | Cutoff (MFI) | Negative vs. Positive | Negative vs. Eradicated | Positive vs. Eradicated | Specificity (%) | Sensitivity (%) | |
| infection status | FliD | 0.20 | 5.72 × 10−29 | 4.80 × 10−14 | 5.84 × 10−14 | 96.8 | 95.7 |
| gGT | 0.95 | 4.73 × 10−23 | 1.69 × 10−10 | 3.39 × 10−10 | 95.2 | 82.0 | |
| GroEL | 2.76 | 2.02 × 10−24 | 3.58 × 10−7 | 5.99 × 10−17 | 96.8 | 87.1 | |
| HpaA | 0.66 | 3.16 × 10−25 | 1.32 × 10−10 | 2.32 × 10−12 | 97.4 | 84.2 | |
| HP lysate | 1.08 | 2.42 × 10−30 | 5.02 × 10−18 | 1.04 × 10−20 | 100.0 | 99.3 | |
| stratification | CagA | 3.80 | 1.69 × 10−16 | 9.84 × 10−15 | 2.30 × 10−4 | 95.2 | 70.5 |
| VacA m1 | 5.66 | 1.15 × 10−9 | 4.13 × 10−2 | 2.11 × 10−4 | 100.0 | 21.6 | |
| HP0231 | 2.86 | 2.21 × 10−11 | 1.39 × 10−2 | 2.69 × 10−6 | 98.4 | 84.2 | |
| Risk Factor (n Patients) | |||||||
|---|---|---|---|---|---|---|---|
| Diseased | not Diseased | ||||||
| Groups | Antigen | yes | no | yes | no | OR | 95% CI |
| asymptomatic (n = 17) vs. atrophic (n = 17) | CagA | 14 | 3 | 8 | 9 | 5.3 | 1.1–25.2 |
| VacA m1 | 8 | 9 | 2 | 15 | 6.7 | 1.2–38.6 | |
| HP0231 | 10 | 7 | 6 | 11 | 2.6 | 0.7–10.5 | |
| asymptomatic (n = 40) vs. intestinal metaplasia (n = 40) | CagA | 36 | 4 | 24 | 16 | 6.0 | 1.8–20.2 |
| VacA m1 | 9 | 31 | 6 | 34 | 1.7 | 0.5–5.2 | |
| HP0231 | 14 | 26 | 19 | 21 | 0.6 | 0.2–1.5 | |
| asymptomatic (n = 51) vs. premalignant changes (n = 57) | CagA | 50 | 7 | 28 | 23 | 5.9 | 2.2–15.4 |
| VacA m1 | 17 | 40 | 8 | 43 | 2.3 | 0.9–5.9 | |
| HP0231 | 24 | 33 | 24 | 27 | 0.8 | 0.4–1.8 | |
| asymptomatic (n = 14) vs. ulcer (n = 14) | CagA | 10 | 4 | 7 | 7 | 2.5 | 0.5–11.9 |
| VacA m1 | 3 | 11 | 12 | 2 | 1.6 | 0.2–11.7 | |
| HP0231 | 5 | 9 | 7 | 7 | 0.6 | 0.1–2.5 | |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Filomena, A.; Guenther, A.; Planatscher, H.; Topin, F.; She, J.; Formichella, L.; Terradot, L.; Gerhard, M.; Joos, T.O.; Meyer, H.; et al. Performance of a Multiplex Serological Helicobacter pylori Assay on a Novel Microfluidic Assay Platform. Proteomes 2017, 5, 24. https://doi.org/10.3390/proteomes5040024
Filomena A, Guenther A, Planatscher H, Topin F, She J, Formichella L, Terradot L, Gerhard M, Joos TO, Meyer H, et al. Performance of a Multiplex Serological Helicobacter pylori Assay on a Novel Microfluidic Assay Platform. Proteomes. 2017; 5(4):24. https://doi.org/10.3390/proteomes5040024
Chicago/Turabian StyleFilomena, Angela, Anna Guenther, Hannes Planatscher, Francois Topin, Joseph She, Luca Formichella, Laurent Terradot, Markus Gerhard, Thomas O. Joos, Hannelore Meyer, and et al. 2017. "Performance of a Multiplex Serological Helicobacter pylori Assay on a Novel Microfluidic Assay Platform" Proteomes 5, no. 4: 24. https://doi.org/10.3390/proteomes5040024
APA StyleFilomena, A., Guenther, A., Planatscher, H., Topin, F., She, J., Formichella, L., Terradot, L., Gerhard, M., Joos, T. O., Meyer, H., & Schneiderhan-Marra, N. (2017). Performance of a Multiplex Serological Helicobacter pylori Assay on a Novel Microfluidic Assay Platform. Proteomes, 5(4), 24. https://doi.org/10.3390/proteomes5040024





