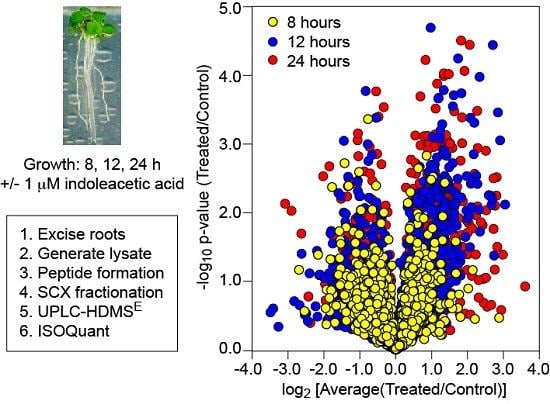
Exogenous Auxin Elicits Changes in the Arabidopsis thaliana Root Proteome in a Time-Dependent Manner
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Material and Hormone Treatment
2.2. Protein Extraction and Processing
2.3. LC-MS Analysis
2.4. Data Analysis
3. Results and Discussion
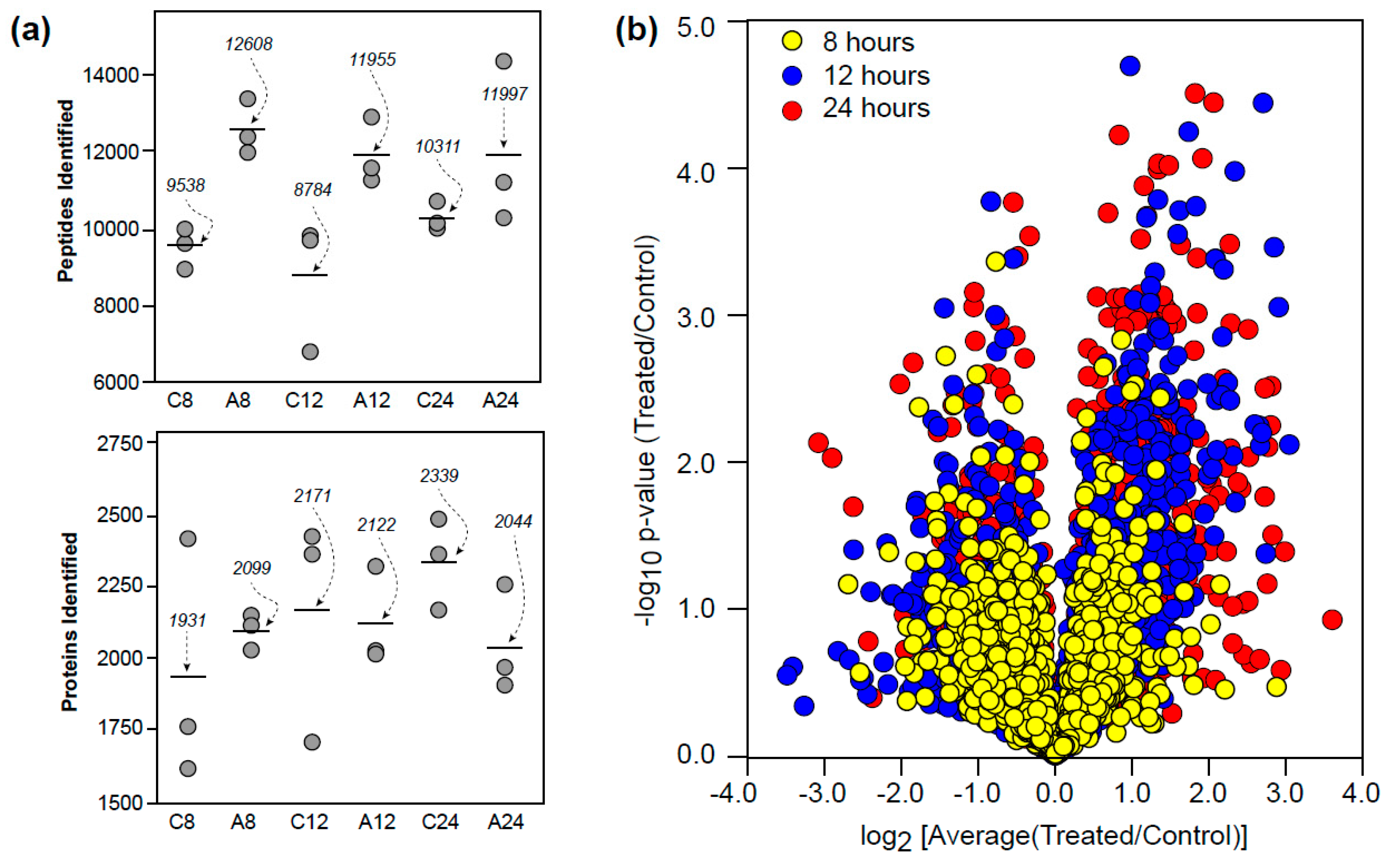
3.1. Proteomic Overview
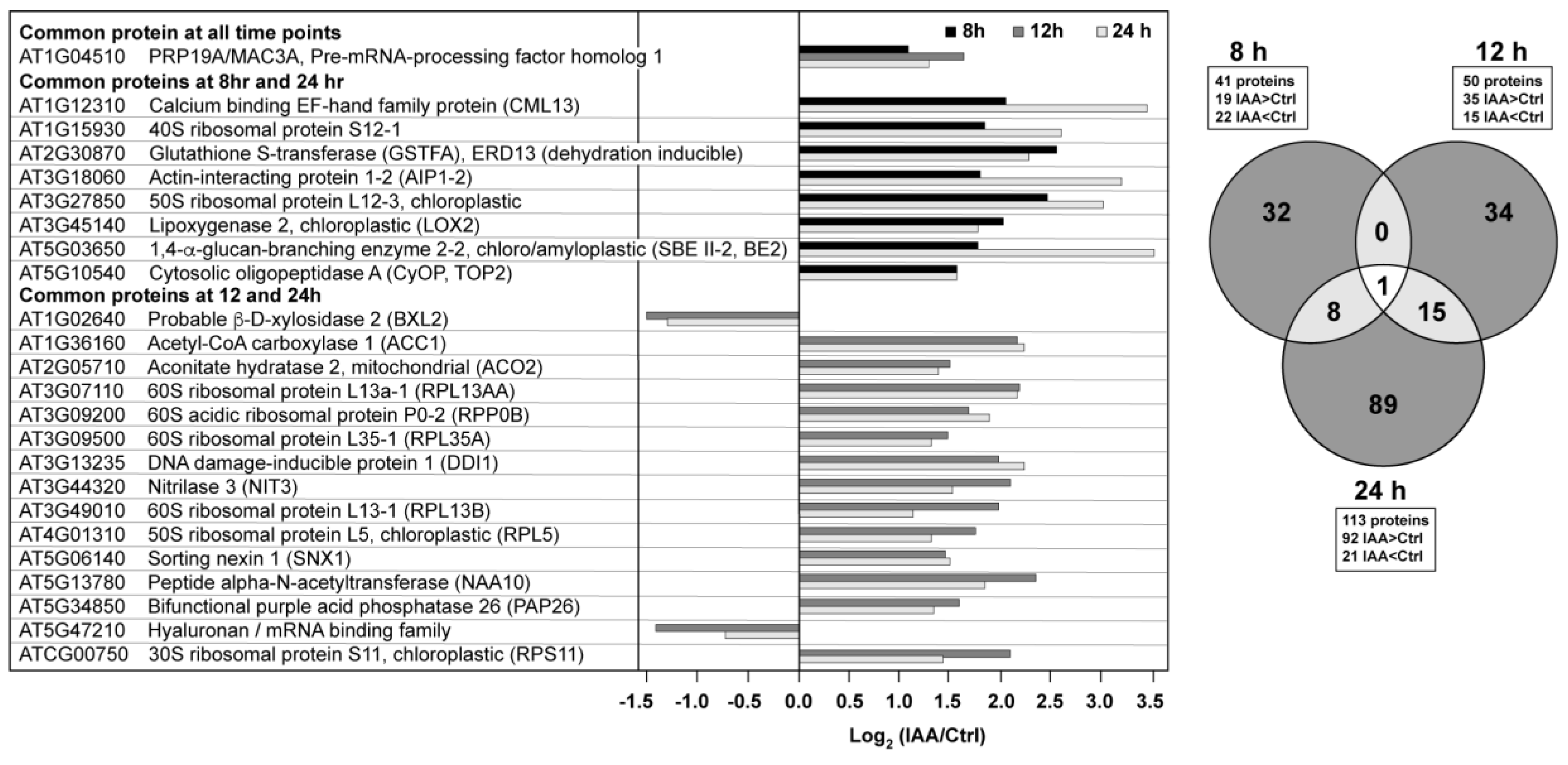
3.2. Temporal Differences
3.3. Small Secretory Peptides (SSPs) and Putative Cell Wall Synthesis-Related Proteins
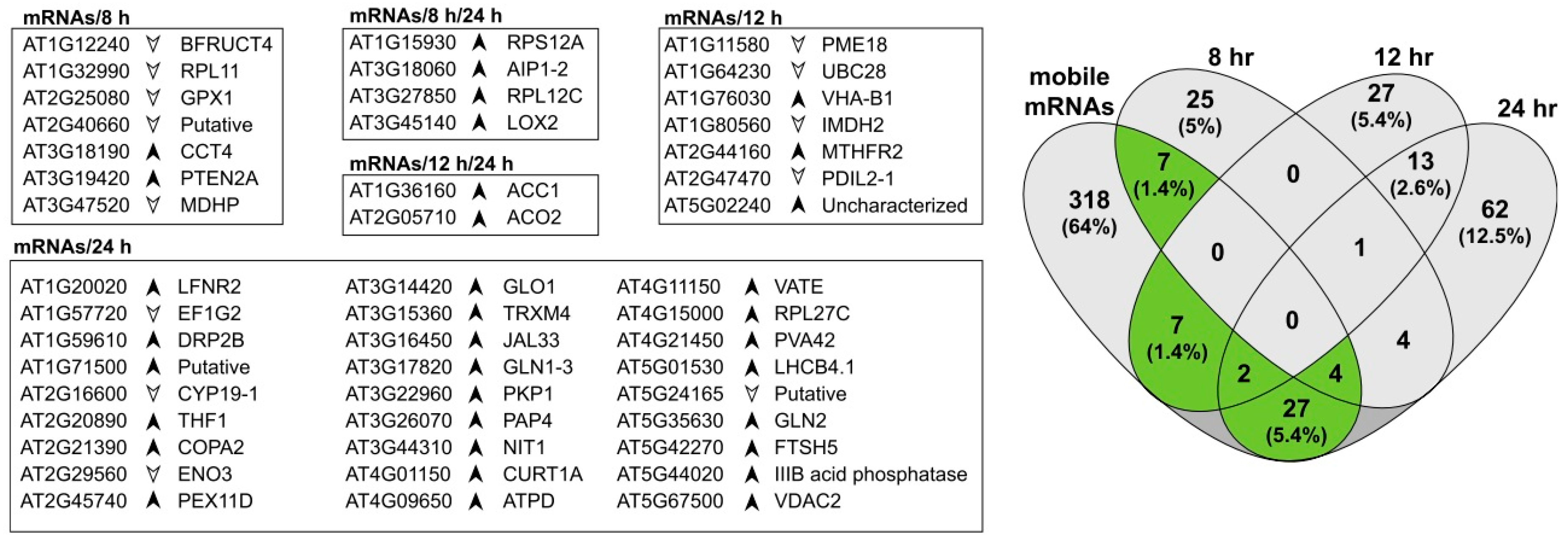
3.4. Proteins Associated with Mobile mRNAs
3.5. Future Directions
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Buer, C.S.; Muday, G.K. The transparent testa4 mutation prevents flavonoid synthesis and alters auxin transport and the response of Arabidopsis roots to gravity and light. Plant Cell 2005, 16, 1191–1205. [Google Scholar] [CrossRef] [PubMed]
- Overvoorde, P.; Fukaki, H.; Beeckman, T. Auxin control of root development. CSH Perspect. Biol. 2010, 2, a001537. [Google Scholar] [CrossRef] [PubMed]
- Casimiro, I.; Marchant, A.; Bhalerao, R.P.; Beeckman, T.; Dhooge, S.; Swarup, R.; Graham, N.; Inze, D.; Sandberg, G.; Casero, P.J.; et al. Auxin transport promotes Arabidopsis lateral root initiation. Plant Cell 2001, 13, 843–852. [Google Scholar] [CrossRef] [PubMed]
- Vilches-Barro, A.; Maizel, A. Talking through walls: Mechanisms of lateral root emergence in Arabidopsis thaliana. Curr. Opin. Plant Biol. 2015, 23, 31–38. [Google Scholar] [CrossRef] [PubMed]
- Mattei, B.; Sabatini, S.; Schinina, M.E. Proteomics in deciphering the auxin commitment in the Arabidopsis thaliana root growth. J. Proteome Res. 2013, 12, 4685–4701. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Zhou, H.; Berke, L.; Heck, A.J.; Mohammed, S.; Scheres, B.; Menke, F.L. Quantitative phosphoproteomics after auxin-stimulated lateral root induction identifies a SNX1 phosphorylation site required for growth. Molec. Cell Proteom. 2013, 1158–1169. [Google Scholar] [CrossRef] [PubMed]
- Slade, W.O.; Ray, W.K.; Williams, P.M.; Winkel, B.S.; Helm, R.F. Effects of exogenous auxin and ethylene on the Arabidopsis root proteome. Phytochemistry 2012, 84, 18–23. [Google Scholar] [CrossRef] [PubMed]
- Michalski, A.; Cox, J.; Mann, M. More than 100,000 detectable peptide species elute in single shotgun proteomics runs but the majority is inaccessible to data-dependent LC-MS/MS. J. Proteome Res. 2011, 10, 1785–1793. [Google Scholar] [CrossRef] [PubMed]
- Shliaha, P.V.; Bond, N.J.; Gatto, L.; Lilley, K.S. Effects of traveling wave ion mobility separation on data independent acquisition in proteomics studies. J. Proteome Res. 2013, 12, 2323–2339. [Google Scholar] [CrossRef] [PubMed]
- Baker, E.S.; Livesay, E.A.; Orton, D.J.; Moore, R.J.; Danielson III, W.F.; Prior, D.C.; Ibrahim, Y.M.; LaMarche, B.L.; Mayampurath, A.M.; Schepmoes, A.A.; et al. An LC-IMS-MS platform providing increased dynamic range for high-throughput proteomic studies. J. Proteome Res. 2010, 9, 997–1006. [Google Scholar] [CrossRef] [PubMed]
- Bond, N.J.; Shliaha, P.V.; Lilley, K.S.; Gatto, L. Improving qualitative and quantitative performance for mse-based label-free proteomics. J. Proteome Res. 2013, 12, 2340–2353. [Google Scholar] [CrossRef] [PubMed]
- Distler, U.; Kuharev, J.; Navarro, P.; Tenzer, S. Label-free quantification in ion mobility-enhanced data-independent acquisition proteomics. Nat. Protoc 2016, 11, 795–812. [Google Scholar] [CrossRef] [PubMed]
- Lewis, D.R.; Olex, A.L.; Lundy, S.R.; Turkett, W.H.; Fetrow, J.S.; Muday, G.K. A kinetic analysis of the auxin transcriptome reveals cell wall remodeling proteins that modulate lateral root development in Arabidopsis. Plant Cell 2013, 25, 3329–3346. [Google Scholar] [CrossRef] [PubMed]
- Lewis, D.R.; Ramirez, M.V.; Miller, N.D.; Vallabhaneni, P.; Ray, W.K.; Helm, R.F.; Winkel, B.S.J.; Muday, G.K. Auxin and ethylene induce flavonol accumulation through distinct transcriptional networks. Plant Physiol. 2011, 156, 144–164. [Google Scholar] [CrossRef] [PubMed]
- Vizcaino, J.A.; Csordas, A.; Del-Toro, N.; Dianes, J.A.; Griss, J.; Lavidas, I.; Mayer, G.; Perez-Riverol, Y.; Reisinger, F.; Ternent, T.; et al. 2016 update of the pride database and its related tools. Nucleic. Acids Res. 2016, 44, 11033. [Google Scholar] [CrossRef] [PubMed]
- Geromanos, S.J.; Hughes, C.; Ciavarini, S.; Vissers, J.P.; Langridge, J.I. Using ion purity scores for enhancing quantitative accuracy and precision in complex proteomics samples. Anal. Bioanal. Chem. 2012, 404, 1127–1139. [Google Scholar] [CrossRef] [PubMed]
- Geromanos, S.J.; Vissers, J.P.; Silva, J.C.; Dorschel, C.A.; Li, G.Z.; Gorenstein, M.V.; Bateman, R.H.; Langridge, J.I. The detection, correlation, and comparison of peptide precursor and product ions from data independent lc-ms with data dependant LC-MS/MS. Proteomics 2009, 9, 1683–1695. [Google Scholar] [CrossRef] [PubMed]
- Monaghan, J.; Xu, F.; Gao, M.; Zhao, Q.; Palma, K.; Long, C.; Chen, S.; Zhang, Y.; Li, X. Two prp19-like u-box proteins in the mos4-associated complex play redundant roles in plant innate immunity. PLoS Pathog 2009, 5, e1000526. [Google Scholar] [CrossRef] [PubMed]
- Deng, X.; Lu, T.; Wang, L.; Gu, L.; Sun, J.; Kong, X.; Liu, C.; Cao, X. Recruitment of the nineteen complex to the activated spliceosome requires atprmt5. Proc. Natl. Acad. Sci. USA 2016, 113, 5447–5452. [Google Scholar] [CrossRef] [PubMed]
- Zhou, C.; Yin, Y.; Dam, P.; Xu, Y. Identification of novel proteins involved in plant cell-wall synthesis based on protein-protein interaction data. J. Proteome Res. 2010, 9, 5025–5037. [Google Scholar] [CrossRef] [PubMed]
- Guerriero, G.; Hausman, J.F.; Ezcurra, I. WD40-repeat proteins in plant cell wall formation: Current evidence and research prospects. Front. Plant Sci. 2015, 6, 1112. [Google Scholar] [CrossRef] [PubMed]
- Linster, E.; Stephan, I.; Bienvenut, W.V.; Maple-Grodem, J.; Myklebust, L.M.; Huber, M.; Reichelt, M.; Sticht, C.; Moller, S.G.; Meinnel, T.; et al. Downregulation of N-terminal acetylation triggers ABA-mediated drought responses in Arabidopsis. Nat. Commun. 2015, 6, 7640. [Google Scholar] [CrossRef] [PubMed]
- Marino, G.; Eckhard, U.; Overall, C.M. Protein termini and their modifications revealed by positional proteomics. ACS Chem. Biol. 2015, 10, 1754–1764. [Google Scholar] [CrossRef] [PubMed]
- Roudier, F.; Gissot, L.; Beaudoin, F.; Haslam, R.; Michaelson, L.; Marion, J.; Molino, D.; Lima, A.; Bach, L.; Morin, H.; et al. Very-long-chain fatty acids are involved in polar auxin transport and developmental patterning in Arabidopsis. Plant Cell 2010, 22, 364–375. [Google Scholar] [CrossRef] [PubMed]
- Bartel, B.; Fink, G.R. Differential regulation of an auxin-producing nitrilase gene family in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 1994, 91, 6649–6653. [Google Scholar] [CrossRef] [PubMed]
- Jaillais, Y.; Fobis-Loisy, I.; Miege, C.; Rollin, C.; Gaude, T. AtSNX1 defines an endosome for auxin-carrier trafficking in Arabidopsis. Nature 2006, 443, 106–109. [Google Scholar] [CrossRef] [PubMed]
- Cui, Y.; Li, X.; Chen, Q.; He, X.; Yang, Q.; Zhang, A.; Yu, X.; Chen, H.; Liu, N.; Xie, Q.; et al. BLOS1, a putative bLOC-1 subunit, interacts with SNX1 and modulates root growth in Arabidopsis. J. Cell Sci. 2010, 123, 3727–3733. [Google Scholar] [CrossRef] [PubMed]
- Ambrose, C.; Ruan, Y.; Gardiner, J.; Tamblyn, L.M.; Catching, A.; Kirik, V.; Marc, J.; Overall, R.; Wasteneys, G.O. CLASP interacts with sorting nexin 1 to link microtubules and auxin transport via PIN2 recycling in Arabidopsis thaliana. Dev. Cell 2013, 24, 649–659. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Sun, R.; Hicks, G.R.; Raikhel, N.V. Arabidopsis ribosomal proteins control vacuole trafficking and developmental programs through the regulation of lipid metabolism. Proc. Natl. Acad. Sci. USA 2015, 112, E89–E98. [Google Scholar] [CrossRef] [PubMed]
- Moeder, W.; Del Pozo, O.; Navarre, D.A.; Martin, G.B.; Klessig, D.F. Aconitase plays a role in regulating resistance to oxidative stress and cell death in Arabidopsis and nicotiana benthamiana. Plant Mol. Biol. 2007, 63, 273–287. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.L.; Sung, S.C.; Tsai, H.L.; Yu, T.T.; Radjacommare, R.; Usharani, R.; Fatimababy, A.S.; Lin, H.Y.; Wang, Y.Y.; Fu, H. The defective proteasome but not substrate recognition function is responsible for the null phenotypes of the Arabidopsis proteasome subunit RPN10. Plant Cell 2011, 23, 2754–2773. [Google Scholar] [CrossRef] [PubMed]
- Robinson, W.D.; Carson, I.; Ying, S.; Ellis, K.; Plaxton, W.C. Eliminating the purple acid phosphatase AtPAP26 in Arabidopsis thaliana delays leaf senescence and impairs phosphorus remobilization. New Phytol. 2012, 196, 1024–1029. [Google Scholar] [CrossRef] [PubMed]
- Robinson, W.D.; Park, J.; Tran, H.T.; Del Vecchio, H.A.; Ying, S.; Zins, J.L.; Patel, K.; McKnight, T.D.; Plaxton, W.C. The secreted purple acid phosphatase isozymes AtPAP12 and AtPAP26 play a pivotal role in extracellular phosphate-scavenging by Arabidopsis thaliana. J. Exp. Bot. 2012, 63, 6531–6542. [Google Scholar] [CrossRef] [PubMed]
- Veljanovski, V.; Vanderbeld, B.; Knowles, V.L.; Snedden, W.A.; Plaxton, W.C. Biochemical and molecular characterization of AtPAP26, a vacuolar purple acid phosphatase up-regulated in phosphate-deprived Arabidopsis suspension cells and seedlings. Plant Physiol. 2006, 142, 1282–1293. [Google Scholar] [CrossRef] [PubMed]
- Marmagne, A.; Ferro, M.; Meinnel, T.; Bruley, C.; Kuhn, L.; Garin, J.; Barbier-Brygoo, H.; Ephritikhine, G. A high content in lipid-modified peripheral proteins and integral receptor kinases features in the Arabidopsis plasma membrane proteome. Mol. Cell Proteom. 2007, 6, 1980–1996. [Google Scholar] [CrossRef] [PubMed]
- Minic, Z.; Jamet, E.; Negroni, L.; Arsene der Garabedian, P.; Zivy, M.; Jouanin, L. A sub-proteome of Arabidopsis thaliana mature stems trapped on concanavalin a is enriched in cell wall glycoside hydrolases. J. Exp. Bot. 2007, 58, 2503–2512. [Google Scholar] [CrossRef] [PubMed]
- Arsovski, A.A.; Popma, T.M.; Haughn, G.W.; Carpita, N.C.; McCann, M.C.; Western, T.L. AtBXL1 encodes a bifunctional β-d-xylosidase/α-l-arabinofuranosidase required for pectic arabinan modification in Arabidopsis mucilage secretory cells. Plant Physiol. 2009, 150, 1219–1234. [Google Scholar] [CrossRef] [PubMed]
- Kmiec, B.; Teixeira, P.F.; Berntsson, R.P.; Murcha, M.W.; Branca, R.M.; Radomiljac, J.D.; Regberg, J.; Svensson, L.M.; Bakali, A.; Langel, U.; et al. Organellar oligopeptidase (oop) provides a complementary pathway for targeting peptide degradation in mitochondria and chloroplasts. Proc. Natl. Acad. Sci. USA 2013, 110, E3761–3769. [Google Scholar] [CrossRef] [PubMed]
- Bianchi, M.W.; Roux, C.; Vartanian, N. Drought regulation of GST8, encoding the Arabidopsis homologue of ParC/Nt107 glutathione transferase/peroxidase. Physiol Plant 2002, 116, 96–105. [Google Scholar] [CrossRef] [PubMed]
- McCormack, E.; Tsai, Y.C.; Braam, J. Handling calcium signaling: Arabidopsis cams and cmls. Trends. Plant Sci. 2005, 10, 383–389. [Google Scholar] [CrossRef] [PubMed]
- Bannenberg, G.; Martinez, M.; Hamberg, M.; Castresana, C. Diversity of the enzymatic activity in the lipoxygenase gene family of Arabidopsis thaliana. Lipids 2009, 44, 85–95. [Google Scholar] [CrossRef] [PubMed]
- Allwood, E.G.; Anthony, R.G.; Smertenko, A.P.; Reichelt, S.; Drobak, B.K.; Doonan, J.H.; Weeds, A.G.; Hussey, P.J. Regulation of the pollen-specific actin-depolymerizing factor LIADF1. Plant Cell 2002, 14, 2915–2927. [Google Scholar] [CrossRef] [PubMed]
- Dumez, S.; Wattebled, F.; Dauvillee, D.; Delvalle, D.; Planchot, V.; Ball, S.G.; D'Hulst, C. Mutants of Arabidopsis lacking starch branching enzyme ii substitute plastidial starch synthesis by cytoplasmic maltose accumulation. Plant Cell 2006, 18, 2694–2709. [Google Scholar] [CrossRef] [PubMed]
- Ghorbani, S.; Lin, Y.C.; Parizot, B.; Fernandez, A.; Njo, M.F.; Van de Peer, Y.; Beeckman, T.; Hilson, P. Expanding the repertoire of secretory peptides controlling root development with comparative genome analysis and functional assays. J. Exp. Bot. 2015, 66, 5257–5269. [Google Scholar] [CrossRef] [PubMed]
- Salminen, T.A.; Blomqvist, K.; Edqvist, J. Lipid transfer proteins: Classification, nomenclature, structure, and function. Planta 2016, 244, 971–997. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Sun, Y.; Chang, J.; Zheng, F.; Pei, H.; Yi, Y.; Chang, C.; Dong, C.H. Regulatory function of Arabidopsis lipid transfer protein 1 (LTP1) in ethylene response and signaling. Plant Mol. Biol. 2016, 91, 471–484. [Google Scholar] [CrossRef] [PubMed]
- Potocka, I.; Baldwin, T.C.; Kurczynska, E.U. Distribution of lipid transfer protein 1 (LTP1) epitopes associated with morphogenic events during somatic embryogenesis of Arabidopsis thaliana. Plant Cell Rep. 2012, 31, 2031–2045. [Google Scholar] [CrossRef] [PubMed]
- Thieme, C.J.; Rojas-Triana, M.; Stecyk, E.; Schudoma, C.; Zhang, W.; Yang, L.; Minambres, M.; Walther, D.; Schulze, W.X.; Paz-Ares, J.; et al. Endogenous Arabidopsis messenger rnas transported to distant tissues. Nat. Plants 2015, 1, 15025. [Google Scholar] [CrossRef] [PubMed]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using david bioinformatics resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef] [PubMed]
- De-la-Pena, C.; Badri, D.V.; Vivanco, J.M. Novel role for pectin methylesterase in Arabidopsis: A new function showing ribosome-inactivating protein (RIP) activity. Biochim. Biophys. Acta 2008, 1780, 773–783. [Google Scholar] [CrossRef] [PubMed]
- Ishikawa, H.; Evans, M.L. Novel software for analysis of root gravitropism: Comparative response patterns of Arabidopsis wild-type and axr1 seedlings. Plant Cell Environ. 1997, 20, 919–928. [Google Scholar] [CrossRef] [PubMed]
- Laskowski, M.; Biller, S.; Stanley, K.; Kajstura, T.; Prusty, R. Expression profiling of auxin-treated Arabidopsis roots: Toward a molecular analysis of lateral root emergence. Plant Cell Physiol. 2006, 47, 788–792. [Google Scholar] [CrossRef] [PubMed]
- Nemhauser, J.L.; Mockler, T.C.; Chory, J. Interdependency of brassinosteroid and auxin signaling in Arabidopsis. PloS Biol. 2004, 2, 1460–1471. [Google Scholar] [CrossRef] [PubMed]
- Stepanova, A.N.; Yun, J.; Likhacheva, A.V.; Alonso, J.M. Multilevel interactions between ethylene and auxin in Arabidopsis roots. Plant Cell 2007, 19, 2169–2185. [Google Scholar] [CrossRef] [PubMed]
- Chapman, E.J.; Estelle, M. Mechanism of auxin-regulated gene expression in plants. Ann. Rev. Genet. 2009, 43, 265–285. [Google Scholar] [CrossRef] [PubMed]
- Silva-Navas, J.; Moreno-Risueno, M.A.; Manzano, C.; Tellez-Robledo, B.; Navarro-Neila, S.; Carrasco, V.; Pollmann, S.; Gallego, F.J.; Del Pozo, J.C. Flavonols mediate root phototropism and growth through regulation of proliferation-to-differentiation transition. Plant Cell 2016, 28, 1372–1387. [Google Scholar] [CrossRef] [PubMed]
- Cosgrove, D.J. Growth of the plant cell wall. Nat. Rev. Molec. Cell Biol. 2005, 6, 850–861. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Rodriguez, C.; Rubio-Somoza, I.; Sibout, R.; Persson, S. Phytohormones and the cell wall in Arabidopsis during seedling growth. Trends Plant Sci. 2010, 15, 291–301. [Google Scholar] [CrossRef] [PubMed]
- Alvarez, S.; Berla, B.M.; Sheffield, J.; Cahoon, R.E.; Jez, J.M.; Hicks, L.M. Comprehensive analysis of the Brassica juncea root proteome in response to cadmium exposure by complementary proteomic approaches. Proteomics 2009, 9, 2419–2431. [Google Scholar] [CrossRef] [PubMed]
- Alvarez, S.; Choudhury, S.R.; Hicks, L.M.; Pandey, S. Quantitative proteomics-based analysis supports a significant role of GTG proteins in regulation of ABA response in Arabidopsis roots. J. Proteome Res. 2013, 12, 1487–1501. [Google Scholar] [CrossRef] [PubMed]
- Walley, J.W.; Sartor, R.C.; Shen, Z.; Schmitz, R.J.; Wu, K.J.; Urich, M.A.; Nery, J.R.; Smith, L.G.; Schnable, J.C.; Ecker, J.R.; et al. Integration of omic networks in a developmental atlas of maize. Science 2016, 353, 814–818. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Yamada, M.; Han, X.; Ohler, U.; Benfey, P.N. High-resolution expression map of the Arabidopsis root reveals alternative splicing and lincrna regulation. Dev. Cell 2016, 39, 508–522. [Google Scholar] [CrossRef] [PubMed]
| TAIR ID | LogFC 1 | UniProt | Protein Names | 8 h | 24 h |
|---|---|---|---|---|---|
| AT1G06430 | 0.706 | Q8W585 | ATP-dependent Zn metalloprotease FTSH 8 | 0.442 | n.s. 2 |
| AT1G12240 | 0.767 | Q39041 | Acid β-fructofuranosidase 4, vacuolar | 0.584 | n.s. |
| AT3G45140 | 0.741 | P38418 | Lipoxygenase 2, chloroplastic | 2.011 | 1.762 |
| AT4G21650 | 0.694 | Q8GUK4 | Subtilisin-like protease SBT3.13, secreted | 0.224 | n.s. |
| AT4G28400 | 0.609 | Q93YW5 | Protein phosphatase 2C 58 (AtPP2C58) | n.s. | 2.361 |
| AT5G22880 | 0.618 | Q9FFC0 | Histone H2B.10 (HTB2) | n.s. | 3.504 |
| AT5G24165 | 0.537 | Q8LDQ8 | At5g24165 (uncharacterized protein) | n.s. | 0.244 |
| AT5G48030 | 0.533 | Q8GWW8 | Chaperone protein dnaJ GFA2, mitochondrial | 1.703 | n.s. |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Slade, W.O.; Ray, W.K.; Hildreth, S.B.; Winkel, B.S.J.; Helm, R.F. Exogenous Auxin Elicits Changes in the Arabidopsis thaliana Root Proteome in a Time-Dependent Manner. Proteomes 2017, 5, 16. https://doi.org/10.3390/proteomes5030016
Slade WO, Ray WK, Hildreth SB, Winkel BSJ, Helm RF. Exogenous Auxin Elicits Changes in the Arabidopsis thaliana Root Proteome in a Time-Dependent Manner. Proteomes. 2017; 5(3):16. https://doi.org/10.3390/proteomes5030016
Chicago/Turabian StyleSlade, William O., W. Keith Ray, Sherry B. Hildreth, Brenda S. J. Winkel, and Richard F. Helm. 2017. "Exogenous Auxin Elicits Changes in the Arabidopsis thaliana Root Proteome in a Time-Dependent Manner" Proteomes 5, no. 3: 16. https://doi.org/10.3390/proteomes5030016
APA StyleSlade, W. O., Ray, W. K., Hildreth, S. B., Winkel, B. S. J., & Helm, R. F. (2017). Exogenous Auxin Elicits Changes in the Arabidopsis thaliana Root Proteome in a Time-Dependent Manner. Proteomes, 5(3), 16. https://doi.org/10.3390/proteomes5030016