Introducing and Comparing Recent Clustering Methods for Massive Data Management in the Internet of Things
Abstract
1. Introduction
Motivation and Contribution
- All the main important techniques which have been devised in the literature, including the necessary details for their implementation in the network, are described.
- A thorough comparison between the main important techniques which have been devised in the literature, as well as their respective advantages and disadvantages, is provided.
- Numerical experiments illustrating the performance of the methods are presented.
- An example with real data for a gas leak detection sensor network is analyzed to show how these various clustering techniques can respond in different ways to data received at the sink level.
- We demonstrate that choosing the correct method makes a difference via performance comparisons on a massive dataset, where some methods show excellent accuracy and scalability, whereas others appear completely inappropriate for the task.
2. An Overview of Modern Clustering Methods Applicable to the IoT
2.1. Introducing Clustering for IoT
2.2. Ellipsoid-Shaped Clusters
2.2.1. K-Means
Generalities
- is the set of possible subsets of .
- is the center of the cluster.
- Assign each to its nearest center
- Recalculate each center as an average of the closest to it.
2.2.2. K-Means++
- The initial centers are chosen randomly, but with a non-uniform probability law.
- The probability of choosing a point as the initial center is proportional to the square of the distance from that point to the already selected centers.
| Algorithm 1 K-means++ initialization |
| Require:K: Number of clusters |
| Require:m elements to be clustered |
| for do |
| for do |
| end for |
| pick r randomly in |
| find l such that r belongs in |
| end for |
2.2.3. K-Medoids
- Initialization: choose k from n points for medoids.
- Associate each point with its closest medoid.
- As long as the cost of configuration decreases:
- -
- For each medoid m and each non-medoid o:
- ∗
- Exchange m and o, associate each point with its closest medoid, and recalculate the cost (sum of the distances of the points to their medoids).
- ∗
- If the total cost of the configuration has increased in the previous step, cancel the exchange.
2.2.4. Gaussian Mixture Model
- We can, for example, have two circular clusters of the same center and different radii, which K-means will not be able to capture.
- Similarly, if the clusters are better modelled using an ellipsoid-shaped subset, K-means might not converge to the correct solution.
- We select the number of clusters and randomly initialize the parameters of the Gaussian distributions of each cluster;
- Given these Gaussian distributions for each cluster, we calculate the probability that each point belongs to each cluster: The closer a point is to a Gaussian center, the more likely it is to belong to the associated cluster (Expectation part);
- Based on these probabilities, we re-estimate the Gaussians: We calculate a new set of parameters of Gaussian distributions, in order to maximize the probability of the points to be in the clusters (Maximization part). These new parameters are calculated using a weighted sum of the point positions, the weights being the probabilities that the points belong to this cluster;
- We repeat this until the distributions no longer change, or almost: We evaluate the log-likelihood of the data to test convergence (this is the objective function to be increased).
2.3. Density-Based Clustering
2.3.1. Mean-Shift
- For nucleus, we consider a sliding ball centered at a random point C and of radius r. Through a hill climbing approach, this nucleus is iteratively moved to an area of higher density, until convergence;
- This is done by moving the center of the ball towards the center of gravity of the ball’s points, which causes the ball to move to denser areas;
- -
- As the ball has been moved, there are potentially new points, and, therefore, a new center of gravity;
- -
- Otherwise, we end up converging.
- Points 1 and 2 are operated with various randomly placed balls, or according to a grid adapted to the data, and when several balls overlap, the less dense one is removed.
2.3.2. DBSCAN
- We start from a point P of the data, not yet visited;
- -
- If there are enough neighbors (min_samples-1) at from this point, clustering starts with P as the first (core) point of the cluster;
- -
- Otherwise, the point is labeled as noise (which can later integrate a cluster) and visited;
- The points at distance of P integrate the cluster of P, then all the points at of these points, etc., until it is no longer possible to expand the cluster;
- We start again with a point not visited, and this until the points are exhausted. At the end, any point will either be in a cluster or labeled as noise.
2.4. Tree-Based Clustering
2.4.1. Hierarchical Clustering
- Each data point is treated as a single cluster, and a distance between clusters is fixed—for example, the average linkage: the average distance between the points of two clusters;
- At each iteration, the two clusters with the shortest distance are merged, and the height of the new node in the dendrogram is the similarity between said clusters;
- Repeat 2 until you only get one cluster left;
- If a predefined number of clusters is wanted, step number 2 is stopped when the number is reached.
- single linkage: The distance between two clusters is the distance corresponding to the two most similar points.
- complete linkage: As above, but with the least similar points.
- average linkage: The average distance.
- Ward: Minimization of the sum of the squares of distances within each cluster. It is an approach of the variance minimization type, and therefore a kind of K-means coupled with a hierarchical agglomerative approach.
2.4.2. Birch
- The threshold: The radius of the subcluster obtained by merging a new individual and the nearest subcluster must be less than this threshold. If this is not the case, a new subcluster is initialized. A low value of this threshold therefore multiplies the number of node breakdowns and, thus, the clusters;
- The branching factor: The maximum number of subclusters in each node. If a new individual entering a node causes the number of subclusters to exceed this branching factor, then the node is split into two, which distribute the subclusters. The parent subgroup of this node is deleted, and two new subclusters are added as parents of these two cut nodes;
- Number of clusters after the last clustering step.
2.5. Other Methods
2.5.1. Spectral Clustering
- Either a kernel function, for example, the RBF or the heat kernel , where d is the distance between individuals, while and are hyperparameters to set up;
- Or a connectivity matrix in the k-nearest neighbors;
- Or, via the precomputed parameter, an affinity matrix provided by the user.
2.5.2. Affinity Propagation
- The R matrix of competence (responsibility), in which the element quantifies how much is justified as a model for , compared to the other candidates.
- The availability matrix A, in which the element represents how appropriate it would be for to take as a model, once all the other points have been considered as a model.
- First, we circulate the skill updates: ;
- Then the availability is updated by:
- –
- for ; and
- –
- .
2.5.3. Constrained Clustering
3. Clustering Evaluation: State-Of-The-Art
3.1. External Validation
3.1.1. Homogeneity, Completeness, and V-Measure
- Homogeneity: Each cluster contains only members of a single class;
- Completeness: All members of a given class are in the same cluster.
- is the conditional entropy of classes knowing clusters, i.e., ,
- is the entropy of the class, equal to ,
3.1.2. Adjusted Rand Index
- Random labeling leads to an ARI close to 0, regardless of the number of points and clusters, which is not the case for RI or V-measure;
- The ARI varies between −1 and 1 included. Negative values are for independent labeling, when similar clusterings have a positive ARI (1 for an exact match);
- No hypothesis on the structure of the clusters: we can therefore compare K-means to spectral clustering, leading a priori to very different structures.
3.1.3. Fowlkes–Mallows Score
- TP is the number of true positives: The number of pairs of points in the same cluster in the real and predicted labeling;
- FP is the number of false positives: Pairs with the same real labeling, but in different predicted clusters;
- FN is the number of false negatives: Pairs in the same predicted clusters, but with different actual labels.
- Unlike mutual information or V-measure, a random clustering will have an FMI score of 0, regardless of the number of clusters or individuals;
- A value close to 0 indicates largely independent labeling, while a value close to 1 indicates clustering agreement;
- Two equal labels (with permutation) have an FMI of 1;
- Finally, there is no hypothesis on the structure of clusters.
3.1.4. Based on Mutual Information
Mutual Information
Normalized Mutual Information
Adjusted Mutual Information
3.2. Internal Validation
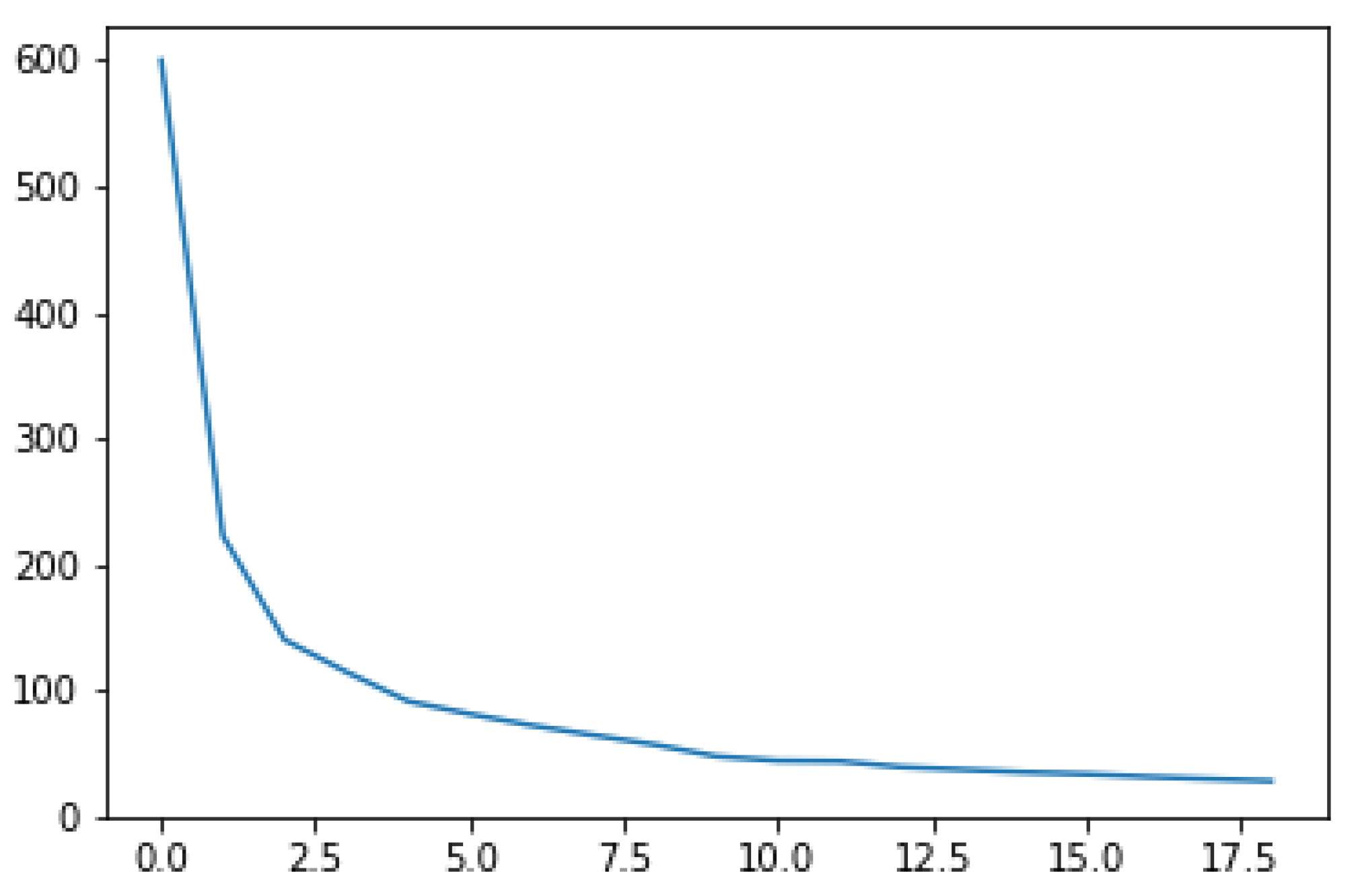
3.2.1. Elbow Method
- We sum the intracluster variances, which we divide by the overall variance;
- The sum of squared errors (SSE) is calculated: the sum, on all clusters, of the square distances between the points and their centroids.
3.2.2. Indices
Calinski-Harabaz
- is the intracluster dispersion;
- is the dispersion matrix between groups;
Davies–Bouldin
- The diameter of the cluster : average distance between each point of this cluster and its centroid;
- : the distance between the centroids of the i and j clusters.
- It can be calculated without the knowledge of true clustering;
- The calculation of the DB is simpler than the Silhouette scores, described hereafter;
- The index is calculated only from distances.
- It does not evaluate very well in a situation of nonconvexity;
- A good value produced by this index does not mean that the information collected is maximum.
Dunn
- For each cluster, the distance between each point of the cluster and the points of the other clusters is calculated. The minimum of these distances corresponds to the intercluster separation (min.separation);
- The distance between the points of each cluster is calculated, its maximum diameter being the intracluster distance reflecting the compact nature of the clustering.
3.2.3. Silhouette
- Close to 1, the object fits very well with its own cluster and very badly with other clusters;
- Near 0, the data are at the border of two clusters;
- Close to −1, the object would fit better in the neighboring cluster: probably a bad assignment.
- be the average distance between i point and the other points of its cluster, measuring how much i fits well with its own cluster (value is the smaller the better the assignment is); and
- be the smallest average distance between i and the points of all other clusters, the average dissimilarity of a point i to a cluster C being seen as the average of the distances between i and the points of C; the cluster with the smallest average dissimilarity is therefore the cluster close to i.
4. Comparison of Modern Clustering Techniques for the IoT
4.1. Experimental Protocol: A Network of Gas Sensors
4.2. Results of the Survey
- Blue: The values highlighted in blue are the highest recorded quality metrics with no outlier removal;
- Green: The values highlighted in green are the ones showing improvement as compared to the values obtained with no outlier removal;
- Red: The values highlighted in red are the ones showing degraded performance as compared to the values obtained with no outlier removal.
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Farhat, A.; Guyeux, C.; Makhoul, A.; Jaber, A.; Tawil, R. On the coverage effects in wireless sensor networks based prognostic and health management. Int. J. Sens. Netw. IJSNET 2018, 28, 125–138. [Google Scholar] [CrossRef]
- Farhat, A.; Guyeux, C.; Makhoul, A.; Jaber, A.; Tawil, R.; Hijazi, A. Impacts of wireless sensor networks strategies and topologies on prognostics and health management. J. Intell. Manuf. 2017, 30, 2129–2155. [Google Scholar] [CrossRef]
- Boudargham, N.; Makhoul, A.; Bou Abdo, J.; Demerjian, J.; Guyeux, C. Efficient Hybrid Emergency Aware MAC Protocol for Wireless Body Sensor. Sensors 2018, 18, 3572. [Google Scholar] [CrossRef] [PubMed]
- Boudargham, N.; Bou Abdo, J.; Demerjian, J.; Makhoul, A.; Guyeux, C. Efficient Cluster Based Routing Protocol for Collaborative Body Sensor Networks. In Proceedings of the Sensornets 2019, 8th International Conference on Sensor Networks, Prague, Czech Republic, 26–27 February 2019. [Google Scholar]
- Xie, Q.Y.; Cheng, Y. K-Centers Mean-shift Reverse Mean-shift clustering algorithm over heterogeneous wireless sensor networks. In Proceedings of the 2014 Wireless Telecommunications Symposium, Washington, DC, USA, 9–11 April 2014; pp. 1–6. [Google Scholar] [CrossRef]
- Zhou, Q.; Li, X.; Xu, Y. Mean Shift Based Collaborative Localization with Dynamically Clustering for Wireless Sensor Networks. In Proceedings of the 2009 WRI International Conference on Communications and Mobile Computing, Yunnan, China, 6–8 January 2009; Volume 2, pp. 66–70. [Google Scholar] [CrossRef]
- Saeedi Emadi, H.; Mazinani, S.M. A Novel Anomaly Detection Algorithm Using DBSCAN and SVM in Wireless Sensor Networks. Wirel. Pers. Commun. 2018, 98, 2025–2035. [Google Scholar] [CrossRef]
- Pan, D.; Zhao, L. Uncertain data cluster based on DBSCAN. In Proceedings of the 2011 International Conference on Multimedia Technology, Hangzhou, China, 26–28 July 2011; pp. 3781–3784. [Google Scholar] [CrossRef]
- Narasegouda, S. Fault Detection in Sensor Network Using DBSCAN and Statistical Models. In Proceedings of the International Conference on Frontiers of Intelligent Computing: Theory and Applications (FICTA) 2013; Satapathy, S.C., Udgata, S.K., Biswal, B.N., Eds.; Springer International Publishing: Cham, Switzerland, 2014; pp. 443–450. [Google Scholar]
- Hu, H.; Wang, X.; Yang, Z.; Zheng, B. A Spectral Clustering Approach to Identifying Cuts in Wireless Sensor Networks. IEEE Sens. J. 2015, 15, 1838–1848. [Google Scholar] [CrossRef]
- Kung, H.T.; Vlah, D. A Spectral Clustering Approach to Validating Sensors via Their Peers in Distributed Sensor Networks. In Proceedings of the 18th International Conference on Computer Communications and Networks, San Francisco, CA, USA, 3–6 August 2009; pp. 1–7. [Google Scholar] [CrossRef]
- Muniraju, G.; Zhang, S.; Tepedelenlioglu, C.; Banavar, M.K.; Spanias, A.; Vargas-Rosales, C.; Villalpando-Hernandez, R. Location Based Distributed Spectral Clustering for Wireless Sensor Networks. In Proceedings of the 2017 Sensor Signal Processing for Defence Conference (SSPD), London, UK, 6–7 December 2017; pp. 1–5. [Google Scholar] [CrossRef]
- Sohn, I.; Lee, J.; Lee, S.H. Low-Energy Adaptive Clustering Hierarchy Using Affinity Propagation for Wireless Sensor Networks. IEEE Commun. Lett. 2016, 20, 558–561. [Google Scholar] [CrossRef]
- ElGammal, M.; Eltoweissy, M. Location-Aware Affinity Propagation Clustering in Wireless Sensor Networks. In Proceedings of the 2009 IEEE International Conference on Wireless and Mobile Computing, Networking and Communications, Marrakech, Morocco, 12–14 October 2009; pp. 471–475. [Google Scholar] [CrossRef]
- Wang, J.; Gao, Y.; Wang, K.; Sangaiah, A.K.; Lim, S.J. An Affinity Propagation-Based Self-Adaptive Clustering Method for Wireless Sensor Networks. Sensors 2019, 19, 2579. [Google Scholar] [CrossRef]
- Sakthidasan, K.; Vasudevan, N.; Diderot, P.K.G.; Kadhiravan, C. WOAPR: An affinity propagation based clustering and optimal path selection for time-critical wireless sensor networks. IET Netw. 2018, 8, 100–106. [Google Scholar] [CrossRef]
- Zhang, T.; Zhao, Q.; Shin, K.; Nakamoto, Y. Bayesian-Optimization-Based Peak Searching Algorithm for Clustering in Wireless Sensor Networks. J. Sens. Actuator Netw. 2018, 7, 2. [Google Scholar] [CrossRef]
- Amaxilatis, D.; Chatzigiannakis, I. Design and analysis of adaptive hierarchical low-power long-range networks. J. Sens. Actuator Netw. 2018, 7, 51. [Google Scholar] [CrossRef]
- Yang, X.; Yan, Y.; Deng, D. Research on clustering routing algorithm based on K-means++ for WSN. In Proceedings of the 2017 6th International Conference on Computer Science and Network Technology (ICCSNT), Dalian, China, 21–22 October 2017; pp. 330–333. [Google Scholar] [CrossRef]
- Li, L.; Li, D.; Li, D. An Efficient Routing Algorithm based on K-means++ Clustering and Fuzzy for Wireless Sensor Network*. In Proceedings of the 2018 13th World Congress on Intelligent Control and Automation (WCICA), Changsha, China, 4–8 July 2018; pp. 716–720. [Google Scholar] [CrossRef]
- Rajasegarar, S.; Leckie, C.; Palaniswami, M.; Bezdek, J.C. Distributed Anomaly Detection in Wireless Sensor Networks. In Proceedings of the 2006 10th IEEE Singapore International Conference on Communication Systems, Singapore, 30 October–1 November 2006; pp. 1–5. [Google Scholar] [CrossRef]
- Bakaraniya, P.; Mehta, S. K-LEACH: An improved LEACH protocol for lifetime improvement in WSN. Int. J. Eng. Trends Technol. 2013, 4, 1521–1526. [Google Scholar]
- Pavithra, M.; Ghuli, P. A novel approach for reducing energy consumption using k-medoids in clustering based WSN. Int. J. Sci. Res. IJSR 2015, 4, 2279–2282. [Google Scholar]
- Mittal, R.; Bhatia, M.P.S. Wireless sensor networks for monitoring the environmental activities. In Proceedings of the 2010 IEEE International Conference on Computational Intelligence and Computing Research, Coimbatore, India, 28–29 December 2010; pp. 1–5. [Google Scholar] [CrossRef]
- Tan, L.; Gong, Y.; Chen, G. A Balanced Parallel Clustering Protocol for Wireless Sensor Networks Using K-Means Techniques. In Proceedings of the 2008 Second International Conference on Sensor Technologies and Applications (Sensorcomm 2008), Cap Esterel, France, 25–31 August 2008; pp. 300–305. [Google Scholar] [CrossRef]
- Park, H.S.; Jun, C.H. A simple and fast algorithm for K-medoids clustering. Expert Syst. Appl. 2009, 36, 3336–3341. [Google Scholar] [CrossRef]
- McLachlan, G.J.; Lee, S.X.; Rathnayake, S.I. Finite mixture models. Annu. Rev. Stat. Its Appl. 2019, 6, 355–378. [Google Scholar] [CrossRef]
- McLachlan, G.; Krishnan, T. The EM Algorithm and Extensions; John Wiley & Sons: Hoboken, NJ, USA, 2007; Volume 382. [Google Scholar]
- Chrétien, S.; Hero, A.O. Kullback proximal algorithms for maximum-likelihood estimation. IEEE Trans. Inf. Theory 2000, 46, 1800–1810. [Google Scholar] [CrossRef]
- Chrétien, S.; Hero, A.O. On EM algorithms and their proximal generalizations. ESAIM Probab. Stat. 2008, 12, 308–326. [Google Scholar] [CrossRef]
- Celeux, G.; Chrétien, S.; Forbes, F.; Mkhadri, A. A component-wise EM algorithm for mixtures. J. Comput. Graph. Stat. 2001, 10, 697–712. [Google Scholar] [CrossRef]
- Biernacki, C.; Chrétien, S. Degeneracy in the maximum likelihood estimation of univariate Gaussian mixtures with EM. Stat. Probab. Lett. 2003, 61, 373–382. [Google Scholar] [CrossRef]
- Chrétien, S.; Hero, A.; Perdry, H. Space alternating penalized Kullback proximal point algorithms for maximizing likelihood with nondifferentiable penalty. Ann. Inst. Stat. Math. 2012, 64, 791–809. [Google Scholar] [CrossRef]
- Comaniciu, D.; Meer, P. Mean shift: A robust approach toward feature space analysis. IEEE Trans. Pattern Anal. Mach. Intell. 2002, 5, 603–619. [Google Scholar] [CrossRef]
- Ester, M.; Kriegel, H.P.; Sander, J.; Xu, X. A density-based algorithm for discovering clusters in large spatial databases with noise. Kdd 1996, 96, 226–231. [Google Scholar]
- McInnes, L.; Healy, J.; Astels, S. hdbscan: Hierarchical density based clustering. J. Open Source Softw. 2017, 2. [Google Scholar] [CrossRef]
- Zhang, T.; Ramakrishnan, R.; Livny, M. BIRCH: An efficient data clustering method for very large databases. In Proceedings of the ACM Sigmod Record, Montreal, QC, Canada, 4–6 June 1996; Volume 25, pp. 103–114. [Google Scholar]
- Von Luxburg, U. A tutorial on spectral clustering. Stat. Comput. 2007, 17, 395–416. [Google Scholar] [CrossRef]
- Chretien, S.; Jagan, K.; Barton, E. Clustering on low-dimensional latent manifolds via Laplacianeigenmaps when clusters overlap. Meas. Sci. Technol. 2019. submitted. [Google Scholar]
- Frey, B.J.; Dueck, D. Clustering by passing messages between data points. Science 2007, 315, 972–976. [Google Scholar] [CrossRef] [PubMed]
- Wagstaff, K.; Cardie, C.; Rogers, S.; Schrödl, S. Constrained k-means clustering with background knowledge. Icml 2001, 1, 577–584. [Google Scholar]
- Active-Semi-Supervised-Clustering Repository. Available online: https://github.com/datamole-ai/active-semi-supervised-clustering (accessed on 29 May 2019).
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Rosenberg, A.; Hirschberg, J. V-measure: A conditional entropy-based external cluster evaluation measure. In Proceedings of the 2007 Joint Conference on Empirical Methods in Natural Language Processing and Computational Natural Language Learning (EMNLP-CoNLL), Prague, Czech Republic, 28–30 June 2007. [Google Scholar]
- Fowlkes, E.B.; Mallows, C.L. A method for comparing two hierarchical clusterings. J. Am. Stat. Assoc. 1983, 78, 553–569. [Google Scholar] [CrossRef]
- Meilă, M. Comparing clusterings—An information based distance. J. Multivar. Anal. 2007, 98, 873–895. [Google Scholar] [CrossRef]
- Vinh, N.X.; Epps, J.; Bailey, J. Information theoretic measures for clusterings comparison: Variants, properties, normalization and correction for chance. J. Mach. Learn. Res. 2010, 11, 2837–2854. [Google Scholar]
- Thorndike, R.L. Who belongs in the family? Psychometrika 1953, 18, 267–276. [Google Scholar] [CrossRef]
- Caliński, T.; Harabasz, J. A dendrite method for cluster analysis. Commun. Stat. Theory Methods 1974, 3, 1–27. [Google Scholar] [CrossRef]
- Davies, D.L.; Bouldin, D.W. A cluster separation measure. IEEE Trans. Pattern Anal. Mach. Intell. 1979, 2, 224–227. [Google Scholar] [CrossRef]
- Dunn, J.C. Well-separated clusters and optimal fuzzy partitions. J. Cybern. 1974, 4, 95–104. [Google Scholar] [CrossRef]
- Rousseeuw, P.J. Silhouettes: A graphical aid to the interpretation and validation of cluster analysis. J. Comput. Appl. Math. 1987, 20, 53–65. [Google Scholar] [CrossRef]
- UC Irvine Machine Learning Repository. Available online: https://archive.ics.uci.edu/ml/index.php (accessed on 17 July 2019).
- Liu, F.T.; Ting, K.M.; Zhou, Z.H. Isolation Forest. In Proceedings of the 2008 Eighth IEEE International Conference on Data Mining, ICDM ’08, Pisa, Italy, 15–19 December 2008; IEEE Computer Society: Washington, DC, USA, 2008; pp. 413–422. [Google Scholar] [CrossRef]
- Hariri, S.; Carrasco Kind, M.; Brunner, R.J. Extended Isolation Forest. arXiv 2018, arXiv:1811.02141. [Google Scholar] [CrossRef]
- Zou, H.; Hastie, T.; Tibshirani, R. Sparse principal component analysis. J. Comput. Graph. Stat. 2006, 15, 265–286. [Google Scholar] [CrossRef]
- Maaten, L.V.D.; Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 2008, 9, 2579–2605. [Google Scholar]
| Method | Parameters | Cluster Size | Nb of Clusters | Geometry |
|---|---|---|---|---|
| Affinity | damping | distinct | large nb of clusters; does not scale | non flat |
| propagation | sample preference | up with the nb of individuals | ||
| Birch | numerous | large data sets | large nb of clusters | non flat |
| DBSCAN | neighborhood size | distinct | Very large set of individuals | non flat |
| average number of clusters | ||||
| GMM | nb of clusters | not scalable | not scalable | flat |
| Hierarchical | nb of clusters | distinct | many clusters | hierarchical |
| clustering | link type, distance | many samples | ||
| K-means | nb of clusters | regular | not too many | flat |
| K-Medoids | nb of clusters | regular | not too many | flat |
| Mean-shift | bandwidth | distinct | many | non flat |
| Spectral | nb of clusters | distinct | small | non flat |
| Methods | Advantages | Disadvantages |
|---|---|---|
| Affinity | - Nb of clusters not required | - Quadratic complexity in the nb of points |
| propagation | - One main parameter: the damping factor 1 | - There may not be convergence |
| Birch | - Outlier deletion, data reduction | - Does not adapt very well to large data 2 |
| - If large number of subclusters is desired | ||
| DBSCAN | - No need to provide a priori the nb of clusters | - Works less well than others when the clusters |
| - Useful for identifying outliers, noise | to be found have different densities 3 | |
| - Can find clusters of arbitrary size and shape | - Does not work well in very high dimension | |
| - Works well for clusters of homogeneous density | ||
| GMM | - Intuitive algorithm associated with maximum likelihood maximisation | - Not scalable |
| - Soft clustering: each point belongs to all clusters, but with different probabilities | ||
| - Allows an estimation of density | ||
| - model based approach that comes with easy to implement information theoretic | ||
| penalties for model selection (number of clusters, etc.) | ||
| Hierarchical | - Nb of clusters not required | - Its specificity for hierarchical data |
| clustering | - The most appropriate cluster nb can be chosen afterwards | - High impact of outliers |
| - Choice of distance not critical | - Slow: | |
| - Works well when data is naturally hierarchical allows you to recover this hierarchy | ||
| K-means | - Fast (linear complexity) | - We have to choose the nb of clusters |
| - Easy to implement | - Not very good, outside of spherical clusters | |
| - Proven in many applications | - Consistency problem: | |
| - Scalable: | different runs produce different | |
| for a large number of individuals | clustering, due to its random initialization | |
| and a reasonable number of clusters 4 | ||
| K-Medoids | - Robust against noise and presence of outliers | - We have to choose the nb of clusters |
| Mean-shift | - No need to provide a priori the nb of clusters | - Window size can be difficult to fix |
| - The centers of the balls converge towards the places of highest density | ||
| Spectral | - Useful when cluster structure is highly non-convex | - Nb of clusters must be provided |
| - Very effective with sparse affinity matrix | - Not recommended for large nb of clusters |
| Outliers | Clustering | Homogeneity | Completeness | ARI | V-Measure | Fowlkes–Mallows | AMI |
|---|---|---|---|---|---|---|---|
| None | kmeans | 0.1983 | 0.2471 | 0.0797 | 0.2201 | 0.2936 | 0.2196 |
| kmeans++ | 0.1739 | 0.2409 | 0.0772 | 0.2020 | 0.3021 | 0.2015 | |
| AC ward | 0.1757 | 0.2337 | 0.0869 | 0.2006 | 0.3006 | 0.2001 | |
| AC complete | 0.0003 | 0.1554 | −3.324 | 0.0006 | 0.4189 | −5.143 | |
| AC average | 0.0003 | 0.1554 | −3.324 | 0.0006 | 0.4189 | −5.143 | |
| GMM | 0.3196 | 0.3613 | 0.2449 | 0.3392 | 0.4012 | 0.3388 | |
| Birch | 0.1811 | 0.3273 | 0.0916 | 0.2332 | 0.3564 | 0.2327 | |
| Isolation Forest | kmeans | 0.2059 | 0.2619 | 0.0850 | 0.2306 | 0.3020 | 0.2301 |
| kmeans++ | 0.1608 | 0.2233 | 0.0772 | 0.1869 | 0.3031 | 0.1865 | |
| AC ward | 0.1795 | 0.2341 | 0.0854 | 0.2032 | 0.2981 | 0.2028 | |
| AC complete | 0.0003 | 0.1550 | −3.581 | 0.0006 | 0.4184 | −5.419 | |
| AC average | 0.0003 | 0.1550 | −3.581 | 0.0006 | 0.4184 | −5.419 | |
| GMM | 0.2783 | 0.3034 | 0.1669 | 0.2903 | 0.3291 | 0.2899 | |
| Birch | 0.1856 | 0.3274 | 0.0922 | 0.2369 | 0.3557 | 0.2365 | |
| Extended IF | kmeans | 0.1962 | 0.2447 | 0.0794 | 0.2177 | 0.2934 | 0.2173 |
| kmeans++ | 0.1731 | 0.2401 | 0.0768 | 0.2012 | 0.3018 | 0.2007 | |
| AC ward | 0.1792 | 0.2363 | 0.0889 | 0.2038 | 0.3017 | 0.2034 | |
| AC complete | 0.0003 | 0.1551 | −3.503 | 0.0006 | 0.4185 | −5.334 | |
| AC average | 0.0003 | 0.1551 | −3.503 | 0.0006 | 0.4185 | −5.334 | |
| GMM | 0.2728 | 0.3460 | 0.1826 | 0.3051 | 0.3661 | 0.3047 | |
| Birch | 0.1806 | 0.3293 | 0.0939 | 0.2333 | 0.3581 | 0.2328 |
| Outliers | Clustering | Homogeneity | Completeness | ARI | V-Measure | Fowlkes–Mallows | AMI |
|---|---|---|---|---|---|---|---|
| None | kmeans | 0.1676 | 0.1893 | 0.0822 | 0.1778 | 0.2682 | 0.1773 |
| kmeans++ | 0.1674 | 0.1891 | 0.0820 | 0.1776 | 0.2681 | 0.1771 | |
| AC ward | 0.2302 | 0.2716 | 0.1131 | 0.2492 | 0.3072 | 0.2488 | |
| AC complete | 0.0848 | 0.1794 | 0.0690 | 0.1152 | 0.3474 | 0.1146 | |
| AC average | 0.0108 | 0.2496 | −0.002 | 0.0208 | 0.4115 | 0.0199 | |
| GMM | 0.2071 | 0.2580 | 0.0950 | 0.2298 | 0.3043 | 0.2294 | |
| Birch | 0.1006 | 0.2026 | 0.0115 | 0.1345 | 0.3206 | 0.1339 | |
| Isolation forest | kmeans | 0.1752 | 0.1920 | 0.0863 | 0.1833 | 0.2665 | 0.1828 |
| kmeans++ | 0.1754 | 0.1923 | 0.0862 | 0.1835 | 0.2667 | 0.1830 | |
| AC ward | 0.1889 | 0.2335 | 0.0763 | 0.2088 | 0.2915 | 0.2084 | |
| AC complete | 0.0745 | 0.2994 | −0.001 | 0.1194 | 0.3746 | 0.1186 | |
| AC average | 0.0410 | 0.1675 | −0.006 | 0.0659 | 0.3601 | 0.0651 | |
| GMM | 0.2006 | 0.2346 | 0.0954 | 0.2162 | 0.2920 | 0.2158 | |
| Birch | 0.1282 | 0.2751 | 0.0178 | 0.1749 | 0.3351 | 0.1743 | |
| Extended IF | kmeans | 0.1746 | 0.1916 | 0.0855 | 0.1827 | 0.2664 | 0.1822 |
| kmeans++ | 0.1750 | 0.1924 | 0.0857 | 0.1833 | 0.2668 | 0.1829 | |
| AC ward | 0.1479 | 0.1761 | 0.0485 | 0.1608 | 0.2558 | 0.1603 | |
| AC complete | 0.1315 | 0.3068 | 0.0219 | 0.1841 | 0.3423 | 0.1835 | |
| AC average | 0.0080 | 0.2333 | -0.001 | 0.0155 | 0.4131 | 0.0146 | |
| GMM | 0.2181 | 0.2553 | 0.1022 | 0.2352 | 0.3011 | 0.2348 | |
| Birch | 0.1522 | 0.2705 | 0.0696 | 0.1948 | 0.3361 | 0.1943 |
| Outliers | Clustering | Homogeneity | Completeness | ARI | V-Measure | Fowlkes–Mallows | AMI |
|---|---|---|---|---|---|---|---|
| None | kmeans | 0.1975 | 0.2441 | 0.0796 | 0.2184 | 0.2937 | 0.2179 |
| kmeans++ | 0.2039 | 0.2515 | 0.0885 | 0.2252 | 0.2989 | 0.2248 | |
| AC ward | 0.2198 | 0.2851 | 0.0852 | 0.2482 | 0.3102 | 0.2478 | |
| AC complete | 0.0706 | 0.3567 | 0.0062 | 0.1179 | 0.3887 | 0.1171 | |
| AC average | 0.0129 | 0.2305 | −0.002 | 0.0244 | 0.4098 | 0.0235 | |
| GMM | 0.2249 | 0.2754 | 0.1017 | 0.2476 | 0.3095 | 0.2472 | |
| Birch | −6.291 | 1.0 | 0.0 | −1.258 | 0.4190 | −2.293 | |
| Isolation forest | kmeans | 0.2297 | 0.2666 | 0.0979 | 0.2468 | 0.2981 | 0.2463 |
| kmeans++ | 0.2297 | 0.2666 | 0.0979 | 0.2468 | 0.2981 | 0.2464 | |
| AC ward | 0.2247 | 0.2793 | 0.0764 | 0.2490 | 0.3023 | 0.2486 | |
| AC complete | 0.0732 | 0.2810 | 0.0168 | 0.1162 | 0.3781 | 0.1155 | |
| AC average | 0.0074 | 0.2255 | -0.001 | 0.0143 | 0.4133 | 0.0134 | |
| GMM | 0.2388 | 0.2839 | 0.0990 | 0.2594 | 0.3074 | 0.2590 | |
| Birch | -3.143 | 1.0 | 0.0 | −6.287 | 0.4186 | −6.287 | |
| Extended IF | kmeans | 0.2313 | 0.2685 | 0.0981 | 0.2485 | 0.2985 | 0.2481 |
| kmeans++ | 0.2083 | 0.2522 | 0.0893 | 0.2281 | 0.2981 | 0.2277 | |
| AC ward | 0.2484 | 0.3249 | 0.1045 | 0.2816 | 0.3294 | 0.2812 | |
| AC complete | 0.1168 | 0.3113 | 0.0084 | 0.1699 | 0.3485 | 0.1693 | |
| AC average | 0.0079 | 0.2364 | -0.001 | 0.0154 | 0.4133 | 0.0145 | |
| GMM | 0.2082 | 0.2560 | 0.0943 | 0.2296 | 0.3030 | 0.2292 | |
| Birch | 3.1444 | 1.0 | 0.0 | 6.2889 | 0.4187 | 1.1054 |
| Outliers | Clustering | Homogeneity | Completeness | ARI | V-Measure | Fowlkes–Mallows | AMI |
|---|---|---|---|---|---|---|---|
| None | kmeans | 0.3519 | 0.3476 | 0.2418 | 0.3497 | 0.3723 | 0.3494 |
| kmeans++ | 0.3537 | 0.3495 | 0.2427 | 0.3516 | 0.3730 | 0.3513 | |
| AC ward | 0.3552 | 0.3593 | 0.2223 | 0.3572 | 0.3618 | 0.3569 | |
| AC complete | 0.3123 | 0.3571 | 0.2209 | 0.3332 | 0.3793 | 0.3328 | |
| AC average | 0.2922 | 0.3467 | 0.1994 | 0.3171 | 0.3684 | 0.3168 | |
| GMM | 0.3443 | 0.3556 | 0.2138 | 0.3498 | 0.3587 | 0.3495 | |
| Birch | 0.3547 | 0.3608 | 0.2370 | 0.3578 | 0.3749 | 0.3574 | |
| Isolation forest | kmeans | 0.3883 | 0.3857 | 0.2904 | 0.3870 | 0.4134 | 0.3867 |
| kmeans++ | 0.3888 | 0.3862 | 0.2908 | 0.3875 | 0.4137 | 0.3872 | |
| AC ward | 0.3405 | 0.3527 | 0.2436 | 0.3465 | 0.3840 | 0.3462 | |
| AC complete | 0.2992 | 0.3311 | 0.2338 | 0.3143 | 0.3801 | 0.3140 | |
| AC average | 0.4221 | 0.5147 | 0.3666 | 0.4638 | 0.5066 | 0.4635 | |
| GMM | 0.3608 | 0.3617 | 0.2666 | 0.3613 | 0.3957 | 0.3610 | |
| Birch | 0.3648 | 0.3696 | 0.2820 | 0.3672 | 0.4104 | 0.3669 | |
| Extended IF | kmeans | 0.3272 | 0.3233 | 0.2238 | 0.3253 | 0.3572 | 0.3249 |
| kmeans++ | 0.3272 | 0.3233 | 0.2238 | 0.3253 | 0.3572 | 0.3249 | |
| AC ward | 0.4051 | 0.4041 | 0.2972 | 0.4046 | 0.4201 | 0.4043 | |
| AC complete | 0.2394 | 0.2432 | 0.1434 | 0.2413 | 0.2974 | 0.2409 | |
| AC average | 0.3611 | 0.3723 | 0.2281 | 0.3666 | 0.3705 | 0.3663 | |
| GMM | 0.3723 | 0.3802 | 0.2343 | 0.3762 | 0.3732 | 0.3759 | |
| Birch | 0.4109 | 0.4201 | 0.2926 | 0.4154 | 0.4208 | 0.4151 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guyeux, C.; Chrétien, S.; Bou Tayeh, G.; Demerjian, J.; Bahi, J. Introducing and Comparing Recent Clustering Methods for Massive Data Management in the Internet of Things. J. Sens. Actuator Netw. 2019, 8, 56. https://doi.org/10.3390/jsan8040056
Guyeux C, Chrétien S, Bou Tayeh G, Demerjian J, Bahi J. Introducing and Comparing Recent Clustering Methods for Massive Data Management in the Internet of Things. Journal of Sensor and Actuator Networks. 2019; 8(4):56. https://doi.org/10.3390/jsan8040056
Chicago/Turabian StyleGuyeux, Christophe, Stéphane Chrétien, Gaby Bou Tayeh, Jacques Demerjian, and Jacques Bahi. 2019. "Introducing and Comparing Recent Clustering Methods for Massive Data Management in the Internet of Things" Journal of Sensor and Actuator Networks 8, no. 4: 56. https://doi.org/10.3390/jsan8040056
APA StyleGuyeux, C., Chrétien, S., Bou Tayeh, G., Demerjian, J., & Bahi, J. (2019). Introducing and Comparing Recent Clustering Methods for Massive Data Management in the Internet of Things. Journal of Sensor and Actuator Networks, 8(4), 56. https://doi.org/10.3390/jsan8040056






