A Fresh Look at Grape Powdery Mildew (Erysiphe necator) A and B Genotypes Revealed Frequent Mixed Infections and Only B Genotypes in Flag Shoot Samples
Abstract
:1. Introduction
2. Results
2.1. Direct Sequencing of the Target TUB2, ITS, and IGS Regions
2.2. Sequencing Following Cloning of Selected TUB2 and IGS PCR Products
2.3. Genotypes Identified in Flag Shoot Samples
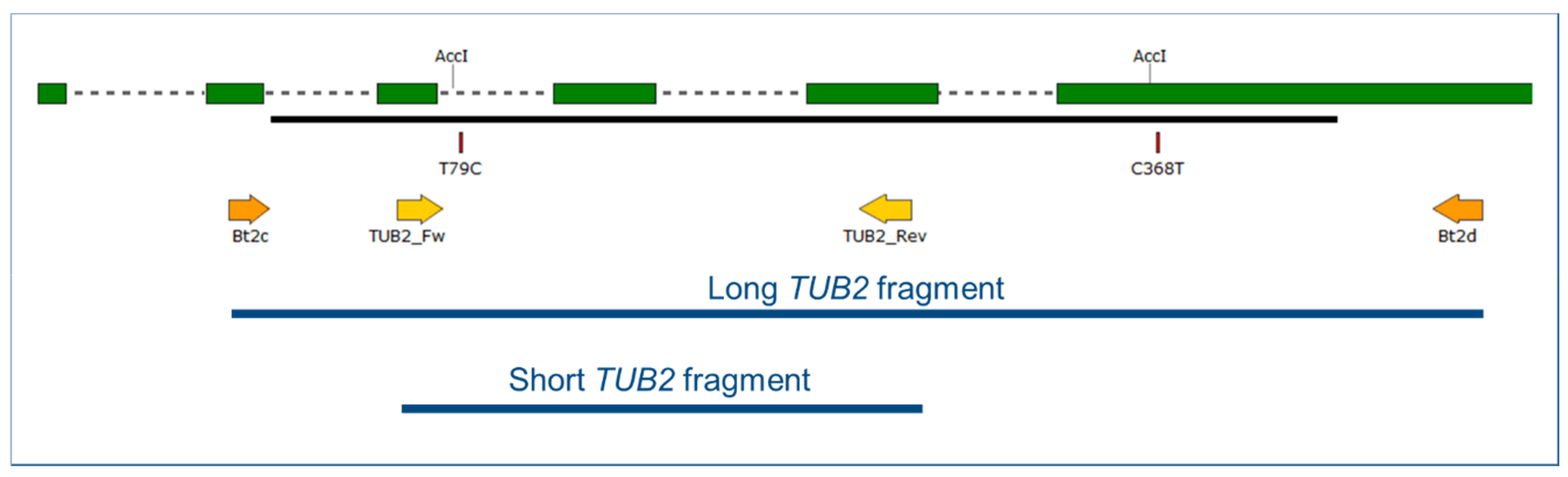
2.4. Cleaved Amplified Polymorphic Sequence (CAPS) Analysis of the TUB2 and IGS Fragments
3. Discussion
4. Materials and Methods
4.1. Erysiphe necator Sample Collection from 2009 to 2011
4.2. Erysiphe necator Samples Collected from Flag Shoot Sites in 2015
4.3. PCR Amplification of the Target TUB2, ITS, and IGS Regions in E. necator Samples
4.4. Direct Sequencing of the Target TUB2, ITS, and IGS Regions
4.5. Sequencing Following Cloning of Selected TUB2 PCR Products
4.6. Sequence Analyses
4.7. Cleaved Amplified Polymorphic Sequence (CAPS) Analysis of the TUB2 and IGS Fragments
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Gadoury, D.M.; Cadle-Davidson, L.; Wilcox, W.F.; Dry, I.B.; Seem, R.C.; Milgroom, M.G. Grapevine powdery mildew (Erysiphe necator): A fascinating system for the study of the biology, ecology and epidemiology of an obligate biotroph. Mol. Plant Pathol. 2012, 13, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Délye, C.; Laigret, F.; Corio-Costet, M.-F. RAPD analysis provides insight into the biology and epidemiology of Uncinula necator. Phytopathology 1997, 87, 670–677. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Evans, K.; Whisson, D.; Stummer, B.; Scott, E. DNA markers identify variation in Australian populations of Uncinula necator. Mycol. Res. 1997, 101, 923–932. [Google Scholar] [CrossRef]
- Délye, C.; Ronchi, V.; Laigret, F.; Corio-Costet, M.-F. Nested allele-specific PCR primers distinguish genetic groups of Uncinula necator. Appl. Environ. Microbiol. 1999, 65, 3950–3954. [Google Scholar] [CrossRef] [Green Version]
- Amrani, L.; Corio-Costet, M.F. A single nucleotide polymorphism in the β-tubulin gene distinguishing two genotypes of Erysiphe necator expressing different symptoms on grapevine. Plant Pathol. 2006, 55, 505–512. [Google Scholar] [CrossRef]
- Brewer, M.T.; Milgroom, M.G. Phylogeography and population structure of the grape powdery mildew fungus, Erysiphe necator, from diverse Vitis species. BMC Evol. Biol. 2010, 10, 268. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Péros, J.-P.; Troulet, C.; Guerriero, M.; Michel-Romiti, C.; Notteghem, J.-L. Genetic variation and population structure of the grape powdery mildew fungus, Erysiphe necator, in Southern France. Eur. J. Plant Pathol. 2005, 113, 407–416. [Google Scholar] [CrossRef]
- Montarry, J.; Cartolaro, P.; Richard-Cervera, S.; Delmotte, F. Spatio-temporal distribution of Erysiphe necator genetic groups and their relationship with disease levels in vineyards. Eur. J. Plant Pathol. 2008, 123, 61–70. [Google Scholar] [CrossRef] [Green Version]
- Miazzi, M.; Hajjeh, H.; Faretra, F. Occurence and distribution of two distinct genetic groups in populations of Erysiphe necator Schw. in Southern Italy. J. Plant Pathol. 2008, 90, 563–573. [Google Scholar]
- Montarry, J.; Cartolaro, P.; Delmotte, F.; Jolivet, J.; Willocquet, L. Genetic structure and aggressiveness of Erysiphe necator populations during grapevine powdery mildew epidemics. Appl. Environ. Microbiol. 2008, 74, 6327–6332. [Google Scholar] [CrossRef] [Green Version]
- Miazzi, M.; Hajjeh, H. Differential sensitivity to triadimenol of Erysiphe necator isolates belonging to different genetic groups. J. Plant Pathol. 2011, 93, 729–735. [Google Scholar]
- Cortesi, P.; Mazzoleni, A.; Pizzatti, C.; Milgroom, M.G. Genetic similarity of flag shoot and ascospore subpopulations of Erysiphe necator in Italy. Appl. Environ. Microbiol. 2005, 71, 7788–7791. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Núñez, Y.; Gallego, J.; Ponz, F.; Raposo, R. Analysis of population structure of Erysiphe necator using AFLP markers. Plant Pathol. 2006, 55, 650–656. [Google Scholar] [CrossRef]
- Bulit, J.; Lafon, R. Powdery mildew of the vine. In The Powdery Mildews; Spencer, D.M., Ed.; Academic Press: London, UK, 1978; pp. 525–548. [Google Scholar]
- Araya, C.; Rosales, I.; Mendez, M.A.; Delmotte, F. Identification and geographic distribution of genetic groups of Erysiphe necator in Chilean vineyards. Vitis 2014, 53, 163–165. [Google Scholar]
- Cooper, J.; Park, D.; Johnston, P. An initial genetic characterisation of the grape powdery mildew (Erysiphe necator) in New Zealand associated with recent reports of the sexual stage. N. Z. Plant Prot. 2015, 68, 389–395. [Google Scholar] [CrossRef] [Green Version]
- Sawant, I.; Ghule, S.; Sawant, S. Molecular analysis reveals that lack of chasmothecia formation in Erysiphe necator in Maharashtra, India is due to presence of only MAT1-2 mating type idiomorph. Vitis 2015, 54, 87–90. [Google Scholar]
- Oliveira, M.; Cunha, M. Study of the Portuguese populations of powdery mildew fungus from diverse grapevine cultivars (Vitis vinifera). J. Int. Sci. Vigne Vin. 2015, 49, 173–182.19. [Google Scholar] [CrossRef] [Green Version]
- Rügner, A.; Rumbolz, J.; Huber, B.; Bleyer, G.; Gisi, U.; Kassemeyer, H.H.; Guggenheim, R. Formation of overwintering structures of Uncinula necator and colonization of grapevine under field conditions. Plant Pathol. 2002, 51, 322–330. [Google Scholar] [CrossRef]
- Gadoury, D.M.; Pearson, R.C. Initiation, development, dispersal and survival of cleistothecia of Uncinula necator in New York vineyards. Phytopathology 1988, 78, 1413–1421. [Google Scholar] [CrossRef]
- Cortesi, P.; Bisiach, M.; Ricciolini, M.; Gadoury, D.M. Cleistothecia of Uncinula necator—An additional source of inoculum in Italian vineyards. Plant Dis. 1997, 81, 922–926. [Google Scholar] [CrossRef] [Green Version]
- Rossi, V.; Caffi, T.; Legler, S.E. Dynamics of ascospore maturation and discharge in Erysiphe necator, the causal agent of grape powdery mildew. Phytopathology 2010, 100, 1321–1329. [Google Scholar] [CrossRef] [PubMed]
- Vági, P.; Caffi, T.; Váczy, K.Z.; Németh, M.Z.; Kiss, L. Refining a method for ascospore viability testing in overwintering chasmothecia of Erysiphe necator. Eur. J. Plant Pathol. 2016, 144, 799–802. [Google Scholar] [CrossRef] [Green Version]
- Délye, C.; Corio-Costet, M.-F. Origin of primary infections of grape by Uncinula necator: RAPD analysis discriminates two biotypes. Mycol. Res. 1998, 102, 283–288. [Google Scholar] [CrossRef]
- Miazzi, M.; Hajjeh, H.; Faretra, F. Observations on the population biology of the grape powdery mildew fungus Uncinula necator. J. Plant Pathol. 2003, 85, 123–129. [Google Scholar]
- Stummer, B.; Scott, E. Detection of novel genotypes in progeny from a controlled cross between isolates of Uncinula necator belonging to distinct phenetic groups. Australas. Plant Path. 2003, 32, 213–218. [Google Scholar] [CrossRef]
- Pintye, A.; Németh, M.Z.; Molnár, O.; Horváth, Á.N.; Spitzmüller, Z.; Szalóki, N.; Pál, K.; Váczy, K.Z.; Kovács, G.M. Improved DNA extraction and quantitative real-time PCR for genotyping Erysiphe necator and detecting the DMI fungicide resistance marker A495T, using single ascocarps. Phytopathol. Mediterr. 2020, 59, 97–106. [Google Scholar]
- Hoffmann, P.; Virányi, F. The occurrence of cleistothecia of Erysiphe necator (Grapevine powdery mildew) and their epidemiological significance in some vine-growing regions of Hungary. Acta Phytopathol. Entomol. Hung. 2007, 42, 9–16. [Google Scholar] [CrossRef]
- Frenkel, O.; Portillo, I.; Brewer, M.; Peros, J.-P.; Cadle-Davidson, L.; Milgroom, M. Development of microsatellite markers from the transcriptome of Erysiphe necator for analysing population structure in North America and Europe. Plant Pathol. 2012, 61, 106–119. [Google Scholar] [CrossRef]
- Schoch, C.L.; Seifert, K.A.; Huhndorf, S.; Robert, V.; Spouge, J.L.; Levesque, C.A.; Chen, W.; Fungal Barcoding, C.; Fungal Barcoding Consortium Author, L. Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc. Natl. Acad. Sci. USA 2012, 109, 6241–6246. [Google Scholar] [CrossRef] [Green Version]
- Jones, L.; Riaz, S.; Morales-Cruz, A.; Amrine, K.C.; McGuire, B.; Gubler, W.D.; Walker, M.A.; Cantu, D. Adaptive genomic structural variation in the grape powdery mildew pathogen, Erysiphe necator. BMC Genom. 2014, 15, 1081. [Google Scholar] [CrossRef] [Green Version]
- Carbone, I.; Kohn, L.M. A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia 1999, 91, 553–556. [Google Scholar] [CrossRef]
- Inoue, H.; Nojima, H.; Okayama, H. High efficiency transformation of Escherichia coli with plasmids. Gene 1990, 96, 23–28. [Google Scholar] [CrossRef]
- Lesemann, S.; Schimpke, S.; Dunemann, F.; Deising, H. Mitochondrial heteroplasmy for the cytochrome b gene controls the level of strobilurin resistance in the apple powdery mildew fungus Podosphaera leucotricha (Ell. & Ev.) ES Salmon. J. Plant. Dis. Prot. 2006, 113, 259–266. [Google Scholar]
- Kovács, G.M.; Jankovics, T.; Kiss, L. Variation in the nrDNA ITS sequences of some powdery mildew species: Do routine molecular identification procedures hide valuable information? Eur. J. Plant Pathol. 2011, 131, 135–141. [Google Scholar] [CrossRef]
- Staden, R.; Beal, K.F.; Bonfield, J.K. The Staden package, 1998. Methods Mol. Biol. 2000, 132, 115–130. [Google Scholar]
- McWilliam, H.; Li, W.; Uludag, M.; Squizzato, S.; Park, Y.M.; Buso, N.; Cowley, A.P.; Lopez, R. Analysis Tool Web Services from the EMBL-EBI. Nucleic Acids Res. 2013, 41, W597–W600. [Google Scholar] [CrossRef] [Green Version]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [Green Version]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [Green Version]
- Feau, N.; Decourcelle, T.; Husson, C.; Desprez-Loustau, M.-L.; Dutech, C. Finding single copy genes out of sequenced genomes for multilocus phylogenetics in non-model fungi. PLoS ONE 2011, 6, e18803. [Google Scholar] [CrossRef]

| Target | GenBank Acc. No. of the Reference Sequence | Primer Sequence (5′ → 3′) | Amplicon Size (bp) | Restriction Enzyme | Enzyme Recognition Site | SNP Position | Detected SNPs | Geno-Type | Expected Sizes (bp) |
|---|---|---|---|---|---|---|---|---|---|
| short TUB2 fragment | GQ255475 | TUB2_Fw: AATTGGGGCTGCTTTCTGTA | 213 | AccI | GTMKAC | 79 | T | A | 213 (not digested) |
| TUB2_Rev: AACATTCATCCTTTCGAGCTG | C | B | 190, 23 | ||||||
| long TUB2 fragment | Bt2c: CAGACTGGCCAATGCGTA | 520 | 79, 368 | T, C | A | 32, 138 | |||
| Bt2d: AGTTCAGCACCCTCGGTGTA | C, C | B | 289, 138, 93 | ||||||
| C, T | B2 | 427, 93 | |||||||
| IGS fragment | GQ255476 | IGS_Fw: GTTGGGATCCTCCTCCAG | 295 | XhoI | CTCGAG | 108 | C | A | 260, 35 |
| NS1R: GAGACAAGCATATGACTAC | T | B | 295 (not digested) |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Csikós, A.; Németh, M.Z.; Frenkel, O.; Kiss, L.; Váczy, K.Z. A Fresh Look at Grape Powdery Mildew (Erysiphe necator) A and B Genotypes Revealed Frequent Mixed Infections and Only B Genotypes in Flag Shoot Samples. Plants 2020, 9, 1156. https://doi.org/10.3390/plants9091156
Csikós A, Németh MZ, Frenkel O, Kiss L, Váczy KZ. A Fresh Look at Grape Powdery Mildew (Erysiphe necator) A and B Genotypes Revealed Frequent Mixed Infections and Only B Genotypes in Flag Shoot Samples. Plants. 2020; 9(9):1156. https://doi.org/10.3390/plants9091156
Chicago/Turabian StyleCsikós, Anett, Márk Z. Németh, Omer Frenkel, Levente Kiss, and Kálmán Zoltán Váczy. 2020. "A Fresh Look at Grape Powdery Mildew (Erysiphe necator) A and B Genotypes Revealed Frequent Mixed Infections and Only B Genotypes in Flag Shoot Samples" Plants 9, no. 9: 1156. https://doi.org/10.3390/plants9091156
APA StyleCsikós, A., Németh, M. Z., Frenkel, O., Kiss, L., & Váczy, K. Z. (2020). A Fresh Look at Grape Powdery Mildew (Erysiphe necator) A and B Genotypes Revealed Frequent Mixed Infections and Only B Genotypes in Flag Shoot Samples. Plants, 9(9), 1156. https://doi.org/10.3390/plants9091156








