Molecular Diagnostics and Detection of Oomycetes on Fiber Crops
Abstract
1. Introduction
2. Fiber Crops
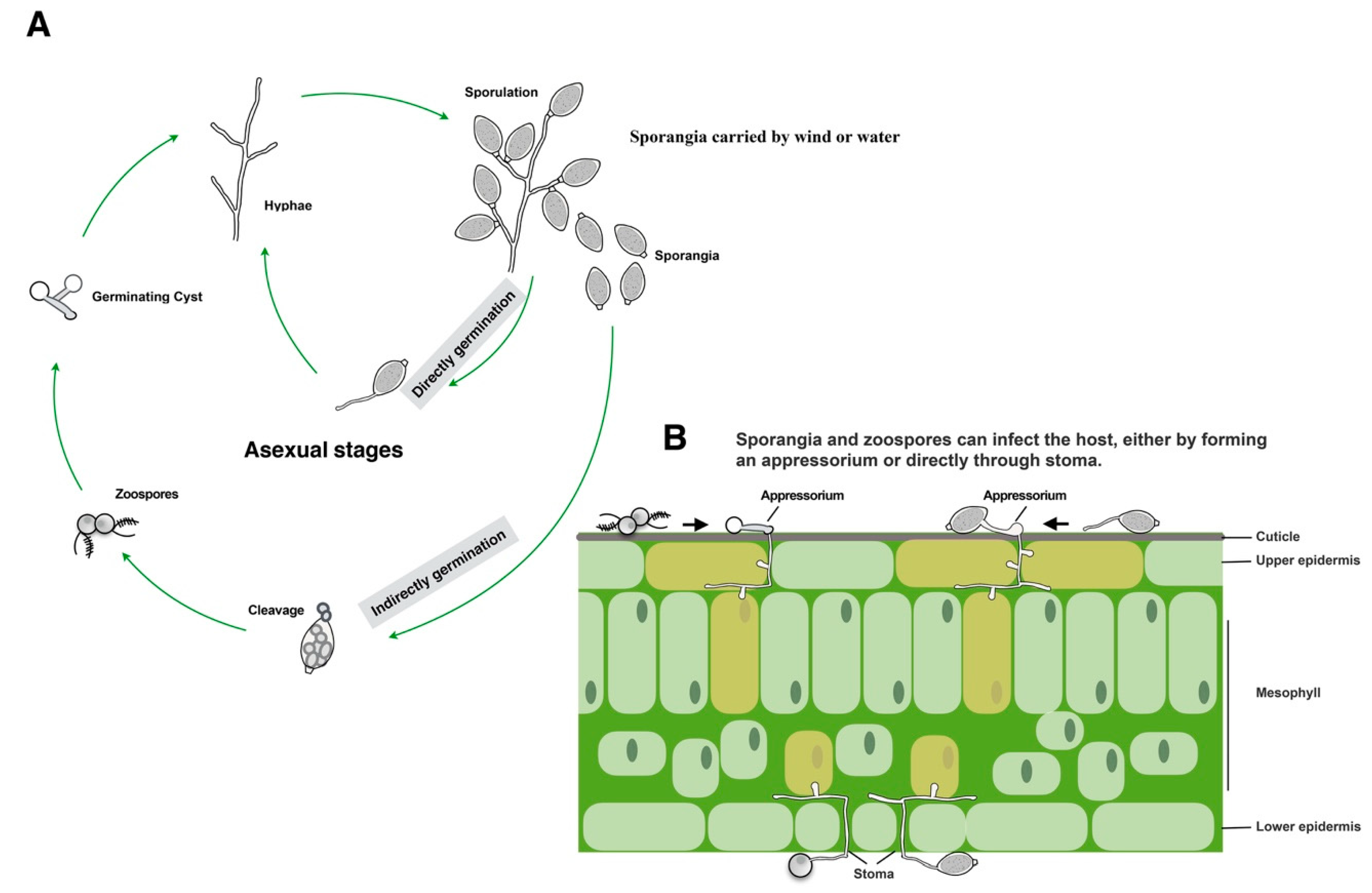
3. Oomycetes
4. Oomycete Pathogens of Fiber Crops
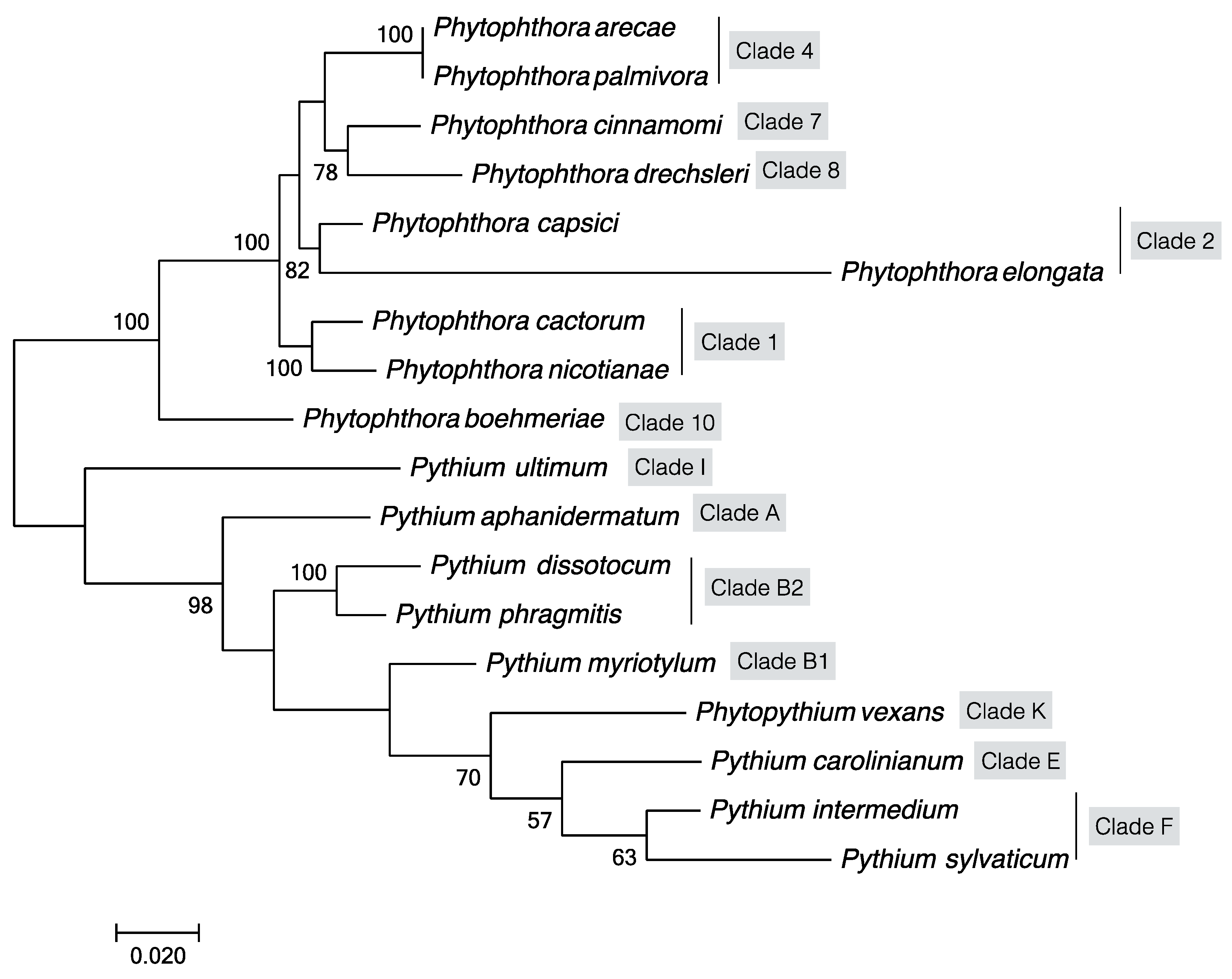
5. Target DNA Selection, Molecular Assays and Phylogeny of Oomycete Pathogens on Fiber Crop
5.1. ITS—Conventional PCR
5.2. Non-ITS Nuclear Genes—Conventional PCR
5.3. Mitochondrial Genes—Conventional PCR
5.4. mtDNA-RT-qPCR Technology
6. Molecular Identification of Oomycete Pathogens in Other Crops
7. Conclusions and Future Prospects
Author Contributions
Funding
Conflicts of Interest
References
- Oerke, E.C. Crop losses to pests. J. Agric. Sci. 2006, 144, 31–43. [Google Scholar] [CrossRef]
- Xu, J. Fungal species concepts in the genomics era. Genome 2020. [Google Scholar] [CrossRef]
- Ray, M.; Ray, A.; Dash, S.; Mishra, A.; Achary, K.G.; Nayak, S.; Singh, S. Fungal disease detection in plants: Traditional assays, novel diagnostic techniques and biosensors. Biosens. Bioelectron. 2017, 87, 708–723. [Google Scholar] [CrossRef]
- Tsui, C.K.; Woodhall, J.; Chen, W.; Levesque, C.A.; Lau, A.; Schoen, C.D.; Baschien, C.; Najafzadeh, M.J.; de Hoog, G.S. Molecular techniques for pathogen identification and fungus detection in the environment. IMA Fungus 2011, 2, 177–189. [Google Scholar] [CrossRef]
- Tör, M.; Woods-Tör, A. Fungal and oomycete diseases. In Encyclopedia of Applied Plant Sciences, 2nd ed.; Thomas, B., Murray, B.G., Murphy, D.J., Eds.; Academic Press: Cambridge, MA, USA, 2017; Volume 3, pp. 77–82. [Google Scholar]
- Kumar, P.; Akhtar, J.; Kandan, A.; Kumar, S.; Batra, R.; Dubey, S.C. Advance detection techniques of phytopathogenic fungi: Curent trends and future perspectives. In Current Trends in Plant Disease Diagnostics and Management Practices; 2016; pp. 265–298. [Google Scholar]
- Schroeder, K.L.; Martin, F.N.; de Cock, A.W.A.M.; Lévesque, C.A.; Paulitz, T.C. Molecular detection and quantification of Pythium species: Evolving taxonomy, new tools, and challenges. Plant Dis. 2013, 97, 4–20. [Google Scholar] [CrossRef]
- Ramadan, A.A.; Kenta, S. Technical review of molecular markers and next-generation sequencing technology to manage plant pathogenic oomycetes. Afr. J. Biotechnol. 2018, 17, 369–379. [Google Scholar] [CrossRef]
- Cheng, Y.; Tang, X.; Gao, C.; Li, Z.; Chen, J.; Guo, L.; Wang, T.; Xu, J. Molecular diagnostics and pathogenesis of fungal pathogens on bast fiber crops. Pathogens 2020, 9, 223. [Google Scholar] [CrossRef]
- Gowda, B. Fibres, Rubber, Firewood, Timber and Bamboo; Univerity of Agricultural Sciences: Bangalore, India, 2007. [Google Scholar]
- Dempsey, J.M. Fiber Crops; Rose Printing Company: Tallahassee, FL, USA, 1975; pp. 203–233. [Google Scholar]
- Kipriotis, E.; Heping, X.; Vafeiadakis, T.; Kiprioti, M.; Alexopoulou, E. Ramie and kenaf as feed crops. Ind. Crop. Prod. 2015, 68, 126–130. [Google Scholar] [CrossRef]
- Tan, L.T.; Yu, C.M.; Chen, P.; Wang, Y.Z.; Chen, J.K.; Wen, L.; Xiong, H.P. Research status and prospective development of bast fiber crops for multi-purpose (In Chinese). Plant Fiber Sci. China 2012, 34, 94–99. [Google Scholar]
- Chen, B.F.; Chen, J.H.; Mu, B.; Zeng, M.; Zhang, H.; Yu, J.; Zhao, J.; Luan, M.B. Advances in medicinal health protection studies of Boehmeria Jacq. spp. (In Chinese). Plant Fiber Sci. China 2016, 38, 237–241. [Google Scholar]
- Li, Y.L.; Cui, Z.G.; Tang, C.X.; Zhang, Z.H.; Gou, Y. Innovative utilization value of ramie fiber mulch (In Chinese). Sichuan Agric. Sci. Technol. 2018, 5, 21–22. [Google Scholar]
- Pittaway, P.; Chakrabarty, S.; Yusaf, T.; Chen, G.; Seneweera, S.; Al-Lwayzy, S.; Bennett, J.; Hopf, J. Bioenergy from cotton industry wastes: A review and potential. Renew. Sustain. Energ. Rev. 2016, 66, 435–448. [Google Scholar]
- Sandler, L.N.; Gibson, K.A.; Borger, C. A call for weed research in industrial hemp (Cannabis sativa L). Weed Res. 2019, 59, 255–259. [Google Scholar] [CrossRef]
- Paridah, M.D.T.A.; Ahmed, B.; Azry, S.O.A.; Ahmed, Z. Review of bast fiber retting. Biol. Resour. 2011, 6, 5260–5281. [Google Scholar]
- Cheng, Z. Ramie industry: Some development opportunities appear from structural adjustment and industrial upgrading (In Chinese). China Fiber Insp. 2018, 10, 113–116. [Google Scholar]
- Baldauf, S.L.; Roger, A.J.; Wenk-Siefert, I.; Doolittle, W.F. A kingdom-level phylogeny of eukaryotes based on combined protein data. Science 2000, 290, 972–977. [Google Scholar] [CrossRef]
- Yoon, H.S.; Hackett, J.D.; Bhattacharya, D. A single origin of the peridinin- and fucoxanthin- containing plastids in dinoflagellates through tertiary endosymbiosis. PNAS 2002, 99, 11724–11729. [Google Scholar] [CrossRef]
- Bhadauria, V.; Banniza, S.; Wei, Y.; Peng, Y.L. Reverse genetics for functional genomics of phytopathogenic fungi and oomycetes. Comp. Funct. Genom. 2009, 2009, 380719. [Google Scholar] [CrossRef]
- Sparrow, F.K. Aquatic Phycomycetes; University of Michigan Press: Ann Arbor, MI, USA, 1960. [Google Scholar]
- Karling, J.S. Predominantly Holocarpic and Eucarpic Simple Biflagellate Phycomycetes; J. Cramer: Stuttgart, Germany, 1981. [Google Scholar]
- Fawke, S.; Doumane, M.; Schornack, S. Oomycete interactions with plants: Infection strategies and resistance principles. Microbiol. Mol. Biol. Rev. 2015, 79, 263–280. [Google Scholar] [CrossRef]
- Erwin, D.C.; Ribeiro, O.K. Phytophthora Diseases Worldwide; The American Phytopathological Society Press: St. Paul, MN, USA, 1996. [Google Scholar]
- Hardham, A.R. The cell biology behind Phytophthora pathogenicity. Australas. Plant Pathol. 2001, 30, 91–98. [Google Scholar] [CrossRef]
- Nowicki, M.; Lichocka, M.; Nowakowska, M.; Kłosińska, U.; Kozik, E.U. A simple dual stain for detailed investigations of plant-fungal pathogen interactions. Veg. Crops Res. Bull. 2012, 77, 61–74. [Google Scholar] [CrossRef][Green Version]
- Lacy, M.L.; Hammerschmidt, R. Diseases of Potato: Late Blight; Michigan State University, Cooperative Extension Service: East Lansing, MI, USA, 1984. [Google Scholar]
- Blair, J.E.; Coffey, M.D.; Park, S.Y.; Geiser, D.M.; Kang, S. A multi-locus phylogeny for Phytophthora utilizing markers derived from complete genome sequences. Fungal Genet. Biol. 2008, 45, 266–277. [Google Scholar] [CrossRef] [PubMed]
- Shen, G.; Wang, Y.C.; Zhang, W.L.; Zheng, X.B. Development of a PCR assay for the molecular detection of Phytophthora boehmeriae in infected cotton. J. Phytopathol. 2005, 153, 291–296. [Google Scholar] [CrossRef]
- dos Santos, Á.F. Phytophthora boehmeriae. For. Phytophthoras 2016, 6. [Google Scholar] [CrossRef]
- Yu, Y.T.; Chen, J.; Gao, C.S.; Zeng, L.B.; Li, Z.M.; Zhu, T.T.; Sun, K.; Cheng, Y.; Sun, X.P.; Yan, L.; et al. First report of brown root rot caused by Pythium vexans on ramie in Hunan, China. Can. J. Plant Pathol. 2016, 38, 405–410. [Google Scholar] [CrossRef]
- Zhang, Y.M.; Zhao, Y.L.; Zhou, W.Z. Research progress of zebra leaf disease on sisal (In Chinese). Chin. J. Trop. Crops 2016, 37, 1627–1633. [Google Scholar]
- He, H.; Zheng, X.B.; Cao, Y.Q.; Lu, J.Y. Identification of Phytophthora species from Boehmeria nivea (In Chinese). China's Fiber Crops 1993, 2, 38–40. [Google Scholar]
- He, H. Studies on Biology of Phytophthora Boehmeriae and Disease Cycle of Cotton Phytophthora Blight. Master’s Dissertation, Nanjing Agricultural University, Nanjing, China, 1992. 51p. [Google Scholar]
- Kousik, C.S.; Parada, C.; Quesada-Ocampo, L. First report of Phytophthora fruit rot on bitter gourd (Mormodica charantia) and sponge gourd (Luffa cylindrica) caused by Phytophthora capsici. Plant Health Prog. 2015, 16, 93–94. [Google Scholar] [CrossRef]
- Kunadiya, M.; Dunstan, W.; White, D.; Hardy, G.E.S.J.; Grigg, A.; Burgess, T. A qPCR assay for the detection of Phytophthora cinnamomi including an mRNA protocol designed to establish propagule viability in environmental samples. Plant Dis. 2019, 103, 2443–2450. [Google Scholar] [CrossRef]
- Dell, B.; Malajczuk, N. Jarrah dieback—A disease caused by Phytophthora cinnamomi. In the Jarrah Forest; Springer: Dordrecht, The Netherlands, 1989; pp. 67–87. [Google Scholar]
- Shearer, B.L.; Tippett, J.T. Jarrah Dieback: The Dynamics and Management of Phytophthora cinnamomi in the Jarrah (Eucalyptus marginata) Forest of South-Western Australia; Department of Conservation and Land Management: Perth, Australia, 1989; Volume 3. [Google Scholar]
- Rea, A.J.; Jung, T.; Burgess, T.I.; Stukely, M.J.C.; Hardy, G.E.S.J. Phytophthora elongate sp. nov., a novel pathogen from the Eucalyptus marginata forest of Western Australia. Australas. Plant Pathpol. 2010, 39, 477–491. [Google Scholar] [CrossRef]
- Cacciola, S.O.; Pane, A.; Faedda, R.; Rizza, C.; Badala, F.; di San Lio, G.M. Bud and root rot of windmill palm (Trachycarpus fortunei) caused by simultaneous infections of Phytophthora palmivora and P. nicotianae in Sicily. Plant Dis. 2011, 95, 769. [Google Scholar] [CrossRef] [PubMed]
- Abdelzaher, H.M.; Imam, M.M.; Shoulkamy, M.A.; Gherbawy, Y.M.A. Biological control of Pythium damping-off of bush okra using rhizosphere strains of Pseudomonas fluorescens. Mycobiology 2004, 32, 139–147. [Google Scholar] [CrossRef]
- Punja, Z.K.; Scott, C.; Chen, S. Root and crown rot pathogens causing wilt symptoms on field-grown marijuana (Cannabis sativa L.) plants. Can. J. Plant Pathol. 2018, 40, 528–541. [Google Scholar] [CrossRef]
- Beckerman, J.; Nisonson, H.; Albright, N.; Creswell, T. First report of Pythium aphanidermatum crown and root rot of industrial hemp in the United States. Plant Dis. 2017, 101, 1038–1039. [Google Scholar] [CrossRef]
- Hashem, M.; Ali, E. Epicoccum nigrum as biocontrol agent of Pythium damping-off and root-rot of cotton seedlings. Arch. Phytopathol. Plant Prot. 2004, 37, 283–297. [Google Scholar] [CrossRef]
- Abdelzaher, H.M.; Elnaghy, M.A. Identification of Pythium carolinianum causing ‘root rot’ of cotton in Egypt and its possible biological control by Pseudomonas fluorescens. Mycopathologia 1998, 142, 143–151. [Google Scholar] [CrossRef]
- Punja, Z.K.; Rodriguez, G. Fusarium and Pythium species infecting roots of hydroponically grown marijuana (Cannabis sativa L.) plants. Can. J. Plant Pathol 2018, 40, 498–513. [Google Scholar] [CrossRef]
- Brown, A.E.; Mercer, P.C. Root-Rot Pathogens of Flax and Microbial Retting. Annual Report on Research and Technical Work of the Department of Agriculture for Northern Ireland. Great Britain. Department of Agriculture for Northern Ireland. 1984, 133–134. [Google Scholar]
- McGehee, C.S.; Apicella, P.; Raudales, R.; Berkowitz, G.; Ma, Y.; Durocher, S.; Lubell, J. First report of root rot and wilt caused by Pythium myriotylum on hemp (Cannabis sativa) in the United States. Plant Dis. 2019, 103, 3288. [Google Scholar] [CrossRef]
- Nechwatal, J.; Wielgoss, A.; Mendgen, K. Pythium phragmitis sp. nov., a new species close to P. arrhenomanes as a pathogen of common reed (Phragmites australis). Mycol. Res. 2005, 109, 1337–1346. [Google Scholar] [CrossRef]
- Ahonsi, M.O.; Agindotan, B.O.; Gray, M.E.; Bradley, C.A. First report of basal stem rot and foliar blight caused by Pythium sylvaticum on Miscanthus sinensis in Illinois. Plant Dis. 2011, 95, 616. [Google Scholar] [CrossRef] [PubMed]
- Gaur, R.; Pandey, L.; Dubey, R.C. Linum usitatissimum, a new host of Pythium ultimum from India. Opt. Lett. 1990, 23, 1829–1831. [Google Scholar]
- Beckerman, J.; Stone, J.; Ruhl, G.; Creswell, T. First report of Pythium ultimum crown and root rot of industrial hemp in the United States. Plant Dis. 2018, 102, 2045. [Google Scholar] [CrossRef] [PubMed]
- Arndt, C.H. Pythium ultimum and the damping-off of cotton seedlings. Phytopathology 1943, 33, 607–611. [Google Scholar]
- Xu, Y.P.; Yang, M.; Guo, H.Y.; Hu, X.L.; Chen, Y.; Yin, P.X.; Li, X.Y.; Li, H.J. Industrial cannabis diseases and insect pests and their control techniques in Kunming (In Chinese). Yunnan Nongye Keji 2004, 4, 46–48. [Google Scholar]
- Kamoun, S.; Furzer, O.; Jones, J.D.; Judelson, H.S.; Ali, G.S.; Dalio, R.J.; Roy, S.G.; Schena, L.; Zambounis, A.; Panabieres, F.; et al. The Top 10 oomycete pathogens in molecular plant pathology. Mol. Plant Pathol. 2015, 16, 413–434. [Google Scholar] [CrossRef]
- Hardham, A.R.; Blackman, L.M. Phytophthora cinnamomi. Mol. Plant Pathol. 2018, 19, 260–285. [Google Scholar] [CrossRef] [PubMed]
- Hausbeck, M.K.; Lamour, K.H. Phytophthora capsici on vegetable crops: Research progress and management challenges. Plant Dis. 2004, 88, 1292–1303. [Google Scholar] [CrossRef]
- Tahi, G.M.; Kébé, B.I.; Sangare, A.; Mondeil, F.; Cilas, C.; Eskes, A.B. Foliar resistance of cacao (Theobroma cacao) to Phytophthora palmivora as an indicator of pod resistance in the field: Interaction of cacao genotype, leaf age and duration of incubation. Plant Pathol. 2006, 55, 776–782. [Google Scholar] [CrossRef]
- Farr, D.F.; Rossman, A.Y. Fungal Databases, Systematic Mycology and Microbiology Laboratory, ARS, USDA. Available online: http://nt.ars-grin.gov/fungaldatabases/ (accessed on 6 October 2014).
- Jeffers, S.; Martin, S.B. Comparison of two selective media for Phytophthora and Pythium species. Plant Dis. 1986, 70, 1038–1043. [Google Scholar] [CrossRef]
- Yin, L.; Tian, X.; Huo, W. Overview of identification techniques for Pythium species based on morphology and molecular biology (In Chinese). China Veg. 2013, 12, 15–22. [Google Scholar]
- Ward, L.; Immanuel, T.M.; Khan, S.; Liefting, L.W.; Delmiglio, C. Conventional PCR. In Molecular Methods in Plant Disease Diagnostics; Boonham, N., Tomlinson, J., Mumford, R., Eds.; CABI: Wallingford, Oxfordshire, UK; Boston, MA, USA, 2016. [Google Scholar]
- Villa-Carvajal, M.; Querol, A.; Belloch, C. Identification of species in the genus Pichia by restriction of the internal transcribed spacers (ITS1 and ITS2) and the 5.8S ribosomal DNA gene. Anton Leeuw 2006, 90, 171–181. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, C.; Kageyama, K.; Suga, H.; Hyakumachi, M. Phylogenetic relationships of Pythium species based on ITS and 5.8S sequences of the ribosomal DNA. Mycoscience 1999, 40, 321–331. [Google Scholar] [CrossRef]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J., White, T.J., Eds.; Academic Press: London, UK, 1990; pp. 315–322. [Google Scholar]
- Maizatul-Suriza, M.; Dickinson, M.; Idris, A.S. Molecular characterization of Phytophthora palmivora responsible for bud rot disease of oil palm in Colombia. World, J. Microbiol. Biotechnol. 2019, 35, 44. [Google Scholar] [CrossRef]
- Xu, J. Fungal DNA barcoding. Genome 2016, 59, 913–932. [Google Scholar] [CrossRef]
- Hughes, K.W.; Petersen, R.H.; Lickey, E.B. Using heterozygosity to estimate a percentage DNA sequence similarity for environmental species’ delimitation across basidiomycete fungi. New Phytol. 2009, 182, 795–798. [Google Scholar] [CrossRef]
- Lara, E.; Belbahri, L. SSU rRNA reveals major trends in oomycete evolution. Fungal Divers. 2011, 49, 93–100. [Google Scholar] [CrossRef]
- Hebert, P.D.N.; Cywinska, A.; Ball, S.L.; DeWaard, J.R. Biological identifications through DNA barcodes. Proc. R. Soc. B Biol. Sci. 2003, 270, 313–321. [Google Scholar] [CrossRef]
- Sandor, S.; Zhang, Y.J.; Xu, J. Fungal mitochondrial genomes and genetic polymorphisms. Appl. Microbiol. Biotech. 2018, 102, 9433–9448. [Google Scholar] [CrossRef]
- Bullerwell, C.E.; Lang, B.F. Fungal evolution: The case of the vanishing mitochondrion. Curr. Opin. Microbiol. 2005, 8, 362–369. [Google Scholar] [CrossRef]
- Tooley, P.W.; Martin, F.N.; Carras, M.M.; Frederick, R.D. Real-time fluorescent polymerase chain reaction detection of Phytophthora ramorum and Phytophthora pseudosyringae using mitochondrial gene regions. Phytopathology 2006, 96, 336–345. [Google Scholar] [CrossRef] [PubMed]
- Robideau, G.P.; De Cock, A.W.A.M.; Co_ey, M.D.; Voglmayr, H.; Brouwer, H.; Bala, K.; Chitty, D.W.; Désaulniers, N.; Eggertson, Q.A.; Gachon, C.M.M.; et al. DNA barcoding of oomycetes with cytochrome c oxidase subunit I and internal transcribed spacer. Mol. Ecol. Resour. 2011, 11, 1002–1011. [Google Scholar] [CrossRef] [PubMed]
- Kulik, T.; Bilska, K.; Zelechowski, M. Promising perspectives for detection, identification, and quantification of plant pathogenic fungi and oomycetes through targeting mitochondrial DNA. Int. J. Mol. Sci. 2020, 21, 2645. [Google Scholar] [CrossRef] [PubMed]
- Boehm, J.; Hahn, A.; Schubert, R.; Bahnweg, G.; Adler, N.; Nechwatal, J.; Oehlmann, R.; Obszwald, W. Real–time quantitative PCR: DNA determination in isolated spores of the mycorrhizal fungus glomus mosseae and monitoring of Phytophthora infestans and Phytophthora citricola in their respective host plants. J. Phytopathol. 1999, 147, 409–416. [Google Scholar] [CrossRef]
- Schaad, N.W.; Frederick, R.D. Real-time PCR and its application for rapid plant disease diagnostics. Can. J. Plant Pathol. 2002, 24, 250–258. [Google Scholar] [CrossRef]
- Martin, F.N. Phylogenetic relationships among some Pythium species inferred from sequence analysis of the mitochondrially encoded cytochrome oxidase II gene. Mycologia 2000, 92, 711–727. [Google Scholar] [CrossRef]
- Briard, M.; Dutertre, M.; Rouxel, F.; Brygoo, Y. Ribosomal RNA sequence divergence within the Pythiaceae. Mycol. Res. 1995, 99, 1119–1127. [Google Scholar] [CrossRef]
- Cooke, D.E.L.; Drenth, A.; Duncan, J.M.; Wagels, G.; Brasier, C.M. A molecular phylogeny of Phytophthora and related oomycetes. Fungal Genet. Biol. 2000, 30, 17–32. [Google Scholar] [CrossRef]
- Lévesque, C.A.; de Cock, A.W.A.M. Molecular phylogeny and taxonomy of the genus Pythium. Mycol. Res. 2004, 108, 1363–1383. [Google Scholar] [CrossRef]
- Bonants, P.; Weerdt, M.H.-d.; Gent-Pelzer, M.v.; Lacourt, I.; Cooke, D.; Duncan, J. Detection and identification of Phytophthora fragariae Hickman by the polymerase chain reaction. Eur. J. Plant Pathol. 1997, 103, 345–355. [Google Scholar] [CrossRef]
- Spies, C.F.J.; Mazzola, M.; McLeod, A. Characterisation and detection of Pythium and Phytophthora species associated with grapevines in South Africa. Eur. J. Plant Pathol. 2011, 131, 103–119. [Google Scholar] [CrossRef]
- Schena, L.; Hughes, K.J.; Cooke, D.E. Detection and quantification of Phytophthora ramorum, P. kernoviae, P. citricola and P. quercina in symptomatic leaves by multiplex real-time PCR. Mol Plant Pathol. 2006, 7, 365–379. [Google Scholar] [CrossRef] [PubMed]
- Engelbrecht, J.; Duong, T.A.; van den Berg, N. Development of a nested quantitative real-time PCR for detecting Phytophthora cinnamomi in Persea americana rootstocks. Plant. Dis. 2013, 97, 1012–1017. [Google Scholar] [CrossRef]
- Bent, E.; Loffredo, A.; Yang, J.I.; McKenry, M.V.; Becker, J.O.; Borneman, J. Investigations into peach seedling stunting caused by a replant soil. FEMS Microbiol. Ecol. 2009, 68, 192–200. [Google Scholar] [CrossRef] [PubMed]
- Li, M.M.; Zhao, W.; Wang, T.; Gu, Y.; Gao, Z.M.; Qi, R.D. Rapid molecular detection of Phytophthora capsici based on its Ypt1 gene (In Chinese). Acta Phytopathol. Sin. 2014, 44, 546–551. [Google Scholar]
- Yuan, X.; Feng, C.; Zhang, Z.; Zhang, C. Complete mitochondrial genome of Phytophthora nicotianae and identification of molecular markers for the oomycetes. Front. Microbiol. 2017, 8, 1484. [Google Scholar] [CrossRef]
- Dong, Z.; Liu, P.; Li, B.; Chen, G.; Chen, Q. Loop-mediated isothermal amplification assay for sensitive and rapid detection of Phytophthora capsici. Can. J. Plant Pathol. 2015, 37, 485–494. [Google Scholar] [CrossRef]
- Shen, D.; Li, Q.; Yu, J.; Zhao, Y.; Zhu, Y.; Xu, H.; Dou, D. Development of a loop-mediated isothermal amplification method for the rapid detection of Pythium ultimum. Australas. Plant Path 2017, 46, 571–576. [Google Scholar] [CrossRef]
- Cheng, Y.C.; Kang, H.J.; Shi, Y.X.; Chai, A.L.; Zhang, H.J.; Xie, X.W.; Li, B.J. Development and application of real-time fluorescent quantitative PCR for detection of Phytophthora capsici (In Chinese). Acta Hortic. Sin. 2018, 45, 997–1006. [Google Scholar]
- Ahonsi, M.O.; Ling, Y.; Kageyama, K. Development of SCAR markers and PCR assays for single or simultaneous species-specific detection of Phytophthora nicotianae and Pythium helicoides in ebb-and-flow irrigated kalanchoe. J. Microbiol. Methods 2010, 83, 260–265. [Google Scholar] [CrossRef]
- Kernaghan, G.; Reeleder, R.D.; Hoke, S.M.T. Quantification of Pythium populations in ginseng soils by culture dependent and real-time PCR methods. Appl. Soil Ecol. 2008, 40, 447–455. [Google Scholar] [CrossRef]
- Cullen, D.W.; Toth, I.K.; Boonham, N.; Walsh, K.; Barker, I.; Lees, A.K. Development and validation of conventional and quantitative polymerase chain reaction assays for the detection of storage rot potato pathogens, Phytophthora erythroseptica, Pythium ultimum and Phoma foveata. J. Phytopathol. 2007, 155, 309–315. [Google Scholar] [CrossRef]
- Ristaino, J.B.; Saville, A.C.; Paul, R.; Cooper, D.; Wei, Q. Detection of Phytophthora infestans by LAMP, real-time LAMP and droplet digital PCR. Plant Dis. 2019, 104, 708–716. [Google Scholar] [CrossRef]
- Notomi, T.; Okayama, H.; Masubuchi, H.; Yonekawa, T.; Watanabe, K.; Amino, N.; Hase, T. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 2000, 28, e63. [Google Scholar] [CrossRef]
- Ramirez-Castillo, F.Y.; Loera-Muro, A.; Jacques, M.; Garneau, P.; Avelar-Gonzalez, F.J.; Harel, J.; Guerrero-Barrera, A.L. Waterborne pathogens: Detection methods and challenges. Pathogens 2015, 4, 307–334. [Google Scholar] [CrossRef]
- Paul, R.; Saville, A.C.; Hansel, J.C.; Ye, Y.; Ball, C.; Williams, A.; Chang, X.; Chen, G.; Gu, Z.; Ristaino, J.B.; et al. Extraction of plant DNA by microneedle patch for rapid detection of plant diseases. ACS Nano 2019, 13, 6540–6549. [Google Scholar] [CrossRef]
- Li, Z.; Paul, R.; Ba Tis, T.; Saville, A.C.; Hansel, J.C.; Yu, T.; Ristaino, J.B.; Wei, Q. Non-invasive plant disease diagnostics enabled by smartphone-based fingerprinting of leaf volatiles. Nat. Plants 2019, 5, 856–866. [Google Scholar] [CrossRef] [PubMed]
- Grunwald, N.J.; McDonald, B.M.; Milgroom, M.G. Population genomics of fungal and oomycete pathogens. Annu. Rev. Phytopathol. 2016, 54, 323–346. [Google Scholar] [CrossRef] [PubMed]
| Group | Crop | Main Distribution | Growth Habitat | Main Applications | Main Oomycete Diseases |
|---|---|---|---|---|---|
| Seed fiber | Cotton (Gossypium hirsutum) | China, USA, India, Brazil, Mexico | Thermophilic plant, sandy loam, loam and light clay with better heat transfer and permeability | Textiles, cottonseed oil | Cotton blight |
| Sponge gourd (Luffa cylindrica) | China, Japan, Korea, India (Kerala, Andhra Pradesh) | Requires 150 to 200 warm days to mature | Used as a bath or kitchen sponge and food | Phytophthora fruit rot | |
| Bast fiber | Hemp (Cannabis sativa) | China, Canada, USA, Europe, East Asia, Nepal | Grows at 16–27 °C, sufficient rain at the first six weeks of growth, short day length. | Textiles, hempseed oil, prescription drug | Hemp blight, hemp root and crown rot wilt |
| Ramie (Boehmeria nivea) | China, Brazil, Philippines, India, Vietnam, Laos, Cambodia | Sandy soil and warm, wet climates, rainfall averaging at least 75 to 130 mm per month | Textiles, soil and water conservation, medicine | Ramie blight, ramie brown root rot | |
| Flax (Linum usitatissimum) | France, Russia, Netherlands, Belarus, Belgium, Canada, Kazakhstan, China, India | Well-drained loam and cool, moist temperate climates | Linen, flax yarn, flax seed, linseed oil | Flax root rot | |
| Leaf fiber | Sisal (Agava sisalana) | Brazil, Tanzania, Kenya, Madagascar, China, Mexico, Haiti, Venezuela, Morocco, South Africa | In the tropical and temperate zones with mean temperature at 25 °C with sufficient sunshine | Making rope, twine, paper, cloth, wall covering and dartboards | Sisal zebra leaf disease |
| Grass fiber | Silvergrass (Miscanthus sinensis) | China, Japan, Korea, USA | In temperate regions around the world | Ornamental plant, bioenergy production | Basal stem rot and foliar blight |
| Reed (Phragmites australis) | Northern Hemisphere | In lakes and rivershores, marshes, coastal brackish swamps, and lagoons | Used in phytoremediation, protecting shoreline from bank erosion, and serving as a food source or habitat protection for arthropods, birds and mammals. | Dieback of reed stands | |
| Palm fiber | Windmill Palm (Trachycarpus fortunei) | China, Japan, India, Burma | Warm and humid climate | Making rope, coir raincoat, brown bandage, carpet, brush and filling material for sofa, medicine, ornament | Windmill Palm bud and root rot |
| Woody fiber | Jarrah (Eucalyptus marginata) | Australia | Rainfall isohyet exceeds 600 mm, grows in soils derived from ironstone | Structural material for bridges, wharves, railway sleepers, ship building and telegraph poles, medicine | Jarrah dieback |
| Pathogens | Disease | Method | Marker | Host Plant | Geographic Region(s) | Reference |
|---|---|---|---|---|---|---|
| Phytophthora spp. | ||||||
| P. arecae | Sisal zebra spot disease | Conventional PCR | ITS | Agava sisalana | China, India | [34] |
| P. boehmeriae (Dominant pathogen) | Cotton blight | Conventional PCR | ITS | Gossypium hirsutum | China | [31] |
| P. boehmeriae | Ramie blight | Conventional PCR | Cox 2, Nad 9, Rps 10, Sec Y | Boehmeria nivea | China (Taiwan), Australia, Greece, South Africa | [32] |
| P. cactorum | Ramie blight | Morphological | Boehmeria nivea | Jiangxi, China | [35,36] | |
| P. cactorum | Cotton blight | Conventional PCR | ITS | Gossypium hirsutum | China | [31] |
| P. capsici | Sponge gourd rot | Conventional PCR | Cox 1, Cox 2, Nad 1, Nad 5, 𝛽-tubulin, EF1, Enolase, HSP090, Ura3, ITS | Luffa cylindrica | USA | [37] |
| P. cinnamomi | Jarrah dieback | Quantitative real-time PCR | Cox 2 | Eucalyptus marginata | Western AustraliaUSA | [38,39,40] |
| P. drechsleri | Cotton blight | Conventional PCR | ITS | Gossypium hirsutum | China | [31] |
| P. elongata | Jarrah dieback | Conventional PCR | ITS andCox 1 | Eucalyptus marginata | Western Australia | [41] |
| P. nicotianae | Cotton blight | Conventional PCR | ITS | Gossypium hirsutum | China | [31] |
| P. nicotianae | Windmill palm bud and root rot | Conventional PCR | ITS | Trachycarpus fortunei | eastern Sicily, Italy | [42] |
| P. nicotianae (Dominant pathogen) | Sisal zebra spot disease | Conventional PCR | ITS | Agava sisalana | China, India | [34] |
| P. palmivora | Cotton blight | Conventional PCR | ITS | Gossypium hirsutum | China | [31] |
| P. palmivora | Sisal zebra spot disease | Conventional PCR | ITS | Agava sisalana | China, India | [34] |
| P. palmivora | Windmill palm bud and root rot | Conventional PCR | ITS | Trachycarpus fortunei | eastern Sicily, Italy | [42] |
| Pythium spp. | ||||||
| P. aphanidermatum | Bush okra damping-off | Morphological | Corchorus olitorius | Egypt | [43] | |
| P. aphanidermatum | Hemp root rot and crown wilt | Conventional PCR | ITS | Cannabis sativa | California, USA | [44] |
| P. aphanidermatum | Hemp crown and root Rot | Conventional PCR | ITS | Cannabis sativa | Indiana, USA | [45] |
| P. baryanum | Cotton damping-off | Morphological | Gossypium hirsutum | Egypt | [46] | |
| P. carolinianum | Cotton root rot | Morphological | Gossypium hirsutum | Egypt | [47] | |
| P. dissotocum | Marijuana root rot | Conventional PCR | ITS, EF-1α | Cannabis sativa | Canada | [48] |
| P. intermedium | Flax root rot | Taxonomic | Linum usitatissimum | UK | [49] | |
| P. myriotylum | Marijuana root rot | Conventional PCR | ITS, EF-1α | Cannabis sativa | Canada | [48] |
| P. myriotylum | Hemp root rot and Wilt | Conventional PCR | ITS, Cox 1, Cox 2 | Cannabis sativa | Connecticut, USA | [50] |
| P. phragmitis | Reed die-back syndrome | Conventional PCR | ITS andCox 2 | Phragmites australis | Lake Constance, Germany | [51] |
| P. sylvaticum | Silvergrass stem rot and blight | Conventional PCR | ITS, Cox 2 | Miscanthus sinensis | Illinois, USA | [52] |
| P. ultimum | Flax damping off | Morphological | Linum usitatissimum | India | [53] | |
| P. ultimum | Hemp crown and root rot | Conventional PCR | ITS | Cannabis sativa | Indiana, USA | [54] |
| P. ultimum | Cotton damping-off | Morphological | Gossypium hirsutum L. | Egypt | [55] | |
| P. vexans | Ramie brown root rot | Conventional PCR | ITS, 18S, 28S | Boehmeria nivea | China | [33] |
| Pseudoperonospora cannabinus | Hemp mildew | Morphological | Cannabis sativa | Austria, Canada, China, Italy | [56] |
| Target DNA | Primer Name and Sequence (5′-3′) | Pathogens | TM (°C) | Product (bp) | Reference | |
|---|---|---|---|---|---|---|
| ITS1-5.8S-ITS2 | DC6 | GAGGGACTTTTGGGTAATCA | Phytophthora spp., Pythium spp. | 62 | 1300 | [31,84] |
| ITS4 | TCCTCCGCTTATTGATATGC | |||||
| PB1 | CGGCTTTCGGGCTGCTGC | P. boehmeriae | 62 | 750 | [31] | |
| PB2 | ATACCCGAAGGCAAAGCGC | |||||
| ITS1-F | CTTGGTCATTTAGAGGAAGTAA | P. aphanidermatum | 60 | 700 | [44,48] | |
| ITS4 | TCCTCCGCTTATTGATATGC | P. dissotocum, P. myriotylum | ||||
| ITS6 | GAAGGTGAAGTCTAACAAGG | P. cinnamomi, P. palmivora, P. elongate | 55 | 796–910 | [42,51] | |
| ITS4 | TCCTCCGCTTATTGATATGC | |||||
| ITS1 | TCCGTAGGTGAACCTGCGG | P. vexans, | 55 | 810–900 | [33,42] | |
| ITS4 | TCCTCCGCTTATTGATATGC | P. nicotianae | ||||
| rDNA 18S | NS3 | GCAAGTCTGGTGCCAGCAGCC | P. vexans | 50–52 | 610 | [33] |
| NS4 | CTTCCGTCAATTCCTTTAAG | |||||
| rDNA 28S | LR0R | GTACCCGCTGAACTTAAGC | P. vexans | 50–52 | 810 | [33] |
| LR3 | CCGTGTTTCAAGACGGG | |||||
| Cox 1 | FM82 | TTGGCAATTAGGTTTTCAAGATCC | P. elongate | 56 | 742 | [41] |
| FM83 | CTCCAATAAAAAATAACCAAAAATG | |||||
| Cox 2 (RT-qPCR) | PCIN147F | CCAGCAACTGTTGTGCATGG | P. cinnamomi | 55–60 | 100 | [38] |
| PCIN249R | AATATAATAAAGCAAATGATGGT | |||||
| PCIN146F | TCCAGCAACTGTTGTGCATG | |||||
| PCIN250R | GAATATAATAAAGCAAATGATGGT | |||||
| PCIN147F | CCAGCAACTGTTGTGCATGG | |||||
| PCIN246R | ATAATAAAGCAAATGATGGT | |||||
| PCIN150F | GCAACTGTTGTGCATGGAGC | |||||
| PCIN247R | TATAATAAAGCAAATGATGGT | |||||
| Cox 2 | FM35 | CAGAACCTTGGCAATTAGG | P. phragmitis | - | 563 | [51] |
| FM58 | CCACAAATTTCACTACATTG | |||||
| FM58 | CCACAAATTTCACTACATTG | P. sylvaticum | 56 | 544 | [52] | |
| FM66 | TAGGATTTCAAGATCCTG | |||||
| EF-1α | EF-1 | ATG GGT AAG GAGGAC AAG AC | P. dissotocum, | 60 | 700 | [48] |
| EF-2 | GGA GGT ACC AGTGAT CAT GTT | P. myriotylum | ||||
| Pathogens | Method | Marker | Primer (5′-3′) | Sample | Hosts | Reference | |
|---|---|---|---|---|---|---|---|
| P. capsici | RT-qPCR | Actin | YM2F | ATTCCTCCTGATAGATAG | Mycelia | [93] | |
| YM2R | CCCTCATCACAGAATGC | ||||||
| P. capsici | Nested PCR | Ypt1 | Ypt1F | ACGGAGAGCTACATCTCGAC | Mycelia | [88] | |
| Ypt1R | GTCAGATCGCTCTTGTTACC | ||||||
| PcYpt1F | AGACTCTGTTGTATAGCAGAG | ||||||
| PcYpt1R | AACGTCTTGAACTTTGGTTG | ||||||
| P. capsici | LAMP | ITS | F3 | GCTGCGGCGTTTAAAGGA | Leaves | Pepper | [91] |
| B3 | AGTGCACACAAAGTTCCCAA | ||||||
| FIP | ACGCCACAGCAGGAAAAGCATTGAGTGTTCGATTCGCGGTA | ||||||
| BIP | GGCTTGGCTTTTGAATCGGCTTTGGATCGACCCTCGACAG | ||||||
| P. cinnamomi | SYBR green (nested PCR) | LPV | LPV3-fwd | GTGCAGACTGTCGATGTG | Avocado | [86] | |
| LPV3-rev | GAACCACAACAGGCACGT | ||||||
| LPV3N-fwd | GTCACGACCATGTTGTTG | ||||||
| LPV3N-rev | GAGGTGAAGGCTGTTGAG | ||||||
| P. nicotianae | Duplex-PCR | SCAR | MPhnic 2F | TTCGAGAAGTACGTGGCGTTT | Leaves | kalanchoe | [94] |
| MPhnic 2R | TTGCAGCGGAGAGTGAGAACT | ||||||
| MPhnic 3F | ATCTCCCAATCGACCGTGAA | ||||||
| MPhnic 3R | CAAGCACGTGACTCGGTTGA | ||||||
| MPhnic 5F | CTCGATACGGACGCAAAGGT | ||||||
| MPhnic 5R | CATGGCTACAGCTGCTGCAA | ||||||
| P. ultimum | Conventional PCR | ITS | PuF | ATGATGGACTAGCTGATGAA | Soils | American ginseng | [95] |
| PuR | TTCCATTACACTTCATAGAA | ||||||
| Pu1F1 | GACGAAGGTTGGTCTGTTG | Tubers | Potato | [96] | |||
| Pu2R1 | CAGAAAAAGAAAGGCAAGTTTG | ||||||
| P. ultimum | TaqMan | ITS | 92F | TGTTTTCATTTTTGGACACTGGA | Tubers | Potato | [96] |
| 166R | TCCATCATAACTTGCATTACAACAGA | ||||||
| 116T | FAM-CGGGAGTCAGCAGGACGAAGGTTG-VIC | ||||||
| P. ultimum | LAMP | ITS | F3 | CAACTGGAAAAGCAAGCGG | Leaf | Wheat, soybean, cucumber, and tobacco | [92] |
| B3 | CCGAAGAACTGTGTCCGC | ||||||
| FIP | GAGCCAGACGGGCCAGTATCAAGTTACAGTGGCGTTGTCA | ||||||
| BIP | TCTCTGTTGCTCGACTGGAGGGTTCCACCTCCTGTAAGACCT | ||||||
| F-Loop | GCTTGCTCCAGTACGAATGC | ||||||
| P. vexans | TaqMan | ITS | PvF1 | TTTCCGTTTTGTGCTTGATG | [84] | ||
| PvR1 | AGCGAACACACCCAATAAGC | ||||||
| VexP1 | HEX™-CCGTGTCTGCTGGCGGGTC-Iowa Black® FQ | ||||||
| P. vexans | RT-qPCR | SSU | VexansF2 | TATACAACCTTGATCGAC | Root tissue | Peach | [87] |
| VexansR2 | GATGGAAAATTGCAACC |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, T.; Gao, C.; Cheng, Y.; Li, Z.; Chen, J.; Guo, L.; Xu, J. Molecular Diagnostics and Detection of Oomycetes on Fiber Crops. Plants 2020, 9, 769. https://doi.org/10.3390/plants9060769
Wang T, Gao C, Cheng Y, Li Z, Chen J, Guo L, Xu J. Molecular Diagnostics and Detection of Oomycetes on Fiber Crops. Plants. 2020; 9(6):769. https://doi.org/10.3390/plants9060769
Chicago/Turabian StyleWang, Tuhong, Chunsheng Gao, Yi Cheng, Zhimin Li, Jia Chen, Litao Guo, and Jianping Xu. 2020. "Molecular Diagnostics and Detection of Oomycetes on Fiber Crops" Plants 9, no. 6: 769. https://doi.org/10.3390/plants9060769
APA StyleWang, T., Gao, C., Cheng, Y., Li, Z., Chen, J., Guo, L., & Xu, J. (2020). Molecular Diagnostics and Detection of Oomycetes on Fiber Crops. Plants, 9(6), 769. https://doi.org/10.3390/plants9060769







