Molecular Verification of the UK National Collection of Cultivated Liriope and Ophiopogon Plants
Abstract
1. Introduction
2. Results
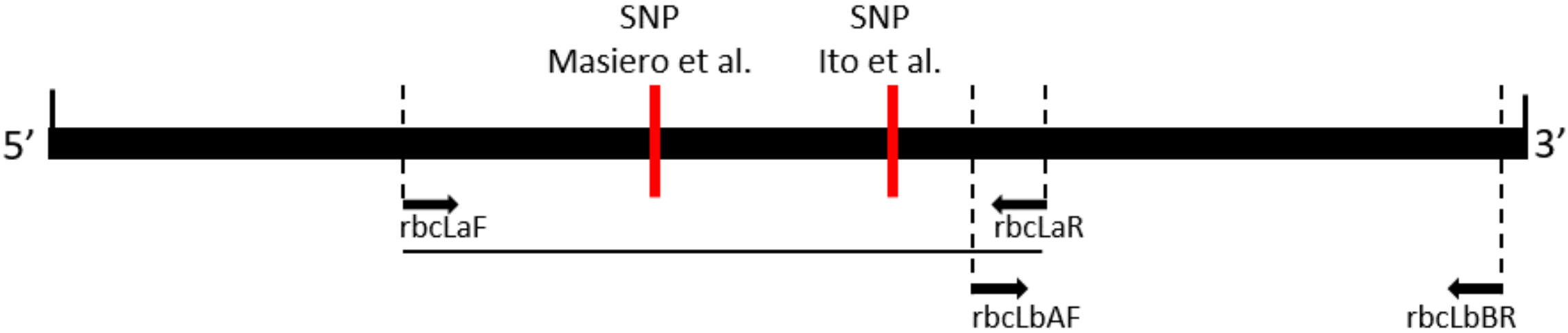
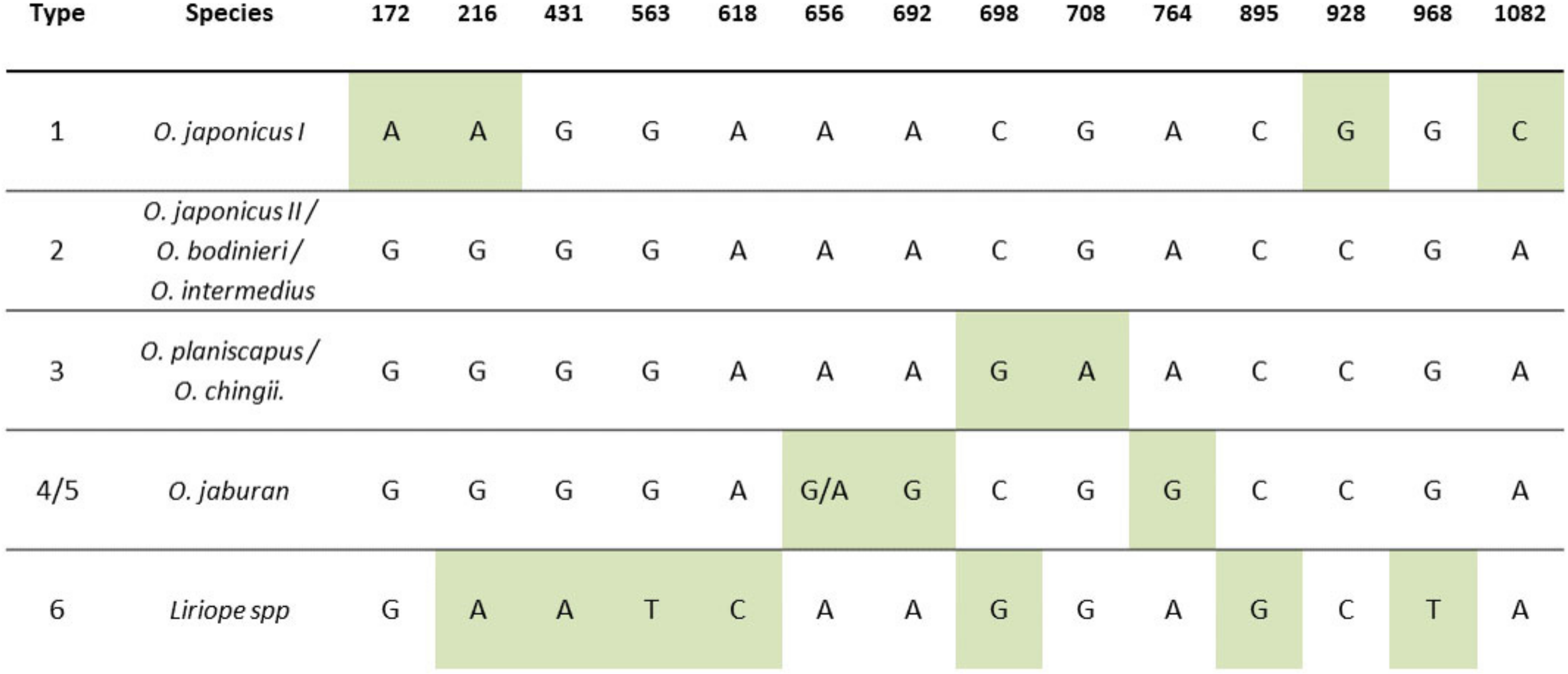
2.1. DNA Barcoding of the rbcLa Region of Liriope and Ophiopogon Accessions
2.2. DNA Barcoding of the rbcLb Region of Liriope and Ophiopogon
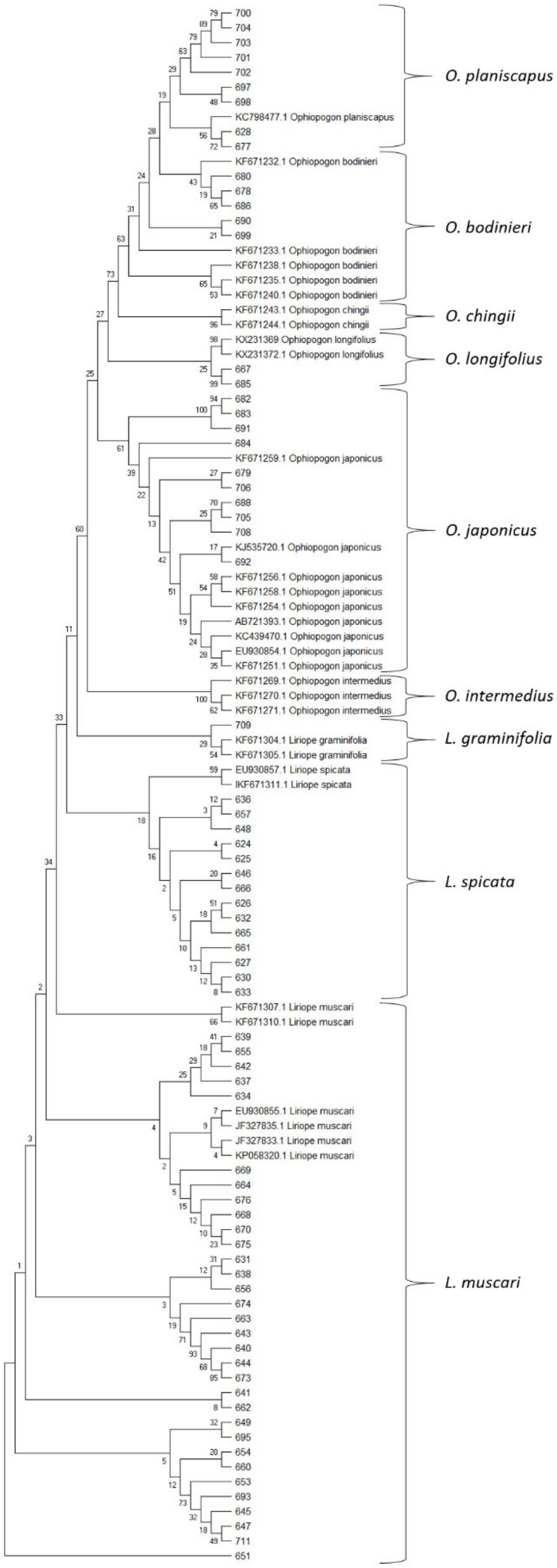
2.3. DNA Barcoding of the nrITS Region of Liriope and Ophiopogon
2.4. Deduction of Individual Taxon Identity
2.4.1. Liriope spp.
2.4.2. O. japonicus
2.4.3. O. bodinieri and O. planiscapus
3. Discussion
4. Materials and Methods
4.1. Plant Collections
4.2. Plant Material and Genomic DNA Extraction
4.3. PCR Protocols
- rbcLa PCR: initial denaturation step of 5 min at 95 °C followed by 35 cycles consisting of 30 s at 95 °C, 20 s at 52 °C, and 50 s at 72 °C, with a final extension period of 5 min at 72 °C.
- rbcLb PCR: initial denaturation step of 5 min at 94 °C followed by 35 cycles consisting of 30 s at 94 °C, 30 s at 56 °C, and 30 s at 72 °C, with a final extension period of 5 min at 72 °C.
- ITS PCR: initial denaturation step of 5 min at 95 °C followed by 30 cycles consisting of 30 s at 95 °C, 30 s at 61 °C, and 30 s at 72 °C, with a final extension period of 5 min at 72 °C.
- ITSP PCR: initial denaturation step of 5 min at 95 °C followed by 30 cycles consisting of 30 s at 95 °C, 30 s at 61 °C, and 30 s at 72 °C, with a final extension period of 5 min at 72 °C.
4.4. DNA Sequence Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| nrITS | nuclear ribosomal internal transcribed spacer |
| rbcL | large subunit of ribulose-1,5-bisphosphate carboxylase/oxygenase gene |
References
- Fantz, P.R. Taxonomic Problems in Cultivated Liriopogons. HortTechnology 1993, 146–150. [Google Scholar] [CrossRef]
- Lattier, J.D.; Ranney, T.G.; Fantz, P.R.; Avent, T. Identification, Nomenclature, Genome Sizes, and Ploidy Levels of Liriope and Ophiopogon Taxa. HortScience 2014, 49, 145–151. [Google Scholar] [CrossRef]
- Nesom, G. Overview of Liriope and Ophiopogon (Ruscaceae) naturalized and commonly cultivated in the USA. Phytoneuron 2010, 56, 1–31. [Google Scholar]
- Wang, G.-Y.; Meng, Y.; Huang, J.-L.; Yang, Y.-P. Molecular Phylogeny of Ophiopogon (Asparagaceae) Inferred from Nuclear and Plastid DNA Sequences. Syst. Bot. 2014, 39, 776–784. [Google Scholar] [CrossRef]
- Fantz, P. Names and species of Ophiopogon cultivated in the southeastern United States. HortTechnology 2009, 19, 385–394. [Google Scholar] [CrossRef]
- Fantz, P.R. Species of Liriope cultivated in the southeastern United States. HortTechnology 2008, 18, 343–348. [Google Scholar] [CrossRef]
- Broussard, M. A Horticultural Study of Liriope and Ophiopogon: Nomenclature, Morphology, and Culture. Ph.D. Thesis, Louisiana State University, Baton Rouge, LA, USA, 2007. Available online: https://digitalcommons.lsu.edu/gradschool_dissertations/4002 (accessed on 6 January 2020).
- Fantz, P.R.; Carey, D.; Avent, T.; Lattier, J. Inventory, Descriptions, and Keys to Segregation and Identification of Liriopogons Cultivated in the Southeastern United States. HortScience 2015, 50, 957–993. [Google Scholar] [CrossRef]
- Mabberley, D. Mabberley’s Plant-Book: A Portable Dictionary of Plants, Their Classifications and Uses; Cambridge University Press: Cambridge, UK, 2008. [Google Scholar]
- Zhou, Q.; Zhou, J.; Chen, J.; Wang, X. Karyotype analysis of medicinal plant Liriope spicata var. prolific (Liliaceae). Biologia 2009, 64, 680–683. [Google Scholar]
- Yang, W.S.; Chen, J.C. Studies and Prospects on Hubei Maidong. Shizhen J. Tradit. Chin. Med. Res. 2006, 10, 1939–1940. [Google Scholar]
- Yamashita, J.; Tamura, M. Molecular phylogeny of the Convallariaceae (Asparagales). In Monocots: Systematics and Evolution; Wilson, K.L., Morrison, D.A., Eds.; CSIRO Publishing: Melbourne, Australia, 2000; pp. 387–400. [Google Scholar]
- Walters, S.M.; Cullen, J. The European Garden Flora: A Manual for the Identification of Plants Cultivated in Europe, Both Out-of-Doors and under Glass; Cambridge University Press: Cambridge, UK, 1984; Volume 6, Part 4. [Google Scholar]
- Cullen, J.; Knees, S.; Cubey, H.; Shaw, J. The European Garden Flora Flowering Plants: A Manual for the Identification of Plants Cultivated in Europe, Both Out-of-Doors and under Glass; Cambridge University Press: Cambridge, UK, 2011. [Google Scholar]
- Thorne, R.F. A phylogenetic classification of the Angiospermae. In Evolutionary Biology; Hecht, M.K., Steere, W.C., Wallace, B., Eds.; Springer: Boston, MA, USA, 1976. [Google Scholar]
- Thorne, R. Synopsis of a putatively phylogenetic classification of the flowering plants. Aliso J. Syst. Evol. Bot. 1968, 6, 57–66. [Google Scholar]
- Takhtajan, A. Flowering Plants; Springer: Dordrecht, The Netherlands, 2009. [Google Scholar] [CrossRef]
- Liang, S.; Dai, K. Pollen morphology and generic phylogenetic relationships in Ophiopogonoideae (Liliaceae). J. Syst. Evol. 1992, 30, 427–437. [Google Scholar]
- Dai, L.; Liang, S. Epidermal features of leaves and their taxonomic significance in subfamily Ophiopogonoideae (Liliaceae). J. Syst. Evol. 1991, 29, 335–346. [Google Scholar]
- Kim, J.-H.; Kim, D.-K.; Forest, F.; Fay, M.F.; Chase, M.W. Molecular phylogenetics of Ruscaceae sensu lato and related families (Asparagales) based on plastid and nuclear DNA sequences. Ann. Bot. 2010, 106. [Google Scholar] [CrossRef]
- Tamura, M.; Yamashita, J.; Fuse, S.; Haraguchi, M. Molecular phylogeny of monocotyledons inferred from combined analysis of plastid matK and rbcL gene sequences. J. Plant Res. 2004, 117, 109–120. [Google Scholar] [CrossRef]
- Rudall, P.J.; Conran, J.G.; Chase, M.W. Systematics of Ruscaceae/Convallariaceae: A combined morphological and molecular investigation. Bot. J. Linn. Soc. 2000, 134, 73–92. [Google Scholar] [CrossRef]
- Conran, J. Cladistic analyses of some net-veined Liliiflorae. Plant Syst. Evol. 1989, 168, 123–141. [Google Scholar]
- Chase, M.W.; Reveal, J.L. A phylogenetic classification of the land plants to accompany APG III. Bot. J. Linn. Soc. 2009, 161, 122–127. [Google Scholar]
- Bremer, B.; Bremer, K.; Chase, M.; Fay, M. An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG III. Bot. J. Linn. Soc. 2009, 161, 105–121. [Google Scholar] [CrossRef]
- Masiero, E.; Banik, D.; Abson, J.; Greene, P.; Slater, A.; Sgamma, T. Genus-Specific Real-Time PCR and HRM Assays to Distinguish Liriope from Ophiopogon Samples. Plants 2017, 6, 53. [Google Scholar] [CrossRef]
- Ito, M.; Sato-Masumoto, N.; Kobayashi, F.; Matsumura, K. Distinguishing Ophiopogon and Liriope tubers based on DNA sequences. J. Nat. Med. 2015, 69, 555–556. [Google Scholar] [CrossRef]
- Shiba, M.; Yamaji, H.; Kondo, K.; Ichiki, H.; Sakakibara, I. Discrimination of Maidong derived from Ophiopogon and Liriope species by rbcL sequences, and their chemical components and tuber anatomy. Nat. Med. 2004, 58, 15–21. [Google Scholar]
- Nei, M.; Kumar, S. Molecular Evolution and Phylogenetics; Oxford University Press: New York, NY, USA, 2000. [Google Scholar]
- Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 1980, 16, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Li, M.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Abson, J. Botany, medicine and mistaken identity. Plant Herit. J. 2017, 25, 41–43. [Google Scholar]
- Kress, J.W.; Erickson, D.L. A Two-Locus Global DNA Barcode for Land Plants: The Coding rbcL Gene Complements the Non-Coding trnH-psbA Spacer Region. PLoS ONE 2007, 2, e508. [Google Scholar] [CrossRef]
- Newmaster, S.G.; Fazekas, A.J.; Ragupathy, S. DNA barcoding in land plants: Evaluation of rbcL in a multigene tiered approach. Can. J. Bot. 2006, 84, 335–341. [Google Scholar] [CrossRef]
- Sgamma, T.; Lockie-Williams, C.; Kreuzer, M.; Williams, S.; Scheyhing, U.; Koch, E.; Slater, A.; Howard, C. DNA Barcoding for Industrial Quality Assurance. Planta Med. 2017, 14, 8–21. [Google Scholar] [CrossRef]
- Korir, N.K.; Han, J.; Shangguan, L.; Wang, C.; Kayesh, E.; Zhang, Y.; Fang, J. Plant variety and cultivar identification: Advances and prospects. Crit. Rev. Biotechnol. 2013, 33, 111. [Google Scholar] [CrossRef]
- Moller, M.; Cronk, Q.C.B. Origin and relationships of Saintpaulia (Gesneriaceae) based on ribosomal DNA internal transcribed spacer (ITS) sequences. Am. J. Bot. 1997, 84, 956–965. [Google Scholar] [CrossRef]
- Bickford, D.; Lohman, D.J.; Sodhi, N.S.; Ng, P.K.L.; Meier, R.; Winker, K.; Ingram, K.K.; Das, I. Cryptic species as a window on diversity and conservation. Trends Ecol. Evol. 2007, 22, 148–155. [Google Scholar] [CrossRef]
- Raclariu, A.C.; Heinrich, M.; Ichim, M.C.; de Boer, H. Benefits and limitations of DNA barcoding and metabarcoding in herbal product authentication. Phytochem. Anal. 2018, 29, 123–128. [Google Scholar] [CrossRef] [PubMed]
- Levin, R.A.; Wagner, W.L.; Hoch, P.C.; Nepokroeff, M.; Pires, J.C.; Zimmer, E.A.; Sytsma, K.J. Family-level relationships of Onagraceae based on chloroplast rbcL and ndhF data. Am. J. Bot. 2003, 90, 107–115. [Google Scholar] [PubMed]
- Kress, W.J.; Erickson, D.L.; Jones, F.A.; Swenson, N.G.; Pérez, R.A.; Sanjur, O.I.; Bermingham, E. Plant DNA barcodes and a community phylogeny of a tropical forest dynamics plot in Panama. Proc. Natl. Acad. Sci. USA 2009, 106, 18621–18626. [Google Scholar] [CrossRef] [PubMed]
- White, T.; Bruns, T.; Lee, S.; Taylor, J.; Innis, M.; Gelfand, D.; Sninsky, J. Amplification and Direct Sequencing of Fungal Ribosomal RNA Genes for Phylogenetics; Academic Press: New York, NY, USA, 1990. [Google Scholar]
| Primers | References | Sequences 5′–3′ | Annealing Temperature | Amplicon Size (bp) |
|---|---|---|---|---|
| rbcLaF | [40] | ATGTCACCACAAACAGAGACTAAAGC | 52 °C | ≃ 500/600 |
| rbcLaR | [41] | GTAAAATCAAGTCCACCRCG | ||
| rbcLbAF | [28] | CGGTGGACTTGATTTTACCA | 56 °C | ≃ 500 |
| rbcLbBR | TCATCACGTAATAAATCAAC | |||
| ITS1 | [42] | TCCGTAGGTGAACCTGCGG | 61 °C | ≃ 600 |
| ITS4 | TCCTCCGCTTATTGATATGC | |||
| ITS5P ITS8P | [37] | GGAAGGAGAAGTCGTAACAAGG CACGCTTCTCCAGACTACA | 61 °C | ≃ 600 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Masiero, E.; Banik, D.; Abson, J.; Greene, P.; Slater, A.; Sgamma, T. Molecular Verification of the UK National Collection of Cultivated Liriope and Ophiopogon Plants. Plants 2020, 9, 558. https://doi.org/10.3390/plants9050558
Masiero E, Banik D, Abson J, Greene P, Slater A, Sgamma T. Molecular Verification of the UK National Collection of Cultivated Liriope and Ophiopogon Plants. Plants. 2020; 9(5):558. https://doi.org/10.3390/plants9050558
Chicago/Turabian StyleMasiero, Eva, Dipanwita Banik, John Abson, Paul Greene, Adrian Slater, and Tiziana Sgamma. 2020. "Molecular Verification of the UK National Collection of Cultivated Liriope and Ophiopogon Plants" Plants 9, no. 5: 558. https://doi.org/10.3390/plants9050558
APA StyleMasiero, E., Banik, D., Abson, J., Greene, P., Slater, A., & Sgamma, T. (2020). Molecular Verification of the UK National Collection of Cultivated Liriope and Ophiopogon Plants. Plants, 9(5), 558. https://doi.org/10.3390/plants9050558





