Genome-Wide Analysis and Expression Profiling of Rice Hybrid Proline-Rich Proteins in Response to Biotic and Abiotic Stresses, and Hormone Treatment
Abstract
1. Introduction
2. Results and Discussion
2.1. Identification and Annotation of Putative OsHyPRPs in Rice
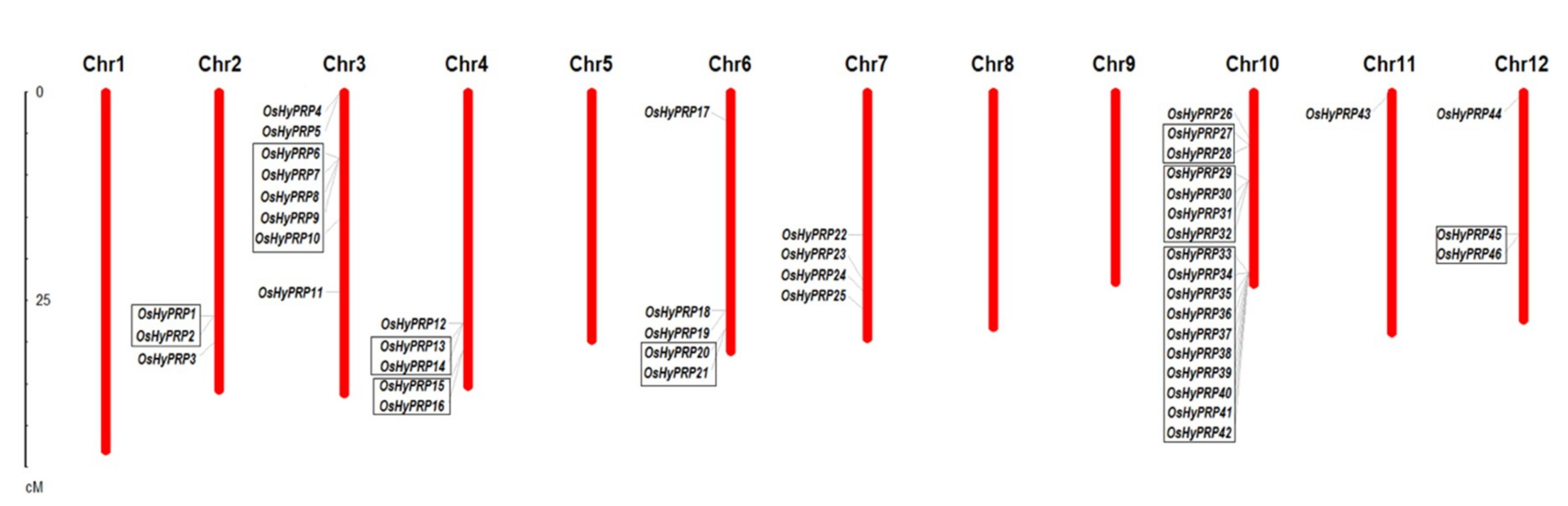
2.2. Chromosomal Location, Gene Duplication and Structural Analysis of OsHyPRPs
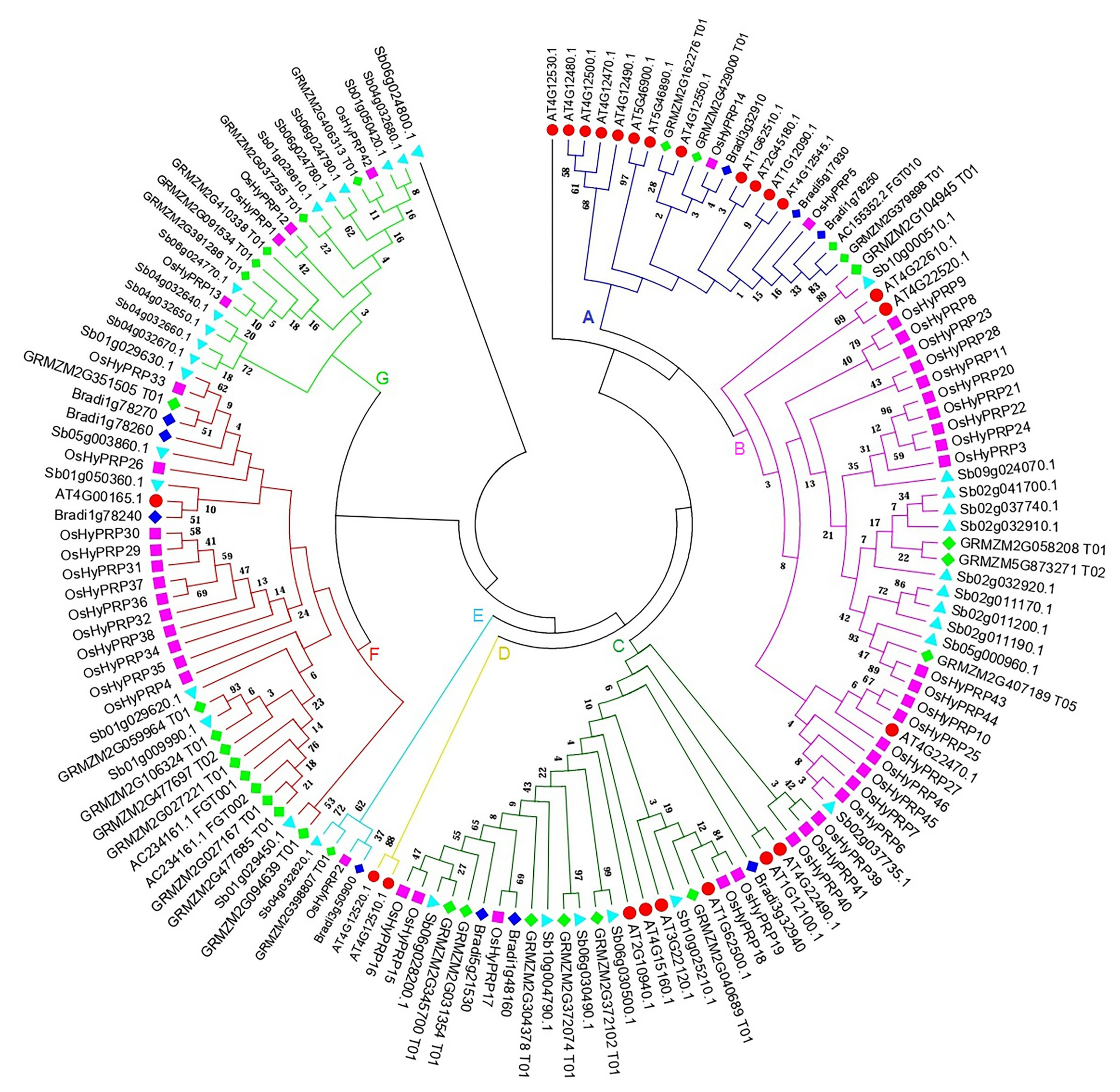
2.3. Phylogenetic Analysis of the OsHyPRP Family among Different Plant Species
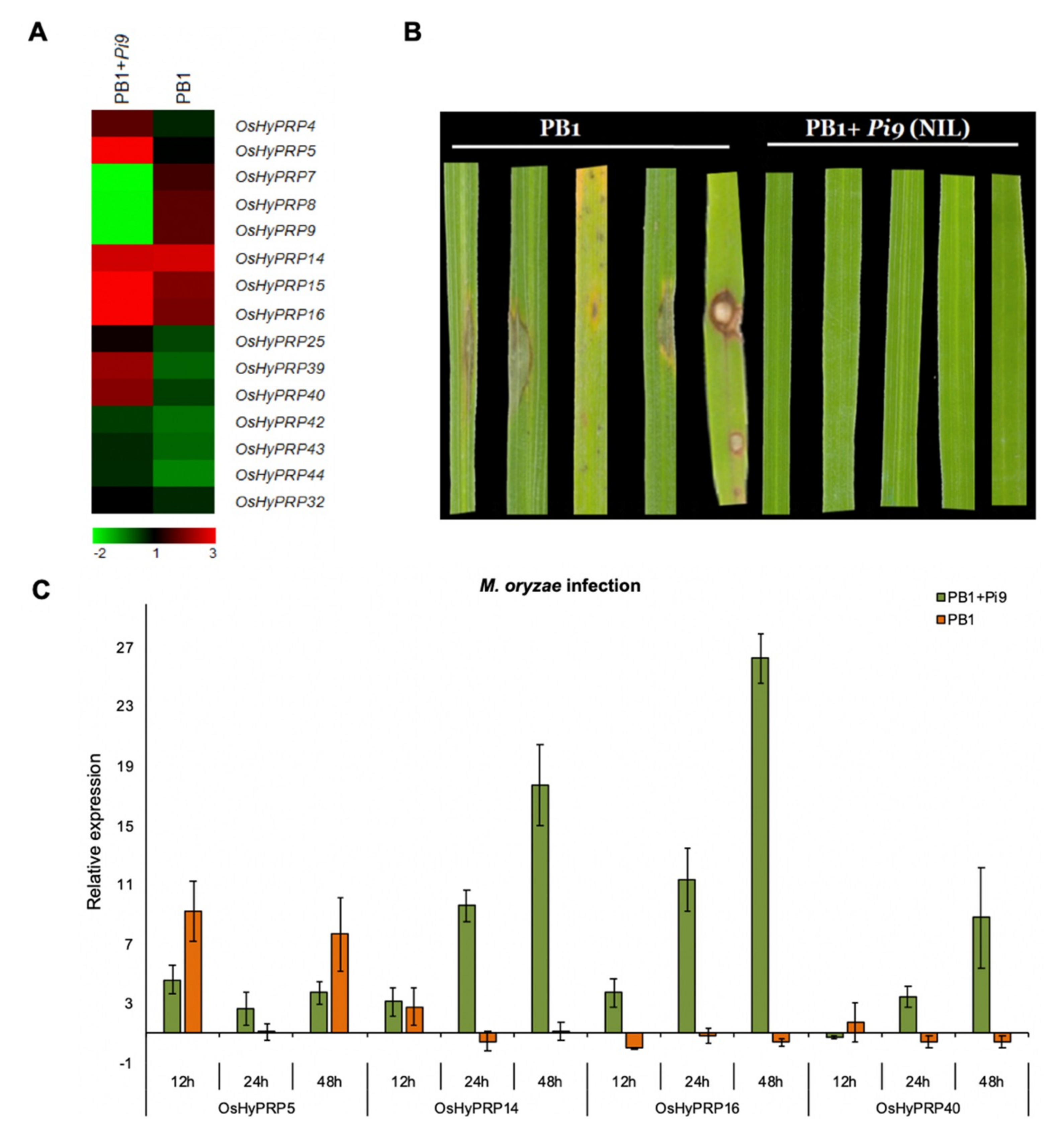
2.4. OsHyPRPs Expression Profile in Response to M. oryzae
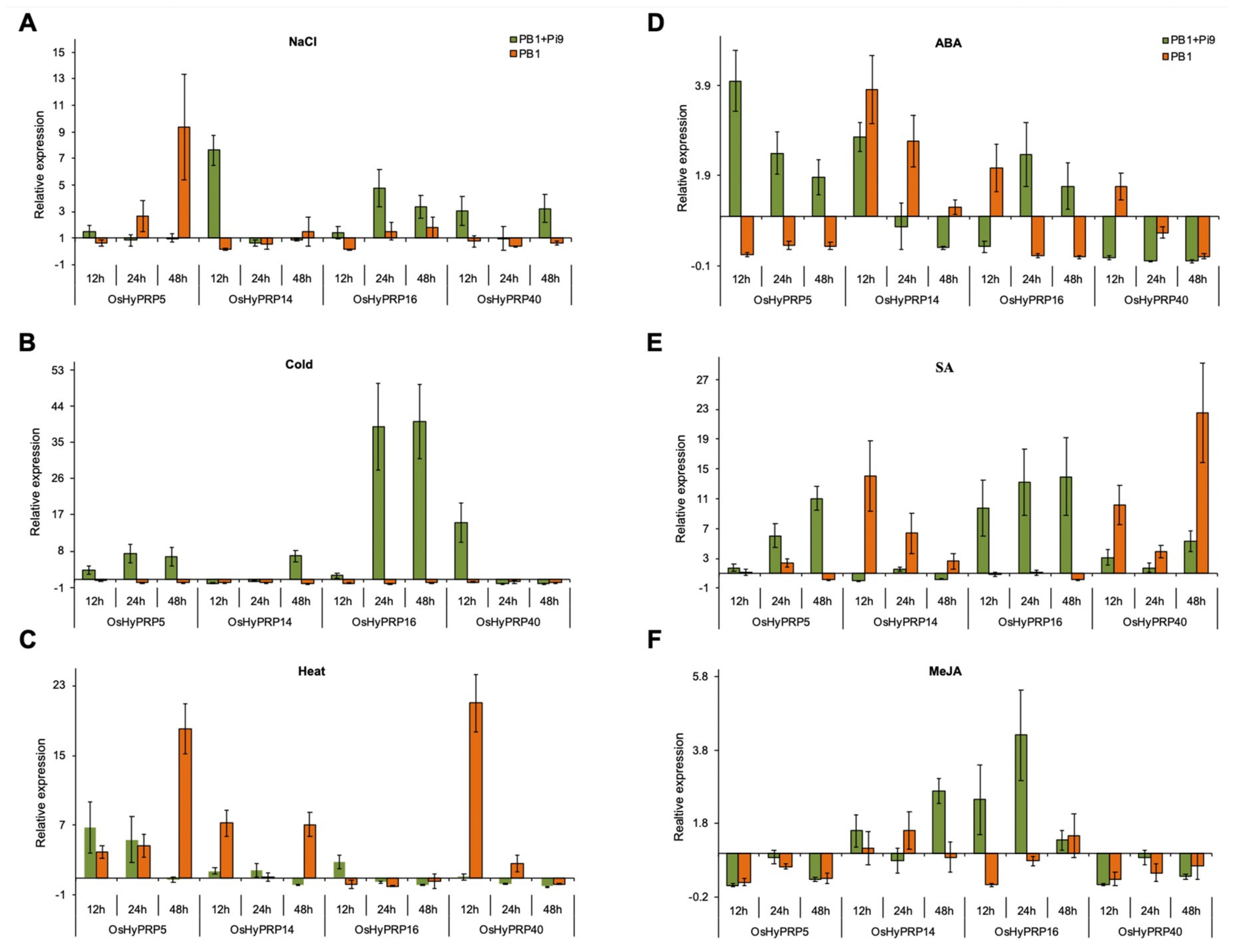
2.5. OsHyPRPs Expression Profile under Abiotic Stress
2.6. Expression Profiling of OsHyPRPs under Phytohormone Treatment
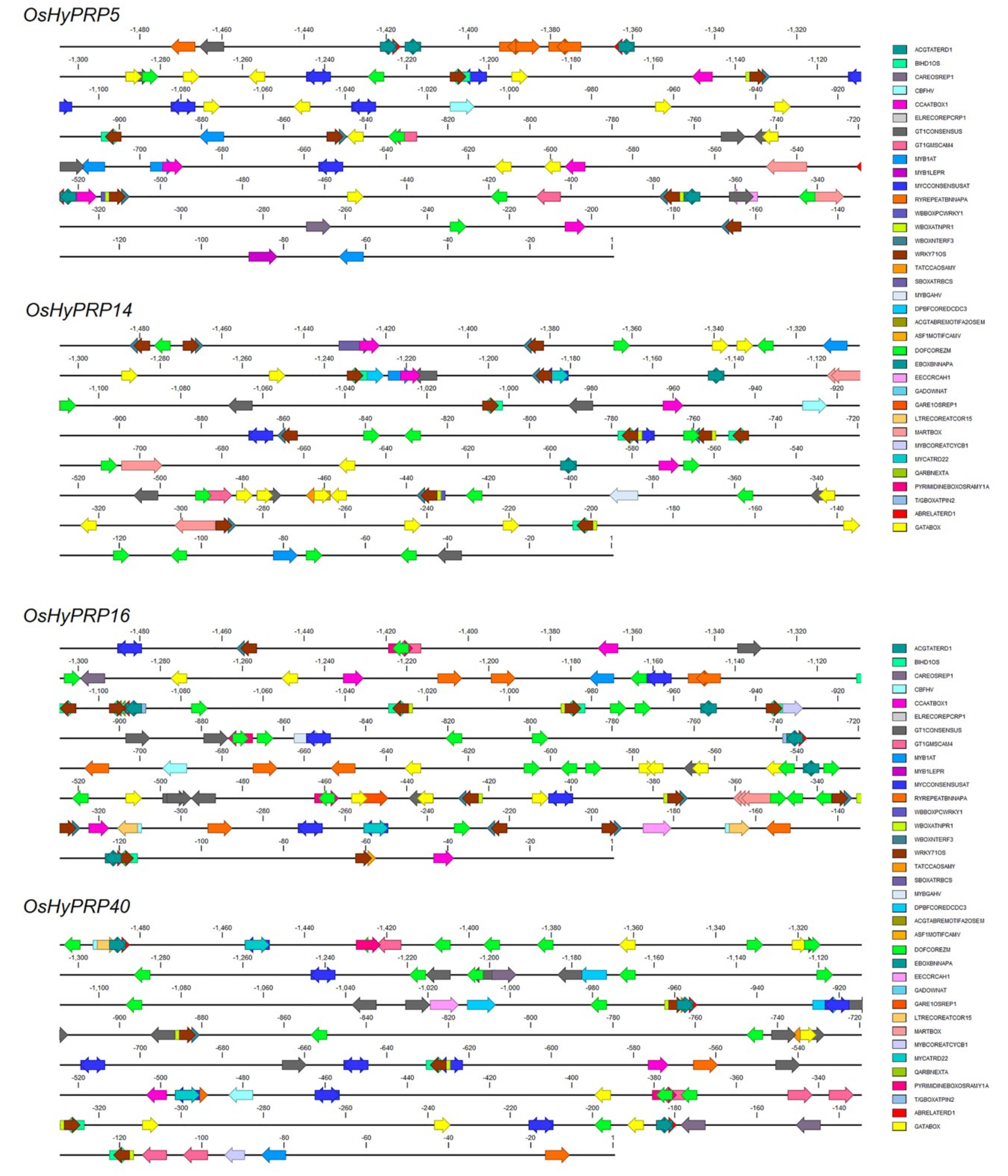
2.7. Analysis of Cis-Regulatory Elements in Four OsHyPRP Promoter Sequences
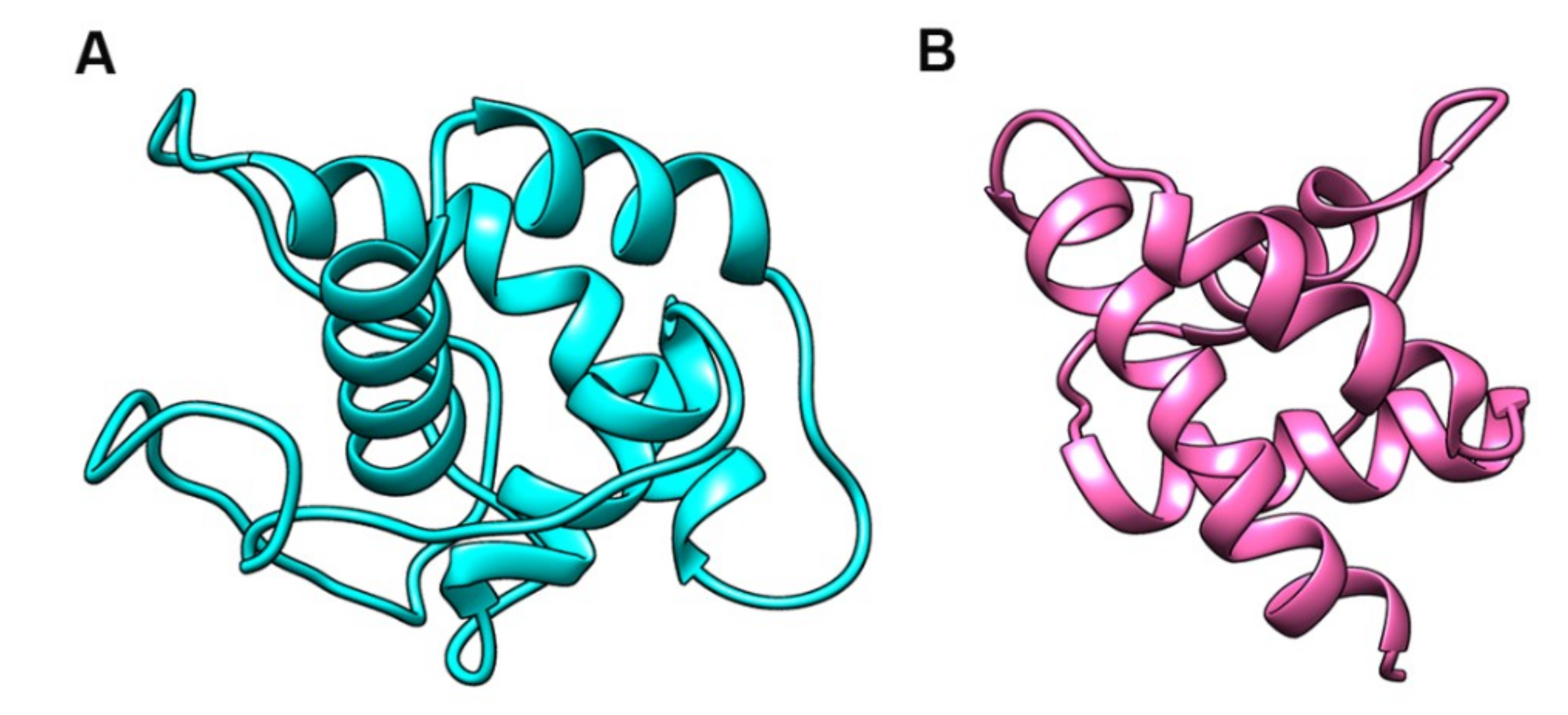
2.8. Protein Structure Prediction and Docking Analysis
3. Materials and Methods
3.1. Identification and Sequence Analysis of OsHyPRPs
3.2. Chromosomal Mapping and Gene Duplication Analysis of OsHyPRP Genes
3.3. Sequence Alignment and Phylogenetic Analysis
3.4. Biotic and Abiotic Stress and Hormone Treatment
3.5. RNA Extraction and Real-Time Quantitative Reverse Transcription PCR
3.6. Promoter Sequence Analysis of Cis-Regulatory Elements
3.7. Protein Structure Prediction and Docking Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Jones, J.D.G.; Dangl, J.L. The plant immune system. Nature 2006, 444, 323. [Google Scholar] [CrossRef] [PubMed]
- Singh, P.; Nag, A.; Arya, P.; Kapoor, R.; Singh, A.; Jaswal, R.; Sharma, T. Prospects of understanding the molecular biology of disease resistance in rice. Int. J. Mol. Sci. 2018, 19, 1141. [Google Scholar] [CrossRef] [PubMed]
- Ali, S.; Ganai, B.A.; Kamili, A.N.; Bhat, A.A.; Mir, Z.A.; Bhat, J.A.; Tyagi, A.; Islam, S.T.; Mushtaq, M.; Yadav, P. Pathogenesis-related proteins and peptides as promising tools for engineering plants with multiple stress tolerance. Microbiol. Res. 2018, 212, 29–37. [Google Scholar] [CrossRef] [PubMed]
- Diaz, I. Plant Defense Genes against Biotic Stresses. Int. J. Mol. Sci. 2018, 19, 2446. [Google Scholar] [CrossRef] [PubMed]
- Van Loon, L.C.; Rep, M.; Pieterse, C.M.J. Significance of inducible defense-related proteins in infected plants. Annu. Rev. Phytopathol. 2006, 44, 135–162. [Google Scholar] [CrossRef] [PubMed]
- Shewry, P.R.; Beaudoin, F.; Jenkins, J.; Griffiths-Jones, S.; Mills, E.N.C. Plant protein families and their relationships to food allergy. Biochem. Soc. Trans. 2002, 30, 906–910. [Google Scholar] [CrossRef] [PubMed]
- Radauer, C.; Breiteneder, H. Evolutionary biology of plant food allergens. J. Allergy Clin. Immunol. 2007, 120, 518–525. [Google Scholar] [CrossRef] [PubMed]
- José-Estanyol, M.; Gomis-Rüth, F.X.; Puigdomènech, P. The eight-cysteine motif, a versatile structure in plant proteins. Plant Physiol. Biochem. 2004, 42, 355–365. [Google Scholar] [CrossRef] [PubMed]
- Dvořáková, L.; Srba, M.; Opatrny, Z.; Fischer, L. Hybrid proline-rich proteins: Novel players in plant cell elongation? Ann. Bot. 2011, 109, 453–462. [Google Scholar] [CrossRef] [PubMed]
- Josè, M.; Puigdomènech, P. Structure and expression of genes coding for structural proteins of the plant cell wall. New Phytol. 1993, 125, 259–282. [Google Scholar] [CrossRef]
- Josè-Estanyol, M.; Puigdomènech, P. Plant cell wall glycoproteins and their genes. Plant Physiol. Biochem. 2000, 38, 97–108. [Google Scholar] [CrossRef]
- Dvořáková, L.; Cvrčková, F.; Fischer, L. Analysis of the hybrid proline-rich protein families from seven plant species suggests rapid diversification of their sequences and expression patterns. BMC Genom. 2007, 8, 412. [Google Scholar] [CrossRef] [PubMed]
- Yeats, T.H.; Rose, J.K.C. The biochemistry and biology of extracellular plant lipid-transfer proteins (LTPs). Protein Sci. 2008, 17, 191–198. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Zhang, X.; Lu, C.; Zeng, X.; Li, Y.; Fu, D.; Wu, G. Non-specific lipid transfer proteins in plants: Presenting new advances and an integrated functional analysis. J. Exp. Bot. 2015, 66, 5663–5681. [Google Scholar] [CrossRef] [PubMed]
- Finkina, E.I.; Melnikova, D.N.; Bogdanov, I.V. Lipid transfer proteins as components of the plant innate immune system: Structure, functions, and applications. Acta Naturae 2016, 8, 29. [Google Scholar] [CrossRef]
- Castonguay, Y.; Laberge, S.; Nadeau, P.; Vézina, L.-P. A cold-induced gene from Medicago sativa encodes a bimodular protein similar to developmentally regulated proteins. Plant Mol. Biol. 1994, 24, 799–804. [Google Scholar] [CrossRef]
- Deutch, C.E.; Winicov, I. Post-transcriptional regulation of a salt-inducible alfalfa gene encoding a putative chimeric proline-rich cell wall protein. Plant Mol. Biol. 1995, 27, 411–418. [Google Scholar] [CrossRef]
- Goodwin, W.; Pallas, J.A.; Jenkins, G.I. Transcripts of a gene encoding a putative cell wall-plasma membrane linker protein are specifically cold-induced in Brassica napus. Plant Mol. Biol. 1996, 31, 771–781. [Google Scholar] [CrossRef]
- Zhang, Y.; Schläppi, M. Cold responsive EARLI1 type HyPRPs improve freezing survival of yeast cells and form higher order complexes in plants. Planta 2007, 227, 233–243. [Google Scholar] [CrossRef]
- Priyanka, B.; Sekhar, K.; Reddy, V.D.; Rao, K.V. Expression of pigeonpea hybrid-proline-rich protein encoding gene (CcHyPRP) in yeast and Arabidopsis affords multiple abiotic stress tolerance. Plant Biotechnol. J. 2010, 8, 76–87. [Google Scholar] [CrossRef]
- Huang, G.; Gong, S.; Xu, W.; Li, P.; Zhang, D.; Qin, L.; Li, W.; Li, X. GhHyPRP4, a cotton gene encoding putative hybrid proline-rich protein, is preferentially expressed in leaves and involved in plant response to cold stress. Acta Biochim. Biophys. Sin. 2011, 43, 519–527. [Google Scholar] [CrossRef] [PubMed]
- Qin, L.-X.; Zhang, D.-J.; Huang, G.-Q.; Li, L.; Li, J.; Gong, S.-Y.; Li, X.-B.; Xu, W.-L. Cotton GhHyPRP3 encoding a hybrid proline-rich protein is stress inducible and its overexpression in Arabidopsis enhances germination under cold temperature and high salinity stress conditions. Acta Physiol. Plant. 2013, 35, 1531–1542. [Google Scholar] [CrossRef]
- Tan, J.; Zhuo, C.; Guo, Z. Nitric oxide mediates cold-and dehydration-induced expression of a novel MfHyPRP that confers tolerance to abiotic stress. Physiol. Plant. 2013, 149, 310–320. [Google Scholar] [PubMed]
- Liu, D.Q.; Han, Q.; Shah, T.; Chen, C.Y.; Wang, Q.; Tang, B.F.; Ge, F. A hybrid proline-rich cell-wall protein gene JsPRP1 from Juglans sigillata Dode confers both biotic and abiotic stresses in transgenic tobacco plants. Trees 2018, 32, 1199–1209. [Google Scholar] [CrossRef]
- Jose-Estanyol, M.; Ruiz-Avila, L.; Puigdomènech, P. A maize embryo-specific gene encodes a proline-rich and hydrophobic protein. Plant Cell 1992, 4, 413–423. [Google Scholar] [CrossRef] [PubMed]
- He, C.-Y.; Zhang, J.-S.; Chen, S.-Y. A soybean gene encoding a proline-rich protein is regulated by salicylic acid, an endogenous circadian rhythm and by various stresses. Theor. Appl. Genet. 2002, 104, 1125–1131. [Google Scholar] [CrossRef] [PubMed]
- Bücker Neto, L.; de Oliveira, R.R.; Wiebke-Strohm, B.; Bencke, M.; Weber, R.L.M.; Cabreira, C.; Abdelnoor, R.V.; Marcelino, F.C.; Zanettini, M.H.B.; Passaglia, L.M.P. Identification of the soybean HyPRP family and specific gene response to Asian soybean rust disease. Genet. Mol. Biol. 2013, 36, 214–224. [Google Scholar] [CrossRef][Green Version]
- Li, B.-C.; Zhang, C.; Chai, Q.-X.; Han, Y.-Y.; Wang, X.-Y.; Liu, M.-X.; Feng, H.; Xu, Z.-Q. Plasmalemma localisation of DOUBLE HYBRID PROLINE-RICH PROTEIN 1 and its function in systemic acquired resistance of Arabidopsis thaliana. Funct. Plant Biol. 2014, 41, 768–779. [Google Scholar] [CrossRef]
- Li, J.; Ouyang, B.; Wang, T.; Luo, Z.; Yang, C.; Li, H.; Sima, W.; Zhang, J.; Ye, Z. HyPRP1 gene suppressed by multiple stresses plays a negative role in abiotic stress tolerance in tomato. Front. Plant Sci. 2016, 7, 967. [Google Scholar] [CrossRef]
- Wu, H.; Zou, J.; May, B.; Gu, Q.; Cheung, A.Y. A tobacco gene family for flower cell wall proteins with a proline-rich domain and a cysteine-rich domain. Proc. Natl. Acad. Sci. USA 1993, 90, 6829–6833. [Google Scholar] [CrossRef]
- Holk, A.; Klumpp, L.; Scherer, G.F.E. A cell wall protein down-regulated by auxin suppresses cell expansion in Daucus carota (L.). Plant Mol. Biol. 2002, 50, 295–305. [Google Scholar] [CrossRef] [PubMed]
- Blanco-Portales, R.; Lopez-Raez, J.A.; Bellido, M.L.; Moyano, E.; Dorado, G.; Gonzalez-Reyes, J.A.; Caballero, J.L.; Munoz-Blanco, J. A strawberry fruit-specific and ripening-related gene codes for a HyPRP protein involved in polyphenol anchoring. Plant Mol. Biol. 2004, 55, 763–780. [Google Scholar] [CrossRef] [PubMed]
- Bouton, S.; Viau, L.; Lelièvre, E.; Limami, A.M. A gene encoding a protein with a proline-rich domain (MtPPRD1), revealed by suppressive subtractive hybridization (SSH), is specifically expressed in the Medicago truncatula embryo axis during germination. J. Exp. Bot. 2005, 56, 825–832. [Google Scholar] [CrossRef] [PubMed]
- Sundaresan, S.; Philosoph-Hadas, S.; Ma, C.; Jiang, C.-Z.; Riov, J.; Mugasimangalam, R.; Kochanek, B.; Salim, S.; Reid, M.S.; Meir, S. The Tomato Hybrid Proline-rich Protein regulates the abscission zone competence to respond to ethylene signals. Hortic. Res. 2018, 5, 28. [Google Scholar] [CrossRef] [PubMed]
- Bubier, J.; Schläppi, M. Cold induction of EARLI1, a putative Arabidopsis lipid transfer protein, is light and calcium dependent. Plant. Cell Environ. 2004, 27, 929–936. [Google Scholar] [CrossRef]
- Peng, T.; Jia, M.-M.; Liu, J.-H. RNAi-based functional elucidation of PtrPRP, a gene encoding a hybrid proline rich protein, in cold tolerance of Poncirus trifoliata. Front. Plant Sci. 2015, 6, 808. [Google Scholar] [CrossRef] [PubMed]
- Mellacheruvu, S.; Tamirisa, S.; Vudem, D.R.; Khareedu, V.R. Pigeonpea hybrid-proline-rich protein (CcHyPRP) confers biotic and abiotic stress tolerance in transgenic rice. Front. Plant Sci. 2016, 6, 1167. [Google Scholar] [CrossRef]
- Jung, H.W.; Tschaplinski, T.J.; Wang, L.; Glazebrook, J.; Greenberg, J.T. Priming in systemic plant immunity. Science 2009, 324, 89–91. [Google Scholar] [CrossRef]
- Yeom, S.; Seo, E.; Oh, S.; Kim, K.W.; Choi, D. A common plant cell-wall protein HyPRP1 has dual roles as a positive regulator of cell death and a negative regulator of basal defense against pathogens. Plant J. 2012, 69, 755–768. [Google Scholar] [CrossRef]
- Yang, J.; Zhang, Y.; Wang, X.; Wang, W.; Li, Z.; Wu, J.; Wang, G.; Wu, L.; Zhang, G.; Ma, Z. HyPRP1 performs a role in negatively regulating cotton resistance to V. dahliae via the thickening of cell walls and ROS accumulation. BMC Plant Biol. 2018, 18, 339. [Google Scholar] [CrossRef]
- Imam, J.; Alam, S.; Mandal, N.P.; Variar, M.; Shukla, P. Molecular screening for identification of blast resistance genes in North East and Eastern Indian rice germplasm (Oryza sativa L.) with PCR based makers. Euphytica 2014, 196, 199–211. [Google Scholar] [CrossRef]
- Khanna, A.; Sharma, V.; Ellur, R.K.; Shikari, A.B.; Krishnan, S.G.; Singh, U.D.; Prakash, G.; Sharma, T.R.; Rathour, R.; Variar, M. Development and evaluation of near-isogenic lines for major blast resistance gene (s) in Basmati rice. Theor. Appl. Genet. 2015, 128, 1243–1259. [Google Scholar] [CrossRef] [PubMed]
- Jain, P.; Singh, P.K.; Kapoor, R.; Khanna, A.; Solanke, A.U.; Krishnan, S.G.; Singh, A.K.; Sharma, V.; Sharma, T.R. Understanding host-pathogen interactions with expression profiling of NILs carrying rice-blast resistance Pi9 gene. Front. Plant Sci. 2017, 8, 93. [Google Scholar] [CrossRef] [PubMed]
- Boutrot, F.; Chantret, N.; Gautier, M.-F. Genome-wide analysis of the rice and Arabidopsis non-specific lipid transfer protein (nsLtp) gene families and identification of wheat nsLtp genes by EST data mining. BMC Genomics 2008, 9, 86. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Wang, J.; Lin, W.; Li, S.; Li, H.; Zhou, J.; Ni, P.; Dong, W.; Hu, S.; Zeng, C. The genomes of Oryza sativa: A history of duplications. PLoS Biol. 2005, 3, e38. [Google Scholar] [CrossRef] [PubMed]
- Seherm, H.; Coakley, S.M. Plant pathogens in a changing world. Australas. Plant Pathol. 2003, 32, 157–165. [Google Scholar] [CrossRef]
- Pandey, P.; Irulappan, V.; Bagavathiannan, M.V.; Senthil-Kumar, M. Impact of combined abiotic and biotic stresses on plant growth and avenues for crop improvement by exploiting physio-morphological traits. Front. Plant Sci. 2017, 8, 537. [Google Scholar] [CrossRef] [PubMed]
- Creissen, H.E.; Jorgensen, T.H.; Brown, J.K.M. Impact of disease on diversity and productivity of plant populations. Funct. Ecol. 2016, 30, 649–657. [Google Scholar] [CrossRef]
- Fernandez, J.; Orth, K. Rise of a cereal killer: The biology of Magnaporthe oryzae biotrophic growth. Trends Microbiol. 2018, 26, 582–597. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Zhang, C.; Xu, D.; Schläppi, M.; Xu, Z.-Q. Expression of recombinant EARLI1, a hybrid proline-rich protein of Arabidopsis, in Escherichia coli and its inhibition effect to the growth of fungal pathogens and Saccharomyces cerevisiae. Gene 2012, 506, 50–61. [Google Scholar] [CrossRef]
- Guy, C.; Kaplan, F.; Kopka, J.; Selbig, J.; Hincha, D.K. Metabolomics of temperature stress. Physiol. Plant. 2008, 132, 220–235. [Google Scholar] [CrossRef] [PubMed]
- Aghamolki, M.T.K.; Yusop, M.K.; Oad, F.C.; Zakikhani, H.; Jaafar, H.Z.; Kharidah, S.; Musa, M.H. Heat stress effects on yield parameters of selected rice cultivars at reproductive growth stages. J. Food Agric. Env. 2014, 12, 741–746. [Google Scholar]
- Hatfield, J.L.; Prueger, J.H. Temperature extremes: Effect on plant growth and development. Weather Clim. Extrem. 2015, 10, 4–10. [Google Scholar] [CrossRef]
- Rodríguez, V.M.; Soengas, P.; Alonso-Villaverde, V.; Sotelo, T.; Cartea, M.E.; Velasco, P. Effect of temperature stress on the early vegetative development of Brassica oleracea L. BMC Plant Biol. 2015, 15, 145. [Google Scholar] [CrossRef] [PubMed]
- Jing, P.; Wang, D.; Zhu, C.; Chen, J. Plant physiological, morphological and yield-related responses to night temperature changes across different species and plant functional types. Front. Plant Sci. 2016, 7, 1774. [Google Scholar] [CrossRef] [PubMed]
- Nievola, C.C.; Carvalho, C.P.; Carvalho, V.; Rodrigues, E. Rapid responses of plants to temperature changes. Temperature 2017, 4, 371–405. [Google Scholar] [CrossRef] [PubMed]
- Srinath, T.; Reddy, V.D.; Rao, K.V. Isolation and functional characterization of a novel stress inducible promoter from pigeonpea (Cajanus cajan L.). Plant Cell Tissue Organ Cult. 2017, 128, 457–468. [Google Scholar] [CrossRef]
- Wilkosz, R.; Schläppi, M. A gene expression screen identifies EARLI1 as a novel vernalization-responsive gene in Arabidopsis thaliana. Plant Mol. Biol. 2000, 44, 777–787. [Google Scholar] [CrossRef]
- Xu, D.; Huang, X.; Xu, Z.-Q.; Schläppi, M. The HyPRP gene EARLI1 has an auxiliary role for germinability and early seedling development under low temperature and salt stress conditions in Arabidopsis thaliana. Planta 2011, 234, 565–577. [Google Scholar] [CrossRef]
- Xu, Z.-Y.; Zhang, X.; Schläppi, M.; Xu, Z.-Q. Cold-inducible expression of AZI1 and its function in improvement of freezing tolerance of Arabidopsis thaliana and Saccharomyces cerevisiae. J. Plant Physiol. 2011, 168, 1576–1587. [Google Scholar] [CrossRef]
- Pitzschke, A.; Datta, S.; Persak, H. Salt stress in Arabidopsis: Lipid transfer protein AZI1 and its control by mitogen-activated protein kinase MPK3. Mol. Plant 2014, 7, 722–738. [Google Scholar] [CrossRef] [PubMed]
- Vickers, C.E.; Gershenzon, J.; Lerdau, M.T.; Loreto, F. A unified mechanism of action for volatile isoprenoids in plant abiotic stress. Nat. Chem. Biol. 2009, 5, 283–291. [Google Scholar] [CrossRef] [PubMed]
- Gray, W.M. Hormonal regulation of plant growth and development. PLoS Biol. 2004, 2, e311. [Google Scholar] [CrossRef] [PubMed]
- Kamiya, Y. Plant hormones: Versatile regulators of plant growth and development. Annu. Rev. Plant Biol. 2010, 60. [Google Scholar] [CrossRef]
- Wang, Y.H.; Irving, H.R. Developing a model of plant hormone interactions. Plant Signal. Behav. 2011, 6, 494–500. [Google Scholar] [CrossRef] [PubMed]
- Munné-Bosch, S.; Müller, M. Hormonal cross-talk in plant development and stress responses. Front. Plant Sci. 2013, 4, 529. [Google Scholar] [CrossRef] [PubMed]
- Nejat, N.; Mantri, N. Plant immune system: Crosstalk between responses to biotic and abiotic stresses the missing link in understanding plant defence. Curr. Issues Mol. Biol. 2017, 23, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Ku, Y.-S.; Sintaha, M.; Cheung, M.-Y.; Lam, H.-M. Plant hormone signaling crosstalks between biotic and abiotic stress responses. Int. J. Mol. Sci. 2018, 19, 3206. [Google Scholar] [CrossRef] [PubMed]
- Kunkel, B.N.; Brooks, D.M. Cross talk between signaling pathways in pathogen defense. Curr. Opin. Plant Biol. 2002, 5, 325–331. [Google Scholar] [CrossRef]
- Kohli, A.; Sreenivasulu, N.; Lakshmanan, P.; Kumar, P.P. The phytohormone crosstalk paradigm takes center stage in understanding how plants respond to abiotic stresses. Plant Cell Rep. 2013, 32, 945–957. [Google Scholar] [CrossRef]
- Pandey, G.K.; Grant, J.J.; Cheong, Y.H.; Kim, B.G.; Li, L.; Luan, S. ABR1, an APETALA2-domain transcription factor that functions as a repressor of ABA response in Arabidopsis. Plant Physiol. 2005, 139, 1185–1193. [Google Scholar] [CrossRef] [PubMed]
- Hernandez-Garcia, C.M.; Finer, J.J. Identification and validation of promoters and cis-acting regulatory elements. Plant Sci. 2014, 217, 109–119. [Google Scholar] [CrossRef] [PubMed]
- Buhot, N.; Gomes, E.; Milat, M.-L.; Ponchet, M.; Marion, D.; Lequeu, J.; Delrot, S.; Coutos-Thévenot, P.; Blein, J.-P. Modulation of the biological activity of a tobacco LTP1 by lipid complexation. Mol. Biol. Cell 2004, 15, 5047–5052. [Google Scholar] [CrossRef] [PubMed]
- Williamson, M.P. The structure and function of proline-rich regions in proteins. Biochem. J. 1994, 297, 249. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Tang, H.; DeBarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.; Jin, H.; Marler, B.; Guo, H. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
- Bonman, J.M.; De Dios, T.I.V.; Khin, M.M. Physiologic specialization of Pyricularia oryzae in the Philippines. Plant Dis. 1986, 70, 767–769. [Google Scholar] [CrossRef]
- Yang, J.; Yan, R.; Roy, A.; Xu, D.; Poisson, J.; Zhang, Y. The I-TASSER Suite: Protein structure and function prediction. Nat. Methods 2015, 12, 7–8. [Google Scholar] [CrossRef]
- Xu, D.; Zhang, J.; Roy, A.; Zhang, Y. Automated protein structure modeling in CASP9 by I-TASSER pipeline combined with QUARK-based ab initio folding and FG-MD-based structure refinement. Proteins Struct. Funct. Bioinform. 2011, 79, 147–160. [Google Scholar] [CrossRef]
- Huey, R.; Morris, G.M.; Olson, A.J.; Goodsell, D.S. A semiempirical free energy force field with charge-based desolvation. J. Comput. Chem. 2007, 28, 1145–1152. [Google Scholar] [CrossRef] [PubMed]
- Bikadi, Z.; Hazai, E. Application of the PM6 semi-empirical method to modeling proteins enhances docking accuracy of AutoDock. J. Cheminform. 2009, 1, 15. [Google Scholar] [CrossRef] [PubMed]
- O’Boyle, N.M.; Banck, M.; James, C.A.; Morley, C.; Vandermeersch, T.; Hutchison, G.R. Open Babel: An open chemical toolbox. J. Cheminform. 2011, 3, 33. [Google Scholar] [CrossRef] [PubMed]

| Locus IDs | Nomenclature | Signal Peptide | Mature Protein | |||
|---|---|---|---|---|---|---|
| Amino Acid | Target a | Amino Acid | Mass (kDa) | pI | ||
| LOC_Os02g44310.1 | OsHyPRP1 | 22 | S | 111 | 11.27 | 8.14 |
| LOC_Os02g44320.1 | OsHyPRP2 | 22 | S | 104 | 10.65 | 6.24 |
| LOC_Os02g49280.1 | OsHyPRP3 | 28 | S | 233 | 23.6 | 5.43 |
| LOC_Os03g01300.1 | OsHyPRP4 | 29 | S | 110 | 11.32 | 5.04 |
| LOC_Os03g01320.1 | OsHyPRP5 | 24 | S | 160 | 13.97 | 8.37 |
| LOC_Os03g14615.1 | OsHyPRP6 | 25 | S | 404 | 41.07 | 8.91 |
| LOC_Os03g14630.1 | OsHyPRP7 | 25 | S | 103 | 10.66 | 8.50 |
| LOC_Os03g14642.1 | OsHyPRP8 | 26 | S | 122 | 12.29 | 8.15 |
| LOC_Os03g14654.1 | OsHyPRP9 | 26 | S | 132 | 13.1 | 8.44 |
| LOC_Os03g26800.1 | OsHyPRP10 | 27 | S | 112 | 11.16 | 6.20 |
| LOC_Os03g43050.1 | OsHyPRP11 | 20 | S | 221 | 23 | 4.69 |
| LOC_Os04g46810.1 | OsHyPRP12 | 23 | S | 105 | 10.8 | 8.65 |
| LOC_Os04g46820.1 | OsHyPRP13 | 23 | S | 106 | 10.91 | 8.43 |
| LOC_Os04g46830.1 | OsHyPRP14 | 22 | S | 115 | 7.4 | 8.02 |
| LOC_Os04g52250.1 | OsHyPRP15 | 27 | S | 175 | 17.56 | 4.88 |
| LOC_Os04g52260.1 | OsHyPRP16 | 32 | S | 122 | 12.18 | 8.04 |
| LOC_Os06g07220.1 | OsHyPRP17 | 22 | S | 224 | 23.35 | 8.96 |
| LOC_Os06g43600.1 | OsHyPRP18 | 27 | S | 228 | 23.89 | 8.86 |
| LOC_Os06g43600.2 | OsHyPRP19 | 27 | S | 228 | 23.89 | 8.86 |
| LOC_Os06g46780.1 | OsHyPRP20 | 28 | S | 240 | 23.79 | 5.51 |
| LOC_Os06g46870.1 | OsHyPRP21 | 28 | S | 242 | 23.98 | 5.51 |
| LOC_Os07g29230.1 | OsHyPRP22 | 26 | S | 247 | 25.26 | 5.24 |
| LOC_Os07g37385.1 | OsHyPRP23 | 26 | S | 283 | 28.86 | 9.89 |
| LOC_Os07g39640.1 | OsHyPRP24 | 19 | M | 186 | 19.14 | 4.72 |
| LOC_Os07g43290.2 | OsHyPRP25 | 22 | S | 141 | 14.61 | 8.40 |
| LOC_Os10g09920.1 | OsHyPRP26 | 25 | S | 103 | 10.36 | 3.97 |
| LOC_Os10g11370.1 | OsHyPRP27 | 21 | S | 336 | 35.39 | 8.88 |
| LOC_Os10g11730.1 | OsHyPRP28 | 23 | S | 143 | 15.18 | 5.15 |
| LOC_Os10g20830.1 | OsHyPRP29 | 23 | S | 114 | 11.75 | 6.44 |
| LOC_Os10g20840.1 | OsHyPRP30 | 23 | S | 114 | 11.82 | 6.86 |
| LOC_Os10g20860.1 | OsHyPRP31 | 23 | S | 110 | 11.51 | 5.98 |
| LOC_Os10g20890.1 | OsHyPRP32 | 28 | S | 98 | 10.17 | 7.08 |
| LOC_Os10g40420.1 | OsHyPRP33 | 34 | S | 128 | 12.58 | 8.68 |
| LOC_Os10g40430.1 | OsHyPRP34 | 23 | S | 123 | 12.27 | 7.07 |
| LOC_Os10g40440.1 | OsHyPRP35 | 23 | S | 119 | 12.05 | 6.62 |
| LOC_Os10g40460.1 | OsHyPRP36 | 25 | S | 106 | 10.96 | 6.85 |
| LOC_Os10g40470.1 | OsHyPRP37 | 24 | S | 107 | 10.99 | 6.92 |
| LOC_Os10g40480.1 | OsHyPRP38 | 24 | S | 112 | 11.58 | 7.72 |
| LOC_Os10g40510.1 | OsHyPRP39 | 21 | S | 112 | 11.49 | 7.38 |
| LOC_Os10g40520.1 | OsHyPRP40 | 22 | S | 100 | 10.42 | 8.00 |
| LOC_Os10g40530.1 | OsHyPRP41 | 21 | S | 111 | 11.55 | 8.05 |
| LOC_Os10g40614.1 | OsHyPRP42 | 26 | S | 141 | 13.17 | 7.91 |
| LOC_Os11g02165.1 | OsHyPRP43 | 29 | C | 222 | 22.95 | 9.45 |
| LOC_Os12g02105.1 | OsHyPRP44 | 28 | C | 219 | 22.43 | 9.35 |
| LOC_Os12g28880.1 | OsHyPRP45 | 24 | C | 148 | 15.28 | 8.74 |
| LOC_Os12g29040.1 | OsHyPRP46 | 20 | M | 190 | 20.01 | 8.26 |
| Ligand | Estimated Free Energy of Binding (kcal/mol) | vdW + Hbond + Desolvation Energy (kcal/mol) | Electrostatic Energy (kcal/mol) | Total Intermolecular Energy (kcal/mol) | Frequency (%) | Interaction Surface | |
|---|---|---|---|---|---|---|---|
| Mature OsHyPRP16 | |||||||
| Sterols | Lanosterol | −5.65 | −7.37 | −0.01 | −7.38 | 75 | 869.90 |
| Desmosterol | −8.88 | −10.61 | −0.04 | −10.65 | 67 | 809.07 | |
| Cholesterol | −9.02 | −10.68 | −0.01 | −10.69 | 57 | 817.93 | |
| Fungisterol | −7.28 | −9.42 | −0.08 | −9.5 | 23 | 819.20 | |
| Ergosterol | −8.08 | −9.52 | −0.01 | −9.53 | 64 | 832.67 | |
| Lipids and plant hormones | Linolenic acid | −6.36 | −9.15 | −0.32 | −9.48 | 12 | 677.53 |
| JA | −6.09 | −6.41 | −0.9 | −7.31 | 90 | 526.62 | |
| MeJA | −5.56 | −6.91 | −0.01 | −6.92 | 33 | 540.28 | |
| SA | −5.54 | −5.66 | −0.11 | −5.77 | 21 | 484.46 | |
| Pathogen cell wall components | NAG | −5.5 | −5.64 | −0.11 | −5.75 | 21 | 484.33 |
| NAM | −6.31 | −6.33 | −0.44 | −6.77 | 48 | 609.37 | |
| Fragment Deleted OsHyPRP16 | |||||||
| Sterols | Lanosterol | 0.7 | −0.96 | −0.01 | −0.97 | 50 | 867.84 |
| Desmosterol | 4.86 | 1.84 | −0.05 | −4.89 | 30 | 616.49 | |
| Cholesterol | 8.4 | −3.13 | −0.02 | −315% | 40 | 722.06 | |
| Fungisterol | 2.7 | −4.84 | −0.05 | −4.89 | 30 | 716.49 | |
| Ergosterol | 4.08 | 1.26 | 0.06 | 1.32 | 40 | 743.26 | |
| Lipids and plant hormones | Linolenic acid | 2.2 | −0.95 | −0.46 | −1.42 | 1 | 600.68 |
| JA | −5.35 | −6.31 | −0.64 | −6.95 | 70 | 538.91 | |
| MeJA | −2.63 | −4.28 | −0.04 | −4.32 | 24 | 447.63 | |
| SA | −2.3 | −2.78 | −0.13 | −2.91 | 19 | 339.79 | |
| Pathogen cell wall components | NAG | −2.38 | −2.72 | −0.05 | −2.76 | 15 | 381.84 |
| NAM | −2.21 | −3.1 | −0.22 | −3.32 | 4 | 422.58 | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kapoor, R.; Kumar, G.; Arya, P.; Jaswal, R.; Jain, P.; Singh, K.; Sharma, T.R. Genome-Wide Analysis and Expression Profiling of Rice Hybrid Proline-Rich Proteins in Response to Biotic and Abiotic Stresses, and Hormone Treatment. Plants 2019, 8, 343. https://doi.org/10.3390/plants8090343
Kapoor R, Kumar G, Arya P, Jaswal R, Jain P, Singh K, Sharma TR. Genome-Wide Analysis and Expression Profiling of Rice Hybrid Proline-Rich Proteins in Response to Biotic and Abiotic Stresses, and Hormone Treatment. Plants. 2019; 8(9):343. https://doi.org/10.3390/plants8090343
Chicago/Turabian StyleKapoor, Ritu, Gulshan Kumar, Preeti Arya, Rajdeep Jaswal, Priyanka Jain, Kashmir Singh, and Tilak Raj Sharma. 2019. "Genome-Wide Analysis and Expression Profiling of Rice Hybrid Proline-Rich Proteins in Response to Biotic and Abiotic Stresses, and Hormone Treatment" Plants 8, no. 9: 343. https://doi.org/10.3390/plants8090343
APA StyleKapoor, R., Kumar, G., Arya, P., Jaswal, R., Jain, P., Singh, K., & Sharma, T. R. (2019). Genome-Wide Analysis and Expression Profiling of Rice Hybrid Proline-Rich Proteins in Response to Biotic and Abiotic Stresses, and Hormone Treatment. Plants, 8(9), 343. https://doi.org/10.3390/plants8090343






