Contribution of Root Hair Development to Sulfate Uptake in Arabidopsis
Abstract
:1. Introduction
2. Results
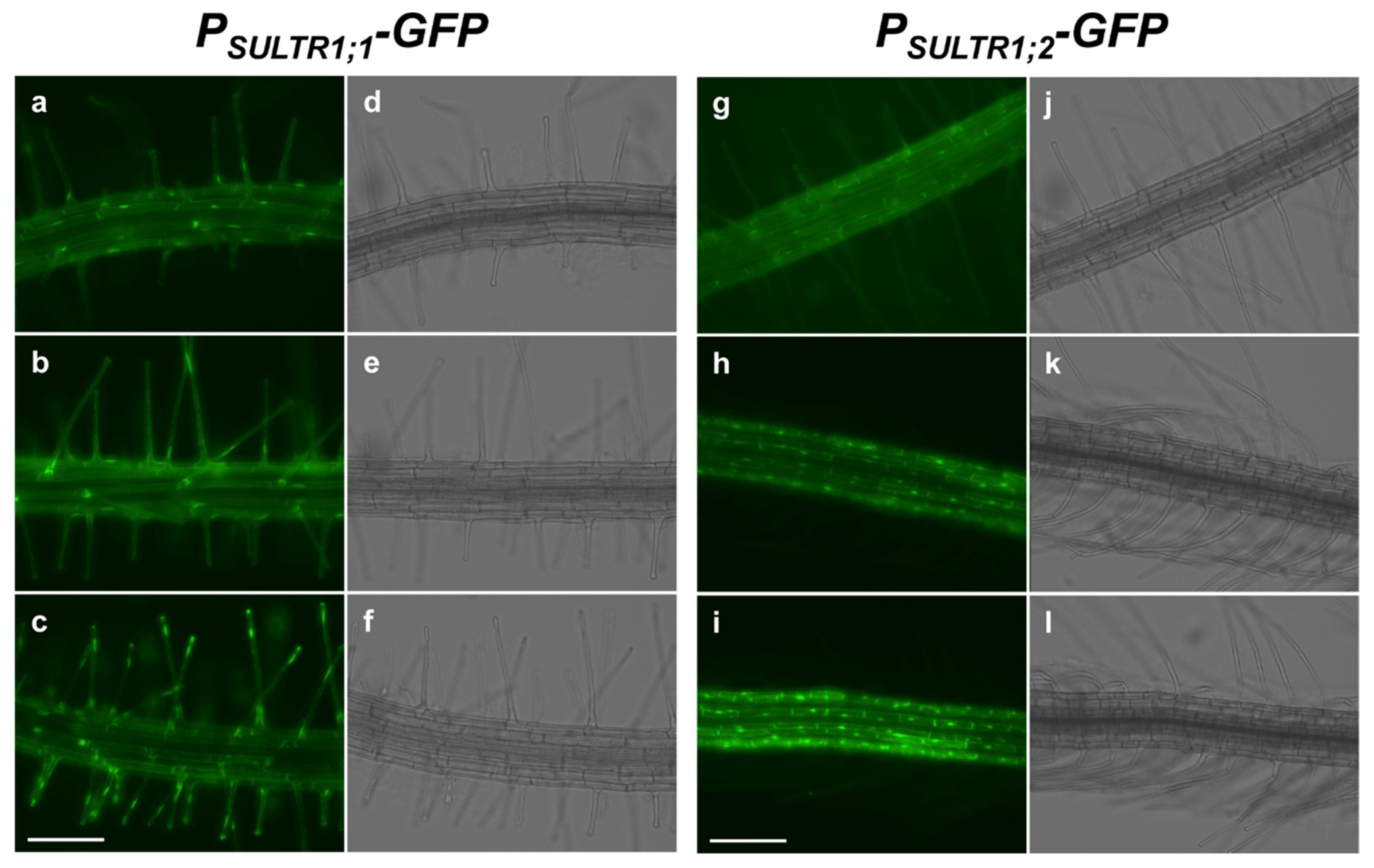
2.1. SULTR1;1 and SULTR1;2 Were Differentially Expressed in the Root Epidermal Cells under −S
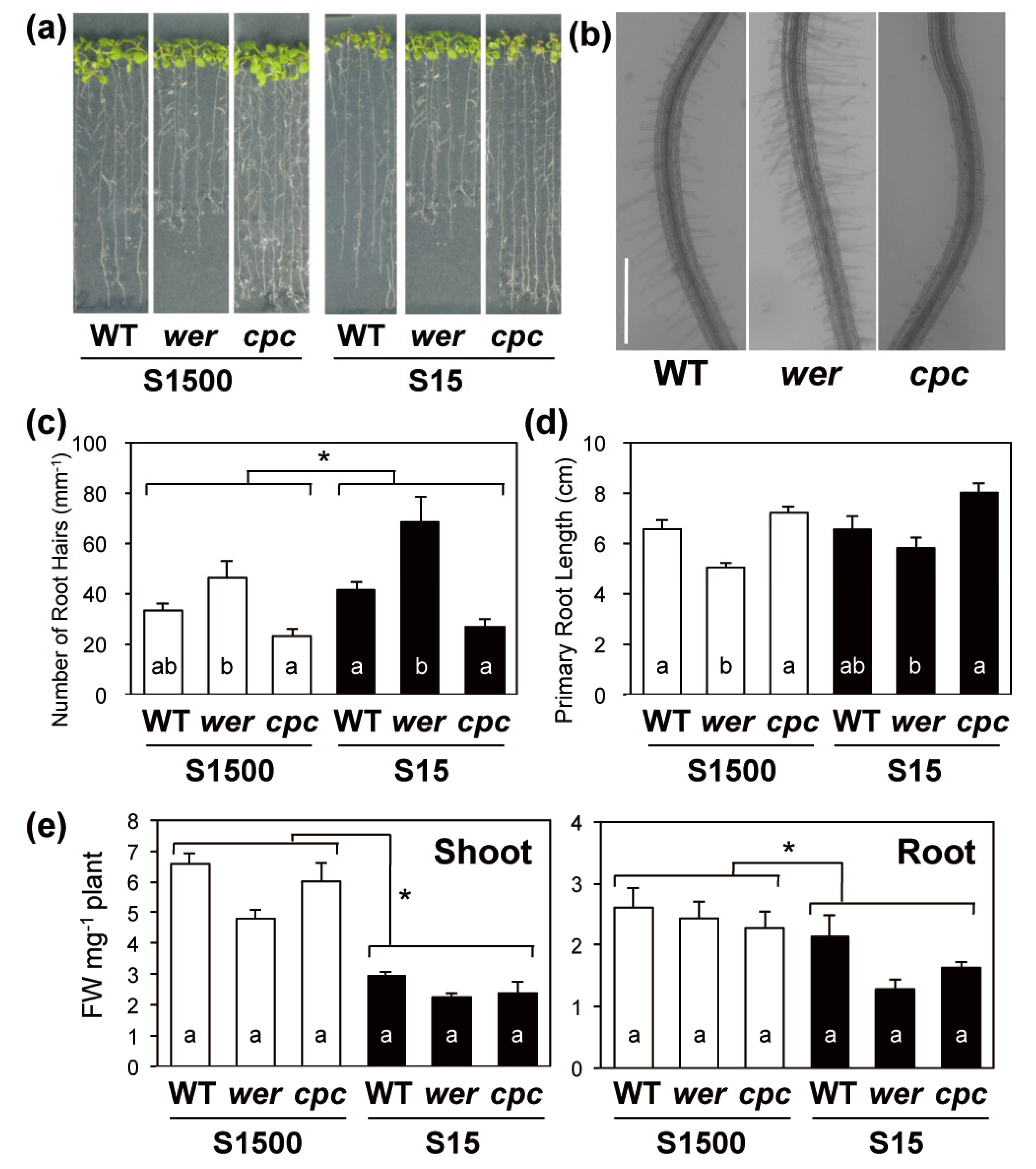
2.2. Primary Root Length and Numbers of Root Hairs Compensated for Each Other in wer and cpc
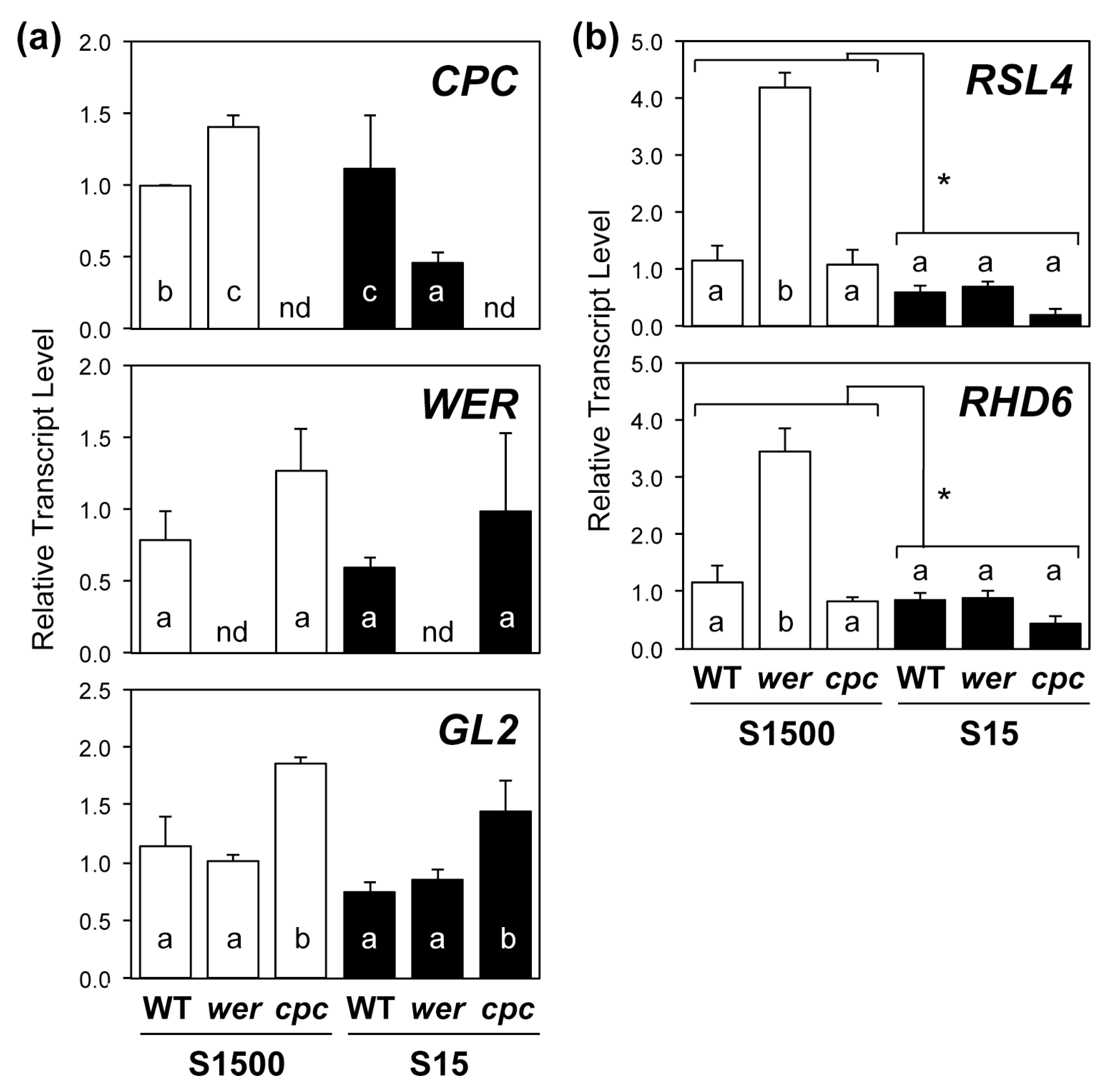
2.3. Regulatory Gene Expression in Root Hair Development Was Consistent with Root Hair Numbers under +S but Not –S Conditions
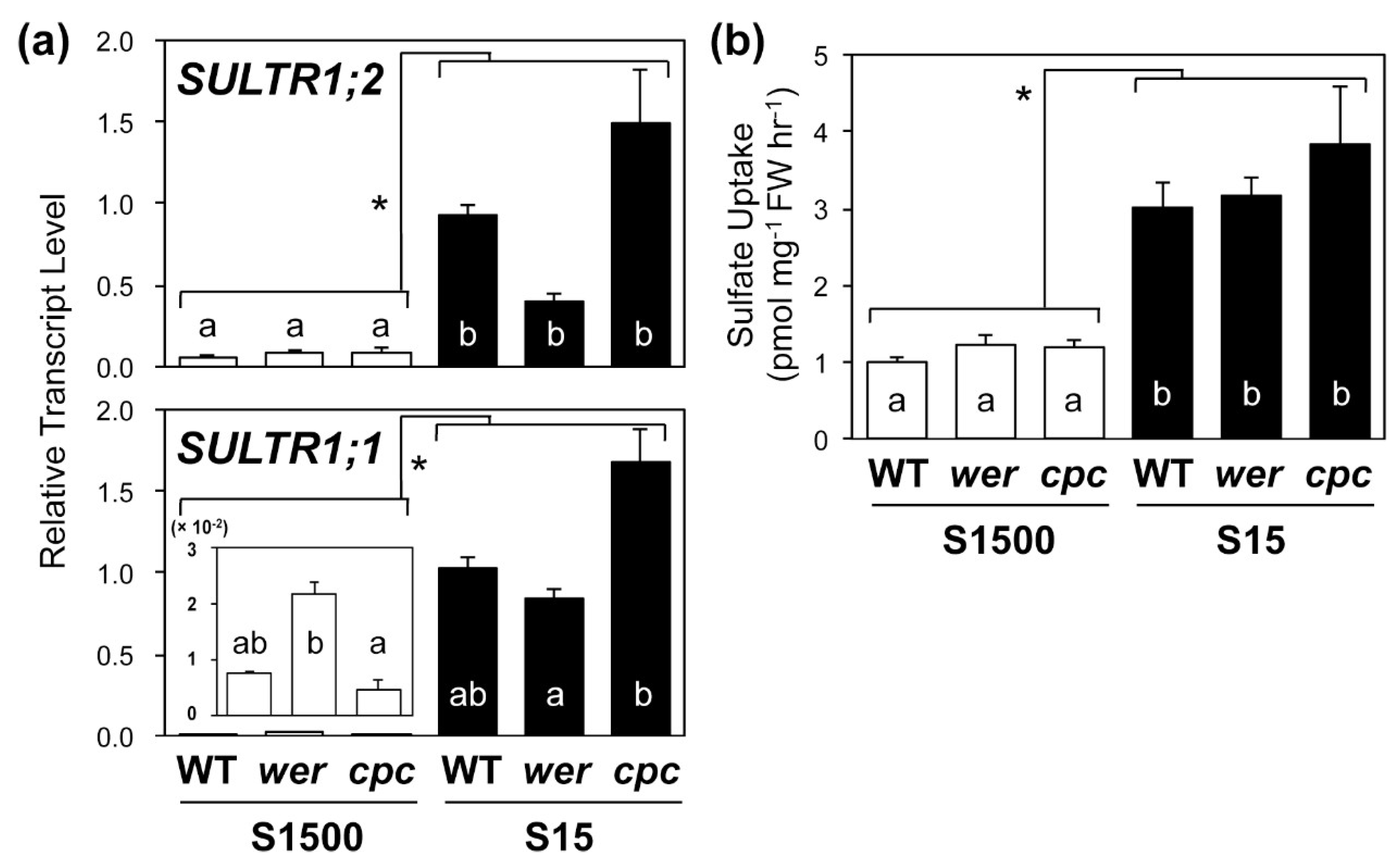
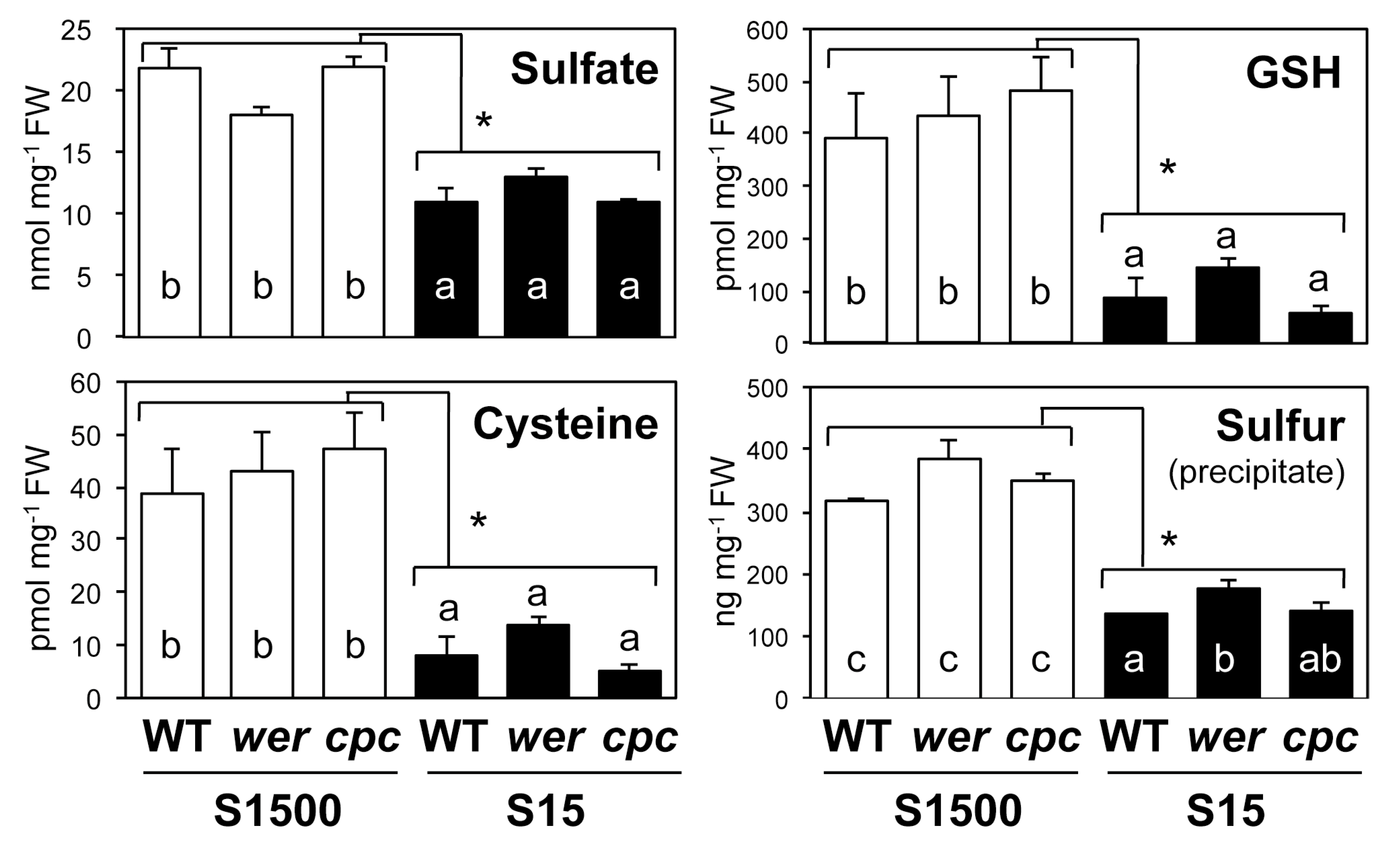
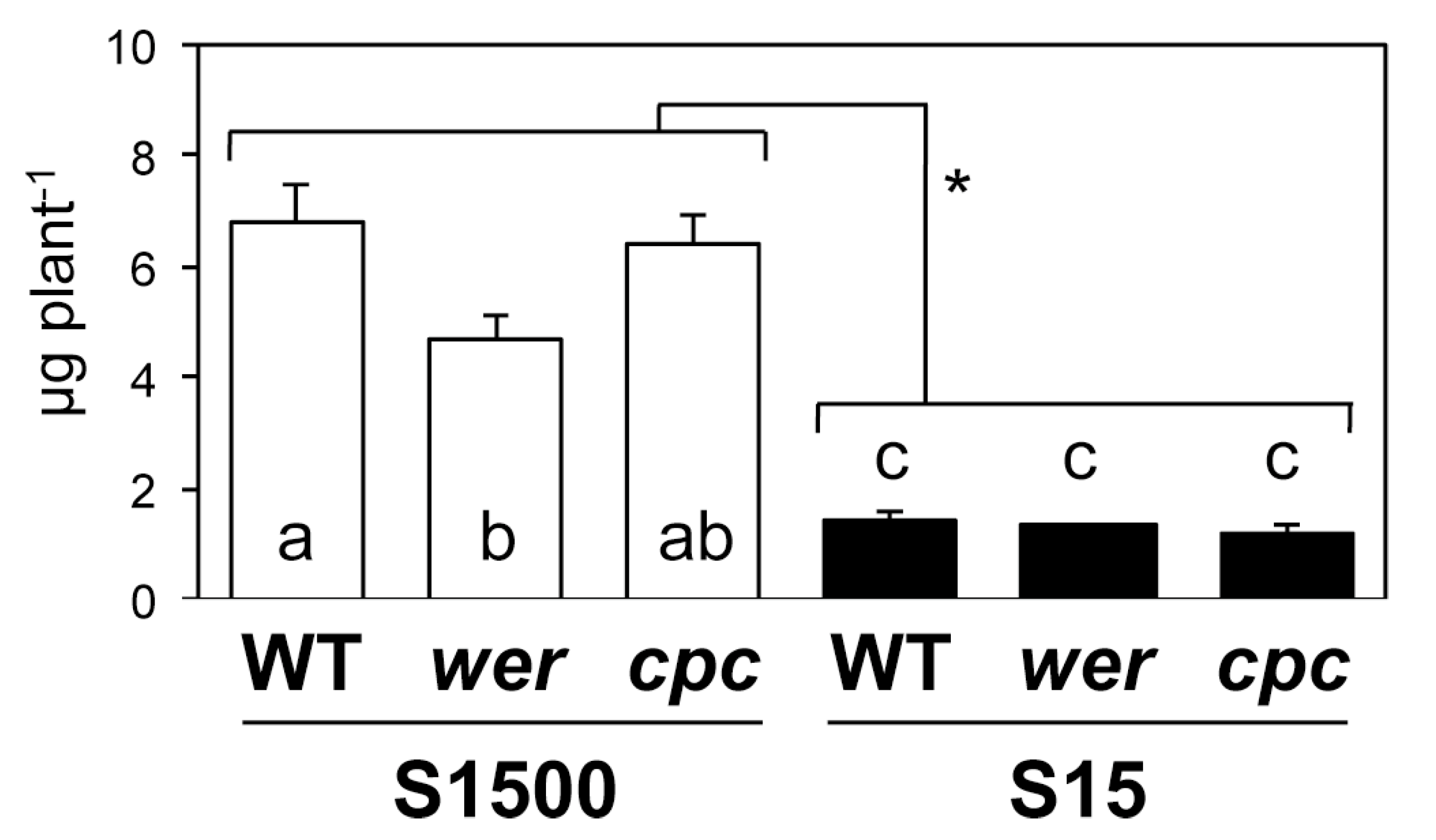
2.4. Sulfate Uptake Was Constant among WT, wer, and cpc
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Observation of GFP Fluorescence
4.3. Observation of Root Development
4.4. Quantitative Real-Time RT-PCR Analysis
4.5. Sulfate Uptake Assay
4.6. Measurement of Sulfate, Cysteine and Glutathione, and Total Sulfur Levels
4.7. Statistical Analysis
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- White, P. Ion uptake mechanisms of individual cells and roots: Short-distance transport. In Mineral Nutrition of Higher Plants, 3rd ed.; Marschner, P., Ed.; Academic Press: Oxford, UK, 2012; pp. 7–48. [Google Scholar]
- Patrick, J.W.; Tyerman, S.D.; van Bel, A.J.E. Long-Distance Transport. In Biochemistry & Molecular Biology of Plants, 2nd ed.; Buchana, B.B., Gruissem, W., Jones, R.L., Eds.; Wiley Blackwell: Oxford, UK, 2015; pp. 658–710. [Google Scholar]
- Lopez-Bucio, J.; Cruz-Ramirez, A.; Herrera-Estrella, L. The role of nutrient availability in regulating root architecture. Curr. Opin. Plant Biol. 2003, 6, 280–287. [Google Scholar] [CrossRef]
- Delhaize, E.; Schachtman, D.; Kochian, L.; Ryan, P.R. Mineral nutrient acquisition, transport, and utilization. In Biochemistry & Molecular Biology of Plants, 2nd ed.; Buchana, B.B., Gruissem, W., Jones, R.L., Eds.; Wiley Blackwell: Oxford, UK, 2015; pp. 1101–1131. [Google Scholar]
- Long, S.R.; Kahn, M.; Seefeldt, L.; Tsay, Y.F.; Kopriva, S. Nitrogen and Sulfur. In Biochemistry & Molecular Biology of Plants, 2nd ed.; Buchana, B.B., Gruissem, W., Jones, R.L., Eds.; Wiley Blackwell: Oxford, UK, 2015; pp. 746–768. [Google Scholar]
- Schiefelbein, J.; Kwak, S.-H.; Wieckowski, Y.; Barron, C.; Bruex, A. The gene regulatory network for root epidermal cell-type pattern formation in Arabidopsis. J. Exp. Bot. 2009, 60, 1515–1521. [Google Scholar] [CrossRef] [PubMed]
- Libault, M.; Brechenmacher, L.; Cheng, J.; Xu, D.; Stacey, G. Root hair systems biology. Trends Plant Sci. 2010, 15, 641–650. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, N.; Kato, M.; Tomioka, R.; Kurata, R.; Fukao, Y.; Aoyama, T.; Maeshima, M. Characteristics of a root hair-less line of Arabidopsis thaliana under physiological stresses. J. Exp. Bot. 2014, 65, 1497–1512. [Google Scholar] [CrossRef] [PubMed]
- Gilroy, S.; Jones, D.L. Through form to function: Root hair development and nutrient uptake. Trends Plant Sci. 2000, 5, 56–60. [Google Scholar] [CrossRef]
- Gahoonia, T.S.; Nielsen, N.E.; Joshi, P.A.; Jahoor, A. A root hairless barley mutant for elucidating genetic of root hairs and phosphorus uptake. Plant Soil 2001, 235, 211–219. [Google Scholar] [CrossRef]
- Schmidt, W.; Tittel, J.; Schikora, A. Role of hormones in the induction of iron deficiency responses in Arabidopsis roots. Plant Physiol. 2000, 122, 1109–1118. [Google Scholar] [CrossRef]
- Ma, J.F.; Goto, S.; Tamai, K.; Ichii, M. Role of root hairs and lateral roots in silicon uptake by rice. Plant Phys. 2001, 127, 1773–1780. [Google Scholar] [CrossRef]
- Ma, Z.; Bielenberg, D.G.; Brown, K.M.; Lynch, J.P. Regulation of root hair density by phosphorus availability in Arabidopsis thaliana. Plant Cell Environ. 2001, 24, 459–467. [Google Scholar] [CrossRef]
- Mueller, M.; Schmidt, W. Environmentally induced plasticity of root hair development in Arabidopsis. Plant Phys. 2004, 134, 409–419. [Google Scholar] [CrossRef] [PubMed]
- Yang, T.J.W.; Perry, P.J.; Ciani, S.; Pandian, S.; Schmidt, W. Manganese deficiency alters the patterning and development of root hairs in Arabidopsis. J. Exp. Bot. 2008, 59, 3453–3464. [Google Scholar] [CrossRef] [PubMed]
- Song, L.; Yu, H.; Dong, J.; Che, X.; Jiao, Y.; Liu, D. The molecular mechanism of ethylene-mediated root hair development induced by phosphate starvation. PLoS Genet. 2016, 12, e1006194. [Google Scholar] [CrossRef]
- Feng, Y.; Xu, P.; Li, B.; Li, P.; Wen, X.; An, F.; Gong, Y.; Xin, Y.; Zhu, Z.; Wang, Y.; et al. Ethylene promotes root hair growth through coordinated EIN3/EIL1 and RHD6/RSL1 activity in Arabidopsis. Proc. Natl. Acad. Sci. USA 2017, 114, 13834–13839. [Google Scholar] [CrossRef]
- Takahashi, H.; Kopriva, S.; Giordano, M.; Saito, K.; Hell, R. Sulfur assimilation in photosynthetic organisms: Molecular functions and regulations of transporters and assimilatory enzymes. Annu. Rev. Plant Biol. 2011, 62, 157–184. [Google Scholar] [CrossRef]
- Leustek, T.; Martin, M.N.; Bick, J.; Davies, J.P. Pathways and regulation of sulfur metabolism revealed through molecular genetic studies. Annu. Rev. Plant Physiol. Plant Mol. Biol. 2000, 51, 141–166. [Google Scholar] [CrossRef]
- Saito, K. Sulfur assimilatory metabolism. The long and smelling road. Plant Physiol. 2004, 136, 2443–2450. [Google Scholar] [CrossRef]
- Takahashi, H.; Watanabe-Takahashi, A.; Smith, F.W.; Blake-Kalff, M.; Hawkesford, M.J.; Saito, K. The roles of three functional sulfate transporters involved in uptake and translocation of sulfate in Arabidopsis thaliana. Plant J. 2000, 23, 171–182. [Google Scholar] [CrossRef] [PubMed]
- Vidmar, J.J.; Tagmount, A.; Cathala, N.; Touraine, B.; Davidian, J.C.E. Cloning and characterization of a root specific high-affinity sulfate transporter from Arabidopsis thaliana. FEBS Lett. 2000, 475, 65–69. [Google Scholar] [CrossRef]
- Shibagaki, N.; Rose, A.; McDermott, J.P.; Fujiwara, T.; Hayashi, H.; Yoneyama, T.; Davies, J.P. Selenate-resistant mutants of Arabidopsis thaliana identify Sultr1;2, a sulfate transporter required for efficient transport of sulfate into roots. Plant J. 2002, 29, 475–486. [Google Scholar] [CrossRef] [PubMed]
- Yoshimoto, N.; Takahashi, H.; Smith, F.W.; Yamaya, T.; Saito, K. Two distinct high-affinity sulfate transporters with different inducibilities mediate uptake of sulfate in Arabidopsis roots. Plant J. 2002, 29, 465–473. [Google Scholar] [CrossRef] [PubMed]
- Yoshimoto, N.; Inoue, E.; Watanabe-Takahashi, A.; Saito, K.; Takahashi, H. Posttranscriptional regulation of high-affinity sulfate transporters in Arabidopsis by sulfur nutrition. Plant Physiol. 2007, 145, 378–388. [Google Scholar] [CrossRef] [PubMed]
- Maruyama-Nakashita, A.; Nakamura, Y.; Watanabe-Takahashi, A.; Yamaya, T.; Takahashi, H. Induction of SULTR1;1 sulfate transporter in Arabidopsis roots involves protein phosphorylation/dephosphorylation circuit for transcriptional regulation. Plant Cell Physiol. 2004, 45, 340–345. [Google Scholar] [CrossRef]
- Maruyama-Nakashita, A.; Nakamura, Y.; Yamaya, T.; Takahashi, H. A novel regulatory pathway of sulfate uptake in Arabidopsis roots: Implication of CRE1/WOL/AHK4-mediated cytokinin-dependent regulation. Plant J. 2004, 38, 779–789. [Google Scholar] [CrossRef]
- Maruyama-Nakashita, A.; Nakamura, Y.; Watanabe, A.; Inoue, E.; Yamaya, T.; Takahashi, H. Identification of a novel cis-acting element conferring sulfur deficiency response in Arabidopsis roots. Plant J. 2005, 42, 305–314. [Google Scholar] [CrossRef] [PubMed]
- Maruyama-Nakashita, A. Metabolic changes sustain the plant life in low-sulfur environments. Curr. Opin. Plant Biol. 2017, 39, 144–151. [Google Scholar] [CrossRef]
- Maruyama-Nakashita, A.; Inoue, E.; Watanabe-Takahashi, A.; Yamaya, T.; Takahashi, H. Transcriptome profiling of sulfur-responsive genes in Arabidopsis reveals global effect on sulfur nutrition on multiple metabolic pathways. Plant Physiol. 2003, 132, 597–605. [Google Scholar] [CrossRef]
- Rouached, H.; Wirtz, M.; Alary, R.; Hell, R.; Arpat, A.B.; Davidian, J.C.; Fourcroy, P.; Berthomieu, P. Differential regulation of the expression of two high-affinity sulfate transporters, SULTR1.1 and SULTR1.2, in Arabidopsis. Plant Physiol. 2008, 147, 897–911. [Google Scholar] [CrossRef]
- Grebe, M. The patterning of epidermal hairs in Arabidopsis—updated. Curr. Opin. Plant Biol. 2012, 15, 31–37. [Google Scholar] [CrossRef]
- Salazar-Henao, J.E.; Veélez-Bermuédez, I.C.; Schmidt, W. The regulation and plasticity of root hair patterning and morphogenesis. Development 2016, 143, 1848–1858. [Google Scholar] [CrossRef]
- Ishida, T.; Kurata, T.; Okada, K.; Wada, T. A genetic regulatory network in the development of trichomes and root hairs. Annu. Rev. Plant Biol. 2008, 59, 365–386. [Google Scholar] [CrossRef] [PubMed]
- Shibata, M.; Sugimoto, K. A gene regulatory network for root hair development. J. Plant Res. 2019. [Google Scholar] [CrossRef] [PubMed]
- Walker, A.R.; Davison, P.A.; Bolognesi-Winfield, A.C.; James, C.M.; Srinivasan, N.; Blundell, T.L.; Esch, J.J.; Marks, M.D.; Gray, J.C. The TRANSPARENT TESTA GLABRA1 Locus, which regulates trichome differentiation and anthocyanin biosynthesis in Arabidopsis, encodes a WD40 repeat protein. Plant Cell 1999, 11, 1337–1349. [Google Scholar] [CrossRef]
- Payne, C.T.; Zhang, F.; Lloyd, A.M. GL3 encodes a bHLH protein that regulates trichome development in Arabidopsis through interaction with GL1 and TTG1. Genetics 2000, 156, 1349–1362. [Google Scholar]
- Bernhardt, C.; Lee, M.M.; Gonzalez, A.; Zhang, F.; Lloyd, A.; Schiefelbein, J. The bHLH genes GLABRA3 (GL3) and ENHANCER OF GLABRA3 (EGL3) specify epidermal cell fate in the Arabidopsis root. Development 2003, 130, 6431–6439. [Google Scholar] [CrossRef]
- Lee, M.M.; Schiefelbein, J. WERWOLF, a MYB-related protein in Arabidopsis, is a position-dependent regulator of epidermal cell patterning. Cell 1999, 99, 473–483. [Google Scholar] [CrossRef]
- Rerie, W.G.; Feldmann, K.A.; Marks, M.D. The GLABRA2 gene encodes a homeo domain protein required for normal trichome development in Arabidopsis. Genes Dev. 1994, 8, 1388–1399. [Google Scholar] [CrossRef] [PubMed]
- Wada, T.; Tachibana, T.; Shimura, Y.; Okada, K. Epidermal cell differentiation in Arabidopsis determined by a Myb-homolog, CPC. Science 1997, 277, 1113–1116. [Google Scholar] [CrossRef] [PubMed]
- Schellmann, S.; Schnittger, A.; Kirik, V.; Wada, T.; Okada, K.; Beermann, A.; Thumfahrt, J.; Jürgens, G.; Hülskamp, M. TRIPTYCHON and CAPRICE mediate lateral inhibition during trichome and root hair patterning in Arabidopsis. EMBO J. 2002, 21, 5036–5046. [Google Scholar] [CrossRef] [PubMed]
- Kirik, V.; Simon, M.; Huelskamp, M.; Schiefelbein, J. The ENHANCER OF TRY AND CPC1 gene acts redundantly with TRIPTYCHON and CAPRICE in trichome and root hair cell patterning in Arabidopsis. Dev. Biol. 2004, 268, 506–513. [Google Scholar] [CrossRef]
- Tominaga, R.; Iwata, M.; Sano, R.; Okada, K.; Wada, T. Arabidopsis CAPRICE-like Myb 3 (CPL3) controls endoreduplication and flowering development in addition to trichome and root-hair formation. Development 2008, 135, 1335–1345. [Google Scholar] [CrossRef]
- Tominaga, R.; Iwata, M.; Okada, K.; Wada, T. Functional analysis of the epidermal-specific MYB genes CAPRICE and WEREWOLF in Arabidopsis. Plant Cell 2007, 19, 2264–2277. [Google Scholar] [CrossRef] [PubMed]
- Song, S.K.; Ryu, K.H.; Kang, Y.H.; Song, J.H.; Cho, Y.H.; Yoo, S.D.; Schiefelbein, J.; Lee, M.M. Cell fate in the Arabidopsis root epidermis is determined by competition between WEREWOLF and CAPRICE. Plant Physiol. 2011, 157, 1196–1208. [Google Scholar] [CrossRef]
- Masucci, J.D.; Schiefelbein, J.W. The rhd6 mutation of Arabidopsis thaliana alters root-hair initiation through an auxin- and ethylene-associated process. Plant Physiol. 1994, 106, 1335–1346. [Google Scholar] [CrossRef]
- Menand, B.; Yi, K.; Jouannic, S.; Hoffmann, L.; Ryan, E.; Linstead, P.; Schaefer, D.G.; Dolan, L. An ancient mechanism controls the development of cells with a rooting function in land plants. Science 2007, 316, 1477–1480. [Google Scholar] [CrossRef] [PubMed]
- Lin, Q.; Ohashi, Y.; Kato, M.; Tsuge, T.; Gu, H.; Qu, L.-J.; Aoyama, T. GLABRA2 directly suppresses basic helix-loop-helix transcription factor genes with diverse functions in root hair development. Plant Cell 2015, 27, 2894–2906. [Google Scholar] [CrossRef]
- Yi, K.; Menand, B.; Bell, E.; Dolan, L. A basic helix-loop-helix transcription factor controls cell growth and size in root hairs. Nat. Genet. 2010, 42, 264–267. [Google Scholar] [CrossRef]
- Rubio, V.; Linhares, F.; Solano, R.; Martin, A.C.; Iglesias, J.; Leyva, A.; Paz-Ares, J. A conserved MYB transcription factor involved in phosphate starvation signaling both in vascular plants and in unicellular algae. Genes Dev. 2001, 15, 2122–2133. [Google Scholar] [CrossRef] [PubMed]
- Bustos, R.; Castrillo, G.; Linhares, F.; Puga, M.I.; Rubio, V.; Pérez-Pérez, J.; Solano, R.; Leyva, A.; Paz-Ares, J. A central regulatory system largely controls transcriptional activation and repression responses to phosphate starvation in Arabidopsis. PLoS Genet. 2010, 6, e1001102. [Google Scholar] [CrossRef]
- Bruex, A.; Kainkaryam, R.M.; Wieckowski, Y.; Kang, Y.H.; Bernhardt, C.; Xia, Y.; Zheng, X.; Wang, J.Y.; Lee, M.M.; Benfey, P.; et al. A gene regulatory network for root epidermis cell differentiation in Arabidopsis. PLoS Genet. 2012, 8, e1002446. [Google Scholar] [CrossRef] [PubMed]
- Simon, M.; Bruex, A.; Kainkaryam, R.M.; Zheng, X.; Huang, L.; Woolf, P.J.; Schiefelbein, J. Tissue-specific profiling reveals transcriptome alterations in Arabidopsis mutants lacking morphological phenotypes. Plant Cell 2013, 25, 3175–3185. [Google Scholar] [CrossRef] [PubMed]
- Maruyama-Nakashita, A.; Nakamura, Y.; Tohge, T.; Saito, K.; Takahashi, H. Arabidopsis SLIM1 is a central transcriptional regulator of plant sulfur response and metabolism. Plant Cell 2006, 18, 3235–3251. [Google Scholar] [CrossRef]
- Maruyama-Nakashita, A.; Watanabe-Takahashi, A.; Inoue, E.; Yamaya, T.; Saito, K.; Takahashi, H. Sulfur-responsive elements in the 3’-nontranscribed intergenic region are essential for the induction of SULFATE TRANSPORTER 2;1 gene expression in Arabidopsis roots under sulfur deficiency. Plant Cell 2015, 27, 1279–1296. [Google Scholar] [CrossRef] [PubMed]
- Lan, P.; Li, W.; Lin, W.-D.; Santi, S.; Schmidt, W. Mapping gene activity of Arabidopsis root hairs. Genome Biol. 2013, 14, R67. [Google Scholar] [CrossRef]
- Tominaga-Wada, R.; Wada, T. Relationship between root hair formation and primary root length in Arabidopsis thaliana. Res. Rev. Biosci. 2017, 12, 128. [Google Scholar]
- Negi, S.; Ivanchenko, M.G.; Muday, G.K. Ethylene regulates lateral root formation and auxin transport in Arabidopsis thaliana. Plant J. 2008, 55, 175–187. [Google Scholar] [CrossRef]
- Fujiwara, T.; Hirai, M.Y.; Chino, M.; Komeda, Y.; Naito, S. Effects of sulfur nutrition on expression of the soybean seed storage protein genes in transgenic petunia. Plant Physiol. 1992, 99, 263–268. [Google Scholar] [CrossRef] [PubMed]
- Hirai, M.Y.; Fujiwara, T.; Chino, M.; Naito, S. Effects of sulfate concentrations on the expression of a soybean seed storage protein gene and its reversibility in transgenic Arabidopsis thaliana. Plant Cell Physiol. 1995, 36, 1331–1339. [Google Scholar] [PubMed]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH image to ImageJ: 25 years of image analysis. Nat. Methods 2012, 9, 671–675. [Google Scholar] [CrossRef] [PubMed]
- Hartig, S.M. Basic image analysis and manipulation in ImageJ. Curr. Protoc. Mol. Biol. 2013, 102, 14–15. [Google Scholar] [CrossRef]
- Hayashi, N.; Tetsumura, T.; Sawa, S.; Wada, T.; Tominaga-Wada, R. CLE14 peptide signaling in Arabidopsis root hair cell fate determination. Plant Biotechnol. 2018, 35, 17–22. [Google Scholar] [CrossRef]
- Kataoka, T.; Hayashi, N.; Yamaya, T.; Takahashi, H. Root-to-shoot transport of sulfate in Arabidopsis: Evidence for the role of SULTR3; 5 as a component of low-affinity sulfate transport system in the root vasculature. Plant Physiol. 2004, 136, 4198–4204. [Google Scholar] [CrossRef] [PubMed]
- Yoshimoto, N.; Kataoka, T.; Maruyama-Nakashita, A.; Takahashi, H. Measurement of uptake and root-to-shoot distribution of sulfate in Arabidopsis seedlings. Bio-Protocol 2016, 6, e1700. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kimura, Y.; Ushiwatari, T.; Suyama, A.; Tominaga-Wada, R.; Wada, T.; Maruyama-Nakashita, A. Contribution of Root Hair Development to Sulfate Uptake in Arabidopsis. Plants 2019, 8, 106. https://doi.org/10.3390/plants8040106
Kimura Y, Ushiwatari T, Suyama A, Tominaga-Wada R, Wada T, Maruyama-Nakashita A. Contribution of Root Hair Development to Sulfate Uptake in Arabidopsis. Plants. 2019; 8(4):106. https://doi.org/10.3390/plants8040106
Chicago/Turabian StyleKimura, Yuki, Tsukasa Ushiwatari, Akiko Suyama, Rumi Tominaga-Wada, Takuji Wada, and Akiko Maruyama-Nakashita. 2019. "Contribution of Root Hair Development to Sulfate Uptake in Arabidopsis" Plants 8, no. 4: 106. https://doi.org/10.3390/plants8040106
APA StyleKimura, Y., Ushiwatari, T., Suyama, A., Tominaga-Wada, R., Wada, T., & Maruyama-Nakashita, A. (2019). Contribution of Root Hair Development to Sulfate Uptake in Arabidopsis. Plants, 8(4), 106. https://doi.org/10.3390/plants8040106




