Segregation Distortion Observed in the Progeny of Crosses Between Oryza sativa and O. meridionalis Caused by Abortion During Seed Development
Abstract
1. Introduction
2. Results
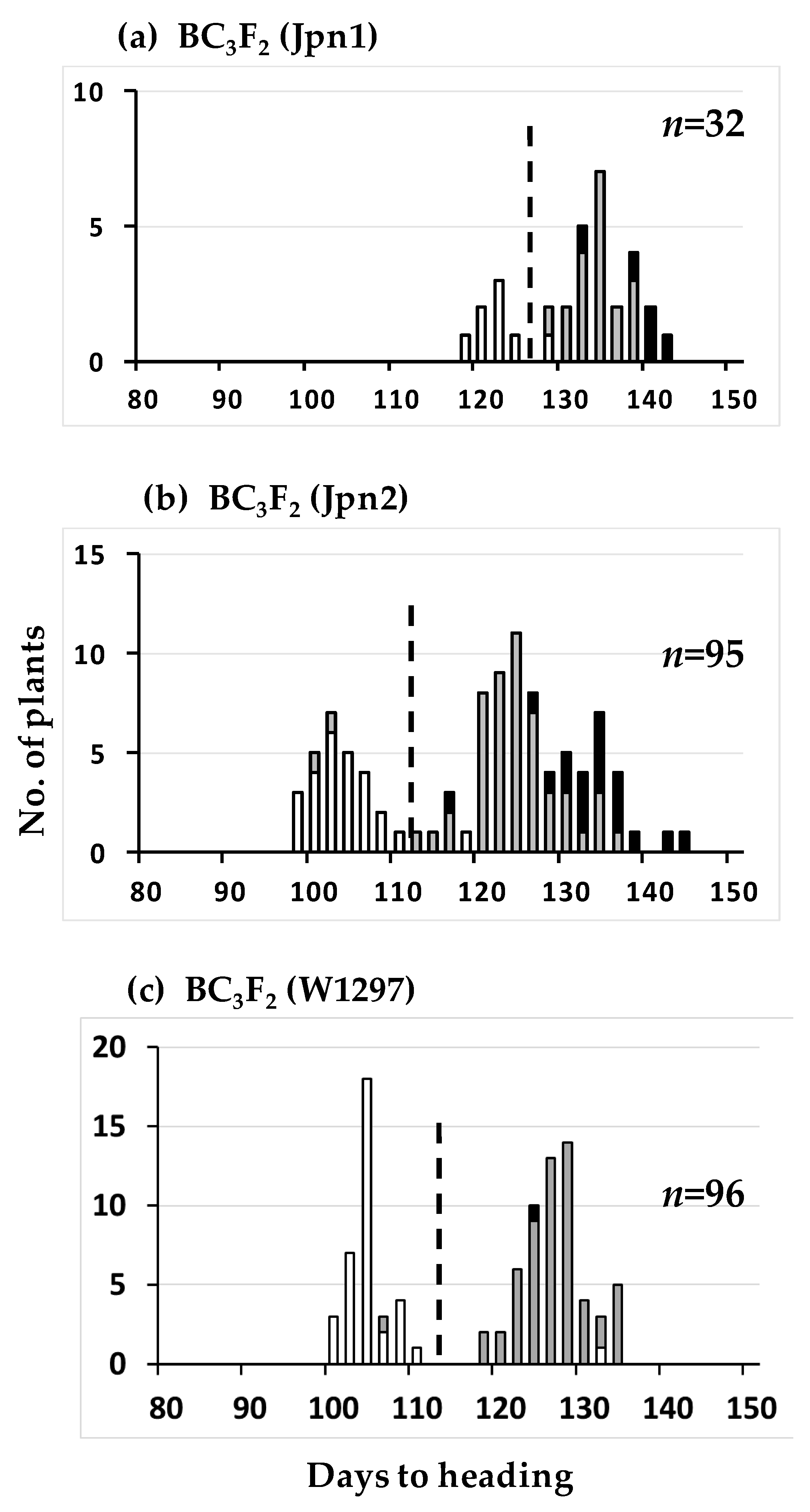
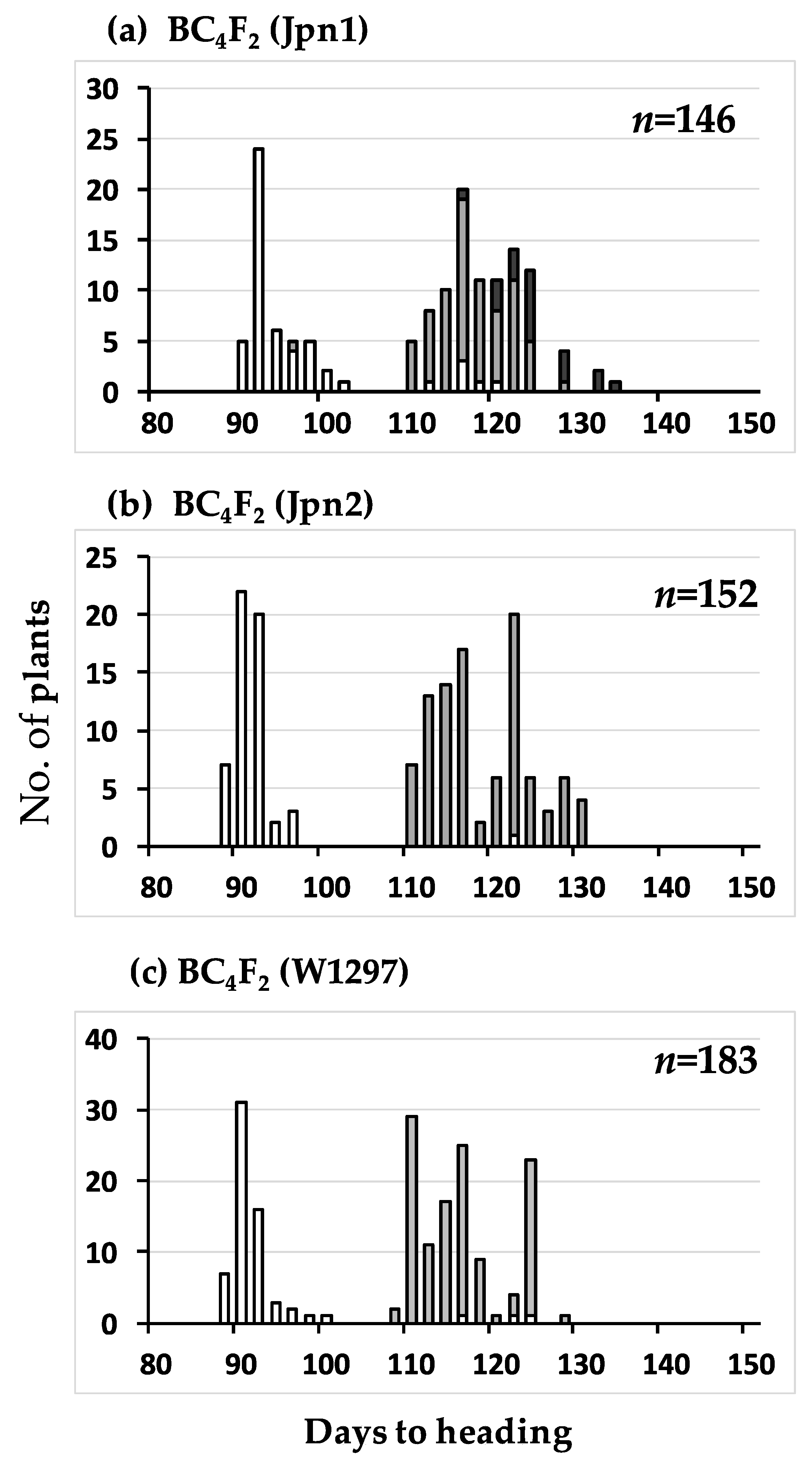
2.1. Mapping of Photoperiod Sensitivity Gene
2.2. Mapping of Segregation Distortion Gene
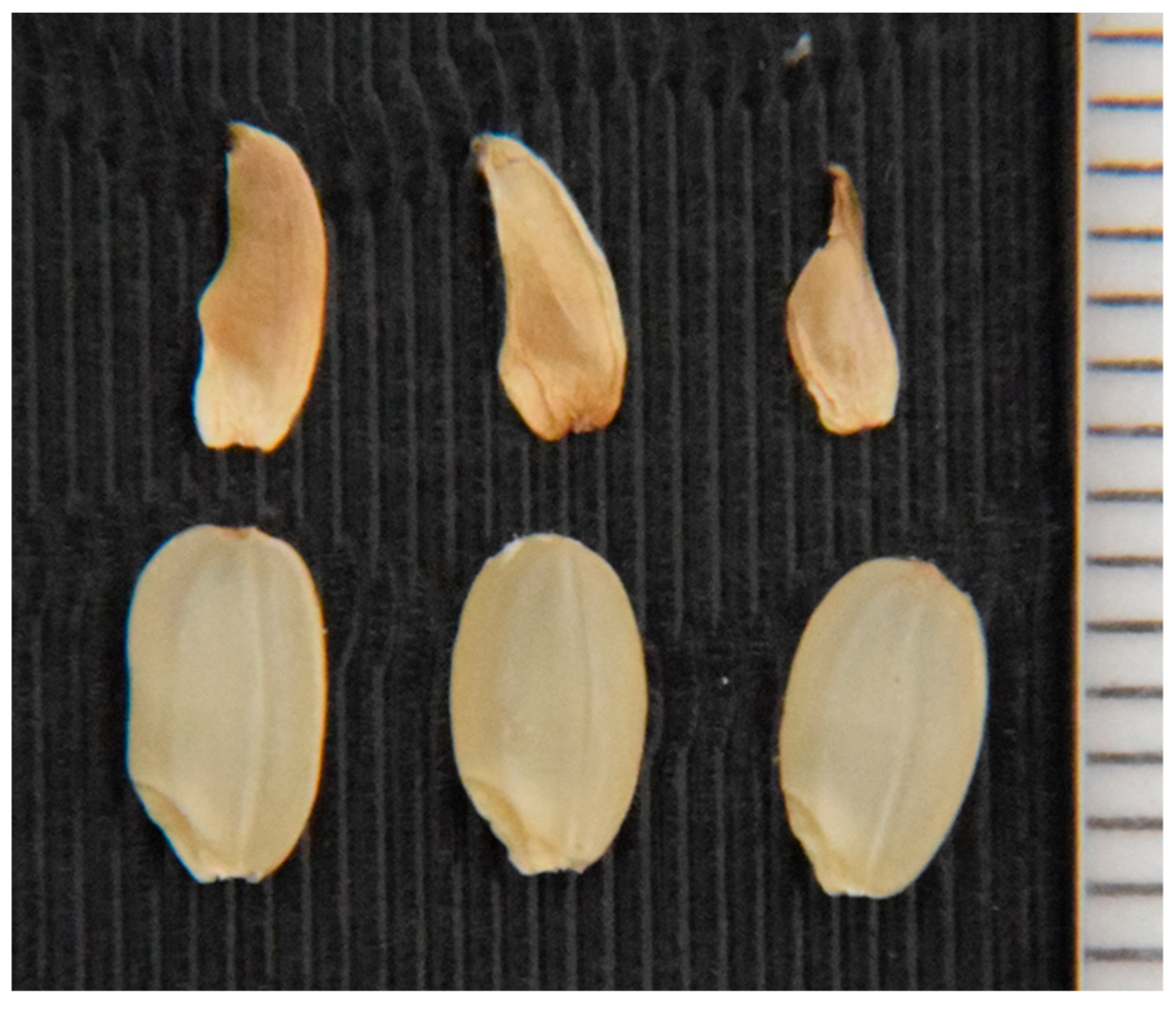
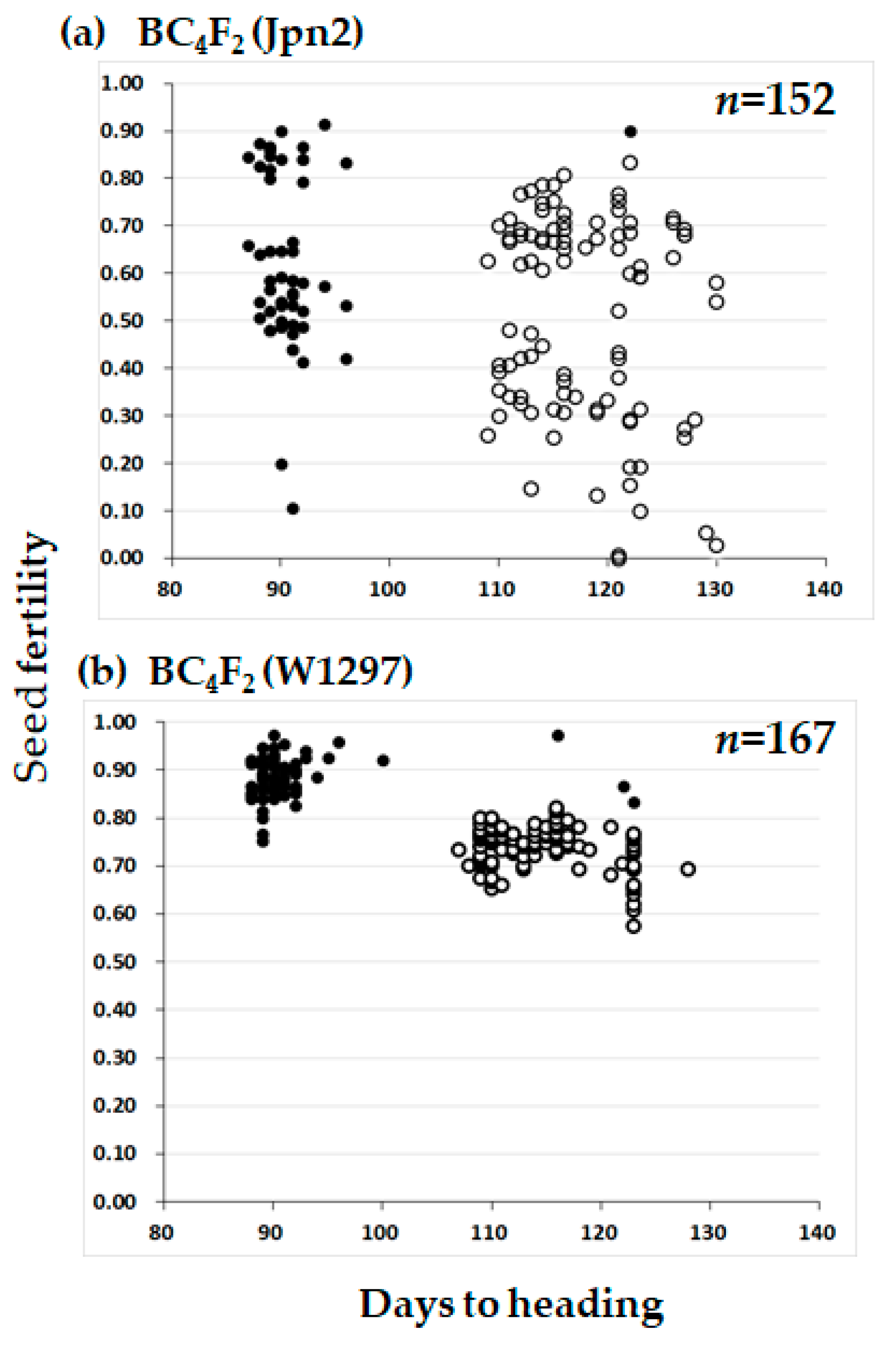
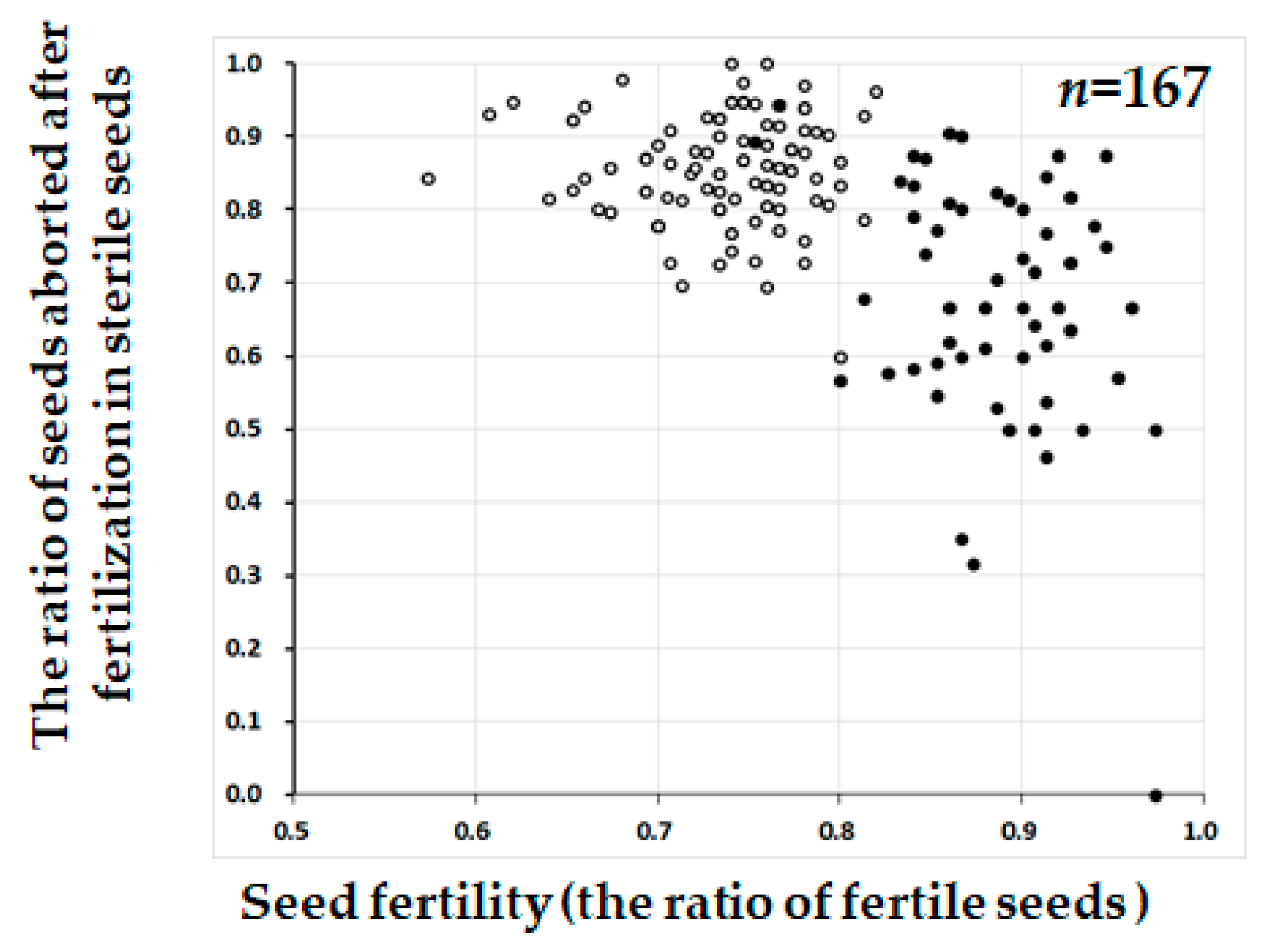
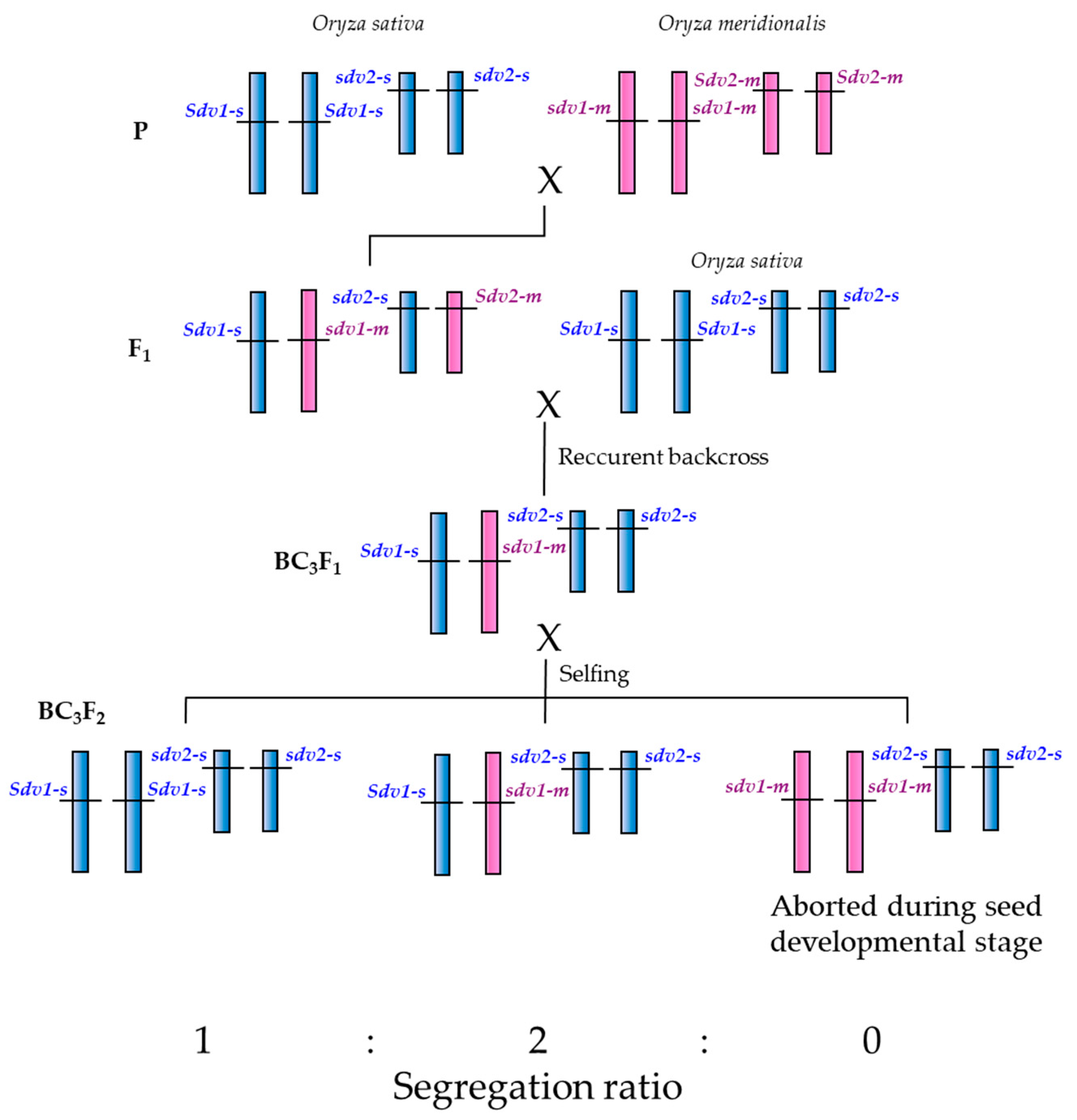
2.3. Segregation Distortion Caused by Abortion During Seed Development
3. Discussion
4. Materials and Methods
4.1. Plant Material
4.2. Trait Evaluation
4.3. DNA Analysis
4.4. DNA Markers
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Henry, R.J.; Rice, N.; Waters, D.L.E.; Kasem, S.; Ishikawa, R.; Hao, Y.; Dillon, S.; Crayn, D.; Wing, R.; Vaughan, D. Australian Oryza: Utility and conservation. Rice 2010, 3, 235–241. [Google Scholar] [CrossRef]
- Huang, X.; Kurata, N.; Wei, X.; Wang, Z.-X.; Wang, A.; Zhao, Q.; Zhao, Y.; Liu, K.; Lu, H.; Li, W.; et al. A map of rice genome variation reveals the origin of cultivated rice. Nature 2012, 490, 497–501. [Google Scholar] [CrossRef] [PubMed]
- Ohtsubo, H.; Cheng, C.; Ohsawa, I.; Tsuchimoto, S.; Ohtsubo, E. Rice retroposon p-SINE1 and origin of cultivated rice. Breed. Sci. 2004, 54, 1–11. [Google Scholar] [CrossRef]
- Cheng, C.; Tsuchimoto, S.; Ohtsubo, H.; Ohtsubo, E. Evolutionary relationships among rice species with AA genome based on SINE insertion analysis. Genes Genet. Syst. 2002, 77, 323–334. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.-H.; Kurata, N.; Akimoto, M.; Ohtsubo, H.; Ohtsubo, E. Identification and characterization of Australian wild rice strains of Oryza meridionalis and Oryza rufipogon by SINE insertion polymorphism. Genes Genet. Syst. 2005, 80, 129–134. [Google Scholar] [CrossRef][Green Version]
- Zhang, Q.-J.; Zhu, T.; Xia, E.-H.; Shi, C.; Liu, Y.-L.; Zhang, Y.; Liu, Y.; Jiang, W.-K.; Zhao, Y.-J.; Mao, S.-Y.; et al. Rapid diversification of five Oryza AA genomes associated with rice adaptation. Proc. Natl. Acad. Sci. USA 2014, 111, E4954–E4962. [Google Scholar] [CrossRef] [PubMed]
- Stein, J.C.; Yu, Y.; Copetti, D.; Zwickl, D.J.; Zhang, L.; Zhang, C.; Chougule, K.; Gao, D.; Iwata, A.; Goicoechea, J.L.; et al. Genomes of 13 domesticated and wild rice relatives highlight genetic conservation, turnover and innovation across the genus Oryza. Nat. Genet. 2018, 50, 285–296. [Google Scholar] [CrossRef] [PubMed]
- Naredo, M.E.B.; Juliano, A.B.; Lu, B.-R.; Jackson, M.T. Hybridization of AA genome rice species from Asia and Australia I. Crosses and development of hybrids. Genet. Resour. Crop Evol. 1997, 44, 17–23. [Google Scholar] [CrossRef]
- Naredo, M.E.B.; Juliano, A.B.; Lu, B.-R.; Jackson, M.T. Taxonomic status of Oryza glumaepatula Steud. II. Hybridization between New World diploids and AA genome species from Asia and Australia. Genet. Resour. Crop Evol. 1998, 45, 205–214. [Google Scholar] [CrossRef]
- Shim, R.A.; Angeles, E.R.; Ashikari, M.; Takashi, T. Development and evaluation of Oryza glaberrima Steud. chromosome segment substitution lines (CSSLs) in the background of O. sativa L. cv. Koshihikari. Breed. Sci. 2010, 60, 613–619. [Google Scholar] [CrossRef]
- Hirabayashi, H.; Sato, H.; Nonoue, Y.; Kuno-Takemoto, Y.; Takeuchi, Y.; Kato, H.; Nemoto, H.; Ogawa, T.; Yano, M.; Imbe, T.; et al. Development of introgression lines derived from Oryza rufipogon and O. glumaepatula in the genetic background of japonica cultivated rice (O. sativa L.) and evaluation of resistance to rice blast. Breed. Sci. 2010, 60, 604–612. [Google Scholar] [CrossRef]
- Yoshimura, A.; Nagayama, H.; Sobrizal; Kurakazu, T.; Sanchez, P.L.; Doi, K.; Yamagata, Y.; Yasui, H. Introgression lines of rice (Oryza sativa L.) carrying a donor genome from the wild species, O. glumaepatula Steud. and O. meridionalis Ng. Breed. Sci. 2010, 60, 597–603. [Google Scholar] [CrossRef]
- Arbelaez, J.D.; Moreno, L.T.; Singh, N.; Tung, C.-W.; Maron, L.G.; Ospina, Y.; Martinez, C.P.; Grenier, C.; Lorieux, M.; McCouch, S. Development and GBS-genotyping of introgression lines (ILs) using two wild species of rice, O. meridionalis and O. rufipogon, in a common recurrent parent, O. sativa cv. Curinga. Mol. Breed. 2015, 35, 81. [Google Scholar] [CrossRef] [PubMed]
- Yano, M.; Katayose, Y.; Ashikari, M.; Yamanouchi, U.; Monna, L.; Fuse, T.; Baba, T.; Yamamoto, K.; Umehara, Y.; Nagamura, Y.; et al. Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the arabidopsis flowering time gene CONSTANS. Plant Cell 2000, 12, 2473–2483. [Google Scholar] [CrossRef]
- Doi, K.; Izawa, T.; Fuse, T.; Yamanouchi, U.; Kubo, T.; Shimatani, Z.; Yano, M.; Yoshimura, A. Ehd1, a B-type response regulator in rice, confers short-day promotion of flowering and controls FT-like gene expression independently of Hd1. Genes Dev. 2004, 18, 926–936. [Google Scholar] [CrossRef] [PubMed]
- Ichitani, K.; Okumoto, Y.; Tanisaka, T. photoperiod sensitivity gene of Se-1 locus found in photoperiod insensitive rice cultivars of the northern limit region of rice cultivation. Jpn. J. Breed. 1997, 47, 145–152. [Google Scholar] [CrossRef][Green Version]
- Ichitani, K.; Okumoto, Y.; Tanisaka, T. Genetic analysis of the rice cultivar Kasalath with special reference to two photoperiod sensitivity loci, E1 and Se-1. Jpn. J. Breed. 1998, 48, 51–57. [Google Scholar] [CrossRef][Green Version]
- Inoue, H.; Nishida, H.; Okumoto, Y.; Tanisaka, T. Identification of an early heading time gene found in the taiwanese rice cultivar Taichung 65. Jpn. J. Breed. 1998, 48, 103–108. [Google Scholar] [CrossRef]
- Temnykh, S.; Park, W.D.; Ayres, N.; Cartinhour, S.; Hauck, N.; Lipovich, L.; Cho, Y.G.; Ishii, T.; McCouch, S.R. Mapping and genome organization of microsatellite sequences in rice (Oryza sativa L.). Theor. Appl. Genet. 2000, 100, 697–712. [Google Scholar] [CrossRef]
- McCouch, S.R.; Teytelman, L.; Xu, Y.; Lobos, K.B.; Clare, K.; Walton, M.; Fu, B.; Maghirang, R.; Li, Z.; Xing, Y.; et al. Development and Mapping of 2240 New SSR Markers for Rice ( Oryza sativa L.). DNA Res. 2002, 9, 199–207. [Google Scholar] [CrossRef]
- Sano, Y. Genetic comparisons of chromosome 6 between wild and cultivated rice. Jpn. J. Breed. 1992, 42, 561–572. [Google Scholar] [CrossRef]
- Matsubara, K.; Khin-Thidar; Sano, Y. A gene block causing cross-incompatibility hidden in wild and cultivated rice. Genetics 2003, 165, 343–352. [Google Scholar] [PubMed]
- Koide, Y.; Ogino, A.; Yoshikawa, T.; Kitashima, Y.; Saito, N.; Kanaoka, Y.; Onishi, K.; Yoshitake, Y.; Tsukiyama, T.; Saito, H.; et al. Lineage-specific gene acquisition or loss is involved in interspecific hybrid sterility in rice. Proc. Natl. Acad. Sci. USA 2018, 115, E1955–E1962. [Google Scholar] [CrossRef] [PubMed]
- Yanagihara, S.; McCouch, S.R.; Ishikawa, K.; Ogi, Y.; Maruyama, K.; Ikehashi, H. Molecular analysis of the inheritance of the S-5 locus, conferring wide compatibility in Indica/Japonica hybrids of rice (O. sativa L.). Theoret. Appl. Genet. 1995, 90, 182–188. [Google Scholar] [CrossRef]
- Liu, Y.S.; Zhu, L.H.; Sun, J.S.; Chen, Y. Mapping QTLs for defective female gametophyte development in an inter-subspecific cross in Oryza Sativa L. Theor. Appl. Genet. 2001, 102, 1243–1251. [Google Scholar] [CrossRef]
- Yang, J.; Zhao, X.; Cheng, K.; Du, H.; Ouyang, Y.; Chen, J.; Qiu, S.; Huang, J.; Jiang, Y.; Jiang, L.; et al. A killer-protector system regulates both hybrid sterility and segregation distortion in rice. Science 2012, 337, 1336–1340. [Google Scholar] [CrossRef]
- Long, Y.; Zhao, L.; Niu, B.; Su, J.; Wu, H.; Chen, Y.; Zhang, Q.; Guo, J.; Zhuang, C.; Mei, M.; et al. Hybrid male sterility in rice controlled by interaction between divergent alleles of two adjacent genes. Proc. Natl. Acad. Sci. USA 2008, 105, 18871–18876. [Google Scholar] [CrossRef]
- McCouch, S.R.; CGSNL (Committee on Gene Symbolization, N. and L., Rice Genetics Cooperative). Gene Nomenclature System for Rice. Rice 2008, 1, 72–84. [Google Scholar] [CrossRef]
- Juliano, A.B.; Naredo, M.E.B.; Lu, B.-R.; Jackson, M.T. Genetic differentiation in Oryza meridionalis Ng based on molecular and crossability analyses. Genet. Resour. Crop Evol. 2005, 52, 435–445. [Google Scholar] [CrossRef]
- Yamagata, Y.; Yamamoto, E.; Aya, K.; Win, K.T.; Doi, K.; Sobrizal; Ito, T.; Kanamori, H.; Wu, J.; Matsumoto, T.; et al. Mitochondrial gene in the nuclear genome induces reproductive barrier in rice. Proc. Natl. Acad. Sci. USA 2010, 107, 1494–1499. [Google Scholar] [CrossRef]
- Nguyen, G.N.; Yamagata, Y.; Shigematsu, Y.; Watanabe, M.; Miyazaki, Y.; Doi, K.; Tashiro, K.; Kuhara, S.; Kanamori, H.; Wu, J.; et al. Duplication and loss of function of genes encoding RNA polymerase III subunit C4 causes hybrid incompatibility in rice. G3 Genes Genom. Genet. 2017, 7, 2565–2575. [Google Scholar] [CrossRef] [PubMed]
- Ichitani, K.; Takemoto, Y.; Iiyama, K.; Taura, S.; Sato, M. Chromosomal location of HCA1 and HCA2, hybrid chlorosis genes in rice. Int. J. Plant Genom. 2012, 2012. [Google Scholar] [CrossRef] [PubMed]
- Bomblies, K.; Weigel, D. Hybrid necrosis: Autoimmunity as a potential gene-flow barrier in plant species. Nat. Rev. Genet. 2007, 8, 382–393. [Google Scholar] [CrossRef] [PubMed]
- Oka, H.I. Phylogenetic differentiation of cultivated rice. XV. Complementary lethal genes in rice. Jpn. J. Genet. 1957, 32, 83–87. [Google Scholar] [CrossRef]
- Harushima, Y.; Yano, M.; Shomura, A.; Sato, M.; Shimano, T.; Kuboki, Y.; Yamamoto, T.; Lin, S.Y.; Antonio, B.A.; Parco, A.; et al. A high-density rice genetic linkage map with 2275 markers using a single F2 population. Genetics 1998, 148, 479–494. [Google Scholar] [PubMed]
- Huang, X.; Peng, X.; Sun, M.-X. OsGCD1 is essential for rice fertility and required for embryo dorsal-ventral pattern formation and endosperm development. New Phytol. 2017, 215, 1039–1058. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Lu, Z.; Wang, X.; Ouyang, Y.; Chen, W.; Xie, K.; Wang, D.; Luo, M.; Luo, J.; Yao, J. Imprinted gene OsFIE1 modulates rice seed development by influencing nutrient metabolism and modifying genome H3K27me3. Plant J. 2016, 87, 305–317. [Google Scholar] [CrossRef] [PubMed]
- Hara, T.; Katoh, H.; Ogawa, D.; Kagaya, Y.; Sato, Y.; Kitano, H.; Nagato, Y.; Ishikawa, R.; Ono, A.; Kinoshita, T.; et al. Rice SNF2 family helicase ENL1 is essential for syncytial endosperm development. Plant J. 2015, 81, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Hong, S.K.; Kitano, H.; Satoh, H.; Nagato, Y. How is embryo size genetically regulated in rice? Development 1996, 122, 2051–2058. [Google Scholar]
- Wambugu, P.W.; Brozynska, M.; Furtado, A.; Waters, D.L.; Henry, R.J. Relationships of wild and domesticated rices (Oryza AA genome species) based upon whole chloroplast genome sequences. Sci. Rep. 2015, 5, 13957. [Google Scholar] [CrossRef]
- Yin, H.; Akimoto, M.; Kaewcheenchai, R.; Sotowa, M.; Ishii, T.; Ishikawa, R. Inconsistent diversities between nuclear and plastid genomes of aa genome species in the genus Oryza. Genes Genet. Syst. 2015, 90, 269–281. [Google Scholar] [CrossRef] [PubMed]
- Sotowa, M.; Ootsuka, K.; Kobayashi, Y.; Hao, Y.; Tanaka, K.; Ichitani, K.; Flowers, J.M.; Purugganan, M.D.; Nakamura, I.; Sato, Y.-I.; et al. Molecular relationships between Australian annual wild rice, Oryza meridionalis, and two related perennial forms. Rice 2013, 6, 26. [Google Scholar] [CrossRef] [PubMed]
- Moner, A.M.; Henry, R.J. Oryza meridionalis Ng. In The Wild Oryza Genomes; Mondal, T.K., Henry, R., Eds.; Springer-Verlag: Heidelberg, Germany, 2018; pp. 177–182. [Google Scholar]
- Brozynska, M.; Copetti, D.; Furtado, A.; Wing, R.A.; Crayn, D.; Fox, G.; Ishikawa, R.; Henry, R.J. Sequencing of australian wild rice genomes reveals ancestral relationships with domesticated rice. Plant Biotechnol. J. 2017, 15, 765–774. [Google Scholar] [CrossRef] [PubMed]
- Moner, A.M.; Furtado, A.; Chivers, I.; Fox, G.; Crayn, D.; Henry, R.J. Diversity and evolution of rice progenitors in australia. Ecol. Evol. 2018, 8, 4360–4366. [Google Scholar] [CrossRef] [PubMed]
- Ichitani, K.; Taura, S.; Sato, M.; Kuboyama, T. Distribution of Hwc2-1, a causal gene of a hybrid weakness, in the World Rice Core collection and the Japanese Rice mini Core collection: Its implications for varietal differentiation and artificial selection. Breed. Sci. 2016, 66, 776–789. [Google Scholar] [CrossRef] [PubMed]
- Muto, C.; Ishikawa, R.; Olsen, K.M.; Kawano, K.; Bounphanousay, C.; Matoh, T.; Sato, Y.-I. Genetic diversity of the wx flanking region in rice landraces in northern Laos. Breed. Sci. 2016, 66, 580–590. [Google Scholar] [CrossRef][Green Version]
- Ichitani, K.; Yamaguchi, D.; Taura, S.; Fukutoku, Y.; Onoue, M.; Shimizu, K.; Hashimoto, F.; Sakata, Y.; Sato, M. Genetic analysis of ion-beam induced extremely late heading mutants in rice. Breed. Sci. 2014, 64, 222–230. [Google Scholar] [CrossRef]
- Wan, J.; Ikehashi, H. Identification of two types of differentiation in cultivated rice (Oryza sativa L.) detected by polymorphism of isozymes and hybrid sterility. Euphytica 1997, 94, 151–161. [Google Scholar] [CrossRef]
- Kawahara, Y.; de la Bastide, M.; Hamilton, J.P.; Kanamori, H.; McCombie, W.R.; Ouyang, S.; Schwartz, D.C.; Tanaka, T.; Wu, J.; Zhou, S.; et al. Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data. Rice 2013, 6, 4. [Google Scholar] [CrossRef]
- Yu, J.; Hu, S.; Wang, J.; Wong, G.K.-S.; Li, S.; Liu, B.; Deng, Y.; Dai, L.; Zhou, Y.; Zhang, X.; et al. A draft sequence of the rice genome (Oryza sativa L. ssp. indica). Science 2002, 296, 79–92. [Google Scholar] [CrossRef]
- Ichitani, K.; Taura, S.; Tezuka, T.; Okiyama, Y.; Kuboyama, T. Chromosomal location of HWA1and HWA2, complementary hybrid weakness genes in rice. Rice 2011, 4, 29–38. [Google Scholar] [CrossRef]
- Busungu, C.; Taura, S.; Sakagami, J.-I.; Ichitani, K. Identification and linkage analysis of a new rice bacterial blight resistance gene from XM14, a mutant line from IR24. Breed. Sci. 2016, 66, 636–645. [Google Scholar] [CrossRef] [PubMed]

| Marker Name | Kind of DNA Marker | Primer Sequence | Location on IRGSP 1.0 pseudomolecule chromosome 6 | |||
|---|---|---|---|---|---|---|
| From | To | Source | ||||
| KGC6_8.73 | Indel | F | GAAGAGGAACATATGTGGTGTAAGC | 8731826 | 8731914 | This study |
| R | AAAATTTATACTCTTGGTGACGTGA | |||||
| RM136 | SSR | F | GAGAGCTCAGCTGCTGCCTCTAGC | 8752461 | 8752562 | Temnykh et al. [19] |
| R | GAGGAGCGCCACGGTGTACGCC | |||||
| KGC6_8.82 | Indel | F | TCTCTACCACACTCATCATCTGC | 8820385 | 8820484 | This study |
| R | CCCTCGAGTAATAAACGATCCAG | |||||
| KGC6_10.09 | Indel | F | TAGTCCTACGAAAACCCCTACTAGA | 10090008 | 10090165 | This study |
| R | TTCCACGCACTAATACTACTACCTC | |||||
| KGC6_12.02 | Indel | F | TTGATTTTGGGAAACATCAGGTAGC | 12020476 | 12020625 | This study |
| R | AGCATGGTAATTTCATCGGATTCAA | |||||
| KGC6_13.00 | Indel | F | CATTCGCATGGTAGCCTTTTCTTAT | 13008017 | 13008169 | This study |
| R | CATAGGTGCCACAAGAGAAATCTTC | |||||
| RM6818 | SSR | F | CGGCGAAGACTTGGAACCT | 16582450 | 16582596 | McCouch et al. [20] |
| R | CCGTCACAAGGCTCGTCC | redesigned in this study | ||||
| RM193 | SSR | F | CAATCAACCAAACCGCGCTC | 18086456 | 18086578 | Temnykh et al. [19] |
| R | CGCGGGCTTCTTCTCCTTC | redesigned in this study | ||||
| KGC6_19.48 | Indel | F | GAAGATAGTTAAGGGGTGTAGTGTGA | 19483821 | 19484065 | This study |
| R | GACCAAAAGTTAAACAACATATTCTTCTAACCTAG | |||||
| KGC6_22.19 | Indel | F | ACAAAATATGCTTTCTTCGTGCGTA | 22191370 | 22191498 | This study |
| R | GCACTCAACTGTATCGTCTTTGAAA | |||||
| Haplotype | Genotype of DNA Marker 1 | No. of Plants | |||||||
|---|---|---|---|---|---|---|---|---|---|
| KGC6_8.73 | KGC6_10.09 | KGC6_12.02 | KGC6_13.00 | RM6818 | RM193 | KGC6_19.48 | KGC6_22.19 | ||
| 1 | H | H | H | H | H | H | H | H | 115 |
| 2 | T | T | T | T | T | T | T | T | 59 |
| 3 | H | H | T | T | T | T | T | T | 2 |
| 4 | T | T | T | T | T | T | T | H | 2 |
| 5 | H | T | T | T | T | T | T | T | 1 |
| 6 | H | H | H | H | H | H | H | H | 1 |
| 7 | W | H | H | H | H | H | H | H | 1 |
| 8 | W | W | H | H | H | H | H | H | 1 |
| 9 | H | H | H | H | H | H | H | W | 1 |
| 183 | |||||||||
| BC4F2 | Genotype of the KGC6_12.02 1 | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Individual Number | BC4F3 | BC5F1 | ||||||||||||||||||
| Plant | Embryo of Fertile Seeds | T65 as Pollen Parent | T65 as Egg Donor | |||||||||||||||||
| P | P | P | P | |||||||||||||||||
| χ2 | χ2 | χ2 | χ2 | χ2 | χ2 | |||||||||||||||
| T | H | W | (1:2:1) | (1:2:0) | T | H | W | (1:2:1) | (1:2:0) | T | H | (1:1) | T | H | (1:1) | |||||
| 1 | 10 | 23 | 0 | 0.004 | 0.712 | 21 | 26 | 0 | 0.000 | 0.099 | 10 | 17 | 0.178 | 12 | 13 | 0.841 | ||||
| 2 | 12 | 14 | 0 | 0.004 | 0.166 | 17 | 27 | 0 | 0.000 | 0.456 | 10 | 17 | 0.178 | 8 | 9 | 0.808 | ||||
| 3 | 12 | 20 | 0 | 0.004 | 0.617 | 16 | 30 | 0 | 0.000 | 0.835 | 17 | 10 | 0.178 | 16 | 7 | 0.061 | ||||
| 4 | 18 | 18 | 0 | 0.000 | 0.034 | 15 | 33 | 0 | 0.000 | 0.759 | 12 | 12 | 1.000 | 14 | 15 | 0.853 | ||||
| 5 | 10 | 15 | 0 | 0.011 | 0.480 | 20 | 27 | 0 | 0.000 | 0.180 | 10 | 14 | 0.414 | 15 | 13 | 0.705 | ||||
| 6 | 12 | 18 | 0 | 0.005 | 0.439 | 21 | 27 | 0 | 0.000 | 0.126 | 14 | 12 | 0.695 | 14 | 9 | 0.297 | ||||
| 7 | 6 | 23 | 0 | 0.002 | 0.149 | 24 | 24 | 0 | 0.000 | 0.014 | 8 | 20 | 0.023 | 21 | 23 | 0.763 | ||||
| Sum | 80 | 131 | 0 | 0.000 | 0.158 | 134 | 194 | 0 | 0.000 | 0.004 | 81 | 102 | 0.121 | 100 | 89 | 0.424 | ||||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Toyomoto, D.; Uemura, M.; Taura, S.; Sato, T.; Henry, R.; Ishikawa, R.; Ichitani, K. Segregation Distortion Observed in the Progeny of Crosses Between Oryza sativa and O. meridionalis Caused by Abortion During Seed Development. Plants 2019, 8, 398. https://doi.org/10.3390/plants8100398
Toyomoto D, Uemura M, Taura S, Sato T, Henry R, Ishikawa R, Ichitani K. Segregation Distortion Observed in the Progeny of Crosses Between Oryza sativa and O. meridionalis Caused by Abortion During Seed Development. Plants. 2019; 8(10):398. https://doi.org/10.3390/plants8100398
Chicago/Turabian StyleToyomoto, Daiki, Masato Uemura, Satoru Taura, Tadashi Sato, Robert Henry, Ryuji Ishikawa, and Katsuyuki Ichitani. 2019. "Segregation Distortion Observed in the Progeny of Crosses Between Oryza sativa and O. meridionalis Caused by Abortion During Seed Development" Plants 8, no. 10: 398. https://doi.org/10.3390/plants8100398
APA StyleToyomoto, D., Uemura, M., Taura, S., Sato, T., Henry, R., Ishikawa, R., & Ichitani, K. (2019). Segregation Distortion Observed in the Progeny of Crosses Between Oryza sativa and O. meridionalis Caused by Abortion During Seed Development. Plants, 8(10), 398. https://doi.org/10.3390/plants8100398







