Efficient Biolistic Transformation of Immature Citrus Rootstocks Using Phosphomannose-isomerase Selection
Abstract
1. Introduction
2. Results
2.1. Preliminary Test of Explant Sensitivity to Mannose and Shoot Regeneration
2.2. Biolistic Transformation of Carrizo Citrange
2.3. Biolistic Transformation of Swingle Citrumelo
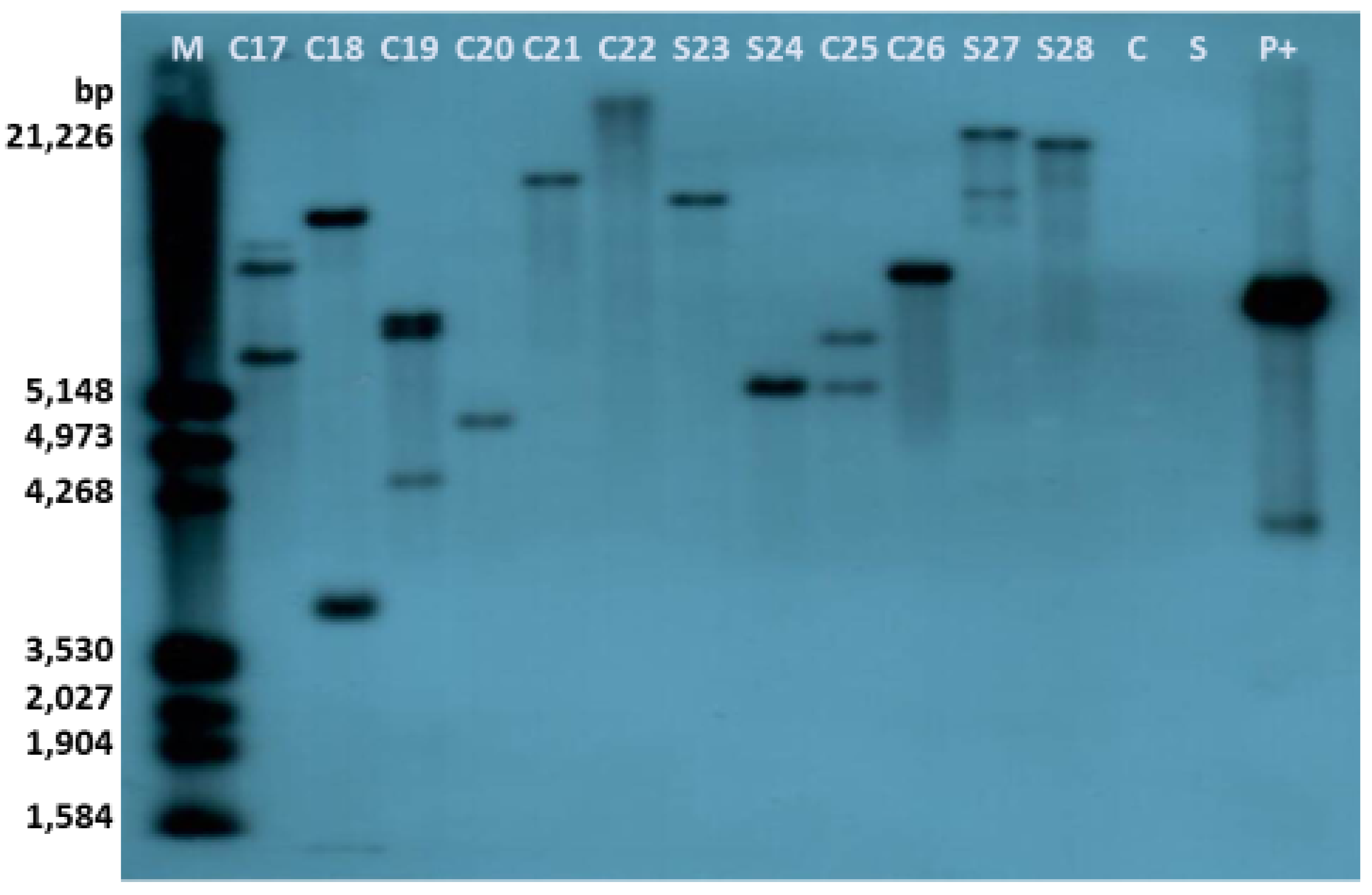
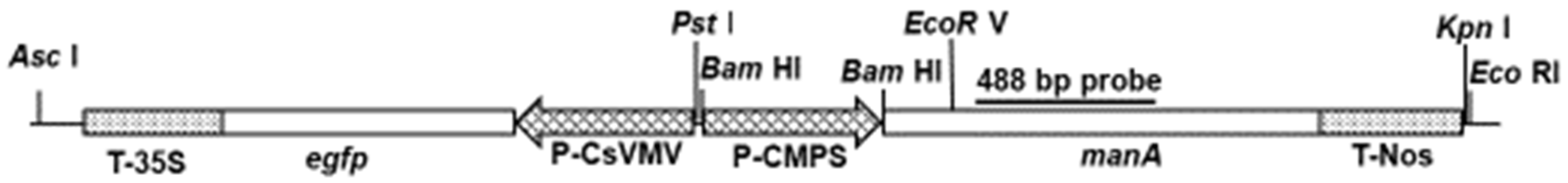
2.4. Molecular Analyses of the Transgenics
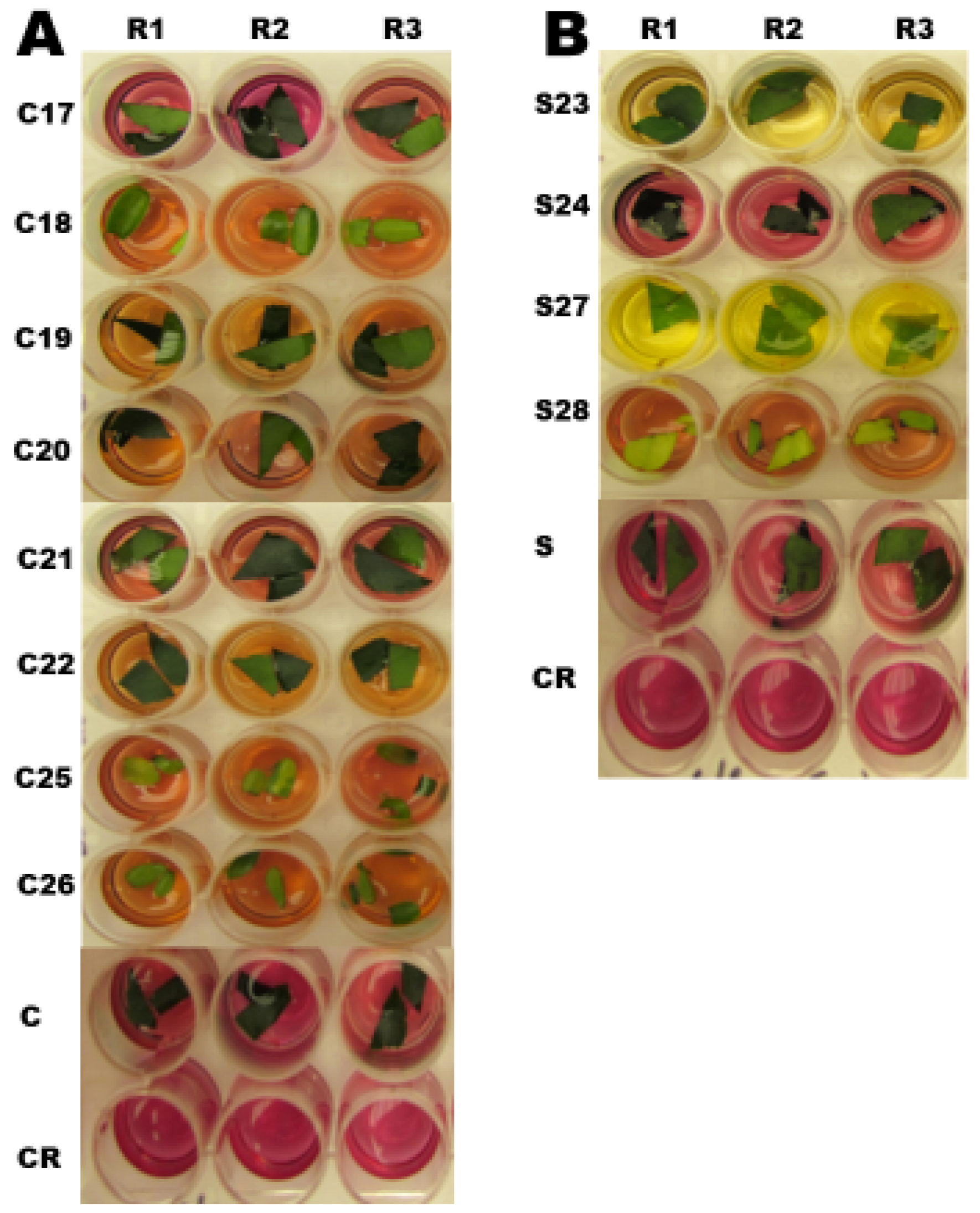
2.5. Chlorophenol Red Assay
3. Discussion
4. Materials and Methods
4.1. Plasmid DNA
4.2. Plant Materials and Tissue Culture
4.3. Preparation of Gold Particles
4.4. Particle Bombardments and Selection on Mannose
4.5. Fluorescence Microscopy and Data Collection
4.6. Micro-Grafting
4.7. Molecular Analyses
4.8. Chlorophenol Red Assay
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- USDA. Florida Citrus Statistics 2016–2017; NAS, Ed.; USDA National Agricultural Statistics Service: Maitland, FL, USA, 2018; p. 14. Available online: https://www.nass.usda.gov/Statistics_by_State/Florida/Publications/Citrus/Citrus_Statistics/2016-17/fcs1617.pdf (accessed on 2 June 2018).
- National Academy of Sciences, Engineering and Medicine. A Review of the Citrus Greening Research and Development Efforts Supported by the Citrus Research and Development Foundation: Fighting a Ravaging Disease; National Academy of Sciences, Engineering and Medicine: Washington, DC, USA, 2018; Available online: https://www.nap.edu/catalog/25026/a-review-of-the-citrus-greening-research-and-development-efforts-supported-by-the-citrus-research-and-development-foundation (accessed on 2 June 2018).
- Ramesh, S.A.; Kaiser, B.N.; Franks, T.; Collins, G.; Sedgley, M. Improved methods in Agrobacterium–mediated transformation of almond using positive (mannose/pmi) or negative (kanamycin resistance) selection-based protocols. Plant Cell Rep. 2006, 25, 821–828. [Google Scholar] [CrossRef] [PubMed]
- Degenhardt, J.; Poppe, A.; Montag, J.; Szankowski, I. The use of the phosphomannose-isomerase/mannose selection system to recover transgenic apple plants. Plant Cell Rep. 2006, 25, 1149–1156. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Potrykus, I.; Puonti-Kaerlas, J. Efficient production of transgenic cassava using negative and positive selection. Transgenic Res. 2000, 9, 405–415. [Google Scholar] [CrossRef] [PubMed]
- Wright, M.; Dawson, J.; Dunder, E.; Suttie, J.; Reed, J.; Kramer, C.; Chang, Y.; Novitzky, R.; Wang, H.; Artim-Moore, L. Efficient biolistic transformation of maize (Zea mays L.) and wheat (Triticum aestivum L.) using the phosphomannose isomerase gene, pmi, as the selectable marker. Plant Cell Rep. 2001, 20, 429–436. [Google Scholar] [CrossRef]
- Duan, Y.; Zhai, C.; Li, H.; Li, J.; Mei, W.; Gui, H.; Ni, D.; Song, F.; Li, L.; Zhang, W. An efficient and high-throughput protocol for Agrobacterium-mediated transformation based on phosphomannose isomerase positive selection in Japonica rice (Oryza sativa L.). Plant Cell Rep. 2012, 31, 1611–1624. [Google Scholar] [CrossRef] [PubMed]
- Gadaleta, A.; Giancaspro, A.; Blechl, A.; Blanco, A. Phosphomannose isomerase, pmi, as a selectable marker gene for durum wheat transformation. J. Cereal Sci. 2006, 43, 31–37. [Google Scholar] [CrossRef]
- Hoa, T.T.C.; Al-Babili, S.; Schaub, P.; Potrykus, I.; Beyer, P. Golden Indica and Japonica rice lines amenable to deregulation. Plant Physiol. 2003, 133, 161–169. [Google Scholar] [CrossRef]
- Hu, L.; Li, H.; Qin, R.; Xu, R.; Li, J.; Li, L.; Wei, P.; Yang, J. Plant phosphomannose isomerase as a selectable marker for rice transformation. Sci. Rep. 2016, 6, 25921. [Google Scholar] [CrossRef]
- Boscariol, R.; Almeida, W.; Derbyshire, M.; Mourao Filho, F.; Mendes, B. The use of the PMI/mannose selection system to recover transgenic sweet orange plants (Citrus sinensis L. Osbeck). Plant Cell Rep. 2003, 22, 122–128. [Google Scholar] [CrossRef]
- Ballester, A.; Cervera, M.; Peña, L. Evaluation of selection strategies alternative to nptII in genetic transformation of citrus plants. Plant Cell Rep. 2008, 27, 1005–1015. [Google Scholar] [CrossRef]
- Dutt, M.; Lee, D.H.; Grosser, J.W. Bifunctional selection–reporter systems for genetic transformation of citrus: Mannose-and kanamycin-based systems. In Vitro Cell. Dev. Biol. Plant 2010, 46, 467–476. [Google Scholar] [CrossRef]
- Febres, V.; Khalaf, A.; Moore, G.A.; Fisher, L. Citrus Transformation: Challenges and Prospects; INTECH Open Access Publisher: London, UK, 2011. [Google Scholar]
- Wu, H.; Acanda, Y.; Shankar, A.; Peeples, M.; Hubbard, C.; Orbović, V.; Zale, J. Genetic transformation of commercially important mature citrus scions. Crop Sci. 2015, 55, 2786–2797. [Google Scholar] [CrossRef]
- Wu, H.; Acanda, Y.; Jia, H.; Wang, N.; Zale, J. Biolistic transformation of Carrizo citrange (Citrus sinensis Osb.× Poncirus trifoliata L. Raf.). Plant Cell Rep. 2016, 35, 1955–1962. [Google Scholar] [CrossRef] [PubMed]
- Dutt, M.; Grosser, J. Evaluation of parameters affecting Agrobacterium-mediated transformation of citrus. Plant Cell Tissue Organ Cult. 2009, 98, 331–340. [Google Scholar] [CrossRef]
- Okkels, F.; Ward, J.; Joersbo, M. Synthesis of cytokinin glucuronides for the selection of transgenic plant cells. Phytochemistry 1997, 46, 801–804. [Google Scholar] [CrossRef]
- Wu, H.; Awan, F.S.; Vilarinho, A.; Zeng, Q.; Kannan, B.; Phipps, T.; McCuiston, J.; Wang, W.; Caffall, K.; Altpeter, F. Transgene integration complexity and expression stability following biolistic or Agrobacterium-mediated transformation of sugarcane. In Vitro Cell. Dev. Biol. Plant 2015, 51, 603–611. [Google Scholar] [CrossRef]
- Jackson, M.A.; Anderson, D.J.; Birch, R.G. Comparison of Agrobacterium and particle bombardment using whole plasmid or minimal cassette for production of high-expressing, low-copy transgenic plants. Transgenic Res. 2013, 22, 143–151. [Google Scholar] [CrossRef] [PubMed]
- Breyer, D.; Kopertekh, L.; Reheul, D. Alternatives to antibiotic resistance marker genes for in vitro selection of genetically modified plants–scientific developments, current use, operational access and biosafety considerations. Crit. Rev. Plant Sci. 2014, 33, 286–330. [Google Scholar] [CrossRef]
- Song, G.-Q.; Sink, K.C.; Ma, Y.; Herlache, T.; Hancock, J.F.; Loescher, W.H. A novel mannose-based selection system for plant transformation using celery mannose-6-phosphate reductase gene. Plant Cell Rep. 2010, 29, 163–172. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Zhang, S.; Zhao, X.; Li, Y.; Liu, T.; Wang, J.; Hou, X.; Li, Y. BcPMI2, isolated from non-heading Chinese cabbage encoding phosphomannose isomerase, improves stress tolerance in transgenic tobacco. Mol. Biol. Rep. 2014, 41, 2207–2216. [Google Scholar] [CrossRef] [PubMed]
- Dutt, M.; Li, Z.; Dhekney, S.; Gray, D. A co-transformation system to produce transgenic grapevines free of marker genes. Plant Sci. 2008, 175, 423–430. [Google Scholar] [CrossRef]
- Dutt, M.; Erpen, L.; Ananthakrishnan, G.; Barthe, G.; Brlansky, R.; Maiti, I.; Grosser, J. Comparative expression analysis of five caulimovirus promoters in citrus. Plant Cell Tissue Organ Cult. 2016, 126, 229–238. [Google Scholar] [CrossRef]
- Hohn, T.; Stavolone, L.; De Haan, P.T.; Ligon, H.T.; Kononova, M. Cestrum Yellow Leaf Curling Virus Promoters. Google Patents: 2007. Available online: https://patents.google.com/patent/WO2001073087A8/en?oq=google+patents+cestrum+yellow (accessed on 2 June 2018).
- Murashige, T.; Skoog, F. A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol. Plant. 1962, 15, 473–497. [Google Scholar] [CrossRef]
- Altpeter, F.; Sandhu, S.; Davey, M.R.; Anthony, P. Genetic transformation–biolistics. Plant Cell Cult. Essent. Methods 2010, 217–239. [Google Scholar] [CrossRef]
- Cervera, M.; Juarez, J.; Navarro, L.; Pena, L. Genetic transformation of mature citrus plants. In Transgenic Plants: Methods and Protocols; Pena, L., Ed.; Humana Press: Totowa, NJ, USA, 2005; Volume 286, pp. 177–188. [Google Scholar]
- Orbović, V.; Shankar, A.; Peeples, M.E.; Hubbard, C.; Zale, J. Citrus transformation using mature tissue explants. In Agrobacterium Protocols; Wang, K., Ed.; Springer: Clifton, NJ, USA, 2015; Volume 1, pp. 259–273. [Google Scholar]
- Porebski, S.; Bailey, L.G.; Baum, B.R. Modification of a CTAB DNA extraction protocol for plants containing high polysaccharide and polyphenol components. Plant Mol. Biol. Rep. 1997, 15, 8–15. [Google Scholar] [CrossRef]
- Kramer, C.; DiMaio, J.; Carswell, G.K.; Shillito, R.D. Selection of transformed protoplast-derived Zea mays colonies with phosphinothricin and a novel assay using the pH indicator chlorophenol red. Planta 1993, 190, 454–458. [Google Scholar] [CrossRef]



| Experiment | Paired Shots | Number of Explants | Number of Shoots | Number of Positive Shoots | Number of Shoots Survived/Number Micro-grafted | PCR Positive Shoots |
|---|---|---|---|---|---|---|
| 1 | 1 | 153 | 294 | 2 | 0/1 | 0 |
| 1 | 2 | 126 | 237 | 3 | 1/2 | 1 |
| 2 | 3 | 165 | 288 | 4 | 2/2 | 2 |
| 2 | 4 | 153 | 284 | 2 | 1/1 | 1 |
| 3 | 5 | 159 | 270 | 1 | 1/1 | 1 |
| 3 | 6 | 166 | 214 | 1 | 0/1 | 0 |
| 4 | 7 | 154 | 346 | 1 | 1/1 | 1 |
| 4 | 8 | 80 | 140 | 0 | 0/0 | 0 |
| 5 | 9 | 149 | 319 | 3 | 1/1 | 1 |
| 5 | 10 | 88 | 173 | 2 | 1/1 | 1 |
| Total | - | 1393 | 2565 | 19 | 8/11 | 8/8 |
| Mean ± SE | - | 139.3 ± 9.9 | 256.5 ± 20.5 | 1.9 ± 0.4 | 72.7% survival | 100% |
| Experiment | Paired Shots | Number of Explants | Number of Shoots | Number of Positive Shoots | Number of Shoots Survived/Number Micro-grafted | PCR Positive Shoots |
|---|---|---|---|---|---|---|
| 1 | 1 | 103 | 121 | 0 | 0/0 | 0 |
| 1 | 2 | 99 | 187 | 2 | 2/2 | 2 |
| 2 | 3 | 97 | 132 | 1 | 1/1 | 1 |
| 2 | 4 | 110 | 164 | 2 | 0/1 | 0 |
| 3 | 5 | 144 | 205 | 3 | 1/2 | 1 |
| Total | - | 553 | 809 | 8 | 4/6 | 4/4 |
| Mean ± SE | - | 110.6 ± 8.6 | 161.8 ± 15.9 | 1.6 ± 0.5 | 66.7% | 100% |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, H.; Acanda, Y.; Canton, M.; Zale, J. Efficient Biolistic Transformation of Immature Citrus Rootstocks Using Phosphomannose-isomerase Selection. Plants 2019, 8, 390. https://doi.org/10.3390/plants8100390
Wu H, Acanda Y, Canton M, Zale J. Efficient Biolistic Transformation of Immature Citrus Rootstocks Using Phosphomannose-isomerase Selection. Plants. 2019; 8(10):390. https://doi.org/10.3390/plants8100390
Chicago/Turabian StyleWu, Hao, Yosvanis Acanda, Michel Canton, and Janice Zale. 2019. "Efficient Biolistic Transformation of Immature Citrus Rootstocks Using Phosphomannose-isomerase Selection" Plants 8, no. 10: 390. https://doi.org/10.3390/plants8100390
APA StyleWu, H., Acanda, Y., Canton, M., & Zale, J. (2019). Efficient Biolistic Transformation of Immature Citrus Rootstocks Using Phosphomannose-isomerase Selection. Plants, 8(10), 390. https://doi.org/10.3390/plants8100390




