Molecular Cloning and Characterization of SaCLCd, SaCLCf, and SaCLCg, Novel Proteins of the Chloride Channel Family (CLC) from the Halophyte Suaeda altissima (L.) Pall
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material
2.2. Total RNA Extraction and First-Strand cDNA Synthesis
2.3. Amplification of SaCLCd, SaCLCf, and SaCLCg cDNA Partial Sequences
2.4. Cloning of the Full-Length SaCLCd, SaCLCf, and SaCLCg cDNA Sequences
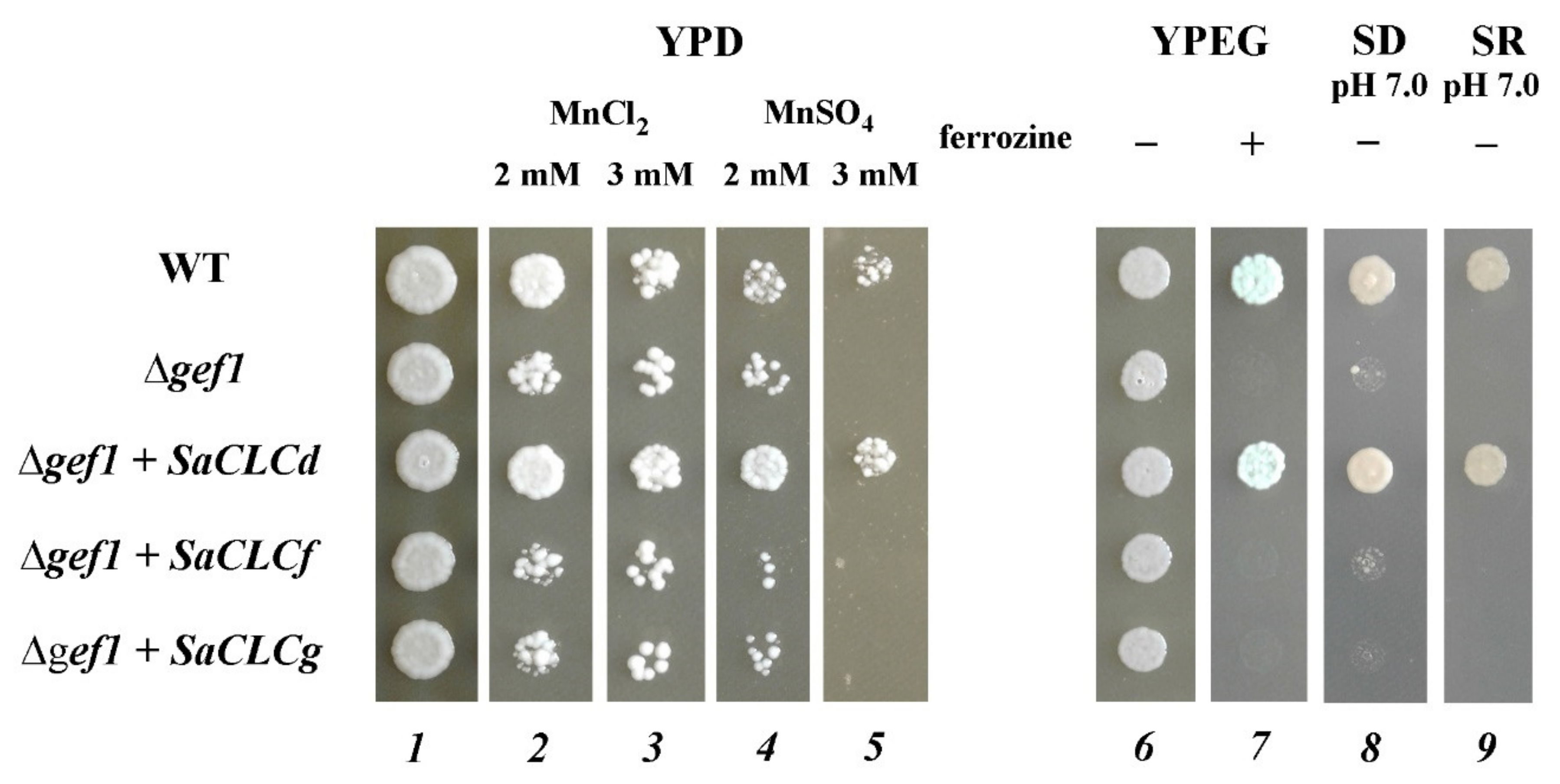
2.5. Heterologous Expression of the SaCLCd, SaCLCf, and SaCLCg Genes in Δgef1 Yeast Mutant
2.6. Quantitative Analysis of SaCLCd, SaCLCf, and SaCLCg Transcripts in S. altissima Organs
2.7. Primer Design
2.8. Bioinformatic Analysis of Amino Acid Sequences
2.9. Determination of Chloride Content in S. altissima Organs
2.10. Statistical Analysis
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Flowers, T.J. Salinisation and horticultural production. Sci. Hortic. 1999, 78, 1–4. [Google Scholar]
- UNESCO. World Water Quality Portal Unesco. Available online: https://en.unesco.org/waterqualitymonitor (accessed on 20 December 2021).
- Ismail, A.M.; Horie, T. Genomics, Physiology, and Molecular Breeding Approaches for Improving Salt Tolerance. Annu. Rev. Plant Biol. 2017, 68, 405–434. [Google Scholar] [CrossRef] [PubMed]
- Qadir, M.; Quillérou, E.; Nangia, V.; Murtaza, G.; Singh, M.; Thomas, R.J.; Drechsel, P.; Noble, A.D. Economics of salt-induced land degradation and restoration. Nat. Resour. Forum 2014, 38, 282–295. [Google Scholar] [CrossRef]
- Wang, B.; Davenport, R.J.; Volkov, V.; Amtmann, A. Low unidirectional sodium influx into root cells restricts net sodium accumulation in Thellungiella halophila, a salt-tolerant relative of Arabidopsis thaliana. J. Exp. Bot. 2006, 57, 1161–1170. [Google Scholar] [CrossRef] [PubMed]
- Apse, M.P.; Blumwald, E. Na+ transport in plants. FEBS Lett. 2007, 581, 2247–2254. [Google Scholar] [CrossRef] [PubMed]
- Munns, R.; Tester, M. Mechanisms of salinity tolerance. Annu. Rev. Plant Biol. 2008, 59, 651–681. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, T.; Hamamoto, S.; Uozumi, N. Sodium transport system in plant cells. Front. Plant Sci. 2013, 4, 410. [Google Scholar] [CrossRef] [PubMed]
- Teakle, N.L.; Tyerman, S.D. Mechanisms of Cl− transport contributing to salt tolerance. Plant Cell Environ. 2010, 33, 566–589. [Google Scholar] [CrossRef]
- Agarwal, P.K.; Shukla, P.S.; Gupta, K.; Jha, B. Bioengineering for salinity tolerance in plants: State of the art. Mol. Biotechnol. 2013, 54, 102–123. [Google Scholar] [CrossRef]
- Mbarki, S.; Sytar, O.; Cerda, A.; Zivcak, M.; Rastogi, A.; He, X.; Zoghlami, A.; Abdelly, C.; Brestic, M. Strategies to mitigate the salt stress effects on photosynthetic apparatus and productivity of crop plants. In Salinity Responses and Tolerance in Plants; Kumar, V., Wani, S.H., Penna, S., Tran, L.S.P., Eds.; Springer: Cham, Switzerland, 2018; Volume 1, pp. 85–136. ISBN 9783319756714. [Google Scholar]
- Flowers, T.J.; Galal, H.K.; Bromham, L. Evolution of halophytes: Multiple origins of salt tolerance in land plants. Funct. Plant Biol. 2010, 37, 604–612. [Google Scholar] [CrossRef]
- Shabala, S. Learning from halophytes: Physiological basis and strategies to improve abiotic stress tolerance in crops. Ann. Bot. 2013, 112, 1209–1221. [Google Scholar] [CrossRef]
- Flowers, T.J.; Munns, R.; Colmer, T.D. Sodium chloride toxicity and the cellular basis of salt tolerance in halophytes. Ann. Bot. 2015, 115, 419–431. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.; Ishitani, M.; Kim, C.; Zhu, J.K. The Arabidopsis thaliana salt tolerance gene SOS1 encodes a putative Na+/H+ antiporter. Proc. Natl. Acad. Sci. USA. 2000, 97, 6896–6901. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.; Quintero, F.J.; Pardo, J.M.; Zhu, J.K. The putative plasma membrane Na+/H+ antiporter SOS1 controls long-distance NA+ transport in plants. Plant Cell 2002, 14, 465–477. [Google Scholar] [CrossRef] [PubMed]
- Tester, M.; Davenport, R. Na+ tolerance and Na+ transport in higher plants. Ann. Bot. 2003, 91, 503–527. [Google Scholar] [CrossRef] [PubMed]
- Davenport, R.J.; Muñoz-Mayor, A.; Jha, D.; Essah, P.A.; Rus, A.; Tester, M. The Na+ transporter AtHKT1;1 controls retrieval of Na+ from the xylem in Arabidopsis. Plant Cell Environ. 2007, 30, 497–507. [Google Scholar] [CrossRef] [PubMed]
- James, R.A.; Blake, C.; Zwart, A.B.; Hare, R.A.; Rathjen, A.J.; Munns, R. Impact of ancestral wheat sodium exclusion genes Nax1 and Nax2 on grain yield of durum wheat on saline soils. Funct. Plant Biol. 2012, 39, 609–618. [Google Scholar] [CrossRef]
- Reguera, M.; Bassil, E.; Blumwald, E. Intracellular NHX-Type cation/H+ antiporters in plants. Mol. Plant 2014, 7, 261–263. [Google Scholar] [CrossRef]
- Wang, Z.; Hong, Y.; Zhu, G.; Li, Y.; Niu, Q.; Yao, J.; Hua, K.; Bai, J.; Zhu, Y.; Shi, H.; et al. Loss of salt tolerance during tomato domestication conferred by variation in a Na+/K+ transporter. EMBO J. 2020, 39, e103256. [Google Scholar] [CrossRef] [PubMed]
- Pantoja, O.; Dainty, J.; Blumwald, E. Cytoplasmic Chloride Regulates Cation Channels in the Vacuolar Membrane of Plant Cells. J. Membr. Biol. 1992, 125, 219–229. [Google Scholar] [CrossRef]
- White, P.J.; Broadley, M.R. Chloride in soils and its uptake and movement within the plant: A review. Ann. Bot. 2001, 88, 967–988. [Google Scholar] [CrossRef]
- Wei, P.; Wang, L.; Liu, A.; Yu, B.; Lam, H.M. GmCLC1 confers enhanced salt tolerance through regulating chloride accumulation in soybean. Front. Plant Sci. 2016, 7, 1082. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Tester, M.; Gilliham, M. Chloride on the move. Trends Plant Sci. 2017, 22, 236–248. [Google Scholar] [CrossRef] [PubMed]
- Flowers, T.J.; Yeo, A.R. Ion relation of salt tolerance. In Solute Transport in Plant Cells and Tissues; Baker, D., Hall, J., Eds.; Longman Scientific and Technical: Harlow, UK, 1988; pp. 392–413. [Google Scholar]
- Rubinigg, M.; Posthumus, F.; Ferschke, M.; Elzenga, J.T.M.; Stulen, I. Effects of NaCl salinity on 15N-nitrate fluxes and specific root length in the halophyte Plantago maritima L. Plant Soil 2003, 250, 201–213. [Google Scholar] [CrossRef]
- Song, J.; Ding, X.; Feng, G.; Zhang, F. Nutritional and osmotic roles of nitrate in a euhalophyte and a xerophyte in saline conditions. New Phytol. 2006, 171, 357–366. [Google Scholar] [CrossRef]
- Kudo, N.; Fujiyama, H. Responses of halophyte Salicornia bigelovii to different forms of nitrogen source. Pedosphere 2010, 20, 311–317. [Google Scholar] [CrossRef]
- Yuan, J.F.; Feng, G.; Ma, H.Y.; Tian, C.Y. Effect of nitrate on root development and nitrogen uptake of Suaeda physophora under NaCl salinity. Pedosphere 2010, 20, 536–544. [Google Scholar] [CrossRef]
- Niu, X.; Bressan, R.A.; Hasegawa, P.M.; Pardo, J.M. Ion homeostasis in NaCI stress environments. Plant Physiol. 1995, 109, 735–742. [Google Scholar] [CrossRef]
- Barbier-Brygoo, H.; Vinauger, M.; Colcombet, J.; Ephritikhine, G.; Frachisse, J.M.; Maurel, C. Anion channels in higher plants: Functional characterization, molecular structure and physiological role. Biochim. Biophys. Acta Biomembr. 2000, 1465, 199–218. [Google Scholar] [CrossRef]
- Zifarelli, G.; Pusch, M. CLC transport proteins in plants. FEBS Lett. 2010, 584, 2122–2127. [Google Scholar] [CrossRef]
- Nedelyaeva, O.I.; Shuvalov, A.V.; Balnokin, Y.V. Chloride Channels and Transporters of the CLC Family in Plants. Russ. J. Plant Physiol. 2020, 67, 767–784. [Google Scholar] [CrossRef]
- Jentsch, T.J.; Friedrich, T.; Schriever, A.; Yamada, H. The CLC chloride channel family. Pflugers Arch. Eur. J. Physiol. 1999, 437, 783–795. [Google Scholar] [CrossRef] [PubMed]
- Miller, C. ClC chloride channels viewed through a transporter lens. Nature 2006, 440, 484–489. [Google Scholar] [CrossRef] [PubMed]
- Marmagne, A.; Vinauger-Douard, M.; Monachello, D.; De Longevialle, A.F.; Charon, C.; Allot, M.; Rappaport, F.; Wollman, F.A.; Barbier-Brygoo, H.; Ephritikhine, G. Two members of the Arabidopsis CLC (chloride channel) family, AtCLCe and AtCLCf, are associated with thylakoid and golgi membranes, respectively. J. Exp. Bot. 2007, 58, 3385–3393. [Google Scholar] [CrossRef]
- Barbier-Brygoo, H.; De Angeli, A.; Filleur, S.; Frachisse, J.-M.; Gambale, F.; Thomine, S.; Wege, S. Anion channels/transporters in plants: From molecular bases to regulatory networks. Annu. Rev. Plant Biol. 2011, 62, 25–51. [Google Scholar] [CrossRef]
- De Angeli, A.; Monachello, D.; Ephritikhine, G.; Frachisse, J.M.; Thomine, S.; Gambale, F.; Barbier-Brygoo, H. The nitrate/proton antiporter AtCLCa mediates nitrate accumulation in plant vacuoles. Nature 2006, 442, 939–942. [Google Scholar] [CrossRef]
- Isayenkov, S.; Isner, J.C.; Maathuis, F.J.M. Vacuolar ion channels: Roles in plant nutrition and signalling. FEBS Lett. 2010, 584, 1982–1988. [Google Scholar] [CrossRef]
- Wege, S.; De Angeli, A.; Droillard, M.J.; Kroniewicz, L.; Merlot, S.; Cornu, D.; Gambale, F.; Martinoia, E.; Barbier-Brygoo, H.; Thomine, S.; et al. Phosphorylation of the vacuolar anion exchanger AtCLCa is required for the stomatal response to abscisic acid. Sci. Signal. 2014, 7, ra65. [Google Scholar] [CrossRef]
- Jentsch, T.J.; Pusch, M. CLC chloride channels and transporters: Structure, function, physiology, and disease. Physiol. Rev. 2018, 98, 1493–1590. [Google Scholar] [CrossRef]
- von der Fecht-Bartenbach, J.; Bogner, M.; Krebs, M.; Stierhof, Y.D.; Schumacher, K.; Ludewig, U. Function of the anion transporter AtCLC-d in the trans-Golgi network. Plant J. 2007, 50, 466–474. [Google Scholar] [CrossRef]
- Guo, W.; Zuo, Z.; Cheng, X.; Sun, J.; Li, H.; Li, L.; Qiu, J.L. The chloride channel family gene CLCd negatively regulates pathogen-associated molecular pattern (PAMP)-triggered immunity in Arabidopsis. J. Exp. Bot. 2014, 65, 1205–1215. [Google Scholar] [CrossRef]
- Scholl, S.; Hillmer, S.; Krebs, M.; Schumacher, K. ClCd and ClCf act redundantly at the trans—Golgi network/early endosome and prevent acidification of the Golgi stack. J. Cell Sci. 2021, 134, jcs258807. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, C.T.; Agorio, A.; Jossier, M.; Depré, S.; Thomine, S.; Filleur, S. Characterization of the chloride channel-like, AtCLCg, involved in chloride tolerance in Arabidopsis thaliana. Plant. Cell Physiol. 2015, 57, 764–775. [Google Scholar] [CrossRef]
- Ashraf, M.; Akram, N.A. Improving salinity tolerance of plants through conventional breeding and genetic engineering: An analytical comparison. Biotechnol. Adv. 2009, 27, 744–752. [Google Scholar] [CrossRef] [PubMed]
- Volkov, V. Salinity tolerance in plants. Quantitative approach to ion transport starting from halophytes and stepping to genetic and protein engineering for manipulating ion fluxes. Front. Plant Sci. 2015, 6, 873. [Google Scholar] [CrossRef] [PubMed]
- Mishra, A.; Tanna, B. Halophytes: Potential resources for salt stress tolerance genes and promoters. Front. Plant Sci. 2017, 8, 829. [Google Scholar] [CrossRef] [PubMed]
- Jha, R.; Patel, J.; Mishra, A.; Jha, B. Introgression of halophytic salt stress-responsive genes for developing stress tolerance in crop plants. In Halophytes and Climate Change: Adaptive Mechanisms and Potential Uses; Hasanuzzaman, M., Shabala, S., Fujita, M., Eds.; CABI: Boston, MA, USA, 2019; pp. 275–286. [Google Scholar]
- Nedelyaeva, O.I.; Shuvalov, A.V.; Mayorova, O.V.; Yurchenko, A.A.; Popova, L.G.; Balnokin, Y.V.; Karpichev, I.V. Cloning and functional analysis of SaCLCc1, a gene belonging to the chloride channel family (CLC), from the halophyte Suaeda altissima (L.) Pall. Dokl. Biochem. Biophys. 2018, 481, 186–189. [Google Scholar] [CrossRef] [PubMed]
- Nedelyaeva, O.I.; Shuvalov, A.V.; Karpichev, I.V.; Beliaev, D.V.; Myasoedov, N.A.; Khalilova, L.A.; Khramov, D.E.; Popova, L.G.; Balnokin, Y.V. Molecular cloning and characterisation of SaCLCa1, a novel protein of the chloride channel (CLC) family from the halophyte Suaeda altissima (L.) Pall. J. Plant Physiol. 2019, 240, 152995. [Google Scholar] [CrossRef] [PubMed]
- López-Rodríguez, A.; Cárabez Trejo, A.; Coyne, L.; Halliwell, R.F.; Miledi, R.; Martínez-Torres, A. The product of the gene GEF1 of Saccharomyces cerevisiae transports Cl—across the plasma membrane. FEMS Yeast Res. 2007, 7, 1218–1229. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Robinson, S.P.; Downton, W.J.S. Potassium, sodium and chloride ion concentrations in leaves and isolated chloroplasts of the halophyte Suaeda australis R. Br. Aust. J. Plant Physiol. 1985, 12, 471–479. [Google Scholar] [CrossRef]
- Yuorieva, N.O.; Voronkov, A.S.; Tereshonok, D.V.; Osipova, E.S.; Platonova, E.V.; Belyaev, D.V. An assay for express screening of potato transformants by GFP fluorescence. Mosc. Univ. Biol. Sci. Bull. 2018, 73, 69–75. [Google Scholar] [CrossRef]
- Shuvalov, A.V.; Yurchenko, A.A.; Nedelyaeva, O.I.; Myasoedov, N.A.; Karpichev, I.V.; Khalilova, L.A.; Popova, L.G.; Balnokin, Y.V. Identification of Some Anion Transporter Genes in the Halophyte Suaeda altissima (L.) Pall. and Their Expression under Nitrate Deficiency and Salinity. Russ. J. Plant Physiol. 2021, 68, 873–882. [Google Scholar] [CrossRef]
- Eldarov, M.A.; Baranov, M.V.; Dumina, M.V.; Shgun, A.A.; Andreeva, N.A.; Trilisenko, L.V.; Kulakovskaya, T.V.; Ryasanova, L.P.; Kulaev, I.S. Polyphosphates and exopolyphosphatase activities in the yeast Saccharomyces cerevisiae under overexpression of homologous and heterologous PPN1 genes. Biochemistry 2013, 78, 946–953. [Google Scholar] [CrossRef] [PubMed]
- Sambrook, J.; Fritsch, E.F.; Maniatis, T. Molecular Cloning: A Laboratory Manual; Cold Spring Harbor Laboratory Press: New York, NY, USA, 1989; p. 1546. [Google Scholar]
- Gaxiola, R.A.; Yuan, D.S.; Klausner, R.D.; Fink, G.R. The yeast CLC chloride channel functions in cation homeostasis. Proc. Natl. Acad. Sci. USA 1998, 95, 4046–4050. [Google Scholar] [CrossRef] [PubMed]
- Sherman, F. Getting started with yeast. In Guide to Yeast Genetics and Molecular Biology; Christine, G., Gerald, R.F., Eds.; Academic Press: San Diego, CA, USA, 1991; pp. 3–21. [Google Scholar]
- Jones, D.T.; Taylor, W.R.; Thornton, J.M. The rapid generation of mutation data matrices from sequences. Bioinformatics 1992, 8, 275–282. [Google Scholar] [CrossRef]
- Hechenberger, M.; Schwappach, B.; Fischer, W.N.; Frommer, W.B.; Jentsch, T.J.; Steinmeyer, K. A family of putative chloride channels from Arabidopsis and functional complementation of a yeast strain with a CLC gene disruption. J. Biol. Chem. 1996, 271, 33632–33638. [Google Scholar] [CrossRef]
- Lurin, C.; Geelen, D.; Barbier-Brygoo, H.; Guern, J.; Maurel, C. Cloning and functional expression of a plant voltage-dependent chloride channel. Plant Cell 1996, 8, 701–711. [Google Scholar] [CrossRef]
- Jentsch, T.J.; Steinmeyer, K.; Schwarz, G. Primary structure of Torpedo marmorata chloride channel isolated by expression cloning in Xenopus oocytes. Nature 1990, 348, 510–514. [Google Scholar] [CrossRef]
- Jentsch, T.J. CLC chloride channels and transporters: From genes to protein structure, pathology and physiology. Crit. Rev. Biochem. Mol. Biol. 2008, 43, 3–36. [Google Scholar] [CrossRef]
- Jentsch, T.J. Discovery of CLC transport proteins: Cloning, structure, function and pathophysiology. J. Physiol. 2015, 593, 4091–4109. [Google Scholar] [CrossRef]
- Dutzler, R. The structural basis of ClC chloride channel function. Trends Neurosci. 2004, 27, 315–320. [Google Scholar] [CrossRef] [PubMed]
- Zifarelli, G.; Pusch, M. Conversion of the 2 Cl−/1 H+ antiporter ClC-5 in a NO3−/H+ antiporter by a single point mutation. EMBO J. 2009, 28, 175–182. [Google Scholar] [CrossRef] [PubMed]
- Wege, S.; Jossier, M.; Filleur, S.; Thomine, S.; Barbier-Brygoo, H.; Gambale, F.; De Angeli, A. The proline 160 in the selectivity filter of the Arabidopsis NO3−/H+ exchanger AtCLCa is essential for nitrate accumulation in planta. Plant J. 2010, 63, 861–869. [Google Scholar] [CrossRef] [PubMed]
- Waterhouse, A.M.; Procter, J.B.; Martin, D.M.A.; Clamp, M.; Barton, G.J. Jalview Version 2—A multiple sequence alignment editor and analysis workbench. Bioinformatics 2009, 25, 1189–1191. [Google Scholar] [CrossRef]
- Bergsdorf, E.Y.; Zdebik, A.A.; Jentsch, T.J. Residues important for nitrate/proton coupling in plant and mammalian CLC transporters. J. Biol. Chem. 2009, 284, 11184–11193. [Google Scholar] [CrossRef]
- Accardi, A.; Walden, M.; Nguitragool, W.; Jayaram, H.; Williams, C.; Miller, C. Separate ion pathways in a Cl−/H+ exchanger. J. Gen. Physiol. 2005, 126, 563–570. [Google Scholar] [CrossRef]
- Miyazaki, H.; Uchida, S.; Takei, Y.; Hirano, T.; Marumo, F.; Sasaki, S. Molecular cloning of CLC chloride channels in Oreochromis mossambicus and their functional complementation of yeast CLC gene mutant. Biochem. Biophys. Res. Commun. 1999, 255, 175–181. [Google Scholar] [CrossRef]
- Kida, Y.; Uchida, S.; Miyazaki, H.; Sasaki, S.; Marumo, F. Localization of mouse CLC-6 and CLC-7 mRNA and their functional complementation of yeast CLC gene mutant. Histochem. Cell Biol. 2001, 115, 189–194. [Google Scholar] [CrossRef]
- Nakamura, A.; Fukuda, A.; Sakai, S.; Tanaka, Y. Molecular cloning, functional expression and subcellular localization of two putative vacuolar voltage-gated chloride channels in rice (Oryza sativa L.). Plant Cell Physiol. 2006, 47, 32–42. [Google Scholar] [CrossRef]
- Wei, P.; Che, B.; Shen, L.; Cui, Y.; Wu, S.; Cheng, C.; Liu, F.; Li, M.W.; Yu, B.; Lam, H.M. Identification and functional characterization of the chloride channel gene, GsCLC-c2 from wild soybean. BMC Plant Biol. 2019, 19, 1–15. [Google Scholar] [CrossRef]
- Accardi, A. Structure and gating of CLC channels and exchangers. J. Physiol. 2015, 593, 4129–4138. [Google Scholar] [CrossRef] [PubMed]
- Lv, Q.D.; Tang, R.; Liu, H.; Gao, X.S.; Li, Y.Z.; Zheng, H.Q.; Zhang, H.X. Cloning and molecular analyses of the Arabidopsis thaliana chloride channel gene family. Plant Sci. 2009, 176, 650–661. [Google Scholar] [CrossRef]
- Flowers, T.J.; Troke, P.F.; Yeo, A.R. The Mechanism of Salt Tolerance in Halophytes. Annu. Rev. Plant Physiol. 1977, 28, 89–121. [Google Scholar] [CrossRef]
- Flowers, T.J.; Colmer, T.D. Plant salt tolerance: Adaptations in halophytes. Ann. Bot. 2015, 115, 327–331. [Google Scholar] [CrossRef] [PubMed]
- Balnokin, Y.V.; Kotov, A.A.; Myasoedov, N.A.; Khailova, G.F.; Kurkova, E.B.; Lun’kov, R.V.; Kotova, L.M. Involvement of long-distance Na+ transport in maintaining water potential gradient in the medium-root-leaf system of a halophyte Suaeda altissima. Russ. J. Plant Physiol. 2005, 52, 489–496. [Google Scholar] [CrossRef]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nedelyaeva, O.I.; Popova, L.G.; Volkov, V.S.; Balnokin, Y.V. Molecular Cloning and Characterization of SaCLCd, SaCLCf, and SaCLCg, Novel Proteins of the Chloride Channel Family (CLC) from the Halophyte Suaeda altissima (L.) Pall. Plants 2022, 11, 409. https://doi.org/10.3390/plants11030409
Nedelyaeva OI, Popova LG, Volkov VS, Balnokin YV. Molecular Cloning and Characterization of SaCLCd, SaCLCf, and SaCLCg, Novel Proteins of the Chloride Channel Family (CLC) from the Halophyte Suaeda altissima (L.) Pall. Plants. 2022; 11(3):409. https://doi.org/10.3390/plants11030409
Chicago/Turabian StyleNedelyaeva, Olga I., Larissa G. Popova, Vadim S. Volkov, and Yurii V. Balnokin. 2022. "Molecular Cloning and Characterization of SaCLCd, SaCLCf, and SaCLCg, Novel Proteins of the Chloride Channel Family (CLC) from the Halophyte Suaeda altissima (L.) Pall" Plants 11, no. 3: 409. https://doi.org/10.3390/plants11030409
APA StyleNedelyaeva, O. I., Popova, L. G., Volkov, V. S., & Balnokin, Y. V. (2022). Molecular Cloning and Characterization of SaCLCd, SaCLCf, and SaCLCg, Novel Proteins of the Chloride Channel Family (CLC) from the Halophyte Suaeda altissima (L.) Pall. Plants, 11(3), 409. https://doi.org/10.3390/plants11030409






