Plant Age and Soil Texture Rather Than the Presence of Root Hairs Cause Differences in Maize Resource Allocation and Root Gene Expression in the Field
Abstract
:1. Introduction
2. Results
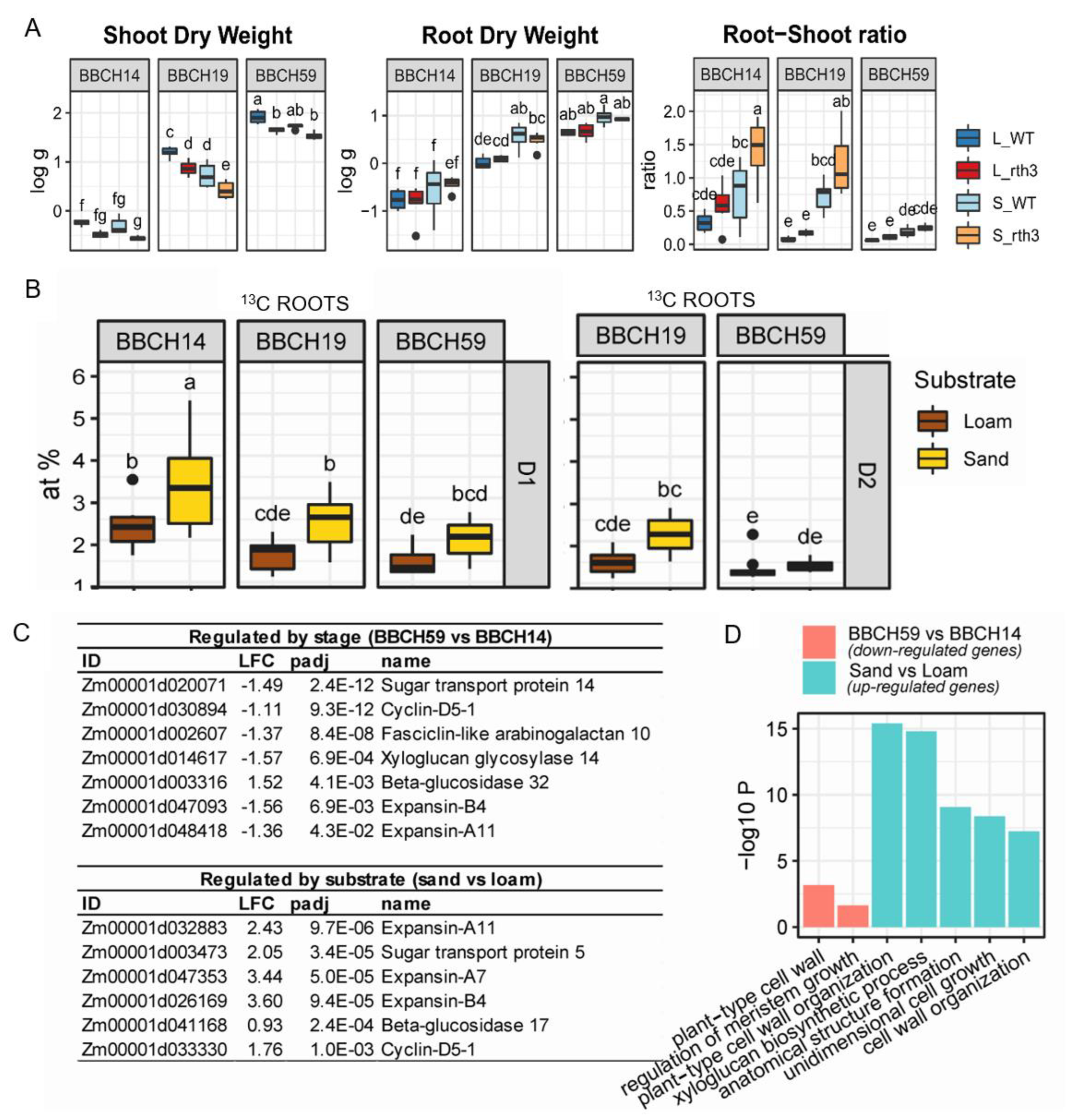
2.1. Maize Growth, Allocation of Recently Assimilated Carbon and Growth-Related Gene Expression
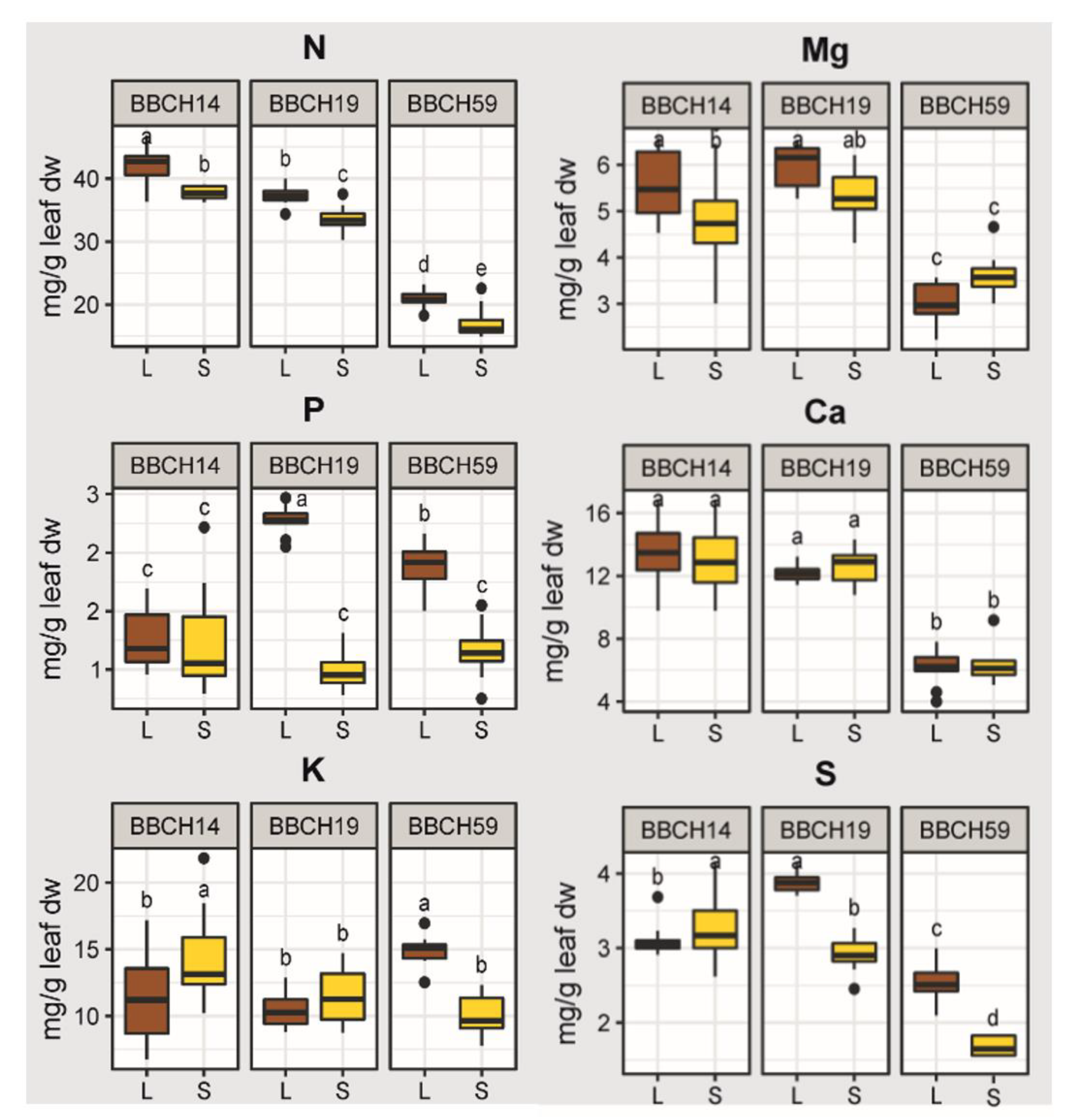
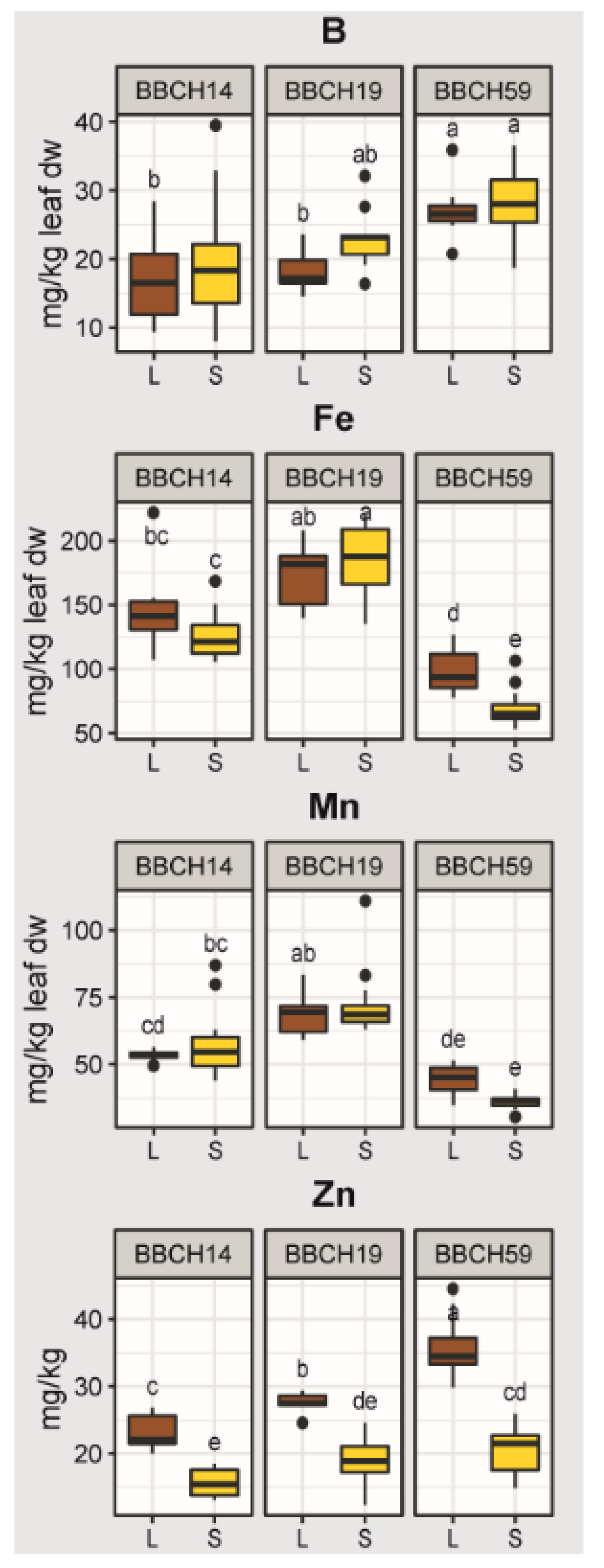
2.2. Effects of Growth Stage, Substrate and Genotype on the Elemental Concentrations of the Youngest Unfolded Leaf of Maize
2.3. Elemental Concentrations in the Remaining Shoot
2.4. Incorporation of Mineral Elements to Maize Shoot Biomass
2.5. The Expression of Root Nutrient Transporter Genes
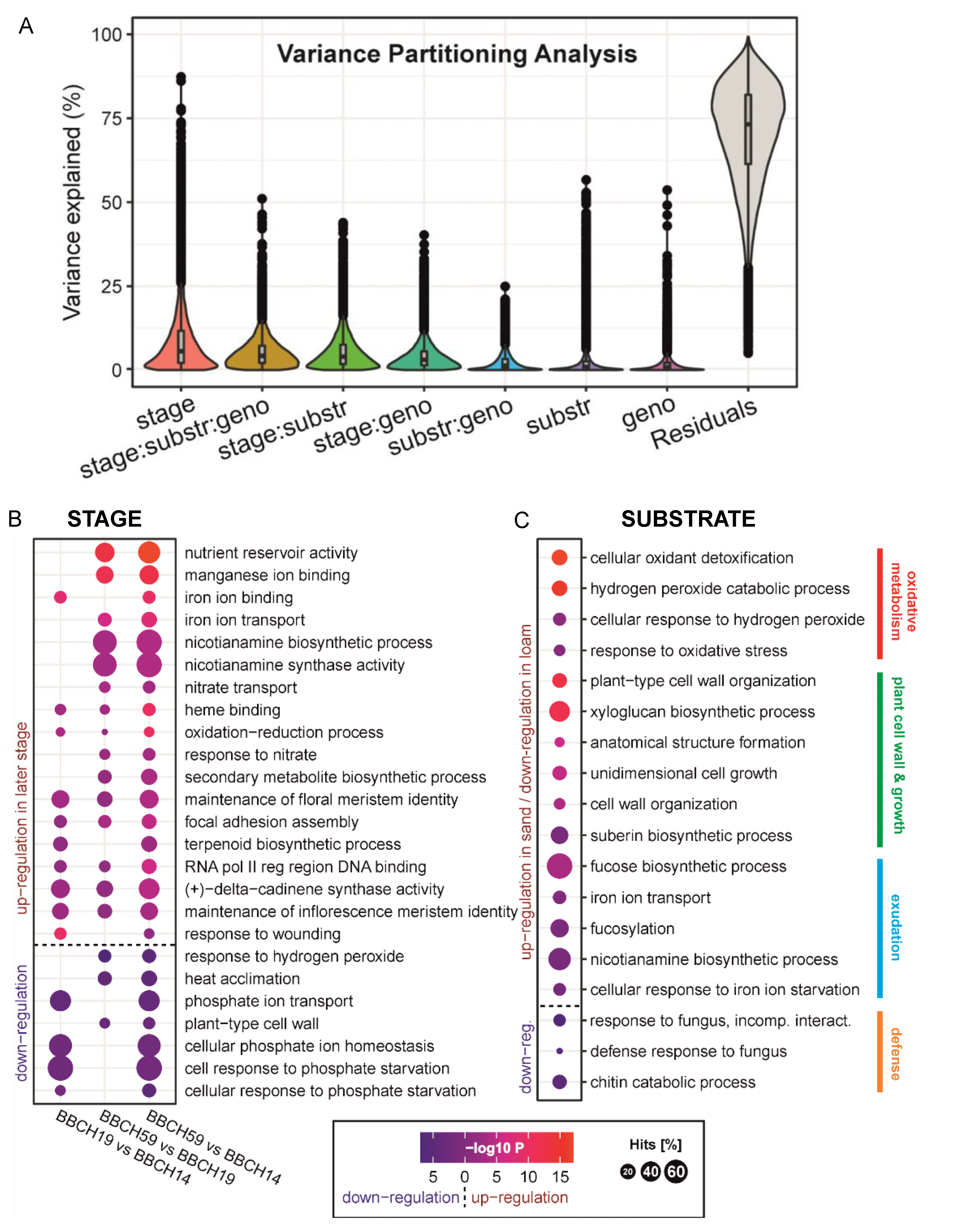
2.6. General Enrichment Patterns of Differentially Expressed Genes (DEGs) in Maize Roots
2.7. Maize Root Gene Expression in Two Depths of the Soil
2.8. The Influence of the Presence of Root Hairs on Maize Root Gene Expression
3. Discussion
3.1. Development, Soil Texture and Drought Drive Maize Gene Expression and Nutritional Status, While the Presence of Root Hairs Supports Shoot Growth
3.2. Lack of Root Hairs Is Reflected by a Small Set of Up-Regulated Genes in the Roots but a Substantial Negative Impact on Maize Shoot Growth
3.3. Growth Stage and Substrate as Drivers of Maize Nutrition
3.4. Decreased Element Concentrations in the Shoot and the Activation of Nutrient Transporters in the Roots
3.5. Young Maize Plants with Higher Carbon Assimilation and Growth Related Gene Expression Rates
3.6. Water Stress towards BBCH59 Leads to Differences between Substrates in Root Plasma Membrane Aquaporin Expression
3.7. Depth Profiles Are More Pronounced at BBCH59 Than BBCH19 and Affect Selected N, P and Aquaporin Genes
4. Materials and Methods
4.1. Experimental Design and Field Study Setup
4.2. Stable Isotope Labeling and Root Sampling
4.3. Root RNA Sequencing
4.4. qPCR Arrays
4.5. Shoot and Leaf Elemental Contents
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Jones, D.L.; Cross, P.; Withers, P.J.A.; DeLuca, T.H.; Robinson, D.A.; Quilliam, R.S.; Harris, I.M.; Chadwick, D.R.; Edwards-Jones, G. Nutrient stripping: The global disparity between food security and soil nutrient stocks. J. Appl. Ecol. 2013, 50, 851–862. [Google Scholar] [CrossRef]
- Cesco, S.; Lucini, L.; Miras-Moreno, B.; Borruso, L.; Mimmo, T.; Pii, Y.; Puglisi, E.; Spini, G.; Taskin, E.; Tiziani, R.; et al. The hidden effects of agrochemicals on Plant metabolism and root-associated microorganisms. Plant Sci. 2021, 311, 111012. [Google Scholar] [CrossRef]
- Biau, A.; Santiveri, F.; Mijangos, I.; Lloveras, J. The impact of organic and mineral fertilizers on soil quality parameters and the productivity of irrigated maize crops in semiarid regions. Eur. J. Soil Biol. 2012, 53, 56–61. [Google Scholar] [CrossRef]
- Lynch, J.P.; Brown, K.M. New roots for agriculture: Exploiting the root phenome. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2012, 367, 1598–1604. [Google Scholar] [CrossRef]
- Cai, G.; Carminati, A.; Abdalla, M.; Ahmed, M.A. Soil textures rather than root hairs dominate water uptake and soil-plant hydraulics under drought. Plant Physiol. 2021, 187, 858–872. [Google Scholar] [CrossRef]
- Hayat, F.; Ahmed, M.A.; Zarebanadkouki, M.; Javaux, M.; Cai, G.; Carminati, A. Transpiration Reduction in Maize (Zea mays L) in Response to Soil Drying. Front. Plant Sci. 2019, 10, 1695. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Körner, C. Paradigm shift in plant growth control. Curr. Opin. Plant Biol. 2015, 25, 107–114. [Google Scholar] [CrossRef] [PubMed]
- Muller, B.; Pantin, F.; Génard, M.; Turc, O.; Freixes, S.; Piques, M.; Gibon, Y. Water deficits uncouple growth from photosynthesis, increase C content, and modify the relationships between C and growth in sink organs. J. Exp. Bot. 2011, 62, 1715–1729. [Google Scholar] [CrossRef] [Green Version]
- Ingestad, T. Relative addition rate and external concentration; Driving variables used in plant nutrition research. Plant Cell Env. 1982, 5, 443–453. [Google Scholar] [CrossRef]
- Huot, B.; Yao, J.; Montgomery, B.L.; He, S.Y. Growth-defense tradeoffs in plants: A balancing act to optimize fitness. Mol. Plant 2014, 7, 1267–1287. [Google Scholar] [CrossRef]
- Schultz, J.C.; Appel, H.M.; Ferrieri, A.P.; Arnold, T.M. Flexible resource allocation during plant defense responses. Front. Plant Sci. 2013, 4, 324. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tarkka, M.T.; Grams, T.E.E.; Angay, O.; Kurth, F.; Maboreke, H.R.; Mailänder, S.; Bönn, M.; Feldhahn, L.; Fleischmann, F.; Ruess, L.; et al. Ectomycorrhizal fungus supports endogenous rhythmic growth and corresponding resource allocation in oak during various below- and aboveground biotic interactions. Sci. Rep. 2021, 11, 23680. [Google Scholar] [CrossRef] [PubMed]
- York, L.M.; Lynch, J.P. Intensive field phenotyping of maize (Zea mays L.) root crowns identifies phenes and phene integration associated with plant growth and nitrogen acquisition. J. Exp. Bot. 2015, 66, 5493–5505. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tai, H.; Lu, X.; Opitz, N.; Marcon, C.; Paschold, A.; Lithio, A.; Nettleton, D.; Hochholdinger, F. Transcriptomic and anatomical complexity of primary, seminal, and crown roots highlight root type-specific functional diversity in maize (Zea mays L.). J. Exp. Bot. 2016, 67, 1123–1135. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yu, P.; Gutjahr, C.; Li, C.; Hochholdinger, F. Genetic Control of Lateral Root Formation in Cereals. Trends Plant Sci. 2016, 21, 951–961. [Google Scholar] [CrossRef]
- Zhan, A.; Lynch, J.P. Reduced frequency of lateral root branching improves N capture from low-N soils in maize. J. Exp. Bot. 2015, 66, 2055–2065. [Google Scholar] [CrossRef] [Green Version]
- Zhan, A.; Schneider, H.; Lynch, J.P. Reduced Lateral Root Branching Density Improves Drought Tolerance in Maize. Plant Physiol. 2015, 168, 1603–1615. [Google Scholar] [CrossRef] [Green Version]
- Hill, C.B.; Cassin, A.; Keeble-Gagnère, G.; Doblin, M.S.; Bacic, A.; Roessner, U. De novo transcriptome assembly and analysis of differentially expressed genes of two barley genotypes reveal root-zone-specific responses to salt exposure. Sci. Rep. 2016, 6, 31558. [Google Scholar] [CrossRef] [Green Version]
- Stelpflug, S.C.; Sekhon, R.S.; Vaillancourt, B.; Hirsch, C.N.; Buell, C.R.; de Leon, N.; Kaeppler, S.M. An Expanded Maize Gene Expression Atlas based on RNA Sequencing and its Use to Explore Root Development. Plant Genome 2016, 9, 1–16. [Google Scholar] [CrossRef]
- Rongsawat, T.; Peltier, J.-B.; Boyer, J.-C.; Véry, A.-A.; Sentenac, H. Looking for Root Hairs to Overcome Poor Soils. Trends Plant Sci. 2021, 26, 83–94. [Google Scholar] [CrossRef]
- Bienert, M.D.; Werner, L.M.; Wimmer, M.A.; Bienert, G.P. Root hairs: The villi of plants. Biochem. Soc. Trans. 2021, 49, 1133–1146. [Google Scholar] [CrossRef] [PubMed]
- Haling, R.E.; Brown, L.K.; Bengough, A.G.; Young, I.M.; Hallett, P.D.; White, P.J.; George, T.S. Root hairs improve root penetration, root-soil contact, and phosphorus acquisition in soils of different strength. J. Exp. Bot. 2013, 64, 3711–3721. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marzec, M.; Melzer, M.; Szarejko, I. Root hair development in the grasses: What we already know and what we still need to know. Plant Physiol. 2015, 168, 407–414. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Carminati, A.; Passioura, J.B.; Zarebanadkouki, M.; Ahmed, M.A.; Ryan, P.R.; Watt, M.; Delhaize, E. Root hairs enable high transpiration rates in drying soils. New Phytol. 2017, 216, 771–781. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marin, M.; Feeney, D.S.; Brown, L.K.; Naveed, M.; Ruiz, S.; Koebernick, N.; Bengough, A.G.; Hallett, P.D.; Roose, T.; Puértolas, J.; et al. Significance of root hairs for plant performance under contrasting field conditions and water deficit. Ann. Bot. 2021, 128, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Stein, R.J.; Höreth, S.; de Melo, J.R.F.; Syllwasschy, L.; Lee, G.; Garbin, M.L.; Clemens, S.; Krämer, U. Relationships between soil and leaf mineral composition are element-specific, environment-dependent and geographically structured in the emerging model Arabidopsis halleri. New Phytol. 2017, 213, 1274–1286. [Google Scholar] [CrossRef] [Green Version]
- Pii, Y.; Cesco, S.; Mimmo, T. Shoot ionome to predict the synergism and antagonism between nutrients as affected by substrate and physiological status. Plant Physiol. Biochem. 2015, 94, 48–56. [Google Scholar] [CrossRef]
- Tiziani, R.; Mimmo, T.; Valentinuzzi, F.; Pii, Y.; Celletti, S.; Cesco, S. Root Handling Affects Carboxylates Exudation and Phosphate Uptake of White Lupin Roots. Front. Plant Sci. 2020, 11, 584568. [Google Scholar] [CrossRef]
- Baxter, I. Should we treat the ionome as a combination of individual elements, or should we be deriving novel combined traits? J. Exp. Bot. 2015, 66, 2127–2131. [Google Scholar] [CrossRef] [Green Version]
- Yu, P.; Wang, C.; Baldauf, J.A.; Tai, H.; Gutjahr, C.; Hochholdinger, F.; Li, C. Root type and soil phosphate determine the taxonomic landscape of colonizing fungi and the transcriptome of field-grown maize roots. New Phytol. 2018, 217, 1240–1253. [Google Scholar] [CrossRef]
- Hu, J.; Lin, X.; Wang, J.; Dai, J.; Cui, X.; Chen, R.; Zhang, J. Arbuscular mycorrhizal fungus enhances crop yield and P-uptake of maize (Zea mays L.): A field case study on a sandy loam soil as affected by long-term P-deficiency fertilization. Soil Biol. Biochem. 2009, 41, 2460–2465. [Google Scholar] [CrossRef]
- Kobayashi, T.; Nishizawa, N.K. Iron sensors and signals in response to iron deficiency. Plant Sci. 2014, 224, 36–43. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sattelmacher, B.; Thoms, K. Root Growth and 14 C-Translocation into the Roots of Maize (Zea mays L.) as influenced by local Nitrate Supply. Z. Pflanzenernaehr. Bodenk. 1989, 152, 7–10. [Google Scholar] [CrossRef]
- Mounier, E.; Pervent, M.; Ljung, K.; Gojon, A.; Nacry, P. Auxin-mediated nitrate signalling by NRT1.1 participates in the adaptive response of Arabidopsis root architecture to the spatial heterogeneity of nitrate availability. Plant Cell Env. 2014, 37, 162–174. [Google Scholar] [CrossRef]
- Vetterlein, D.; Lippold, E.; Schreiter, S.; Phalempin, M.; Fahrenkampf, T.; Hochholdinger, F.; Marcon, C.; Tarkka, M.; Oburger, E.; Ahmed, M.A.; et al. Experimental platforms for the investigation of spatiotemporal patterns in the rhizosphere—Laboratory and field scale. J. Plant Nutr. Soil Sci. 2021, 184, 35–50. [Google Scholar] [CrossRef]
- Vetterlein, D.; Carminati, A.; Kögel-Knabner, I.; Bienert, G.P.; Smalla, K.; Oburger, E.; Schnepf, A.; Banitz, T.; Tarkka, M.T.; Schlüter, S. Rhizosphere Spatiotemporal Organization–A Key to Rhizosphere Functions. Front. Agron. 2020, 2, 8. [Google Scholar] [CrossRef]
- Lippold, E.; Phalempin, M.; Schlüter, S.; Vetterlein, D. Does the lack of root hairs alter root system architecture of Zea mays? Plant Soil 2021, 467, 267–286. [Google Scholar] [CrossRef]
- Ganther, M.; Vetterlein, D.; Heintz-Buschart, A.; Tarkka, M.T. Transcriptome sequencing analysis of maize roots reveals the effects of substrate and root hair formation in a spatial context. Plant Soil 2021, 478, 211–228. [Google Scholar] [CrossRef]
- Vetterlein, D.; Phalempin, M.; Lippold, E.; Schlüter, S.; Schreiter, S.; Ahmed, M.A.; Carminati, A.; Duddek, P.; Jorda, H.; Bienert, P.; et al. Root Hairs Matter at Field Scale for Maize Shoot Growth and Nutrient Uptake, But Root Trait Plasticity is Primarily Triggered by Texture and Drought. Plant Soil 2022, 478, 119–141. [Google Scholar] [CrossRef]
- Bilyera, N.; Hummel, C.; Daudin, G.; Santangeli, M.; Zhang, X.; Santner, J.; Lippold, E.; Schlüter, S.; Bertrand, I.; Wenzel, W.; et al. Co-localised phosphorus mobilization processes in the rhizosphere of field-grown maize jointly contribute to plant nutrition. Soil Biol. Biochem. 2022, 165, 108497. [Google Scholar] [CrossRef]
- Jorda, H.; Ahmed, M.A.; Javaux, M.; Carminati, A.; Duddek, P.; Vetterlein, D.; Vanderborght, J. Field scale plant water relation of maize (Zea mays) under drought—Impact of root hairs and soil texture. Plant Soil 2022, 478, 59–84. [Google Scholar] [CrossRef]
- Acosta-Gamboa, L.M.; Liu, S.; Langley, E.; Campbell, Z.; Castro-Guerrero, N.; Mendoza-Cozatl, D.; Lorence, A. Moderate to severe water limitation differentially affects the phenome and ionome of Arabidopsis. Funct. Plant Biol. 2016, 44, 94–106. [Google Scholar] [CrossRef] [PubMed]
- Peuke, A.D.; Rennenberg, H. Impacts of drought on mineral macro- and microelements in provenances of beech (Fagus sylvatica L.) seedlings. Tree Physiol. 2011, 31, 196–207. [Google Scholar] [CrossRef] [PubMed]
- Bergmann, W. Farbatlas—Ernährungsstörungen bei Kulturpflanzen: Visuelle und Analytische Diagnose; VEB Gustav Fischer Verlag: Jena, Germany, 1986. [Google Scholar]
- Cordero, E.; Longchamps, L.; Khosla, R.; Sacco, D. Spatial management strategies for nitrogen in maize production based on soil and crop data. Sci. Total Environ. 2019, 697, 133854. [Google Scholar] [CrossRef]
- Zhu, J.; Kaeppler, S.M.; Lynch, J.P. Mapping of QTL controlling root hair length in maize (Zea mays L.) under phosphorus deficiency. Plant Soil 2005, 270, 299–310. [Google Scholar] [CrossRef]
- Wang, D.; Lv, S.; Jiang, P.; Li, Y. Roles, Regulation, and Agricultural Application of Plant Phosphate Transporters. Front. Plant Sci. 2017, 8, 817. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Klamer, F.; Vogel, F.; Li, X.; Bremer, H.; Neumann, G.; Neuhäuser, B.; Hochholdinger, F.; Ludewig, U. Estimating the importance of maize root hairs in low phosphorus conditions and under drought. Ann. Bot. 2019, 124, 961–968. [Google Scholar] [CrossRef] [PubMed]
- Earl, H.J.; Liu, W.; Bowley, S.R.; Tollenaar, M. Effects of Abiotic Stress on Respiratory Carbon Loss of Two Maize (Zea mays L.) Inbred Lines and Their Hybrid During Silking and Grain-Filling. Crop Sci. 2012, 52, 1795–1802. [Google Scholar] [CrossRef]
- Benešová, M.; Holá, D.; Fischer, L.; Jedelský, P.L.; Hnilička, F.; Wilhelmová, N.; Rothová, O.; Kočová, M.; Procházková, D.; Honnerová, J.; et al. The physiology and proteomics of drought tolerance in maize: Early stomatal closure as a cause of lower tolerance to short-term dehydration? PLoS ONE 2012, 7, e38017. [Google Scholar] [CrossRef]
- Hochholdinger, F.; Wen, T.-J.; Zimmermann, R.; Chimot-Marolle, P.; Da Costa e Silva, O.; Bruce, W.; Lamkey, K.R.; Wienand, U.; Schnable, P.S. The maize (Zea mays L.) roothairless3 gene encodes a putative GPI-anchored, monocot-specific, COBRA-like protein that significantly affects grain yield. Plant J. 2008, 54, 888–898. [Google Scholar] [CrossRef]
- Brady, S.M.; Song, S.; Dhugga, K.S.; Rafalski, J.A.; Benfey, P.N. Combining expression and comparative evolutionary analysis. The COBRA gene family. Plant Physiol. 2007, 143, 172–187. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bista, D.R.; Heckathorn, S.A.; Jayawardena, D.M.; Mishra, S.; Boldt, J.K. Effects of Drought on Nutrient Uptake and the Levels of Nutrient-Uptake Proteins in Roots of Drought-Sensitive and -Tolerant Grasses. Plants 2018, 7, 28. [Google Scholar] [CrossRef] [Green Version]
- Dodig, D.; Božinović, S.; Nikolić, A.; Zorić, M.; Vančetović, J.; Ignjatović-Micić, D.; Delić, N.; Weigelt-Fischer, K.; Altmann, T.; Junker, A. Dynamics of Maize Vegetative Growth and Drought Adaptability Using Image-Based Phenotyping Under Controlled Conditions. Front. Plant Sci. 2021, 12, 652116. [Google Scholar] [CrossRef] [PubMed]
- He, M.; Dijkstra, F.A. Drought effect on plant nitrogen and phosphorus: A meta-analysis. New Phytol. 2014, 204, 924–931. [Google Scholar] [CrossRef] [PubMed]
- Stich, B.; Benke, A.; Schmidt, M.; Urbany, C.; Shi, R.; von Wirén, N. The maize shoot ionome: Its interaction partners, predictive power, and genetic determinants. Plant Cell Environ. 2020, 43, 2095–2111. [Google Scholar] [CrossRef]
- Baxter, I.R.; Vitek, O.; Lahner, B.; Muthukumar, B.; Borghi, M.; Morrissey, J.; Guerinot, M.L.; Salt, D.E. The leaf ionome as a multivariable system to detect a plant’s physiological status. Proc. Natl. Acad. Sci. USA 2008, 105, 12081–12086. [Google Scholar] [CrossRef] [Green Version]
- Courbet, G.; D’Oria, A.; Lornac, A.; Diquélou, S.; Pluchon, S.; Arkoun, M.; Koprivova, A.; Kopriva, S.; Etienne, P.; Ourry, A. Specificity and Plasticity of the Functional Ionome of Brassica napus and Triticum aestivum Subjected to Macronutrient Deprivation. Front. Plant Sci. 2021, 12, 641648. [Google Scholar] [CrossRef]
- Watanabe, T.; Urayama, M.; Shinano, T.; Okada, R.; Osaki, M. Application of ionomics to plant and soil in fields under long-term fertilizer trials. Springerplus 2015, 4, 781. [Google Scholar] [CrossRef]
- Kumar, S.; Kumar, S.; Mohapatra, T. Interaction Between Macro- and Micro-Nutrients in Plants. Front. Plant Sci. 2021, 12, 665583. [Google Scholar] [CrossRef]
- Briat, J.-F.; Rouached, H.; Tissot, N.; Gaymard, F.; Dubos, C. Integration of P, S, Fe, and Zn nutrition signals in Arabidopsis thaliana: Potential involvement of PHOSPHATE STARVATION RESPONSE 1 (PHR1). Front. Plant Sci. 2015, 6, 290. [Google Scholar] [CrossRef]
- Chen, Q.; Mu, X.; Chen, F.; Yuan, L.; Mi, G. Dynamic change of mineral nutrient content in different plant organs during the grain filling stage in maize grown under contrasting nitrogen supply. Eur. J. Agron. 2016, 80, 137–153. [Google Scholar] [CrossRef] [Green Version]
- Maillard, A.; Diquélou, S.; Billard, V.; Laîné, P.; Garnica, M.; Prudent, M.; Garcia-Mina, J.-M.; Yvin, J.-C.; Ourry, A. Leaf mineral nutrient remobilization during leaf senescence and modulation by nutrient deficiency. Front. Plant Sci. 2015, 6, 317. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bouain, N.; Shahzad, Z.; Rouached, A.; Khan, G.A.; Berthomieu, P.; Abdelly, C.; Poirier, Y.; Rouached, H. Phosphate and zinc transport and signalling in plants: Toward a better understanding of their homeostasis interaction. J. Exp. Bot. 2014, 65, 5725–5741. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Trevisan, S.; Borsa, P.; Botton, A.; Varotto, S.; Malagoli, M.; Ruperti, B.; Quaggiotti, S. Expression of two maize putative nitrate transporters in response to nitrate and sugar availability. Plant Biol. 2008, 10, 462–475. [Google Scholar] [CrossRef] [PubMed]
- Yun, S.J.; Kaeppler, S.M. Induction of maize acid phosphatase activities under phosphorus starvation. Plant Soil 2001, 237, 109–115. [Google Scholar] [CrossRef]
- Takahashi, M.; Terada, Y.; Nakai, I.; Nakanishi, H.; Yoshimura, E.; Mori, S.; Nishizawa, N.K. Role of nicotianamine in the intracellular delivery of metals and plant reproductive development. Plant Cell 2003, 15, 1263–1280. [Google Scholar] [CrossRef]
- Hanikenne, M.; Esteves, S.M.; Fanara, S.; Rouached, H. Coordinated homeostasis of essential mineral nutrients: A focus on iron. J. Exp. Bot. 2021, 72, 2136–2153. [Google Scholar] [CrossRef]
- Nozoye, T.; Nagasaka, S.; Kobayashi, T.; Takahashi, M.; Sato, Y.; Sato, Y.; Uozumi, N.; Nakanishi, H.; Nishizawa, N.K. Phytosiderophore efflux transporters are crucial for iron acquisition in graminaceous plants. J. Biol. Chem. 2011, 286, 5446–5454. [Google Scholar] [CrossRef] [Green Version]
- von Wiren, N.; Marschner, H.; Romheld, V. Roots of Iron-Efficient Maize also Absorb Phytosiderophore-Chelated Zinc. Plant Physiol. 1996, 111, 1119–1125. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gargallo-Garriga, A.; Sardans, J.; Pérez-Trujillo, M.; Rivas-Ubach, A.; Oravec, M.; Vecerova, K.; Urban, O.; Jentsch, A.; Kreyling, J.; Beierkuhnlein, C.; et al. Opposite metabolic responses of shoots and roots to drought. Sci. Rep. 2014, 4, 6829. [Google Scholar] [CrossRef]
- Wu, Y.; Thorne, E.T.; Sharp, R.E.; Cosgrove, D.J. Modification of expansin transcript levels in the maize primary root at low water potentials. Plant Physiol. 2001, 126, 1471–1479. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ren, B.; Dong, S.; Zhao, B.; Liu, P.; Zhang, J. Responses of Nitrogen Metabolism, Uptake and Translocation of Maize to Waterlogging at Different Growth Stages. Front. Plant Sci. 2017, 8, 1216. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Araus, J.L.; Serret, M.D.; Edmeades, G.O. Phenotyping maize for adaptation to drought. Front. Physiol. 2012, 3, 305. [Google Scholar] [CrossRef] [Green Version]
- Çakir, R. Effect of water stress at different development stages on vegetative and reproductive growth of corn. Field Crops Res. 2004, 89, 1–16. [Google Scholar] [CrossRef]
- Pandey, R.; Maranville, J.; Chetima, M. Deficit irrigation and nitrogen effects on maize in a Sahelian environment. Agric. Water Manag. 2000, 46, 15–27. [Google Scholar] [CrossRef]
- Collins, B.; Chapman, S.; Hammer, G.; Chenu, K. Limiting transpiration rate in high evaporative demand conditions to improve Australian wheat productivity. Plants 2021, 3, diab006. [Google Scholar] [CrossRef]
- Bacher, H.; Sharaby, Y.; Walia, H.; Peleg, Z. Modify Root/Shoot ratio Alleviate Root Water Influxes in Wheat under Drought Stress. J. Exp. Bot. 2021, 73, 1643–1654. [Google Scholar] [CrossRef]
- Messina, C.D.; Sinclair, T.R.; Hammer, G.L.; Curan, D.; Thompson, J.; Oler, Z.; Gho, C.; Cooper, M. Limited-Transpiration Trait May Increase Maize Drought Tolerance in the US Corn Belt. Agron. J. 2015, 107, 1978–1986. [Google Scholar] [CrossRef] [Green Version]
- Siefritz, F.; Tyree, M.T.; Lovisolo, C.; Schubert, A.; Kaldenhoff, R. PIP1 plasma membrane aquaporins in tobacco: From cellular effects to function in plants. Plant Cell 2002, 14, 869–876. [Google Scholar] [CrossRef] [Green Version]
- Aroca, R.; Porcel, R.; Ruiz-Lozano, J.M. Regulation of root water uptake under abiotic stress conditions. J. Exp. Bot. 2012, 63, 43–57. [Google Scholar] [CrossRef]
- Šurbanovski, N.; Sargent, D.J.; Else, M.A.; Simpson, D.W.; Zhang, H.; Grant, O.M. Expression of Fragaria vesca PIP aquaporins in response to drought stress: PIP down-regulation correlates with the decline in substrate moisture content. PLoS ONE 2013, 8, e74945. [Google Scholar] [CrossRef] [Green Version]
- Galmés, J.; Pou, A.; Alsina, M.M.; Tomàs, M.; Medrano, H.; Flexas, J. Aquaporin expression in response to different water stress intensities and recovery in Richter-110 (Vitis sp.): Relationship with ecophysiological status. Planta 2007, 226, 671–681. [Google Scholar] [CrossRef]
- Del Carmen Martinez-Ballesta, M.; Del Carmen Rodriguez-Hernandez, M.; Alcaraz-Lopez, C.; Mota-Cadenas, C.; Muries, B.; Carvajal, M. Plant Hydraulic Conductivity: The Aquaporins Contribution. In Hydraulic Conductivity—Issues, Determination and Applications; Elango, L., Ed.; InTech: Rijeka, Croatia, 2011; ISBN 978-953-307-288-3. [Google Scholar]
- Wiesler, F.; Horst, W.J. Root growth of maize cultivars under field conditions as studied by the core and minirhizotron method and relationships to shoot growth. Z. Pflanzenernaehr. Bodenk. 1994, 157, 351–358. [Google Scholar] [CrossRef]
- Zhang, J.; Zhang, X.; Liang, J. Exudation rate and hydraulic conductivity of maize roots are enhanced by soil drying and abscisic acid treatment. New Phytol. 1995, 131, 329–336. [Google Scholar] [CrossRef]
- Nagy, R.; Vasconcelos, M.J.V.; Zhao, S.; McElver, J.; Bruce, W.; Amrhein, N.; Raghothama, K.G.; Bucher, M. Differential regulation of five Pht1 phosphate transporters from maize (Zea mays L.). Plant Biol. 2006, 8, 186–197. [Google Scholar] [CrossRef]
- Liu, F.; Xu, Y.; Jiang, H.; Jiang, C.; Du, Y.; Gong, C.; Wang, W.; Zhu, S.; Han, G.; Cheng, B. Systematic Identification, Evolution and Expression Analysis of the Zea mays PHT1 Gene Family Reveals Several New Members Involved in Root Colonization by Arbuscular Mycorrhizal Fungi. Int. J. Mol. Sci. 2016, 17, 930. [Google Scholar] [CrossRef] [Green Version]
- Lancashire, P.D.; Bleiholder, H.; den van BOOM, T.; Langelüddeke, P.; Stauss, R.; Weber, E.; Witzenberger, A. A uniform decimal code for growth stages of crops and weeds. Ann. Appl. Biol. 1991, 119, 561–601. [Google Scholar] [CrossRef]
- Ganther, M.; Yim, B.; Ibrahim, Z.; Bienert, M.D.; Lippold, E.; Maccario, L.; Sørensen, S.J.; Bienert, G.P.; Vetterlein, D.; Heintz-Buschart, A.; et al. Compatibility of X-ray computed tomography with plant gene expression, rhizosphere bacterial communities and enzyme activities. J. Exp. Bot. 2020, 71, 5603–5614. [Google Scholar] [CrossRef]
- Andrews, S. FastQC—A Quality Control Tool for High Throughput Sequence Data. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 11 August 2020).
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [Green Version]
- Zerbino, D.R.; Achuthan, P.; Akanni, W.; Amode, M.R.; Barrell, D.; Bhai, J.; Billis, K.; Cummins, C.; Gall, A.; Girón, C.G.; et al. Ensembl 2018. Nucleic Acids Res. 2018, 46, D754–D761. [Google Scholar] [CrossRef]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liao, Y.; Smyth, G.K.; Shi, W. featureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2017. [Google Scholar]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. B 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Young, M.D.; Wakefield, M.J.; Smyth, G.K.; Oshlack, A. Gene ontology analysis for RNA-seq: Accounting for selection bias: Accounting for selection bias. Genome Biol. 2010, 11, R14. [Google Scholar] [CrossRef] [Green Version]
- Wimalanathan, K.; Friedberg, I.; Andorf, C.M.; Lawrence-Dill, C.J. Maize GO Annotation-Methods, Evaluation, and Review (maize-GAMER). Plant Direct. 2018, 2, e00052. [Google Scholar] [CrossRef]
- Oksanen, J.; Blanchet, F.G.; Friendly, M.; Kindt, R.; Legendre, P.; McGlinn, D.; Minchin, P.R.; O’Hara, R.B.; Simpson, G.L.; Solymos, P.; et al. Vegan: Community Ecology Package, 2020. Available online: https://CRAN.R-project.org/package=vegan (accessed on 1 November 2021).
- Hoffman, G.E.; Schadt, E.E. VariancePartition: Interpreting drivers of variation in complex gene expression studies. BMC Bioinform. 2016, 17, 483. [Google Scholar] [CrossRef] [Green Version]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative C(T) method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ganther, M.; Lippold, E.; Bienert, M.D.; Bouffaud, M.-L.; Bauer, M.; Baumann, L.; Bienert, G.P.; Vetterlein, D.; Heintz-Buschart, A.; Tarkka, M.T. Plant Age and Soil Texture Rather Than the Presence of Root Hairs Cause Differences in Maize Resource Allocation and Root Gene Expression in the Field. Plants 2022, 11, 2883. https://doi.org/10.3390/plants11212883
Ganther M, Lippold E, Bienert MD, Bouffaud M-L, Bauer M, Baumann L, Bienert GP, Vetterlein D, Heintz-Buschart A, Tarkka MT. Plant Age and Soil Texture Rather Than the Presence of Root Hairs Cause Differences in Maize Resource Allocation and Root Gene Expression in the Field. Plants. 2022; 11(21):2883. https://doi.org/10.3390/plants11212883
Chicago/Turabian StyleGanther, Minh, Eva Lippold, Manuela Désirée Bienert, Marie-Lara Bouffaud, Mario Bauer, Louis Baumann, Gerd Patrick Bienert, Doris Vetterlein, Anna Heintz-Buschart, and Mika Tapio Tarkka. 2022. "Plant Age and Soil Texture Rather Than the Presence of Root Hairs Cause Differences in Maize Resource Allocation and Root Gene Expression in the Field" Plants 11, no. 21: 2883. https://doi.org/10.3390/plants11212883
APA StyleGanther, M., Lippold, E., Bienert, M. D., Bouffaud, M.-L., Bauer, M., Baumann, L., Bienert, G. P., Vetterlein, D., Heintz-Buschart, A., & Tarkka, M. T. (2022). Plant Age and Soil Texture Rather Than the Presence of Root Hairs Cause Differences in Maize Resource Allocation and Root Gene Expression in the Field. Plants, 11(21), 2883. https://doi.org/10.3390/plants11212883







