Phytochemical Analysis of the Methanolic Extract and Essential Oil from Leaves of Industrial Hemp Futura 75 Cultivar: Isolation of a New Cannabinoid Derivative and Biological Profile Using Computational Approaches
Abstract
:1. Introduction
2. Results and Discussion
2.1. Phytochemical Analysis of Methanol Extracts
2.2. Analysis of Volatile Compounds from Leaves of C. sativa L. vr. Futura 75
2.3. Major Phytochemical Components in Futura 75 Water Infusion
2.4. Inverse Virtual Screening
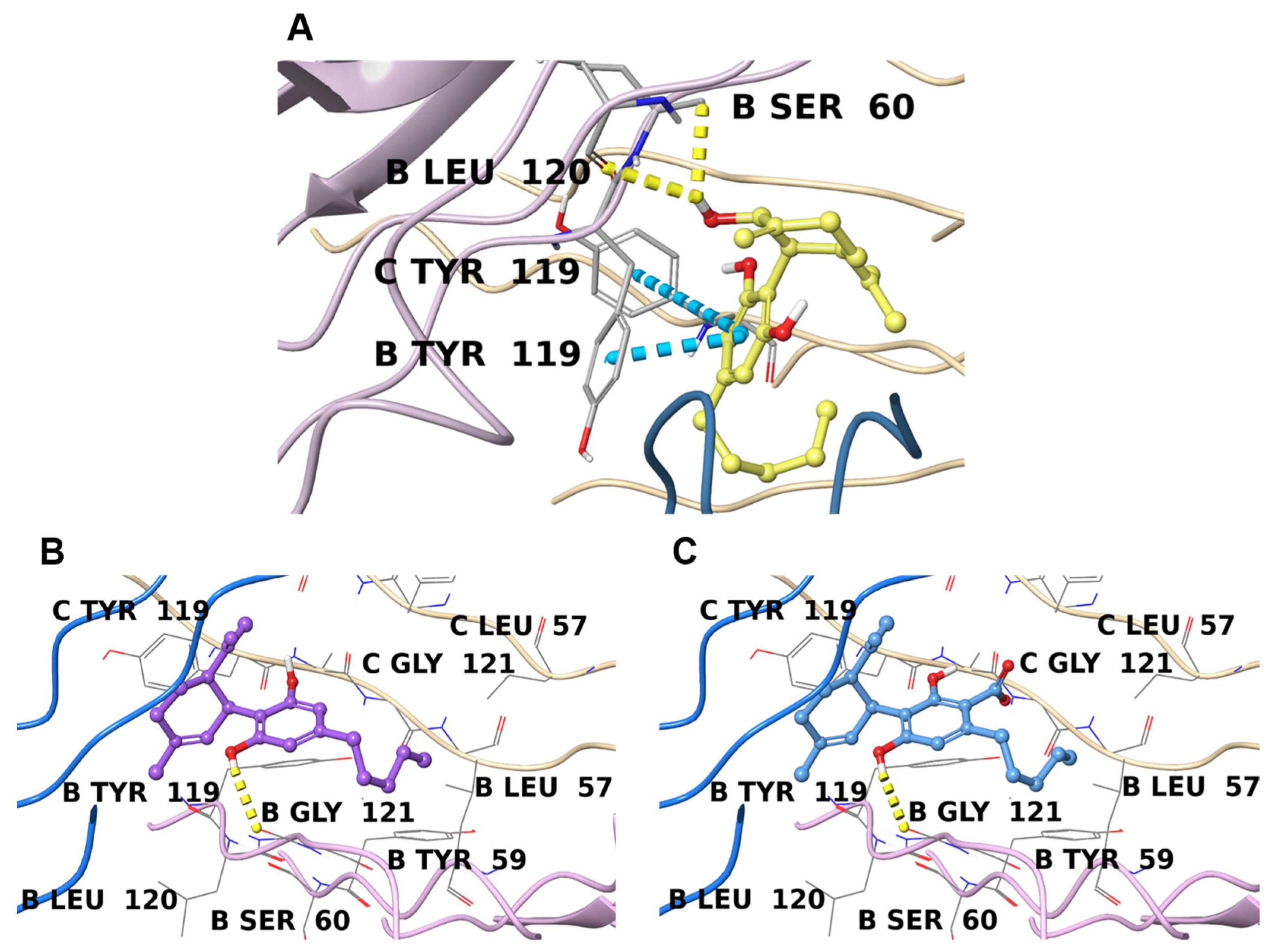
2.4.1. TNFα
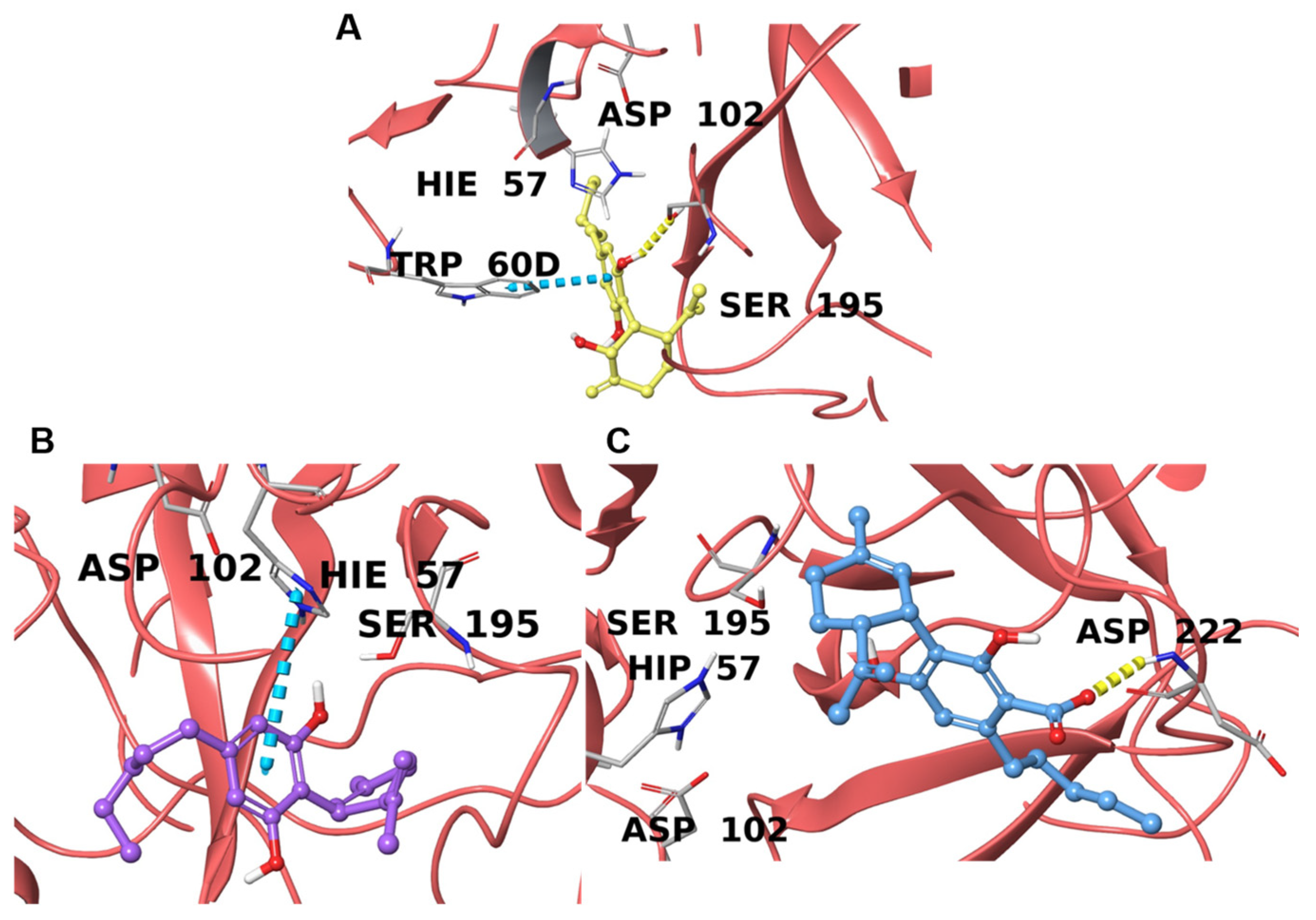
2.4.2. Prothrombin/Thrombin
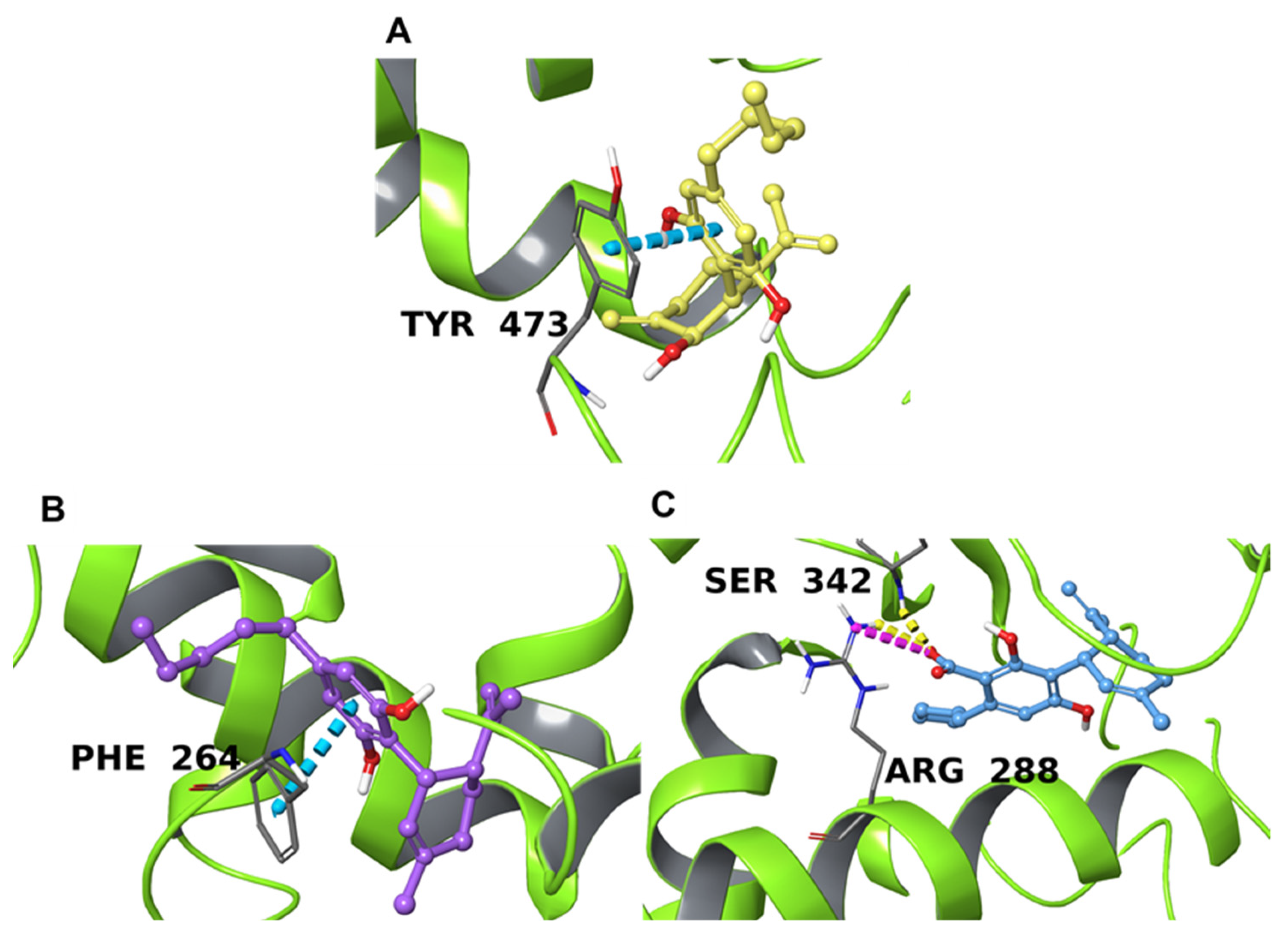
2.4.3. PPARγ
3. Materials and Methods
3.1. General Experimental Procedure
3.2. Plant Material
3.3. Extraction and Isolation
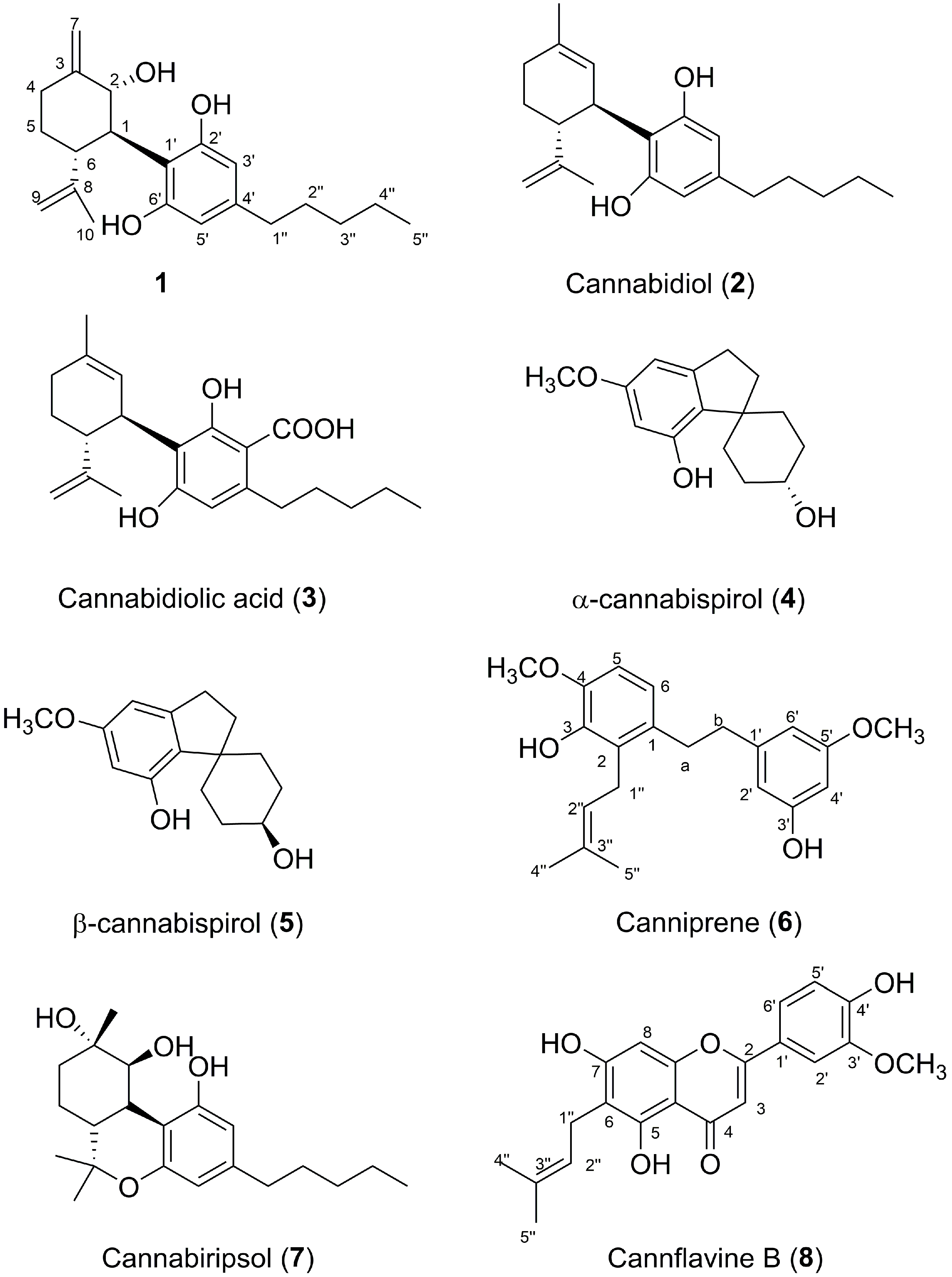
3.3.1. 2α-Hydroxy-Δ3,7-Cannabitriol (1)
3.3.2. Cannabidiol (2)
3.3.3. Cannabidiolic Acid (3)
3.3.4. α-Cannabispiranol (4) and β-Cannabispiranol (5)
3.3.5. Canniprene (6)
3.3.6. Cannabiripsol (7)
3.3.7. Cannflavin B (8)
3.4. Essential Oil Preparation
3.5. GC-FID and Gas Chromatography/Mass Spectrometry (GC/MS) Analysis
3.6. Identification of Essential Oil Components
3.7. Water Infusion Preparation
3.8. Input File Preparation
3.9. Inverse Virtual Screening
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Clarke, R.; Merlin, M. Cannabis: Evolution and Ethnobotany; University of California Press: Oakland, CA, USA, 2016. [Google Scholar]
- Pattnaik, F.; Nanda, S.; Mohanty, S.; Dalai, A.K.; Kumar, V.; Ponnusamy, S.K.; Naik, S. Cannabis: Chemistry, extraction and therapeutic applications. Chemosphere 2022, 289, 133012. [Google Scholar] [CrossRef] [PubMed]
- Iftikhar, A.; Zafar, U.; Ahmed, W.; Shabbir, M.A.; Sameen, A.; Sahar, A.; Bhat, Z.F.; Kowalczewski, P.Ł.; Jarzębski, M.; Aadil, R.M. Applications of Cannabis sativa L. in food and its therapeutic potential: From a prohibited drug to a nutritional supplement. Molecules 2021, 26, 7699. [Google Scholar] [CrossRef] [PubMed]
- Bonini, S.A.; Premoli, M.; Tambaro, S.; Kumar, A.; Maccarinelli, G.; Memo, M.; Mastinu, A. Cannabis sativa: A comprehensive ethnopharmacological review of a medicinal plant with a long history. J. Ethnopharmacol. 2018, 227, 300–315. [Google Scholar] [CrossRef] [PubMed]
- ElSohly, M.A.; Slade, D. Chemical constituents of marijuana: The complex mixture of natural cannabinoids. Life Sci. 2005, 78, 539–548. [Google Scholar] [CrossRef]
- Appendino, G.; Chianese, G.; Taglialatela-Scafati, O. Cannabinoids: Occurrence and medicinal chemistry. Curr. Med. Chem. 2011, 18, 1085–1099. [Google Scholar] [CrossRef]
- De Meijer, E.P.M. The Chemical Phenotypes (Chemotypes) of Cannabis; Oxford University Press: Oxford, UK, 2014; Volume 89, p. 110. [Google Scholar]
- Hartsel, J.A.; Eades, J.; Hickory, B.; Makriyannis, A. Chapter 53—Cannabis sativa and hemp. In Nutraceuticals; Gupta, R.C., Ed.; Academic Press: Boston, MA, USA, 2016; pp. 735–754. [Google Scholar]
- Rules for Direct Payments to Farmers under Support Schemes within the Framework of the Common Agricultural Policy and Repealing Council Regulation (EC) No 637/2008 and Council Regulation (EC) No 73/2009. Available online: https://eur-lex.europa.eu/legal-content/EN/TXT/?uri=celex:32013R1307 (accessed on 1 March 2021).
- Plant Variety Catalogues, Databases & Information Systems. Available online: https://ec.europa.eu/food/plant/plant_propagation_material/plant_variety_catalogues_databases_en (accessed on 1 March 2022).
- Lewis, M.A.; Russo, E.B.; Smith, K.M. Pharmacological foundations of Cannabis chemovars. Planta Med. 2018, 84, 225–233. [Google Scholar] [CrossRef]
- Mechoulam, R.; Hanuš, L.O.; Pertwee, R.; Howlett, A.C. Early phytocannabinoid chemistry to endocannabinoids and beyond. Nat. Rev. Neurosci. 2014, 15, 757–764. [Google Scholar] [CrossRef]
- Brighenti, V.; Pellati, F.; Steinbach, M.; Maran, D.; Benvenuti, S. Development of a new extraction technique and HPLC method for the analysis of non-psychoactive cannabinoids in fibre-type Cannabis sativa L. (hemp). J. Pharm. Biomed. Anal. 2017, 143, 228–236. [Google Scholar] [CrossRef]
- Radwan, M.M.; Chandra, S.; Gul, S.; ElSohly, M.A. Cannabinoids, phenolics, terpenes and alkaloids of Cannabis. Molecules 2021, 26, 2774. [Google Scholar] [CrossRef]
- Morales, P.; Reggio, P.H.; Jagerovic, N. An overview on medicinal chemistry of synthetic and natural derivatives of cannabidiol. Front. Pharmacol. 2017, 8, 422. [Google Scholar] [CrossRef] [Green Version]
- Aizpurua-Olaizola, O.; Elezgarai, I.; Rico-Barrio, I.; Zarandona, I.; Etxebarria, N.; Usobiaga, A. Targeting the endocannabinoid system: Future therapeutic strategies. Drug Discov. Today 2017, 22, 105–110. [Google Scholar] [CrossRef] [PubMed]
- Aliferis, K.A.; Bernard-Perron, D. Cannabinomics: Application of metabolomics in Cannabis (Cannabis sativa L.) research and development. Front. Plant Sci. 2020, 11, 554. [Google Scholar] [CrossRef] [PubMed]
- Di Marzo, V.; Petrosino, S. Endocannabinoids and the regulation of their levels in health and disease. Curr. Opin. Lipidol. 2007, 18, 129–140. [Google Scholar] [CrossRef] [PubMed]
- Alenabi, A.; Malekinejad, H. Cannabinoids pharmacological effects are beyond the palliative effects: CB2 cannabinoid receptor agonist induced cytotoxicity and apoptosis in human colorectal cancer cells (HT-29). Mol. Cell. Biochem. 2021, 476, 3285–3301. [Google Scholar] [CrossRef] [PubMed]
- Pugazhendhi, A.; Suganthy, N.; Chau, T.P.; Sharma, A.; Unpaprom, Y.; Ramaraj, R.; Karuppusamy, I.; Brindhadevi, K. Cannabinoids as anticancer and neuroprotective drugs: Structural insights and pharmacological interactions—A review. Process Biochem. 2021, 111, 9–31. [Google Scholar] [CrossRef]
- Robaina Cabrera, C.L.; Keir-Rudman, S.; Horniman, N.; Clarkson, N.; Page, C. The anti-inflammatory effects of cannabidiol and cannabigerol alone, and in combination. Pulm. Pharmacol. Ther. 2021, 69, 102047. [Google Scholar] [CrossRef]
- Anil, S.M.; Shalev, N.; Vinayaka, A.C.; Nadarajan, S.; Namdar, D.; Belausov, E.; Shoval, I.; Mani, K.A.; Mechrez, G.; Koltai, H. Cannabis compounds exhibit anti-inflammatory activity in vitro in COVID-19-related inflammation in lung epithelial cells and pro-inflammatory activity in macrophages. Sci. Rep. 2021, 11, 1462. [Google Scholar] [CrossRef]
- Cheng, X.; Lin, J.; Chen, Z.; Mao, Y.; Wu, X.; Xu, C.; Du, J.; Dong, Z.; Yang, H.; Zhou, F.; et al. CB2-mediated attenuation of nucleus pulposus degeneration via the amelioration of inflammation and oxidative stress in vivo and in vitro. Mol. Med. 2021, 27, 92. [Google Scholar] [CrossRef]
- Lataliza, A.A.B.; de Assis, P.M.; da Rocha Laurindo, L.; Gonçalves, E.C.D.; Raposo, N.R.B.; Dutra, R.C. Antidepressant-like effect of rosmarinic acid during LPS-induced neuroinflammatory model: The potential role of cannabinoid receptors/PPAR-γ signaling pathway. Phytother. Res. 2021, 35, 6974–6989. [Google Scholar] [CrossRef]
- Rapaka, D.; Bitra, V.R.; Challa, S.R.; Adiukwu, P.C. Potentiation of microglial endocannabinoid signaling alleviates neuroinflammation in Alzheimer’s disease. Neuropeptides 2021, 90, 102196. [Google Scholar] [CrossRef]
- Calvi, L.; Pavlovic, R.; Panseri, S.; Giupponi, L.; Leoni, V.; Giorgi, A. Quality traits of medical Cannabis sativa L. inflorescences and derived products based on comprehensive mass-spectrometry analytical investigation. In Recent Advances in Cannabinoid Research; Intech Open book Series; IntechOpen: London, UK, 2018. [Google Scholar]
- Pellati, F.; Borgonetti, V.; Brighenti, V.; Biagi, M.; Benvenuti, S.; Corsi, L. Cannabis sativa L. and conpsychoactive Cannabinoids: Their chemistry and role against oxidative stress, inflammation, and cancer. BioMed Res. Int. 2018, 2018, 1691428. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Irakli, M.; Tsaliki, E.; Kalivas, A.; Kleisiaris, F.; Sarrou, E.; Cook, C.M. Effect οf genotype and growing year on the nutritional, phytochemical, and antioxidant properties of industrial hemp (Cannabis sativa L.) seeds. Antioxidants 2019, 8, 491. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Borrelli, F.; Fasolino, I.; Romano, B.; Capasso, R.; Maiello, F.; Coppola, D.; Orlando, P.; Battista, G.; Pagano, E.; Di Marzo, V.; et al. Beneficial effect of the non-psychotropic plant cannabinoid cannabigerol on experimental inflammatory bowel disease. Biochem. Pharmacol. 2013, 85, 1306–1316. [Google Scholar] [CrossRef] [PubMed]
- Guzmán, M. Cannabinoids: Potential anticancer agents. Nat. Rev. Cancer 2003, 3, 745–755. [Google Scholar] [CrossRef] [PubMed]
- Pacifici, R.; Marchei, E.; Salvatore, F.; Guandalini, L.; Busardò, F.P.; Pichini, S. Evaluation of cannabinoids concentration and stability in standardized preparations of cannabis tea and cannabis oil by ultra-high performance liquid chromatography tandem mass spectrometry. Clin. Chem. Lab. Med. 2017, 55, 1555–1563. [Google Scholar] [CrossRef] [PubMed]
- Nelson, K.M.; Bisson, J.; Singh, G.; Graham, J.G.; Chen, S.-N.; Friesen, J.B.; Dahlin, J.L.; Niemitz, M.; Walters, M.A.; Pauli, G.F. The essential medicinal chemistry of cannabidiol (CBD). J. Med. Chem. 2020, 63, 12137–12155. [Google Scholar] [CrossRef]
- Hazekamp, A.; Fischedick, J.T. Cannabis—From cultivar to chemovar. Drug Test Anal. 2012, 4, 660–667. [Google Scholar] [CrossRef]
- Nuutinen, T. Medicinal properties of terpenes found in Cannabis sativa and Humulus lupulus. Eur. J. Med. Chem. 2018, 157, 198–228. [Google Scholar] [CrossRef]
- Pollastro, F.; Minassi, A.; Fresu, G.L. Cannabis phenolics and their bioactivities. Curr. Med. Chem. 2018, 25, 1160–1185. [Google Scholar] [CrossRef]
- Kara, Ş.; Gul, V.; Kiralan, M. Fatty acid composition of hempseed oils from different locatins in Turkey. Span. J. Agric. Res. 2010, 8, 385–390. [Google Scholar]
- Vonapartis, E.; Aubin, M.-P.; Seguin, P.; Mustafa, A.F.; Charron, J.-B. Seed composition of ten industrial hemp cultivars approved for production in Canada. J. Food Compost. Anal. 2015, 39, 8–12. [Google Scholar] [CrossRef]
- Callaway, J.C. Hempseed as a nutritional resource: An overview. Euphytica 2004, 140, 65–72. [Google Scholar] [CrossRef]
- Rehman, M.S.U.; Rashid, N.; Saif, A.; Mahmood, T.; Han, J.-I. Potential of bioenergy production from industrial hemp (Cannabis sativa): Pakistan perspective. Renew. Sustain. Energ. Rev. 2013, 18, 154–164. [Google Scholar] [CrossRef]
- Giupponi, L.; Leoni, V.; Carrer, M.; Ceciliani, G.; Sala, S.; Panseri, S.; Pavlovic, R.; Giorgi, A. Overview on Italian hemp production chain, related productive and commercial activities and legislative framework. Ital. J. Agron. 2020, 15, 194–205. [Google Scholar] [CrossRef]
- Del Gatto, A.; Laureti, D.; Crescentin, P. Un biennio di valutazione di varietà di canapa. Inf. Agrar. 2001, 57, 39–44. [Google Scholar]
- Ferrante, C.; Recinella, L.; Ronci, M.; Menghini, L.; Brunetti, L.; Chiavaroli, A.; Leone, S.; Di Iorio, L.; Carradori, S.; Tirillini, B.; et al. Multiple pharmacognostic characterization on hemp commercial cultivars: Focus on inflorescence water extract activity. Food Chem. Toxicol. 2019, 125, 452–461. [Google Scholar] [CrossRef] [PubMed]
- Zengin, G.; Menghini, L.; Di Sotto, A.; Mancinelli, R.; Sisto, F.; Carradori, S.; Cesa, S.; Fraschetti, C.; Filippi, A.; Angiolella, L.; et al. Chromatographic analyses, in vitro biological activities, and cytotoxicity of Cannabis sativa L. essential oil: A multidisciplinary study. Molecules 2018, 23, 3266. [Google Scholar] [CrossRef] [Green Version]
- Benelli, G.; Pavela, R.; Lupidi, G.; Nabissi, M.; Petrelli, R.; Ngahang Kamte, S.L.; Cappellacci, L.; Fiorini, D.; Sut, S.; Dall’Acqua, S.; et al. The crop-residue of fiber hemp cv. Futura 75: From a waste product to a source of botanical insecticides. Environ. Sci. Pollut. Res. 2018, 25, 10515–10525. [Google Scholar] [CrossRef]
- Orlando, G.; Recinella, L.; Chiavaroli, A.; Brunetti, L.; Leone, S.; Carradori, S.; Di Simone, S.; Ciferri, M.C.; Zengin, G.; Ak, G.; et al. Water extract from inflorescences of industrial hemp Futura 75 variety as a source of anti-inflammatory, anti-proliferative and antimycotic agents: Results from in silico, in vitro and ex vivo studies. Antioxidants 2020, 9, 437. [Google Scholar] [CrossRef]
- Isidore, E.; Karim, H.; Ioannou, I. Extraction of phenolic compounds and terpenes from Cannabis sativa L. by-products: From conventional to intensified processes. Antioxidants 2021, 10, 942. [Google Scholar] [CrossRef]
- De Marino, S.; Festa, C.; Zollo, F.; Iorizzi, M. Novel steroidal components from the underground parts of Ruscus aculeatus L. Molecules 2012, 17, 14002–14014. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Choi, Y.H.; Hazekamp, A.; Peltenburg-Looman, A.M.G.; Frédérich, M.; Erkelens, C.; Lefeber, A.W.M.; Verpoorte, R. NMR assignments of the major cannabinoids and cannabiflavonoids isolated from flowers of Cannabis sativa. Phytochem. Anal. 2004, 15, 345–354. [Google Scholar] [CrossRef] [PubMed]
- Radwan, M.M.; ElSohly, M.A.; Slade, D.; Ahmed, S.A.; Wilson, L.; El-Alfy, A.T.; Khan, I.A.; Ross, S.A. Non-cannabinoid constituents from a high potency Cannabis sativa variety. Phytochemistry 2008, 69, 2627–2633. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shoyama, Y.; Nishioka, I. Cannabis. XIII. Two new spiro-compounds, cannabispirol and acetyl cannabispirol. Chem. Pharm. Bull. 1978, 26, 3641–3646. [Google Scholar] [CrossRef]
- Guo, T.; Liu, Q.; Hou, P.; Li, F.; Guo, S.; Song, W.; Zhang, H.; Liu, X.; Zhang, S.; Zhang, J.; et al. Stilbenoids and cannabinoids from the leaves of Cannabis sativa f. sativa with potential reverse cholesterol transport activity. Food Funct. 2018, 9, 6608–6617. [Google Scholar] [CrossRef] [PubMed]
- Boeren, E.G.; Elsohly, M.A.; Turner, C.E. Cannabiripsol: A novel Cannabis constituent. Experientia 1979, 35, 1278–1279. [Google Scholar] [CrossRef] [PubMed]
- Appendino, G.; Taglialatela-Scafati, O. Cannabinoids: Chemistry and Medicine. In Natural Products: Phytochemistry, Botany and Metabolism of Alkaloids, Phenolics and Terpenes; Ramawat, K.G., Mérillon, J.-M., Eds.; Springer: Berlin/Heidelberg, Germany, 2013; pp. 3415–3435. [Google Scholar]
- Radwan, M.M.; ElSohly, M.A.; El-Alfy, A.T.; Ahmed, S.A.; Slade, D.; Husni, A.S.; Manly, S.P.; Wilson, L.; Seale, S.; Cutler, S.J.; et al. Isolation and pharmacological evaluation of minor cannabinoids from high-potency Cannabis sativa. J. Nat. Prod. 2015, 78, 1271–1276. [Google Scholar] [CrossRef] [Green Version]
- Chianese, G.; Sirignano, C.; Benetti, E.; Marzaroli, V.; Collado, J.A.; de la Vega, L.; Appendino, G.; Munoz, E.; Taglialatela-Scafati, O. A Nrf-2 Stimulatory Hydroxylated Cannabidiol Derivative from Hemp (Cannabis sativa). J. Nat. Prod. 2022, 85, 1089–1097. [Google Scholar] [CrossRef]
- Fiorini, D.; Molle, A.; Nabissi, M.; Santini, G.; Benelli, G.; Maggi, F. Valorizing industrial hemp (Cannabis sativa L.) by-products: Cannabidiol enrichment in the inflorescence essential oil optimizing sample pre-treatment prior to distillation. Ind. Crops Prod. 2019, 128, 581–589. [Google Scholar] [CrossRef]
- Adams, R.P. Identification of Essential Oil Components by Gas Chromatography/Mass Spectrometry; Allured Publishing Corporation: Carol Stream, IL, USA, 2007; Volume 456. [Google Scholar]
- Nagy, D.U.; Cianfaglione, K.; Maggi, F.; Sut, S.; Dall’Acqua, S. Chemical characterization of leaves, male and female flowers from spontaneous cannabis (Cannabis sativa L.) growing in Hungary. Chem. Biodivers. 2019, 16, e1800562. [Google Scholar] [CrossRef]
- Ross, S.A.; ElSohly, M.A.; Sultana, G.N.N.; Mehmedic, Z.; Hossain, C.F.; Chandra, S. Flavonoid glycosides and cannabinoids from the pollen of Cannabis sativa L. Phytochem. Anal. 2005, 16, 45–48. [Google Scholar] [CrossRef] [PubMed]
- Calvi, L.; Pentimalli, D.; Panseri, S.; Giupponi, L.; Gelmini, F.; Beretta, G.; Vitali, D.; Bruno, M.; Zilio, E.; Pavlovic, R.; et al. Comprehensive quality evaluation of medical Cannabis sativa L. inflorescence and macerated oils based on HS-SPME coupled to GC–MS and LC-HRMS (q-exactive orbitrap®) approach. J. Pharm. Biomed. Anal. 2018, 150, 208–219. [Google Scholar] [CrossRef] [PubMed]
- Smeriglio, A.; Trombetta, D.; Cornara, L.; Valussi, M.; De Feo, V.; Caputo, L. Characterization and phytotoxicity assessment of essential oils from plant byproducts. Molecules 2019, 24, 2941. [Google Scholar] [CrossRef] [Green Version]
- Bahi, A.; Al Mansouri, S.; Al Memari, E.; Al Ameri, M.; Nurulain, S.M.; Ojha, S. β-Caryophyllene, a CB2 receptor agonist produces multiple behavioral changes relevant to anxiety and depression in mice. Physiol. Behav. 2014, 135, 119–124. [Google Scholar] [CrossRef] [PubMed]
- Chini, M.G.; Lauro, G.; Bifulco, G. Addressing the target identification and accelerating the repositioning of anti-inflammatory/anti-cancer organic compounds by computational approaches. Eur. J. Org. Chem. 2021, 2021, 2966–2981. [Google Scholar] [CrossRef]
- Lauro, G.; Masullo, M.; Piacente, S.; Riccio, R.; Bifulco, G. Inverse Virtual Screening allows the discovery of the biological activity of natural compounds. Biorg. Med. Chem. 2012, 20, 3596–3602. [Google Scholar] [CrossRef]
- Lauro, G.; Romano, A.; Riccio, R.; Bifulco, G. Inverse Virtual Screening of antitumor targets: Pilot study on a small database of natural bioactive compounds. J. Nat. Prod. 2011, 74, 1401–1407. [Google Scholar] [CrossRef] [PubMed]
- De Vita, S.; Lauro, G.; Ruggiero, D.; Terracciano, S.; Riccio, R.; Bifulco, G. Protein preparation automatic protocol for high-throughput inverse virtual screening: Accelerating the target identification by computational methods. J. Chem. Inf. Model. 2019, 59, 4678–4690. [Google Scholar] [CrossRef]
- Dietrich, J.D.; Longenecker, K.L.; Wilson, N.S.; Goess, C.; Panchal, S.C.; Swann, S.L.; Petros, A.M.; Hobson, A.D.; Ihle, D.; Song, D.; et al. Development of orally efficacious allosteric inhibitors of TNFα via fragment-based drug design. J. Med. Chem. 2021, 64, 417–429. [Google Scholar] [CrossRef]
- Lightwood, D.J.; Munro, R.J.; Porter, J.; McMillan, D.; Carrington, B.; Turner, A.; Scott-Tucker, A.; Hickford, E.S.; Schmidt, A.; Fox, D.; et al. A conformation-selective monoclonal antibody against a small molecule-stabilised signalling-deficient form of TNF. Nat. Commun. 2021, 12, 583. [Google Scholar] [CrossRef]
- Hillisch, A.; Gericke, K.M.; Allerheiligen, S.; Roehrig, S.; Schaefer, M.; Tersteegen, A.; Schulz, S.; Lienau, P.; Gnoth, M.; Puetter, V.; et al. Design, synthesis, and pharmacological characterization of a neutral, non-prodrug thrombin inhibitor with good oral pharmacokinetics. J. Med. Chem. 2020, 63, 12574–12594. [Google Scholar] [CrossRef] [PubMed]
- Carter, W.J.; Myles, T.; Gibbs, C.S.; Leung, L.L.; Huntington, J.A. Crystal structure of anticoagulant thrombin variant E217K provides insights into thrombin allostery. J. Biol. Chem. 2004, 279, 26387–26394. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Waku, T.; Shiraki, T.; Oyama, T.; Maebara, K.; Nakamori, R.; Morikawa, K. The nuclear receptor PPARγ individually responds to serotonin- and fatty acid-metabolites. EMBO J. 2010, 29, 3395–3407. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oberfield, J.L.; Collins, J.L.; Holmes, C.P.; Goreham, D.M.; Cooper, J.P.; Cobb, J.E.; Lenhard, J.M.; Hull-Ryde, E.A.; Mohr, C.P.; Blanchard, S.G.; et al. A peroxisome proliferator-activated receptor γ ligand inhibits adipocyte differentiation. Proc. Natl. Acad. Sci. USA 1999, 96, 6102–6106. [Google Scholar] [CrossRef] [Green Version]
- Chu, W.-M. Tumor necrosis factor. Cancer Lett. 2013, 328, 222–225. [Google Scholar] [CrossRef] [Green Version]
- Akash, M.S.H.; Rehman, K.; Liaqat, A. Tumor Necrosis Factor-Alpha: Role in development of insulin resistance and pathogenesis of type 2 diabetes mellitus. J. Cell. Biochem. 2018, 119, 105–110. [Google Scholar] [CrossRef]
- Kalliolias, G.D.; Ivashkiv, L.B. TNF biology, pathogenic mechanisms and emerging therapeutic strategies. Nat. Rev. Rheumatol. 2016, 12, 49–62. [Google Scholar] [CrossRef]
- Dostert, C.; Grusdat, M.; Letellier, E.; Brenner, D. The TNF family of ligands and receptors: Communication modules in the immune system and beyond. Physiol. Rev. 2018, 99, 115–160. [Google Scholar] [CrossRef]
- Jinesh, S. Pharmaceutical aspects of anti-inflammatory TNF-blocking drugs. Inflammopharmacology 2015, 23, 71–77. [Google Scholar] [CrossRef]
- Lis, K.; Kuzawińska, O.; Bałkowiec-Iskra, E. Tumor necrosis factor inhibitors—State of knowledge. Arch. Med. Sci. 2014, 10, 1175–1185. [Google Scholar] [CrossRef]
- Dömling, A.; Li, X. TNF-α: The shape of small molecules to come? Drug Discov. Today 2022, 27, 3–7. [Google Scholar] [CrossRef] [PubMed]
- Ma, H.; Xu, F.; Liu, C.; Seeram, N.P. A Network pharmacology approach to identify potential molecular targets for cannabidiol’s anti-inflammatory activity. Cannabis Cannabinoid Res. 2020, 6, 288–299. [Google Scholar] [CrossRef] [PubMed]
- Fischer-Stenger, K.; Dove Pettit, D.A.; Cabral, G.A. Delta 9-tetrahydrocannabinol inhibition of tumor necrosis factor-alpha: Suppression of post-translational events. J. Pharmacol. Exp. Ther. 1993, 267, 1558. [Google Scholar] [PubMed]
- Suryavanshi, S.V.; Kovalchuk, I.; Kovalchuk, O. Cannabinoids as key regulators of inflammasome signaling: A current perspective. Front. Immunol. 2021, 11, 613613. [Google Scholar] [CrossRef]
- Abd El-Karim, S.S.; Mohamed, H.S.; Abdelhameed, M.F.; El-Galil, E.; Amr, A.; Almehizia, A.A.; Nossier, E.S. Design, synthesis and molecular docking of new pyrazole-thiazolidinones as potent anti-inflammatory and analgesic agents with TNF-α inhibitory activity. Bioorg. Chem. 2021, 111, 104827. [Google Scholar] [CrossRef]
- Ali, Y.; Alam, M.S.; Hamid, H.; Husain, A.; Shafi, S.; Dhulap, A.; Hussain, F.; Bano, S.; Kharbanda, C.; Nazreen, S.; et al. Design and synthesis of butenolide-based novel benzyl pyrrolones: Their TNF-α based molecular docking with in vivo and in vitro anti-inflammatory activity. Chem. Biol. Drug Des. 2015, 86, 619–625. [Google Scholar] [CrossRef]
- O’Connell, J.; Porter, J.; Kroeplien, B.; Norman, T.; Rapecki, S.; Davis, R.; McMillan, D.; Arakaki, T.; Burgin, A.; Fox, D., III; et al. Small molecules that inhibit TNF signalling by stabilising an asymmetric form of the trimer. Nat. Commun. 2019, 10, 5795. [Google Scholar] [CrossRef]
- Dahlbäck, B. Blood coagulation. Lancet 2000, 355, 1627–1632. [Google Scholar] [CrossRef]
- O’Brien, P.J.; Mureebe, L. Direct thrombin inhibitors. J. Cardiovasc. Pharmacol. Ther. 2011, 17, 5–11. [Google Scholar] [CrossRef]
- Shi, C.-C.; Chen, T.-R.; Zhang, Q.-H.; Wei, L.-H.; Huang, C.; Zhu, Y.-D.; Liu, H.-B.; Bai, Y.-K.; Wang, F.-J.; Guo, W.-Z.; et al. Inhibition of human thrombin by the constituents of licorice: Inhibition kinetics and mechanistic insights through in vitro and in silico studies. RSC Adv. 2020, 10, 3626–3635. [Google Scholar] [CrossRef]
- Li, Q.-Q.; Yang, F.-Q.; Wang, Y.-Z.; Wu, Z.-Y.; Xia, Z.-N.; Chen, H. Evaluation of thrombin inhibitory activity of catechins by online capillary electrophoresis-based immobilized enzyme microreactor and molecular docking. Talanta 2018, 185, 16–22. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.-Q.; Yang, Y.-X.; Qv, J.-W.; Hu, G.; Hu, Y.-J.; Xia, Z.-N.; Yang, F.-Q. Investigation of interactions between thrombin and ten phenolic compounds by affinity capillary electrophoresis and molecular docking. J. Anal. Methods Chem. 2018, 2018, 4707609. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, X.; Yang, Z.; Su, F.; Li, J.; Boadi, E.O.; Chang, Y.-X.; Wang, H. Study on structure activity relationship of natural flavonoids against thrombin by molecular docking virtual screening combined with activity evaluation in vitro. Molecules 2020, 25, 422. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Troisi, R.; Balasco, N.; Autiero, I.; Vitagliano, L.; Sica, F. Exosite binding in thrombin: A global structural/dynamic overview of complexes with aptamers and other ligands. Int. J. Mol. Sci. 2021, 22, 10803. [Google Scholar] [CrossRef] [PubMed]
- Jouanjus, E.; Raymond, V.; Lapeyre-Mestre, M.; Wolff, V. What is the Current Knowledge about the Cardiovascular Risk for Users of Cannabis-Based Products? A Systematic Review. Curr. Atheroscler. Rep. 2017, 19, 26. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Wang, J.; Luo, S.; Zhan, Y.; Lu, Q. The roles of PPARγ and its agonists in autoimmune diseases: A comprehensive review. J. Autoimmun. 2020, 113, 102510. [Google Scholar] [CrossRef]
- Mirza, A.Z.; Althagafi, I.I.; Shamshad, H. Role of PPAR receptor in different diseases and their ligands: Physiological importance and clinical implications. Eur. J. Med. Chem. 2019, 166, 502–513. [Google Scholar] [CrossRef]
- Mazumder, M.; Ponnan, P.; Das, U.; Gourinath, S.; Khan, H.A.; Yang, J.; Sakharkar, M.K. Investigations on binding pattern of kinase inhibitors with PPARγ: Molecular docking, molecular dynamic simulations, and free energy calculation studies. PPAR Res. 2017, 2017, 6397836. [Google Scholar] [CrossRef] [Green Version]
- Jian, Y.; He, Y.; Yang, J.; Han, W.; Zhai, X.; Zhao, Y.; Li, Y. Molecular modeling study for the design of novel Peroxisome Proliferator-Activated Receptor Gamma agonists using 3D-QSAR and molecular docking. Int. J. Mol. Sci. 2018, 19, 630. [Google Scholar] [CrossRef] [Green Version]
- Sheu, S.-H.; Kaya, T.; Waxman, D.J.; Vajda, S. Exploring the binding site structure of the PPARγ ligand-binding domain by computational solvent mapping. Biochemistry 2005, 44, 1193–1209. [Google Scholar] [CrossRef]
- Devchand, P.R.; Liu, T.; Altman, R.B.; FitzGerald, G.A.; Schadt, E.E. The pioglitazone trek via human PPAR gamma: From discovery to a medicine at the FDA and beyond. Front. Pharmacol. 2018, 9, 1093. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ashraf, S.A.; Elkhalifa, A.E.O.; Mehmood, K.; Adnan, M.; Khan, M.A.; Eltoum, N.E.; Krishnan, A.; Baig, M.S. Multi-targeted molecular docking, pharmacokinetics, and drug-likeness evaluation of okra-derived ligand abscisic acid targeting signaling proteins involved in the development of diabetes. Molecules 2021, 26, 5957. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, K.; Tamura, T.; Henmi, K.; Kuboyama, T.; Yanagisawa, A.; Matsubara, M.; Takahashi, Y.; Suzuki, M.; Saito, J.-I.; Ueno, K.; et al. Development of dihydrodibenzooxepine Peroxisome Proliferator-Activated Receptor (PPAR) Gamma ligands of a novel binding mode as anticancer agents: Effective mimicry of chiral structures by olefinic E/Z-isomers. J. Med. Chem. 2018, 61, 10067–10083. [Google Scholar] [CrossRef] [PubMed]
- Chandra, V.; Huang, P.; Hamuro, Y.; Raghuram, S.; Wang, Y.; Burris, T.P.; Rastinejad, F. Structure of the intact PPAR-γ–RXR-α nuclear receptor complex on DNA. Nature 2008, 456, 350–356. [Google Scholar] [CrossRef]
- Nichols, J.M.; Kaplan, B.L.F. Immune responses regulated by cannabidiol. Cannabis Cannabinoid Res. 2019, 5, 12–31. [Google Scholar] [CrossRef] [Green Version]
- García-Martín, A.; Garrido-Rodríguez, M.; Navarrete, C.; Caprioglio, D.; Palomares, B.; DeMesa, J.; Rollland, A.; Appendino, G.; Muñoz, E. Cannabinoid derivatives acting as dual PPARγ/CB2 agonists as therapeutic agents for systemic sclerosis. Biochem. Pharmacol. 2019, 163, 321–334. [Google Scholar] [CrossRef]
- O’Sullivan, S.E. An update on PPAR activation by cannabinoids. Br. J. Pharmacol. 2016, 173, 1899–1910. [Google Scholar] [CrossRef] [Green Version]
- Dimmito, M.P.; Stefanucci, A.; Della Valle, A.; Scioli, G.; Cichelli, A.; Mollica, A. An overview on plants cannabinoids endorsed with cardiovascular effects. Biomed. Pharmacother. 2021, 142, 111963. [Google Scholar] [CrossRef]
- Iannotti, F.A.; De Maio, F.; Panza, E.; Appendino, G.; Taglialatela-Scafati, O.; De Petrocellis, L.; Amodeo, P.; Vitale, R.M. Identification and characterization of cannabimovone, a cannabinoid from Cannabis sativa, as a novel PPARγ agonist via a combined computational and functional study. Molecules 2020, 25, 1119. [Google Scholar] [CrossRef] [Green Version]
- Council of Europe; European Pharmacopoeia Commission; European Directorate for the Quality of Medicines. European Pharmacopoeia; EDQM: Strasbourg, France, 2004; p. 217. [Google Scholar]
- McLafferty, F.W. Wiley Registry of Mass Spectral Data, with NIST98 Spectra, 7th ed.; John Wiley & Sons: New York, NY, USA, 2000. [Google Scholar]
- Grob, R.L.; Kaiser, M.A. Qualitative and Quantitative Analysis by Gas Chromatography. In Modern Practice of Gas Chromatography; John Wiley & Sons: New York, NY, USA, 2004. [Google Scholar]
- Schrödinger Release 2020: SiteMap, 2020; Schrödinger, LLC: New York, NY, USA, 2020.
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef] [Green Version]
| Position | δHa | δCa |
|---|---|---|
| 1 | 3.15 t (J = 11.0 Hz) | 48.2 |
| 2 | 4.73 br d (J = 11.0 Hz) | 73.9 |
| 3 | - | 153.1 |
| 4 | 2.20 m 2.45 m | 35.4 |
| 5 | 1.43 dd (J = 13.0, 3.6 Hz) 1.70 m | 34.8 |
| 6 | 3.35 ddd (J = 11.0, 11.2, 3.5 Hz) | 48.2 |
| 7 | 4.76 s 5.00 s | 104.7 |
| 8 | - | 149.4 |
| 9 | 4.37 s 4.59 br s | 110.6 |
| 10 | 1.56 s | 19.1 |
| 1′ | - | 113.0 |
| 2′ | - | 157.9 |
| 3′ | 6.11 s | 107.8 |
| 4′ | - | 142.9 |
| 5′ | 6.09 s | 108.7 |
| 6′ | - | 157.9 |
| 1″ | 2.38 t (J = 7.7) | 36.6 |
| 2″ | 1.55 m | 32.0 |
| 3″ | 1.31 m | 23.6 |
| 4″ | 1.32 m | 32.7 |
| 5″ | 0.90 t (J = 6.8 Hz) | 14.4 |
| No. | Compound | Exp. RI | Ref. RI | Area% ± DS | Abb. |
|---|---|---|---|---|---|
| 1 | α-Pinene | 929 | 939 | 0.56 ± 0.03 | BM |
| 2 | Camphene | 944 | 954 | 0.02 ± 0.00 | BM |
| 3 | Sabinene | 972 | 975 | 0.17 ± 0.02 | BM |
| 4 | Myrcene | 991 | 990 | 0.11 ± 0.01 | AM |
| 5 | α-Phellandrene | 1000 | 1002 | 0.01 ± 0.00 | MM |
| 6 | δ-3-Carene | 1006 | 1011 | 0.01 ± 0.00 | BM |
| 7 | α-Terpinene | 1015 | 1017 | 0.02 ± 0.01 | MM |
| 8 | p-Cymene | 1023 | 1024 | 0.05 ± 0.00 | MM |
| 9 | Limonene | 1027 | 1029 | 0.14 ± 0.05 | MM |
| 10 | 1,8 Cineole | 1031 | 1031 | 0.09 ± 0.01 | BMO |
| 11 | (Z)-β-Ocimene | 1041 | 1037 | 0.01 ± 0.00 | AM |
| 12 | (E)-β-Ocimene | 1052 | 1050 | 0.02 ± 0.00 | AM |
| 13 | γ-Terpinene | 1060 | 1059 | 0.02 ± 0.00 | MM |
| 14 | Terpinolene | 1087 | 1088 | 0.02 ± 0.01 | MM |
| 15 | p-Cymenene | 1089 | 1091 | 0.02 ± 0.00 | MM |
| 16 | Linalool | 1101 | 1096 | 0.10 ± 0.01 | AM |
| 17 | Nonanal | 1106 | 1100 | 0.02 ± 0.00 | OT |
| 18 | endo-Fenchol | 1112 | 1116 | 0.02 ± 0.00 | BMO |
| 19 | trans-Pinene-hydrate | 1121 | 1122 | 0.01 ± 0.00 | BMO |
| 20 | trans-Pinocarveol | 1138 | 1139 | 0.03 ± 0.00 | BMO |
| 21 | Camphor | 1144 | 1146 | 0.01 ± 0.00 | BMO |
| 22 | Borneol | 1166 | 1169 | 0.06 ± 0.00 | BMO |
| 23 | Terpinen-4-ol | 1177 | 1177 | 0.08 ± 0.00 | BMO |
| 24 | p-Cymen-8-ol | 1187 | 1182 | 0.01 ± 0.01 | MMO |
| 25 | α-Terpineol | 1190 | 1188 | 0.03 ± 0.00 | MMO |
| 26 | Myrtenol | 1195 | 1195 | 0.02 ± 0.00 | BMO |
| 27 | Estragole (Methyl chavicol) | 1198 | 1195 | 0.01 ± 0.01 | OT |
| 28 | trans-Pulegol | 1218 | 1214 | 0.06 ± 0.01 | MMO |
| 29 | trans-Carveol | 1220 | 1216 | 0.01 ± 0.00 | MMO |
| 30 | Linalool acetate | 1260 | 1257 | 0.04 ± 0.00 | AMO |
| 31 | Eugenol | 1359 | 1359 | 0.04 ± 0.00 | MMO |
| 32 | α-Ylangene | 1369 | 1375 | 0.12 ± 0.01 | BS |
| 33 | α-Copaene | 1373 | 1376 | 0.09 ± 0.02 | BS |
| 34 | β-Elemene | 1388 | 1390 | 0.11 ± 0.01 | MS |
| 35 | β-Longipinene | 1402 | 1400 | 0.55 ± 0.04 | BS |
| 36 | Z-Caryophyllene | 1407 | 1408 | 0.13 ± 0.03 | BS |
| 37 | β-Caryophyllene | 1419 | 1419 | 13.82 ± 0.47 | BS |
| 38 | α-trans-Bergamotene | 1436 | 1434 | 1.58 ± 0.04 | MS |
| 39 | α-Humulene | 1453 | 1454 | 5.33 ± 0.17 | MS |
| 40 | allo-Aromadendrene | 1459 | 1460 | 1.46 ± 0.10 | BS |
| 41 | dehydro-Aromadendrene | 1460 | 1462 | 0.45 ± 0.02 | BS |
| 42 | γ-Himachalene | 1483 | 1482 | 3.5 ± 0.14 | BS |
| 43 | α-Selinene | 1492 | 1498 | 1.97 ± 0.03 | BS |
| 44 | β-Himacalene | 1508 | 1500 | 0.22 ± 0.03 | BS |
| 45 | δ-Amorphene | 1512 | 1512 | 0.73 ± 0.06 | BS |
| 46 | γ-Cadinene | 1516 | 1513 | 0.32 ± 0.06 | BS |
| 47 | δ-Cadinene | 1523 | 1523 | 0.32 ± 0.03 | BS |
| 48 | trans-Cadina-1,4-diene | 1532 | 1534 | 0.92 ± 0.03 | BS |
| 49 | α-Cadinene | 1539 | 1538 | 1.22 ± 0.08 | BS |
| 50 | α-Calacorene | 1541 | 1545 | 0.1 ± 0.01 | BS |
| 51 | Selina-3,7(11)-diene | 1543 | 1546 | 0.04 ± 0.00 | BS |
| 52 | Italicene epoxide | 1549 | 1548 | 1.0 ± 0.01 | BSO |
| 53 | (E)-Nerolidol | 1567 | 1563 | 1.01 ± 0.04 | ASO |
| 54 | Caryophyllene oxide | 1581 | 1583 | 5.7 ± 0.39 | BSO |
| 55 | Spathulenol | 1583 | 1578 | 0.09 ± 0.01 | BSO |
| 56 | Viridiflorol | 1595 | 1592 | 0.55 ± 0.06 | BSO |
| 57 | Ledol | 1599 | 1602 | 0.45 ± 0.09 | BSO |
| 58 | Humulene epoxide II | 1606 | 1608 | 1.7 ± 0.16 | MSO |
| 59 | Isolongifolan-7-α-ol | 1616 | 1619 | 0.83 ± 0.02 | BSO |
| 60 | allo-Aromadendrene-epoxide | 1634 | 1641 | 4.41 ± 0.37 | BSO |
| 61 | Caryophylla-4(12),8(13)-dien-5α-ol | 1638 | 1640 | 2.43 ± 0.18 | BSO |
| 62 | Selina-3,11-dien-6-α-ol | 1643 | 1644 | 0.63 ± 0.07 | BSO |
| 63 | Desmethoxy encecalin | 1651 | 1647 | 0.94 ± 0.11 | OT |
| 64 | α-Bisabolol oxide B | 1661 | 1658 | 5.12 ± 0.33 | BSO |
| 65 | (Z)-Caryophyllene-14-hydroxy | 1674 | 1667 | 3.44 ± 0.10 | BSO |
| 66 | epi-α-Bisabolol- | 1684 | 1684 | 0.3 ± 0.04 | MSO |
| 67 | Eudesm-7(11)-en-4-ol | 1693 | 1700 | 0.23 ± 0.02 | BSO |
| 68 | (2E,6E)-Farnesyl acetate | 1847 | 1846 | 0.12 ± 0.03 | OT |
| 69 | (5E,9E)-Farnesyl acetone | 1919 | 1913 | 0.15 ± 0.02 | OT |
| 70 | trans-Phytol | 2113 | 2104 | 0.22 ± 0.05 | OT |
| 71 | Linoleic acid | 2137 | 2133 | 0.12 ± 0.01 | OT |
| 72 | Cannabidivarin | 2217 | 0.72 ± 0.08 | CB | |
| 73 | Cannabicitran | 2271 | 1.56 ± 0.1 | CB | |
| 74 | U (314, 299, 271, 258, 243, 231, 174) | 2332 | 0.2 ± 0.01 | U | |
| 75 | Cannabiclyclol | 2367 | 0.26 ± 0.02 | CB | |
| 76 | Cannabidiol (CBD) | 2432 | 2430 | 28.48 ± 3.02 | CB |
| 77 | Cannabichromene | 2437 | 0.61 ± 0.12 | CB | |
| 78 | Dronabinol (Δ8-THC) | 2484 | 0.11 ± 0.01 | CB | |
| 79 | U (330, 312, 247, 205, 148, 135) | 2488 | 0.13 ± 0.01 | U | |
| 80 | Δ-THC | 2521 | 0.15 ± 0.03 | CB | |
| 81 | Cannabigerol | 2587 | 0.08 ± 0.01 | CB | |
| 82 | Cannabinol | 2587 | 0.14 ± 0.03 | CB | |
| 83 | Heptacosane | 2699 | 2700 | 0.2 ± 0.00 | OT |
| 84 | Nonacosane | 2899 | 2900 | 1.04 ± 0.03 | OT |
| Total identified (%) | 97.45 | ||||
| Oil yield (%) | 0.1 | ||||
| Monoterpene Hydrocarbons | 1.28 | ||||
| Oxigenate monoterpenes | 0.52 | ||||
| Sesquiterpene Hydrocarbons | 32.71 | ||||
| Oxigenate sesquiterpenes | 28.04 | ||||
| Cannabinoids | 32.11 | ||||
| Others | 2.79 |
| Compound | Top-Scored Target | PDB | Binding Affinity |
|---|---|---|---|
| 1 | Tumor necrosis factor | 6X83 [67] | −9.2 |
| 2 | Tumor necrosis factor | 7KPA [68] | −9.7 |
| 3 | Tumor necrosis factor | 7KPA [68] | −10.1 |
| UniProt ID | Molecule Name | Binding Affinity (kcal/mol) (PDB ID) | ||
|---|---|---|---|---|
| 1 | 2 | 3 | ||
| P00734 | Prothrombin/Thrombin | −8.4 (6ZUX [69]) | −8.6 (6ZV8 [69]) | −8.5 (1RD3 [70]) |
| P37231 | Peroxisome proliferator-activated receptor gamma | −8.6 (2ZK6 [71]) | −9.2 (4PRG [72]) | −9.4 (4PRG [72]) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
De Vita, S.; Finamore, C.; Chini, M.G.; Saviano, G.; De Felice, V.; De Marino, S.; Lauro, G.; Casapullo, A.; Fantasma, F.; Trombetta, F.; et al. Phytochemical Analysis of the Methanolic Extract and Essential Oil from Leaves of Industrial Hemp Futura 75 Cultivar: Isolation of a New Cannabinoid Derivative and Biological Profile Using Computational Approaches. Plants 2022, 11, 1671. https://doi.org/10.3390/plants11131671
De Vita S, Finamore C, Chini MG, Saviano G, De Felice V, De Marino S, Lauro G, Casapullo A, Fantasma F, Trombetta F, et al. Phytochemical Analysis of the Methanolic Extract and Essential Oil from Leaves of Industrial Hemp Futura 75 Cultivar: Isolation of a New Cannabinoid Derivative and Biological Profile Using Computational Approaches. Plants. 2022; 11(13):1671. https://doi.org/10.3390/plants11131671
Chicago/Turabian StyleDe Vita, Simona, Claudia Finamore, Maria Giovanna Chini, Gabriella Saviano, Vincenzo De Felice, Simona De Marino, Gianluigi Lauro, Agostino Casapullo, Francesca Fantasma, Federico Trombetta, and et al. 2022. "Phytochemical Analysis of the Methanolic Extract and Essential Oil from Leaves of Industrial Hemp Futura 75 Cultivar: Isolation of a New Cannabinoid Derivative and Biological Profile Using Computational Approaches" Plants 11, no. 13: 1671. https://doi.org/10.3390/plants11131671
APA StyleDe Vita, S., Finamore, C., Chini, M. G., Saviano, G., De Felice, V., De Marino, S., Lauro, G., Casapullo, A., Fantasma, F., Trombetta, F., Bifulco, G., & Iorizzi, M. (2022). Phytochemical Analysis of the Methanolic Extract and Essential Oil from Leaves of Industrial Hemp Futura 75 Cultivar: Isolation of a New Cannabinoid Derivative and Biological Profile Using Computational Approaches. Plants, 11(13), 1671. https://doi.org/10.3390/plants11131671











