CottonGen: The Community Database for Cotton Genomics, Genetics, and Breeding Research
Abstract
1. Introduction
2. Contents and Functions
2.1. Data and Web Interface
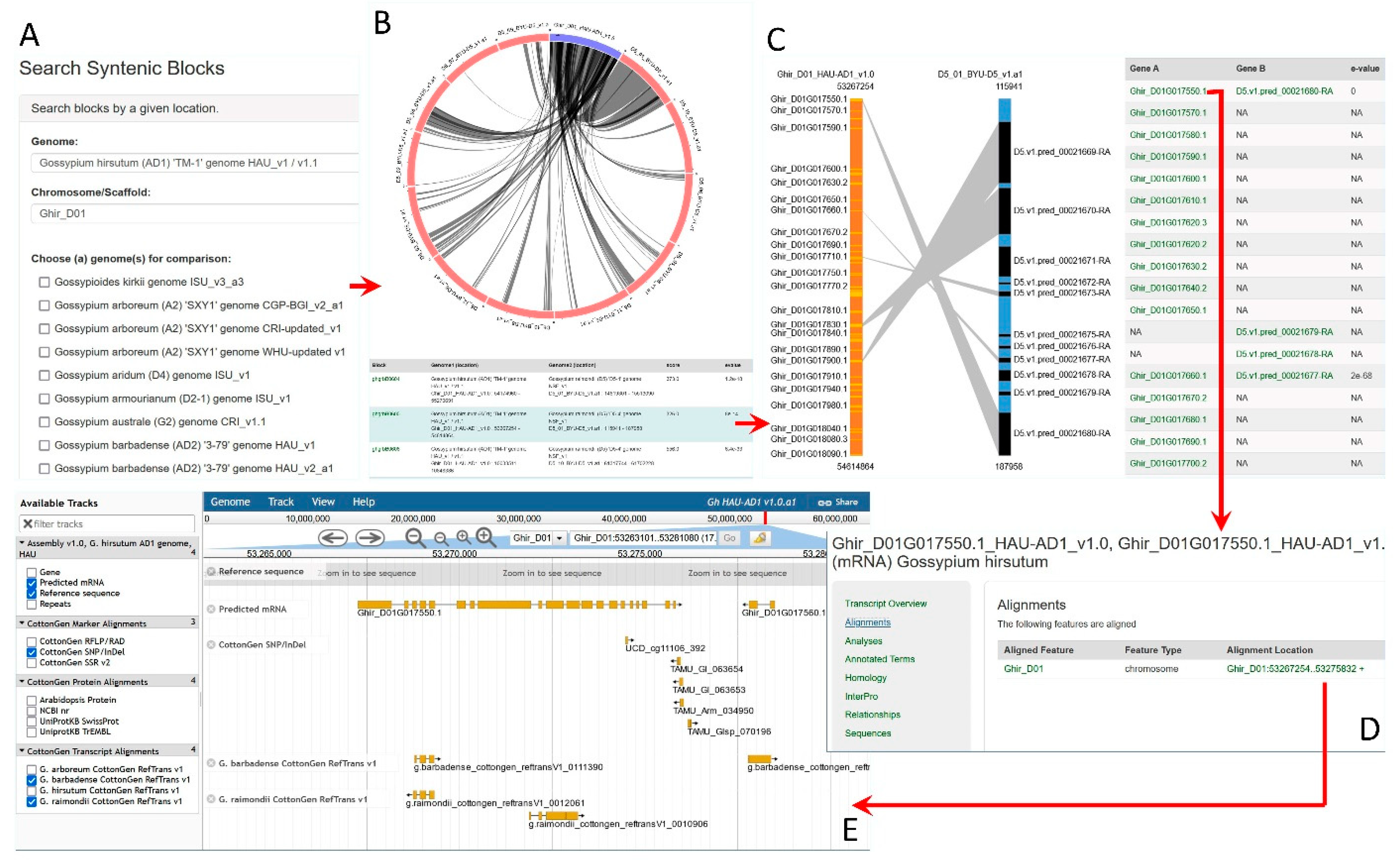
2.2. Genomics Data
2.2.1. Whole Genome Sequence Data
2.2.2. Transcriptome Data
2.2.3. NCBI Genes
2.3. Genetics Data
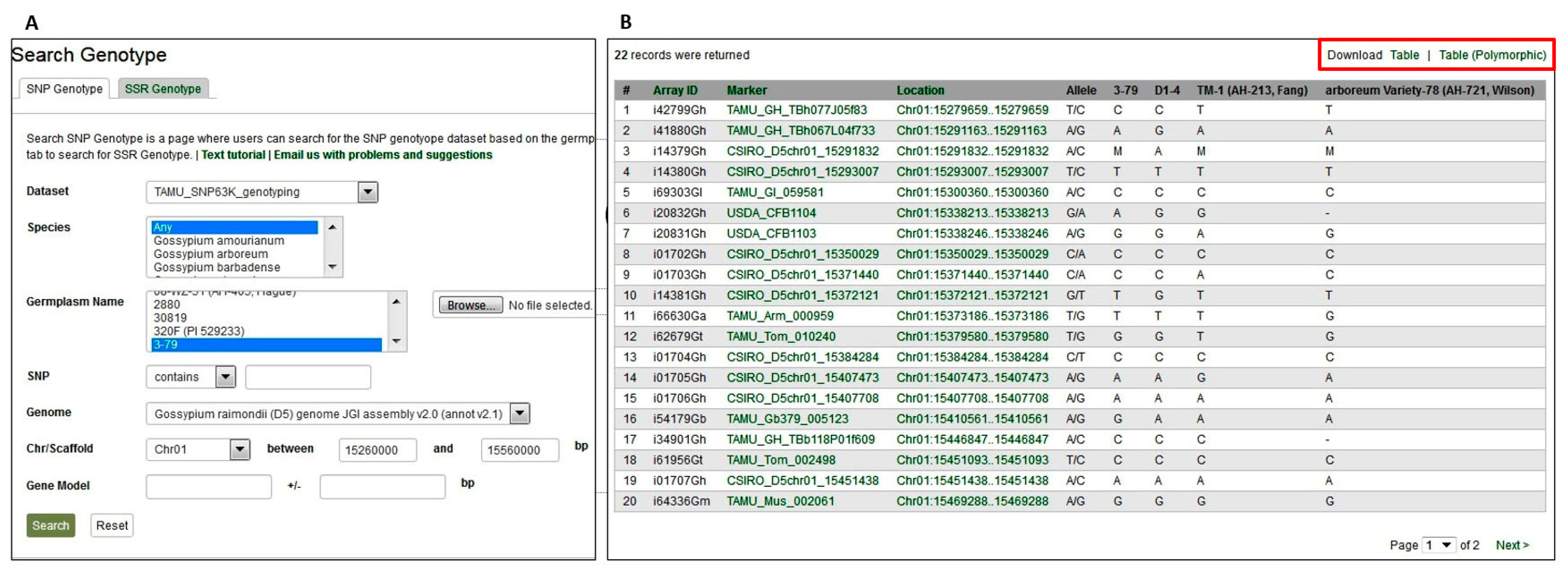
2.3.1. Genetic Marker and SNP Array Data
2.3.2. Genetic Maps and QTLs
2.3.3. Genotypic and Phenotypic Diversity Data
2.3.4. Cotton Trait Ontology
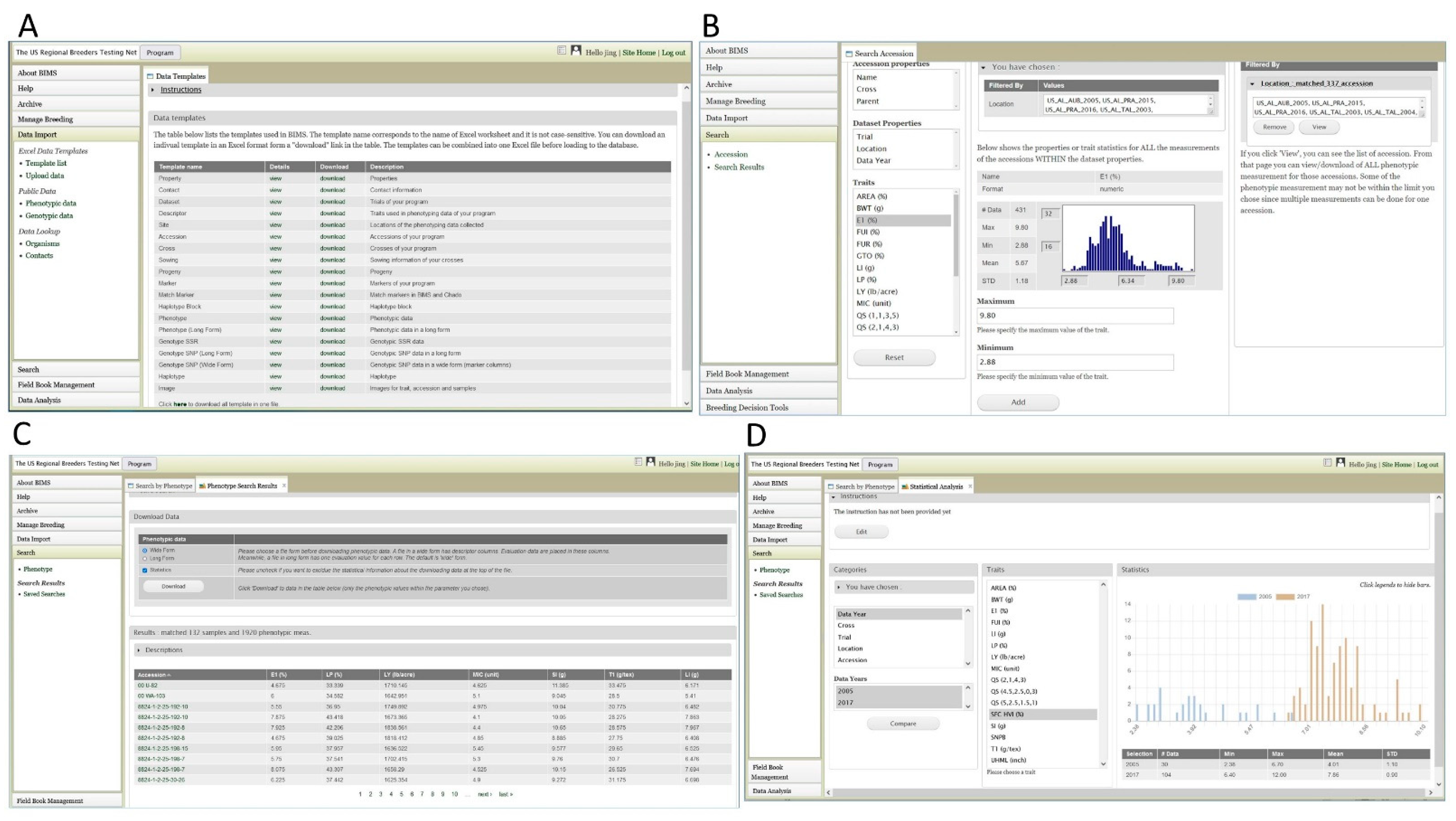
2.4. Breeding Data and Breeding Information Management System
2.5. Community Resources
3. Data Collection, Submission and Download
4. Concluding Remarks and Future Direction
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Yu, J.; Main, D. Role of Bioinformatics Tools and Databases in Cotton Research. In Cotton, 2nd ed.; Agronomy Monograph 57; Fang, D.D., Percy, R.G., Eds.; John Wiley & Sons: New York, NY, USA, 2015; pp. 303–338. [Google Scholar]
- Yu, J.; Jung, S.; Cheng, C.-H.; Ficklin, S.P.; Lee, T.; Zheng, P.; Jones, D.; Percy, R.G.; Main, D. CottonGen: A genomics, genetics and breeding database for cotton research. Nucleic Acids Res. 2014, 42, D1229–D1236. [Google Scholar] [CrossRef]
- Yu, J.; Hinze, L.L.; Yu, J.Z.; Kohel, R.J. CottonDB.org—New website for cotton genome database. In Proceedings of the International Cotton Genome Initiative Research Conference, Brasilia, Brazil, 18–20 September 2006; Available online: https://www.ars.usda.gov/research/publications/publication/?seqNo115=197886 (accessed on 17 November 2021).
- Yu, J.; Kohel, R.; Hinze, L.; Yu, J.Z.; Frelichowski, J.; Ficklin, S.; Main, D.; Percy, R.G. CottonDB. In Proceedings of the International Plant and Animal Genome Conference XX, San Diego, CA, USA, 14–18 January 2012; Available online: https://pag.confex.com/pag/xx/webprogram/Paper1715.html (accessed on 17 November 2021).
- Blenda, A.; Scheffler, J.; Scheffler, B.; Palmer, M.; Lacape, J.-M.; Yu, J.Z.; Jesudurai, C.; Jung, S.; Muthukumar, S.; Yellambalase, P.; et al. CMD: A Cotton Microsatellite Database resource for Gossypium genomics. BMC Genom. 2006, 7, 132. [Google Scholar] [CrossRef]
- Ficklin, S.; Sanderson, L.-A.; Cheng, C.-H.; Staton, M.E.; Lee, T.; Cho, I.-H.; Jung, S.; Bett, K.E.; Main, D. Tripal: A construction toolkit for online genome databases. Database 2011, 2011, bar044. [Google Scholar] [CrossRef]
- Sanderson, L.A.; Ficklin, S.P.; Cheng, C.H.; Jung, S.; Feltus, F.A.; Bett, K.E.; Main, D. Tripal v1.1: A standards-based toolkit for construction of online genetic and genomic databases. Database 2013, 2013, bat075. [Google Scholar] [CrossRef] [PubMed]
- Staton, M.; Cannon, E.; Sanderson, L.A.; Wegrzyn, J.; Anderson, T.; Buehler, S.; Cobo-Simón, I.; Faaberg, K.; Grau, E.; Guignon, V.; et al. Tripal, a community update after 10 years of supporting open source, standards-based genetic, genomic and breeding databases. Briefings Bioinf. 2021, 22, bbab238. [Google Scholar] [CrossRef]
- Mungall, C.J.; Emmert, D.B. The FlyBase Consortium a Chado case study: An ontology-based modular schema for representing genome-associated biological information. Bioinformatics 2007, 23, i337–i346. [Google Scholar] [CrossRef]
- Mungall, C.J.; Batchelor, C.; Eilbeck, K. Evolution of the Sequence Ontology terms and relationships. J. Biomed. Inform. 2011, 44, 87–93. [Google Scholar] [CrossRef]
- Shrestha, R.; Arnaud, E.; Mauleon, R.; Senger, M.; Davenport, G.; Hancock, D.; Morrison, N.; Bruskiewich, R.; McLaren, G. Multifunctional crop trait ontology for breeders’ data: Field book, annotation, data discovery and semantic enrichment of the literature. AoB Plants 2010, 2010, plq008. [Google Scholar] [CrossRef] [PubMed]
- Cooper, L.; Meier, A.; Laporte, M.-A.; Elser, J.L.; Mungall, C.; Sinn, B.T.; Cavaliere, D.; Carbon, S.; Dunn, N.A.; Smith, B.; et al. The Planteome database: An integrated resource for reference ontologies, plant genomics and phenomics. Nucleic Acids Res. 2018, 46, D1168–D1180. [Google Scholar] [CrossRef]
- Sook, J.; Taein, L.; Ksenija, G.; Campbell, T.; Yu, J.; Humann, J.; Ru, S.; Edge-Garza, D.; Hough, H.; Main, D. The Breeding Information Management System (BIMS): An online resource for crop breeding. Database 2021, 2021, baab054. [Google Scholar] [CrossRef]
- Wang, K.; Wang, Z.; Li, F.; Ye, W.; Wang, J.; Song, G.; Yue, Z.; Cong, L.; Shang, H.; Zhu, S.; et al. The draft genome of a diploid cotton Gossypium raimondii. Nat. Genet. 2012, 44, 1098–1103. [Google Scholar] [CrossRef]
- Paterson, A.H.; Wendel, J.F.; Gundlach, H.; Guo, H.; Jenkins, J.; Jin, D.; Llewellyn, D.; Showmaker, K.C.; Shu, S.; Udall, J.; et al. Repeated polyploidization of Gossypium genomes and the evolution of spinnable cotton fibres. Nat. Cell Biol. 2012, 492, 423–427. [Google Scholar] [CrossRef] [PubMed]
- Udall, J.A.; Long, E.; Hanson, C.; Yuan, D.; Ramaraj, T.; Conover, J.L.; Gong, L.; Arick, M.; Grover, C.E.; Peterson, D.G.; et al. De Novo Genome Sequence Assemblies of Gossypium raimondii and Gossypium turneri. G3 Genes Genomes Genet. 2019, 9, 3079–3085. [Google Scholar] [CrossRef]
- Grover, C.E.; Arick, M.; Thrash, A.; Conover, J.L.; Sanders, W.S.; Peterson, D.; Frelichowski, J.E.; Scheffler, J.A.; Scheffler, B.; Wendel, J.F. Insights into the Evolution of the New World Diploid Cottons (Gossypium, SubgenusHouzingenia) Based on Genome Sequencing. Genome Biol. Evol. 2019, 11, 53–71. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Li, J.; Wang, P.; Liu, F.; Liu, Z.; Zhao, G.; Xu, Z.; Pei, L.; Grover, C.E.; Wendel, J.F.; et al. Comparative Genome Analyses Highlight Transposon-Mediated Genome Expansion and the Evolutionary Architecture of 3D Genomic Folding in Cotton. Mol. Biol. Evol. 2021, 38, 3621–3636. [Google Scholar] [CrossRef]
- Li, F.; Fan, G.; Wang, K.; Sun, F.; Yuan, Y.; Song, G.; Li, Q.; Ma, Z.; Lu, C.; Zou, C.; et al. Genome sequence of the cultivated cotton Gossypium arboreum. Nat. Genet. 2014, 46, 567–572. [Google Scholar] [CrossRef]
- Du, X.; Huang, G.; He, S.; Yang, Z.; Sun, G.; Ma, X.; Li, N.; Zhang, X.; Sun, J.; Liu, M.; et al. Resequencing of 243 diploid cotton accessions based on an updated A genome identifies the genetic basis of key agronomic traits. Nat. Genet. 2018, 50, 796–802. [Google Scholar] [CrossRef]
- Huang, G.; Wu, Z.; Percy, R.G.; Bai, M.; Li, Y.; Frelichowski, J.E.; Hu, J.; Wang, K.; John, Z.Y.; Zhu, Y. Genome sequence of Gossypium herbaceum and genome updates of Gossypium arboreum and Gossypium hirsutum provide insights into cotton A-genome evolution. Nat. Genet. 2020, 52, 516–524. [Google Scholar] [CrossRef]
- Grover, C.E.; Yuan, D.; Arick, M.A.; Miller, E.R.; Hu, G.; Peterson, D.G.; Wendel, J.F.; Udall, J.A. The Gossypium anomalum genome as a resource for cotton improvement and evolutionary analysis of hybrid incompatibility. BioRxiv 2021. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Ge, X.; Li, W.; Jin, Y.; Liu, L.; Hu, W.; Liu, F.; Chen, Y.; Peng, S.; Li, F. Cotton D genome assemblies built with long-read data unveil mechanisms of centromere evolution and stress tolerance divergence. BMC Biol. 2021, 19, 115. [Google Scholar] [CrossRef]
- Grover, C.E.; Yuan, D.; Arick, M.A.; Miller, E.R.; Hu, G.; Peterson, D.G.; Wendel, J.F.; Udall, J.A. The Gossypium stocksii genome as a novel resource for cotton improvement. bioRxiv 2021. [Google Scholar] [CrossRef]
- Grover, C.E.; Pan, M.; Yuan, D.; Arick, M.A.; Hu, G.; Brase, L.; Stelly, D.M.; Lu, Z.; Schmitz, R.J.; Peterson, D.G.; et al. The Gossypium longicalyx Genome as a Resource for Cotton Breeding and Evolution. G3 Genes Genomes Genet. 2020, 10, 1457–1467. [Google Scholar] [CrossRef] [PubMed]
- Cai, Y.; Cai, X.; Wang, Q.; Wang, P.; Zhang, Y.; Cai, C.; Xu, Y.; Wang, K.; Zhou, Z.; Wang, C.; et al. Genome sequencing of the Australian wild diploid species Gossypium australe highlights disease resistance and delayed gland morphogenesis. Plant Biotechnol. J. 2019, 18, 814–828. [Google Scholar] [CrossRef] [PubMed]
- Udall, J.A.; Long, E.; Ramaraj, T.; Conover, J.L.; Yuan, D.; Grover, C.E.; Gong, L.; Arick, M.; Masonbrink, R.E.; Peterson, D.G.; et al. The Genome Sequence of Gossypioides kirkii Illustrates a Descending Dysploidy in Plants. Front. Plant Sci. 2019, 10, 1541. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Fan, G.; Lu, C.; Xiao, G.; Zou, C.; Kohel, R.J.; Ma, Z.; Shang, H.; Ma, X.; Wu, J.; et al. Genome sequence of cultivated Upland cotton (Gossypium hirsutum TM-1) provides insights into genome evolution. Nat. Biotechnol. 2015, 33, 524–530. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Hu, Y.; Jiang, W.; Fang, L.; Guan, X.; Chen, J.; Zhang, J.; Saski, C.A.; Scheffler, B.E.; Stelly, D.M.; et al. Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fiber improvement. Nat. Biotechnol. 2015, 33, 531–537. [Google Scholar] [CrossRef]
- Chen, Z.J.; Gossypium Hirsutum v1.1 (Upland Cotton) at Phytozome. 2017. Available online: https://phytozome-next.jgi.doe.gov/info/Ghirsutum_v1_1 (accessed on 17 November 2021).
- Wang, M.; Tu, L.; Yuan, D.; Zhu, D.; Shen, C.; Li, J.; Liu, F.; Pei, L.; Wang, P.; Zhao, G.; et al. Reference genome sequences of two cultivated allotetraploid cottons, Gossypium hirsutum and Gossypium barbadense. Nat. Genet. 2019, 51, 224–229. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Chen, J.; Fang, L.; Zhang, Z.; Ma, W.; Niu, Y.; Ju, L.; Deng, J.; Zhao, T.; Lian, J.; et al. Gossypium barbadense and Gossypium hirsutum genomes provide insights into the origin and evolution of allotetraploid cotton. Nat. Genet. 2019, 51, 739–748. [Google Scholar] [CrossRef]
- Yang, Z.; Ge, X.; Yang, Z.; Qin, W.; Sun, G.; Wang, Z.; Li, Z.; Liu, J.; Wu, J.; Wang, Y.; et al. Extensive intraspecific gene order and gene structural variations in upland cotton cultivars. Nat. Commun. 2019, 10, 2989. [Google Scholar] [CrossRef]
- Chen, Z.J.; Sreedasyam, A.; Ando, A.; Song, Q.; De Santiago, L.M.; Hulse-Kemp, A.M.; Ding, M.; Ye, W.; Kirkbride, R.C.; Jenkins, J.; et al. Genomic diversifications of five Gossypium allopolyploid species and their impact on cotton improvement. Nat. Genet. 2020, 52, 525–533. [Google Scholar] [CrossRef]
- Yuan, D.; Tang, Z.; Wang, M.; Gao, W.; Tu, L.; Jin, X.; Chen, L.; He, Y.; Zhang, L.; Zhu, L.; et al. The genome sequence of Sea-Island cotton (Gossypium barbadense) provides insights into the allopolyploidization and development of superior spinnable fibres. Sci. Rep. 2016, 5, 17662. [Google Scholar] [CrossRef]
- Boutet, E.; Lieberherr, D.; Tognolli, M.; Schneider, M.; Bairoch, A. UniProtKB/Swiss-Prot. In Methods in Molecular Biology v406: Plant Bioinformatics: Methods and Protocols; Edwards, D., Ed.; Humana Press Inc.: Totowa, NJ, USA, 2007; Volume 406, pp. 89–112. [Google Scholar]
- Schneider, M.; Lane, L.; Boutet, E.; Lieberherr, D.; Tognolli, M.; Bougueleret, L.; Bairoch, A. The UniProtKB/Swiss-Prot knowledgebase and its Plant Proteome Annotation Program. J. Proteom. 2008, 72, 567–573. [Google Scholar] [CrossRef]
- Benson, D.A.; Boguski, M.S.; Lipman, D.J.; Ostell, J. GenBank. Nucleic Acids Res. 1997, 25, 1–6. [Google Scholar] [CrossRef]
- Sayers, E.W.; Cavanaugh, M.; Clark, K.; Ostell, J.; Pruitt, K.; Mizrachi, I.K. GenBank. Nucleic Acids Res. 2019, 47, D94–D99. [Google Scholar] [CrossRef]
- Hunter, S.; Jones, P.; Mitchell, A.; Apweiler, R.; Attwood, T.K.; Bateman, A.; Bernard, T.; Binns, D.; Bork, P.; Burge, S.; et al. InterPro in 2011: New developments in the family and domain prediction database. Nucleic Acids Res. 2012, 40, D306–D312. [Google Scholar] [CrossRef]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.; Cherry, J.M.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T.; et al. Gene ontology: Tool for the unification of biology. Nat. Genet. 2000, 25, 25–29. [Google Scholar] [CrossRef]
- Kanehisa, M.; Goto, S.; Kawashima, S.; Okuno, Y.; Hattori, M. The KEGG resource for deciphering the genome. Nucleic Acids Res. 2004, 32, D277–D280. [Google Scholar] [CrossRef]
- Kanehisa, M.; Sato, Y.; Kawashima, M.; Furumichi, M.; Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 2016, 44, D457–D462. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Tang, H.; DeBarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.-H.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef] [PubMed]
- Page, J.T.; Huynh, M.D.; Liechty, Z.S.; Grupp, K.; Stelly, D.; Hulse, A.M.; Ashrafi, H.; Van Deynze, A.; Wendel, J.F.; Udall, J.A. Insights into the evolution of cotton diploids and polyploids from whole-genome re-sequencing. G3 Genes Genomes Genet. 2013, 3, 1809–1818. [Google Scholar] [CrossRef] [PubMed]
- Page, J.T.; Gingle, A.R.; Udall, J.A. PolyCat: A resource for genome categorization of sequencing reads from allopolyploid organisms. G3 Genes Genomes Genet. 2013, 3, 517–525. [Google Scholar] [CrossRef]
- Buels, R.; Yao, E.; Diesh, C.M.; Hayes, R.D.; Munoz-Torres, M.; Helt, G.; Goodstein, D.M.; Elsik, C.G.; Lewis, S.E.; Stein, L.; et al. JBrowse: A dynamic web platform for genome visualization and analysis. Genome Biol. 2016, 17, 66. [Google Scholar] [CrossRef]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T.L. BLAST+: Architecture and applications. BMC Bioinform. 2009, 10, 421. [Google Scholar] [CrossRef]
- Schläpfer, P.; Zhang, P.; Wang, C.; Kim, T.; Banf, M.; Chae, L.; Dreher, K.; Chavali, A.K.; Nilo-Poyanco, R.; Bernard, T.; et al. Genome-wide prediction of metabolic enzymes, pathways, and gene clusters in plants. Plant Physiol. 2017, 173, 2041–2059. [Google Scholar] [CrossRef]
- Caspi, R.; Dreher, K.; Karp, P.D. The challenge of constructing, classifying, and representing metabolic pathways. FEMS Microbiol. Lett 2013, 345, 85–93. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [PubMed]
- Gordon, D.; Abajian, C.; Green, P. Consed: A graphical tool for sequence finishing. Genome Res. 1998, 8, 195–202. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Huang, X.; Madan, A. CAP3: A DNA sequence assembly program. Genome Res. 1999, 9, 868–877. [Google Scholar] [CrossRef]
- Langmead, B.; Trapnell, C.; Pop, M.; Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009, 10, R25. [Google Scholar] [CrossRef]
- Fu, L.; Niu, B.; Zhu, Z.; Wu, S.; Li, W. CD-HIT: Accelerated for clustering the next-generation sequencing data. Bioinformatics 2012, 28, 3150–3152. [Google Scholar] [CrossRef]
- Davidson, N.M.; Oshlack, A. Corset: Enabling differential gene expression analysis for de novo assembled transcriptomes. Genome Biol. 2014, 15, 410. [Google Scholar] [PubMed]
- Jones, P.; Binns, D.; Chang, H.Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; et al. InterProScan 5: Genome-scale protein function classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef]
- Conesa, A.; Götz, S.; García-Gómez, J.M.; Terol, J.; Talón, M.; Robles, M. Blast2GO: A universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 2005, 21, 3674–3676. [Google Scholar] [CrossRef] [PubMed]
- Hulse-Kemp, A.M.; Lemm, J.; Plieske, J.; Ashrafi, H.; Buyyarapu, R.; Fang, D.D.; Frelichowski, J.; Giband, M.; Hague, S.; Hinze, L.L.; et al. Development of a 63K SNP Array for Cotton and High-Density Mapping of Intraspecific and Interspecific Populations of Gossypium spp. G3 Genes Genomes Genet. 2015, 5, 1187–1209. [Google Scholar] [CrossRef]
- Cai, C.; Zhu, G.; Zhang, T.; Guo, W. High-density 80 K SNP array is a powerful tool for genotyping G. hirsutum accessions and genome analysis. BMC Genom. 2017, 18, 654. [Google Scholar] [CrossRef] [PubMed]
- Buble, K.; Jung, S.; Humann, J.L.; Yu, J.; Cheng, C.-H.; Lee, T.; Ficklin, S.P.; Hough, H.; Condon, B.; Staton, M.E.; et al. Tripal MapViewer: A tool for interactive visualization and comparison of genetic maps. Database 2019, 2019, baz100. [Google Scholar] [CrossRef]
- Youens-Clark, K.; Faga, B.; Yap, I.V.; Stein, L.; Ware, D. CMap 1.01: A comparative mapping application for the Internet. Bioinformatics 2009, 25, 3040–3042. [Google Scholar] [CrossRef]
- Rife, T.; Poland, J.A. Field Book: An Open-Source Application for Field Data Collection on Android. Crop. Sci. 2014, 54, 1624–1627. [Google Scholar] [CrossRef]
- Gore, M.; Fang, D.; Poland, J.; Zhang, J.; Percy, R.G.; Cantrell, R.G.; Thyssen, G.; Lipka, A.E. Linkage Map Construction and Quantitative Trait Locus Analysis of Agronomic and Fiber Quality Traits in Cotton. Plant Genome 2014, 7, 1–10. [Google Scholar] [CrossRef]
- Shang, L.; Wang, Y.; Wang, X.; Liu, F.; Abduweli, A.; Cai, S.; Li, Y.; Ma, L.; Wang, K.; Hua, J. Genetic Analysis and Stable QTL Detection on Fiber Quality Traits Using Two Recombinant Inbred Line Populations and Their Backcross Progeny in Upland Cotton. G3 Genes Genomes Genet. 2016. [Google Scholar] [CrossRef]
- Jung, S.; Cheng, C.-H.; Buble, K.; Lee, T.; Humann, J.; Yu, J.; Crabb, J.; Hough, H.; Main, D. Tripal MegaSearch: A tool for interactive and customizable query and download of big data. Database 2021, 2021, baab023. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Henry, N.; Almsaeed, A.; Zhou, X.; Wegrzyn, J.; Ficklin, S.; Staton, M. New extension software modules to enhance searching and display of transcriptome data in Tripal databases. Database 2017, 2017, bax052. [Google Scholar] [CrossRef]
- Harper, L.C.; Campbell, J.D.; Cannon, E.; Jung, S.; Poelchau, M.; Walls, R.; Andorf, C.M.; Arnaud, E.; Berardini, T.; Birkett, C.; et al. AgBioData consortium recommendations for sustainable genomics and genetics databases for agriculture. Database 2018, 2018, bay088. [Google Scholar] [CrossRef] [PubMed]
| Type | By 8/14/13 | By 8/31/21 | Data Details by 31 August 2021 |
|---|---|---|---|
| Genome | G.raimondii (2) | 46 (30 diploids and 16 tetraploids) | Whole genome assemblies and annotations of 30 diploid species: G.anomalum (1), G.arboreum (4), G.aridum (1), G.armourianum (1), G.australe (1), G.davidsonii (2), G.gossypioides(1), G.harknessii (1), G.herbaceum (1), G.kirkii (1), G.klotzschianum (1), G.laxum (1), G.lobatum (1), G.longicalyx (1), G.raimondii (5), G.rotundifolium (1), G.schwendimanii (1), G.stocksii (1), G.thurberi (2), G.trilobum (1), G.turneri (1); and 5 tetraploid species: G.hirsutum (9), G.barbadense (4), G.tomentosum (1), G.mustelinum (1), G.darwinii (1) |
| Gene and mRNA | 119,971 genes | 1,874,940 genes and 2,528,191 mRNAs | Genes and mRNAs from whole genome assemblies and parsed from NCBI nucleotide sequences |
| Transcript | 149,916 | 214,180 RefTrans | RefTrans for G.hirsutum, G.barbadense, G.arboreum, G. raimondii |
| Marker | 26,089 | 587,004 | Including 459,825 SNPs (TAMU63K and NAU80K arrays, and other SNPs), 109,848 SSRs |
| Map | 49 | 115 | 130,533 loci from 110 genetic maps, 2 consensus maps, 2 bin maps, and 1 silico map |
| QTL | 988 | 6772 | Including 4178 quality traits, 1547 agronomical trait, 273 biotic stress traits, and 189 biochemical traits |
| Species | 50 | 85 | Including the 4 cultivated species, 53 wild species, and 28 cross or lab made diploid, tetraploid, and hexaploidy hybrids |
| Germplasm | 14,959 | 19,827 | Including collection and sub-collections from US-NCGC, US-GRIN, China, and Uzbekistan |
| Phenotype data | 118,302 | 539,975 | Phenotypic scores from the US regional breeder’s tests; the trait evaluations from US, Uzbekistan, and China germplasm collections; and the data collected from various QTL studies |
| Genotype data | 68,640 | 25,532,891 | SNP genotype data from 25,213,321 measurements using 71,424 markers, SSR genotype data from 319,570 measurements using 2825 markers |
| Image | 0 | 45,211 | Including 44,998 NCGC digital characterizations |
| Publication | 10,731 | 16,066 | Including journal articles, conference proceedings, patents, book chapters, and theses/dissertations |
| Library | 181 | 181 | Including 135 cDNA, 41 genomic DNA, 2 SNP chip, and 2 unassigned libraries |
| Genome Sequence Name | Germplasm Type | Pub Year (Ref.) |
|---|---|---|
| Gossypium raimondii (D5) ‘D5-3’ genome CGP-BGI_v1 | wild | 2012 [14] |
| Gossypium raimondii (D5) genome JGI_v2_a2.1 | wild | 2012 [15] |
| Gossypium raimondii (D5) ‘D5-4’ genome NSF_v1 | wild | 2019 [16] |
| Gossypium raimondii (D5) ‘D5-8’ genome ISU_v1 | wild | 2019 [17] |
| Gossypium raimondii (D5) ‘D502’ genome HAU_v1 | wild | 2021 [18] |
| Gossypium arboreum (A2) ‘SXY1’ genome CGP-BGI_v2_a1 | cultivar | 2014 [19] |
| Gossypium arboreum (A2) ‘SXY1’ genome CRI-updated_v1 | cultivar | 2018 [20] |
| Gossypium arboreum (A2) ‘SXY1’ genome WHU-updated_v1 | cultivar | 2020 [21] |
| Gossypium arboreum (A2) ‘SXY1’ genome HAU_v1 | cultivar | 2021 [18] |
| Gossypium herbaceum (A1) ‘Mutema’ genome WHU_v1 | cultivar | 2020 [21] |
| Gossypium anomalum (B1) genome NSF_v1 | wild | 2021 [22] |
| Gossypium thurberi (D1-35) genome ISU_v1 | wild | 2019 [17] |
| Gossypium thurberi (D1-5) genome CRI_v1 | wild | 2021 [23] |
| Gossypium armourianum (D2-1) genome ISU_v1 | wild | 2019 [17] |
| Gossypium harknessii (D2-2) genome ISU_v1 | wild | 2019 [17] |
| Gossypium davidsonii (D3d-27) genome ISU_v1 | wild | 2019 [17] |
| Gossypium davidsonii (D3d-8) genome CRI_v1 | wild | 2021 [23] |
| Gossypium klotzschianum (D3-k) genome ISU_v1 | wild | 2019 [17] |
| Gossypium aridum (D4) genome ISU_v1 | wild | 2019 [17] |
| Gossypium gossypioides (D6) genome ISU_v1 | wild | 2019 [17] |
| Gossypium lobatum (D7) genome ISU_v1 | wild | 2019 [17] |
| Gossypium trilobum (D8) genome ISU_v1 | wild | 2019 [17] |
| Gossypium laxum (D9) genome ISU_v1 | wild | 2019 [17] |
| Gossypium turneri (D10) genome NSF_v1_a2 | wild | 2019 [17] |
| Gossypium schwendimanii (D11) genome ISU_v1 | wild | 2019 [17] |
| Gossypium stocksii (E1) genome NSF_v1 | wild | 2021 [24] |
| Gossypium longicalyx (F1) genome NSF_v1 | wild | 2020 [25] |
| Gossypium australe (G2) genome CRI_v1.1 | wild | 2019 [26] |
| Gossypium rotundifolium (K12) ‘Grot K201’ genome HAU_v1 | wild | 2021 [18] |
| Gossypioides kirkii genome ISU_v3 | wild | 2019 [27] |
| Genome Sequence Name | Germplasm Type | Pub Year (ref.) |
|---|---|---|
| Gossypium hirsutum (AD1) ‘TM-1’ genome CGP-BGI_v1 | cultivar | 2015 [28] |
| Gossypium hirsutum (AD1) ‘TM-1’ genome NAU-NBI_v1.1 | cultivar | 2015 [29] |
| Gossypium hirsutum (AD1) ‘TM-1’ Genome UTX-JGI-interim-release_v1.1 | cultivar | 2017 [30] |
| Gossypium hirsutum (AD1) ‘TM-1’ genome HAU_v1 | cultivar | 2018 [31] |
| Gossypium hirsutum (AD1) ‘TM-1’ genome ZJU-improved v2.1_a1 | cultivar | 2019 [32] |
| Gossypium hirsutum (AD1) ‘TM-1’ genome CRI_v1 | cultivar | 2019 [33] |
| Gossypium hirsutum (AD1) ‘ZM24’ genome CRI_v1 | cultivar | 2019 [33] |
| Gossypium hirsutum (AD1) ‘TM-1’ genome WHU_v1 | cultivar | 2020 [21] |
| Gossypium hirsutum (AD1) ‘TM-1’ genome UTX_v2.1 | cultivar | 2020 [34] |
| Gossypium barbadense (AD2) ‘3-79’ genome HAU_v1 | cultivar | 2015 [35] |
| Gossypium barbadense (AD2) ‘3-79’genome HAU_v2_a1 | cultivar | 2018 [31] |
| Gossypium barbadense (AD2) ‘H7124’ genome ZJU_v1.1_a1 | cultivar | 2019 [32] |
| Gossypium barbadense (AD2) ‘3-79’ genome HGS_v1.1 | cultivar | 2020 [34] |
| Gossypium tomentosum (AD3) gnome HGS_v1.1 | wild | 2020 [34] |
| Gossypium mustelinum (AD4) genome JGI_v1.1 | wild | 2020 [34] |
| Gossypium darwinii (AD5) genome HGS_v1.1 | wild | 2020 [34] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yu, J.; Jung, S.; Cheng, C.-H.; Lee, T.; Zheng, P.; Buble, K.; Crabb, J.; Humann, J.; Hough, H.; Jones, D.; et al. CottonGen: The Community Database for Cotton Genomics, Genetics, and Breeding Research. Plants 2021, 10, 2805. https://doi.org/10.3390/plants10122805
Yu J, Jung S, Cheng C-H, Lee T, Zheng P, Buble K, Crabb J, Humann J, Hough H, Jones D, et al. CottonGen: The Community Database for Cotton Genomics, Genetics, and Breeding Research. Plants. 2021; 10(12):2805. https://doi.org/10.3390/plants10122805
Chicago/Turabian StyleYu, Jing, Sook Jung, Chun-Huai Cheng, Taein Lee, Ping Zheng, Katheryn Buble, James Crabb, Jodi Humann, Heidi Hough, Don Jones, and et al. 2021. "CottonGen: The Community Database for Cotton Genomics, Genetics, and Breeding Research" Plants 10, no. 12: 2805. https://doi.org/10.3390/plants10122805
APA StyleYu, J., Jung, S., Cheng, C.-H., Lee, T., Zheng, P., Buble, K., Crabb, J., Humann, J., Hough, H., Jones, D., Campbell, J. T., Udall, J., & Main, D. (2021). CottonGen: The Community Database for Cotton Genomics, Genetics, and Breeding Research. Plants, 10(12), 2805. https://doi.org/10.3390/plants10122805






