The Effect of a Host on the Primary Metabolic Profiling of Cuscuta Campestris’ Main Organs, Haustoria, Stem and Flower
Abstract
:1. Introduction
2. Results
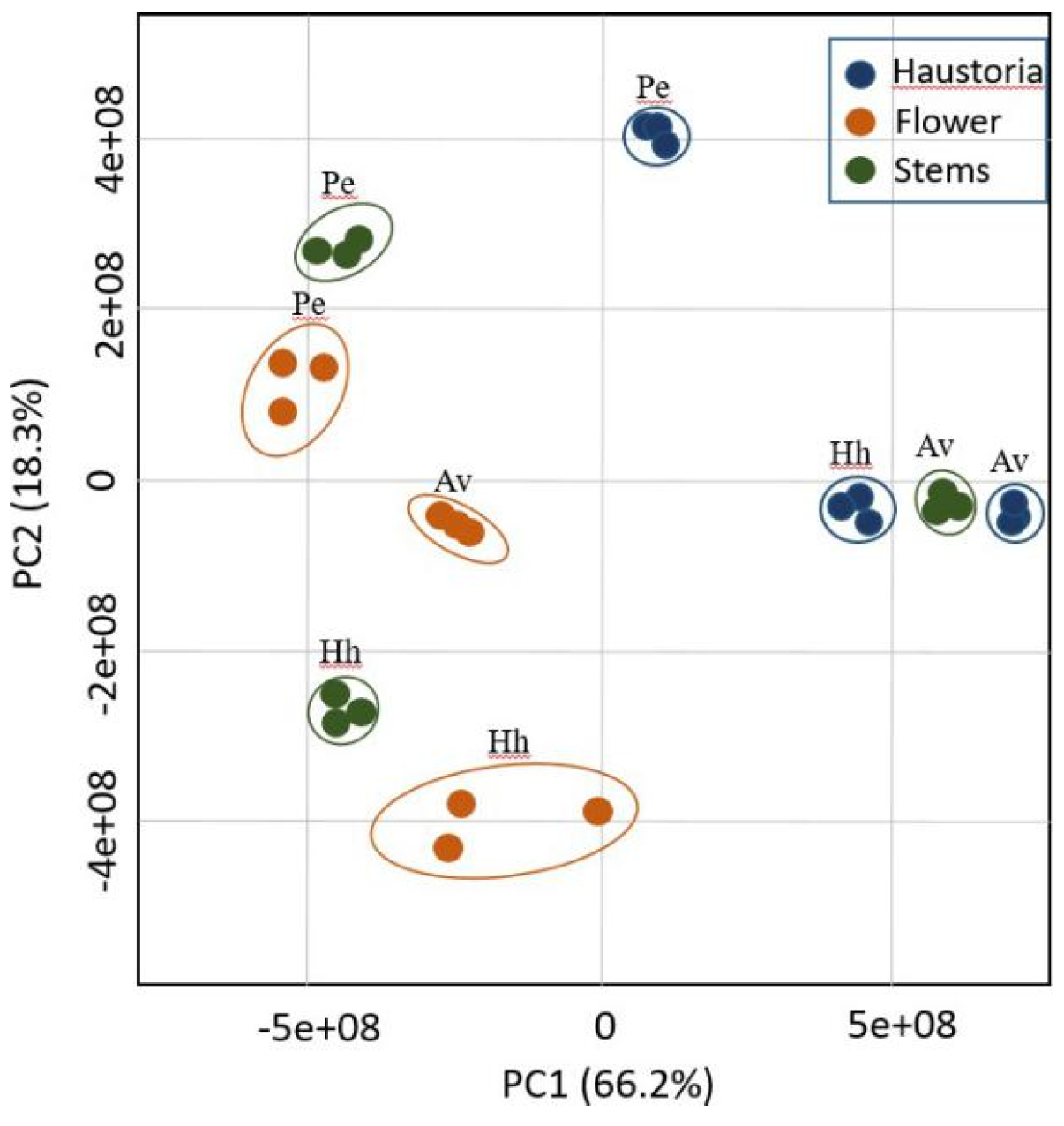
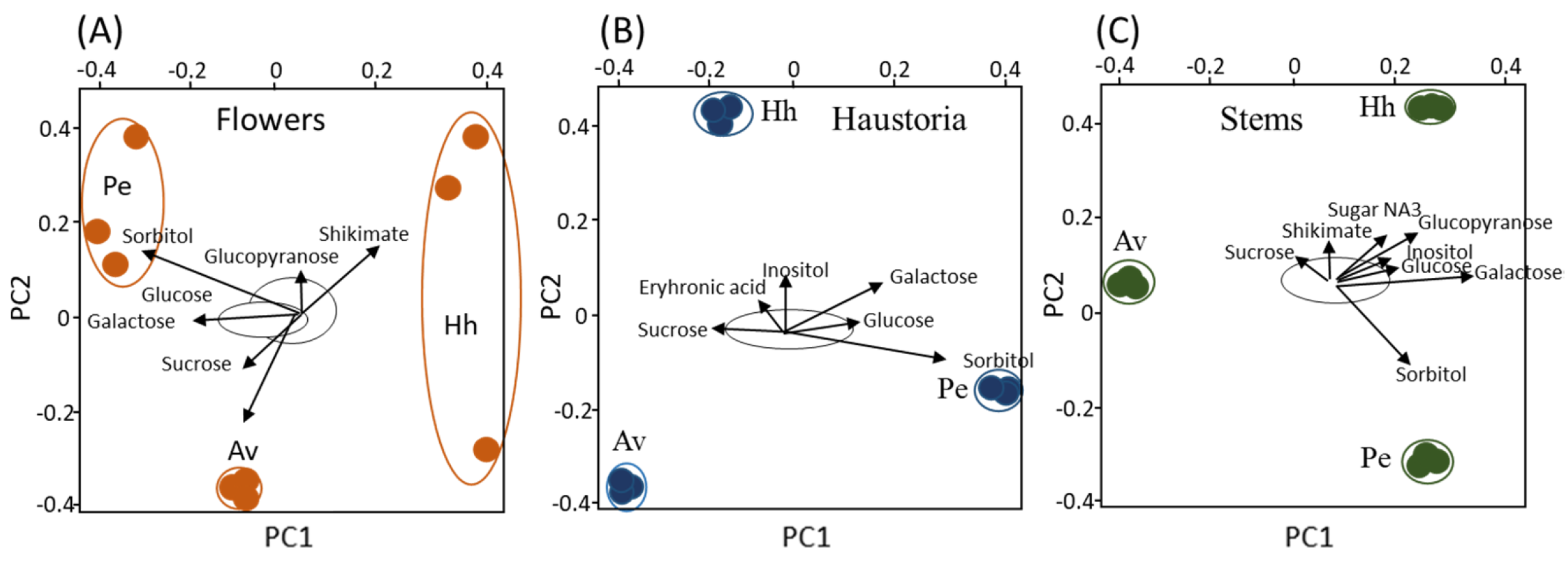
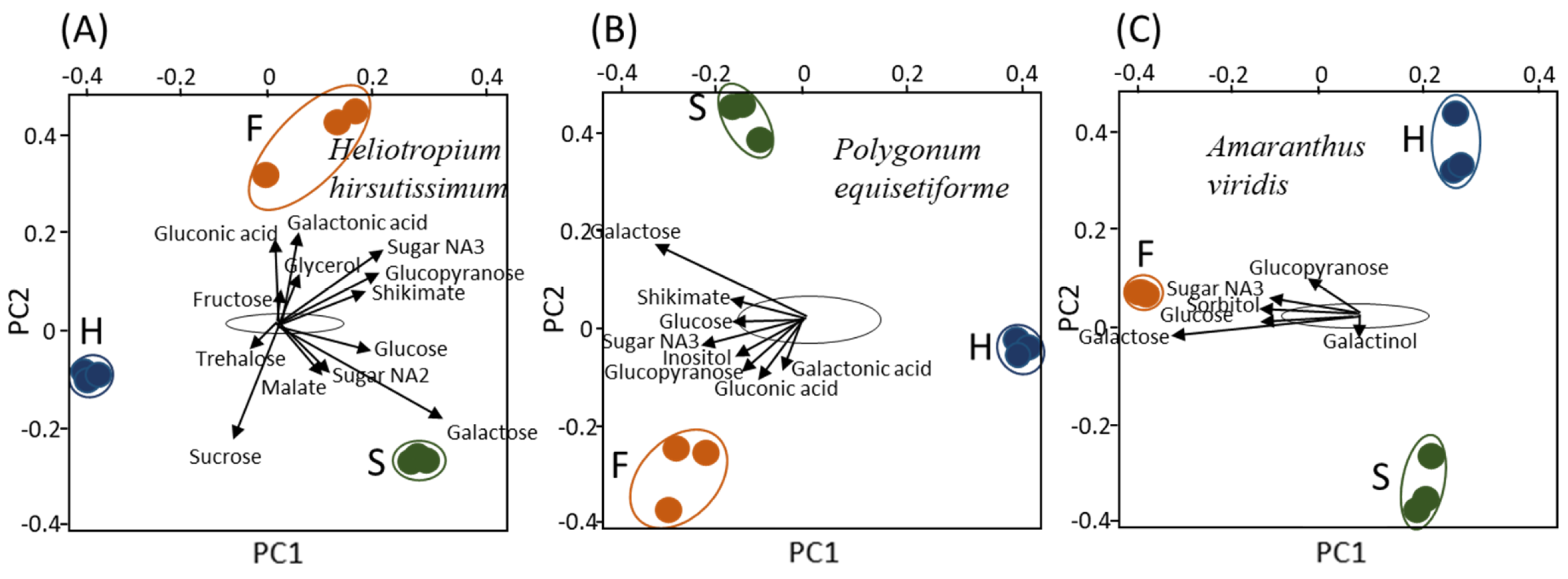
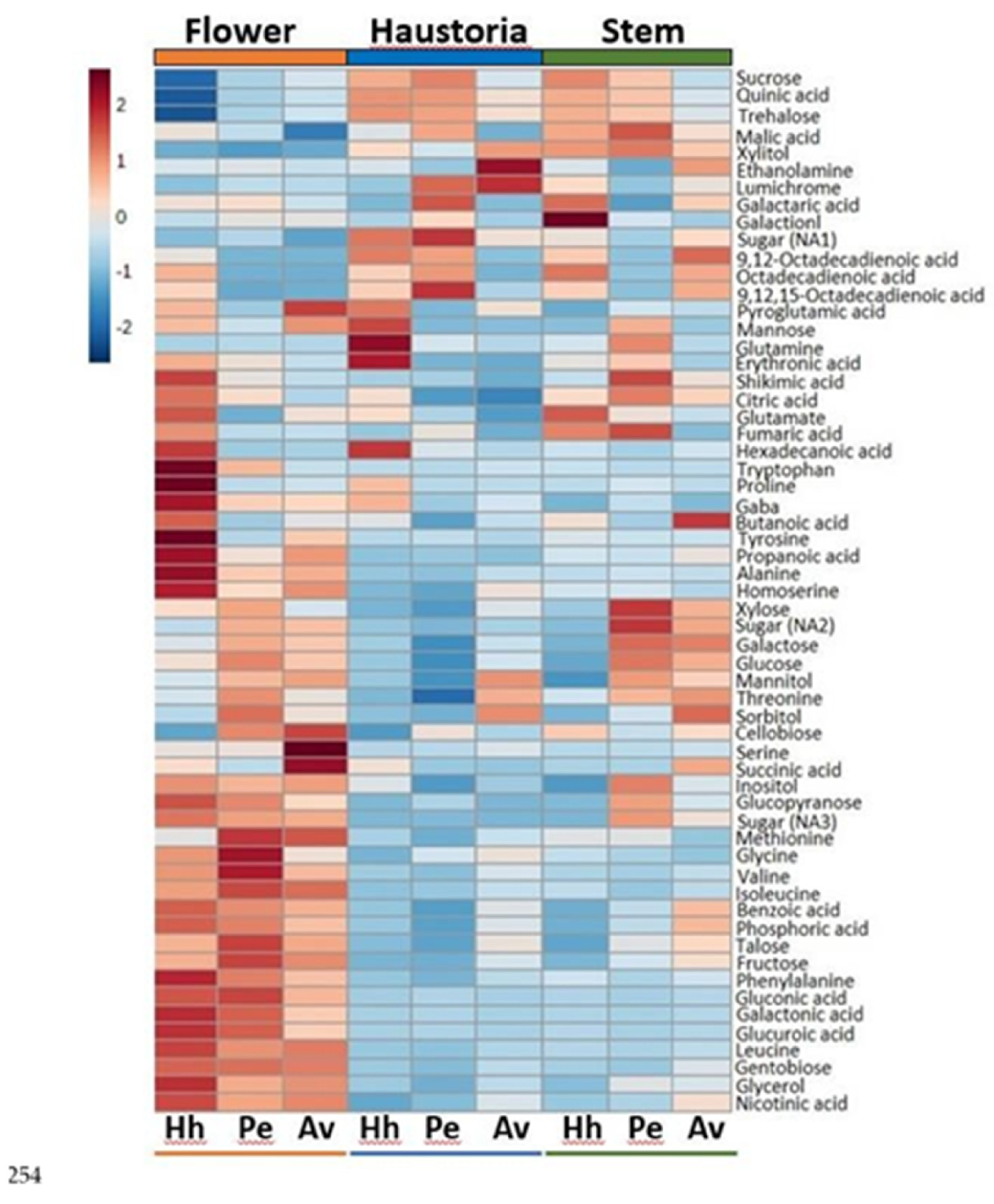
2.1. Primary Metabolic Profiles Analysis Using Gas Chromatography-Mass Spectrometry (GC-MS) Reveals a Distinct Metabolic Profile in Haustoria, Stem and Flowers with Respects to the Hosts
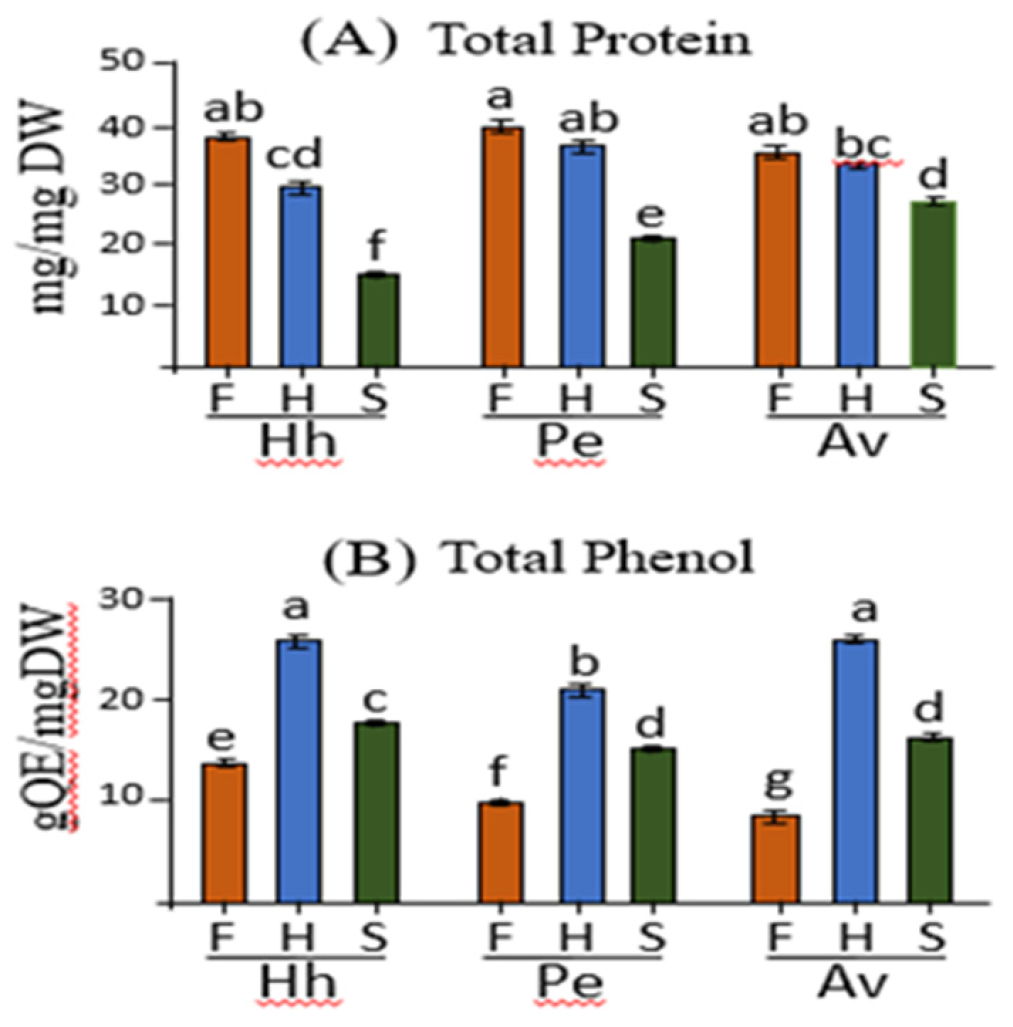
2.2. Determination of Total Soluble Protein and Total Phenolic Compounds
3. Discussion
4. Materials and Methods
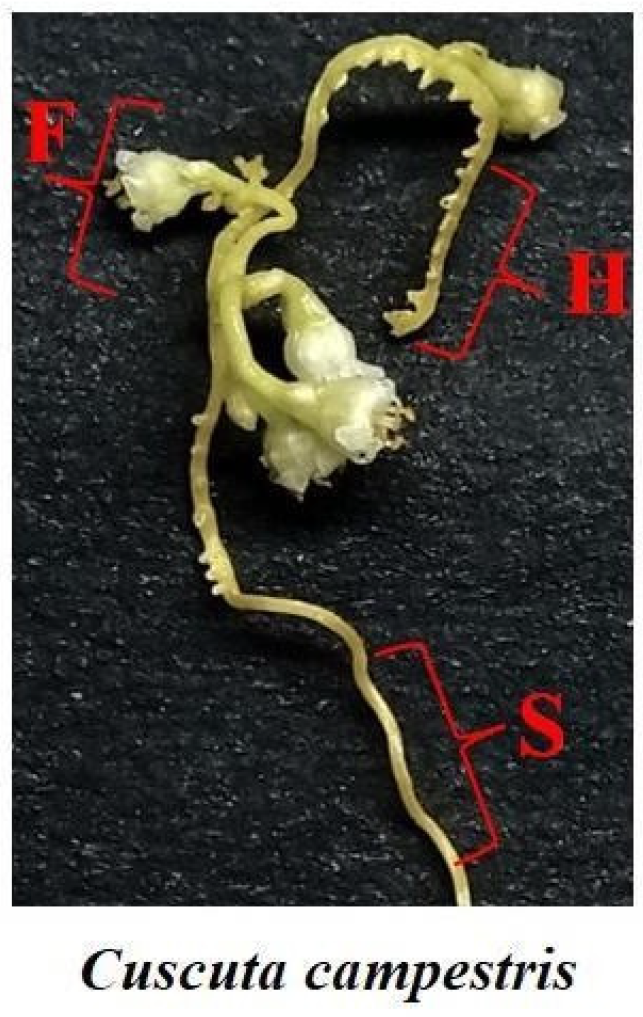
4.1. Plant Material and Sample Collection
4.2. Extraction and Analysis of Primary Metabolites Using GC-MS
4.3. Total Soluble Protein Determination
4.4. Statistical Analyses
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Smith, J.D.; Mescher, M.C.; De Moraes, C.M. Implications of bioactive solute transfer from hosts to parasitic plants. Curr. Opin. Plant. Biol. 2013, 16, 464–472. [Google Scholar] [CrossRef]
- Kim, G.; Westwood, J.H. Macromolecule exchange in Cuscuta-host plant interactions. Curr. Opin. Plant. Biol. 2015, 26, 20–25. [Google Scholar] [CrossRef] [Green Version]
- Saric-Krsmanovic, M.; Vrbnicanin, S. Field dodder-how to control it? Pestic. I Fitomed. 2015, 30, 137–145. [Google Scholar] [CrossRef]
- Ahmad, A.; Tandon, S.; Xuan, T.D.; Nooreen, Z. A Review on Phytoconstituents and Biological activities of Cuscuta species. Biomed. Pharm. 2017, 92, 772–795. [Google Scholar] [CrossRef]
- Furuhashi, T.; Nakamura, T.; Iwase, K. Analysis of metabolites in stem parasitic plant interactions: Interaction of Cuscuta–Momordica versus Cassytha–Ipomoea. Plants 2016, 5, 43. [Google Scholar] [CrossRef] [PubMed]
- Flores-Sánchez, I.J.; Garza-Ortiz, A. Is There a Secondary/Specialized Metabolism in the Genus Cuscuta and which Is the Role of the Host Plant? Phytochem Rev. 2019, 18, 1299–1335. [Google Scholar] [CrossRef]
- Yu, H.; Liu, J.; He, W.M.; Miao, S.L.; Dong, M. Cuscuta australis restrains three exotic invasive plants and benefits native species. Biol Invasions 2011, 13, 747–756. [Google Scholar] [CrossRef]
- Clermont, K.; Wang, Y.; Liu, S.; Yang, Z.; dePamphilis, C.W.; Yoder, J.I. Comparative Metabolomics of Early Development of the Parasitic Plants Phelipanche aegyptiaca and Triphysaria versicolor. Metabolites 2019, 9, 114. [Google Scholar] [CrossRef] [Green Version]
- Nickrent, D.L.; Musselman, L.J. Introduction to Parasitic Flowering Plants. Plant Health Instr. 2004, 13, 300–315. [Google Scholar] [CrossRef]
- Vogel, A.; Schwacke, R.; Denton, A.K.; Usadel, B.; Hollmann, J.; Fischer, K.; Bolger, A.; Schmidt, M.H.W.; Bolger, M.E.; Gundlach, H.; et al. Footprints of parasitism in the genome of the parasitic flowering plant Cuscuta campestris. Nat. Commun. 2018, 9, 2515. [Google Scholar] [CrossRef]
- Clarke, C.R.; Timko, M.P.; Yoder, J.I.; Axtell, M.J.; Westwood, J.H. Molecular Dialog between Parasitic Plants and Their Hosts. Annu. Rev. Phytopathol. 2019, 57, 279–299. [Google Scholar] [CrossRef] [PubMed]
- Hacham, Y.; Hershenhorn, J.; Dor, E.; Amir, R. Primary metabolic profiling of Egyptian broomrape (Phelipanche aegyptiaca) compared to its host tomato roots. J. Plant Physiol. 2016, 205, 11–19. [Google Scholar] [CrossRef] [PubMed]
- Nandula, V.K.; Foster, J.G.; Foy, C.L. Impact of Egyptian broomrape (Orobanche aegyptiaca (Pers.) parasitism on amino acid composition of carrot (Daucus carota L.). J. Agric. Food Chem. 2000, 48, 3930–3934. [Google Scholar] [CrossRef]
- Wakabayashi, T.; Joseph, B.; Yasumoto, S.; Akashi, T.; Aoki, T.; Harada, K.; Muranaka, S.; Bamba, T.; Fukusaki, E.; Takeuchi, Y.; et al. Planteose as a storage carbohydrate required for early stage of germination of Orobanche minor and its metabolism as a possible target for selective control. J. Exp. Bot. 2015, 66, 3085–3097. [Google Scholar] [CrossRef] [Green Version]
- Obata, T.; Fernie, A.R. The use of metabolomics to dissect plant responses to abiotic stresses. Cell Mol. Life Sci. 2012, 69, 3225–3243. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nakabayashi, R.; Saito, K. Integrated metabolomics for abiotic stress responses in plants. Curr. Opin. Plant. Biol. 2015, 24, 10–16. [Google Scholar] [CrossRef] [Green Version]
- Al-Gburi, B.K.H.; Al-Sahaf, F.H.; Al-fadhal, F.A.; Del Monte, J.P. Detection of phytochemical compounds and pigments in seeds and shoots of Cuscuta campestris parasitizing on eggplant. Physiol. Mol. Biol. Plants 2019, 25, 253–261. [Google Scholar] [CrossRef] [PubMed]
- Xia, J.; Sinelnikov, I.V.; Han, B.; Wishart, D.S. MetaboAnalyst 3.0-making metabolomics more meaningful. Nucleic Acids Res. 2015, 43, W251–W257. [Google Scholar] [CrossRef] [Green Version]
- Kaiser, B.; Vogg, G.; Fürst, U.B.; Albert, M. Parasitic plants of the genus Cuscuta and their interaction with susceptible and resistant host plants. Front. Plant. Sci 2015, 6, 45. [Google Scholar] [CrossRef] [Green Version]
- Koch, A.M.; Binder, C.; Sanders, I.R. Does the generalist parasitic plant Cuscuta campestris selectively forage in heterogeneous plant communities? New Phytol. 2004, 162, 147–155. [Google Scholar] [CrossRef] [Green Version]
- Nativ, N.; Hacham, Y.; Hershenhorn, J.; Dor, E.; Amir, R. Metabolic investigation of Phelipanche aegyptiaca reveals significant changes during developmental stages and in its different organs. Front. Plant. Sci. 2017, 8, 491. [Google Scholar] [CrossRef] [Green Version]
- Löffler, C.; Sahm, A.; Wray, V.; Czygan, F.C.; Proksch, P. Soluble phenolic constituents from Cuscuta reflexa and Cuscuta platyloba. Biochem. Syst. Ecol. 1995, 23, 121–128. [Google Scholar] [CrossRef]
- Isah, T. Stress and defense responses in plant secondary metabolites production. Biol. Res. 2019, 52, 39. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Özen, H.Ç.; Surmuş Asan, H. The effect of Cuscuta babylonica Aucher (Cuscuta) parasitism on the phenolic contents of Carthamus glaucus Bieb.subsp. glaucus. J. Inst. Sci. Technol. 2016, 4, 31. [Google Scholar] [CrossRef]
- Furuhashi, T.; Fragner, L.; Furuhashi, K.; Valledor, L.; Sun, X.; Weckwerth, W. Metabolite changes with induction of Cuscuta haustorium and translocation from host plants. J. Plant. Interact. 2012, 7, 84–93. [Google Scholar] [CrossRef]
- Anis, E.; Ullah, N.; Mustafa, G.; Malik, A.; Alza, N.; Badar, Y. Phytochemical studies on Cuscuta reflexa. J. Nat. Prod. 1999, 5, 124–126. [Google Scholar]
- Bais, N.; Kakkar, A. Phytochemical Analysis of Methanolic Extract of Cuscuta reflexa Grown on Cassia fistula and Ficus benghalensis by GC-MS. Int. J. Pharm. Sci. Rev. Res. 2014, 25, 33–36. [Google Scholar]
- Tanruean, K.; Kaewnarin, K.; Suwannarach, N.; Lumyong, S. Comparative evaluation of phytochemicals, and antidiabetic and antioxidant activities of Cuscuta reflexa grown on different hosts in Northern Thailand. Nat. Prod. Commun. 2017, 12, 51–54. [Google Scholar] [CrossRef] [Green Version]
- Hibberd, J.M.; Jeschke, W.D. Solute flux into parasitic plants. J. Exp. Bot. 2001, 52, 2043–2049. [Google Scholar] [CrossRef] [Green Version]
- Cohen, H.; Israeli, H.; Matityahu, I.; Amir, R. Seed-specific expression of a feedback-insensitive form of CYSTATHIONINE-SYNTHASE in arabidopsis stimulates metabolic and transcriptomic responses associated with desiccation stress. Plant. Physiol. 2014, 166, 1575–1592. [Google Scholar] [CrossRef] [Green Version]
- Ben Nasr, C.; Ayed, N.; Metche, M. Quantitative determination of the polyphenolic content of pomegranate peel. Z. Lebensm. Unters. Forsch. 1996, 203, 374–378. [Google Scholar] [CrossRef] [PubMed]
| Flowers | Haustoria | Stem | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Metabolites | Hh | Pe | Av | Hh | Pe | Av | Hh | Pe | Av |
| Sugars | |||||||||
| Sucrose | 229 ± 46 e | 366 ± 16 cd | 2 ± 0.03 f | 710 ± 9 a | 463 ± 17 bc | 763 ± 40 a | 535 ± 8 b | 463 ± 10 bc | 284 ± 1 de |
| Glucose | 373 ± 36 bc | 473 ± 21 ab | 325 ± 5 bc | 183 ± 9 bc | 253 ± 7 bc | 30 ± 1 c | 747 ± 215 a | 485 ± 17 ab | 32 ± 2 c |
| Trehalose | 38 ± 8 e | 78 ± 2 c | 372 ± 6 d | 159 ± 2 a | 110 ± 4 b | 152 ± 7 a | 113 ± 1 b | 107 ± 1 c | 55 ± 1 de |
| Galactose | 828 ± 69 c | 988 ± 36 b | 794 ± 4 cd | 614 ± 18 e | 659 ± 16 de | 284 ± 12 f | 1206 ± 12 a | 1318 ± 30 a | 168 ± 5 f |
| Talose | 54 ± 2 b | 65 ± 1 a | 41 ± 0.2 c | 10 ± 1 e | 32 ± 1 d | 3 ± 0.2 f | 27 ± 1 d | 42 ± 1 c | 1 ± 0.07 f |
| Fructose | 115 ± 5 ab | 137 ± 4 a | 97 ± 1 bc | 14 ± 1 e | 54 ± 1 d | 11 ± 0.5 e | 48 ± 10 d | 81 ± 9 c | 5 ± 0.5 e |
| Glucopyranose | 367 ± 14 a | 255 ± 54 ab | 18 ± 0.9 d | 24 ± 1 d | 18 ± 0.8 d | 76 ± 3 d | 238 ± 38 bc | 122 ± 19 cd | 9 ± 0.5 d |
| Mannose | 2 ± 0.1 bc | 1 ± 0.02 bc | 460 ± 2 a | 4 ± 0 b | ND | ND | 2 ± 0.02 bc | ND | ND |
| Cellobiose | 3 ± 1 c | 39 ± 2 a | 7 ± 0.4 c | 1 ± 0.1 c | 13 ± 1 c | 31 ± 3 ab | 16 ± 7 bc | 30 ± 2 ab | 13 ± 0.9 c |
| Gentobiose | 11 ± 0.5 a | 8 ± 1 b | 2 ± 0.1 de | 1 ± 0.06 de | 2 ± 0.01 d | 1 ± 0.02 e | 1 ± 0.02 de | 4 ± 0.6 c | 1 ± 0.05 e |
| Xylose | 11 ± 1 abc | 11 ± 1 ab | 5 ± 0.3 cd | 3 ± 0.09 de | 7 ± 0.4 bcd | 1 ± 0.01 e | 16 ± 2 a | 13 ± 1 a | 1 ± 0.02 e |
| Sugar (NA1) | 1 ± 0.04 c | 2 ± 0.3 c | ND | 5 ± 3 a | 2 ± 0.05 ab | 6 ± 0.9 a | 1 ± 0.01 c | 2 ± 0.2 ab | 1 ± 0.01 c |
| Sugar (NA2) | 50 ± 1 c | 123 ± 2 b | 98 ± 2 b | 23 ± 1 cd | 32 ± 0.8 cd | 6 ± 0.6 d | 193 ± a | 134 ± 2 b | 4 ± 0.8 d |
| Sugar (NA3) | 513 ± 4 a | 358 ± 9 b | 297 ± 3 b | 5 ± 0.7 d | 2 ± 0.01 d | 13 ± 0.8 d | 379 ± 6 ab | 232 ± 23 c | 1 ± 0.07 d |
| Sugar acids | |||||||||
| Galactonic acid | 119 ± 1.9 b | 83 ± 1 c | 148 ± 4 a | 1 ± 0.02 d | 2 ± 0.1 d | 5 ± 0.4 d | 2 ± 0.4 d | 7 ± 0.1 d | 2 ± 0.06 d |
| Glucuronte | 15 ± 0.9 a | 11 ± 0.5 b | 109 ± 1 c | ND | ND | 1 ± 0.02 d | ND | 1 ± 0.04 d | ND |
| Gluconic acid | 242 ± 38 a | 207 ± 5 a | 40 ± 1 b | 2 ± 0.01 b | ND | 16 ± 1 b | 4 ± 0.6 b | 18 ± 1 b | 5 ± 0.5 b |
| Galactaric acid | 4 ± 0.4 cd | 3 ± 0.4 d | 5 ± 0.8 b | 2 ± 0.06 e | 2 ± 0.06 e | 6 ± 0.7 a | 1 ± 0.05 e | 4 ± 0.9 bc | 2 ± 0.3 e |
| TCA metabolites | |||||||||
| Malic acid | 329 ± 10 b | 244 ± 5 c | 150 ± 1 d | 332 ± 5 b | 226 ± 5 c | 381 ± 8 a | 365 ± 15 ab | 328 ± 3 b | 150 ± 1 d |
| Citric acid | 264 ± 13 a | 191 ± 3 cd | 152 ± 1 f | 241 ± 2 ab | 161 ± 2 ef | 186 ± 5 de | 219 ± 7 bc | 230 ± 2 b | 90 ± 0.9 g |
| Succinic acid | 44 ± 2 abc | 19 ± 0.7 bc | 77 ± 1 a | 43 ± 15 abc | 11 ± 0.8 c | 13 ± 0.6 c | 15 ± 1 c | 61 ± 21 ab | 6 ± 0.4 c |
| Fumaric acid | 73 ± 2 a | 22 ± 0.7 cd | 21 ± 0.9 d | 17 ± 0.9 de | 8 ± 0.5 ef | 40 ± 0.4 b | 76 ± 4 a | 13 ± 0.3 f | 30 ± 1 c |
| Organic acids | |||||||||
| Shikimic acid | 645 ± 45 a | 309 ± 32 bc | 224 ± 3 cd | 279 ± 13 bc | 173 ± 9 de | 278 ± 16 bc | 548 ± 17 a | 380 ± 12 b | 123 ± 1 e |
| Benzoic acid | 16 ± 0.8 a | 12 ± 0.7 b | 9 ± 0.9 c | 5 ± 0.4 e | 8 ± 0.6 d | 3 ± 0.5 f | 6 ± 0.6 e | 12 ± 0.5 b | 1 ± 0.1 g |
| Pyroglutamate | 10 ± 0.7 ab | 5 ± 0.6 de | 9 ± 0.6 ab | 12 ± 1 a | 7 ± 0.5 bcd | 5 ± 0.4 d | 6 ± 0.5 cd | 6 ± 1 bcd | 1 ± 0.04 e |
| Nicotinic acid | 3 ± 0.2 a | 2 ± 0.02 ab | 2 ± 0.3 ab | 1 ± 0 bc | 1 ± 0.01 abc | 1 ± 0.01 bc | 1 ± 0.06 bc | 2 ± 0.02 ab | 1 ± 0.06 c |
| Quinic acid | 1 ± 0.01 d | 1 ± 0.1 cd | 42 ± 1 cd | 4 ± 0 a | 2 ± 0.3 bcd | 4 ± 0.1 a | 2 ± 0.2 bc | 2 ± 0.3 bcd | 1 ± 0.03 cd |
| Butanoic acid | 1 ± 0.2 a | ND | ND | ND | ND | ND | ND | 1 ± 0.01 a | ND |
| Propanoic acid | 23 ± 1 a | 10 ± 0.3 bcd | 12 ± 0.4 b | 6 ± 0.5 def | 5 ± 0.6 ef | 7 ± 0.6 de | 7 ± 1 cde | 11 ± 1 bc | 3 ± 0.02 f |
| Phosphoric acid | 327 ± 4 a | 246 ± 1 b | 174 ± 3 c | 99 ± 6 e | 147 ± 5 d | 49 ± 0.4 f | 110 ± 7 e | 241 ± 4 b | 25 ± 0.4 g |
| Erythronic acid | 86 ± 6 b | 51 ± 1 c | 32 ± 0.6 d | 142 ± 6 a | 148 ± 1 e | 19 ± 0.9 de | 65 ± 2 c | 32 ± 1 d | 23 ± 1 de |
| GABA | 57 ± 1 a | 22 ± 0.2 c | 18 ± 0.9 c | 34 ± 1 b | 13 ± 0.7 d | 6 ± 0.7 e | 10 ± 0.9 d | 2 ± 0.4 f | 1 ± 0.07 f |
| Polyols | |||||||||
| Mannitol | 87 ± 35 bc | 135 ± 2 ab | 134 ± 1 ab | 53 ± 2 cd | 173 ± 4 a | 3 ± 0.8 d | 156 ± 4 a | 139 ± 2 ab | 4 ± 2 d |
| Xylitol | 26 ± 11 cd | 33 ± 1 abcd | 30 ± 0.1 b | 50 ± 1 a | 46 ± 0.9 abc | 45 ± 1 abc | 43 ± 1 abc | 47 ± 2 ab | 18 ± 1 d |
| Inositol | 646 ± 24 a | 488 ± 12 c | 2 ± 0 g | 470 ± 5 cd | 317 ± 5 e | 260 ± 14 e | 563 ± 10 b | 430 ± 9 d | 96 ± 3 f |
| Galactinol | 20 ± 2 c | 33 ± 2 bc | 76 ± 1 ab | 13 ± 1 c | 7 ± 0.4 c | 61 ± 24 ab | 24 ± 1 bc | 7 ± 0.6 c | 63 ± 1 a |
| Sorbitol | 187 ± 80 cd | 686 ± 11 ab | 303 ± 2 c | 75 ± 3 de | 677 ± 16 b | 10 ± 1 e | 229 ± 5 e | 818 ± 9 a | 13 ± 1 e |
| Others | |||||||||
| Ethanolamine | 83 ± 8 cd | 70 ± 1 cd | 56 ± 0 def | 88 ± 2 c | 157 ± 4 a | 63 ± 3 cde | 41 ± 6 ef | 122 ± 12 b | 29 ± 1 f |
| Glycerol | 265 ± 1 a | 148 ± 2 b | 146 ± 2 b | 54 ± 2 e | 69 ± 2 de | 22 ± 0.9 f | 88 ± 3 cd | 96 ± 13 c | 15 ± 1 f |
| Lumichrome | ND | 1 ± 0.07 bc | ND | ND | 2 ± 0.1 a | 2 ± 0.03 a | ND | 1 ± 0.05b | ND |
| Fatty acids | |||||||||
| Hexadecanoate | 10 ± 1 a | ND | 1 ± 0.1 bc | 10 ± 0.07 a | 1 ± 0.01 c | 8 ± 0.8 a | 1 ± 0.02 c | 7 ± 3 a | 7 ± 0.3 ab |
| Octadecanoate | 2 ± 1 a | ND | ND | 1 ± 0.7 a | ND | 2 ± 0.07 a | ND | 2 ± 1 a | 2 ± 0.01 a |
| Octadecadienoate | 3 ± 1 c | ND | ND | 7 ± 0.08 ab | 1 ± 0.01 d | 6 ± 0.7 ab | 1 ± 0.01 d | 7 ± 0.1 a | 5 ± 0.7 bc |
| Octadecatrienoat | 6 ± 0.5 ab | ND | 1 ± 0.03b | 7 ± 2 ab | 2 ± 2 b | 12 ± 0.8 a | 1 ± 1 b | 8 ± 3 ab | 7 ± 0.7 ab |
| Amino acids | |||||||||
| Alanine | 1210 ± 44 a | 888 ± 18 b | 965 ± 74 b | 521 ± 87 c | 665 ± 30 c | 516 ± 3 c | 695 ± 24 c | 665 ± 39 c | 655 ± 15 c |
| Valine | 3782 ± 232 b | 5324 ± 205 a | 3347 ± 48 b | 1124 ± 47 d | 1927 ± 81 c | 851 ± 72 d | 1183 ± 35 cd | 1601 ± 120 cd | 1307 ± 95 cd |
| Serine | 5200 ± 18 8b | 5401 ± 107 b | 9591 ± 1504 a | 2890 ± 173 c | 4495 ± 265 bc | 3031 ± 178 bc | 3194 ± 71 bc | 3704 ± 722 bc | 2893 ± 231 bc |
| Leucine | 2414 ± 70 a | 2113 ± 26 abc | 2180 ± 103 ab | 768 ± 12 abc | 1037 ± 6 cd | 659 ± 36 d | 973 ± 9 d | 1130 ± 19b cd | 1055 ± 39 cd |
| Threonine | 627 ± 19 ab | 737 ± 12 a | 590 ± 167 ab | 551 ± 8 b | 713 ± 27 ab | 439 ± 24 b | 707 ± 5 ab | 729 ± 40 a | 621 ± 24 ab |
| Isoleucine | 1199 ± 37 a | 1374 ± 24 a | 1290 ± 57 a | 566 ± 10 bc | 794 ± 13 b | 600 ± 29 bc | 601 ± 12 c | 751 ± 18 bc | 767 ± 31 b |
| Proline | 1026 ± 58 a | 198 ± 15 cd | 279 ± 22 cd | 639 ± 13 b | 177 ± 41 cd | 127 ± 18 d | 328 ± 35 c | 153 ± 49 cd | 179 ± 11 cd |
| Glycine | 782 ± 53 a | 921 ± 166 a | 637 ± 74 a | 311 ± 47 a | 614 ± 38 a | 544 ± 30 a | 440 ± 81 a | 369 ± 80 a | 510 ± 40 a |
| Homoserine | 327 ± 21 a | 252 ± 8 abc | 267 ± 28 abcd | 176 ± 2 cd | 238 ± 29 ab | 158 ± 14 d | 223 ± 1 abcd | 204 ± 4 bcd | 221 ± 13 abcd |
| Methionine | 270 ± 9 ab | 396 ± 4 a | 367 ± 19 a | 215 ± 21 b | 234 ± 47 b | 162 ± 5 b | 267 ± 5 ab | 197 ± 13 b | 270 ± 7 ab |
| Phenylalanine | 562 ± 16 a | 499 ± 4 ab | 429 ± 26 abc | 241 ± 1 bcd | 300 ± 2 cd | 193 ± 1 d | 281 ± 4 cd | 327 ± 5 bcd | 327 ± 18 bcd |
| Glutamate | 20294 ± 110 a | 7387 ± 243 cd | 14981 ± 789 ab | 15377 ± 295 bcd | 4623 ± 1194 d | 11007 ± 1358 bc | 14600 ± 865 abc | 12221 ± 1286 ab | 20197 ± 402 a |
| Glutamine | 2130 ± 152 d | 7629 ± 895 cd | 5743 ± 901 cd | 41281 ± 3025 a | 8196 ± 661 cd | 15172 ± 1118 b | 35770 ± 1484 a | 8322 ± 614 bc | 13122 ± 1049 bc |
| Tyrosine | 26612 ± 101 a | 2400 ± 197 b | 15255 ± 181 ab | 3395 ± 463 ab | 1318 ± 84 b | 5891 ± 573 ab | 7072 ± 539 ab | 7025 ± 224 ab | 9318 ± 943 ab |
| Tryptophan | 4109 ± 113 a | 2628 ± 87 b | 1047 ± 214 c | 765 ± 193 c | 1251 ± 27 c | 666 ± 70 c | 727 ± 78 c | 833 ± 124 c | 1086 ± 252 c |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kumar, K.; Amir, R. The Effect of a Host on the Primary Metabolic Profiling of Cuscuta Campestris’ Main Organs, Haustoria, Stem and Flower. Plants 2021, 10, 2098. https://doi.org/10.3390/plants10102098
Kumar K, Amir R. The Effect of a Host on the Primary Metabolic Profiling of Cuscuta Campestris’ Main Organs, Haustoria, Stem and Flower. Plants. 2021; 10(10):2098. https://doi.org/10.3390/plants10102098
Chicago/Turabian StyleKumar, Krishna, and Rachel Amir. 2021. "The Effect of a Host on the Primary Metabolic Profiling of Cuscuta Campestris’ Main Organs, Haustoria, Stem and Flower" Plants 10, no. 10: 2098. https://doi.org/10.3390/plants10102098
APA StyleKumar, K., & Amir, R. (2021). The Effect of a Host on the Primary Metabolic Profiling of Cuscuta Campestris’ Main Organs, Haustoria, Stem and Flower. Plants, 10(10), 2098. https://doi.org/10.3390/plants10102098







