Rapid Accumulation of Proline Enhances Salinity Tolerance in Australian Wild Rice Oryza australiensis Domin
Abstract
:1. Introduction
2. Results
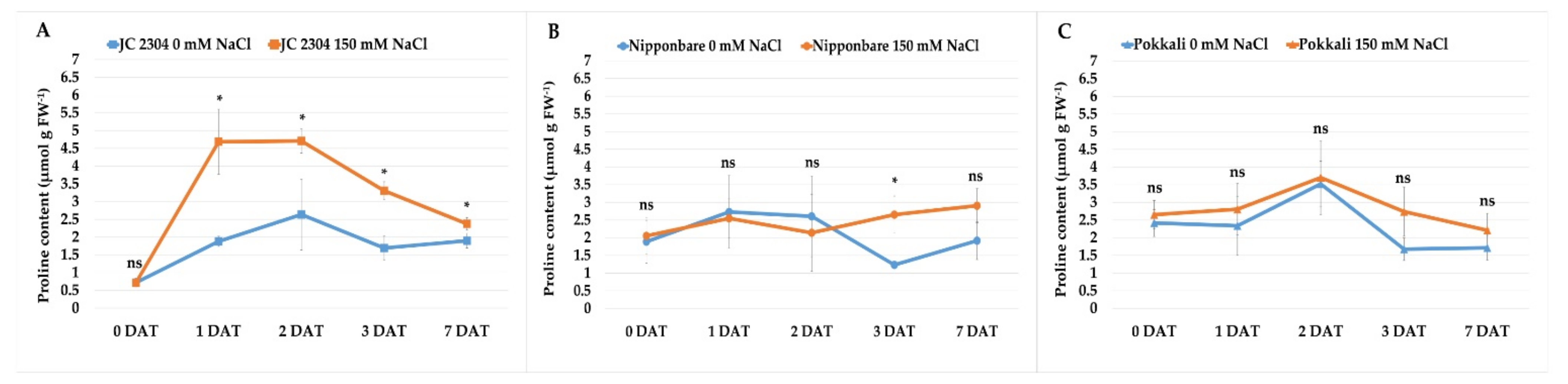
2.1. Wild Rice O. australiensis Rapidly Accumulated Free Proline under Salinity Stress
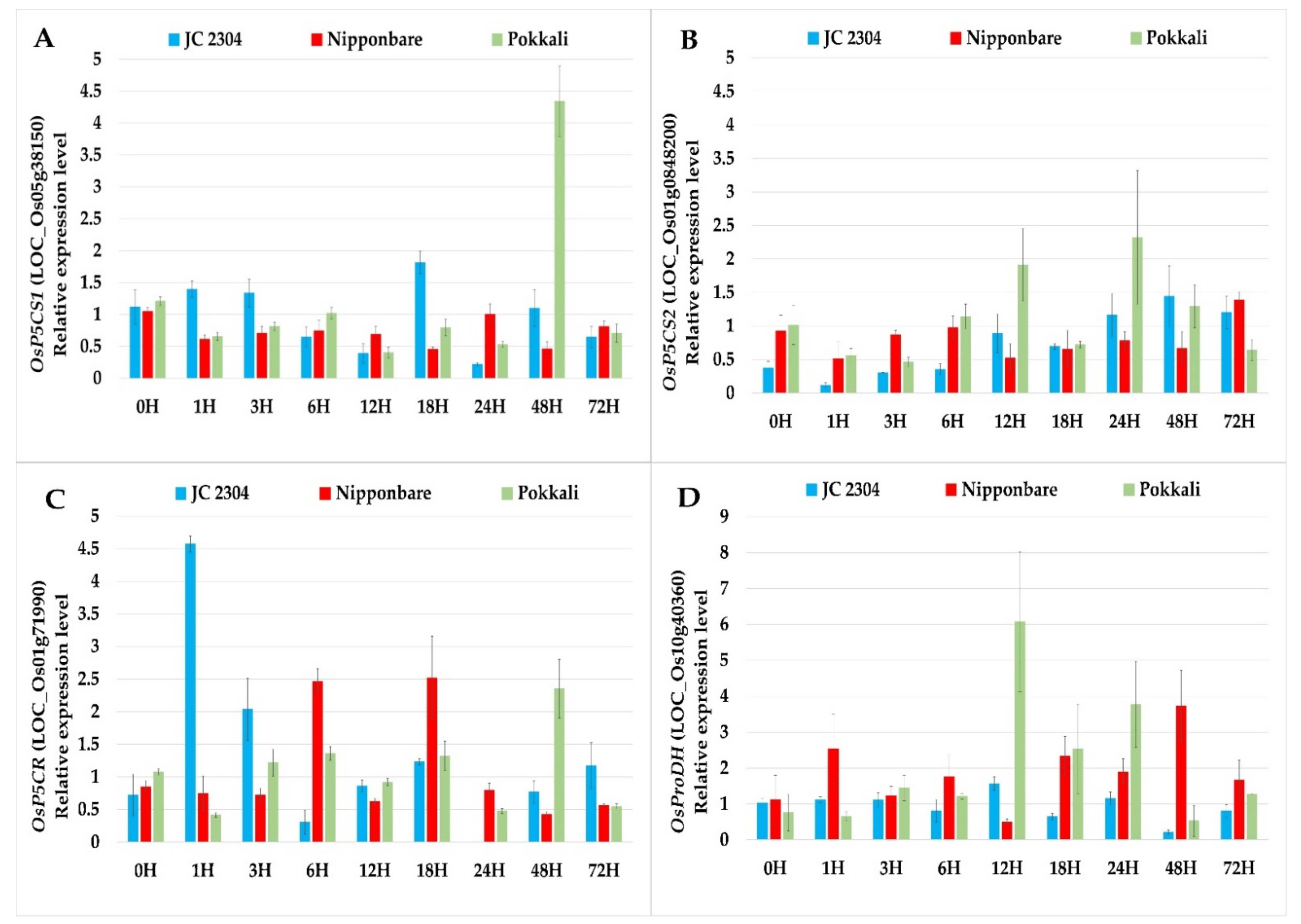
2.2. Wild Rice O. australiensis Activated Proline Synthesis and Depressed Proline Degradation Genes at the Early Stage of Exposure to Salinity Stress
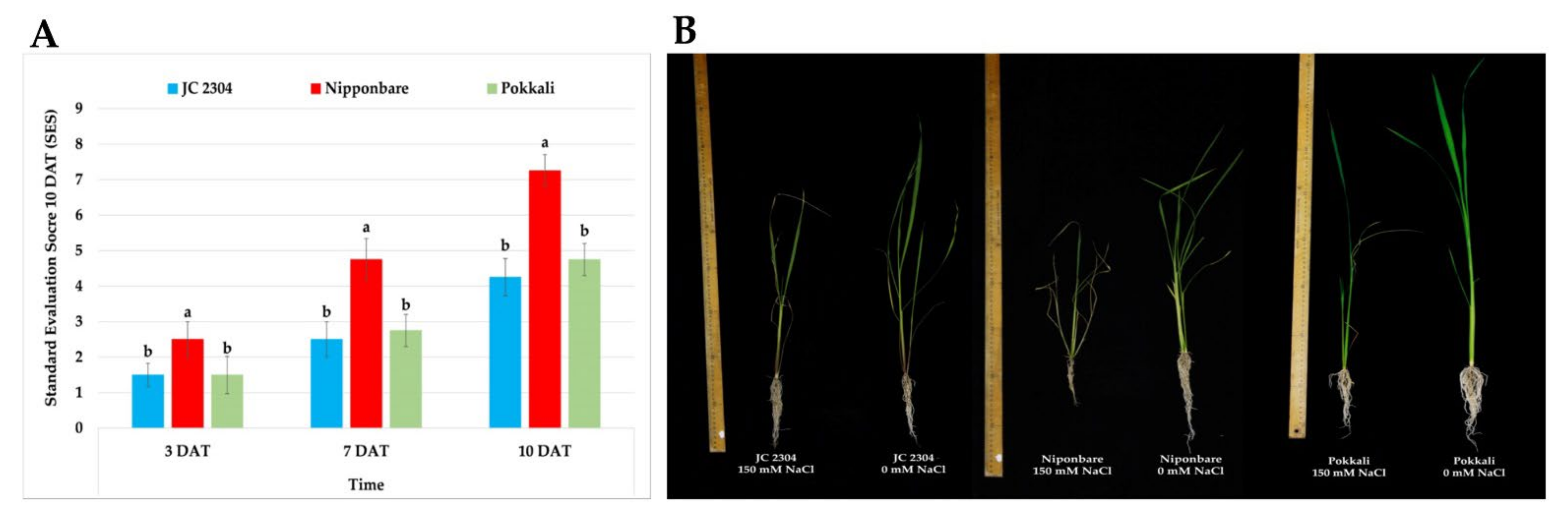
2.3. Wild Rice O. australiensis Showed a High Tolerant Level to Salinity Stress
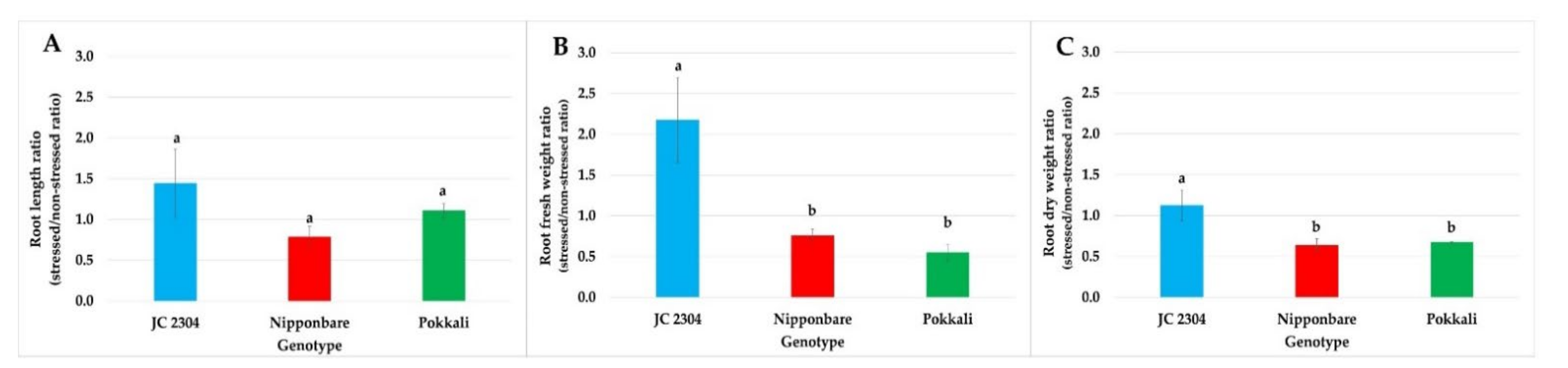
2.4. Wild Rice O. australiensis Exhibited Higher Root Growth and Biomass Than Cultivated Rice under Salinity Stress
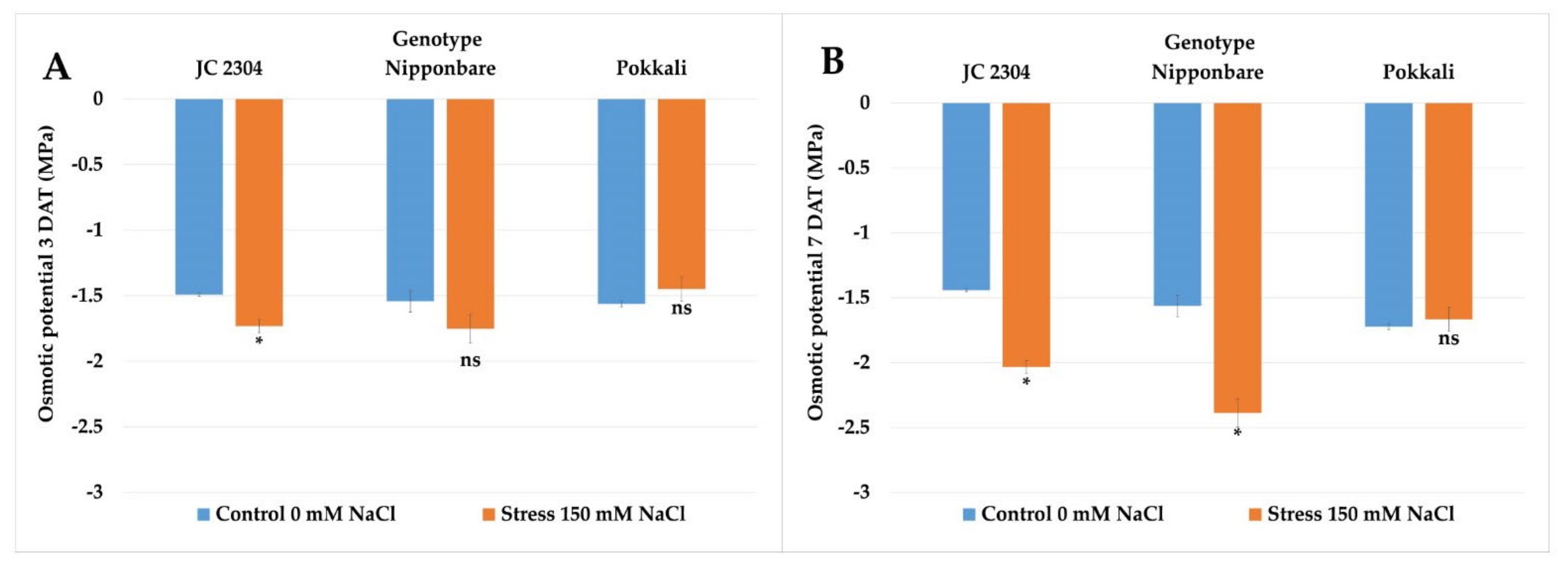
2.5. Wild Rice O. australiensis Lowered Osmotic Potential at the Early Stage of Exposure to Salinity Stress
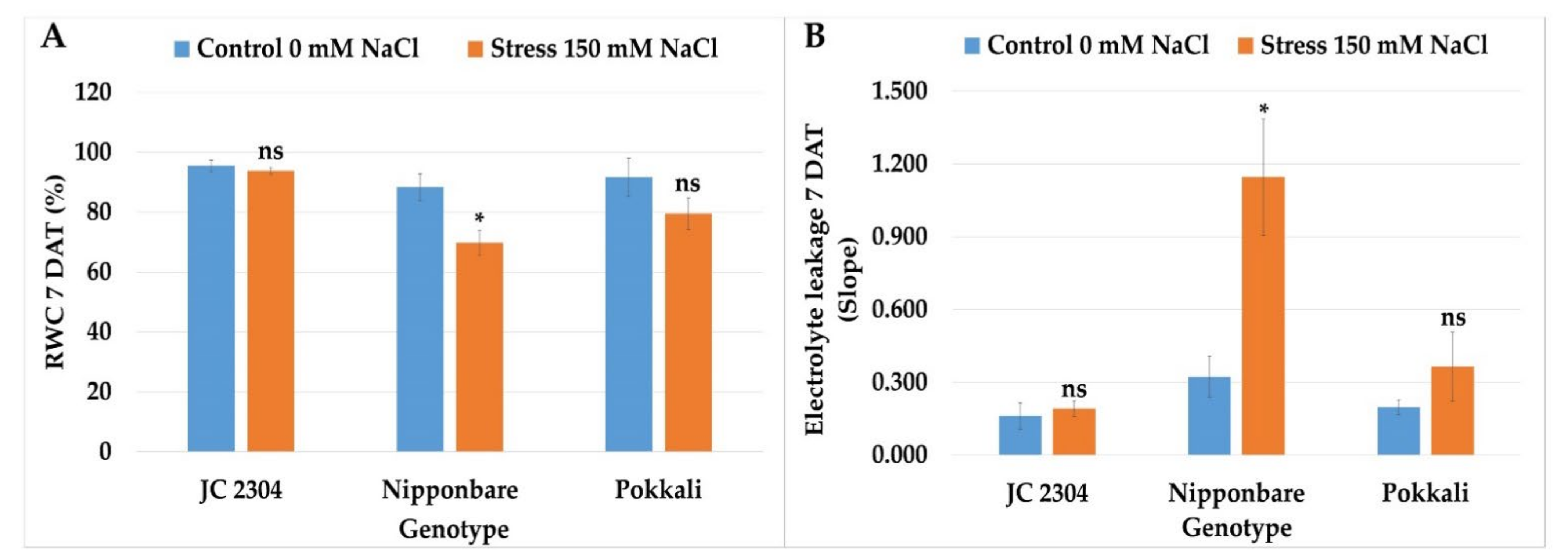
2.6. Wild Rice O. australiensis Maintained Higher Relative Water Content and Cell Membrane Integrity under Salinity Stress Compared to Salt-Sensitive Cultivated Rice
2.7. Wild Rice O. australiensis Accumulated Less Toxic Ion and Maintained a Lower Na+/K+ Ratio in Shoots under Salinity Stress Compared to Cultivated Rice
3. Discussion
4. Materials and Methods
4.1. Plant Material
4.2. Seedling Preparation and Salinity Stress Assay
4.3. Determination of Proline Content
4.4. Gene Expression Analysis
4.5. Determination of Relative Water Content
4.6. Determination of Osmotic Potential
4.7. Determination of Sodium and Potassium Content
4.8. Measurement of Electrolyte Leakage
4.9. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Al Otayk, S.M. Response of some wheat genotypes to different salinity levels of Irrigated water. South Cross J. 2020, 13, 37–45. [Google Scholar] [CrossRef]
- Qadir, M.; Quillérou, E.; Nangia, V.; Murtaza, G.; Singh, M.; Thomas, R.J.; Drechsel, P.; Noble, A.D. Economics of salt-induced land degradation and restoration. Nat. Resour. Forum 2014, 38, 282–295. [Google Scholar] [CrossRef]
- Nguyen, N. Global Climate Changes and Rice Food Security; International Rice Commission, FAO: Rome, Italy, 2002. [Google Scholar]
- Munns, R.; Tester, M. Mechanisms of salinity tolerance. Annu. Rev. Plant Biol. 2008, 59, 651–681. [Google Scholar] [CrossRef] [Green Version]
- Zhou, Y.; Yang, P.; Cui, F.; Zhang, F.; Luo, X.; Xie, J. Transcriptome analysis of salt stress responsiveness in the seedlings of dongxiang wild rice (Oryza rufipogon Griff.). PLoS ONE 2016, 11, e0146242. [Google Scholar] [CrossRef] [Green Version]
- Hoang, T.M.L.; Tran, T.N.; Nguyen, T.K.T.; Williams, B.; Wurm, P.; Bellairs, S.; Mundree, S. Improvement of salinity stress tolerance in rice: Challenges and opportunities. Agronomy 2016, 6, 54. [Google Scholar] [CrossRef]
- Zhang, A.; Liu, Y.; Wang, F.; Li, T.; Chen, Z.; Kong, D.; Bi, J.; Zhang, F.; Luo, X.; Wang, J.; et al. Enhanced rice salinity tolerance via CRISPR/Cas9-targeted mutagenesis of the OsRR22 gene. Mol. Breed. 2019, 39, 47. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ganeshan, P.; Jain, A.; Parmar, B.; Rao, A.; Sreenu, K.; Mishra, P.; Mesapogu, S.; Subrahmanyam, D.; Ram, T.; Sarla, N.; et al. Identification of salt tolerant rice lines among interspecific BILs developed by crossing “Oryza sativa O. rufipogon and O. sativa O. nivara”. South Cross J. 2016, 10, 220–228. [Google Scholar]
- Solis, C.A.; Yong, M.T.; Vinarao, R.; Jena, K.; Holford, P.; Shabala, L.; Zhou, M.; Shabala, S.; Chen, Z.-H. Back to the wild: On a quest for donors toward salinity tolerant rice. Front. Plant Sci. 2020, 11, 323–337. [Google Scholar] [CrossRef]
- Henry, R.J.; Rice, N.F.; Waters, D.; Kasem, S.; Ishikawa, R.; Hao, Y.; Dillon, S.L.; Crayn, D.; Wing, R.; Vaughan, D. Australian oryza: Utility and conservation. Rice 2009, 3, 235–241. [Google Scholar] [CrossRef] [Green Version]
- Scafaro, A.; Gallé, A.; Van Rie, J.; Carmo-Silva, E.; Salvucci, M.E.; Atwell, B.J. Heat tolerance in a wild Oryza species is attributed to maintenance of Rubisco activation by a thermally stable Rubisco activase ortholog. New Phytol. 2016, 211, 899–911. [Google Scholar] [CrossRef] [Green Version]
- Hu, J.; Xiao, C.; Cheng, M.-X.; Gao, G.-J.; Zhang, Q.-L.; He, Y.-Q. A new finely mapped Oryza australiensis-derived QTL in rice confers resistance to brown planthopper. Gene 2015, 561, 132–137. [Google Scholar] [CrossRef] [PubMed]
- Yichie, Y.; Brien, C.; Berger, B.; Roberts, T.H.; Atwell, B.J. Salinity tolerance in Australian wild Oryza species varies widely and matches that observed in O. sativa. Rice 2018, 11, 66. [Google Scholar] [CrossRef] [PubMed]
- Bose, J.; Rodrigo-Moreno, A.; Shabala, S. ROS homeostasis in halophytes in the context of salinity stress tolerance. J. Exp. Bot. 2014, 65, 1241–1257. [Google Scholar] [CrossRef]
- Munns, R.; Gilliham, M. Salinity tolerance of crops—what is the cost? New Phytol. 2015, 208, 668–673. [Google Scholar] [CrossRef] [Green Version]
- McCue, K.F.; Hanson, A.D. Drought and salt tolerance: Towards understanding and application. Trends Biotechnol. 1990, 8, 358–362. [Google Scholar] [CrossRef]
- Delauney, A.J.; Verma, D.P.S. Proline biosynthesis and osmoregulation in plants. Plant J. 1993, 4, 215–223. [Google Scholar] [CrossRef]
- Zulfiqar, F.; Akram, N.A.; Ashraf, M. Osmoprotection in plants under abiotic stresses: New insights into a classical phenomenon. Planta 2019, 251, 3. [Google Scholar] [CrossRef] [Green Version]
- Kishor, P.B.K.; Kumari, P.H.; Sunita, M.S.L.; Sreenivasulu, N. Role of proline in cell wall synthesis and plant development and its implications in plant ontogeny. Front. Plant Sci. 2015, 6, 544. [Google Scholar] [CrossRef] [Green Version]
- Kemble, A.R.; MacPherson, H.T. Liberation of amino acids in perennial rye grass during wilting. Biochem. J. 1954, 58, 46–49. [Google Scholar] [CrossRef] [PubMed]
- Stewart, G.R.; Lee, J.A. The role of proline accumulation in halophytes. Planta 1974, 120, 279–289. [Google Scholar] [CrossRef] [PubMed]
- Hare, P.; Cress, W. Metabolic implications of stress-induced proline accumulation in plants. Plant Growth Regul. 1997, 21, 79–102. [Google Scholar] [CrossRef]
- Kaur, G.; Asthir, B. Proline: A key player in plant abiotic stress tolerance. Biol. Plant 2015, 59, 609–619. [Google Scholar] [CrossRef]
- Meena, M.; Divyanshu, K.; Kumar, S.; Swapnil, P.; Zehra, A.; Shukla, V.; Yadav, M.; Upadhyay, R.S. Regulation of L-proline biosynthesis, signal transduction, transport, accumulation and its vital role in plants during variable environmental conditions. Heliyon 2019, 5, e02952. [Google Scholar] [CrossRef] [Green Version]
- Hasegawa, P.M.; Bressan, R.A.; Zhu, J.K.; Bohnert, H.J. Plant cellular and molecular responses to high salinity. Annu. Rev. Plant. Physiol. Plant. Mol. Biol. 2000, 51, 463–499. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Szabados, L.; Savouré, A. Proline: A multifunctional amino acid. Trends Plant Sci. 2010, 15, 89–97. [Google Scholar] [CrossRef]
- Miller, G.; Honig, A.; Stein, H.; Suzuki, N.; Mittler, R.; Zilberstein, A. Unraveling Δ1-pyrroline-5-carboxylate-proline cycle in plants by uncoupled expression of proline oxidation enzymes. J. Biol. Chem. 2009, 284, 26482–26492. [Google Scholar] [CrossRef] [Green Version]
- Servet, C.; Ghelis, T.; Richard, L.; Zilberstein, A.; Savoure, A. Proline dehydrogenase: A key enzyme in controlling cellular homeostasis. Front. Biosci. 2012, 17, 607–620. [Google Scholar] [CrossRef] [Green Version]
- Hong, Z.; Lakkineni, K.; Zhang, Z.; Verma, D.P.S. Removal of feedback inhibition of Δ1-pyrroline-5-carboxylate synthetase results in increased proline accumulation and protection of plants from osmotic stress. Plant Physiol. 2000, 122, 1129–1136. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ashraf, M.; Foolad, M. Roles of glycine betaine and proline in improving plant abiotic stress resistance. Environ. Exp. Bot. 2007, 59, 206–216. [Google Scholar] [CrossRef]
- Carillo, P.; Mastrolonardo, G.; Nacca, F.; Parisi, D.; Verlotta, A.; Fuggi, A. Nitrogen metabolism in durum wheat under salinity: Accumulation of proline and glycine betaine. Funct. Plant Biol. 2008, 35, 412–426. [Google Scholar] [CrossRef] [PubMed]
- Fichman, Y.; Gerdes, S.Y.; Kovács, H.; Szabados, L.; Zilberstein, A.; Csonka, L.N. Evolution of proline biosynthesis: Enzymology, bioinformatics, genetics, and transcriptional regulation. Biol. Rev. 2014, 90, 1065–1099. [Google Scholar] [CrossRef]
- Hu, C.A.; Delauney, A.J.; Verma, D.P. A bifunctional enzyme (delta 1-pyrroline-5-carboxylate synthetase) catalyzes the first two steps in proline biosynthesis in plants. Proc. Natl. Acad. Sci. USA 1992, 89, 9354–9358. [Google Scholar] [CrossRef] [Green Version]
- Fougère, F.; Le Rudulier, D.; Streeter, J.G. Effects of salt stress on amino acid, organic acid, and carbohydrate composition of roots, bacteroids, and cytosol of alfalfa (Medicago sativa L.). Plant Physiol. 1991, 96, 1228–1236. [Google Scholar] [CrossRef] [Green Version]
- Khanna-Chopra, R.; Semwal, V.; Lakra, N.; Pareek, A. Proline—A key regulator conferring plant tolerance to salinity and drought. In Plant Tolerance to Environmental Stress; CRC Press: Boca Raton, FL, USA, 2019; pp. 59–80. [Google Scholar]
- Hayat, S.; Hayat, Q.; Alyemeni, M.; Wani, D.A.; Pichtel, J.; Ahmad, A. Role of proline under changing environment: A review. Plant Signal. Behav. 2012, 7, 1456–1466. [Google Scholar] [CrossRef] [Green Version]
- Huang, Z.; Zhao, L.; Chen, D.; Liang, M.; Liu, Z.; Shao, H.; Long, X. Salt stress encourages proline accumulation by regulating proline biosynthesis and degradation in Jerusalem artichoke plantlets. PLoS ONE 2013, 8, e62085. [Google Scholar] [CrossRef] [PubMed]
- Gregorio, G.; Senadhira, D.; Mendoza, R. Screening rice for salinity tolerance. In IRRI Discussion Paper Series; International Rice Research Institute: Los Baños, Philippines, 1997; Volume 22. [Google Scholar]
- Dionisio-Sese, M.L.; Tobita, S. Antioxidant responses of rice seedlings to salinity stress. Plant Sci. 1998, 135, 1–9. [Google Scholar] [CrossRef]
- Cha-um, S.; Trakulyingcharoen, T.; Smitamana, P.; Kirdmanee, C. Salt Tolerance in two rice cultivars differing salt tolerant abilities in responses to iso-osmotic stress. Aust. J. Crop Sci. 2009, 3, 221–230. [Google Scholar]
- Hoang, T.M.L.; Williams, B.; Khanna, H.; Dale, J.; Mundree, S.G. Physiological basis of salt stress tolerance in rice expressing the antiapoptotic gene SfIAP. Funct. Plant Biol. 2014, 41, 1168–1177. [Google Scholar] [CrossRef] [Green Version]
- Qin, H.; Li, Y.; Huang, R. Advances and Challenges in the Breeding of Salt-Tolerant Rice. Int. J. Mol. Sci. 2020, 21, 8385. [Google Scholar] [CrossRef]
- Brar, D.S.; Khush, G.S. Transferring Genes from Wild Species into Rice; International Rice Research Institute: Metro Manila, Philippines, 2002; Volume 14, pp. 197–217. [Google Scholar]
- Menguer, P.K.; Sperotto, R.A.; Ricachenevsky, F.K. A walk on the wild side: Oryza species as source for rice abiotic stress tolerance. Genet. Mol. Biol. 2017, 40, 238–252. [Google Scholar] [CrossRef] [Green Version]
- Henry, R.J. Australian wild rice populations: A key resource for global food security. Front. Plant Sci. 2019, 10, 1354. [Google Scholar] [CrossRef]
- Khush, G.S.; Brar, D.S. Alien introgression in rice. Nucleus 2017, 60, 251–261. [Google Scholar] [CrossRef]
- Gupta, B.; Huang, B. Mechanism of salinity tolerance in plants: Physiological, biochemical, and molecular characterization. Int. J. Genom. 2014, 2014, 701596. [Google Scholar] [CrossRef] [PubMed]
- Reddy, I.N.B.L.; Kim, B.-K.; Yoon, I.-S.; Kim, K.-H.; Kwon, T.-R. Salt tolerance in rice: Focus on mechanisms and approaches. Rice Sci. 2017, 24, 123–144. [Google Scholar] [CrossRef]
- Yichie, Y.; Hasan, M.T.; Tobias, P.A.; Pascovici, D.; Goold, H.; Van Sluyter, S.C.; Roberts, T.H.; Atwell, B. Salt-treated roots of Oryza australiensis seedlings are enriched with proteins involved in energetics and transport. Proteomics 2019, 19, e1900175. [Google Scholar] [CrossRef] [PubMed]
- Kavi Kishor, P.B.; Sreenivasulu, N. Is proline accumulation per se correlated with stress tolerance or is proline homeostasis a more critical issue? Plant Cell Environ. 2014, 37, 300–311. [Google Scholar] [CrossRef]
- Misra, N.; Gupta, A.K. Effect of salt stress on proline metabolism in two high yielding genotypes of green gram. Plant Sci. 2005, 169, 331–339. [Google Scholar] [CrossRef]
- Poustini, K.; Siosemardeh, A.; Ranjbar, M. Proline accumulation as a response to salt stress in 30 wheat (Triticum aestivum L.) cultivars differing in salt tolerance. Genet. Resour. Crop. Evol. 2006, 54, 925–934. [Google Scholar] [CrossRef]
- Surabhi, G.-K.; Reddy, A.M.; Kumari, G.J.; Sudhakar, C. Modulations in key enzymes of nitrogen metabolism in two high yielding genotypes of mulberry (Morus alba L.) with differential sensitivity to salt stress. Environ. Exp. Bot. 2008, 64, 171–179. [Google Scholar] [CrossRef]
- Xue, X.; Liu, A.; Hua, X. Proline accumulation and transcriptional regulation of proline biosynthesis and degradation in Brassica napus. BMB Rep. 2009, 42, 28–34. [Google Scholar] [CrossRef] [Green Version]
- Roy, D.; Bhunia, A.; Basu, N.; Banerjee, S.K. Effect of nacl- salinity on metabolism of proline in salt- sensitive and salt- resistant cultivars of rice. Biol. Plant. 1992, 34, 159–162. [Google Scholar] [CrossRef]
- Roychoudhury, A.; Basu, S.; Sarkar, S.N.; Sengupta, D.N. Comparative physiological and molecular responses of a common aromatic indica rice cultivar to high salinity with non-aromatic indica rice cultivars. Plant Cell Rep. 2008, 27, 1395–1410. [Google Scholar] [CrossRef] [PubMed]
- Cha-Um, S.; Singh, H.P.; Samphumphuang, T.; Kirdmanee, C. Calcium-alleviated salt tolerance in indica rice (Oryza sativa L. spp. indica): Physiological and morphological changes. Aust. J. Crop Sci. 2012, 6, 176–182. [Google Scholar]
- Abdelaziz, M.N.; Xuan, T.D.; Mekawy, A.; Wang, H.; Khanh, T.D. Relationship of salinity tolerance to Na+ exclusion, proline accumulation, and antioxidant enzyme activity in rice seedlings. Agriculture 2018, 8, 166. [Google Scholar] [CrossRef] [Green Version]
- Hildebrandt, T.M. Synthesis versus degradation: Directions of amino acid metabolism during Arabidopsis abiotic stress response. Plant Mol. Biol. 2018, 98, 121–135. [Google Scholar] [CrossRef]
- Negrão, S.; Schmöckel, S.M.; Tester, M. Evaluating physiological responses of plants to salinity stress. Ann. Bot. 2017, 119, 1–11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, N.L.; Lah, W.A.C.; Kadir, N.A.; Mustaqim, M.; Rahmat, Z.; Ahmad, A.; Lam, S.D.; Ismail, M.R. Susceptibility and tolerance of rice crop to salt threat: Physiological and metabolic inspections. PLoS ONE 2018, 13, e0192732. [Google Scholar] [CrossRef]
- Thomson, M.J.; de Ocampo, M.; Egdane, J.; Rahman, M.A.; Sajise, A.G.; Adorada, D.L.; Tumimbang-Raiz, E.; Blumwald, E.; Seraj, Z.I.; Singh, R.K.; et al. Characterizing the saltol quantitative trait locus for salinity tolerance in rice. Rice 2010, 3, 148–160. [Google Scholar] [CrossRef] [Green Version]
- Miller, G.; Suzuki, N.; Ciftci-Yilmaz, S.; Mittler, R. Reactive oxygen species homeostasis and signalling during drought and salinity stresses. Plant Cell Environ. 2010, 33, 453–467. [Google Scholar] [CrossRef]
- Mansour, M.M.F. Plasma membrane permeability as an indicator of salt tolerance in plants. Biol. Plant 2013, 57, 1–10. [Google Scholar] [CrossRef]
- Ueda, A.; Yahagi, H.; Fujikawa, Y.; Nagaoka, T.; Esaka, M.; Calcaño, M.; González, M.M.; Martich, J.D.H.; Saneoka, H. Comparative physiological analysis of salinity tolerance in rice. Soil Sci. Plant Nutr. 2013, 59, 896–903. [Google Scholar] [CrossRef]
- Mansour, M. Nitrogen containing compounds and adaptation of plants to salinity stress. Biol. Plant. 2000, 43, 491–500. [Google Scholar] [CrossRef]
- Salama, K.H.A.; Mansour, M.M.F.; Ali, F.Z.M.; Abou-hadid, A.F. NaCl-induced changes in plasma membrane lipids and proteins of Zea mays L. cultivars differing in their response to salinity. Acta Physiol. Plant. 2007, 29, 351. [Google Scholar] [CrossRef]
- Alvarez-Pizarro, J.C.; Gomes-Filho, E.; de Lacerda, C.F.; Alencar, N.L.M.; Prisco, J.T. Salt-induced changes on H+-ATPase activity, sterol and phospholipid content and lipid peroxidation of root plasma membrane from dwarf-cashew (Anacardium occidentale L.) seedlings. Plant Growth Regul. 2009, 59, 125–135. [Google Scholar] [CrossRef]
- Serrano, R.; Mulet, J.; Rios, G.; Marquez, J.; de Larrinoa, I.; Leube, M.; Mendizabal, I.; Pascual-Ahuir, A.; Proft, M.; Ros, R.; et al. A Glimpse of the Mechanisms of Ion Homeostasis during Salt Stress. J. Exp. Bot. 1999, 50. [Google Scholar] [CrossRef]
- Garcia, A.B.; Engler, J.; Iyer, S.; Gerats, T.; Van Montagu, M.; Caplan, A.B. Effects of osmoprotectants upon NaCl stress in rice. Plant Physiol. 1997, 115, 159–169. [Google Scholar] [CrossRef] [Green Version]
- Zhu, J.-K. Regulation of ion homeostasis under salt stress. Curr. Opin. Plant Biol. 2003, 6, 441–445. [Google Scholar] [CrossRef]
- Lin, H.X.; Zhu, M.Z.; Yano, M.; Gao, J.P.; Liang, Z.W.; Su, W.A.; Hu, X.H.; Ren, Z.; Chao, D.Y. QTLs for Na+ and K+ uptake of the shoots and roots controlling rice salt tolerance. Theor. Appl. Genet. 2004, 108, 253–260. [Google Scholar] [CrossRef]
- Platten, J.D.; Egdane, J.; Ismail, A.M. Salinity tolerance, Na+ exclusion and allele mining of HKT1;5 in Oryza sativa and O. glaberrima: Many sources, many genes, one mechanism? BMC Plant Biol. 2013, 13, 32. [Google Scholar] [CrossRef] [Green Version]
- Ferdose, J.; Kawasaki, M.; Taniguchi, M.; Miyake, H. Differential sensitivity of rice cultivars to salinity and its relation to ion accumulation and root tip structure. Plant Prod. Sci. 2009, 12, 453–461. [Google Scholar] [CrossRef]
- Zhang, Y.; Fang, J.; Wu, X.; Dong, L. Na+/K+ balance and transport regulatory mechanisms in weedy and cultivated rice (Oryza sativa L.) under salt stress. BMC Plant Biol. 2018, 18, 375. [Google Scholar] [CrossRef]
- Theerawitaya, C.; Tisarum, R.; Samphumphuang, T.; Takabe, T.; Cha-Um, S. Expression levels of the Na+/K+ transporter OsHKT2;1 and vacuolar Na+/H+ exchanger OsNHX1, Na enrichment, maintaining the photosynthetic abilities and growth performances of indica rice seedlings under salt stress. Physiol. Mol. Biol. Plants 2020, 26, 513–523. [Google Scholar] [CrossRef] [PubMed]
- Neang, S.; Goto, I.; Skoulding, N.S.; Cartagena, J.A.; Kano-Nakata, M.; Yamauchi, A.; Mitsuya, S. Tissue-specific expression analysis of Na+ and Cl− transporter genes associated with salt removal ability in rice leaf sheath. BMC Plant Biol. 2020, 26, 513–523. [Google Scholar] [CrossRef] [PubMed]
- Cuin, T.A.; Zhou, M.; Parsons, D.; Shabala, S. Genetic behaviour of physiological traits conferring cytosolic K+/ Na+ homeostasis in wheat. Plant Biol. 2012, 14, 438–446. [Google Scholar] [CrossRef] [PubMed]
- Bates, L.S.; Waldren, R.P.; Teare, I.D. Rapid determination of free proline for water-stress studies. Plant Soil 1973, 39, 205–207. [Google Scholar] [CrossRef]
- Forlani, G.; Bertazzini, M.; Zarattini, M.; Funck, D. Functional characterization and expression analysis of rice δ1-pyrroline-5-carboxylate dehydrogenase provide new insight into the regulation of proline and arginine catabolism. Front. Plant Sci. 2015, 6, 591. [Google Scholar] [CrossRef] [Green Version]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nguyen, H.T.T.; Das Bhowmik, S.; Long, H.; Cheng, Y.; Mundree, S.; Hoang, L.T.M. Rapid Accumulation of Proline Enhances Salinity Tolerance in Australian Wild Rice Oryza australiensis Domin. Plants 2021, 10, 2044. https://doi.org/10.3390/plants10102044
Nguyen HTT, Das Bhowmik S, Long H, Cheng Y, Mundree S, Hoang LTM. Rapid Accumulation of Proline Enhances Salinity Tolerance in Australian Wild Rice Oryza australiensis Domin. Plants. 2021; 10(10):2044. https://doi.org/10.3390/plants10102044
Chicago/Turabian StyleNguyen, Ha Thi Thuy, Sudipta Das Bhowmik, Hao Long, Yen Cheng, Sagadevan Mundree, and Linh Thi My Hoang. 2021. "Rapid Accumulation of Proline Enhances Salinity Tolerance in Australian Wild Rice Oryza australiensis Domin" Plants 10, no. 10: 2044. https://doi.org/10.3390/plants10102044
APA StyleNguyen, H. T. T., Das Bhowmik, S., Long, H., Cheng, Y., Mundree, S., & Hoang, L. T. M. (2021). Rapid Accumulation of Proline Enhances Salinity Tolerance in Australian Wild Rice Oryza australiensis Domin. Plants, 10(10), 2044. https://doi.org/10.3390/plants10102044





