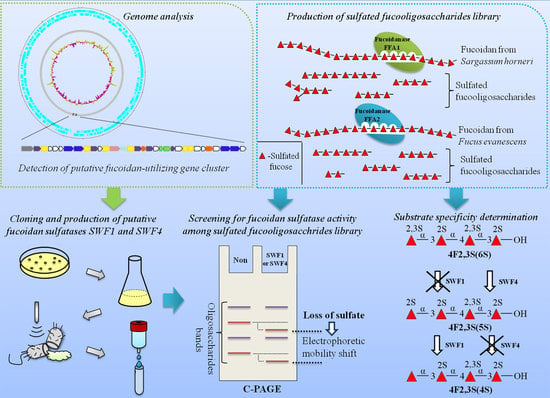
Fucoidan Sulfatases from Marine Bacterium Wenyingzhuangia fucanilytica CZ1127T
Abstract
1. Introduction
2. Materials and Methods
2.1. Reagents
2.2. General Methods
2.3. Identification and Amino Acid Sequences Analysis of Sulfatases
2.4. Homology Modeling of SWF1 and SWF4
2.5. Cloning of Fucoidan Sulfatases SWF1 and SWF4
2.6. Production of Recombinant Fucoidan Sulfatases SWF1 and SWF4
2.7. Purification of Sulfatases SWF1 and SWF4
2.8. Sodium Dodecyl Sulfate Polyacrylamide Gel Electrophoresis of Proteins
2.9. Fucoidan Sulfatase Activity Assay
2.10. Determination of Substrate Specificity of Sulfatases SWF1 and SWF4
2.11. Determination of the pH Optimum for Sulfatases SWF1 and SWF4 Activity
2.12. Determination of the Optimal Temperature for Sulfatases SWF1 and SWF4 Activity
2.13. Influence of Different Compounds on SWF1 and SWF4 Activity
2.14. Preparation of Reaction Products for Nuclear Magnetic Resonance Analysis
2.15. Nuclear Magnetic Resonance Spectroscopy
3. Results
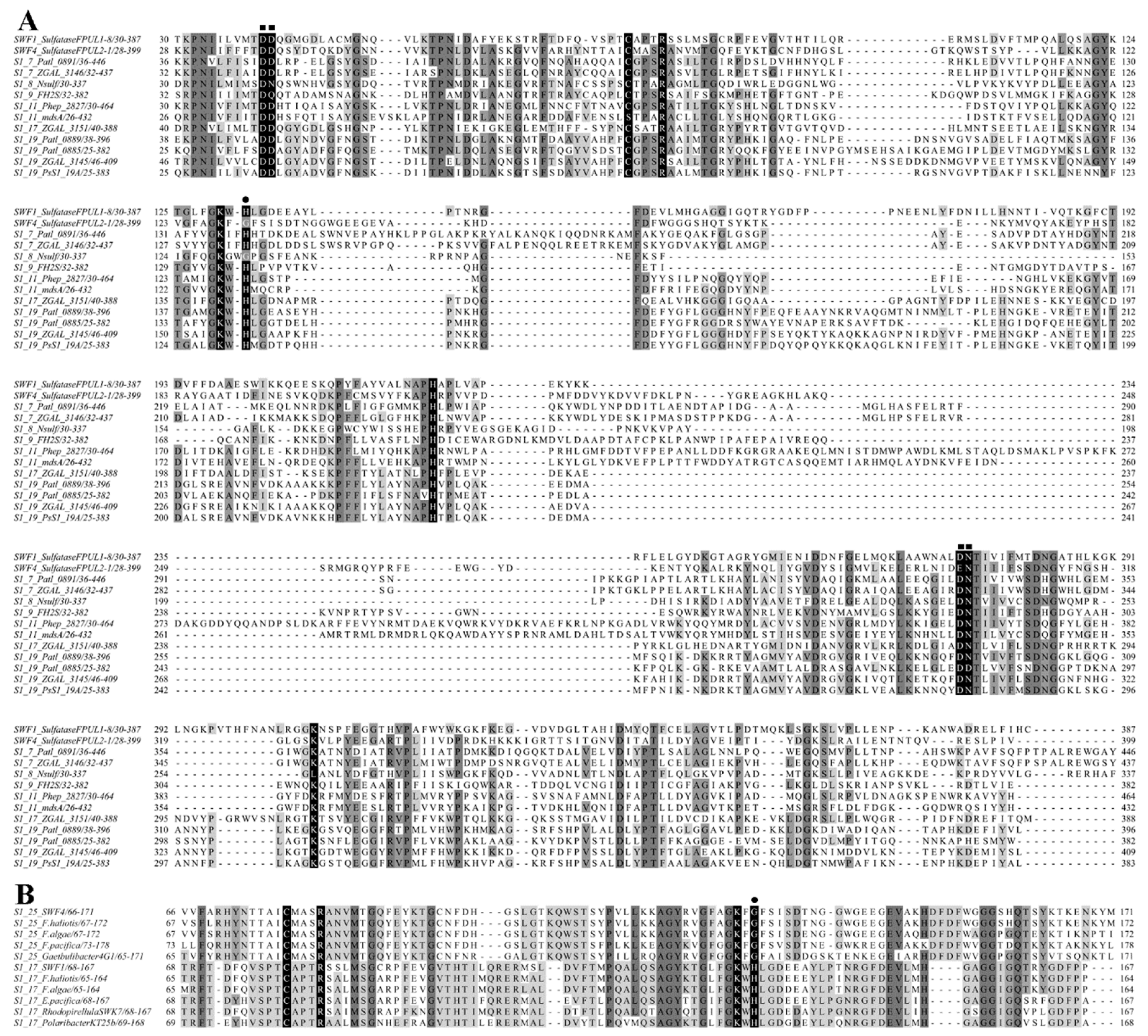
3.1. Amino Acid Sequence Analysis of SWF1 and SWF4
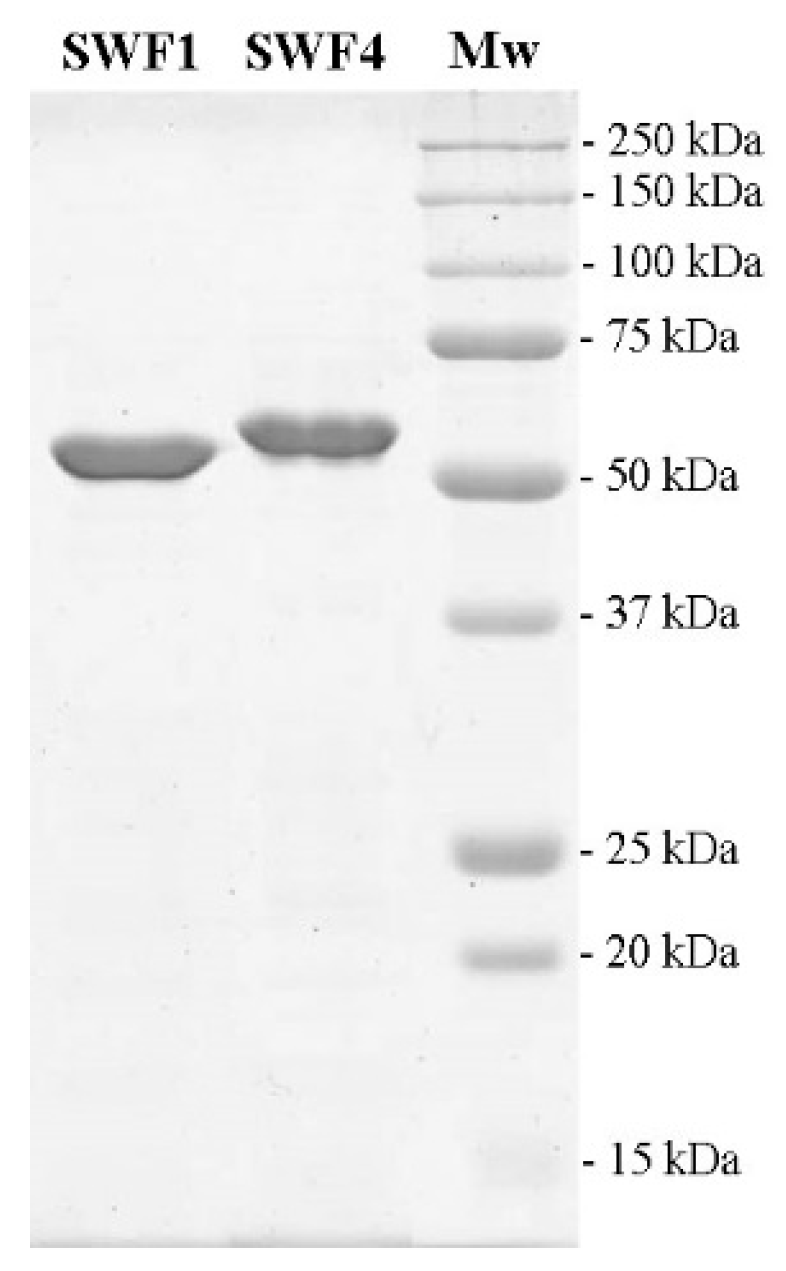
3.2. Expression and Purification of Fucoidan Sulfatases SWF1 and SWF4
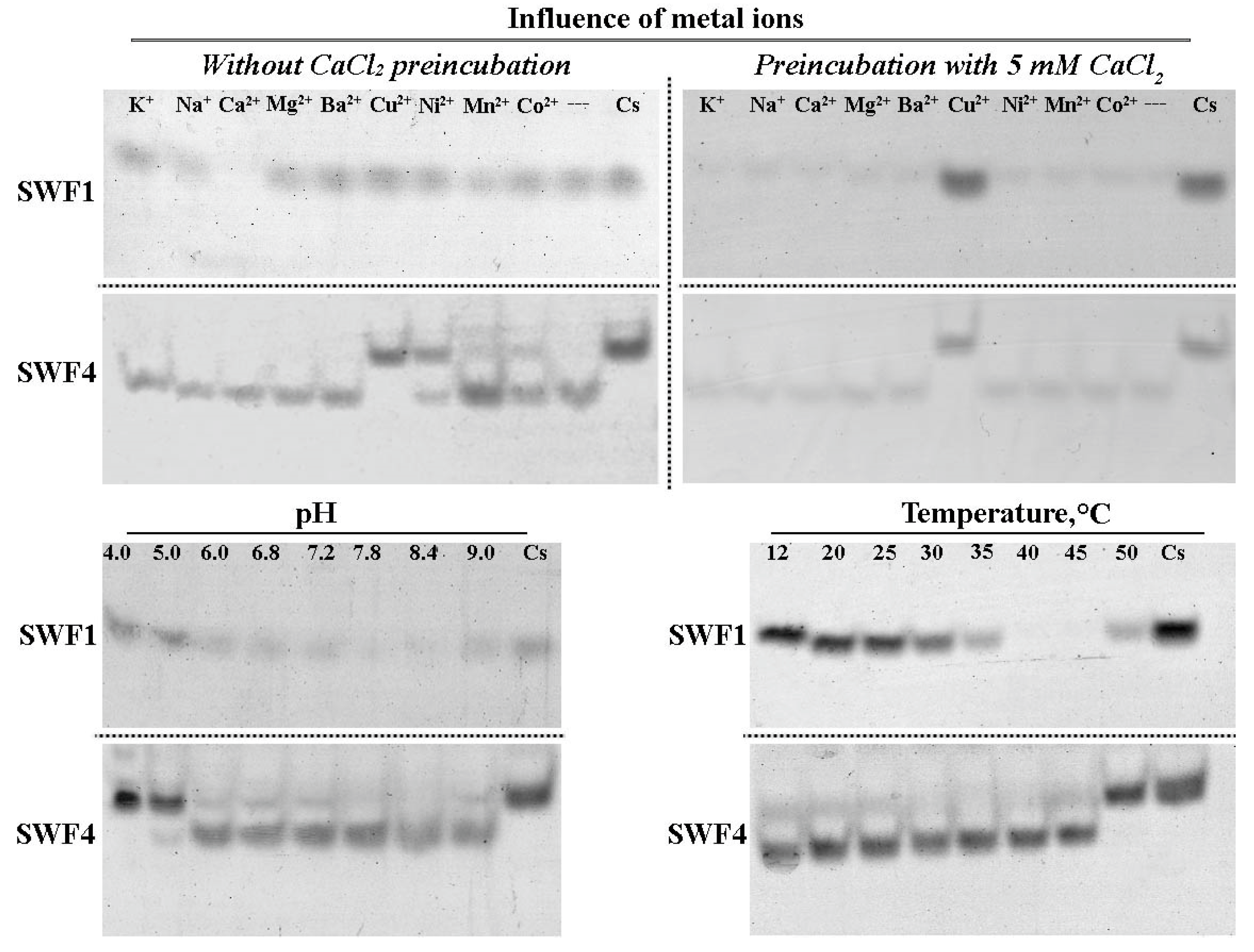
3.3. Optimal Conditions for Catalytic Activity of SWF1 and SWF4
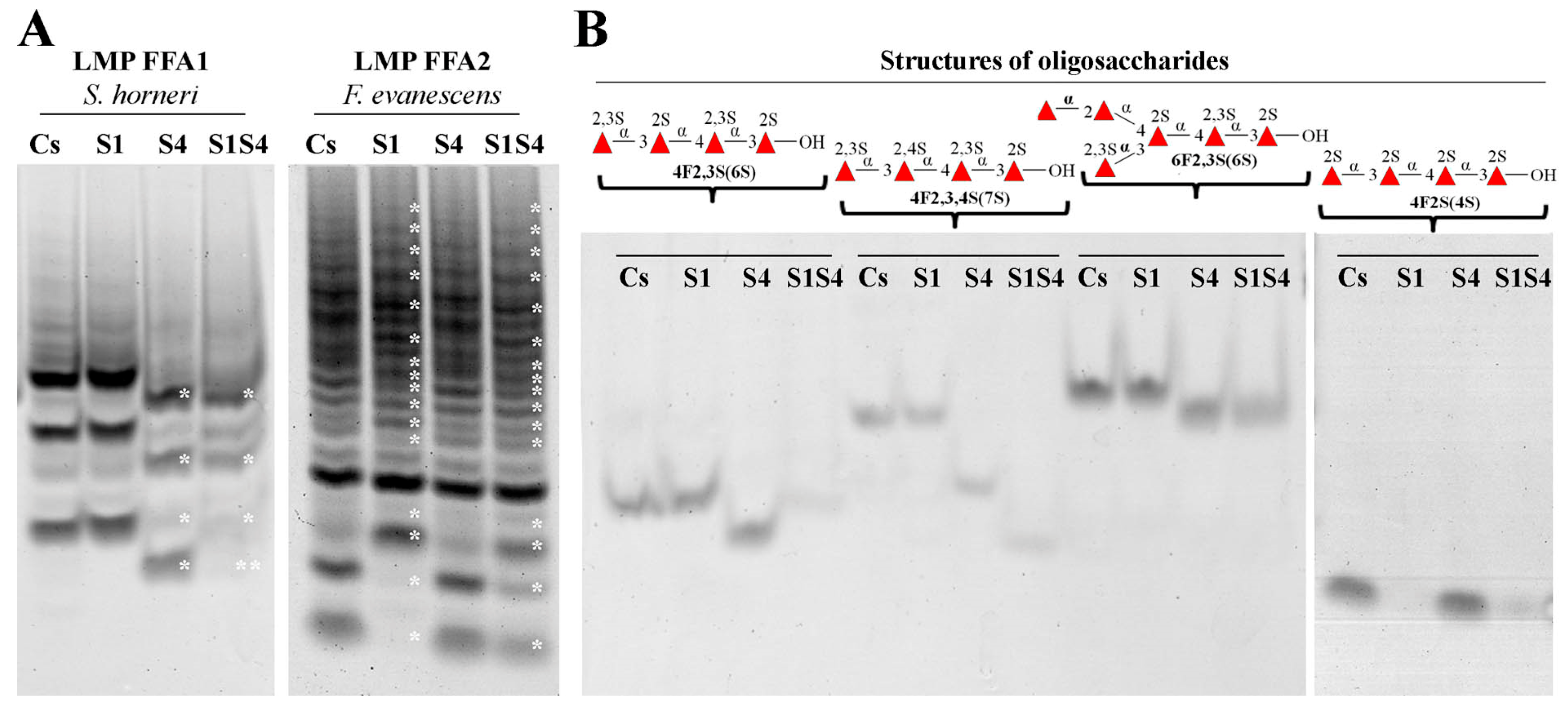
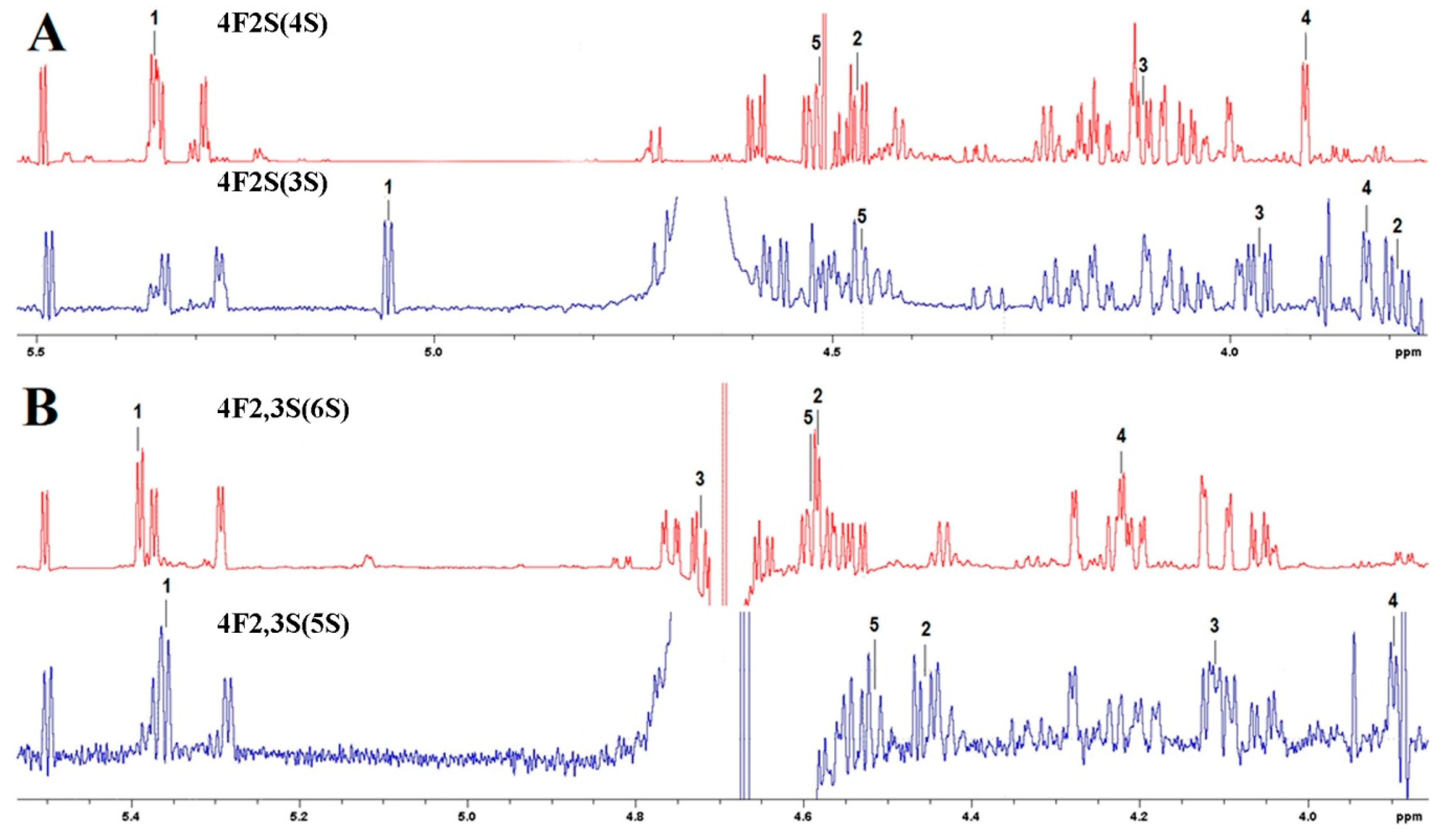
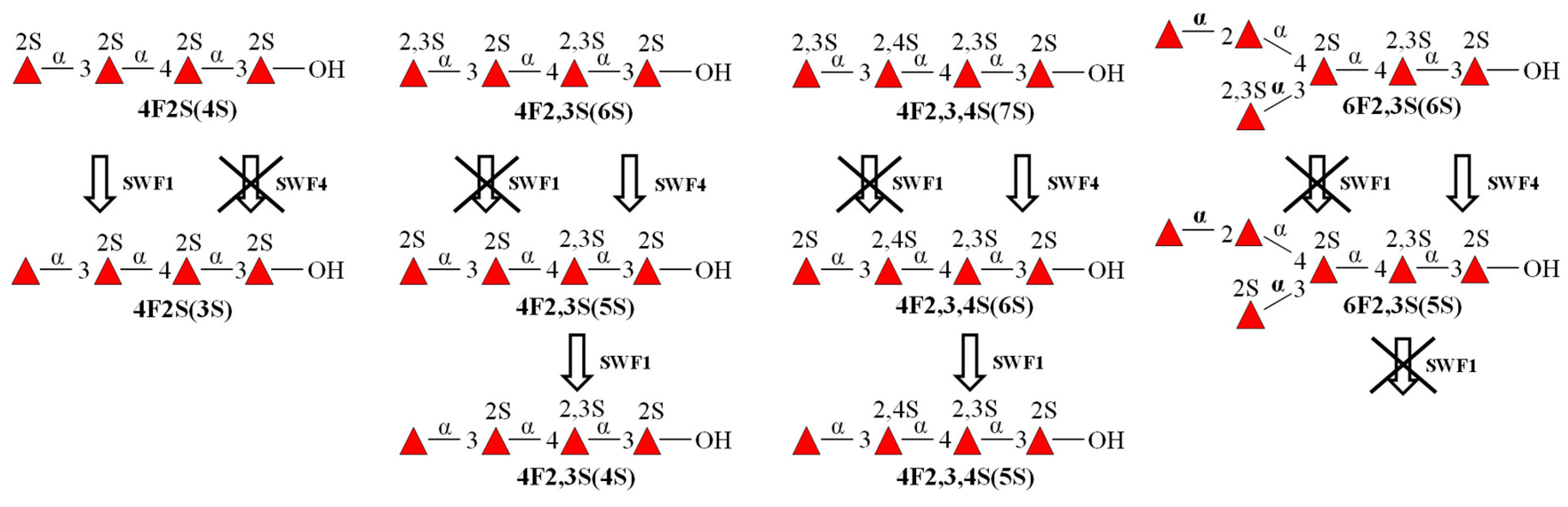
3.4. Substrate Specificity and Mode of Action of SWF1 and SWF4
4. Discussion
4.1. Identification of Putative Fucoidan Sulfatases in Marine Bacteria Wenyingzhuangia fucanilytica CZ1127T
4.2. Analysis of Amino Acid Sequences of Putative Fucoidan Sulfatases SWF1 and SWF4
4.3. Conservativeness of Sulfate-Binding S-Subsites of SWF1 and SWF4
4.4. Role of a Calcium Cation in Enzyme Activities of SWF1 and SWF4
4.5. Substrate Specificity of Sulfatases SWF1 and SWF4
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Chi, S.; Liu, T.; Wang, X.; Wang, R.; Wang, S.; Wang, G.; Shan, G.; Liu, C. Functional genomics analysis reveals the biosynthesis pathways of important cellular components (alginate and fucoidan) of Saccharina. Curr. Genet. 2018, 64, 259–273. [Google Scholar] [CrossRef] [PubMed]
- Deniaud-Bouët, E.; Hardouin, K.; Potin, P.; Kloareg, B.; Hervé, C. A review about brown algal cell walls and fucose-containing sulfated polysaccharides: Cell wall context, biomedical properties and key research challenges. Carbohydr. Polym. 2017, 175, 395–408. [Google Scholar] [CrossRef] [PubMed]
- Ale, M.T.; Mikkelsen, J.D.; Meyer, A.S. Important determinants for fucoidan bioactivity: A critical review of structure-function relations and extraction methods for fucose-containing sulfated polysaccharides from brown seaweeds. Mar. Drugs 2011, 9, 2106–2130. [Google Scholar] [CrossRef] [PubMed]
- Colin, S.; Deniaud, E.; Jam, M.; Descamps, V.; Chevolot, Y.; Kervarec, N.; Yvin, J.-C.; Barbeyron, T.; Michel, G.; Kloareg, B. Cloning and biochemical characterization of the fucanase FcnA: Definition of a novel glycoside hydrolase family specific for sulfated fucans. Glycobiology 2006, 16, 1021–1032. [Google Scholar] [CrossRef] [PubMed]
- Silchenko, A.S.; Ustyuzhanina, N.E.; Kusaykin, M.I.; Krylov, V.B.; Shashkov, A.S.; Dmitrenok, A.S.; Usoltseva, R.V.; Zueva, A.O.; Nifantiev, N.E.; Zvyagintseva, T.N. Expression and biochemical characterization and substrate specificity of the fucoidanase from Formosa algae. Glycobiology 2017, 27, 254–263. [Google Scholar] [CrossRef] [PubMed]
- Silchenko, A.S.; Rasin, A.B.; Kusaykin, M.I.; Kalinovsky, A.I.; Miansong, Z.; Changheng, L.; Malyarenko, O.; Zueva, A.O.; Zvyagintseva, T.N.; Ermakova, S.P. Structure, enzymatic transformation, anticancer activity of fucoidan and sulphated fucooligosaccharides from Sargassum horneri. Carbohydr. Polym. 2017, 175, 654–660. [Google Scholar] [CrossRef] [PubMed]
- Nagao, T.; Kumabe, A.; Komatsu, F.; Yagi, H.; Suzuki, H.; Ohshiro, T. Gene identification and characterization of fucoidan deacetylase for potential application to fucoidan degradation and diversification. J. Biosci. Bioeng. 2017, 124, 277–282. [Google Scholar] [CrossRef] [PubMed]
- Silchenko, A.S.; Kusaykin, M.I.; Zakharenko, A.M.; Menshova, R.V.; Khanh, H.H.N.; Dmitrenok, P.S.; Isakov, V.V.; Zvyagintseva, T.N. Endo-1,4-fucoidanase from Vietnamese marine mollusk Lambis sp. which producing sulphated fucooligosaccharides. J. Mol. Catal. B Enzym. 2014, 102, 154–160. [Google Scholar] [CrossRef]
- Silchenko, A.S.; Rasin, A.B.; Kusaykin, M.I.; Malyarenko, O.S.; Shevchenko, N.M.; Zueva, A.O.; Kalinovsky, A.I.; Zvyagintseva, T.N.; Ermakova, S.P. Modification of native fucoidan from Fucus evanescens by recombinant fucoidanase from marine bacteria Formosa algae. Carbohydr. Polym. 2018, 193, 189–195. [Google Scholar] [CrossRef] [PubMed]
- Kusaykin, M.I.; Silchenko, A.S.; Zakharenko, A.M.; Zvyagintseva, T.N. Fucoidanases. Glycobiology 2015, 26, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Barbeyron, T.; Brillet-Guéguen, L.; Carré, W.; Carrière, C.; Caron, C.; Czjzek, M.; Hoebeke, M.; Michel, G. Matching the diversity of sulfated biomolecules: Creation of a classification database for sulfatases reflecting their substrate specificity. PLoS ONE 2016, 11, e0164846. [Google Scholar] [CrossRef] [PubMed]
- Helbert, W. Marine Polysaccharide sulfatases. Front. Mar. Sci. 2017, 4, 6. [Google Scholar] [CrossRef]
- Furukawa, S.; Fujikawa, T.; Koga, D.; Ide, A. Production of fucoidan-degrading enzymes, fucoidanase, and fucoidan sulfatase by Vibrio sp. N-5. Nippon SUISAN GAKKAISHI 1992, 58, 1499–1503. [Google Scholar] [CrossRef]
- Furukawa, S.; Fujikawa, T. Growth and fucoidan sulfatase production in fucoidan-utilizing bacteria from sea sand. J. Agric. Chem. Soc. Jpn. 1984, 58, 1123–1126. [Google Scholar] [CrossRef]
- Shimanaka, K.; Ikai, K.; Kato, I.; Sakai, T.; Ishizuka, K. Structures of oligosaccharides derived from Cladosiphon okamuranus fucoidan by digestion with marine bacterial enzymes. Mar. Biotechnol. 2003, 5, 536–544. [Google Scholar] [CrossRef] [PubMed]
- Wegner, C.E.; Richter-Heitmann, T.; Klindworth, A.; Klockow, C.; Richter, M.; Achstetter, T.; Glöckner, F.O.; Harder, J. Expression of sulfatases in Rhodopirellula baltica and the diversity of sulfatases in the genus Rhodopirellula. Mar. Genom. 2013, 9, 51–61. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.; Xue, C.; Tang, Q.; Li, D.; Wu, X.; Wang, J. Isolation and characterization of a sea cucumber fucoidan-utilizing marine bacterium. Lett. Appl. Microbiol. 2010, 50, 301–307. [Google Scholar] [CrossRef] [PubMed]
- Thanassi, N.M.; Nakada, H.I. Enzymic degradation of fucoidan by enzymes from the hepatopancreas of abalone, Haliotus species. Arch. Biochem. Biophys. 1967, 118, 172–177. [Google Scholar] [CrossRef]
- Daniel, R.; Berteau, O.; Chevolot, L.; Varenne, A.; Gareil, P.; Goasdoue, N. Regioselective desulfation of sulfated L-fucopyranoside by a new sulfoesterase from the marine mollusk Pecten maximus: Application to the structural study of algal fucoidan (Ascophyllum nodosum). Eur. J. Biochem. 2001, 268, 5617–5626. [Google Scholar] [CrossRef] [PubMed]
- Lloyd, P.F.; Forrester, P.F. Desulphation of l-fucose monosulphates by an enzyme from Patella vulgata. Biochem. J. 1971, 124, 21P. [Google Scholar] [CrossRef] [PubMed]
- Zvyagintseva, T.N.; Shevchenko, N.M.; Popivnich, I.B.; Isakov, V.V.; Scobun, A.S.; Sundukova, E.V.; Elyakova, L.A. A new procedure for the separation of water-soluble polysaccharides from brown seaweeds. Carbohydr. Res. 1999, 322, 32–39. [Google Scholar] [CrossRef]
- Dubois, M.; Gilles, K.A.; Hamilton, J.K.; Rebers, P.A.; Smith, F. Colorimetric Method for determination of sugars and related Subtances. Anal. Chem. 1956, 28, 350–356. [Google Scholar] [CrossRef]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Quevillon, E.; Silventoinen, V.; Pillai, S.; Harte, N.; Mulder, N.; Apweiler, R.; Lopez, R. InterProScan: Protein domains identifier. Nucleic Acids Res. 2005, 33, W116–W120. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Yohe, T.; Huang, L.; Entwistle, S.; Wu, P.; Yang, Z.; Busk, P.K.; Xu, Y.; Yin, Y. dbCAN2: A meta server for automated carbohydrate-active enzyme annotation. Nucleic Acids Res. 2018. [Google Scholar] [CrossRef] [PubMed]
- Sievers, F.; Wilm, A.; Dineen, D.; Gibson, T.J.; Karplus, K.; Li, W.; Lopez, R.; McWilliam, H.; Remmert, M.; Söding, J.; et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 2011, 7, 539. [Google Scholar] [CrossRef] [PubMed]
- Waterhouse, A.M.; Procter, J.B.; Martin, D.M.A.; Clamp, M.; Barton, G.J. Jalview Version 2—A multiple sequence alignment editor and analysis workbench. Bioinformatics 2009, 25, 1189–1191. [Google Scholar] [CrossRef] [PubMed]
- Bendtsen, J.D.; Jensen, L.J.; Blom, N.; von Heijne, G.; Brunak, S. Feature-based prediction of non-classical and leaderless protein secretion. Protein Eng. Des. Sel. 2004, 17, 349–356. [Google Scholar] [CrossRef] [PubMed]
- Carver, T.; Harris, S.R.; Berriman, M.; Parkhill, J.; McQuillan, J.A. Artemis: An integrated platform for visualization and analysis of high-throughput sequence-based experimental data. Bioinformatics 2012, 28, 464–469. [Google Scholar] [CrossRef] [PubMed]
- Waterhouse, A.; Bertoni, M.; Bienert, S.; Studer, G.; Tauriello, G.; Gumienny, R.; Heer, F.T.; de Beer, T.A.P.; Rempfer, C.; Bordoli, L.; et al. SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Res. 2018, 46, W296–W303. [Google Scholar] [CrossRef] [PubMed]
- Krieger, E.; Joo, K.; Lee, J.; Lee, J.; Raman, S.; Thompson, J.; Tyka, M.; Baker, D.; Karplus, K. Improving physical realism, stereochemistry, and side-chain accuracy in homology modeling: Four approaches that performed well in CASP8. Proteins Struct. Funct. Bioinform. 2009, 77, 114–122. [Google Scholar] [CrossRef] [PubMed]
- Benkert, P.; Biasini, M.; Schwede, T. Toward the estimation of the absolute quality of individual protein structure models. Bioinformatics 2011, 27, 343–350. [Google Scholar] [CrossRef] [PubMed]
- Bond, S.R.; Naus, C.C. RF-Cloning.org: An online tool for the design of restriction-free cloning projects. Nucleic Acids Res. 2012, 40, W209–W213. [Google Scholar] [CrossRef] [PubMed]
- Restriction Free Cloning. Available online: http://rf-cloning.org (accessed on 14 September 2018).
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Chang, Y.; Dong, S.; Xue, C. Wenyingzhuangia fucanilytica sp. nov., a sulfated fucan utilizing bacterium isolated from shallow coastal seawater. Int. J. Syst. Evol. Microbiol. 2016, 66, 3270–3275. [Google Scholar] [CrossRef] [PubMed]
- Hanson, S.R.; Best, M.D.; Wong, C.-H. Sulfatases: Structure, mechanism, biological activity, inhibition, and synthetic utility. Angew. Chem. Int. Ed. 2004, 43, 5736–5763. [Google Scholar] [CrossRef] [PubMed]
- Hettle, A.G.; Vickers, C.; Robb, C.S.; Liu, F.; Withers, S.G.; Hehemann, J.-H.; Boraston, A.B. The molecular basis of polysaccharide sulfatase activity and a nomenclature for catalytic subsites in this class of enzyme. Structure 2018, 26, 747–758. [Google Scholar] [CrossRef] [PubMed]
- Benjdia, A.; Dehò, G.; Rabot, S.; Berteau, O. First evidences for a third sulfatase maturation system in prokaryotes from E. coli aslB and ydeM deletion mutants. FEBS Lett. 2007, 58, 1009–1014. [Google Scholar] [CrossRef] [PubMed]
- Rashid, A.; Mackie, W.; Colquhoun, I.J.; Lamba, D. Novel synthesis of monosulphated methyl α-d-galactopyranosides. Can. J. Chem. 1990, 68, 1122–1127. [Google Scholar] [CrossRef]
- Terrapon, N.; Lombard, V.; Gilbert, H.J.; Henrissat, B. Automatic prediction of polysaccharide utilization loci in Bacteroidetes species. Bioinformatics 2015, 31, 647–655. [Google Scholar] [CrossRef] [PubMed]
- Ficko-Blean, E.; Préchoux, A.; Thomas, F.; Rochat, T.; Larocque, R.; Zhu, Y.; Stam, M.; Génicot, S.; Jam, M.; Calteau, A.; et al. Carrageenan catabolism is encoded by a complex regulon in marine heterotrophic bacteria. Nat. Commun. 2017, 8, 1685. [Google Scholar] [CrossRef] [PubMed]
- Appel, M.J.; Bertozzi, C.R. Formylglycine, a post-translationally generated residue with unique catalytic capabilities and biotechnology applications. ACS Chem. Biol. 2015, 10, 72–84. [Google Scholar] [CrossRef] [PubMed]
- Myette, J.R.; Soundararajan, V.; Behr, J.; Shriver, Z.; Raman, R.; Sasisekharan, R. Heparin/heparan sulfate N-sulfamidase from Flavobacterium heparinum: Structural and biochemical investigation of catalytic nitrogen-sulfur bond cleavage. J. Biol. Chem. 2009, 284, 35189–35200. [Google Scholar] [CrossRef] [PubMed]
- Myette, J.R.; Soundararajan, V.; Shriver, Z.; Raman, R.; Sasisekharan, R. Heparin/heparan sulfate 6-O-sulfatase from Flavobacterium heparinum: Integrated structural and biochemical investigation of enzyme active site and substrate specificity. J. Biol. Chem. 2009, 284, 35177–35188. [Google Scholar] [CrossRef] [PubMed]
- Myette, J.R.; Shriver, Z.; Claycamp, C.; McLean, M.W.; Venkataraman, G.; Sasisekharan, R. The heparin/heparan sulfate 2-O-sulfatase from Flavobacterium heparinum: Molecular cloning, recombinant expression, and biochemical characterization. J. Biol. Chem. 2003, 278, 12157–12166. [Google Scholar] [CrossRef] [PubMed]
- Cartmell, A.; Lowe, E.C.; Baslé, A.; Firbank, S.J.; Ndeh, D.A.; Murray, H.; Terrapon, N.; Lombard, V.; Henrissat, B.; Turnbull, J.E.; et al. How members of the human gut microbiota overcome the sulfation problem posed by glycosaminoglycans. Proc. Natl. Acad. Sci. USA 2017, 114, 7037–7042. [Google Scholar] [CrossRef] [PubMed]
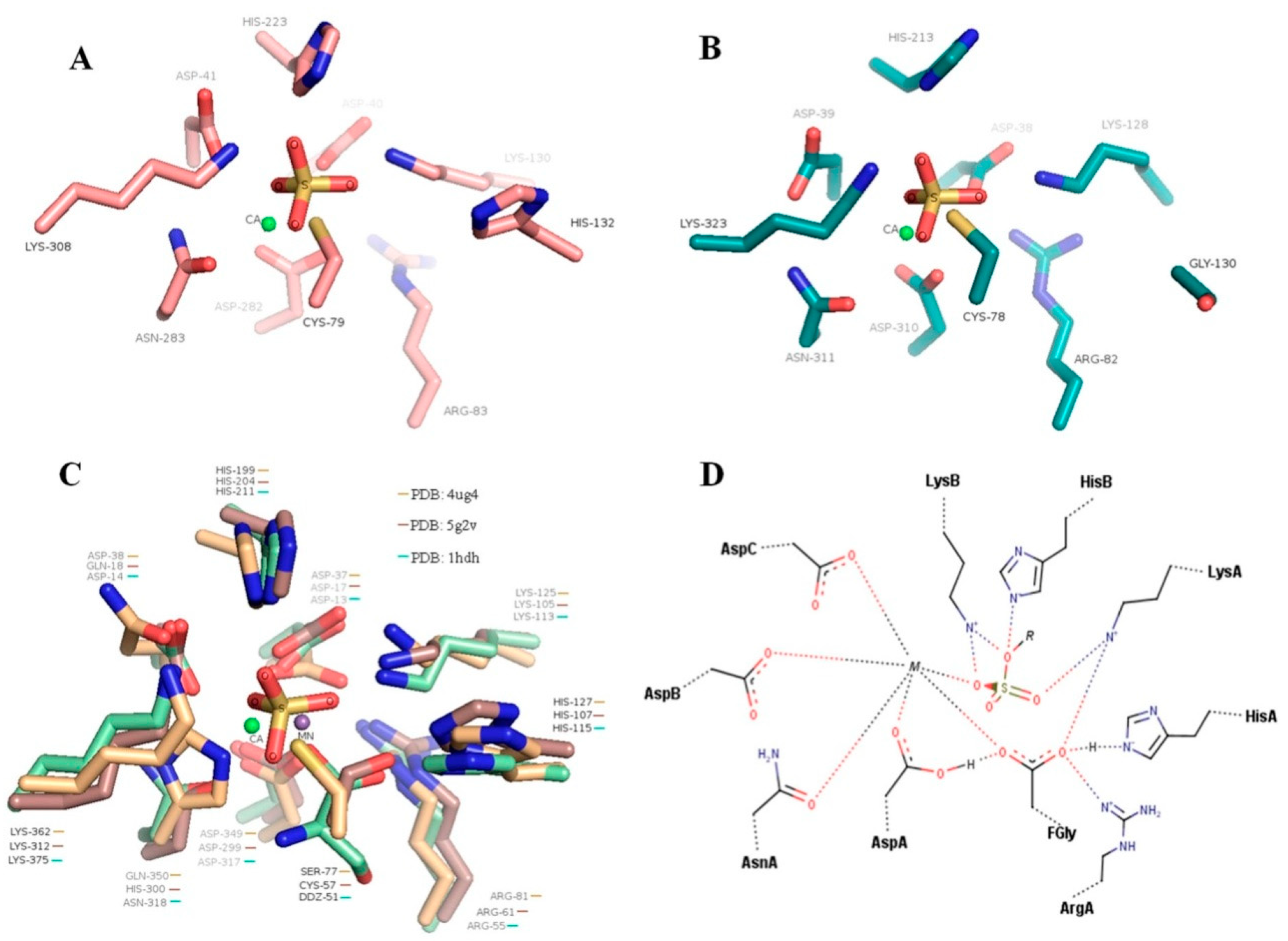
| Residue * | PDB: 5g2v | PDB: 4ug4 | PDB: 1hdh | SWF1 | SWF4 | Proposed Function |
|---|---|---|---|---|---|---|
| FGly/(modified residue) | Ser 77 | Cys 57 | Cys 51 | Cys 79 | Cys 78 | Catalytic nucleophile. Formation of sulfoenzyme intermediate complex. |
| M | Ca2+ | Mn2+ | Ca2 | Ca2 | Ca2 | Coordination and stabilization of sulfate group of substrate. |
| AsnA | Gln 350 | His 300 | Asn 318 | Asn 283 | Asn 311 | Coordination of metal ion. |
| AspA | Asp 349 | Asp 299 | Asp 317 | Asp 282 | Asp 310 | Coordination of metal ion and activation of FGly. |
| AspB | Asp 37 | Asp 17 | Asp 13 | Asp 40 | Asp 38 | Coordination of metal ion. |
| AspC | Asp 38 | Gln 18 | Asp14 | Asp 41 | Asp 39 | Coordination of metal ion. |
| LysA | Lys 125 | Lys 105 | Lys 113 | Lys 130 | Lys 128 | Sulfate binding and stabilisation of FGly. |
| LysB | Lys 362 | Lys 312 | Lys 375 | Lys 308 | Lys 323 | Sulfate binding and protonation of ester group of substrate. |
| ArgA | Arg 81 | Arg 61 | Arg 55 | Arg 83 | Arg 82 | Stabilization of FGly. |
| HisA | His 127 | His 107 | His 115 | His 132 | (Gly 130) | Deprotonation of FGly, elimination of sulfoenzyme intermediate complex. |
| HisB | His 199 | His 204 | His 211 | His 223 | His 213 | Sulfate binding and protonation of ester group of substrate. |
| Position | δC, Type | δH (J in Hz) | ROESY | HMBCα |
|---|---|---|---|---|
| 1 | 96.8, CH | 5.06, d (3.9) | 2, 9, 10 | 3, 5, 9 |
| 2 | 69.4, CH | 3.79, dd (9.9, 3.8) | 1 | 3 |
| 3 | 70.7, CH | 3.96, dd (10.3, 3.4) | 4, 5 | 1, 2, 4 |
| 4 | 73.3, CH | 3.83, d (3.4) | 3, 5, 6 | 3, 5 |
| 5 | 68.1, CH | 4.46, m | 3, 4, 6 | 1, 4, 6 |
| 6 | 16.5, CH3 | 1.21, d (6.6) | 4, 5 | 5 |
| 7 | 100.3, CH | 5.27, d (3.3) | 8, 16, 18 | 9, 11, 16 |
| 8 | 74.6, CH | 4.57, dd (10.3, 3.6) | 7 | 9 |
| 9 | 72.9, CH | 4.18, dd (10.8, 3.1) | 1, 10, 11 | 1, 7, 8, 10 |
| 10 | 70.1, CH | 4.11, d (2.2) | 1, 9, 11, 12 | 9, 11 |
| 11 | 68.6, CH | 4.44, m | 9, 10, 12 | 7, 10, 12 |
| 12 | 16.5, CH3 | 1.24, d (6.6) | 10, 11 | 11 |
| 13 | 95.4, CH | 5.34, d (3.9) | 14, 21, 22 | 15, 17, 21 |
| 14 | 76.6, CH | 4.49, dd (10.5, 3.8) | 13 | 15, 16 |
| 15 | 68.6, CH | 4.16, dd (10.9, 2.9) | 16, 17 | 13, 14, 16 |
| 16 | 83.7, CH | 3.99, d (2.6) | 7, 15, 17, 18 | 7, 14, 15, 16 |
| 17 | 68.9, CH | 4.52, m | 15, 16, 18 | 13, 17, 18 |
| 18 | 16.8, CH3 | 1.38, d (6.8) | 7, 16, 17 | 17 |
| 19 | 91.7, CH | 5.48, d (3.9) | 20 | 21, 23 |
| 20 | 74.7, CH | 4.51, dd (10.2, 3.2) | 19 | 21 |
| 21 | 73.9, CH | 4.05, dd (9.9, 3.1) | 13, 23 | 13, 19, 20 |
| 22 | 69.9, CH | 4.08, d (3.2) | 13, 23, 24 | 23 |
| 23 | 67.2, CH | 4.23, q (13.2, 6.6) | 21, 22, 24 | 19, 22, 24 |
| 24 | 16.7, CH3 | 1.24, d (6.6) | 22, 23 | 23 |
| Position | δH (J in Hz) | ROESY |
|---|---|---|
| 1 | 5.36, d (4.6) | 2, 9, 10 |
| 2 | 4.46, dd (9.5, 4.1) | 1 |
| 3 | 4.11, m | 5 |
| 4 | 3.90, d (4.2) | 5 |
| 5 | 4.51, m | 3, 4 |
| 6 | 1.23, d (6.0) | |
| 7 | 5.28, d (3.7) | 8, 16, 18 |
| 8 | 4.57, m | 7 |
| 9 | 4.19, dd (10.4, 3.2) | 11 |
| 10 | 4.11, m | 11 |
| 11 | 4.43, m | 9, 10 |
| 12 | 1.29, d (6.5) | |
| 13 | 5.37, d (4.4) | 21, 22 |
| 14 | 4.65, m | |
| 15 | 4.76, dd (11.2, 2.8) | 16, 17 |
| 16 | 4.28, d (2.9) | 7, 15, 17 |
| 17 | 4.55, m | 15, 16 |
| 18 | 1.40, d (6.8) | 7 |
| 19 | 5.50, d (3.9) | 20 |
| 20 | 4.54, m | 19 |
| 21 | 4.06, dd (10.2, 3.2) | 13, 23 |
| 22 | 4.09, d (4.8) | 13, 23 |
| 23 | 4.23, m | 21, 22 |
| 24 | 1.24, d (6.4) |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Silchenko, A.S.; Rasin, A.B.; Zueva, A.O.; Kusaykin, M.I.; Zvyagintseva, T.N.; Kalinovsky, A.I.; Kurilenko, V.V.; Ermakova, S.P. Fucoidan Sulfatases from Marine Bacterium Wenyingzhuangia fucanilytica CZ1127T. Biomolecules 2018, 8, 98. https://doi.org/10.3390/biom8040098
Silchenko AS, Rasin AB, Zueva AO, Kusaykin MI, Zvyagintseva TN, Kalinovsky AI, Kurilenko VV, Ermakova SP. Fucoidan Sulfatases from Marine Bacterium Wenyingzhuangia fucanilytica CZ1127T. Biomolecules. 2018; 8(4):98. https://doi.org/10.3390/biom8040098
Chicago/Turabian StyleSilchenko, Artem S., Anton B. Rasin, Anastasiya O. Zueva, Mikhail I. Kusaykin, Tatiana N. Zvyagintseva, Anatoly I. Kalinovsky, Valeriya V. Kurilenko, and Svetlana P. Ermakova. 2018. "Fucoidan Sulfatases from Marine Bacterium Wenyingzhuangia fucanilytica CZ1127T" Biomolecules 8, no. 4: 98. https://doi.org/10.3390/biom8040098
APA StyleSilchenko, A. S., Rasin, A. B., Zueva, A. O., Kusaykin, M. I., Zvyagintseva, T. N., Kalinovsky, A. I., Kurilenko, V. V., & Ermakova, S. P. (2018). Fucoidan Sulfatases from Marine Bacterium Wenyingzhuangia fucanilytica CZ1127T. Biomolecules, 8(4), 98. https://doi.org/10.3390/biom8040098