Local Attraction of Substrates and Co-Substrates Enhances Weak Acid and Base Transmembrane Transport
Abstract
:1. Introduction
2. Electrostatic Attraction and Neutralization of Substrate Ions by the Transport Protein
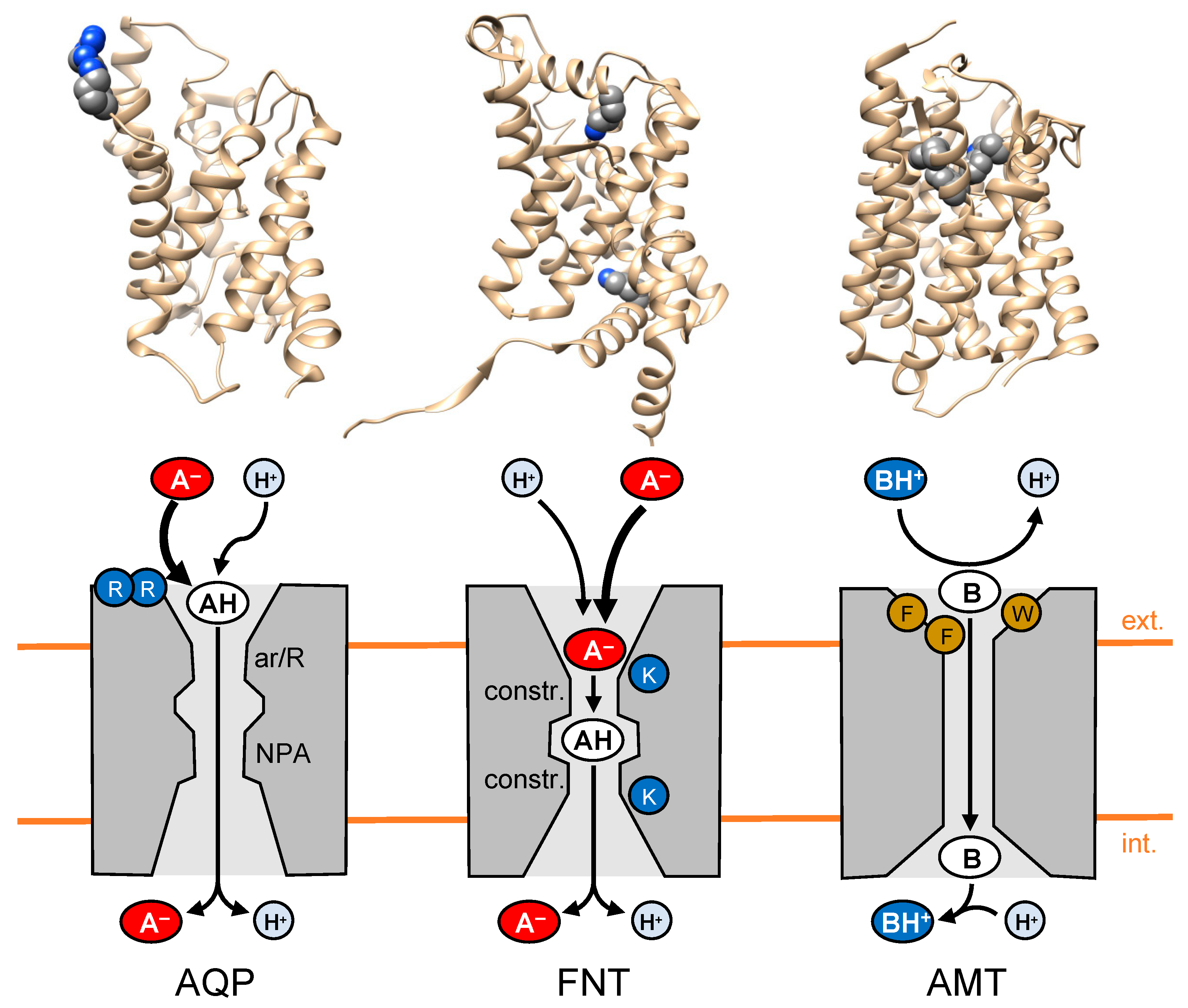
2.1. Substrate Attraction by Lactic Acid-Facilitating Aquaporins
2.2. The Next Step in Evolution: Channel-like Formate-Nitrite Transporters
2.3. Weak Base Transport: Opposite Prerequisites and Requirements
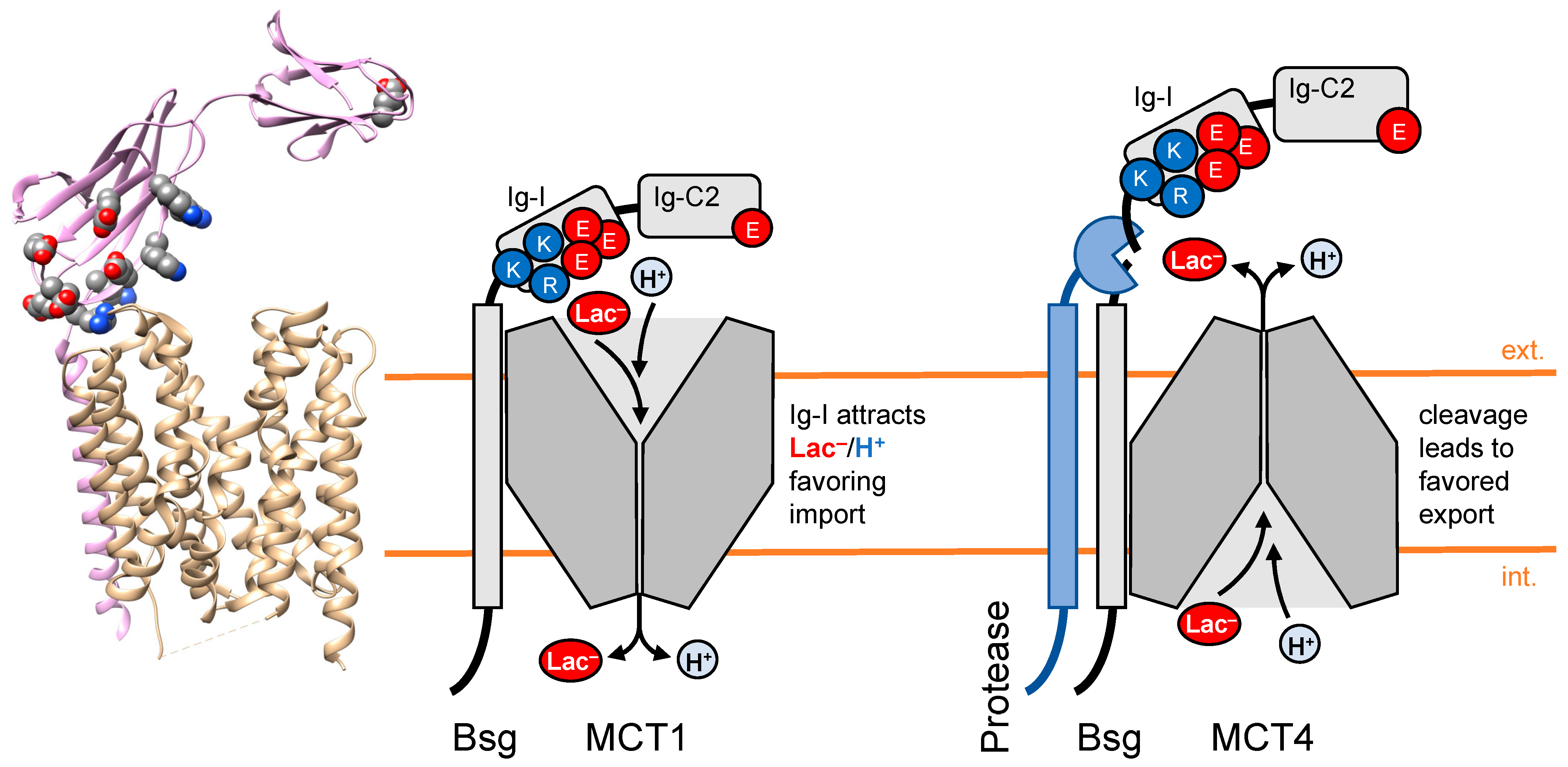
3. Chaperones of Transport Proteins Act as Local Attractors for Substrates
4. Carbonic Anhydrases Contribute Non-Catalytically to Proton-Driven Transport
4.1. Extracellular CAIV
4.2. Extracellular CAIX
4.3. Intracellular CAII
4.4. “Push and Pull Principle” of Fully CA-Decorated MCT
5. Transporter-Associated Enzymes Provide Substrates in Place, Enhancing Transport
5.1. CA Activity Increases Activity of Proton-Driven Lactate Transport by Locally Generating Protons
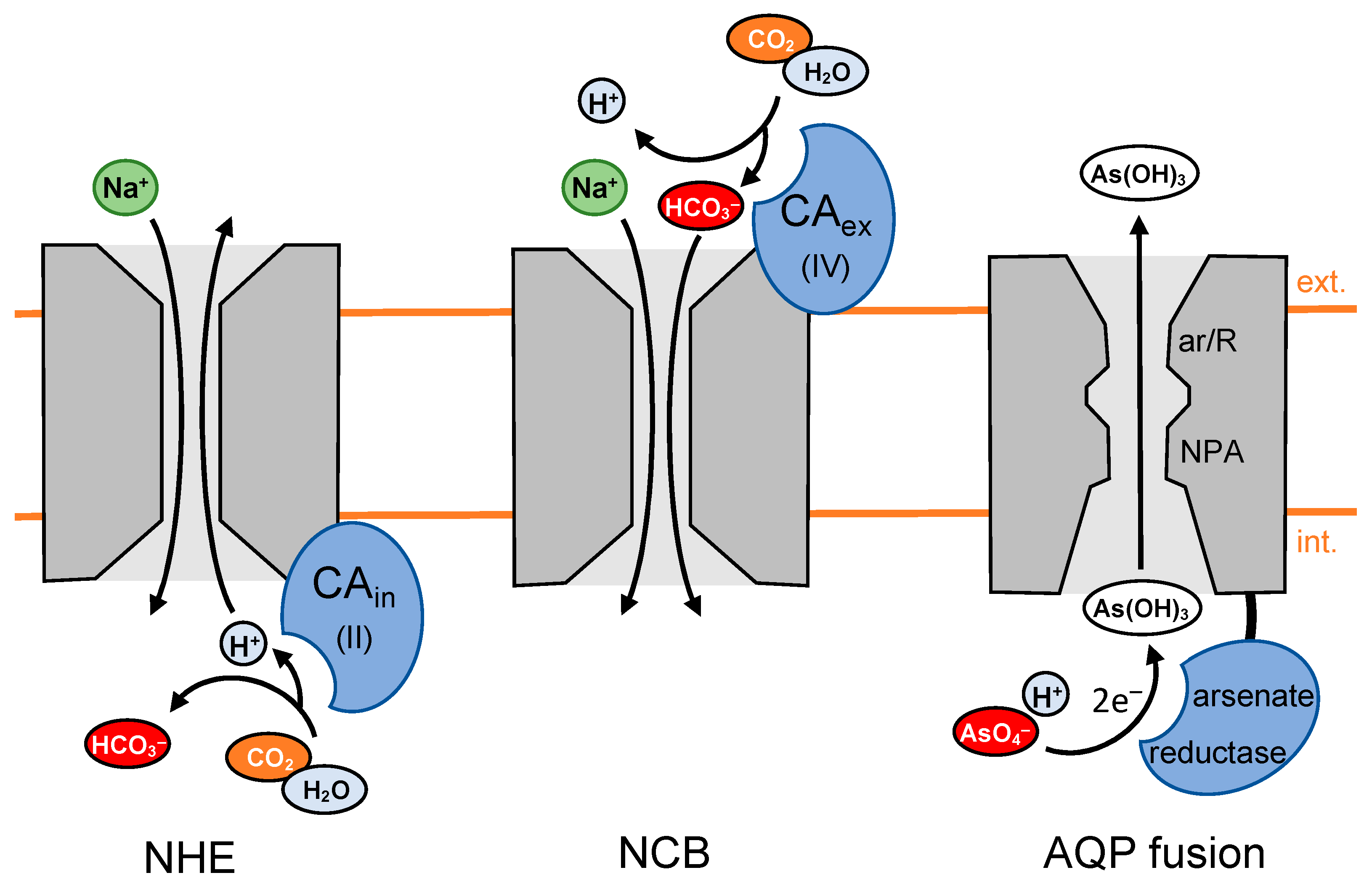
5.2. CA Activity Increases Activity of the Na+/H+ Exchanger by Locally Generating Protons
5.3. CA Activity Increases HCO3− Transport by Locally Generating Bicarbonate
5.4. Channel-Enzyme Fusion Proteins Generate and Compartmentalize Substrates as a Single Entity
6. Lipids of the Cell Membranes Facilitate Proton-Coupled Transport
7. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Barros, L.P.; Martínez, C. An enquiry into metabolite domains. Biophys. J. 2007, 92, 3878–3884. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ovens, M.J.; Davies, A.J.; Wilson, M.C.; Murray, C.M.; Halestrap, A.P. AR-C155858 is a potent inhibitor of monocarboxylate transporters MCT1 and MCT2 that binds to an intracellular site involving transmembrane helices 7–10. Biochem. J. 2010, 425, 523–530. [Google Scholar] [CrossRef] [PubMed]
- Martínez, C.; Kalise, D.; Barros, L.P. General requirement for harvesting antennae at Ca2+ and H+ channels and transporters. Front. Neuroenergetics 2010, 10, 27. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Murata, K.; Mitsuoka, K.; Hirai, T.; Walz, T.; Agre, P.; Heymann, J.B.; Engel, A.; Fujiyoshi, Y. Structural determinants of water permeation through aquaporin-1. Nature 2000, 407, 599–605. [Google Scholar] [CrossRef] [PubMed]
- Fu, D.; Libson, A.; Miercke, L.J.W.; Weitzman, C.; Nollert, P.; Krucinski, J.; Stroud, R.M. Structure of a glycerol-conducting channel and the basis for its selectivity. Science 2000, 290, 481–486. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zardoya, R. Phylogeny and evolution of the major intrinsic protein family. Biol. Cell 2005, 97, 397–414. [Google Scholar] [CrossRef] [Green Version]
- Beitz, E.; Wu, B.; Holm, L.M.; Schultz, J.E.; Zeuthen, T. Point mutations in the aromatic/arginine region in aquaporin 1 allow passage of urea, glycerol, ammonia, and protons. Proc. Natl. Acad. Sci. USA 2006, 103, 269–274. [Google Scholar] [CrossRef] [Green Version]
- Wree, D.; Wu, B.; Zeuthen, T. Requirement for asparagine in the aquaporin NPA sequence signature motifs for cation exclusion. FEBS J. 2011, 278, 740–748. [Google Scholar] [CrossRef]
- Beitz, E.; Golldack, A.; Rothert, M.; von Bülow, J. Challenges and achievements in the therapeutic modulation of aquaporin functionality. Pharmacol. Ther. 2015, 155, 22–35. [Google Scholar] [CrossRef]
- Froger, A.; Rolland, J.-P.; Bron, P.; Lagrée, V.; Le Cahérec, F.; Deschamps, S.; Hubert, J.-F.; Pellerin, I.; Thomas, D.; Delamarche, C. Functional characterization of a microbial aquaglyceroporin. Microbiology 2001, 147, 1129–1135. [Google Scholar] [CrossRef]
- Tsukaguchi, H.; Shayakul, C.; Berger, U.V.; Mackenzie, B.; Devidas, S.; Guggino, W.B.; van Hoek, A.N.; Hediger, M.A. Molecular characterization of a broad selectivity neutral solute channel. J. Biol. Chem. 1998, 273, 24737–24743. [Google Scholar] [CrossRef] [Green Version]
- Rothert, M.; Rönfeldt, D.; Beitz, E. Electrostatic attraction of weak monoacid anions increases probability for protonation and passage through aquaporins. J. Biol. Chem. 2017, 292, 9358–9364. [Google Scholar] [CrossRef] [Green Version]
- Geistlinger, K.; Schmidt, J.D.R.; Beitz, E. Lactic acid permeability of aquaporin-9 enables cytoplasmic lactate accumulation via an ion trap. Life 2022, 12, 120. [Google Scholar] [CrossRef]
- Lü, W.; Du, J.; Schwarzer, N.J.; Wacker, T.; Andrade, S.L.; Einsle, O. The formate/nitrite transporter family of anion channels. Biol. Chem. 2013, 394, 715–727. [Google Scholar] [CrossRef]
- Wu, B.; Rambow, J.; Bock, S.; Holm-Bertelsen, J.; Wiechert, M.; Soares, A.B.; Spielmann, T.; Beitz, E. Identity of a Plasmodium lactate/H+ symporter structurally unrelated to human transporters. Nat. Commun. 2015, 6, 6284. [Google Scholar] [CrossRef] [Green Version]
- Wang, Y.; Huang, Y.; Wang, J.; Cheng, C.; Huang, W.; Lu, P.; Xu, Y.; Wang, P.; Yan, N.; Shi, Y. Structure of the formate transporter FocA reveals a pentameric aquaporin-like channel. Nature 2009, 462, 467–472. [Google Scholar] [CrossRef]
- Helmstetter, F.; Arnold, P.; Höger, B.; Petersen, L.M.; Beitz, E. Formate–nitrite transporters carrying nonprotonatable amide amino acids instead of a central histidine maintain pH-dependent transport. J. Biol. Chem. 2019, 294, 623–631. [Google Scholar] [CrossRef] [Green Version]
- Czyzewski, B.K.; Wang, D.-N. Identification and characterization of a bacterial hydrosulphide ion channel. Nature 2012, 483, 494–497. [Google Scholar] [CrossRef] [Green Version]
- Nerlich, C.; Epalle, N.H.; Seick, P.; Beitz, E. Discovery and development of inhibitors of the plasmodial FNT-type lactate transporter as novel antimalarials. Pharmaceuticals 2021, 14, 1191. [Google Scholar] [CrossRef]
- Peng, X.; Wang, N.; Zhu, A.; Xu, H.; Li, J.; Zhou, Y.; Wang, C.; Xiao, Q.; Guo, L.; Liu, F.; et al. Structural characterization of the Plasmodium falciparum lactate transporter PfFNT alone and in complex with antimalarial compound MMV007839 reveals its inhibition mechanism. PLoS Biol. 2021, 19, e3001386. [Google Scholar] [CrossRef]
- Golldack, A.; Henke, B.; Bergmann, B.; Wiechert, M.; Erler, H.; Blancke Soares, A.; Spielmann, T.; Beitz, E. Substrate-analogous inhibitors exert antimalarial action by targeting the Plasmodium lactate transporter PfFNT at nanomolar scale. PLoS Pathog. 2017, 13, e1006172. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Walloch, P.; Henke, B.; Häuer, S.; Bergmann, B.; Spielmann, T.; Beitz, E. Introduction of scaffold nitrogen atoms renders inhibitors of the malarial L-lactate transporter, PfFNT, effective against the Gly107Ser resistance mutation. J. Med. Chem. 2020, 63, 9731–9741. [Google Scholar] [CrossRef] [PubMed]
- Walloch, P.; Hansen, C.; Priegann, T.; Schade, D.; Beitz, E. Pentafluoro-3-hydroxy-pent-2-en-1-ones potently inhibit FNT-type lactate transporters from all five human-pathogenic Plasmodium species. ChemMedChem 2021, 16, 1283–1289. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, J.D.R.; Beitz, E. Mutational widening of constrictions in a formate-nitrite/H+ transporter enables aquaporin-like water permeability and proton conductance. J. Biol. Chem. 2022, 298, 101513. [Google Scholar] [CrossRef] [PubMed]
- Bader, A.; Beitz, E. Transmembrane facilitation of lactate/H+ instead of lactic acid is not a question of semantics but of cell viability. Membranes 2020, 10, 236. [Google Scholar] [CrossRef]
- Wiechert, M.; Beitz, E. Mechanism of formate-nitrite transporters by dielectric shift of substrate acidity. EMBO J. 2017, 36, 949–958. [Google Scholar] [CrossRef] [Green Version]
- Wiechert, M.; Beitz, E. Formate-nitrite transporters: Monoacids ride the dielectric slide. Channels 2017, 11, 365–367. [Google Scholar] [CrossRef] [Green Version]
- Winkler, F.K. Amt/MEP/Rh proteins conduct ammonia. Pflug. Arch. 2006, 451, 701–707. [Google Scholar] [CrossRef] [Green Version]
- Khademi, S.; O’Connell, J., III; Remis, J.; Robles-Colmenares, Y.; Miercke, L.J.W.; Stroud, M. Mechanism of ammonia transport by Amt/MEP/Rh: Structure of AmtB at 1.35 A. Science 2004, 305, 1587–1594. [Google Scholar] [CrossRef] [Green Version]
- Javelle, A.; Lupo, D.; Ripoche, P.; Fulford, T.; Merrick, M.; Winkler, F.K. Substrate binding, deprotonation, and selectivity at the periplasmic entrance of the Escherichia coli ammonia channel AmtB. Proc. Natl. Acad. Sci. USA 2008, 105, 5040–5045. [Google Scholar] [CrossRef]
- Hall, J.A.; Kustu, S. The pivotal twin histidines and aromatic triad of the Escherichia coli ammonium channel AmtB can be replaced. Proc. Natl. Acad. Sci. USA 2011, 108, 13270–13274. [Google Scholar] [CrossRef] [Green Version]
- Pantoja, O. High affinity ammonium transporters: Molecular mechanism of action. Front. Plant Sci. 2012, 3, 34. [Google Scholar] [CrossRef] [Green Version]
- Halestrap, A.P. The SLC16 gene family—Structure, role and regulation in health and disease. Mol. Asp. Med. 2013, 34, 337–349. [Google Scholar] [CrossRef]
- Felmlee, M.A.; Jones, R.S.; Rodriguez-Cruz, V.; Follman, K.E.; Morris, M.E. Monocarboxylate transporters (SLC16): Function, regulation, and role in health and disease. Pharmacol. Rev. 2020, 72, 466–485. [Google Scholar] [CrossRef]
- Bouzier-Sore, A.-K.; Voisin, P.; Bouchaud, V.; Bezancon, E.; Franconi, J.-M.; Pellerin, L. Competition between glucose and lactate as oxidative energy substrates in both neurons and astrocytes: A comparative NMR study. Eur. J. Neurosci. 2006, 24, 1687–1694. [Google Scholar] [CrossRef]
- Wang, N.; Jiang, X.; Zhang, S.; Zhu, A.; Yuan, Y.; Xu, H.; Lei, J.; Yan, C. Structural basis of human monocarboxylate transporter 1 inhibition by anti-cancer drug candidates. Cell 2021, 184, 370–383.e13. [Google Scholar] [CrossRef]
- Kirk, P.; Wilson, M.C.; Heddle, C.; Brown, M.H.; Barclay, A.N.; Halestrap, A.P. CD147 is tightly associated with lactate transporters MCT1 and MCT4 and facilitates their cell surface expression. EMBO J. 2000, 19, 3896–3904. [Google Scholar] [CrossRef]
- Wilson, C.M.; Meredith, D.; Fox, J.E.M.; Manoharam, C.; Davies, A.J.; Halestrap, A.P. Basigin (CD147) Is the target for organomercurial inhibition of monocarboxylate transporter isoforms 1 and 4. J. Biol. Chem. 2005, 280, 27213–27221. [Google Scholar] [CrossRef] [Green Version]
- Klier, M.; Schüler, C.; Halestrap, A.P.; Sly, W.S.; Deitmer, J.W.; Becker, H.M. Transport activity of the high-affinity monocarboxylate transporter MCT2 is enhanced by extracellular carbonic anhydrase IV but not by intracellular carbonic anhydrase II. J. Biol. Chem. 2011, 286, 27781–27791. [Google Scholar] [CrossRef] [Green Version]
- Köpnick, A.-L.; Jansen, A.; Geistlinger, K.; Epalle, N.H.; Beitz, E. Basigin drives intracellular accumulation of L-lactate by harvesting protons and substrate anions. PLoS ONE 2021, 16, e0249110. [Google Scholar] [CrossRef]
- Redzic, J.S.; Armstrong, C.S.; Isern, N.G.; Jones, D.N.M.; Kieft, J.S.; Eisenmesser, E.Z. The retinal specific CD147 Ig0 domain: From molecular structure to biological activity. J. Mol. Biol. 2011, 411, 68–82. [Google Scholar] [CrossRef] [PubMed]
- Updegraff, B.L.; Zhou, X.; Guo, Y.; Padanad, M.S.; Chen, P.-H.; Yang, C.; Sudderth, J.; Rodriguez-Tirado, C.; Girard, L.; Minna, J.D.; et al. Transmembrane protease TMPRSS11B promotes lung cancer growth by enhancing lactate export and glycolytic metabolism. Cell Rep. 2018, 25, 2223–2233. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Becker, H.M.; Deitmer, J.W. Proton transport in cancer cells: The role of carbonic anhydrases. Int. J. Mol. Sci. 2021, 22, 3171. [Google Scholar] [CrossRef] [PubMed]
- Okuyama, T.; Waheed, A.; Kusumoto, W.; Zhu, X.L.; Sly, W.S. Carbonic anhydrase IV: Role of removal of C-terminal domain in glycosylphosphatidylinositol anchoring and realization of enzyme activity. Arch. Biochem. Biophys. 1995, 320, 315–322. [Google Scholar] [CrossRef] [PubMed]
- Forero-Quintero, L.S.; Ames, S.; Schneider, H.-P.; Thyssen, A.; Boone, C.D.; Andring, J.T.; McKenna, R.; Casey, J.R.; Deitmer, J.W.; Becker, H.M. Membrane-anchored carbonic anhydrase IV interacts with monocarboxylate transporters via their chaperones CD147 and GP70. J. Biol. Chem. 2019, 294, 593–607. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Klier, M.; Andes, T.; Deitmer, J.; Becker, M.H. Intracellular and extracellular carbonic anhydrases cooperate non-enzymatically to enhance activity of monocarboxylate transporters. J. Biol. Chem. 2014, 289, 2765–2775. [Google Scholar] [CrossRef] [Green Version]
- Noor, S.I.; Pouyssegur, J.; Deitmer, J.W.; Becker, H.M. Integration of a ‘proton antenna’ facilitates transport activity of the monocarboxylate transporter MCT4. FEBS J. 2017, 284, 149–162. [Google Scholar] [CrossRef] [Green Version]
- Jamali, S.; Klier, M.; Ames, S.; Barros, L.F.; McKenna, R.; Dietmer, J.D.; Beker, M. Hypoxia-induced carbonic anhydrase IX facilitates lactate flux in human breast cancer cells by non-catalytic function. Sci. Rep. 2015, 5, 13605. [Google Scholar] [CrossRef] [Green Version]
- Ames, S.; Pastorekova, S.; Becker, H.B. The proteoglycan-like domain of carbonic anhydrase IX mediates non-catalytic facilitation of lactate transport in cancer cells. Oncotarget 2018, 9, 27940–27957. [Google Scholar] [CrossRef] [Green Version]
- Kaluz, S.; Kaluzová, M.; Liao, S.-Y.; Lerman, M.; Stanbridge, E.J. Transcriptional control of the tumor- and hypoxia-marker carbonic anhydrase 9: A one transcription factor (HIF-1) show? Biochim. Biophys. Acta 2009, 1795, 162–172. [Google Scholar] [CrossRef]
- Stridh, M.H.; Alt, M.D.; Wittmann, S.; Heidtmann, H.; Aggarwal, M.; Reiderer, B.; Seidler, U.; Wennemuth, G.; McKenna, R.; Dietmer, J.W.; et al. Lactate flux in astrocytes is enhanced by a non-catalytic action of carbonic anhydrase II. J. Physiol. 2012, 510, 2333–2351. [Google Scholar] [CrossRef]
- Noor, S.I.; Dietz, S.; Heidtmann, H.; Boone, C.D.; McKenna, R.; Deitmer, J.W.; Becker, H.M. Analysis of the binding moiety mediating the interaction between monocarboxylate transporters and carbonic anhydrase II. J. Biol. Chem. 2015, 90, 4476–4486. [Google Scholar] [CrossRef] [Green Version]
- Noor, S.I.; Jamali, S.; Ames, S.; Langer, S.; Deitmer, J.W.; Becker, H.M. A surface proton antenna in carbonic anhydrase II supports lactate transport in cancer cells. Elife 2018, 7, e35176. [Google Scholar] [CrossRef]
- Becker, H.M.; Deitmer, J.W. Nonenzymatic proton handling by carbonic anhydrase II during H+-lactate cotransport via monocarboxylate transporter 1. J. Biol. Chem. 2008, 283, 21655–21667. [Google Scholar] [CrossRef]
- Shinobu, A.; Agmon, N. Mapping proton wires in proteins: Carbonic anhydrase and GFP chromophore biosynthesis. J. Phys. Chem. A 2009, 113, 7253–7266. [Google Scholar] [CrossRef]
- Heberle, J.; Riesle, J.; Thiedmann, G.; Oesterhelt, D.; Dencher, N.A. Proton migration along the membrane surface and retarded surface to bulk transfer. Nature 1994, 70, 379–382. [Google Scholar] [CrossRef]
- Brändén, M.; Sandén, T.; Brzezinski, P.; Widengren, J. Localized proton microcircuits at the biological membrane-water interface. Proc. Natl. Acad. Sci. USA 2006, 103, 19766–19770. [Google Scholar] [CrossRef] [Green Version]
- Svichar, N.; Chesler, M. Surface Carbonic anhydrase activity on astrocytes and neurons facilitates lactate transport. Glia 2003, 41, 415–419. [Google Scholar] [CrossRef]
- Wetzel, P.; Hasse, A.; Papadopoulos, S.; Voipio, J.; Kaila, K.; Gros, G. Extracellular carbonic anhydrase activity facilitates lactic acid transport in rat skeletal muscle fibres. J. Physiol. 2001, 531, 743–756. [Google Scholar] [CrossRef]
- Li, X.; Alvarez, B.; Casey, J.R.; Reithmeier, R.A.F.; Fliegel, L. Carbonic anhydrase II binds to and enhances activity of the Na+ /H+ exchanger. J. Biol. Chem. 2002, 277, 36085–36091. [Google Scholar] [CrossRef]
- Krishnan, D.; Liu, L.; Wiebe, S.A.; Casey, J.R.; Cordat, E.; Alexander, R.T. Carbonic anhydrase II binds to and increases the activity of the epithelial sodium-proton exchanger, NHE3. Am. J. Physiol. 2015, 309, 383–392. [Google Scholar] [CrossRef] [Green Version]
- Wu, Q.; Pierce, V.M.; Delamere, N.A. Cytoplasmic pH responses to carbonic anhydrase inhibitors in cultured rabbit nonpigmented ciliary epithelium. J. Membr. Biol. 1998, 162, 31–38. [Google Scholar] [CrossRef] [PubMed]
- Liskova, V.; Hudecova, S.; Lencesova, L.; Iuliano, F.; Sirova, M.; Ondrias, K.; Pasterkova, S.; Krizanova, O. Type 1 Sodium calcium exchanger forms a complex with carbonic anhydrase IX and via reverse mode activity contributes to pH control in hypoxic tumors. Cancers 2019, 11, 1139. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Becker, H.M. Carbonic anhydrase IX and acid transport in cancer. Br. J. Cancer 2020, 122, 157–167. [Google Scholar] [CrossRef] [PubMed]
- Dietmer, J.W.; Becker, H.M. Transport metabolons with carbonic anhydrases. Front. Physiol. 2013, 4, 291. [Google Scholar] [CrossRef] [Green Version]
- Becker, H.M.; Deitmer, J.W. Transport metabolons and acid/base balance in tumor cells. Cancers 2020, 12, 899. [Google Scholar] [CrossRef] [Green Version]
- Alvarez, B.V.; Loiselle, F.B.; Supuran, C.T.; Schwartz, G.J.; Casey, J.R. Direct extracellular interaction between carbonic anhydrase IV and the human NBC1 sodium/bicarbonate co-transporter. Biochemistry 2003, 42, 12321–12329. [Google Scholar] [CrossRef]
- Gross, E.; Pushkin, A.; Abuladze, N.; Fedotoff, O.; Kurtz, I. Regulation of the sodium bicarbonate cotransporter kNBC1 function: Role of Asp986, Asp988 and kNBC1—carbonic anhydrase II binding. J. Physiol. 2002, 544, 679–685. [Google Scholar] [CrossRef]
- Loiselle, F.B.; Morgan, P.E.; Alvarez, B.V.; Casey, J.R. Regulation of the human NBC3 Na+/HCO3− cotransporter by carbonic anhydrase II and PKA. Am. J. Physiol. 2004, 86, 1423–1433. [Google Scholar] [CrossRef] [Green Version]
- Morgan, P.E.; Pastoreková, S.; Stuart-Tilley, A.K.; Alper, S.L.; Casey, J.R. Interactions of transmembrane carbonic anhydrase, CAIX, with bicarbonate transporters. Am. J. Physiol. 2007, 293, 738–748. [Google Scholar] [CrossRef]
- Vince, J.W.; Reithmeier, R.A.F. Carbonic anhydrase II binds to the carboxyl terminus of human band 3, the erythrocyte Cl−/HCO3+ exchanger. J. Biol. Chem. 1998, 273, 28430–28437. [Google Scholar] [CrossRef] [Green Version]
- Vince, J.W.; Carlsson, U.; Reithmeier, R.A.F. Localization of the Cl−/HCO3− anion exchanger binding site to the amino-terminal region of carbonic anhydrase II. Biochemistry 2000, 39, 13344–13349. [Google Scholar] [CrossRef]
- Sterling, D.; Reithmeier, R.A.F.; Casey, J.R. A transport metabolon. Functional interaction of carbonic anhydrase II and chloride/bicarbonate exchangers. J. Biol. Chem. 2001, 276, 47886–47894. [Google Scholar] [CrossRef] [Green Version]
- Piermarini, P.M.; Kim, E.; Boron, W.F. Evidence against a direct interaction between intracellular carbonic anhydrase II and pure C-terminal domains of SLC4 bicarbonate transporters. J. Biol. Chem. 2007, 282, 1409–1421. [Google Scholar] [CrossRef] [Green Version]
- Al-Samir, S.; Papadopoulos, S.; Scheibe, R.J.; Meißne, J.D.; Cartron, J.-P.; Sly, W.S.; Alper, S.L.; Gros, G.; Endeward, V. Activity and distribution of intracellular carbonic anhydrase II and their effects on the transport activity of anion exchanger AE1/SLC4A1. J. Physiol. 2013, 591, 4963–4982. [Google Scholar] [CrossRef]
- Yamada, H.; Horita, S.; Suzuki, M.; Fujita, T.; Seki, G. Functional role of a putative carbonic anhydrase II-binding domain in the electrogenic Na+-HCO3− cotransporter NBCe1 expressed in Xenopus oocytes. Channels 2011, 5, 106–109. [Google Scholar] [CrossRef] [Green Version]
- Lu, J.; Daly, C.M.; Parker, M.D.; Gill, H.S.; Piermarini, P.M.; Pelletier, M.F.; Boron, W.F. Effect of human carbonic anhydrase II on the activity of the human electrogenic Na+/HCO3− cotransporter NBCe1-A in Xenopus oocytes. J. Biol. Chem. 2006, 281, 19241–19250. [Google Scholar] [CrossRef] [Green Version]
- McMurtrie, H.; Cleary, H.J.; Alvarez, B.V.; Loiselle, F.B.; Sterling, D.; Morgan, P.E.; Johnson, D.E.; Casey, J.R. The bicarbonate transport metabolon. J. Enzym. Inhib. Med. Chem. 2004, 19, 231–236. [Google Scholar] [CrossRef] [Green Version]
- Song, J.; Wu, B.; Beitz, E. Functional and evolutional implications of natural channel-enzyme fusion proteins. Biomol. Concepts 2011, 2, 439–444. [Google Scholar] [CrossRef]
- Wu, B.; Song, J.; Beitz, E. Novel channel enzyme fusion proteins confer arsenate resistance. J. Biol. Chem. 2010, 285, 40081–40087. [Google Scholar] [CrossRef]
- Xu, L.; Öjemyr, L.N.; Bergstrand, J.; Brzezinski, P.; Widengren, J. Protonation dynamics on lipid nanodiscs: Influence of the membrane surface area and external buffers. Biophys. J. 2016, 110, 1993–2003. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gutman, M.; Nachliel, E. Time-resolved dynamics of proton transfer in proteinous systems. Annu. Rev. Phys Chem. 1997, 48, 329–356. [Google Scholar] [CrossRef] [PubMed]
- Wolf, M.G.; Grubmüller, H.; Groenhof, G. Anomalous surface diffusion of protons on lipid membranes. Biophys. J. 2014, 107, 76–87. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Flenner, E.; Das, J.; Rheinstädter, M.C.; Kosztin, I. Subdiffusion and lateral diffusion coefficient of lipid atoms and molecules in phospholipid bilayers. Phys. Rev. E Stat. Nonlin. Soft Matter Phys. 2009, 79, 011907. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Springer, A.; Hagen, V.; Cherepanov, D.A.; Antonenko, Y.N.; Pohl, P. Protons migrate along interfacial water without significant contributions from jumps between ionizable groups on the membrane surface. Proc. Natl. Acad. Sci. USA 2011, 108, 14461–14466. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Epalle, N.H.; Beitz, E. Local Attraction of Substrates and Co-Substrates Enhances Weak Acid and Base Transmembrane Transport. Biomolecules 2022, 12, 1794. https://doi.org/10.3390/biom12121794
Epalle NH, Beitz E. Local Attraction of Substrates and Co-Substrates Enhances Weak Acid and Base Transmembrane Transport. Biomolecules. 2022; 12(12):1794. https://doi.org/10.3390/biom12121794
Chicago/Turabian StyleEpalle, Nathan Hugo, and Eric Beitz. 2022. "Local Attraction of Substrates and Co-Substrates Enhances Weak Acid and Base Transmembrane Transport" Biomolecules 12, no. 12: 1794. https://doi.org/10.3390/biom12121794
APA StyleEpalle, N. H., & Beitz, E. (2022). Local Attraction of Substrates and Co-Substrates Enhances Weak Acid and Base Transmembrane Transport. Biomolecules, 12(12), 1794. https://doi.org/10.3390/biom12121794





