Rational Development and Characterization of a Ubiquitin Variant with Selectivity for Ubiquitin C-Terminal Hydrolase L3
Abstract
:1. Introduction
2. Materials and Methods
2.1. Computational Interaction Analysis and Positional Scanning
2.2. Plasmids and Cloning
2.3. Recombinant Expression of UCHL3 and UCHL1 Proteins
2.4. Recombinant Expression of UbV Proteins
2.5. Ubiquitin Intein Chitin Binding Domain Expression and Purification for Ub-ABPs
2.6. Binding Affinity Analysis
2.7. Inhibition Assays
2.8. Quantification of UbV-ABP Inactivation Efficiency
2.9. Cellular Target Engagement Assays
3. Results
3.1. Characterization of T66K/V70F Double Mutant UbV
3.1.1. Inactivation Efficiency of Double Mutants against UCHL3 and UCHL1
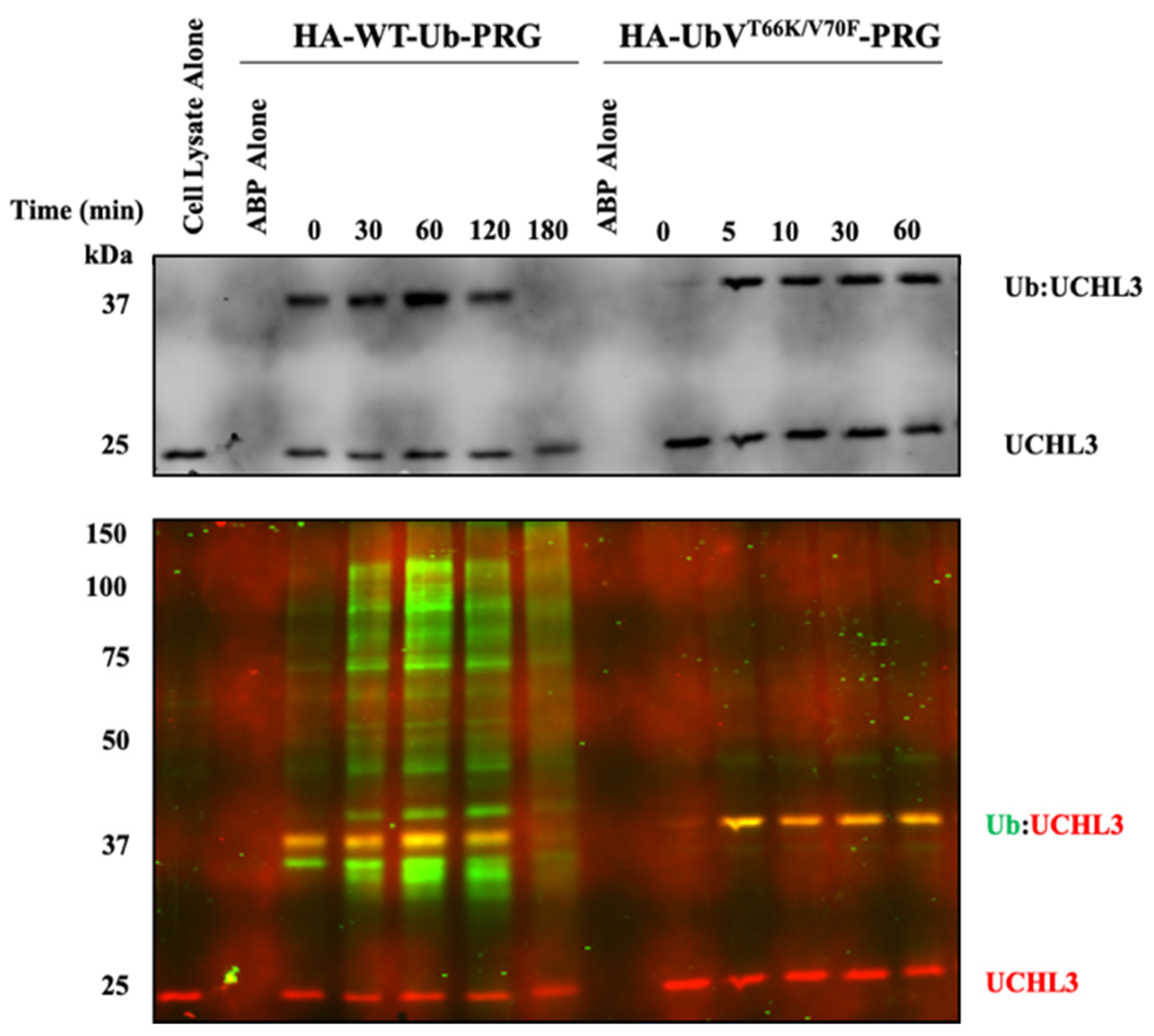
3.1.2. Characterization of UbVT66K/V70F-PRG in HEK293 Cell Lysates
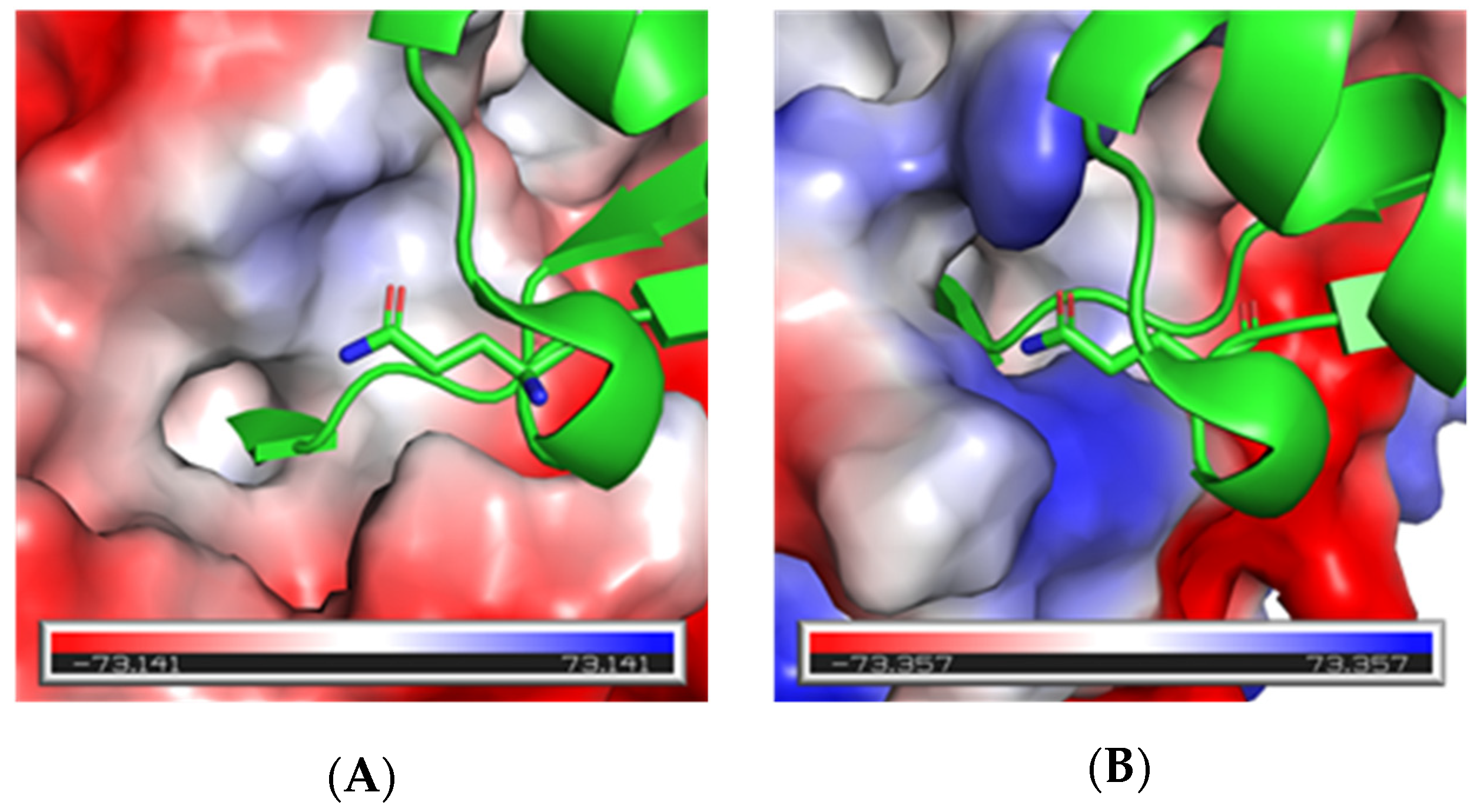
3.2. Computational Binding Analysis and Positional Scanning
3.3. In Vitro Analysis of UbV toward UCHL3 and UCHL1
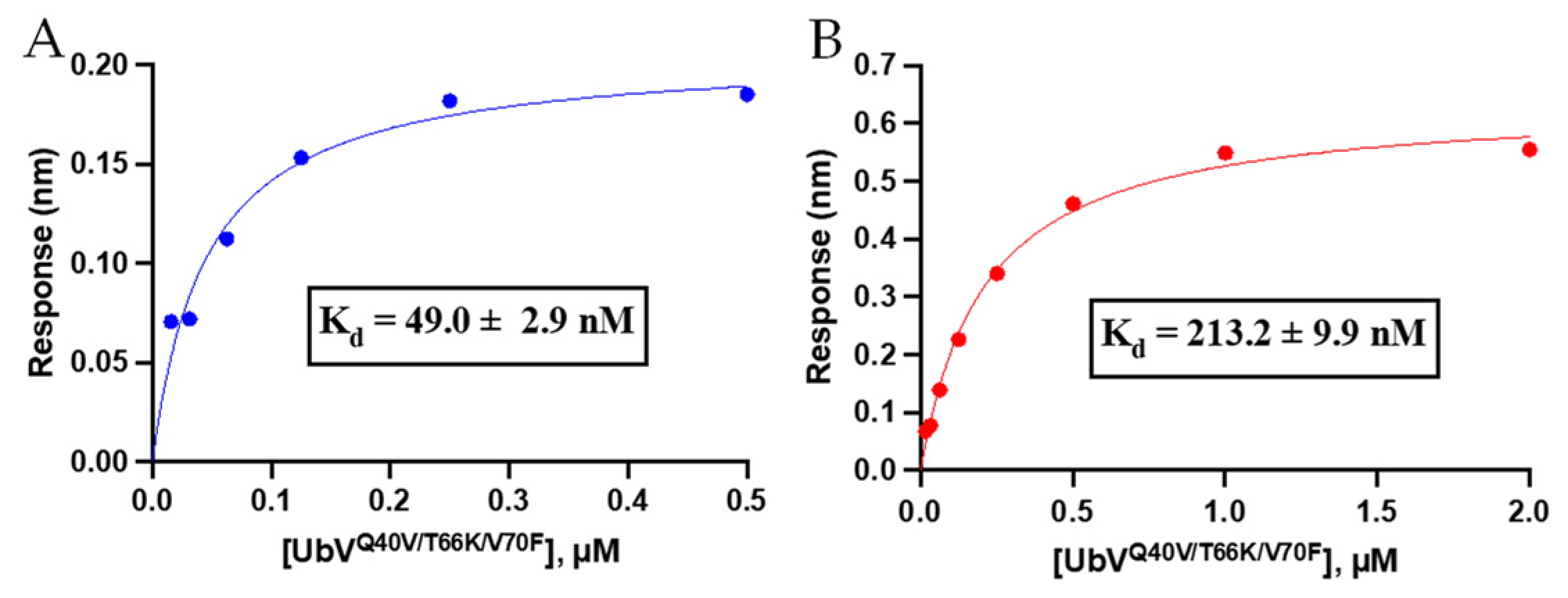
3.3.1. Binding Affinity Analysis of Triple-Mutant UbV Using Biolayer Interferometry
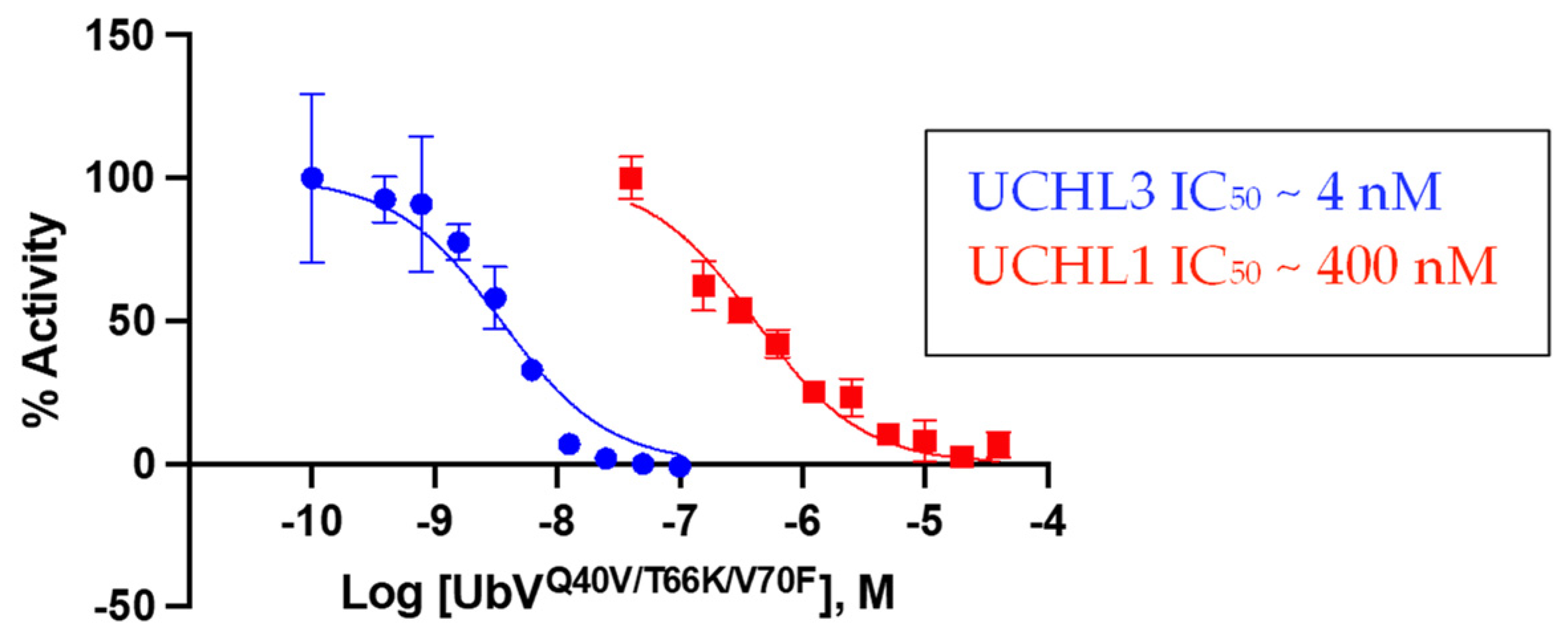
3.3.2. Biochemical Inhibition of DUB Activity by the Triple-Mutant UbV
3.3.3. Inactivation Efficiency of UbV Triple-mutant Activity-Based Probes
3.4. Evaluation of UbVQ40V/T66K/V70F-PRG in MDA-MB-231 Cell Lysates
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hewings, D.S.; Heideker, J.; Ma, T.P.; AhYoung, A.P.; El Oualid, F.; Amore, A.; Costakes, G.T.; Kirchhofer, D.; Brasher, B.; Pillow, T.; et al. Reactive-site-centric chemoproteomics identifies a distinct class of deubiquitinase enzymes. Nat. Commun. 2018, 9, 1162. [Google Scholar] [CrossRef] [PubMed]
- Kwasna, D.; Rehman, S.A.A.; Natarajan, J.; Matthews, S.; Madden, R.; De Cesare, V.; Weidlich, S.; Virdee, S.; Ahel, I.; Gibbs-Seymour, I. Discovery and characterization of ZUFSP/ZUP1, a distinct deubiquitinase class important for genome stability. Mol. Cell 2018, 70, 150–164. [Google Scholar] [CrossRef] [Green Version]
- Abdul Rehman, S.A.; Kristariyanto, Y.A.; Choi, S.Y.; Nkosi, P.J.; Weidlich, S.; Labib, K.; Hofmann, K.; Kulathu, Y. MINDY-1 Is a Member of an Evolutionarily Conserved and Structurally Distinct New Family of Deubiquitinating Enzymes. Mol. Cell 2016, 63, 146–155. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liao, C.; Beveridge, R.; Hudson, J.J.R.; Parker, J.D.; Chiang, S.C.; Ray, S.; Ashour, M.E.; Sudbery, I.; Dickman, M.J.; El-Khamisy, S.F. UCHL3 Regulates Topoisomerase-Induced Chromosomal Break Repair by Controlling TDP1 Proteostasis. Cell Rep. 2018, 23, 3352–3365. [Google Scholar] [CrossRef]
- Nishi, R.; Wijnhoven, P.W.G.; Kimura, Y.; Matsui, M.; Konietzny, R.; Wu, Q.; Nakamura, K.; Blundell, T.L.; Kessler, B.M. The deubiquitylating enzyme UCHL3 regulates Ku80 retention at sites of DNA damage. Sci. Rep. 2018, 8, 17891. [Google Scholar] [CrossRef]
- Ishida, N.; Nakagawa, T.; Iemura, S.-I.; Yasui, A.; Shima, H.; Katoh, Y.; Nagasawa, Y.; Natsume, T.; Igarashi, K.; Nakayama, K. Ubiquitylation of Ku80 by RNF126 Promotes Completion of Nonhomologous End Joining-Mediated DNA Repair. Mol. Cell. Biol. 2017, 37, e00347-16. [Google Scholar] [CrossRef] [Green Version]
- Postow, L.; Ghenoiu, C.; Woo, E.M.; Krutchinsky, A.N.; Chait, B.T.; Funabiki, H. Ku80 removal from DNA through double strand break—Induced ubiquitylation. J. Cell Biol. 2008, 182, 467–479. [Google Scholar] [CrossRef] [Green Version]
- Yang, R.; Liu, N.; Chen, L.; Jiang, Y.; Shi, Y.; Mao, C.; Liu, Y.; Wang, M.; Lai, W.; Tang, H.; et al. GIAT4RA functions as a tumor suppressor in non-small cell lung cancer by counteracting Uchl3–mediated deubiquitination of LSH. Oncogene 2019, 38, 7133–7145. [Google Scholar] [CrossRef]
- Luo, K.; Li, L.; Li, Y.; Wu, C.; Yin, Y.; Chen, Y.; Deng, M.; Nowsheen, S.; Yuan, J.; Lou, Z. A phosphorylation–deubiquitination cascade regulates the BRCA2–RAD51 axis in homologous recombination. Genes Dev. 2016, 30, 2581–2595. [Google Scholar] [CrossRef] [PubMed]
- Song, Z.; Li, J.; Zhang, L.; Deng, J.; Fang, Z.; Xiang, X.; Xiong, J. UCHL3 promotes pancreatic cancer progression and chemo-resistance through FOXM1 stabilization. Am. J. Cancer Res. 2019, 9, 1970–1981. [Google Scholar]
- Zhang, M.H.; Zhang, H.H.; Du, X.H.; Gao, J.; Li, C.; Shi, H.R.; Li, S.Z. UCHL3 promotes ovarian cancer progression by stabilizing TRAF2 to activate the NF-κB pathway. Oncogene 2020, 39, 322–333. [Google Scholar] [CrossRef]
- Song, Z.; Tu, X.; Zhou, Q.; Huang, J.; Chen, Y.; Liu, J.; Lee, S.B.; Kim, W.; Nowsheen, S.; Luo, K.; et al. A novel UCHL3 inhibitor, perifosine, enhances PARP inhibitor cytotoxicity through inhibition of homologous recombination-mediated DNA double strand break repair. Cell Death Dis. 2019, 10, 398. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, Y.; Lashuel, H.; Choi, S.; Xing, X.; Case, A.; Ni, J.; Yeh, L.-A.; Cuny, G.D.; Stein, R.L.; Lansbury, P.T. Discovery of Inhibitors that Elucidate the Role of UCH-L1 Activity in the H1299 Lung Cancer Cell Line. Chem. Biol. 2003, 10, 837–846. [Google Scholar] [CrossRef] [Green Version]
- Chiarini, F.; Del Sole, M.; Mongiorgi, S.; Gaboardi, G.C.; Cappellini, A.; Mantovani, I.; Follo, M.Y.; McCubrey, J.A.; Martelli, A.M. The novel Akt inhibitor, perifosine, induces caspase-dependent apoptosis and downregulates P-glycoprotein expression in multidrug-resistant human T-acute leukemia cells by a JNK-dependent mechanism. Leukemia 2008, 22, 1106–1116. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kondapaka, S.B.; Singh, S.S.; Dasmahapatra, G.P.; Sausville, E.A.; Roy, K.K. Perifosine, a novel alkylphospholipid, inhibits protein kinase B activation. Mol. Cancer Ther. 2003, 2, 1093–1103. [Google Scholar] [PubMed]
- Ernst, A.; Avvakumov, G.; Tong, J.; Fan, Y.; Zhao, Y.; Alberts, P.; Persaud, A.; Walker, J.R.; Neculai, A.; Neculai, D.; et al. A strategy for modulation of enzymes in the Ubiquitin System. Science 2013, 209, 590–596. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Leung, I.; Jarvik, N.; Sidhu, S.S. A Highly Diverse and Functional Naïve Ubiquitin Variant Library for Generation of Intracellular Affinity Reagents. J. Mol. Biol. 2017, 429, 115–127. [Google Scholar] [CrossRef]
- Zhang, W.; Sartori, M.A.; Makhnevych, T.; Federowicz, K.E.; Dong, X.; Liu, L.; Nim, S.; Dong, A.; Yang, J.; Li, Y.; et al. Generation and Validation of Intracellular Ubiquitin Variant Inhibitors for USP7 and USP10. J. Mol. Biol. 2017, 429, 3546–3560. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhou, L.; Rouge, L.; Phillips, A.H.; Lam, C.; Liu, P.; Sandoval, W.; Helgason, E.; Murray, J.M.; Wertz, I.E.; et al. Conformational stabilization of ubiquitin yields potent and selective inhibitors of USP7. Nat. Chem. Biol. 2012, 9, 51–58. [Google Scholar] [CrossRef] [PubMed]
- Hewitt, C.S.; Krabill, A.D.; Das, C.; Flaherty, D.P. Development of Ubiquitin Variants with Selectivity for Ubiquitin C-Terminal Hydrolase Deubiquitinases. Biochemistry 2020, 59, 3447–3462. [Google Scholar] [CrossRef]
- Gjonaj, L.; Sapmaz, A.; González-Prieto, R.; Vertegaal, A.C.O.; Flierman, D.; Ovaa, H. USP7: Combining tools towards selectivity. Chem. Commun. 2019, 55, 5075–5078. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gjonaj, L.; Sapmaz, A.; Flierman, D.; Janssen, G.M.C.; van Veelen, P.A.; Ovaa, H. Development of a DUB-selective fluorogenic substrate. Chem. Sci. 2019, 10, 10290–10296. [Google Scholar] [CrossRef]
- Rut, W.; Zmudzinski, M.; Snipas, S.J.; Bekes, M.; Huang, T.T.; Drag, M.; Drag, M. Engineered unnatural ubiquitin for optimal detection of deubiquitinating enzymes. Chem. Sci. 2020, 11, 6058–6069. [Google Scholar] [CrossRef]
- Misaghi, S.; Galardy, P.J.; Meester, W.J.N.; Ovaa, H.; Ploegh, H.L.; Gaudet, R. Structure of the ubiquitin hydrolase UCH-L3 complexed with a suicide substrate. J. Biol. Chem. 2005, 280, 1512–1520. [Google Scholar] [CrossRef] [Green Version]
- Boudreaux, D.A.; Maiti, T.K.; Davies, C.W.; Das, C. Ubiquitin vinyl methyl ester binding orients the misaligned active site of the ubiquitin hydrolase UCHL1 into productive conformation. Proc. Natl. Acad. Sci. USA. 2010, 107, 9117–9122. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shelley, J.C.; Cholleti, A.; Frye, L.L.; Greenwood, J.R.; Timlin, M.R.; Uchimaya, M. Epik: A software program for pK a prediction and protonation state generation for drug-like molecules. J. Comput. Aided. Mol. Des. 2007, 21, 681–691. [Google Scholar] [CrossRef] [PubMed]
- Beard, H.; Cholleti, A.; Pearlman, D.; Sherman, W.; Loving, K.A. Applying physics-based scoring to calculate free energies of binding for single amino acid mutations in protein-protein complexes. PLoS ONE 2013, 8, e82849. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sirin, S.; Pearlman, D.A.; Sherman, W. Physics-based enzyme design: Predicting binding affinity and catalytic activity. Proteins Struct. Funct. Bioinforma. 2014, 82, 3397–3409. [Google Scholar] [CrossRef]
- Krabill, A.D.; Chen, H.; Hussain, S.; Feng, C.; Abdullah, A.; Das, C.; Aryal, U.K.; Post, C.B.; Wendt, M.K.; Galardy, P.J.; et al. Ubiquitin C-Terminal Hydrolase L1: Biochemical and Cellular Characterization of a Covalent Cyanopyrrolidine-Based Inhibitor. ChemBioChem 2020, 21, 712–721. [Google Scholar] [CrossRef]
- Krabill, A.D.; Chen, H.; Hussain, S.; Hewitt, C.S.; Imhoff, R.D.; Muli, C.S.; Das, C.; Galardy, P.J.; Wendt, M.K.; Flaherty, D.P. Optimization and Anti-Cancer Properties of Fluoromethylketones as covalent inhibitors for Ubiquitin C-terminal Hydrolase L1. Molecules 2021, 26, 1227. [Google Scholar] [CrossRef]
- Sheedlo, M.J.; Kenny, S.; Podkorytov, I.S.; Brown, K.; Ma, J.; Iyer, S.; Hewitt, C.S.; Arbough, T.; Mikhailovskii, O.; Flaherty, D.P.; et al. Insights into Ubiquitin Product Release in Hydrolysis Catalyzed by the Bacterial Deubiquitinase SdeA. Biochemistry 2021, 60, 584–596. [Google Scholar] [CrossRef]
- Hassiepen, U.; Eidhoff, U.; Meder, G.; Bulber, J.F.; Hein, A.; Bodendorf, U.; Lorthiois, E.; Martoglio, B. A sensitive fluorescence intensity assay for deubiquitinating proteases using ubiquitin-rhodamine110-glycine as substrate. Anal. Biochem. 2007, 371, 201–207. [Google Scholar] [CrossRef] [PubMed]
- Resnick, E.; Bradley, A.; Gan, J.; Douangamath, A.; Krojer, T.; Sethi, R.; Geurink, P.P.; Aimon, A.; Amitai, G.; Bellini, D.; et al. Rapid Covalent-Probe Discovery by Electrophile-Fragment Screening. J. Am. Chem. Soc. 2019, 141, 8951–8968. [Google Scholar] [CrossRef] [Green Version]
- Griffin, P.; Sexton, A.; Macneill, L.; Iizuka, Y.; Lee, M.K.; Bazzaro, M. Method for measuring the activity of deubiquitinating enzymes in cell lines and tissue samples. J. Vis. Exp. 2015, 99, 52784. [Google Scholar] [CrossRef] [Green Version]
- Routenberg Love, K.; Pandya, R.K.; Spooner, E.; Ploegh, H.L. Ubiquitin C-Terminal Electrophiles Are Activity-Base Probes for Identification and Mechanistic Study of Ubiquitin Conjugating Machinery. ACS Chem. Biol. 2009, 4, 275. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ekkebus, R.; Van Kasteren, S.I.; Kulathu, Y.; Scholten, A.; Berlin, I.; Geurink, P.P.; De Jong, A.; Goerdayal, S.; Neefjes, J.; Heck, A.J.R.; et al. On terminal alkynes that can react with active-site cysteine nucleophiles in proteases. J. Am. Chem. Soc. 2013, 135, 2867–2870. [Google Scholar] [CrossRef]
- Nishikawa, K.; Li, H.; Kawamura, R.; Osaka, H.; Wang, Y.L.; Hara, Y.; Hirokawa, T.; Manago, Y.; Amano, T.; Noda, M.; et al. Alterations of structure and hydrolase activity of parkinsonism-associated human ubiquitin carboxyl-terminal hydrolase L1 variants. Biochem. Biophys. Res. Commun. 2003, 304, 176–183. [Google Scholar] [CrossRef]
- Gui, W.; Ott, C.A.; Yang, K.; Chung, J.S.; Shen, S.; Zhuang, Z. Cell-Permeable Activity-Based Ubiquitin Probes Enable Intracellular Profiling of Human Deubiquitinases. J. Am. Chem. Soc. 2018, 140, 12424–12433. [Google Scholar] [CrossRef]
- Gui, W.; Shen, S.; Zhuang, Z. Photocaged Cell-Permeable Ubiquitin Probe for Temporal Profiling of Deubiquitinating Enzymes. J. Am. Chem. Soc. 2020, 142, 19493–19501. [Google Scholar] [CrossRef]
- Mandal, S.; Mann, G.; Satish, G.; Brik, A. Enhanced Live-Cell Delivery of Synthetic Proteins Assisted by Cell-Penetrating Peptides Fused to DABCYL. Angew. Chemie Int. Ed. 2021, 60, 7333–7343. [Google Scholar] [CrossRef] [PubMed]

| DUB | Electrophile | Ubiquitin | Kinact/KI (M−1s−1) a,b |
|---|---|---|---|
| UCHL3 | VME | WT-Ub | 1.6 ± 0.07 × 106 |
| UbVT66K/V70F | 1.7 ± 0.4 × 106 | ||
| PRG | WT-Ub | 4.8 ± 0.2 × 106 | |
| UbVT66K/V70F | 4.1 ± 0.4 × 104 | ||
| UCHL1 | VME | WT-Ub | 6.7 ± 0.3 × 103 |
| UbVT66K/V70F | 1.4 ± 0.04 × 102 | ||
| PRG | WT-Ub | 1.3 ± 0.2 × 102 | |
| UbVT66K/V70F | 3.8 ± 4 × 100 |
| DUB | Electrophile | Ubiquitin | Kinact/KI (M−1s−1) a,b |
|---|---|---|---|
| UCHL3 | VME | WT-Ub | 1.6 ± 0.07 × 106 |
| UbVQ40V/T66K/V70F | 2.2 ± 0.8 × 106 | ||
| PRG | WT-Ub | 4.8 ± 0.2 × 106 | |
| UbVQ40V/T66K/V70F | 1.8 ± 0.2 × 105 | ||
| UCHL1 | VME | WT-Ub | 6.7 ± 0.3 × 103 |
| UbVQ40V/T66K/V70F | 5.4 ± 0.3 × 102 | ||
| PRG | WT-Ub | 1.3 ± 0.2 × 102 | |
| UbVQ40V/T66K/V70F | 9.1 ± 1.6 × 100 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hewitt, C.S.; Das, C.; Flaherty, D.P. Rational Development and Characterization of a Ubiquitin Variant with Selectivity for Ubiquitin C-Terminal Hydrolase L3. Biomolecules 2022, 12, 62. https://doi.org/10.3390/biom12010062
Hewitt CS, Das C, Flaherty DP. Rational Development and Characterization of a Ubiquitin Variant with Selectivity for Ubiquitin C-Terminal Hydrolase L3. Biomolecules. 2022; 12(1):62. https://doi.org/10.3390/biom12010062
Chicago/Turabian StyleHewitt, Chad S., Chittaranjan Das, and Daniel P. Flaherty. 2022. "Rational Development and Characterization of a Ubiquitin Variant with Selectivity for Ubiquitin C-Terminal Hydrolase L3" Biomolecules 12, no. 1: 62. https://doi.org/10.3390/biom12010062
APA StyleHewitt, C. S., Das, C., & Flaherty, D. P. (2022). Rational Development and Characterization of a Ubiquitin Variant with Selectivity for Ubiquitin C-Terminal Hydrolase L3. Biomolecules, 12(1), 62. https://doi.org/10.3390/biom12010062






