Protein Intrinsic Disorder and Evolvability of MERS-CoV
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Zaki, A.M.; van Boheemen, S.; Bestebroer, T.M.; Osterhaus, A.D.; Fouchier, R.A. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N. Engl. J. Med. 2012, 367, 1814–1820. [Google Scholar] [CrossRef]
- Al Sulayyim, H.J.; Khorshid, S.M.; Al Moummar, S.H. Demographic, clinical, and outcomes of confirmed cases of Middle East Respiratory Syndrome coronavirus (MERS-CoV) in Najran, Kingdom of Saudi Arabia (KSA); A retrospective record based study. J. Infect. Public Health 2020, 13, 1342–1346. [Google Scholar] [CrossRef] [PubMed]
- Abroug, F.; Slim, A.; Ouanes-Besbes, L.; Hadj Kacem, M.A.; Dachraoui, F.; Ouanes, I.; Lu, X.; Tao, Y.; Paden, C.; Caidi, H.; et al. Family cluster of Middle East respiratory syndrome coronavirus infections, Tunisia, 2013. Emerg. Infect. Dis. 2014, 20, 1527–1530. [Google Scholar] [CrossRef] [PubMed]
- Omrani, A.S.; Matin, M.A.; Haddad, Q.; Al-Nakhli, D.; Memish, Z.A.; Albarrak, A.M. A family cluster of Middle East Respiratory Syndrome Coronavirus infections related to a likely unrecognized asymptomatic or mild case. Int. J. Infect. Dis. 2013, 17, e668–e672. [Google Scholar] [CrossRef]
- Memish, Z.A.; Zumla, A.I.; Al-Hakeem, R.F.; Al-Rabeeah, A.A.; Stephens, G.M. Family cluster of Middle East respiratory syndrome coronavirus infections. N. Engl. J. Med. 2013, 368, 2487–2494. [Google Scholar] [CrossRef] [PubMed]
- Drosten, C.; Gunther, S.; Preiser, W.; van der Werf, S.; Brodt, H.R.; Becker, S.; Rabenau, H.; Panning, M.; Kolesnikova, L.; Fouchier, R.A.; et al. Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N. Engl. J. Med. 2003, 348, 1967–1976. [Google Scholar] [CrossRef]
- Du, L.; He, Y.; Zhou, Y.; Liu, S.; Zheng, B.J.; Jiang, S. The spike protein of SARS-CoV--a target for vaccine and therapeutic development. Nat. Rev. Microbiol. 2009, 7, 226–236. [Google Scholar] [CrossRef]
- Anthony, S.J.; Gilardi, K.; Menachery, V.D.; Goldstein, T.; Ssebide, B.; Mbabazi, R.; Navarrete-Macias, I.; Liang, E.; Wells, H.; Hicks, A.; et al. Further Evidence for Bats as the Evolutionary Source of Middle East Respiratory Syndrome Coronavirus. mBio 2017, 8, e00373-17. [Google Scholar] [CrossRef]
- Memish, Z.A.; Mishra, N.; Olival, K.J.; Fagbo, S.F.; Kapoor, V.; Epstein, J.H.; Alhakeem, R.; Durosinloun, A.; Al Asmari, M.; Islam, A.; et al. Middle East respiratory syndrome coronavirus in bats, Saudi Arabia. Emerg. Infect. Dis. 2013, 19, 1819–1823. [Google Scholar] [CrossRef] [PubMed]
- Lau, S.K.; Li, K.S.; Tsang, A.K.; Lam, C.S.; Ahmed, S.; Chen, H.; Chan, K.H.; Woo, P.C.; Yuen, K.Y. Genetic characterization of Betacoronavirus lineage C viruses in bats reveals marked sequence divergence in the spike protein of pipistrellus bat coronavirus HKU5 in Japanese pipistrelle: Implications for the origin of the novel Middle East respiratory syndrome coronavirus. J. Virol. 2013, 87, 8638–8650. [Google Scholar] [CrossRef]
- Reusken, C.B.; Haagmans, B.L.; Muller, M.A.; Gutierrez, C.; Godeke, G.J.; Meyer, B.; Muth, D.; Raj, V.S.; Smits-De Vries, L.; Corman, V.M.; et al. Middle East respiratory syndrome coronavirus neutralising serum antibodies in dromedary camels: A comparative serological study. Lancet Infect. Dis. 2013, 13, 859–866. [Google Scholar] [CrossRef]
- Perera, R.A.; Wang, P.; Gomaa, M.R.; El-Shesheny, R.; Kandeil, A.; Bagato, O.; Siu, L.Y.; Shehata, M.M.; Kayed, A.S.; Moatasim, Y.; et al. Seroepidemiology for MERS coronavirus using microneutralisation and pseudoparticle virus neutralisation assays reveal a high prevalence of antibody in dromedary camels in Egypt, June 2013. Euro Surveill. 2013, 18, pii=20574. [Google Scholar] [CrossRef] [PubMed]
- Haagmans, B.L.; Al Dhahiry, S.H.; Reusken, C.B.; Raj, V.S.; Galiano, M.; Myers, R.; Godeke, G.J.; Jonges, M.; Farag, E.; Diab, A.; et al. Middle East respiratory syndrome coronavirus in dromedary camels: An outbreak investigation. Lancet Infect. Dis. 2014, 14, 140–145. [Google Scholar] [CrossRef]
- Li, Y.H.; Hu, C.Y.; Wu, N.P.; Yao, H.P.; Li, L.J. Molecular Characteristics, Functions, and Related Pathogenicity of MERS-CoV Proteins. Engineering 2019, 5, 940–947. [Google Scholar] [CrossRef] [PubMed]
- Jiaming, L.; Yanfeng, Y.; Yao, D.; Yawei, H.; Linlin, B.; Baoying, H.; Jinghua, Y.; Gao, G.F.; Chuan, Q.; Wenjie, T. The recombinant N-terminal domain of spike proteins is a potential vaccine against Middle East respiratory syndrome coronavirus (MERS-CoV) infection. Vaccine 2017, 35, 10–18. [Google Scholar] [CrossRef]
- Perrier, A.; Bonnin, A.; Desmarets, L.; Danneels, A.; Goffard, A.; Rouille, Y.; Dubuisson, J.; Belouzard, S. The C-terminal domain of the MERS coronavirus M protein contains a trans-Golgi network localization signal. J. Biol. Chem. 2019, 294, 14406–14421. [Google Scholar] [CrossRef] [PubMed]
- Alshehri, M.A.; Manee, M.M.; Alqahtani, F.H.; Al-Shomrani, B.M.; Uversky, V.N. On the Prevalence and Potential Functionality of an Intrinsic Disorder in the MERS-CoV Proteome. Viruses 2021, 13, 339. [Google Scholar] [CrossRef]
- Goh, G.K.; Dunker, A.K.; Uversky, V. Prediction of Intrinsic Disorder in MERS-CoV/HCoV-EMC Supports a High Oral-Fecal Transmission. PLoS Curr. 2013, 5. [Google Scholar] [CrossRef]
- Hatcher, E.L.; Zhdanov, S.A.; Bao, Y.; Blinkova, O.; Nawrocki, E.P.; Ostapchuck, Y.; Schaffer, A.A.; Brister, J.R. Virus Variation Resource - improved response to emergent viral outbreaks. Nucleic Acids Res. 2017, 45, D482–D490. [Google Scholar] [CrossRef]
- Hamers-Casterman, C.; Atarhouch, T.; Muyldermans, S.; Robinson, G.; Hamers, C.; Songa, E.B.; Bendahman, N.; Hamers, R. Naturally occurring antibodies devoid of light chains. Nature 1993, 363, 446–448. [Google Scholar] [CrossRef]
- Dumoulin, M.; Conrath, K.; Van Meirhaeghe, A.; Meersman, F.; Heremans, K.; Frenken, L.G.; Muyldermans, S.; Wyns, L.; Matagne, A. Single-domain antibody fragments with high conformational stability. Protein Sci. 2002, 11, 500–515. [Google Scholar] [CrossRef] [PubMed]
- Ebrahimizadeh, W.; Mousavi Gargari, S.; Rajabibazl, M.; Safaee Ardekani, L.; Zare, H.; Bakherad, H. Isolation and characterization of protective anti-LPS nanobody against V. cholerae O1 recognizing Inaba and Ogawa serotypes. Appl. Microbiol. Biotechnol. 2013, 97, 4457–4466. [Google Scholar] [CrossRef] [PubMed]
- De Vos, J.; Devoogdt, N.; Lahoutte, T.; Muyldermans, S. Camelid single-domain antibody-fragment engineering for (pre)clinical in vivo molecular imaging applications: Adjusting the bullet to its target. Expert Opin. Biol. Ther. 2013, 13, 1149–1160. [Google Scholar] [CrossRef] [PubMed]
- de Marco, A. Biotechnological applications of recombinant single-domain antibody fragments. Microb. Cell Fact. 2011, 10, 44. [Google Scholar] [CrossRef]
- Obradovic, Z.; Peng, K.; Vucetic, S.; Radivojac, P.; Dunker, A.K. Exploiting heterogeneous sequence properties improves prediction of protein disorder. Proteins 2005, 61 (Suppl. S7), 176–182. [Google Scholar] [CrossRef]
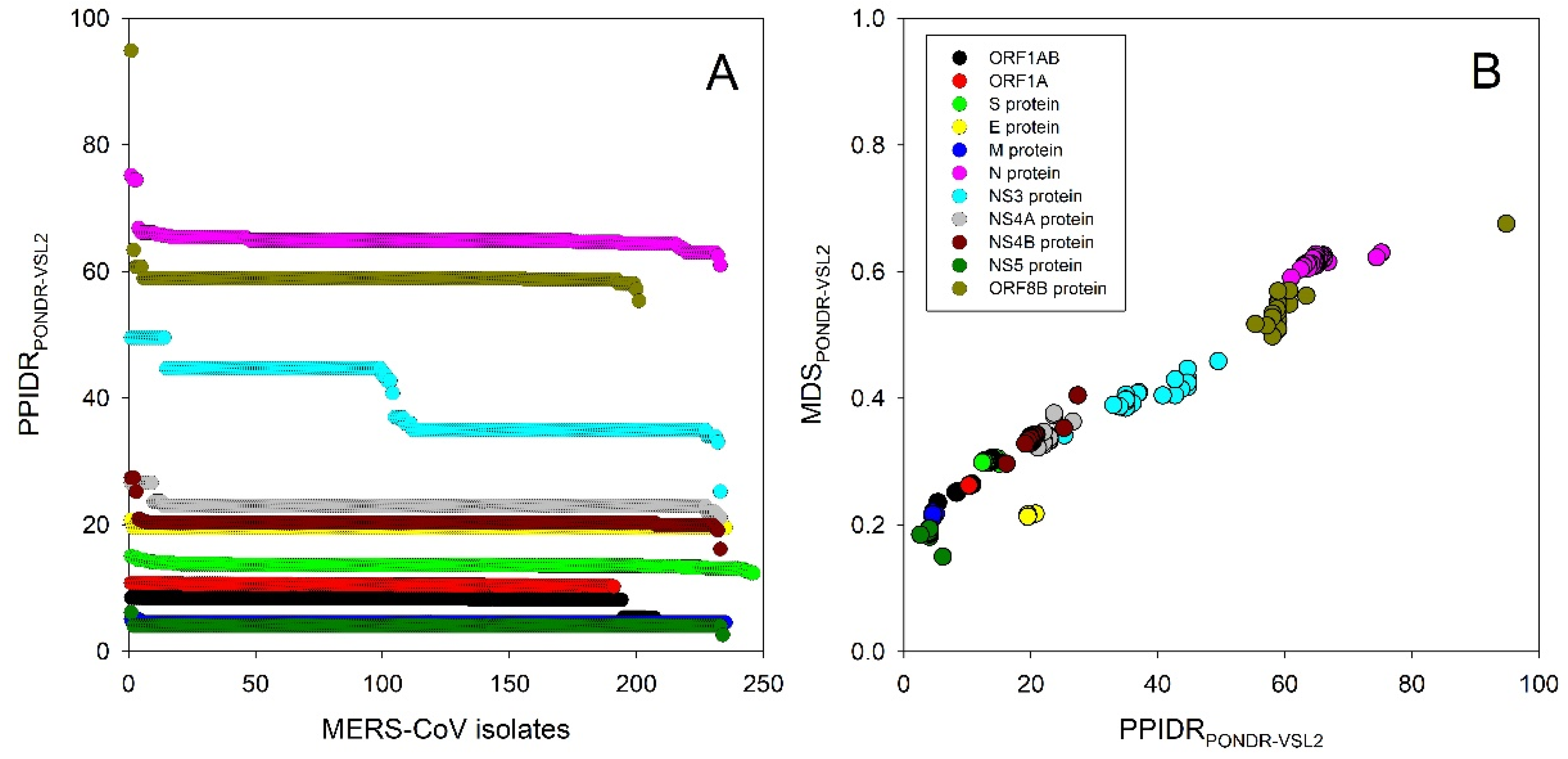
| Protein | ID | PPIDR | Time | Protein | ID | PPIDR | Time |
|---|---|---|---|---|---|---|---|
| ORF1AB | AGV08406 | 8.42% | 2012/06/19 | NS3 | AGV08409 | 44.66% | 2012/06/19 |
| AIZ74438 | 5.30% | 2013/05/07 | AHE78109 | 33.01% | 2013/11/05 | ||
| AFS88944 | 8.48% | 2012/06/13 | AGN70930 | 49.52% | 2013/05/01 | ||
| ORF1A | AGV08407 | 10.66% | 2012/06/19 | NS4A | AGV08410 | 22.94% | 2012/06/19 |
| AIL23988 | 10.25% | 2014/04/22 | AMQ49006 | 21.10% | 2014/11/04 | ||
| AWM99581 | 10.79% | 2016/08/01 | AKN24778 | 26.61% | 2014/05/12 | ||
| S protein | AGV08408 | 13.52% | 2012/06/19 | NS4B | AGV08411 | 27.39% | 2012/06/19 |
| AHC74088 | 12.34% | 2013/10/13 | AIZ74435 | 16.15% | 2013/05/07 | ||
| AMQ49004 | 14.98% | 2014/11/04 | AGV08382 | 27.39% | 2012/10/23 | ||
| E protein | AGV08413 | 19.51% | 2012/06/19 | NS5 | AGV08412 | 4.02% | 2012/06/19 |
| AIL23939 | 19.51% | 2014/04/07 | AIZ74454 | 2.56% | 2013/05/07 | ||
| AGV08472 | 20.73% | 2013/05/13 | AMQ49019 | 6.12% | 2015/07/12 | ||
| M protein | AGV08414 | 4.57% | 2012/06/19 | ORF8B | AGV08416 | 58.93% | 2012/06/19 |
| AKK52618 | 4.57% | 2015/03/01 | AXN92235 | 53.36% | 2018/05/05 | ||
| ARQ84741 | 5.02% | 2015/08/25 | AIL23997 | 94.87% | 2014/04/22 | ||
| N protein | AGV08415 | 64.89% | 2012/06/19 | ||||
| AIZ74430 | 60.99% | 2013/05/07 | |||||
| AHI48709 | 75.18% | 2013/08/06 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Uversky, V.N.; Redwan, E.M.; Aljadawi, A.A. Protein Intrinsic Disorder and Evolvability of MERS-CoV. Biomolecules 2021, 11, 608. https://doi.org/10.3390/biom11040608
Uversky VN, Redwan EM, Aljadawi AA. Protein Intrinsic Disorder and Evolvability of MERS-CoV. Biomolecules. 2021; 11(4):608. https://doi.org/10.3390/biom11040608
Chicago/Turabian StyleUversky, Vladimir N., Elrashdy M. Redwan, and Abdullah A. Aljadawi. 2021. "Protein Intrinsic Disorder and Evolvability of MERS-CoV" Biomolecules 11, no. 4: 608. https://doi.org/10.3390/biom11040608
APA StyleUversky, V. N., Redwan, E. M., & Aljadawi, A. A. (2021). Protein Intrinsic Disorder and Evolvability of MERS-CoV. Biomolecules, 11(4), 608. https://doi.org/10.3390/biom11040608






