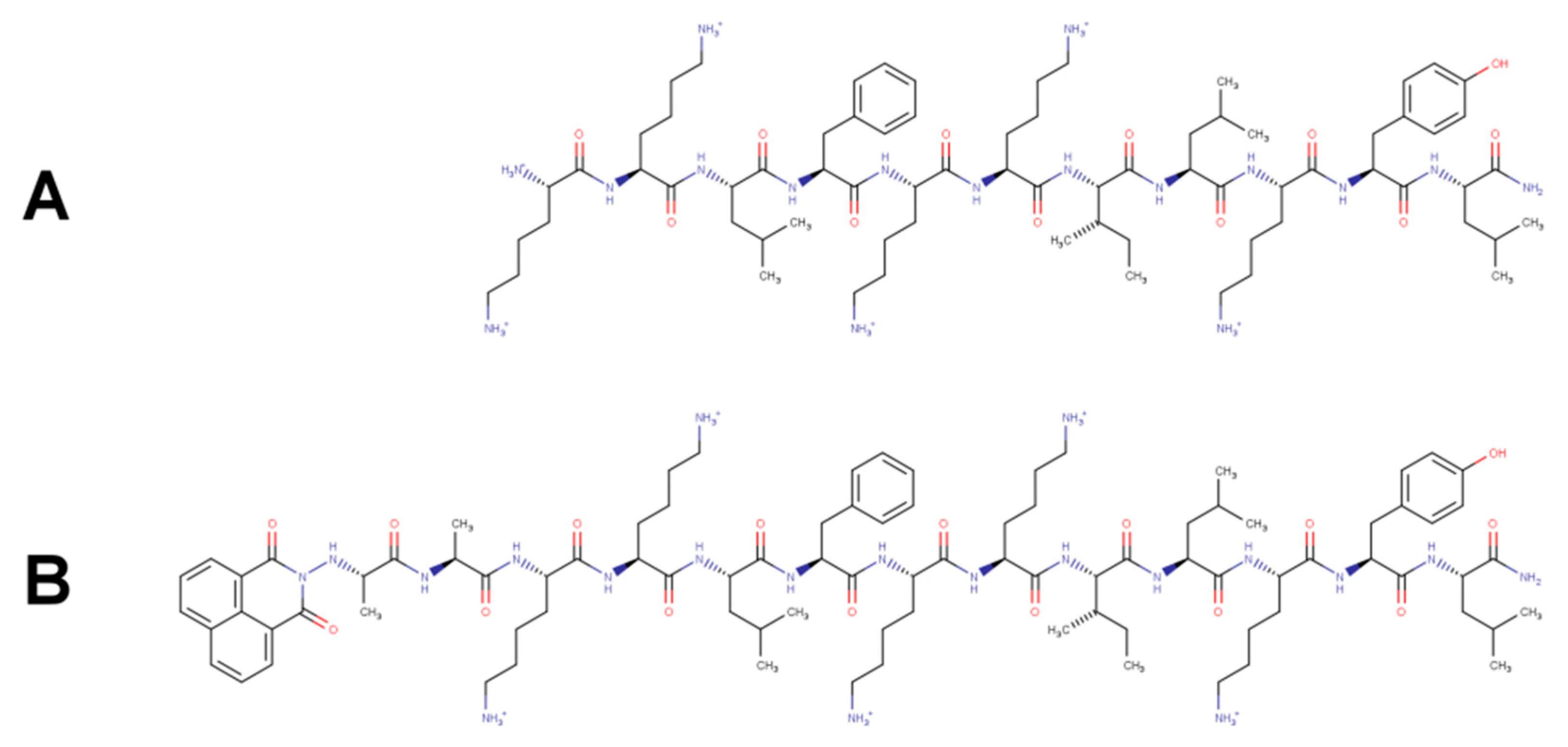
Naphthalimide-Containing BP100 Leads to Higher Model Membranes Interactions and Antimicrobial Activity
Abstract
1. Introduction
2. Materials and Methods
2.1. Reagents
2.2. Peptide Synthesis
2.3. Peptide and ds-DNA Solutions
2.4. Model Membrane Preparation
2.5. Fluorescence
2.6. Circular Dichroism
2.7. Dynamic Light Scattering (DLS)
2.8. Isothermal Titration Calorimetry (ITC)
2.9. Differential Scanning Calorimetry
2.10. Model Membrane Permeabilization
2.11. Minimum Inhibitory Concentration Assay
2.12. Hemolytic Activity
3. Results
3.1. NAPHT-BP100 Membrane Interaction—Binding Extent and Secondary Structure
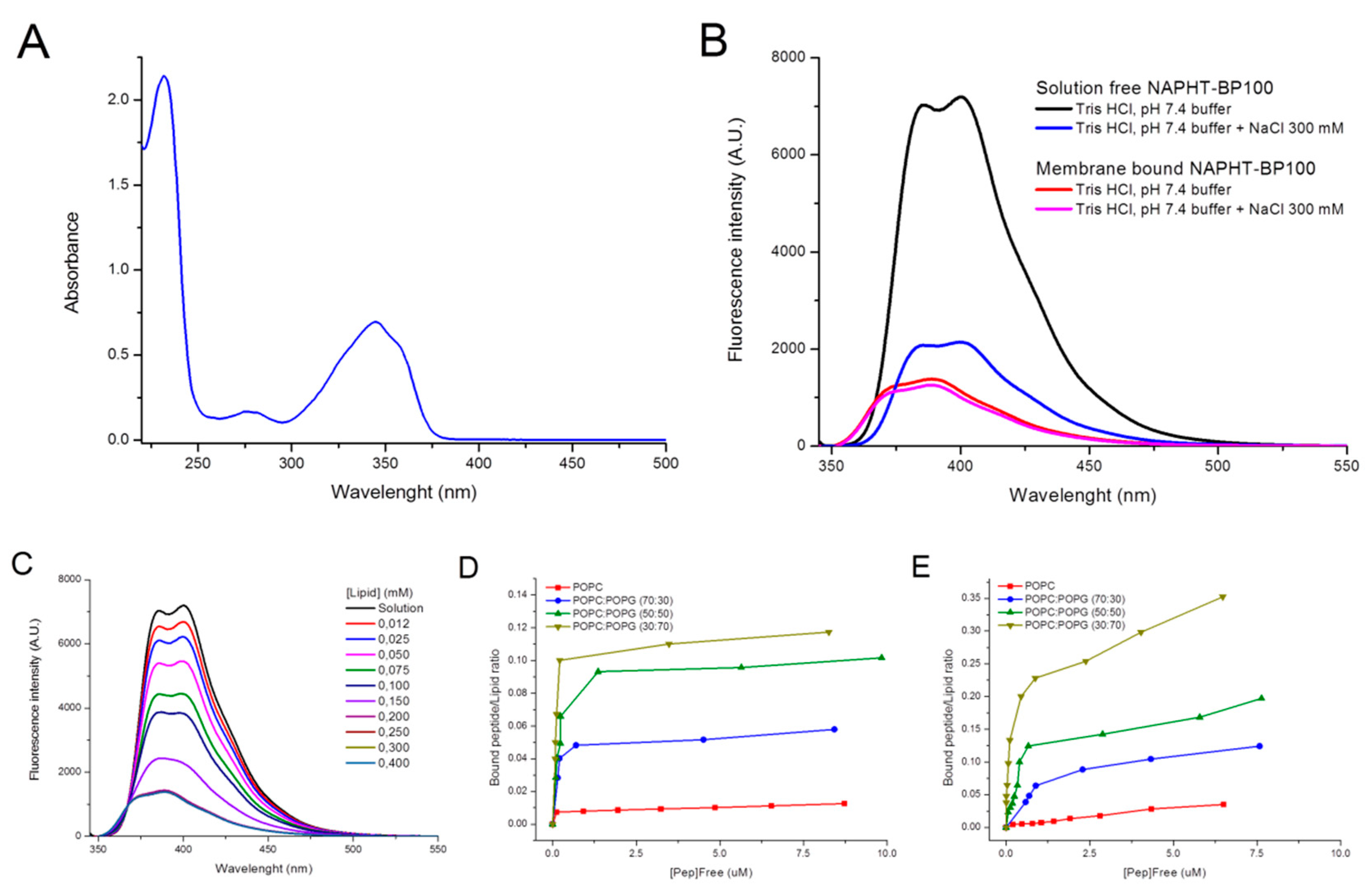
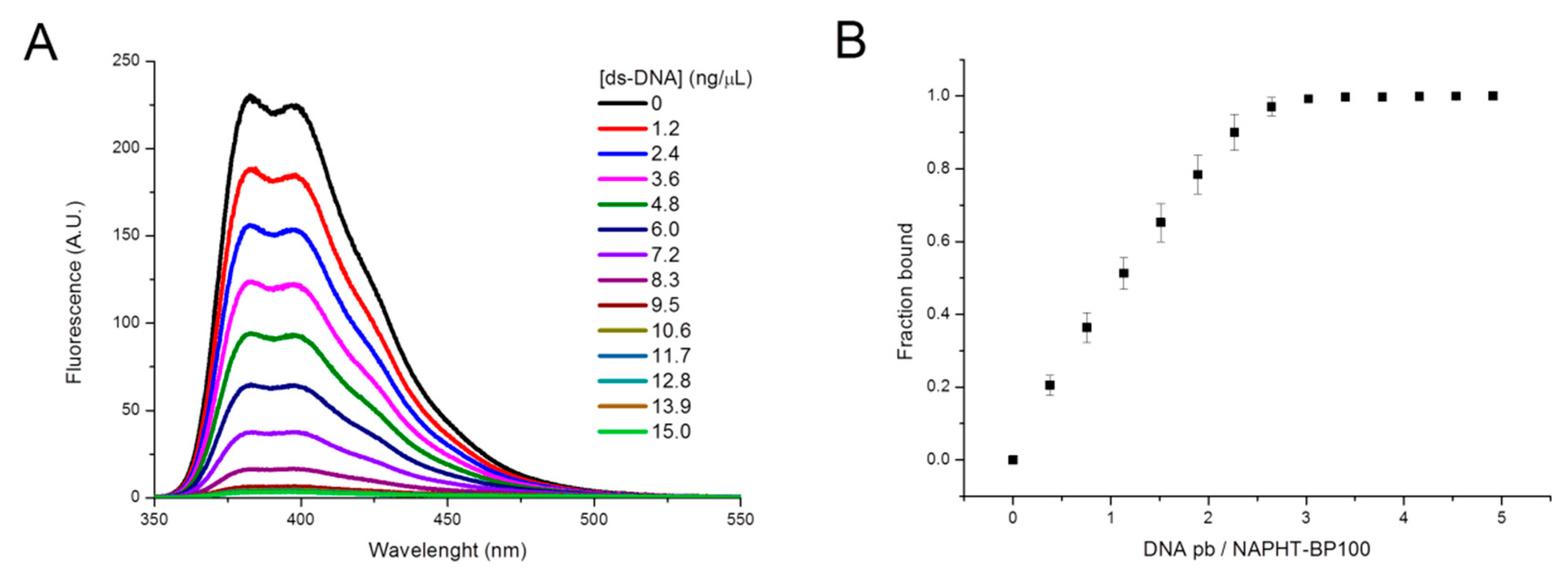
3.1.1. UV-Absorption and Fluorescence—NAPHT-BP100 Membrane and ds-DNA Binding
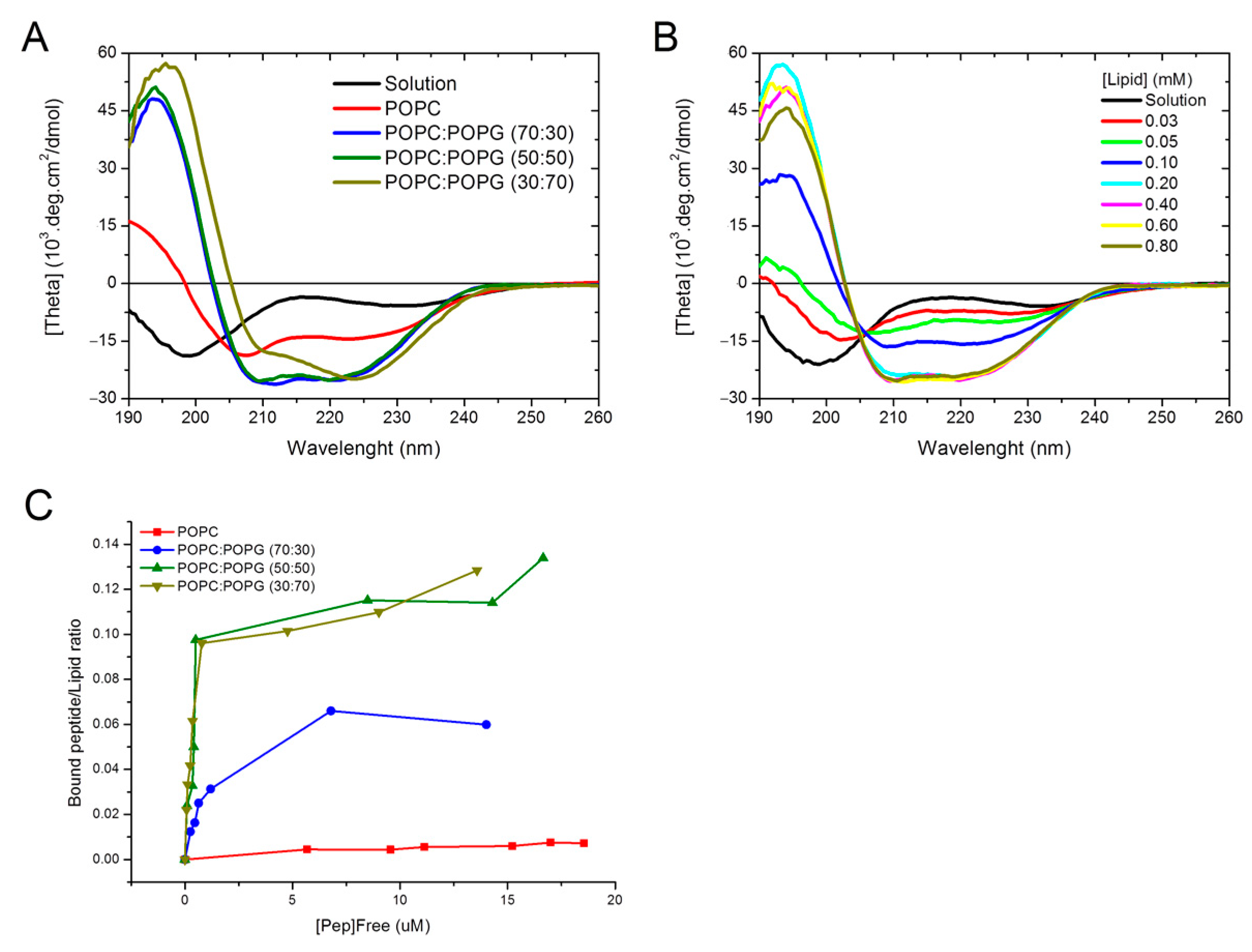
3.1.2. Circular Dichroism—Secondary Structure Analysis
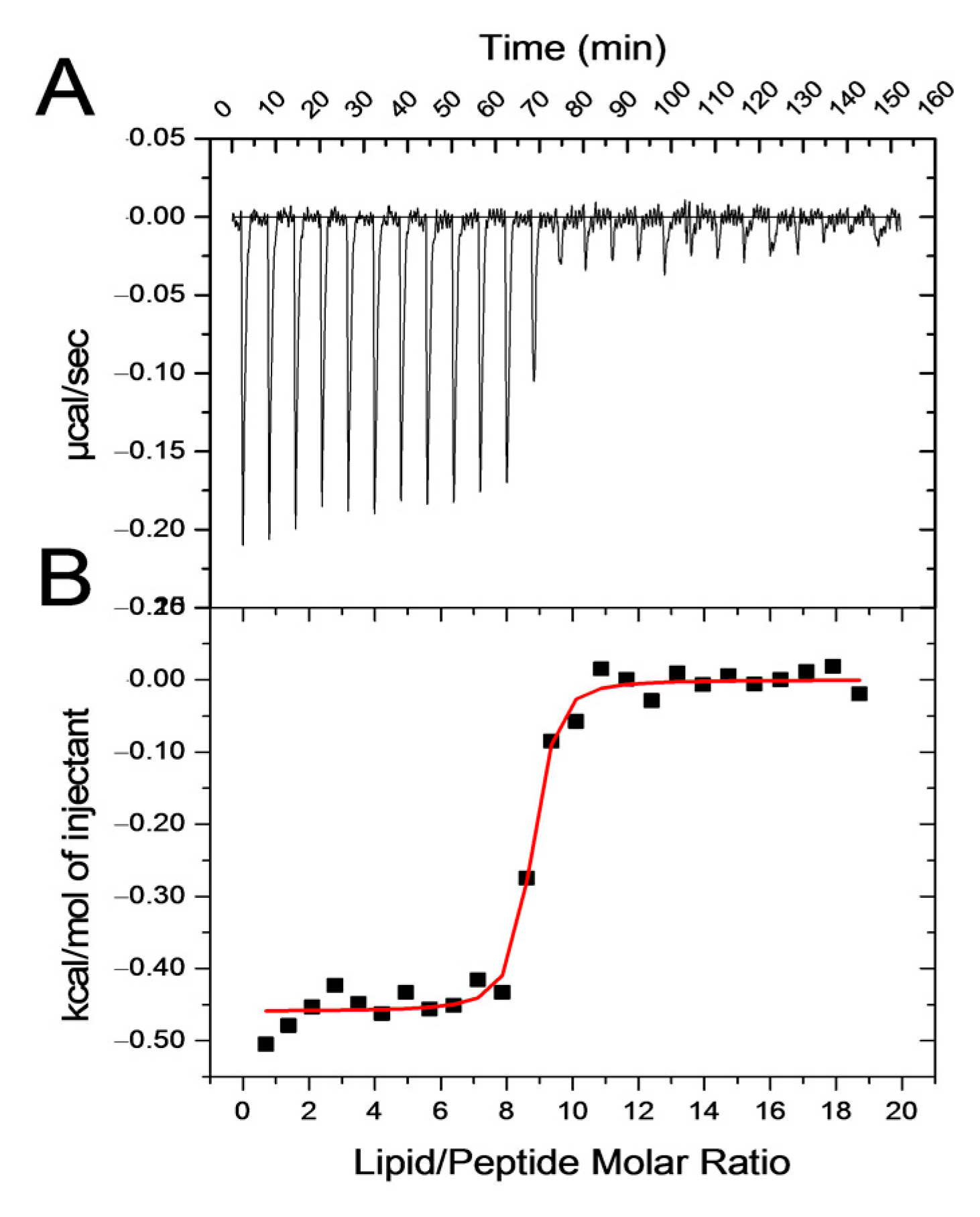
3.1.3. Isothermal Titration Calorimetry—Interaction Thermodynamics
3.2. NAPHT-BP100 Effect Over the Membrane—Action Mechanism
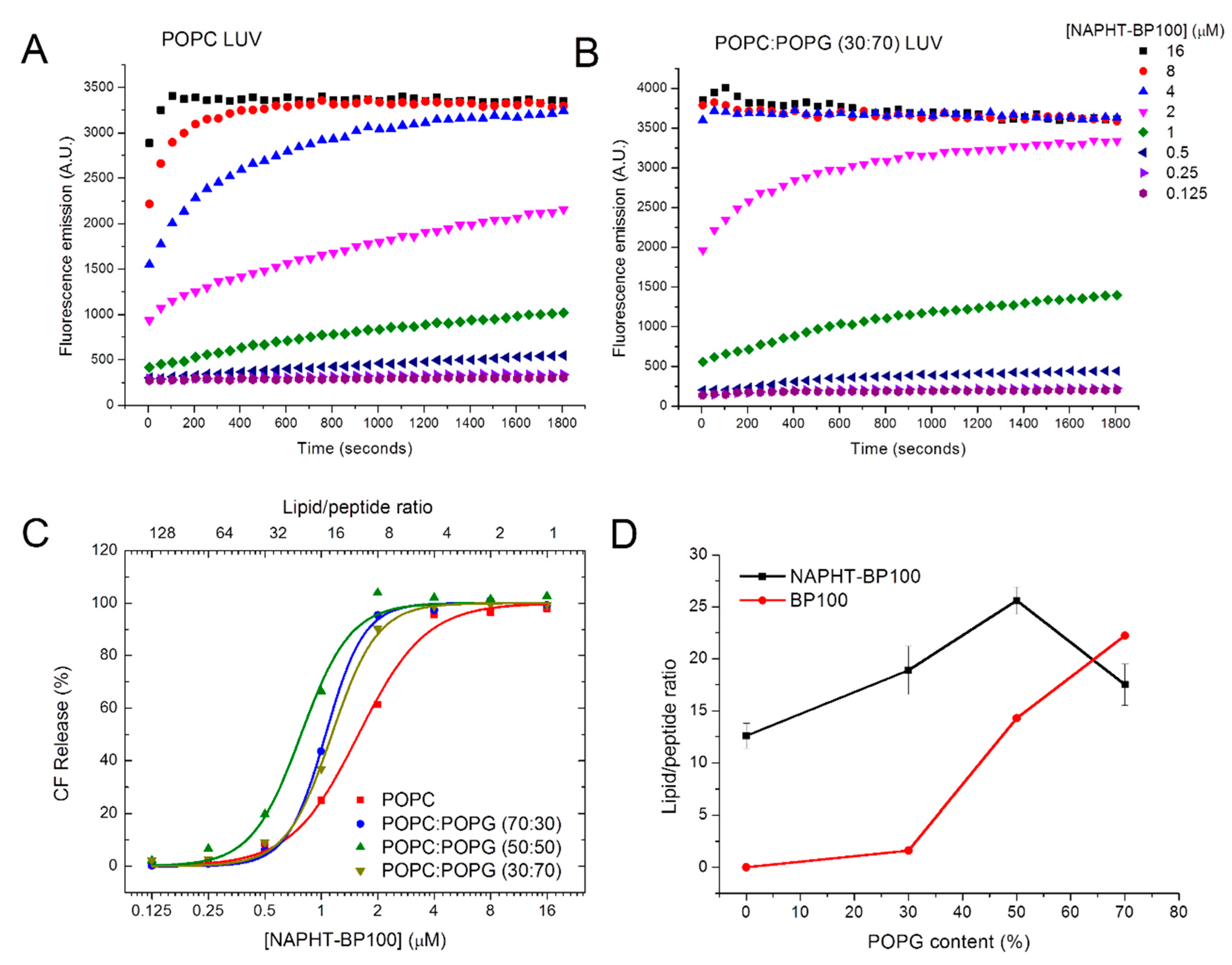
3.2.1. Carboxyfluorescein Leakage—Peptide Permeabilizing Activity
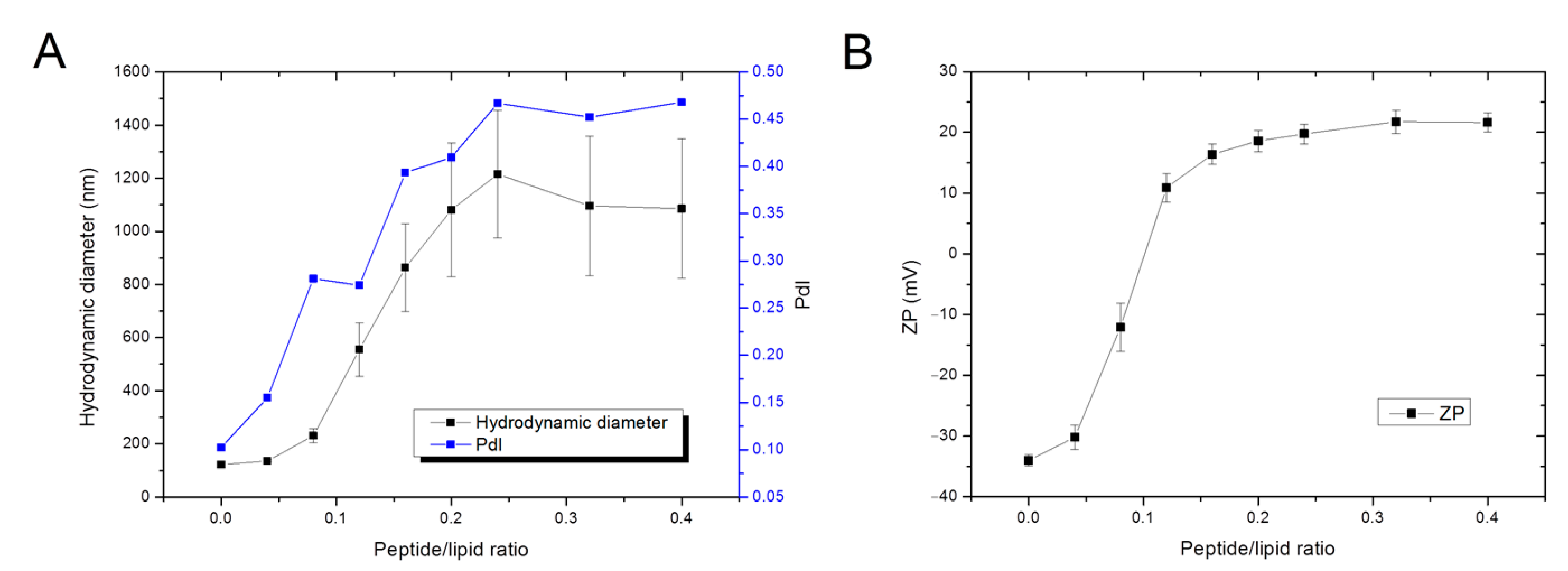
3.2.2. Dynamic Light Scattering—Peptide Effect over Liposome’s Size and Surface Charge
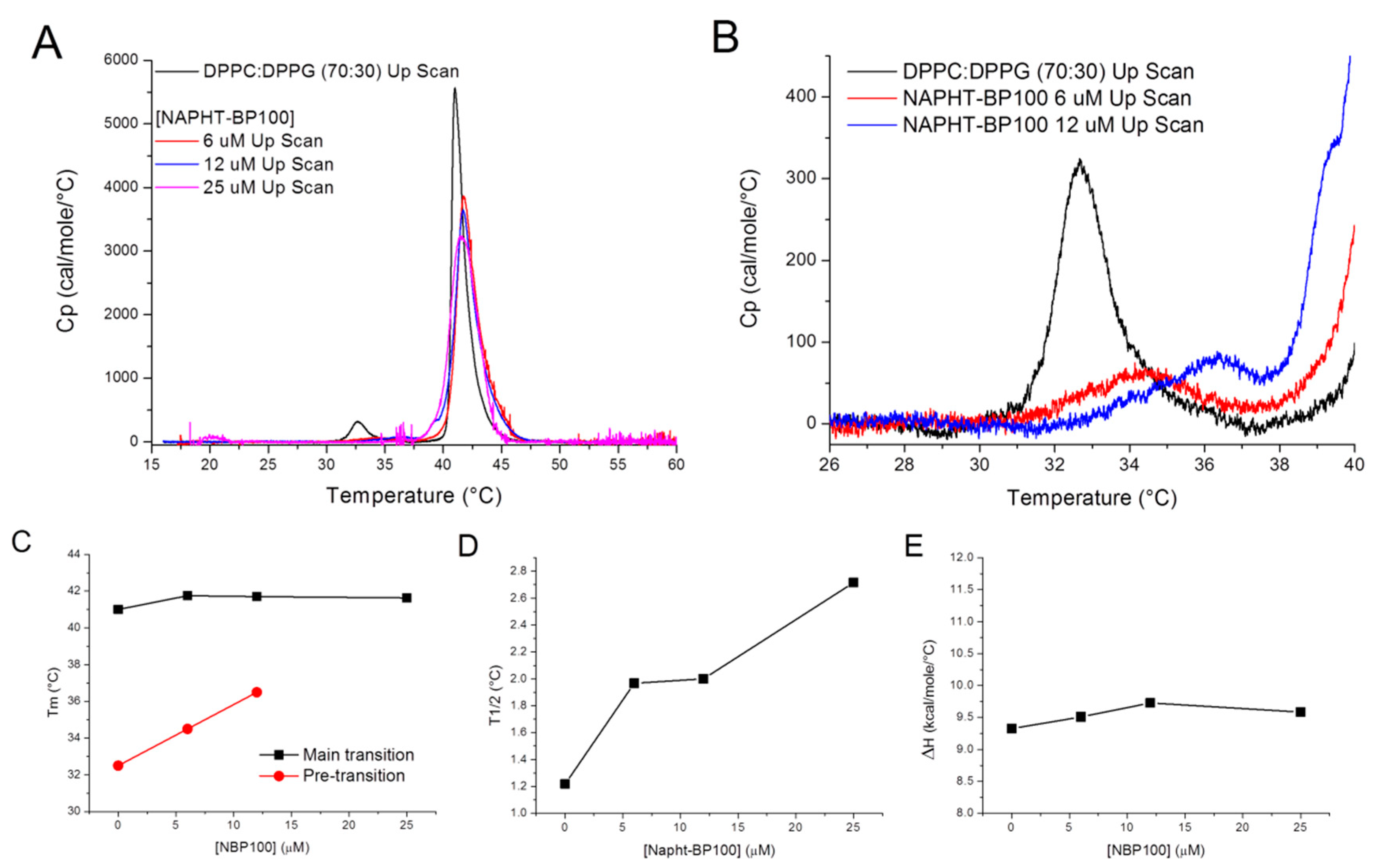
3.2.3. Differential Scanning Calorimetry—Peptide Effect Over Lipid Packing
3.3. Biological Activity
3.3.1. Minimum Inhibitory Concentration
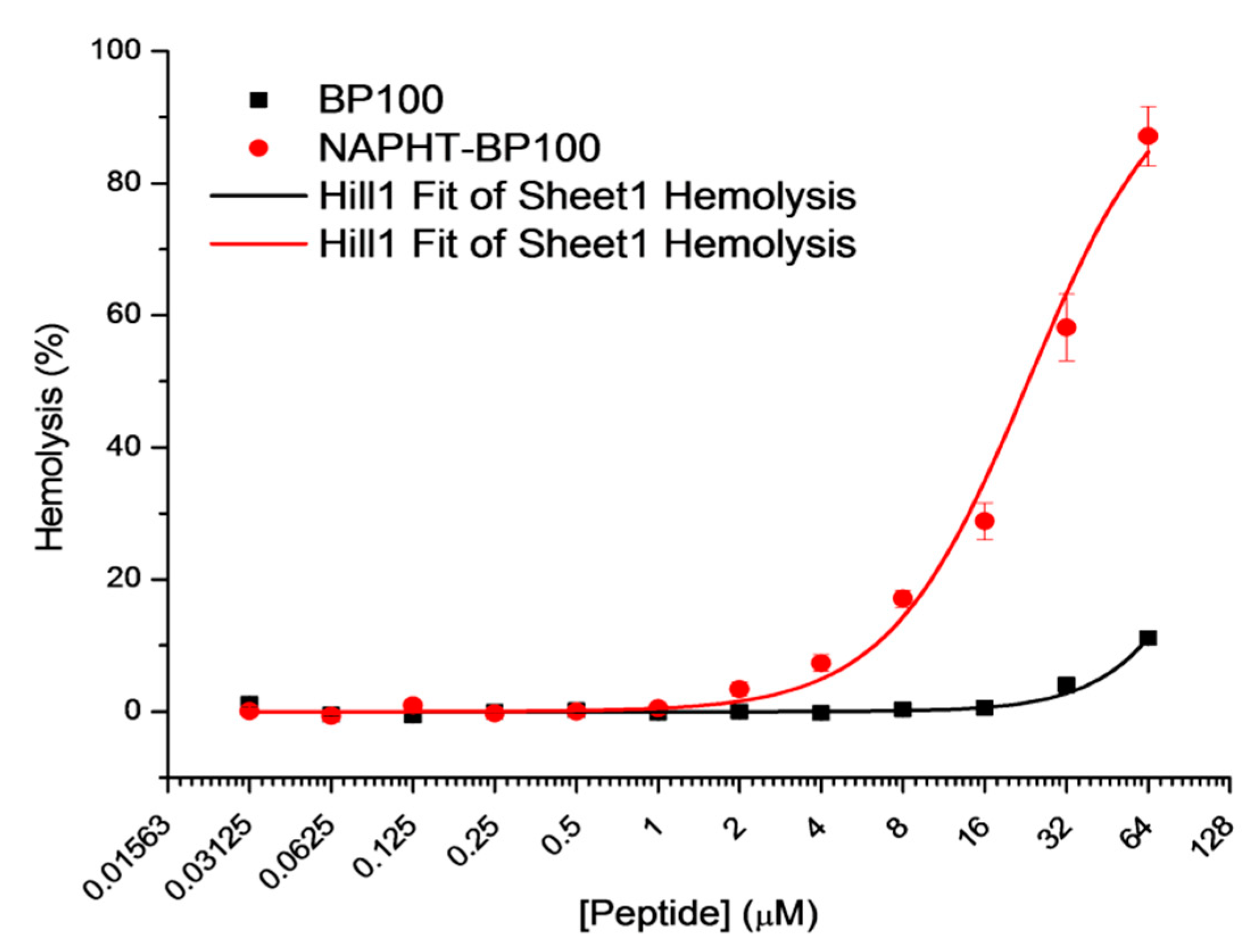
3.3.2. Hemolytic Activity
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Chen, C.H.; Lu, T.K. Development and Challenges of Antimicrobial Peptides for Therapeutic Applications. Antibiotics 2020, 9, 24. [Google Scholar] [CrossRef] [PubMed]
- Lohner, K. Membrane-active antimicrobial peptides as template structures for novel antibiotic agents. Curr. Top. Med. Chem. 2017, 17, 508–519. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Koh, J.J.; Liu, S.; Lakshminarayanan, R.; Verma, C.S.; Beuerman, R.W. Membrane active antimicrobial peptides: Translating mechanistic insights to design. Front. Neurosci. 2017, 11, 73. [Google Scholar] [CrossRef] [PubMed]
- Jenssen, H.; Hamill, P.; Hancock, R.E.W. Peptide Antimicrobial Agents. Clin. Microbiol. Rev. 2006, 19, 491–511. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, L.; Haney, E.; Vogel, H. The expanding scope of antimicrobial peptide structures and their modes of action. Trends Biotechnol. 2011, 29, 464–472. [Google Scholar] [CrossRef]
- Powers, J.; Hancock, R. The relationship between peptide structure and antibacterial activity. Peptides 2003, 24, 1681–1691. [Google Scholar] [CrossRef]
- Yeaman, M.R.; Yount, N.Y.; Hauger, R.L.; Grigoriadis, D.E.; Dallman, M.F.; Plotsky, P.M.; Vale, W.W.; Dautzenberg, F.M. Mechanisms of Antimicrobial Peptide Action and Resistance. Pharmacol. Rev. 2003, 55, 27–55. [Google Scholar] [CrossRef]
- Park, P.; Franco, L.R.; Chaimovich, H.; Coutinho, K.; Cuccovia, I.M.; Lima, F.S. Binding and Flip as Initial Steps for BP-100 Antimicrobial Actions. Sci. Rep. 2019, 9, 8622. [Google Scholar] [CrossRef]
- Killian, J.; von Heijne, G. How proteins adapt to a membrane-water interface. Trends Biochem. Sci. 2000, 25, 429–434. [Google Scholar] [CrossRef]
- Lima, F.S.; Andrade, M.F.C.; Mortara, L.; Dias, L.G.; Cuccovia, I.M.; Chaimovich, H. Ion dehydration controls adsorption at the micellar interface: Hydrotropic ions. Phys. Chem. Chem. Phys. 2017, 19, 30658–30666. [Google Scholar] [CrossRef]
- Badosa, E.; Ferre, R.; Planas, M.; Feliu, L.; Besalú, E.; Cabrefiga, J.; Bardají, E.; Montesinos, E. A library of linear undecapeptides with bactericidal activity against phytopathogenic bacteria. Peptides 2007, 28, 2276–2285. [Google Scholar] [CrossRef] [PubMed]
- Ferre, R.; Melo, M.N.; Correia, A.D.; Feliu, L.; Bardají, E.; Planas, M.; Castanho, M. Synergistic Effects of the Membrane Actions of Cecropin-Melittin Antimicrobial Hybrid Peptide BP100. Biophys. J. 2009, 96, 1815–1827. [Google Scholar] [CrossRef]
- Alves, C.S.; Melo, M.N.; Franquelim, H.G.; Ferre, R.; Planas, M.; Feliu, L.; Bardají, E.; Kowalczyk, W.; Andreu, D.; Santos, N.C.; et al. Escherichia coli Cell Surface Perturbation and Disruption Induced by Antimicrobial Peptides BP100 and pepR. J. Biol. Chem. 2010, 285, 27536–27544. [Google Scholar] [CrossRef]
- Torcato, I.M.; Huang, Y.-H.; Franquelim, H.G.; Gaspar, D.; Craik, D.J.; Castanho, M.A.; Henriques, S.T. Design and characterization of novel antimicrobial peptides, R-BP100 and RW-BP100, with activity against Gram-negative and Gram-positive bacteria. Biochim. Biophys. Acta Biomembr. 2013, 1828, 944–955. [Google Scholar] [CrossRef] [PubMed]
- Manzini, M.C.; Perez, K.R.; Riske, K.A.; Bozelli, J.C.; Santos, T.L.; Da Silva, M.A.; Saraiva, G.K.; Politi, M.J.; Valente, A.P.; Almeida, F.C.; et al. Peptide:lipid ratio and membrane surface charge determine the mechanism of action of the antimicrobial peptide BP100. Conformational and functional studies. Biochim. Biophys. Acta Biomembr. 2014, 1838, 1985–1999. [Google Scholar] [CrossRef] [PubMed]
- Wadhwani, P.; Strandberg, E.; Berg, J.V.D.; Mink, C.; Bürck, J.; Ciriello, R.A.; Ulrich, A.S. Dynamical structure of the short multifunctional peptide BP100 in membranes. Biochim. Biophys. Acta Biomembr. 2014, 1838, 940–949. [Google Scholar] [CrossRef]
- Carretero, G.P.; Saraiva, G.K.; Cauz, A.C.; Rodrigues, M.A.; Kiyota, S.; Riske, K.A.; Dos Santos, A.A.; Pinatto-Botelho, M.F.; Bemquerer, M.P.; Gueiros-Filho, F.J.; et al. Synthesis, biophysical and functional studies of two BP100 analogues modified by a hydrophobic chain and a cyclic peptide. Biochim. Biophys. Acta Biomembr. 2018, 1860, 1502–1516. [Google Scholar] [CrossRef]
- Zhang, B.; Gu, H.; Shi, W.; Li, H.; Ma, G.; Chen, X.; Qian, H.; Lin, H.; Huang, W.; Ge, L. Synthesis and biological evaluation of novel aliphatic acid-conjugated antimicrobial peptides as potential agents with anti-tumor, multidrug resistance-reversing activity and enhanced stability. Amino Acids 2017, 49, 1831–1841. [Google Scholar] [CrossRef]
- Kamal, A.; Bolla, N.R.; Srikanth, P.S.; Srivastava, A.K. Naphthalimide derivatives with therapeutic characteristics: A patent review. Expert Opin. Ther. Patents 2013, 23, 299–317. [Google Scholar] [CrossRef]
- Braña, M.F.; Ramos, A. Naphthalimides as anti-cancer agents: Synthesis and biological activity. Curr. Med. Chem. Anti. Cancer Agents 2001, 1, 237–255. [Google Scholar] [CrossRef]
- Min, L.; Hui, X. Overview of Naphthalimide Analogs as Anticancer Agents. Curr. Med. Chem. 2009, 16, 4797–4813. [Google Scholar]
- Ingrassia, L.; Lefranc, F.; Kiss, R.; Mijatovic, T. Naphthalimides and azonafides as promising anti-cancer agents. Curr. Med. Chem. 2009, 16, 1192–1213. [Google Scholar] [CrossRef]
- Tandon, R.; Luxami, V.; Kaur, H.; Tandon, N.; Paul, K. 1,8-Naphthalimide: A Potent DNA Intercalator and Target for Cancer Therapy. Chem. Rec. 2017, 17, 956–993. [Google Scholar] [CrossRef] [PubMed]
- Kang, J.; Gopala, L.; Reddy-Tangadanchu, V.; Gao, W.; Zhou, C.H. Novel naphthalimide nitroimidazoles as multitargeting antibacterial agents against resistant Acinetobacter baumanni. Future Med. Chem. 2018, 10, 711–724. [Google Scholar] [CrossRef]
- Marinov, M.; Kostova, I.; Naydenova, E.; Stoyanov, N. Synthesis and antimicrobial activity of 1,8-naphthalimide derivatives of nalidixic acid. J. Chem. Technol. Metal. 2019, 54, 1146–1156. [Google Scholar]
- Shaki, H.; Khosravi, A.; Gharanjig, J.; Mahboubi, A. Investigation of synthesis, characterization, photophysical and biological properties of novel antimicrobial fluorescent naphthalimide derivatives. Mater. Technol. 2016, 31, 322–331. [Google Scholar] [CrossRef]
- Banerjee, S.; Veale, E.B.; Phelan, C.M.; Murphy, S.A.; Tocci, G.M.; Gillespie, L.J.; Frimannsson, D.O.; Kelly, J.M.; Gunnlaugsson, T. Recent advances in the development of 1,8-naphthalimide based DNA targeting binders, anticancer and fluorescent cellular imaging agents. Chem. Soc. Rev. 2013, 42, 1601–1618. [Google Scholar] [CrossRef]
- Hickey, S.M.; Ashton, T.D.; Boer, G.; Bader, C.A.; Thomas, M.; Elliott, A.G.; Schmuck, C.; Yu, H.Y.; Li, J.; Nation, R.L.; et al. Norbornane-based cationic antimicrobial peptidomimetics targeting the bacterial membrane. Eur. J. Med. Chem. 2018, 160, 9–22. [Google Scholar] [CrossRef]
- Wu, A.; Xu, Y.; Qian, X. Novel naphthalimide–amino acid conjugates with flexible leucine moiety as side chain: Design, synthesis and potential antitumor activity. Bioorg. Med. Chem. 2009, 17, 592–599. [Google Scholar] [CrossRef]
- Chan, W.; White, P. FMOC Solid Phase Peptide Synthesis; Oxford University Press: Oxford, UK, 2000. [Google Scholar]
- Friedman, M. Applications of the Ninhydrin Reaction for Analysis of Amino Acids, Peptides, and Proteins to Agricultural and Biomedical Sciences. J. Agric. Food Chem. 2004, 52, 385–406. [Google Scholar] [CrossRef]
- Abraham, B.; Kelly, L.A. Photooxidation of Amino Acids and Proteins Mediated by Novel 1,8-Naphthalimide Derivatives. J. Phys. Chem. B 2003, 107, 12534–12541. [Google Scholar] [CrossRef]
- Aveline, B.; Matsugo, S.; Redmond, R. Photochemical Mechanisms Responsible for the Versatile Application of Naphthalimides and Naphthaldiimides in Biological Systems. J. Am. Chem. Soc. 1997, 119, 11785–11795. [Google Scholar] [CrossRef]
- Shibue, M.; Mant, C.; Hodges, R. Effect of anionic ion-pairing reagent concentration (1–60 mm) on reversed-phase liquid chromatography elution behaviour of peptides. J. Chromatogr. A 2005, 1080, 58–67. [Google Scholar] [CrossRef]
- Rouser, G.; Fleischer, S.; Yamamoto, A. Two dimensional thin layer chromatographic separation of polar lipids and determination of phospholipids by phosphorus analysis of spots. Lipids 1970, 5, 494–496. [Google Scholar] [CrossRef]
- Beschiaschvili, G.; Seelig, J. Peptide binding to lipid bilayers. Binding isotherms and zeta-potential of a cyclic somatostatin analog. Biochemistry 1990, 29, 10995–11000. [Google Scholar] [CrossRef] [PubMed]
- Dathe, M.; Schümann, M.; Wieprecht, T.; Winkler, A.; Beyermann, M.; Krause, E.; Matsuzaki, K.; Murase, O.; Bienert, M. Peptide Helicity and Membrane Surface Charge Modulate the Balance of Electrostatic and Hydrophobic Interactions with Lipid Bilayers and Biological Membranes. Biochemistry 1996, 35, 12612–12622. [Google Scholar] [CrossRef]
- Domingues, T.M.; Mattei, B.; Seelig, J.; Perez, K.R.; Miranda, A.; Riske, K.A. Interaction of the Antimicrobial Peptide Gomesin with Model Membranes: A Calorimetric Study. Langmuir 2013, 29, 8609–8618. [Google Scholar] [CrossRef]
- Wiegand, I.; Hilpert, K.; Hancock, R.E.W. Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat. Protoc. 2008, 3, 163–175. [Google Scholar] [CrossRef]
- Mojsoska, B.; Carretero, G.; Larsen, S.; Mateiu, R.V.; Jenssen, H. Peptoids successfully inhibit the growth of gram negative E. coli causing substantial membrane damage. Sci. Rep. 2017, 7, srep42332. [Google Scholar] [CrossRef]
- Magalhães, J.; Pereira, R.; Triboni, E.; Filho, P.B.; Gehlen, M.; Nart, F. Solvent effect on the photophysical properties of 4-phenoxy-N-methyl-1,8-naphthalimide. J. Photochem. Photobiol. A Chem. 2006, 183, 165–170. [Google Scholar] [CrossRef]
- Banerjee, S.; Kitchen, J.A.; Gunnlaugsson, T.; Kelly, J.M. Synthesis and photophysical evaluation of a pyridinium 4-amino-1,8-naphthalimide derivative that upon intercalation displays preference for AT-rich double-stranded DNA. Org. Biomol. Chem. 2012, 10, 3033–3043. [Google Scholar] [CrossRef] [PubMed]
- Misiewicz, J.; Afonin, S.; Grage, S.L.; Berg, J.V.D.; Strandberg, E.; Wadhwani, P.; Ulrich, A.S. Action of the multifunctional peptide BP100 on native biomembranes examined by solid-state NMR. J. Biomol. NMR 2015, 61, 287–298. [Google Scholar] [CrossRef]
- Micsonai, A.; Wien, F.; Bulyáki, É.; Kun, J.; Moussong, É.; Lee, Y.-H.; Goto, Y.; Réfrégiers, M.; Kardos, J. BeStSel: A web server for accurate protein secondary structure prediction and fold recognition from the circular dichroism spectra. Nucleic Acids Res. 2018, 46, W315–W322. [Google Scholar] [CrossRef]
- Mortara, L.; Chaimovich, H.; Cuccovia, I.M.; Horinek, D.; Lima, F.S. Dehydration Determines Hydrotropic Ion Affinity for Zwitterionic Micelles. J. Chem. Inf. Model. 2019, 60, 604–610. [Google Scholar] [CrossRef]
- Cauz, A.C.G.; Carretero, G.P.B.; Saraiva, G.K.V.; Park, P.; Mortara, L.; Cuccovia, I.M.; Brocchi, M.; Gueiros-Filho, F.J. Violacein Targets the Cytoplasmic Membrane of Bacteria. ACS Infect. Dis. 2019, 5, 539–549. [Google Scholar] [CrossRef] [PubMed]
- Garidel, P.; Johann, C.; Mennicke, L.; Blume, A. The mixing behavior of pseudobinary phosphatidylcholine-phosphatidylglycerol mixtures as a function of pH and chain length. Eur. Biophys. J. 1997, 26, 447–459. [Google Scholar] [CrossRef]
- Hildebrand, A.; Beyer, K.; Neubert, R.; Garidel, P.; Blume, A. Solubilization of negatively charged DPPC/DPPG liposomes by bile salts. J. Colloid Interface Sci. 2004, 279, 559–571. [Google Scholar] [CrossRef]
- Eggenberger, K.; Mink, C.; Wadhwani, P.; Ulrich, A.S.; Nick, P. Using the Peptide Bp100 as a Cell-Penetrating Tool for the Chemical Engineering of Actin Filaments within Living Plant Cells. Chem. Bio. Chem. 2010, 12, 132–137. [Google Scholar] [CrossRef] [PubMed]
- Reinhardt, A.; Neundorf, I. Design and Application of Antimicrobial Peptide Conjugates. Int. J. Mol. Sci. 2016, 17, 701. [Google Scholar] [CrossRef] [PubMed]
- Paschoal, J.; Yamaguchi, J.; Miranda, J.; Carretero, G.; Melo, R.; Santos, R.; Xavier, C.; Schreier, S.; Camargo, A.; Ianzer, D. Insights into cardiovascular effects of proline-rich oligopeptide (Bj-PRO-10c) revealed by structure–activity analyses: Dissociation of antihypertensive and bradycardic effects. Amino Acids 2014, 46, 401–413. [Google Scholar] [CrossRef]
- Rohl, C.A.; Fiori, W.; Baldwin, R.L. Alanine is helix-stabilizing in both template-nucleated and standard peptide helices. Proc. Natl. Acad. Sci. USA 1999, 96, 3682–3687. [Google Scholar] [CrossRef]
- Almeida, P.F.; Ladokhin, A.S.; White, S.H. Hydrogen-bond energetics drive helix formation in membrane interfaces. Biochim. Biophys. Acta Biomembr. 2012, 1818, 178–182. [Google Scholar] [CrossRef] [PubMed]
- Gautier, R.; Douguet, D.; Antonny, B.; Drin, G. HELIQUEST: A web server to screen sequences with specific α-helical properties. Bioinformatics 2008, 24, 2101–2102. [Google Scholar] [CrossRef] [PubMed]
- Eisenberg, D.; Schwarz, E.; Komaromy, M.; Wall, R. Analysis of membrane and surface protein sequences with the hydrophobic moment plot. J. Mol. Biol. 1984, 179, 125–142. [Google Scholar] [CrossRef]
- Carretero, G.; Vicente, E.; Cilli, E.; Alvarez, C.; Jenssen, H.; Schreier, S. Dissecting the mechanism of action of actinoporins. Role of the N-terminal amphipathic α-helix in membrane binding and pore activity of sticholysins I and II. PLoS ONE 2018, 13, 0202981. [Google Scholar] [CrossRef] [PubMed]
- Ros, U.; Carretero, G.; Paulino, J.; Crusca, E., Jr.; Pazos, F.; Cilli, E.M.; Lanio, M.; Schreier, S.; Alvarez, C. Self-association and folding in membrane determine the mode of action of peptides from the lytic segment of sticholysins. Biochimie 2019, 156, 109–117. [Google Scholar] [CrossRef] [PubMed]
| Method | Fluorescence | CD | ||
|---|---|---|---|---|
| LUV POPG Content (%) | No Salt | 0.3 M NaCl | No Salt | |
| 0 | 0.01 | 0.05 | 0.005 | |
| 30 | 0.8 | 0.7 | 0.5 | |
| 50 | 2.4 | 2.0 | 1.5 | |
| 70 | 5.1 | 5.2 | 2.8 | |
| Lipid. | α-Helix (%) | β-Sheet (%) | Flexible (%) |
|---|---|---|---|
| None | 19 | 0 | 81 |
| POPC:POPG 70:30 | 76 | 6 | 18 |
| POPC:POPG 50:50 | 83 | 0 | 17 |
| POPC:POPG 30:70 | 75 | 17 | 8 |
| PG% | N (Lip/NAPHT-BP100) | ΔH (kcal/mol) | ΔS (cal/mol/K) | −T ΔS (kcal/mol) | ΔG (kcal/mol) | K (M−1) |
|---|---|---|---|---|---|---|
| 30 | 21.7 | −5.4 ± 0.3 | 9.8 ± 2.5 | −2.9 | −8.3 | 1.4 ± 1.0 × 106 |
| 50 | 10.6 | −5.1 ± 1.3 | 14.1 ± 4.8 | −4.2 | −9.3 | 3.2 ± 0.6 × 107 |
| BP100 | NAPHT-BP100 | |
|---|---|---|
| E. coli | 2 µM | 1 µM |
| S. aureus | 2 µM | 1 µM |
| B. subtilis | 2 µM | 2 µM |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Carretero, G.P.B.; Saraiva, G.K.V.; Rodrigues, M.A.; Kiyota, S.; Bemquerer, M.P.; Chaimovich, H.; Cuccovia, I.M. Naphthalimide-Containing BP100 Leads to Higher Model Membranes Interactions and Antimicrobial Activity. Biomolecules 2021, 11, 542. https://doi.org/10.3390/biom11040542
Carretero GPB, Saraiva GKV, Rodrigues MA, Kiyota S, Bemquerer MP, Chaimovich H, Cuccovia IM. Naphthalimide-Containing BP100 Leads to Higher Model Membranes Interactions and Antimicrobial Activity. Biomolecules. 2021; 11(4):542. https://doi.org/10.3390/biom11040542
Chicago/Turabian StyleCarretero, Gustavo Penteado Battesini, Greice Kelle Viegas Saraiva, Magali Aparecida Rodrigues, Sumika Kiyota, Marcelo Porto Bemquerer, Hernan Chaimovich, and Iolanda Midea Cuccovia. 2021. "Naphthalimide-Containing BP100 Leads to Higher Model Membranes Interactions and Antimicrobial Activity" Biomolecules 11, no. 4: 542. https://doi.org/10.3390/biom11040542
APA StyleCarretero, G. P. B., Saraiva, G. K. V., Rodrigues, M. A., Kiyota, S., Bemquerer, M. P., Chaimovich, H., & Cuccovia, I. M. (2021). Naphthalimide-Containing BP100 Leads to Higher Model Membranes Interactions and Antimicrobial Activity. Biomolecules, 11(4), 542. https://doi.org/10.3390/biom11040542






