Functional Role of AKNA: A Scoping Review
Abstract
:1. Introduction
2. Materials and Methods
2.1. Search Strategy
2.2. Study Selection
2.3. Eligibility Criteria
2.4. Inclusion Criteria
2.4.1. Study Design
2.4.2. Language
2.4.3. Publication Issue
2.5. Exclusion Criteria
2.6. Data Extraction
2.7. Outcome Measurement
2.8. Quality Assessment
2.9. Synthesis of Results
2.10. Statistical Analysis
3. Results
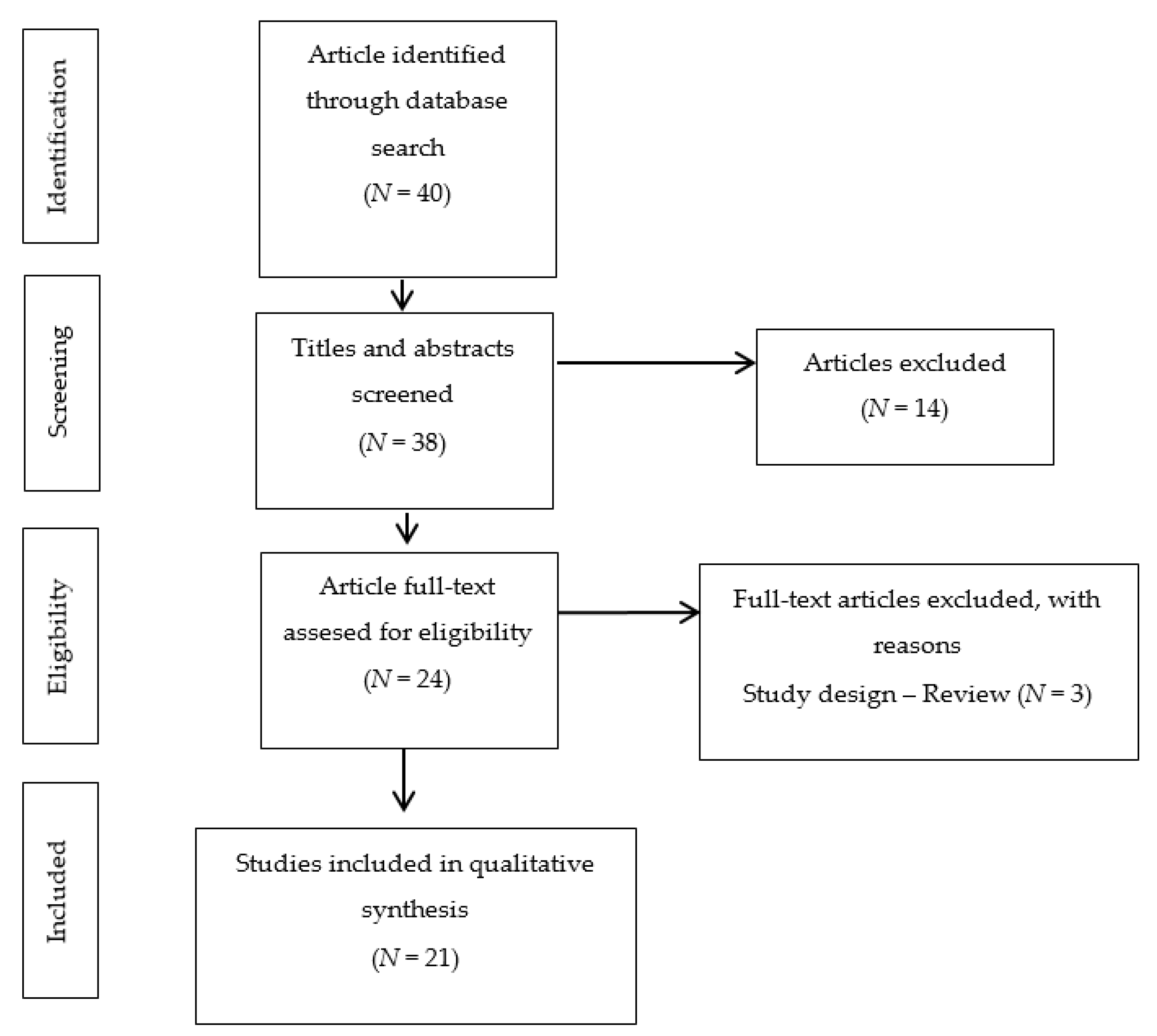
3.1. Included Studies
3.1.1. Quality Assessment of Included Studies
3.1.2. General Characteristics of Included Studies
3.1.3. Main Findings
4. Discussion
5. Conclusions
- i.
- Inflammation: Expression of proinflammatory cytokines.
- ii.
- Immune response: CD40 and CD40L regulation and B-cell maturation.
- iii.
- Development: Centrosomal microtubule reorganization and delamination of certain types of neural stem cells by an EMT-like process.
- iv.
- Molecular mechanisms associated to pathologic processes (gastric and cervical cancer, Vogt-Koyanagi-Harada syndrome, knee osteoarthritis, primary Sjögren’s syndrome, primary ciliary dyskinesia, head and neck squamous cell carcinoma, and acute lymphoblastic leukemia) being a key transcription factor that regulates genes involved in immunity, inflammation, and cancer.
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sims-Mourtada, J.C.; Bruce, S.; McKeller, M.R.; Rangel, R.; Guzman-Rojas, L.; Cain, K.; Lopez, C.; Zimonjic, D.B.; Popescu, N.C.; Gordon, J.; et al. The human AKNA gene expresses multiple transcripts and protein isoforms as a result of alternative promoter usage, splicing, and polyadenylation. DNA Cell Biol. 2005, 24, 325–338. [Google Scholar] [CrossRef] [PubMed]
- Thye, T.; Burchard, G.D.; Nilius, M.; Muller-Myhsok, B.; Horstmann, R.D. Genomewide linkage analysis identifies polymorphism in the human interferon-gamma receptor affecting Helicobacter pylori infection. Am. J. Hum. Genet. 2003, 72, 448–453. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Landvik, N.E.; Hart, K.; Skaug, V.; Stangeland, L.B.; Haugen, A.; Zienolddiny, S. A specific interleukin-1B haplotype correlates with high levels of IL1B mRNA in the lung and increased risk of non-small cell lung cancer. Carcinogenesis 2009, 30, 1186–1192. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Savas, S.; Liu, G. Genetic variations as cancer prognostic markers: Review and update. Hum. Mutat. 2009, 30, 1369–1377. [Google Scholar] [CrossRef] [PubMed]
- Siddiqa, A.; Sims-Mourtada, J.C.; Guzman-Rojas, L.; Rangel, R.; Guret, C.; Madrid-Marina, V.; Sun, Y.; Martinez-Valdez, H. Regulation of CD40 and CD40 ligand by the AT-hook transcription factor AKNA. Nature 2001, 410, 383–387. [Google Scholar] [CrossRef] [PubMed]
- Perales, G.; Burguete-García, A.I.; Dimas, J.; Bahena-Román, M.; Bermúdez-Morales, V.H.; Moreno, J.; Madrid-Marina, V. A polymorphism in the AT-hook motif of the transcriptional regulator AKNA is a risk factor for cervical cancer. Biomarkers 2010, 15, 470–474. [Google Scholar] [CrossRef] [PubMed]
- Munn, Z.; Peters, M.D.J.; Stern, C.; Tufanaru, C.; McArthur, A.; Aromataris, E. Systematic review or scoping review? Guidance for authors when choosing between a systematic or scoping review approach. BMC Med. Res. Methodol. 2018, 18, 143. [Google Scholar] [CrossRef]
- Tricco, A.C.; Lillie, E.; Zarin, W.; O’Brien, K.K.; Colquhoun, H.; Levac, D.; Moher, D.; Peters, M.D.J.; Horsley, T.; Weeks, L.; et al. PRISMA Extension for Scoping Reviews (PRISMA-ScR): Checklist and Explanation. Ann. Intern. Med. 2018, 169, 467–473. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vandenbroucke, J.P.; Von-Elm, E.; Altman, D.G.; Gøtzsche, P.C.; Mulrow, C.D.; Pocock, S.J.; Poole, C.; Schlesselman, J.J.; Egger, M. Iniciativa STROBE. Mejorar la comunicación de estudios observacionales en epidemiología (STROBE): Explicación y elaboración [Strengthening the reporting of observational studies in epidemiology (STROBE): Explanation and elaboration]. Gac. Sanit. 2009, 23, 158. [Google Scholar] [CrossRef]
- Kilkenny, C.; Browne, W.J.; Cuthill, I.C.; Emerson, M.; Altman, D.G. Improving bioscience research reporting: The ARRIVE guidelines for reporting animal research. PLoS Biol. 2010, 8, e1000412. [Google Scholar] [CrossRef]
- Ma, W.; Ortiz-Quintero, B.; Rangel, R.; McKeller, M.R.; Herrera-Rodriguez, S.; Castillo, E.F.; Schluns, K.S.; Hall, M.; Zhang, H.; Suh, W.K.; et al. Coordinate activation of in ammatory gene networks, alveolar destruction and neonatal death in AKNA de cient mice. Cell Res. 2011, 21, 1564–1577. [Google Scholar] [CrossRef] [Green Version]
- Suram, S.; Silveira, L.J.; Mahaffey, S.; Brown, G.D.; Bonventre, J.V.; Williams, D.L.; Gow, N.A.; Bratton, D.L.; Murphy, R.C.; Leslie, C.C. Cytosolic phospholipase A(2)α and eicosanoids regulat e expression of genes in macrophages involved in host defense and inflammation. PLoS ONE 2013, 8, e69002. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Piulats, J.M.; Vidal, A.; García-Rodríguez, F.J.; Muñoz, C.; Nadal, M.; Moutinho, C.; Martínez-Iniesta, M.; Mora, J.; Figueras, A.; Guinó, E.; et al. Orthoxenografts of Testicular Germ Cell Tumors Demonstrate Genomic Changes Associated with Cisplatin Resistance and Identify PDMP as a Resensitizing Agent. Clin. Cancer Res. 2018, 24, 3755–3766. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hug, P.; Anderegg, L.; Kehl, A.; Jagannathan, V.; Leeb, T. AKNA frameshift variant in three dogs with recurrent in ammatory pulmonary disease. Genes 2019, 10, 567. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pereira, G.L.; Malheiros, J.M.; Ospina, A.M.T.; Chardulo, L.A.L.; Curi, R.A. Exome sequencing in genomic regions related to racing performance of Quarter Horses. J. Appl. Genet. 2019, 60, 79–86. [Google Scholar] [CrossRef] [PubMed]
- MacDonald, K.P.; Le-Texier, L.; Zhang, P.; Morris, H.; Kuns, R.D.; Lineburg, K.E.; Leveque, L.; Don, A.L.; Markey, K.A.; Vuckovic, S.; et al. Modification of T cell responses by stem cell mobilization requires direct signaling of the T cell by G-CSF and IL-10. J. Immunol. 2014, 192, 3180–3189. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, X.; Huang, D.; Guo, P.; Wu, Q.; Dai, M.; Cheng, G.; Hao, H.; Xie, S.; Yuan, Z.; Wang, X. PKA/CREB and NF-κB pathway regulates AKNA transcription: A novel insight into T-2 toxin-induced inflammation and GH deficiency in GH3 cells. Toxicology 2017, 392, 81–95. [Google Scholar] [CrossRef]
- Camargo-Ortega, G.; Falk, S.; Johansson, P.A.; Peyre, E.; Broix, L.; Sahu, S.K.; Hirst, W.; Schlichthaerle, T.; De Juan Romero, C.; Draganova, K.; et al. The centrosome protein AKNA regulates neurogenesis via microtubule organization. Nature 2019, 567, 113–117. [Google Scholar] [CrossRef] [PubMed]
- Soriano-Valero, S.; Román-Rodriguez, J.L.; Agustín-Panadero, R.; Bellot-Arcís, C.; Fons-Font, A.; Fernández-Estevan, L. Systematic review of chewing simulators: Reality and reproducibility of in vitro studies. J. Clin. Exp. Dent. 2020, 12, e1189–e1195. [Google Scholar] [CrossRef]
- Manzo-Merino, J.; Lagunas-Martínez, A.; Contreras-Ochoa, C.O.; Lizano, M.; Castro-Muñoz, L.J.; Calderón-Corona, C.; Torres-Poveda, K.; Román-Gonzalez, A.; Hernández-Pando, R.; Bahena-Román, M.; et al. The Human Papillomavirus (HPV) E6 Oncoprotein Regulates CD40 Expression via the AT-Hook Transcription Factor AKNA. Cancers 2018, 10, 521. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dixon-Woods, M.; Agarwal, S.; Jones, D.; Young, B.; Sutton, A. Synthesising qualitative and quantitative evidence: A review of possible methods. J. Health Serv. Res. Policy 2005, 10, 45–53. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Bartenhagen, C.; Gombert, M.; Okpanyi, V.; Binder, V.; Röttgers, S.; Bradtke, J.; Teigler-Schlegel, A.; Harbott, J.; Ginzel, S.; et al. Next-generation-sequencing of recurrent childhood high hyperdiploid acute lymphoblastic leukemia reveals mutations typically associated with high risk patients. Leuk. Res. 2015, 39, 990–1001. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.; Pan, Y.; Liu, J. The relevance between the immune response-related gene module and clinical traits in head and neck squamous cell carcinoma. Cancer Manag. Res. 2019, 11, 7455–7472. [Google Scholar] [CrossRef] [Green Version]
- Shamseldin, H.E.; Al-Mogarri, I.; Alqwaiee, M.M.; Alharbi, A.S.; Baqais, K.; AlSaadi, M.; AlAnzi, T.; Alhashem, A.; Saghier, A.; Ameen, W.; et al. An exome-first approach to aid in the diagnosis of primary ciliary dyskinesia. Hum. Genet. 2020, 139, 1273–1283. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Sun, D.; Li, W.; Xin, Y. AKNA Is a Potential Prognostic Biomarker in Gastric Cancer and Function as a Tumor Suppressor by Modulating EMT-Related Pathways. BioMed Res. Int. 2020, 2020, 6726759. [Google Scholar] [CrossRef] [PubMed]
- Mao, L.; Yang, P.; Hou, S.; Li, F.; Kijlstra, A. Label-free proteomics reveals decreased expression of CD18 and AKNA in peripheral CD4+ T cells from patients with Vogt-Koyanagi-Harada syndrome. PLoS ONE 2011, 6, e14616. [Google Scholar] [CrossRef]
- Martínez-Nava, G.A.; Torres-Poveda, K.; Lagunas-Martínez, A.; Bahena-Román, M.; Zurita-Díaz, M.A.; Ortíz-Flores, E.; García-Carrancá, A.; Madrid-Marina, V.; Burguete-García, A.I. Cervical cancer-associated promoter polymorphism affects akna expression levels. Genes Immun. 2015, 16, 43–53. [Google Scholar] [CrossRef] [PubMed]
- Martínez-Nava, G.A.; Fernández-Torres, J.; Martínez-Flores, K.; Zamudio-Cuevas, Y.; Clavijo-Cornejo, D.; Espinosa-Morales, R.; Lozada, C.A.; Gutierrez, M.; Granados, J.; Pineda, C.; et al. The association of AKNA gene polymorphisms with knee osteoarthritis suggests the relevance of this immune response regulator in the disease genetic susceptibility. Mol. Biol. Rep. 2018, 45, 151–161. [Google Scholar] [CrossRef]
- Hernández-Molina, G.; Rodríguez-Pérez, J.M.; Fernández-Torres, J.; Lima, G.; Pérez-Hernández, N.; López-Reyes, A.; Martínez-Nava, G.A. HIF1A (rs11549465) and AKNA (rs10817595) Gene Polymorphisms Are Associated with Primary Sjögren’s Syndrome. BioMed Res. Int. 2017, 2017, 5845849. [Google Scholar] [CrossRef] [Green Version]
- Zhao, T.; Ma, C.; Xie, B.; Zhao, B.; Wang, W.; Liu, J. Evaluation of Common Variants in the AKNA Gene and Susceptibility to Knee Osteoarthritis among the Han Chinese. Genet. Test. Mol. Biomark. 2020, 24, 425–430. [Google Scholar] [CrossRef]
- Kawaguchi, A. Neuronal Delamination and Outer Radial Glia Generation in Neocortical Development. Front. Cell Dev. Biol. 2021, 8, 623573. [Google Scholar] [CrossRef] [PubMed]
- Waseem, S.S.; Moawia, A.; Budde, B.; Tariq, M.; Khan, A.; Ali, Z.; Khan, S.; Iqbal, M.; Malik, N.A.; Altmüller, J.; et al. A Homozygous AKNA Frameshift Variant Is Associated with Microcephaly in a Pakistani Family. Genes 2021, 12, 1494. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Zou, X.; Wang, K.; Wang, Y.; Wang, M.; Guo, T.; Zhong, B.; Jiang, N. Comprehensive analysis of Transcription Factors identified novel prognostic biomarker in human bladder cancer. J. Cancer 2021, 12, 5605–5621. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Pan, Q.; Ge, H.; Xing, L.; Hong, Y.; Chen, P. Deep learning-based clustering robustly identified two classes of sepsis with both prognostic and predictive values. EBioMedicine 2020, 62, 103081. [Google Scholar] [CrossRef] [PubMed]

| Section | Question |
|---|---|
| Introduction | Q1 Is the scientific context clearly explained? |
| Q2 Are the objectives clearly stated? | |
| Methods | Q3 What is the design of the study? (1 cross-sectional; 2 case and control) |
| Q4 Are inclusion criteria and the selection of participants clearly explained? Q5 Sample size (0 if <20, 1 if 20–100, 2 if >100) Q6 Is the method (validity) described? Q7 Are statistical analyses appropriate? | |
| Results | Q8 Are subjects’ characteristics described? |
| Q9 Are the results interpretable? | |
| Discussion | Q10 Are the study findings discussed with related studies published in the literature? |
| Q11 Are study limitations discussed? |
| Studies | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 | 20 | Score |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Ma et al. (2011) [11] | * | * | * | * | * | * | * | * | * | * | * | * | * | * | * | * | * | 17 | |||
| Suram et al. (2013) [12] | * | * | * | * | * | * | * | * | * | * | * | * | * | 13 | |||||||
| Piulats et al. (2018) [13] | * | * | * | * | * | * | * | * | * | * | * | * | * | * | 14 | ||||||
| Hug et al. (2019) [14] | * | * | * | * | * | * | * | * | * | * | * | * | * | * | 14 | ||||||
| Pereira et al. (2019) [15] | * | * | * | * | * | * | * | * | * | * | * | * | * | * | 14 | ||||||
| MacDonald et al. (2014) [16] | * | * | * | * | * | * | * | * | * | * | * | * | * | 13 | |||||||
| Liu et al. (2017) [17] | * | * | * | * | * | * | * | * | * | * | * | * | * | * | * | * | 17 | ||||
| Camargo et al. (2019) [18] | * | * | * | * | * | * | * | * | * | * | * | * | * | * | * | * | * | * | 18 |
| Studies | 1 | 2a | 2b | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | Score |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Siddiqa et al. (2001) [5] | * | * | * | * | * | * | 6 | |||||||||
| Sims et al. 2005 [1] | * | * | * | * | * | * | * | 6 | ||||||||
| Manzo-Merino et al. (2018) [20] | * | * | * | * | * | * | * | 7 | ||||||||
| MacDonald et al. (2014) [16] | * | * | * | * | * | * | * | * | 8 | |||||||
| Liu et al. (2017) [17] | * | * | * | * | * | * | * | 7 | ||||||||
| Camargo et al. (2019) [18] | * | * | * | * | * | * | * | 7 |
| Studies | Q1 | Q2 | Q3 | Q4 | Q5 | Q6 | Q7 | Q8 | Q9 | Q10 | Q11 | Total |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Perales et al. (2010) [6] | 2 | 2 | 1 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 1 | 20 |
| Mao et al. (2011) [26] | 2 | 2 | 2 | 2 | 1 | 2 | 2 | 2 | 2 | 2 | 1 | 20 |
| Martínez et al. (2015) [27] | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 22 |
| Chen et al. (2015) [22] | 2 | 2 | 1 | 1 | 1 | 2 | 2 | 2 | 2 | 2 | 0 | 17 |
| Hernández et al. (2017) [29] | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 22 |
| Martínez et al. (2018) [28] | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 22 |
| Song et al. (2019) [23] | 2 | 2 | 1 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 21 |
| Shamseldin et al. (2020) [24] | 2 | 2 | 1 | 1 | 1 | 2 | 2 | 2 | 2 | 2 | 2 | 19 |
| Wang et al. (2020) [25] | 2 | 2 | 1 | 1 | 1 | 2 | 2 | 2 | 2 | 2 | 1 | 18 |
| Zhao et al. (2020) [30] | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 22 |
| Author, Year | Study Design | No. of Participants | Principal Study Criteria | Main Finding |
|---|---|---|---|---|
| Perales et al. (2010) [6] | Cross-sectional | 47 human papillomavirus (HPV)-positive biopsies from Mexican women diagnosed with squamous intraepithelial lesion (SIL) (N = 21) or CC (cervical cancer) (N = 26) and samples of apparently healthy women (N = 50) non cervical lesion (NCL) and with HPV-negative status | To investigate the allelic frequency of arginine (R)-glutamine (Q) as potential risk factors for HPV-associated CC | AKNA Q/Q homozygosis is a risk factor for CC associated with HPV infection |
| Mao et al. (2011) [26] | Cases and controls | N = 21, 10 cases and 11 controls | To identify differentially expressed membrane proteins in patients with active Vogt-Koyanagi-Harada syndrome (VKH) and controls | A possible regulation of the CD18 molecule (integrin B2, which participates in cell adhesion, neutrophil chemotaxis, and cell signaling) was found, and a role in apoptosis of CD4 + T cells and a decreased level of AKNA expression could be observed, along with a possible down-regulation in the expression of CD40L in T cells |
| Martínez et al. (2015) [27] | Cross-sectional | 420 HPV+ women, 109 NCL, 149 SIL, and 162 CC | To assess the association of single nucleotide polymorphisms (SNPs) of AKNA F1 isoform promoter region with SIL and CC, as well as their effect on akna mRNA expression levels in peripheral blood mononuclear cell (PBMCs) in both stages of the disease, by simultaneously measuring the number of transcripts from each allele and the feature known as allelic expression imbalance (AEI) | Two polymorphisms were associated with SIL and CC, and an association between high akna expression levels and CC and SIL was identified, although its effects differed in each disease stage. To show the potential existence of a cis-acting polymorphism, the akna allelic expression imbalance was evaluated for the alleles of the−1372C4A polymorphism, and the number of transcripts derived from the A allele was found to be significantly higher than those from the C allele |
| Chen et al. (2015) [22] | Cross-sectional | Five pediatric acute lymphoblastic leukemia (ALL) patients and validation cohorts with non-recurrent high hyperdiploid ALL (N = 6), recurrent ETV6-RUNX1-positive ALL (N = 7), non-recurrent Down-syndrome associated (N = 6), and TCF3-PBX1-rearranged (N = 5) ALL | To identify relapse-associated mutations in patients with hyperdiploid acute lymphoblastic leukemia by sequencing | Somatic mutations were also detected in signaling molecules (AKNA, PPP1R3C, NLRP4, GLIS1, BAX) involved in B cell the differentiation, proliferation and in the relapse samples of recurrent hyperdiploid ALL |
| Hernández et al. (2017) [29] | Cases and controls | 110 patients with primary Sjögren’s syndrome (pSS) and 141 ethnically matched healthy controls | To evaluate the allele and genotype frequencies of polymorphic sites of the HIF1A and AKNA genes in pSS | The HIF1A Pro582Ser (rs11549465) T allele and C/T genotype, as well as the AKNA−1372C>A (rs10817595) A/A genotype were identified as susceptibility genetic factors for pSS, conferring the former a decreased and the latter an increased risk of pSS in a Mexican mestizo population |
| Martínez et al. (2018) [28] | Cases and controls | 81 knee osteoarthritis (KOA) patients and 140 healthy controls | To assess the potential association of AKNA polymorphisms with KOA susceptibility in a Mexican population | Regulatory and coding polymorphisms of akna can influence the development of KOA |
| Song et al. (2019) [23] | Cross-sectional | 250 samples from head and neck squamous cell carcinoma (HNSCC) patients | To predict an intrinsic relationship or correlation between the akna expressions and HNSCC; to validate these hub genes (genes with a high correlation in candidate modules) in the Cancer Genome Atlas (TCGA) dataset to confirm that they have a biological significance in the tumorigenesis of HNSCC | akna is one of the 16 hub genes that play an important role in HNSCC tumorigenesis, and could be used as a biomarker in the future |
| Shamseldin et al. (2020) [24] | Cross-sectional | 81 patients in 56 families | To identify pathogenic gene variants associated with primary ciliary dyskinesia (PCD) | Homozygous nonsense variation in akna 1990C>T:p.(Glna664*) was associated to PCD and AKNA is proposed as a novel candidate in a lung phenotype that overlaps clinically with PCD and a potentially multiciliated cell-specific role |
| Wang et al. (2020) [25] | Cross-sectional | 32 fresh primary (gastric cancer) GC and 32 matched normal gastric epithelial tissues from patients with GC undergoing resection in the First Affiliated Hospital of China Medical University | To investigate the role of AKNA in GC | AKNA could act as a tumor suppressor by modulating epithelial-mesenchymal transition (EMT)-related pathways in GC. AKNA could serve as a potential biomarker and an effective target for GC diagnosis and therapy |
| Zhao et al. (2020) [30] | Cases and controls | 2500 Han Chinese subjects comprising 824 KOA patients and 1676 controls | To investigate the association between the AKNA gene and susceptibility to KOA in a Han Chinese population | This study is the first to provide evidence of a potential link between the risk of KOA and akna gene polymorphism among subjects with Han Chinese ancestry |
| Author, Year | Study Model | Experimental Units | Principal Study Criteria | Main Finding |
|---|---|---|---|---|
| Ma et al. (2011) [11] | Wild type and AKNA-KO mice | AKNA-KO and AKNA-KO2 mice compared to wild-type mice | To provide experimental support to the hypothesis that AKNA expression plays an important role in the mechanisms that regulate the magnitude of inflammatory responses to pathogens | AKNA plays a role in mechanisms that regulate the magnitude of acute inflammatory responses by coordinately repressing genes involved in neutrophil activation, mobilization, and function. The authors hypothesize that the increased expression of cytokines, chemotactic factors, and MMP9 resulting from AKNA deficiency may indeed reflect a loss of the repressive function of the gene |
| Suram et al. (2013) [12] | Pathogen-free Balb/c mice | Cytosolic phospholipase A2 (cPLA2)α−/− mice, TLR4 mutant mouse strain C3H/HeJ, control strain C3H/HeOuJ, TLR2−/− (C57BL/6), MyD88−/− mice (C57BL/6/129), MyD88+/− C57BL/6/129 mice were crossed to produce MyD88−/− mice and MyD88+/+ littermate controls, C57BL/6 control mice and Dectin-1−/− mice (129sv/ev) | To investigate the functional consequences of cPLA2α activation and the effect of endogenously produced eicosanoids on gene expression in response to C. albicans | Genes for interferon regulatory factor (Irf)1 and Irf4 and AKNA were expressed at lower levels in A2α cPLA2α+/+ mouse peritoneal macrophages stimulated with C. albicans than in cPLA2α−/− macrophages, suggesting a correlation between a low AKNA expression and CD40 expression level |
| Piulats et al. (2018) [13] | Murine model (nude mice) | One nude mouse strain | To investigate the genetic basis of cisplatin resistance, as the efficacy of 82 cisplatin-based chemotherapy in the treatment of distinct malignancies is often hampered by intrinsic or acquired drug resistance of tumor cells | akna was negatively regulated in cisplatin resistant tumors with gains of 9q32-q33.1 |
| Hug et al. (2019) [14] | Rough collie dogs and dogs of various other breeds | 88 rough collie dogs, 539 dogs of various other breeds as controls | To identify the genetic origin of recurrent pulmonary disease in rough collie dogs | A variant with a 4-bp deletion (c.2717_2720delACAG) in the akna gene was identified as a causative variant candidate for an inherited, autosomal recessive, recurrent inflammatory pulmonary disease in dogs. Genetic analysis showed a genotype-phenotype correlation of the AKNA variant with inflammatory pulmonary disease |
| Pereira et al. (2019) [15] | Horses | Two groups, high- vs. low-performance in racing; 360 specimens of racing quarter horses, 78 males and 282 females | To analyze exomes and UTRs in regions previously associated to racing performance by GWAS in quarter horse racehorses | AKNA was related to racing performance in the quarter horse breed, with a positive regulation of transcription from polymerase II RNA promoter |
| Siddiqa et al. (2001) [5] | Cells of the immune system, lymphoid and non-lymphoid tissues | Human B lymphocytes, lymphoid tissues, and other kinds of non-lymphoid tissues. The article does not indicate the number of experiments performed | To evaluate the role of the AT-hook transcription factor as a regulator of CD40 and CD40L gene expression | AKNA is a human nuclear protein that contains multiple PEST protein-cleavage motifs, it is mainly expressed by B and T lymphocytes, NK, and dendritic cells. During B-lymphocyte differentiation, it is expressed by germinal center B lymphocytes. AKNA acts as an AT-hook protein that binds the A/T-rich regulatory elements of the promoters of CD40 and CD40L and coordinately regulates their expression |
| Sims et al. 2005 [1] | Human cell lines and tissues | Normal human mononuclear cells. Various cell lines including the PreB acute lymphocyte leukemia (PreB ALL) cell line Blin-1, Nalm 16, the Burkitt lymphoma cell line Raji, the T-cell line Jurkat, the erythroleukemia cell line K562, and the lymphoblastoid cell line JY. Primary B-lymphocytes purified from fresh tonsils. Tonsils or human thymus. The article does not indicate the number of experiments performed | To analyze the expression of multiple AKNA transcripts | Many of the AKNA transcripts originate from alternative splicing; others derive from differential polyadenylation and promoter usage. The alternative AKNA transcripts encode overlapping protein isoforms (p70 and p100), which can have a common function The AKNA PEST-dependent cleavage occurs in mature B cells and is required for CD40 upregulation. Multiple isoforms possibly expressed in a tissue-specific manner. The AKNA F1 and F2 isoform bind to the promoter of costimulatory molecules CD40 and CD40L (CD154) |
| Manzo-Merino et al. (2018) [20] | Human cell lines and tissues | HPV-positive cell lines, including SiHa (positive for HPV16) and HeLa (positive for HPV18), as well as the HPV-negative cell lines HaCaT and HEK293T. Primary human keratinocytes. Cervical tissue from 10 cases of hysterectomy for squamous cell carcinoma and four samples of normal cervical epithelium | To identify the effect of the HPV E6 oncoprotein on AKNA | AKNA promotes CD40 and IL-8 expression in keratinocytes |
| MacDonald et al. (2014) [16] | Murine model and cells | 8–12 weeks-old mice. T cells purified of non-T cell splenocytes. Splenic CD3+CD4+GFP+ Tregs. Data pooled from two- and three-replicate experiments | To evaluate the effect of human granulocyte colony-stimulating factor (G-CSF) on the modification of T cell responses | AKNA is upregulated in Treg cells upon stimulation with G-CSF |
| Liu et al. (2017) [17] | Murine model and GH3 cell lines | 16 specific pathogen-free female Wistar rats (Rattus norvegicus, 5–6-weeks-old). GH3 cells from the Cell Bank of the Academy of Sciences (Beijing, China). All experiments were performed at least in triplicate on three separate occasions | To evaluate the role of AKNA in the inflammatory response mediated by the T-2 toxin-induced and growth hormone deficiency | AKNA was found to be a key regulator of T-2 toxin-mediated growth hormone deficiency through a p-p65/p-CREB dependent mechanism |
| Camargo et al. (2019) [18] | Murine model and BAC-transgenic cell lines | Female and male mice at embryonic stages E9, E11, E13, E14, E15, and E18. 2–12 months-old C57BL/6J and RjHan:NMRI mice. Primary E14 cortical cells and Neuro2A, NMuMG, human iPS, A20, and Mpf cells. All experiments in this study, except proteomic analyses, were replicated multiple times with the same experimental protocol, followed by the same analysis | To search for candidate regulators of neural cell differentiation and the fate of cells | AKNA was found to be essential to organize centrosomal microtubules and promote their nucleation and growth, showing that, depending on the levels of AKNA, this protein controlled the delamination process in the formation of the subventricular zone, or the exit from the subventricular zone, revealing the critical role of AKNA in the organization of the centrosomal microtubules |
| Author, Year | Finding | Functional Interpretation |
|---|---|---|
| Siddiqa et al. (2001) [5] | Predominant expression of AKNA in secondary lymphoid organs | AKNA could be important in antigen-dependent immune responses |
| AKNA is predominantly expressed by germinal center B lymphocytes | AKNA may be important in the physiology of germinal center reaction | |
| AKNA can bind CD40 regulatory elements and to AT-rich CD40L promoter elements and upregulates the expression of CD40L | AKNA is a key transcription factor that regulates the expression of the CD40-CD40L receptor/ligand pair highlighting the physiological significance of AKNA during immune responses | |
| Sims et al. 2005 [1] | The AT-hook transcription factor AKNA is encoded by a single gene locus mapping to chromosome 9q32 | The p70 and p100 isoforms of AKNA originating from distinct alternatively processed mRNA and translated from different translation-initiation sites exhibit similar functions, pointing at the role of AKNA in controlling the amplitude of the immune response |
| AKNA expresses at least nine distinct transcripts, some of which are expressed in a tissue-specific manner | AKNA can be post-translationally processed to regulate the expression of CD40 | |
| Perales et al. (2010) [6] | AKNA Q/Q homozygosis is a risk factor for human papillomavirus (HPV)-associated cervical cancer (CC) | AKNA appears to be an important genetic factor associated with the risk of CC |
| Ma et al. (2011) [11] | akna−/− mice die postnatally due to severe and destructive lung infections | AKNA expression plays an important role in the mechanisms that regulate the magnitude of inflammatory responses to pathogens |
| Mao et al. (2011) [26] | AKNA expression is diminished in CD4+ T cells from patients with the autoimmune disease Vogt-Koyanagi-Harada (VKH) syndrome | A role for AKNA in the pathogenesis of the autoimmune disease VKH syndrome is suggested |
| Suram et al. (2013) [12] | The AT-hook transcription factor AKNA is expressed at lower levels in cytosolic phospholipase A2α cytosolic phospholipase A2 (cPLA2)α+/+ mouse peritoneal macrophages stimulated with C. albicans than cPLA2α−/− macrophages | AKNA came up as a molecule that dampens the inflammatory response downstream of the cPLA2α signaling elicited by C. Albicans in mouse peritoneal macrophages |
| MacDonald et al. (2014) [16] | Human granulocyte colony-stimulating factor (G-CSF) upregulated AKNA mRNA in nTreg cells | Although AKNA was reported to be expressed by CD4 T cells, no studies thus far have characterized the expression or function of AKNA in Tregs cells |
| There is a significant impairment in Treg development and maintenance in CD40-deficient mice | A novel role for CD40-CD40L interactions in the generation and maintenance of Treg cells was demonstrated. Further investigations determine the mechanism by which CD40 signaling contributes to Treg cells biology are necessary | |
| Martínez et al. (2015) [27] | (i) Significant negative association between squamous intraepithelial lesion (SIL) and CC with akna −392C4T and −1372C4A polymorphisms, (ii) significant association of AKNA expression levels at the cervix and in PBMC with CC diagnosis, and (iii) AKNA expression differs across SNP genotypes in CC patients | akna is a CC susceptibility genetic factor and the akna transcriptional regulation has a role in this disease |
| Chen et al. (2015) [22] | Somatic mutations in signaling molecules such as AKNA are involved in the differentiation, proliferation, and death of B cells in the relapse samples of recurrent hyperdiploid acute lymphoblastic leukemia (ALL) | Mutations found in akna may play key roles in the pathogenesis of recurrent disease in high hyperdiploid ALL |
| Hernández et al. (2017) [29] | The HIF1A Pro582Ser T allele and C/T genotype and akna −1372C > A polymorphism A/A genotype are proposed as genetic factors associated with primary Sjögren’s syndrome (pSS), in the Mexican mestizo population | AKNA function as a transcription regulator of CD40L and CD40 could be relevant in pSS pathology, in the Mexican population |
| Liu et al. (2017) [17] | AKNA is suggested to be a key regulator of T-2 toxin-mediated inflammation and growth hormone deficiency | AKNA has a significant role in T-2 toxin-induced inflammatory response and growth inhibition. The PKA/CREB and NF-κB pathway participate in the signaling regulating AKNA expression |
| Manzo-Merino et al. (2018) [20] | AKNA expression is lower in CC tissue than in normal tissues | Inherent AKNA deficiency in neoplastic epithelial cells, not found in immune cells that conform the tumoral inflammatory infiltrate |
| The HPV E6 oncoprotein de-regulate AKNA and CD40 expression. This event involves the action of p53 | The E6/p53/AKNA axis might play an important role in the de-regulation of the immune system during HR-HPV-induced carcinogenesis in CC promoting epithelial-mesenchymal transition (EMT) | |
| Restoration of AKNA levels in HeLa cells induces the expression of interleukin 8 (IL-8) | The inflammatory status promoted by HPV via p53/AKNA dampens immune response favoring tumor growth | |
| Martínez et al. (2018) [28] | Carriers of the minor allele homozygous genotype of the two AKNA SNPs (-1372C > A and Pro624Leu) have higher possibilities of knee osteoarthritis (KOA) than carriers of the heterozygous or ancestral homozygous genotypes of each SNP | This study represents the first evidence of a potential new KOA susceptibility gene. The deregulation in the co-stimulation of the immune system cells may be the mechanism underlying such association |
| Piulats et al. (2018) [13] | akna was negatively regulated in cisplatin-resistant tumors with gains in the 9q32-q33.1 region, but not in cisplatin-sensitive tumors altered, and could be related to tumor resistance to cisplatin | AKNA could be a prognostic marker to identify cisplatin refractory tumors |
| Camargo et al. (2019) [18] | AKNA is a centriolar protein that localizes to the subdistal appendages of the mother centriole in neurons | Low levels of AKNA cause stem cells to remain in the stem cell niche; higher levels promote their detachment from the niche, favoring migration |
| Hug et al. (2019) [14] | Detected a spontaneous variant in akna resulting of a 4-bp deletion and a truncated variant of AKNA as the cause of a severe autosomal recessive lung disease in dogs, resembling primary ciliary dyskinesia | As an important anti-inflammatory regulator of the immune response, AKNA should be considered as a candidate gene for human patients with unexplained recurrent inflammatory pulmonary disease |
| Pereira et al. (2019) [15] | AKNA was related to racing performance in the quarter horse breed, with a positive regulation of transcription from polymerase II RNA promoter | AKNA could be a gene related to speed for the racing line of quarter horses |
| Song et al. (2019) [23] | AKNA was one of the 16 hub genes identified in head and neck squamous cell carcinoma (HNSCC) tumorigenesis | AKNA plays an important role in HNSCC tumorigenesis |
| Shamseldin et al. (2020) [24] | A homozygous nonsense variation in akna 1990C > T:p.(Glna664*) was associated to primary ciliary dyskinesia (PCD)-like disease | AKNA is proposed as a novel candidate in a lung phenotype that overlaps clinically with PCD and potentially in ciliated cell function |
| Wang et al. (2020) [25] | AKNA is downregulated in gastric cancer (GC). AKNA is proposed as a potential tumor suppressor, affecting EMT-related pathways including chemokines and cytokines signaling pathways. AKNA might be regulated by circTRNC18/miR-762 axis | AKNA could serve as a potential biomarker and an effective target for GC diagnosis and therapy |
| Zhao et al. (2020) [30] | An intronic SNP of akna (SNP rs10817595) is significantly associated with the risk of KOA in a sample based on the Chinese Han population | A potential link between the risk of KOA and AKNA in subjects with Chinese Han ancestry. This association signal might be explained by the upregulation of the immune response and inflammation resulting of decreased AKNA expression. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ramírez-González, A.; Manzo-Merino, J.; Contreras-Ochoa, C.O.; Bahena-Román, M.; Aguilar-Villaseñor, J.M.; Lagunas-Martínez, A.; Rosenstein, Y.; Madrid Marina, V.; Torres-Poveda, K. Functional Role of AKNA: A Scoping Review. Biomolecules 2021, 11, 1709. https://doi.org/10.3390/biom11111709
Ramírez-González A, Manzo-Merino J, Contreras-Ochoa CO, Bahena-Román M, Aguilar-Villaseñor JM, Lagunas-Martínez A, Rosenstein Y, Madrid Marina V, Torres-Poveda K. Functional Role of AKNA: A Scoping Review. Biomolecules. 2021; 11(11):1709. https://doi.org/10.3390/biom11111709
Chicago/Turabian StyleRamírez-González, Abrahán, Joaquín Manzo-Merino, Carla Olbia Contreras-Ochoa, Margarita Bahena-Román, José Manasés Aguilar-Villaseñor, Alfredo Lagunas-Martínez, Yvonne Rosenstein, Vicente Madrid Marina, and Kirvis Torres-Poveda. 2021. "Functional Role of AKNA: A Scoping Review" Biomolecules 11, no. 11: 1709. https://doi.org/10.3390/biom11111709








