RP-UHPLC-MS Chemical Profiling, Biological and In Silico Docking Studies to Unravel the Therapeutic Potential of Heliotropium crispum Desf. as a Novel Source of Neuroprotective Bioactive Compounds
Abstract
1. Introduction
2. Results and Discussion
2.1. Total Bioactive Contents
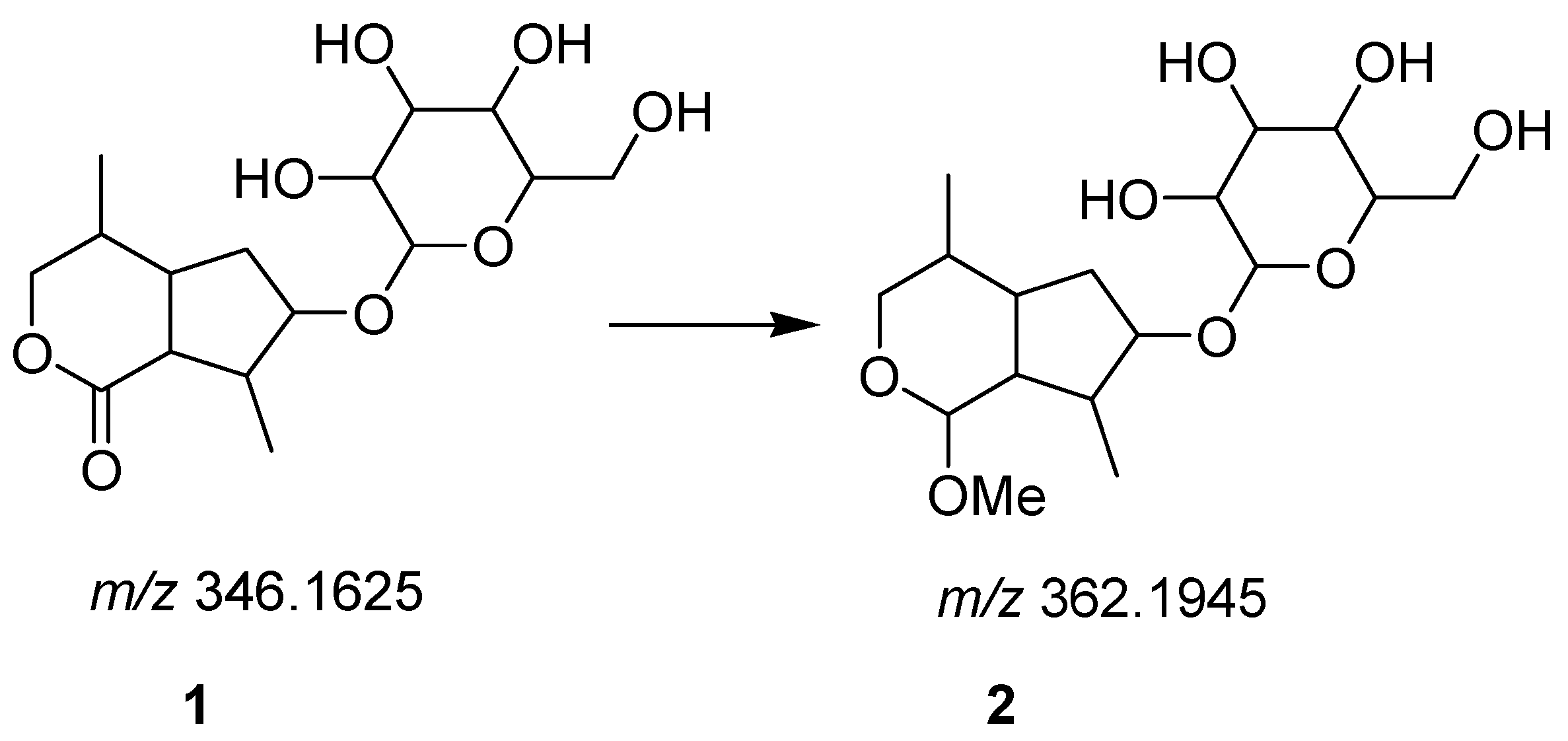
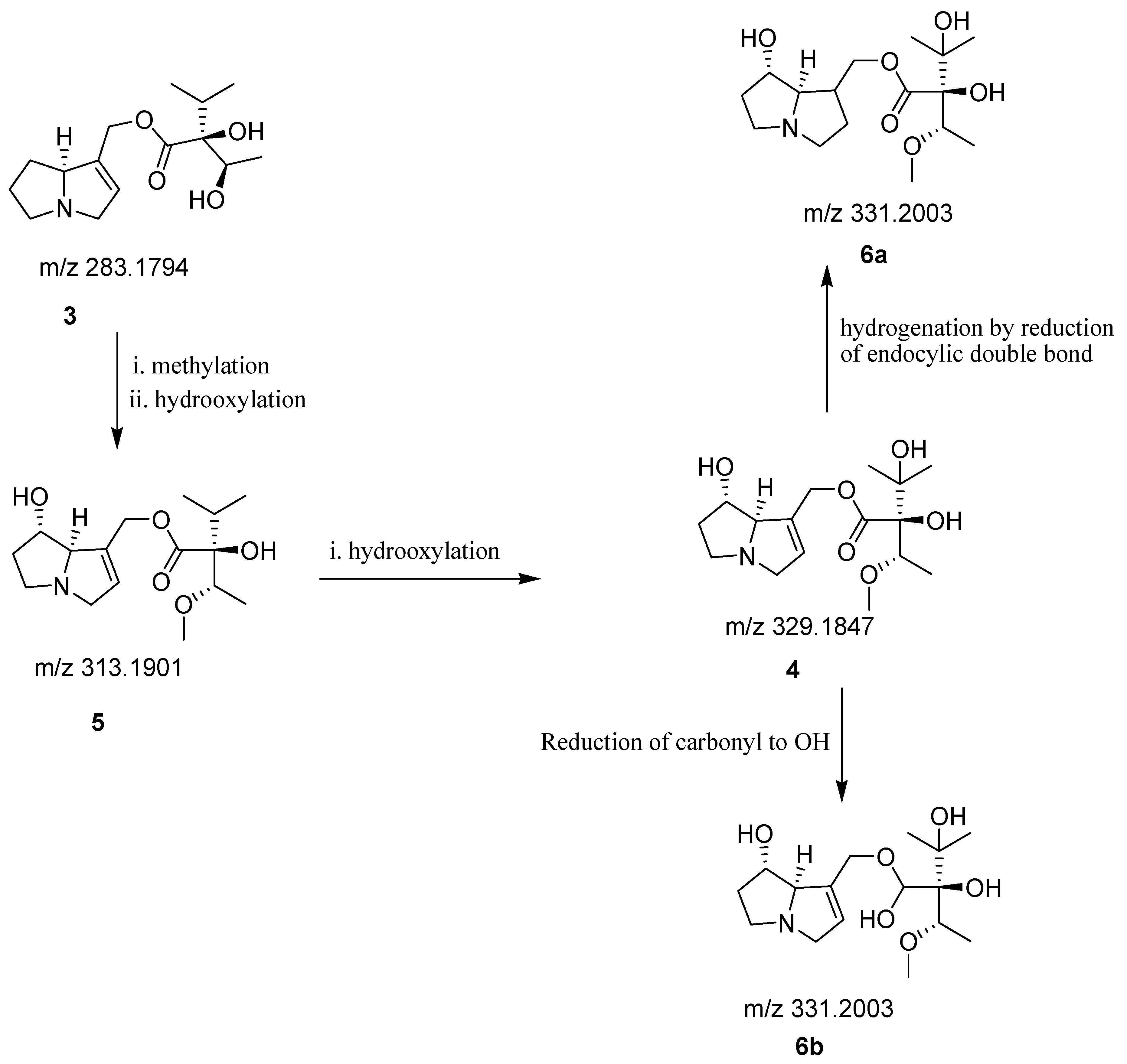
2.2. Identification of Secondary Metabolites by Reverse Phase Ultra-High Performance Liquid Chromatography Mass Spectrometry (RP-UHPLC-QTOF-MS)
2.3. Antioxidant Assays
2.4. Enzyme Inhibition Potential
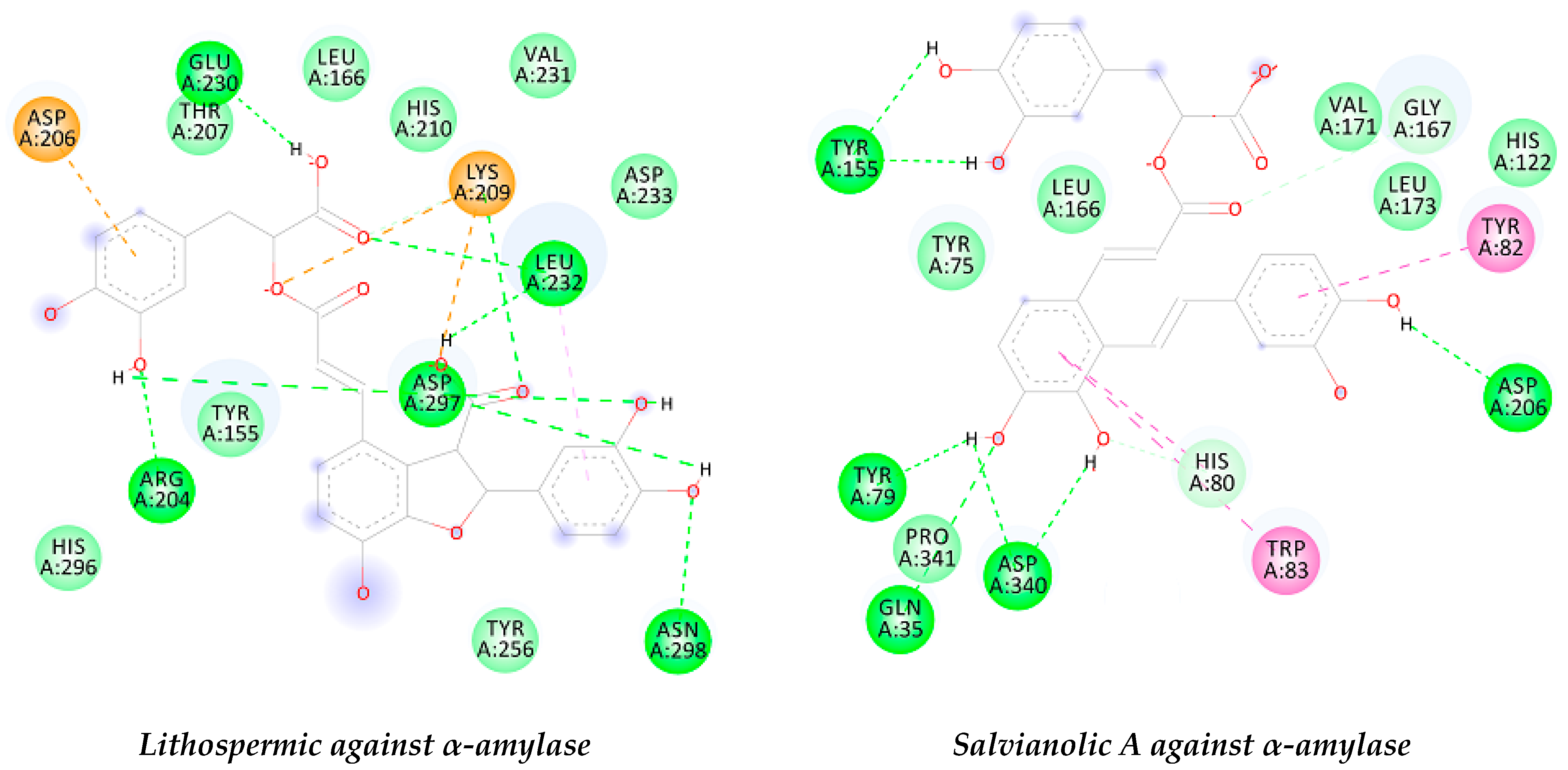
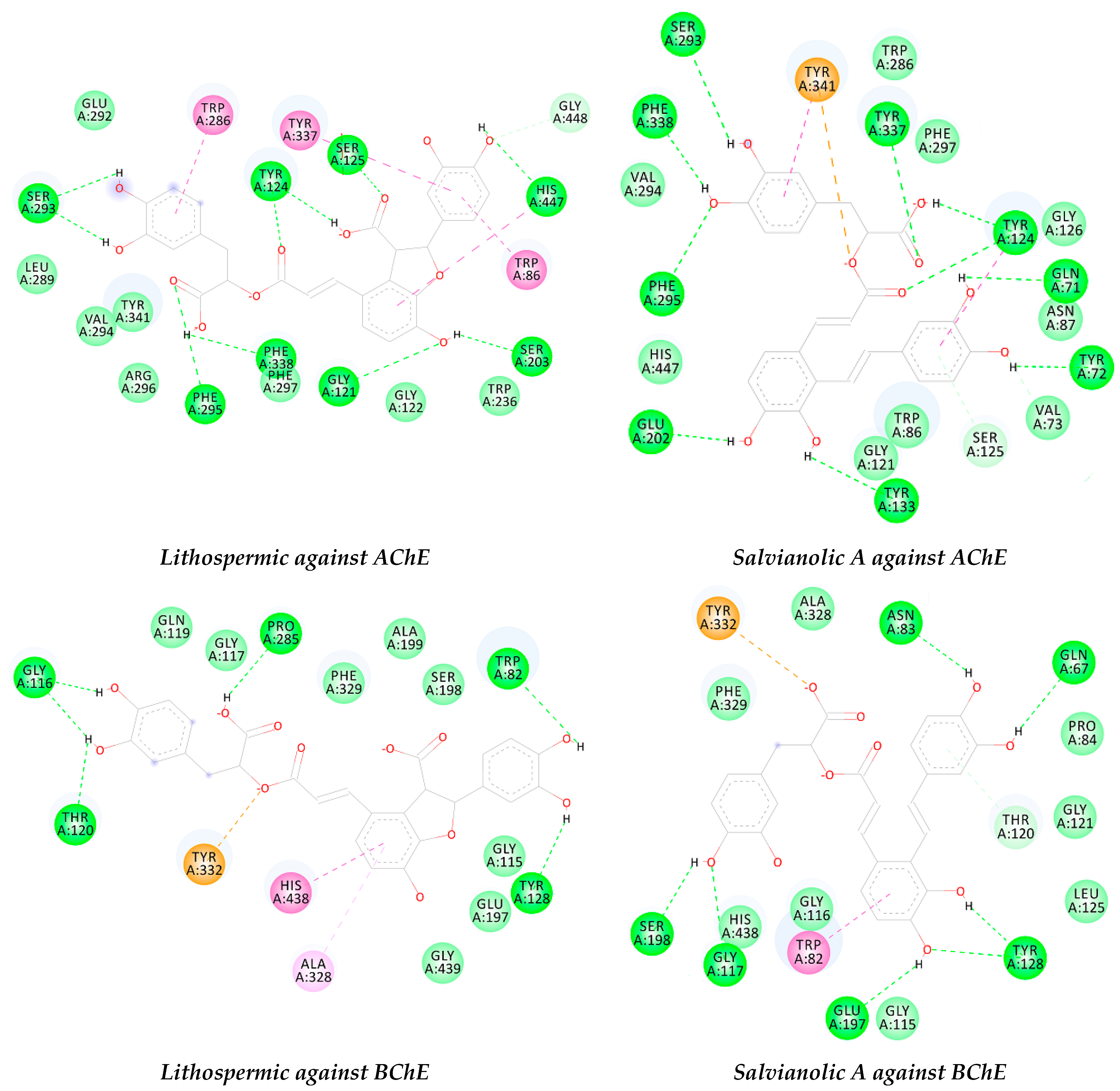
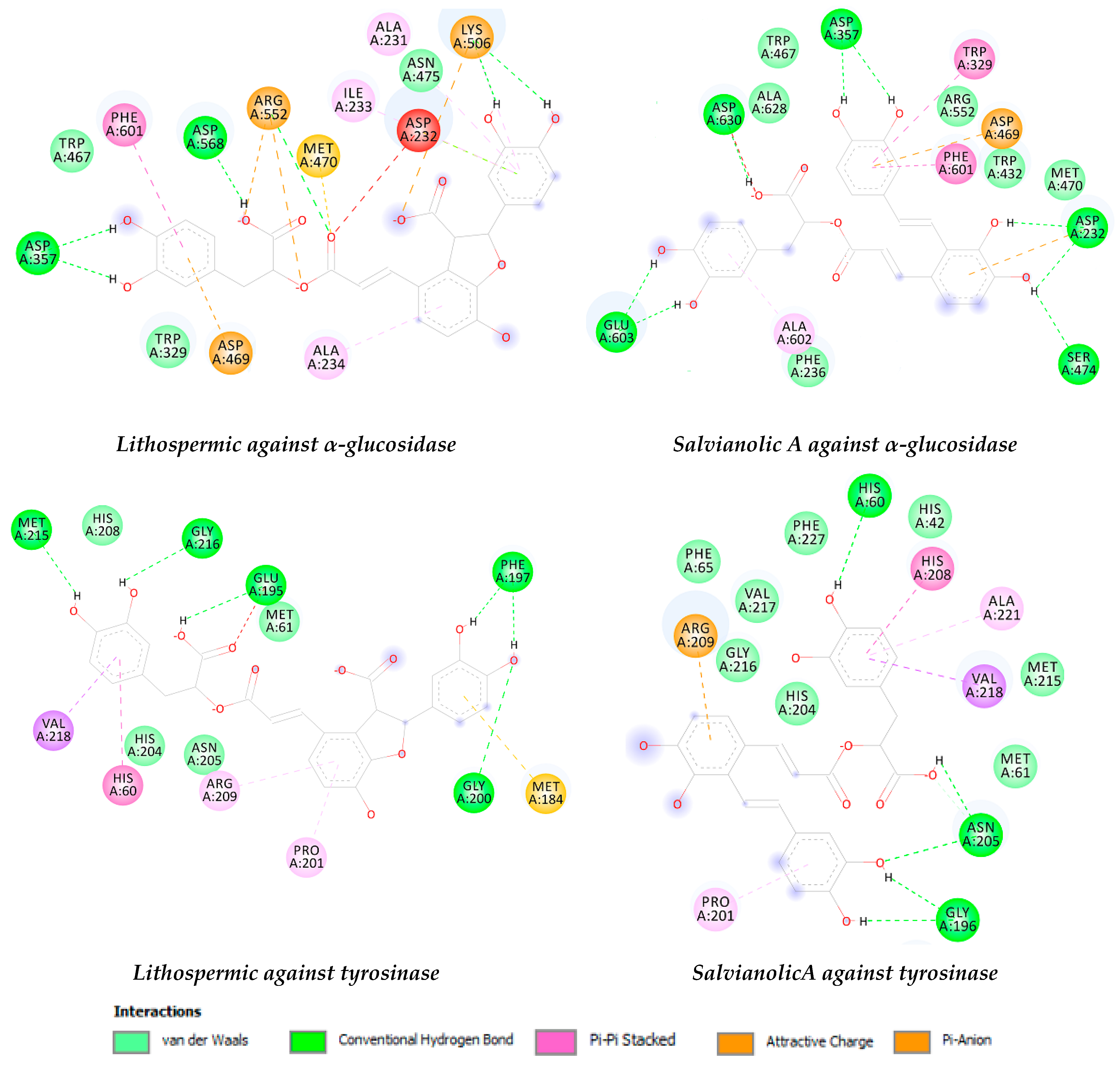
2.5. Docking Results
3. Materials and Methods
3.1. Plant Material and Extraction
3.2. Total Bioactive Contents
3.3. RP-UHPLC-MS Analysis
3.4. Antioxidant Assays
3.5. Enzyme Inhibition Assays
3.6. Docking Calculations
3.7. Statistical Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ahmad, S.; AbdEl-Salam, N.M.; Ullah, R. In vitro antimicrobial bioassays, DPPH radical scavenging activity, and FTIR spectroscopy analysis of Heliotropium bacciferum. BioMed Res. Int. 2016, 2016. [Google Scholar] [CrossRef] [PubMed]
- Cuthbertson, D.J.; Johnson, S.R.; Piljac-Žegarac, J.; Kappel, J.; Schäfer, S.; Wüst, M.; Ketchum, R.E.; Croteau, R.B.; Marques, J.V.; Davin, L.B. Accurate mass–time tag library for LC/MS-based metabolite profiling of medicinal plants. Phytochemistry 2013, 91, 187–197. [Google Scholar] [CrossRef] [PubMed]
- Shanmugam, S.; Bhavani, P. Studies on the comparison of phytochemical constituents and antimicrobial activity of Curcuma longa varieties. Int. J. Curr. Microbiol. Appl. Sci. 2014, 3, 573–581. [Google Scholar]
- Khurm, M.; Chaudhry, B.; Uzair, M.; Janbaz, K. Antimicrobial, Cytotoxic, Phytotoxic and Antioxidant Potential of Heliotropium strigosum Willd. Medicines 2016, 3, 20. [Google Scholar] [CrossRef]
- Ghori, M.K.; Ghaffari, M.A.; Hussain, S.N.; Manzoor, M.; Aziz, M.; Sarwer, W. Ethnopharmacological, phytochemical and pharmacognostic potential of genus Heliotropium L. Turk. J. Pharm. Sci. 2016, 13, 143–168. [Google Scholar] [CrossRef]
- Selvi, F.; Bigazzi, M. Leaf surface and anatomy in Boraginaceae tribe Boragineae with respect to ecology and taxonomy. Flora 2001, 196, 269–285. [Google Scholar] [CrossRef]
- Yeo, D.; Attioua, B.; Lehalle, C.; Kossi, M.; N’guessan, J.D.; Djaman, A.J.; Lobstein, A.; Frossard, N. Isolation of wound healing compounds from Heliotropium indicum. J. Appl. Pharm. Sci. 2011, 1, 12. [Google Scholar]
- Khurm, M.; Chaudhry, B.A.; Uzair, M.; Hussain, K. Antiglycation and Insecticidal Potential of Heliotropium strigosum Willd. J. Nat. Prod. Plant Resour. 2016, 6, 1–7. [Google Scholar]
- Qureshi, R.; Bhatti, G.R.; Memon, R.A. Ethnomedicinal uses of herbs from northern part of Nara desert, Pakistan. Pak. J. Bot. 2010, 42, 839–851. [Google Scholar]
- Qureshi, R.; Bhatti, G.R. Ethnobotany of plants used by the Thari people of Nara Desert, Pakistan. Fitoterapia 2008, 79, 468–473. [Google Scholar] [CrossRef]
- Hameed, M.; Ashraf, M.; Al-Quriany, F.; Nawaz, T.; Ahmad, M.S.A.; Younis, A.; Naz, N. Medicinal flora of the Cholistan desert: A review. Pak. J. Bot. 2011, 43, 39–50. [Google Scholar]
- Khan, F.; Hashmi, M.U.; Khalid, N.; Hayat, M.Q.; Ikram, A.; Janjua, H.A. Controlled assembly of silver nano-fluid in Heliotropium crispum extract: A potent anti-biofilm and bactericidal formulation. Appl. Surf. Sci. 2016, 387, 317–331. [Google Scholar] [CrossRef]
- Singh, B.; Sahu, P.M.; Sharma, R.A. Flavonoids from Heliotropium subulatum exudate and their evaluation for antioxidant, antineoplastic and cytotoxic activities II. Cytotechnology 2017, 69, 103–115. [Google Scholar] [CrossRef] [PubMed]
- Qayyum, A.; Sarfraz, R.A.; Ashraf, A.; Adil, S. Phenolic composition and biological (anti diabetic and antioxidant) activities of different solvent extracts of an endemic plant (heliotropium strigosum). J. Chil. Chem. Soc. 2016, 61, 2828–2831. [Google Scholar] [CrossRef]
- Pragada, R.; Rao Ethadi, S.; Yasodhara, B.; Praneeth Dasari, V.; Mallikarjuna, R.T. In-vitro antioxidant and antibacterial activities of different fractions of Heliotropium indicum L. J. Pharm. Res. 2012, 5, 1051. [Google Scholar]
- Xie, S.; Uesato, S.; Inouye, H.; Fujita, T.; Murai, F.; Tagawa, M.; Shingu, T. Absolute structure of nepetaside, a new iridoid glucoside from Nepeta cataria. Phytochemistry 1988, 27, 469–472. [Google Scholar] [CrossRef]
- Mori, A.; Ohkusu, H.; Kohsaka, M.; Kurono, M. Isolation of hypoxanthine from calf brain. J. Neurochem. 1973, 20, 1291–1292. [Google Scholar] [CrossRef]
- Ringeard, J.M.; Griesmar, P.; Caplain, E.; Michiel, M.; Serfaty, S.; Huerou, J.Y.L.; Marinkova, D.; Yotova, L. Design of poly (N-acryloylglycine) materials for incorporation of microorganisms. J. Appl. Polym. Sci. 2013, 130, 835–841. [Google Scholar] [CrossRef]
- Sauter, H.; Steglich, W.; Anke, T. Strobilurins: Evolution of a new class of active substances. Angew. Chem. Int. Ed. 1999, 38, 1328–1349. [Google Scholar] [CrossRef]
- Chang, H.S.; Cheng, M.J.; Chen, I.S. Secondary metabolites from magnolia kachirachirai. Helv. Chim. Acta 2011, 94, 703–710. [Google Scholar] [CrossRef]
- Khan, S.; Tousif, M.I.; Raiz, N.; Nazir, M.; Mukhtar, M.; Ali, L.; Tareen, R.B.; Saleem, M. Rarely Occurring Natural Products Isolated from Vincetoxicum stocksii. J. Chem. Soc. Pak. 2019, 41, 695–700. [Google Scholar]
- Ranilla, L.G.; Kwon, Y.-I.; Apostolidis, E.; Shetty, K. Phenolic compounds, antioxidant activity and in vitro inhibitory potential against key enzymes relevant for hyperglycemia and hypertension of commonly used medicinal plants, herbs and spices in Latin America. Bioresour. Technol. 2010, 101, 4676–4689. [Google Scholar] [CrossRef] [PubMed]
- Guntern, A.; Ioset, J.-R.; Queiroz, E.F.; Sandor, P.; Foggin, C.; Hostettmann, K. Heliotropamide, a Novel Oxopyrrolidine-3-carboxamide from Heliotropium o valifolium. J. Nat. Prod. 2003, 66, 1550–1553. [Google Scholar] [CrossRef] [PubMed]
- Bender, O.; Llorent-Martínez, E.J.; Zengin, G.; Mollica, A.; Ceylan, R.; Molina-García, L.; Fernández-de Córdova, M.L.; Atalay, A. Integration of in vitro and in silico perspectives to explain chemical characterization, biological potential and anticancer effects of Hypericum salsugineum: A pharmacologically active source for functional drug formulations. PLoS ONE 2018, 13, e0197815. [Google Scholar] [CrossRef]
- EL-Manawaty, M.; Gohar, L. In vitro alpha-glucosidase inhibitory activity of Egyptian plant extracts as an indication for their antidiabetic activity. Asian J. Pharm. Clin. Res. 2018, 11, 360–367. [Google Scholar] [CrossRef]
- Grochowski, D.M.; Uysal, S.; Aktumsek, A.; Granica, S.; Zengin, G.; Ceylan, R.; Locatelli, M.; Tomczyk, M. In vitro enzyme inhibitory properties, antioxidant activities, and phytochemical profile of Potentilla thuringiaca. Phytochem. Lett. 2017, 20, 365–372. [Google Scholar] [CrossRef]
- Saleem, H.; Htar, T.T.; Naidu, R.; Nawawi, N.S.; Ahmad, I.; Ashraf, M.; Ahemad, N. Biological, chemical and toxicological perspectives on aerial and roots of Filago germanica (L.) huds: Functional approaches for novel phyto-pharmaceuticals. Food Chem. Toxicol. 2019, 123, 363–373. [Google Scholar] [CrossRef]
- Irwin, J.J.; Sterling, T.; Mysinger, M.M.; Bolstad, E.S.; Coleman, R.G. ZINC: A free tool to discover chemistry for biology. J. Chem. Inf. Modeling 2012, 52, 1757–1768. [Google Scholar] [CrossRef]
- Frisch, M.; Trucks, G.; Schlegel, H.B.; Scuseria, G.E.; Robb, M.A.; Cheeseman, J.R.; Scalmani, G.; Barone, V.; Mennucci, B.; Petersson, G. Gaussian 09, Revision d. 01; Gaussian. Inc.: Wallingford, CT, USA, 2009; Volume 201. [Google Scholar]
| Total Bioactive Contents | |
| Total phenolic content (mg GAE/g extract) | 24.84 ± 0.82 |
| Total flavonoid content (mg RE/g extract) | 19.73 ± 0.45 |
| Antioxidant assays | |
| DPPH (mg TE/g extract) | 43.19 ± 0.97 |
| ABTS (mg TE/g extract) | 41.80 ± 3.85 |
| FRAP (mg TE/g extract) | 63.11 ± 1.51 |
| CUPRAC (mg TE/g extract) | 116.11 ± 3.39 |
| Phosphomolybdenum (mg TE/g extract) | 2.11 ± 0.16 |
| Metal chelating (mg EDTAE/g extract) | 46.72 ± 2.27 |
| Compound Class | RT | Mass | Name | Formula |
|---|---|---|---|---|
| Peptide | 0.63 | 226.0711 | 5-Acetylamino-6-formylamino-3-methyluracil (7) | C8H10N4O4 |
| Pyruvic acids | 0.639 | 216.0414 | Isobergaptene (27) | C12H8O4 |
| Peptide | 0.658 | 166.0487 | 1-Methylxanthine (8) | C6H6N4O2 |
| Peptide | 0.664 | 136.0387 | Hypoxanthine (9) | C5H4N4O |
| Pyruvic acids | 0.885 | 104.0113 | β-Hydroxypyruvic acid (25) | C3H4O4 |
| Peptide | 1.002 | 129.0431 | N-Acryloylglycine (10) | C5H7NO3 |
| Pyruvic acids | 3.339 | 198.0530 | 2-Hydroxy-3,4-dimethoxybenzoic Acid (28) | C9H10O5 |
| Pyruvic acids | 3.388 | 278.0100 | 3-Methoxymandelic acid-4-O-sulfate (29) | C9H10O8S |
| Peptide | 6.329 | 391.1849 | Orysastrobin (11) | C18H25N5O5 |
| Pyruvic acids | 6.89 | 138.0317 | p-Salicylic acid (30) | C7H6O3 |
| Polyphenolic | 6.977 | 350.0676 | Benzenebutanoic acid, 2-hydroxy-3,4-dimethoxy-6-methyl-5-(sulfooxy) (33) | C13H18O9S |
| Peptide | 7.175 | 308.0937 | Mecarbinzid (12) | C13H16N4O3S |
| Polyphenolic | 7.464 | 396.1987 | 1-Hexanol arabinosylglucoside (34) | C17H32O10 |
| Peptide | 7.82 | 176.1047 | L-Cladinose (21) | C8H16O4 |
| Peptide | 8.214 | 411.1563 | Oxidized Oplophorus luciferin (13) | C25H21N3O3 |
| Peptide | 8.521 | 454.187 | Biotin-X-NHS (14) | C20H30N4O6S |
| Polyphenolic | 8.796 | 304.0951 | (S)-Chiral alcohol (35) | C16H16O6 |
| Polyphenolic | 8.87 | 306.1111 | Cimifugin (36) | C16H18O6 |
| Polyphenolic | 8.993 | 450.2090 | Rivenprost (37) | C24H34O6S |
| Polyphenolic | 9.649 | 440.0418 | Eupatin 3-O-sulfate (38) | C18H16O11S |
| Pyruvic acids | 9.702 | 188.1049 | Nonic Acid (26) | C9H16O4 |
| Polyphenolic | 9.772 | 360.0850 | Tamadone (39) | C18H16O8 |
| Coumaric acid | 9.772 | 650.2034 | Cis-ferulic acid [arabinosyl-(1->3)-[glucosyl-(1->6)]-glucosyl] ester (31) | C27H38O18 |
| Carbamate | 9.778 | 290.1171 | Carbinoxamine (15) | C16H19ClN2O |
| Coumaric acid | 10.054 | 194.0581 | Ferulic acid (32) | C10H10O4 |
| Polyphenolic | 10.333 | 344.09 | Wightin (40) | C18H16O7 |
| Polyphenolic | 10.456 | 494.1219 | Salvianolic acid A (41) | C26H22O10 |
| Polyphenolic | 10.704 | 538.1115 | Lithospermic acid (42) | C27H22O12 |
| Irodide | 10.711 | 362.19457 | Nepetaside A (2) | C17H30O8 |
| Fatty acids | 11.183 | 328.2252 | 11-hydroperoxy-12,13-epoxy-9-octadecenoic acid (43) | C18H32O5 |
| Irodide | 11.203 | 346.1625 | Nepetaside (1) | C16H26O8 |
| Fatty acids | 11.586 | 330.2411 | 5,8,12-trihydroxy-9-octadecenoic acid (44) | C18H34O5 |
| Fatty acids | 11.921 | 288.2302 | 9,16-dihydroxy-palmitic acid (45) | C16H32O4 |
| Fatty acids | 14.275 | 314.2458 | 9,10-Epoxy-18-hydroxystearate (46) | C18H34O4 |
| Fatty acids | 15.238 | 294.2195 | α-9(10)-EpODE (47) | C18H30O3 |
| Fatty acids | 16.052 | 296.2352 | 12-oxo-10Z-octadecenoic acid (48) | C18H32O3 |
| Fatty acids | 18.599 | 278.2246 | 9Z,12Z,15E-octadecatrienoic acid (49) | C18H30O2 |
| Fatty acids | 19.493 | 280.2407 | 6E,9E-octadecadienoic acid (50) | C18H32O2 |
| Compound Class | RT | Mass | Name | Formula |
|---|---|---|---|---|
| Peptide | 0.7 | 157.1101 | Tranexamic acid (16) | C8H15NO2 |
| Peptide | 0.715 | 139.0999 | 5-Pentyloxazole (17) | C8H13NO |
| Peptide | 0.715 | 169.1105 | Homoarecoline (18) | C9H15NO2 |
| Peptide | 6.865 | 315.1686 | Alizapride (20) | C16H21N5O2 |
| Pyrrolizidine alkaloids | 7.338 | 283.1794 | Supinine (3) | C15H25NO4 |
| Pyrrolizidine alkaloids | 7.414 | 329.1847 | Europine (4) | C16H27NO6 |
| Pyrrolizidine alkaloids | 7.486 | 331.2003 | Europine A (6) | C16H29NO6 |
| Pyrrolizidine alkaloids | 7.677 | 313.1901 | Heliotrine (5) | C16H27NO5 |
| Peptide | 8.282 | 297.1953 | N-(3-Oxododecanoyl) homoserine lactone (19) | C16H27NO4 |
| Peptide | 8.852 | 437.2331 | N1, N10-Dicoumaroylspermidine (22) | C25H31N3O4 |
| Peptide | 17.871 | 292.1684 | 9-Acetoxyfukinanolide (23) | C17H24O4 |
| Peptide | 19.586 | 281.2728 | Dodemorph (24) | C18H35NO |
| Enzyme Inhibition Assays | |
|---|---|
| AChE (mg GALAE/g extract) | 3.80 ± 0.89 |
| BChE (mg GALAE/g extract) | 3.44 ± 0.13 |
| α-Amylase (mmol ACAE/g extract) | 0.57 ± 0.02 |
| α -Glucosidase (mmolACAE/g) | 1.86 ± 0.01 |
| Tyrosinase (mg KAE/g extract) | 129.65 ± 0.82 |
| Compound | α-Amylase | AChE | BChE | α-Glucosidase | Tyrosinase |
|---|---|---|---|---|---|
| Cimifugin (36) | −6.96 | −8.04 | −8.28 | −6.32 | −5.85 |
| (7.94 uM) | (1.29 uM) | (855.31 nM) | (23.45 uM) | (51.22 uM) | |
| Europine (4) | −5.17 | −7.63 | −6.86 | −6.59 | −6.18 |
| (161.82 uM) | (2.56 uM) | (9.33 uM) | (14.82 uM) | (29.51 uM) | |
| Heliotrine (5) | −5.75 | −7.55 | −7.21 | −7.17 | −5.88 |
| (60.66 uM) | (2.94 uM) | (5.22 uM) | (5.59 uM) | (48.76 uM) | |
| 11-hydroperoxy-12,13-epoxy-9-octadecenoic acid (43) | −6.48 | −8.23 | −7.52 | −6.75 | −5.6 |
| (17.80 uM) | (931.96 nM) | (3.08 uM) | (11.30 uM) | (77.96 uM) | |
| Lithospermic (42) | −8.07 | −12.1 | −10.7 | −8.11 | −7.34 |
| (1.21 uM) | (1.35 nM) | (14.33 nM) | (1.14 uM) | (4.17 uM) | |
| Salvianolic A (41) | −7.7 | −12.26 | −10.3 | −8.77 | −6.89 |
| (2.27 uM) | (1.03 nM) | (28.00 nM) | (372.12 nM) | (8.95 uM) | |
| Supinine (3) | −5.93 | −7.73 | −7.27 | −7.17 | −6.69 |
| (45.08 uM) | (2.17 uM) | (4.65 uM) | (5.59 uM) | (12.53 uM) | |
| 5,8,12-TH-9-OD (44) | −4.47 | −6.94 | −6.52 | −6.37 | −5.86 |
| (532.97 uM) | (8.13 uM) | (16.58 uM) | (21.39 uM) | (50.72 uM) |
| Enzyme | Lithospermic (42) | Salvianolic A (41) | ||
|---|---|---|---|---|
| Docking Free Energy in kcal/mol | Interactions | Docking Free Energy in kcal/mol | Interactions | |
| α-amylase | −8.07 | HB: Asn298, Asp297, Arg204, Leu232, and Gly230. | −7.7 | HB: Tyr155, Asp206, Asp340, Glu35, and Tyr79. |
| π-π: Tyr82 and Trp83. | ||||
| AChE | −12.1 | HB: His447, Ser203, Gly121, Phe383, Phe295, Ser293, Tyr124, and Ser125. | −12.26 | HB: Tyr337, Gln71, Tyr72, Tyr133, Glu202, Phe383, Phe295, Ser293, and Tyr124. |
| π-π: Trp86, Trp286 and Tyr337. | π-π: Tyr124 and Tyr341. | |||
| BChE | −10.7 | HB: Trp82, Tyr128, Thr120, Gly116, and Pro285. | −10.3 | HB: Glu67, Tyr128, Glu197, Gly117, Ser198, and Asn83. |
| π-π: His438. | π-π: Trp82. | |||
| α-glucosidase | −8.11 | HB: Asp568 and Asp357. | −8.77 | HB: Asp357, Asp232, Ser474, Glu603, Asp630, and Asp357. |
| π-π: Phe601. | π-π: Trp329 and Phe601. | |||
| tyrosinase | −7.34 | HB: Met215, Gly216, Glu195, Phe197, and Gly200. | −6.89 | HB: His60, Asn205, and Gly196. |
| π-π: His60 and Pro201. | π-π: Pro201 | |||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Arshad, A.; Ahemad, S.; Saleem, H.; Saleem, M.; Zengin, G.; Abdallah, H.H.; Tousif, M.I.; Ahemad, N.; Fawzi Mahomoodally, M. RP-UHPLC-MS Chemical Profiling, Biological and In Silico Docking Studies to Unravel the Therapeutic Potential of Heliotropium crispum Desf. as a Novel Source of Neuroprotective Bioactive Compounds. Biomolecules 2021, 11, 53. https://doi.org/10.3390/biom11010053
Arshad A, Ahemad S, Saleem H, Saleem M, Zengin G, Abdallah HH, Tousif MI, Ahemad N, Fawzi Mahomoodally M. RP-UHPLC-MS Chemical Profiling, Biological and In Silico Docking Studies to Unravel the Therapeutic Potential of Heliotropium crispum Desf. as a Novel Source of Neuroprotective Bioactive Compounds. Biomolecules. 2021; 11(1):53. https://doi.org/10.3390/biom11010053
Chicago/Turabian StyleArshad, Adeel, Saeed Ahemad, Hammad Saleem, Muhammad Saleem, Gokhan Zengin, Hassan H. Abdallah, Muhammad Imran Tousif, Nafees Ahemad, and Mohamad Fawzi Mahomoodally. 2021. "RP-UHPLC-MS Chemical Profiling, Biological and In Silico Docking Studies to Unravel the Therapeutic Potential of Heliotropium crispum Desf. as a Novel Source of Neuroprotective Bioactive Compounds" Biomolecules 11, no. 1: 53. https://doi.org/10.3390/biom11010053
APA StyleArshad, A., Ahemad, S., Saleem, H., Saleem, M., Zengin, G., Abdallah, H. H., Tousif, M. I., Ahemad, N., & Fawzi Mahomoodally, M. (2021). RP-UHPLC-MS Chemical Profiling, Biological and In Silico Docking Studies to Unravel the Therapeutic Potential of Heliotropium crispum Desf. as a Novel Source of Neuroprotective Bioactive Compounds. Biomolecules, 11(1), 53. https://doi.org/10.3390/biom11010053







