Lipid Phosphate Phosphatases and Cancer
Abstract
1. Introduction
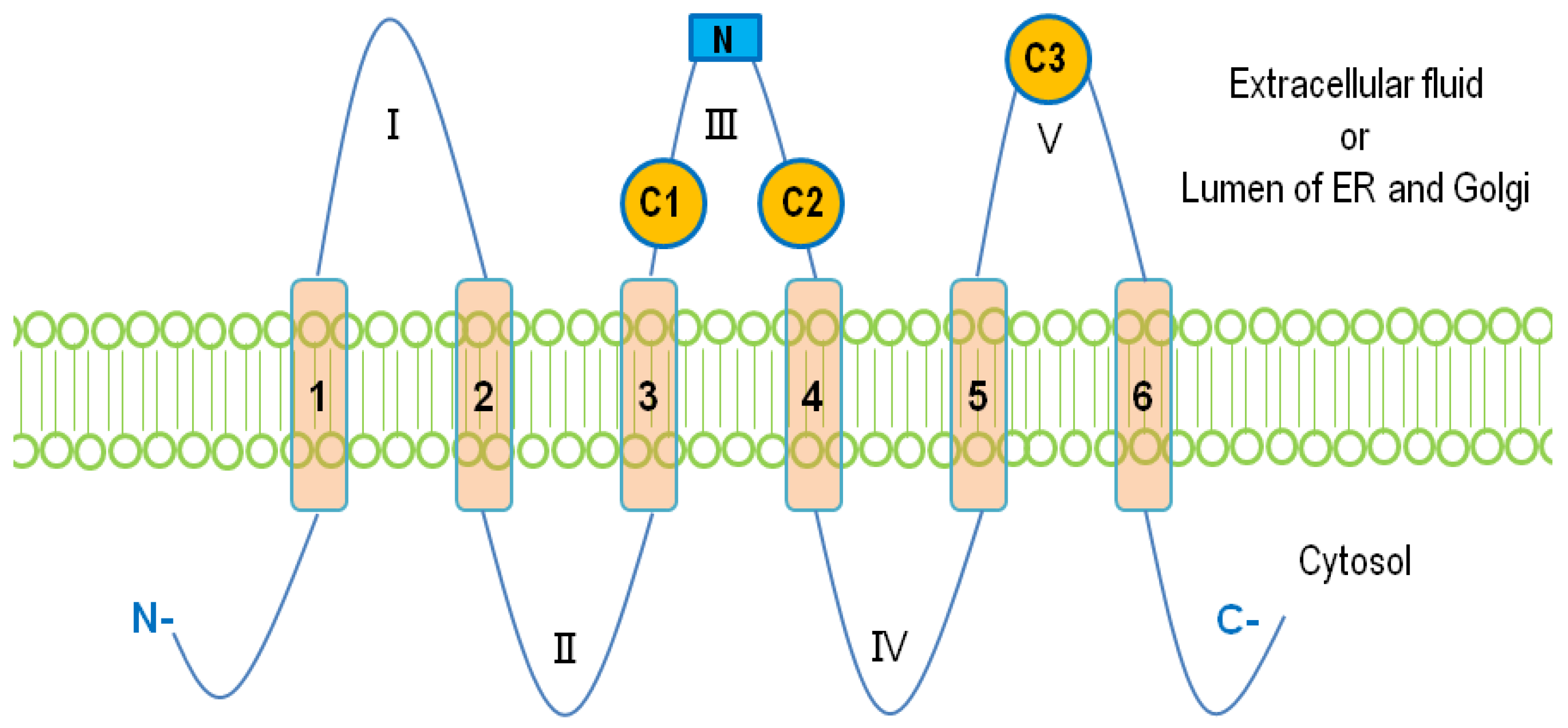
2. Structure and Membrane Topology of LPP
3. Ecto-Activity of LPPs
4. Intracellular Activities of the LPPs
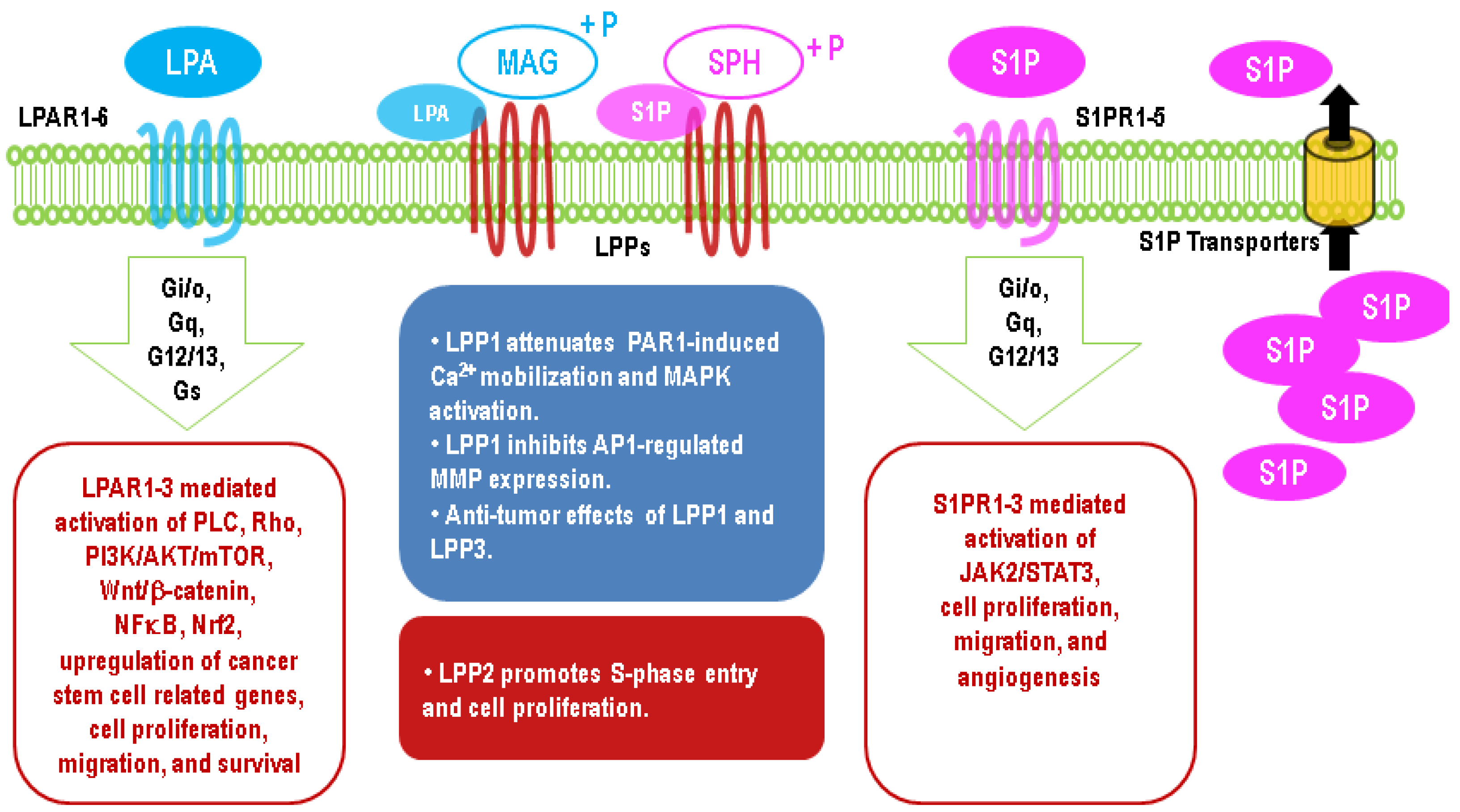
5. Upregulation of LPA Signaling in Cancers
6. Upregulation of S1P Signaling in Cancers
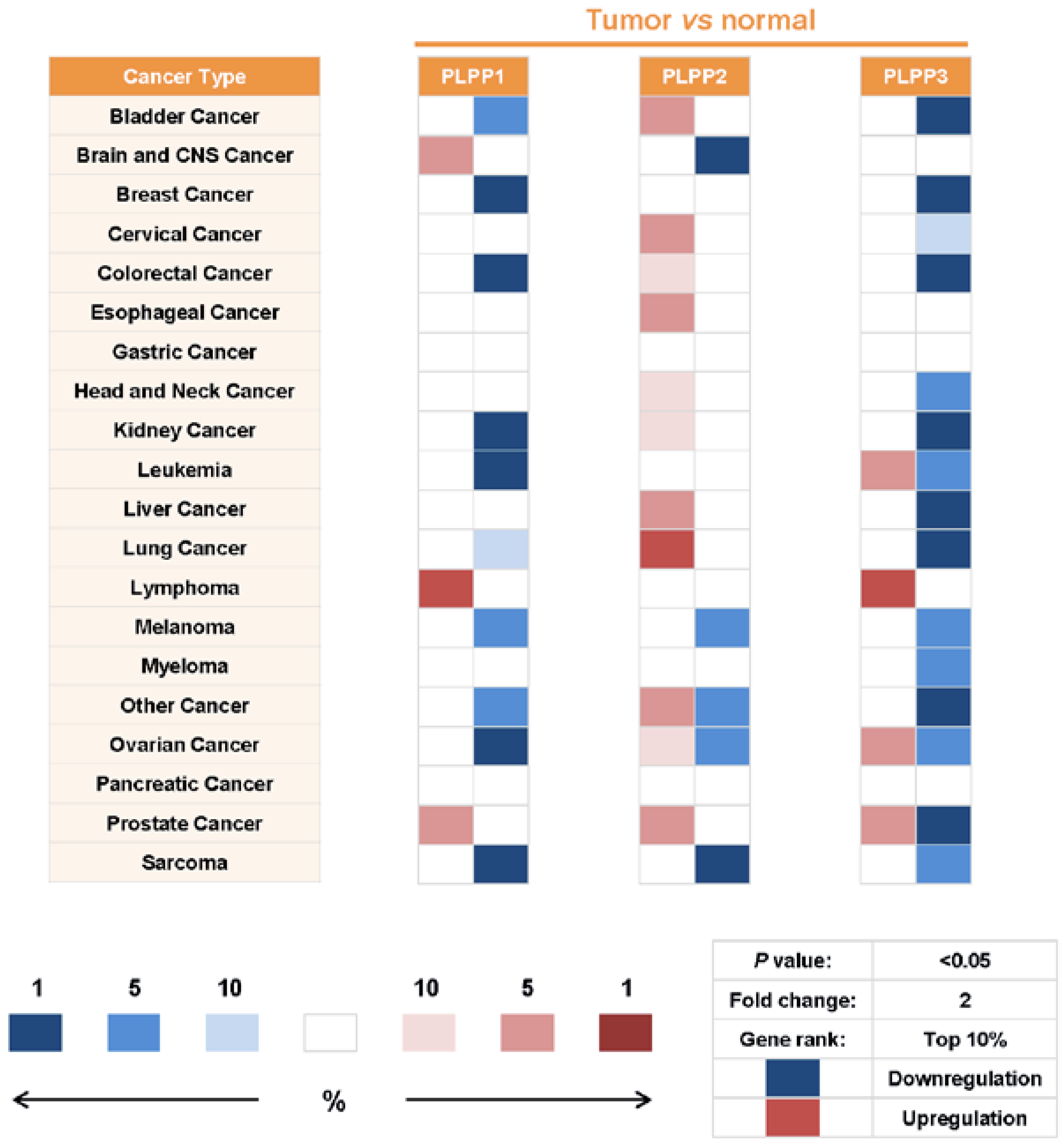
7. Alterations of LPP Expression in Cancers
8. Effects of LPPs in Cancers
9. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Brindley, D.N.; Waggoner, D.W. Phosphatidate phosphohydrolase and signal transduction. Chem. Phys. Lipids 1996, 80, 45–57. [Google Scholar] [CrossRef]
- Dillon, D.A.; Chen, X.; Zeimetz, G.M.; Wu, W.I.; Waggoner, D.W.; Dewald, J.; Brindley, D.N.; Carman, G.M. Mammalian Mg2+-independent phosphatidate phosphatase (PAP2) displays diacylglycerol pyrophosphate phosphatase activity. J. Biol. Chem. 1997, 272, 10361–10366. [Google Scholar] [CrossRef]
- Takeuchi, M.; Harigai, M.; Momohara, S.; Ball, E.; Abe, J.; Furuichi, K.; Kamatani, N. Cloning and characterization of DPPL1 and DPPL2, representatives of a novel type of mammalian phosphatidate phosphatase. Gene 2007, 399, 174–180. [Google Scholar] [CrossRef]
- Fukunaga, K.; Arita, M.; Takahashi, M.; Morris, A.J.; Pfeffer, M.; Levy, B.D. Identification and functional characterization of a presqualene diphosphate phosphatase. J. Biol. Chem. 2006, 281, 9490–9497. [Google Scholar] [CrossRef]
- Carlo, T.; Petasis, N.A.; Levy, B.D. Activation of polyisoprenyl diphosphate phosphatase 1 remodels cellular presqualene diphosphate. Biochemistry 2009, 48, 2997–3004. [Google Scholar] [CrossRef][Green Version]
- Ren, H.; Panchatcharam, M.; Mueller, P.; Escalante-Alcalde, D.; Morris, A.J.; Smyth, S.S. Lipid phosphate phosphatase (LPP3) and vascular development. Biochim. Biophys. Acta 2013, 1831, 126–132. [Google Scholar] [CrossRef]
- Morris, K.E.; Schang, L.M.; Brindley, D.N. Lipid phosphate phosphatase-2 activity regulates S-phase entry of the cell cycle in Rat2 fibroblasts. J. Biol. Chem. 2006, 281, 9297–9306. [Google Scholar] [CrossRef]
- Chandra, M.; Escalante-Alcalde, D.; Bhuiyan, M.S.; Orr, A.W.; Kevil, C.; Morris, A.J.; Nam, H.; Dominic, P.; McCarthy, K.J.; Miriyala, S.; et al. Cardiac-specific inactivation of LPP3 in mice leads to myocardial dysfunction and heart failure. Redox Biol. 2018, 14, 261–271. [Google Scholar] [CrossRef]
- Tang, X.; Benesch, M.G.; Brindley, D.N. Lipid phosphate phosphatases and their roles in mammalian physiology and pathology. J. Lipid Res. 2015, 56, 2048–2060. [Google Scholar] [CrossRef]
- Huitema, K.; van den Dikkenberg, J.; Brouwers, J.F.; Holthuis, J.C. Identification of a family of animal sphingomyelin synthases. EMBO J. 2004, 23, 33–44. [Google Scholar] [CrossRef]
- Liu, G.H.; Guan, T.; Datta, K.; Coppinger, J.; Yates, J., 3rd; Gerace, L. Regulation of myoblast differentiation by the nuclear envelope protein NET39. Mol. Cell Biol. 2009, 29, 5800–5812. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Smith, S.W.; Weiss, S.B.; Kennedy, E.P. The enzymatic dephosphorylation of phosphatidic acids. J. Biol. Chem. 1957, 228, 915–922. [Google Scholar] [PubMed]
- Coleman, R.; Huebscher, G. Metabolism of phospholipids. V. Studies of phosphatidic acid phosphatase. Biochim. Biophys. Acta 1962, 56, 479–490. [Google Scholar] [CrossRef]
- Kennedy, E.P.; Weiss, S.B. The function of cytidine coenzymes in the biosynthesis of phospholipides. J. Biol. Chem. 1956, 222, 193–214. [Google Scholar]
- Lamb, R.G.; Fallon, H.J. Glycerolipid formation from sn-glycerol-3-phosphate by rat liver cell fractions. The role of phosphatidate phosphohydrolase. Biochim. Biophys. Acta 1974, 348, 166–178. [Google Scholar] [CrossRef]
- Jamdar, S.C.; Fallon, H.J. Glycerolipid synthesis in rat adipose tissue. II. Properties and distribution of phosphatidate phosphatase. J. Lipid Res. 1973, 14, 517–524. [Google Scholar]
- Sturton, R.G.; Brindley, D.N. Factors controlling the metabolism of phosphatidate by phosphohydrolase and phospholipase A-type activities. Effects of magnesium, calcium and amphiphilic cationic drugs. Biochim. Biophys. Acta 1980, 619, 494–505. [Google Scholar] [CrossRef]
- Jamdar, S.C.; Osborne, L.J.; Wells, G.N. Glycerolipid biosynthesis in rat adipose tissue 12. Properties of Mg2+-dependent and -independent phosphatidate phosphohydrolase. Arch. Biochem. Biophys. 1984, 233, 370–377. [Google Scholar] [CrossRef]
- Smith, M.E.; Hubscher, G. The biosynthesis of glycerides by mitochondria from rat liver. The requirement for a soluble protein. Biochem. J. 1966, 101, 308–316. [Google Scholar] [CrossRef]
- Jelsema, C.L.; Morre, D.J. Distribution of phospholipid biosynthetic enzymes among cell components of rat liver. J. Biol. Chem. 1978, 253, 7960–7971. [Google Scholar]
- Han, G.S.; Wu, W.I.; Carman, G.M. The Saccharomyces cerevisiae Lipin homolog is a Mg2+-dependent phosphatidate phosphatase enzyme. J. Biol. Chem. 2006, 281, 9210–9218. [Google Scholar] [CrossRef]
- Kok, B.P.; Venkatraman, G.; Capatos, D.; Brindley, D.N. Unlike two peas in a pod: Lipid phosphate phosphatases and phosphatidate phosphatases. Chem. Rev. 2012, 112, 5121–5146. [Google Scholar] [CrossRef] [PubMed]
- Jamal, Z.; Martin, A.; Gomez-Munoz, A.; Brindley, D.N. Plasma membrane fractions from rat liver contain a phosphatidate phosphohydrolase distinct from that in the endoplasmic reticulum and cytosol. J. Biol. Chem. 1991, 266, 2988–2996. [Google Scholar] [PubMed]
- Fleming, I.N.; Yeaman, S.J. Purification and characterization of N-ethylmaleimide-insensitive phosphatidic acid phosphohydrolase (PAP2) from rat liver. Biochem. J. 1995, 308, 983–989. [Google Scholar] [CrossRef] [PubMed]
- Waggoner, D.W.; Martin, A.; Dewald, J.; Gomez-Munoz, A.; Brindley, D.N. Purification and characterization of novel plasma membrane phosphatidate phosphohydrolase from rat liver. J. Biol. Chem. 1995, 270, 19422–19429. [Google Scholar] [CrossRef] [PubMed]
- Siess, E.A.; Hofstetter, M.M. Identification of phosphatidate phosphohydrolase purified from rat liver membranes on SDS-polyacrylamide gel electrophoresis. FEBS Lett. 1996, 381, 169–173. [Google Scholar] [CrossRef]
- Brindley, D.N.; Waggoner, D.W. Mammalian lipid phosphate phosphohydrolases. J. Biol. Chem. 1998, 273, 24281–24284. [Google Scholar] [CrossRef]
- Kai, M.; Wada, I.; Imai, S.; Sakane, F.; Kanoh, H. Cloning and characterization of two human isozymes of Mg2+-independent phosphatidic acid phosphatase. J. Biol. Chem. 1997, 272, 24572–24578. [Google Scholar] [CrossRef]
- Kai, M.; Wada, I.; Imai, S.; Sakane, F.; Kanoh, H. Identification and cDNA cloning of 35-kDa phosphatidic acid phosphatase (type 2) bound to plasma membranes. Polymerase chain reaction amplification of mouse H2O2-inducible hic53 clone yielded the cDNA encoding phosphatidic acid phosphatase. J. Biol. Chem. 1996, 271, 18931–18938. [Google Scholar] [CrossRef]
- Hooks, S.B.; Ragan, S.P.; Lynch, K.R. Identification of a novel human phosphatidic acid phosphatase type 2 isoform. FEBS Lett. 1998, 427, 188–192. [Google Scholar] [CrossRef]
- Tate, R.J.; Tolan, D.; Pyne, S. Molecular cloning of magnesium-independent type 2 phosphatidic acid phosphatases from airway smooth muscle. Cell. Signal. 1999, 11, 515–522. [Google Scholar] [CrossRef]
- Burnett, C.; Howard, K. Fly and mammalian lipid phosphate phosphatase isoforms differ in activity both in vitro and in vivo. EMBO Rep. 2003, 4, 793–799. [Google Scholar] [CrossRef] [PubMed]
- Stukey, J.; Carman, G.M. Identification of a novel phosphatase sequence motif. Protein Sci. 1997, 6, 469–472. [Google Scholar] [CrossRef] [PubMed]
- Toke, D.A.; Bennett, W.L.; Oshiro, J.; Wu, W.I.; Voelker, D.R.; Carman, G.M. Isolation and characterization of the Saccharomyces cerevisiae LPP1 gene encoding a Mg2+-independent phosphatidate phosphatase. J. Biol. Chem. 1998, 273, 14331–14338. [Google Scholar] [CrossRef] [PubMed]
- Fagerberg, L.; Hallstrom, B.M.; Oksvold, P.; Kampf, C.; Djureinovic, D.; Odeberg, J.; Habuka, M.; Tahmasebpoor, S.; Danielsson, A.; Edlund, K.; et al. Analysis of the human tissue-specific expression by genome-wide integration of transcriptomics and antibody-based proteomics. Mol. Cell Proteom. 2014, 13, 397–406. [Google Scholar] [CrossRef]
- Uhlen, M.; Fagerberg, L.; Hallstrom, B.M.; Lindskog, C.; Oksvold, P.; Mardinoglu, A.; Sivertsson, A.; Kampf, C.; Sjostedt, E.; Asplund, A.; et al. Proteomics. Tissue-based map of the human proteome. Science 2015, 347, 1260419. [Google Scholar] [CrossRef]
- Jasinska, R.; Zhang, Q.X.; Pilquil, C.; Singh, I.; Xu, J.; Dewald, J.; Dillon, D.A.; Berthiaume, L.G.; Carman, G.M.; Waggoner, D.W.; et al. Lipid phosphate phosphohydrolase-1 degrades exogenous glycerolipid and sphingolipid phosphate esters. Biochem. J. 1999, 340, 677–686. [Google Scholar] [CrossRef]
- Sciorra, V.A.; Morris, A.J. Sequential actions of phospholipase D and phosphatidic acid phosphohydrolase 2b generate diglyceride in mammalian cells. Mol. Biol. Cell 1999, 10, 3863–3876. [Google Scholar] [CrossRef]
- Nanjundan, M.; Possmayer, F. Pulmonary lipid phosphate phosphohydrolase in plasma membrane signalling platforms. Biochem. J. 2001, 358, 637–646. [Google Scholar] [CrossRef]
- Jia, Y.J.; Kai, M.; Wada, I.; Sakane, F.; Kanoh, H. Differential localization of lipid phosphate phosphatases 1 and 3 to cell surface subdomains in polarized MDCK cells. FEBS Lett. 2003, 552, 240–246. [Google Scholar] [CrossRef]
- Barilà, D.; Plateroti, M.; Nobili, F.; Muda, A.O.; Xie, Y.; Morimoto, T.; Perozzi, G. The Dri 42 gene, whose expression is up-regulated during epithelial differentiation, encodes a novel endoplasmic reticulum resident transmembrane protein. J. Biol. Chem. 1996, 271, 29928–29936. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.X.; Pilquil, C.S.; Dewald, J.; Berthiaume, L.G.; Brindley, D.N. Identification of structurally important domains of lipid phosphate phosphatase-1: Implications for its sites of action. Biochem. J. 2000, 345, 181–184. [Google Scholar] [CrossRef] [PubMed]
- Sigal, Y.J.; McDermott, M.I.; Morris, A.J. Integral membrane lipid phosphatases/phosphotransferases: Common structure and diverse functions. Biochem. J. 2005, 387, 281–293. [Google Scholar] [CrossRef] [PubMed]
- Messerschmidt, A.; Wever, R. X-ray structure of a vanadium-containing enzyme: Chloroperoxidase from the fungus Curvularia inaequalis. Proc. Natl. Acad. Sci. USA 1996, 93, 392–396. [Google Scholar] [CrossRef]
- Messerschmidt, A.; Prade, L.; Wever, R. Implications for the catalytic mechanism of the vanadium-containing enzyme chloroperoxidase from the fungus Curvularia inaequalis by X-ray structures of the native and peroxide form. Biol. Chem. 1997, 378, 309–315. [Google Scholar] [CrossRef]
- Miriyala, S.; Subramanian, T.; Panchatcharam, M.; Ren, H.; McDermott, M.I.; Sunkara, M.; Drennan, T.; Smyth, S.S.; Spielmann, H.P.; Morris, A.J. Functional characterization of the atypical integral membrane lipid phosphatase PDP1/PPAPDC2 identifies a pathway for interconversion of isoprenols and isoprenoid phosphates in mammalian cells. J. Biol. Chem. 2010, 285, 13918–13929. [Google Scholar] [CrossRef]
- Fan, J.; Jiang, D.; Zhao, Y.; Liu, J.; Zhang, X.C. Crystal structure of lipid phosphatase Escherichia coli phosphatidylglycerophosphate phosphatase B. Proc. Natl. Acad. Sci. USA 2014, 111, 7636–7640. [Google Scholar] [CrossRef]
- Umezu-Goto, M.; Kishi, Y.; Taira, A.; Hama, K.; Dohmae, N.; Takio, K.; Yamori, T.; Mills, G.B.; Inoue, K.; Aoki, J.; et al. Autotaxin has lysophospholipase D activity leading to tumor cell growth and motility by lysophosphatidic acid production. J. Cell Biol. 2002, 158, 227–233. [Google Scholar] [CrossRef]
- Tokumura, A.; Majima, E.; Kariya, Y.; Tominaga, K.; Kogure, K.; Yasuda, K.; Fukuzawa, K. Identification of human plasma lysophospholipase D, a lysophosphatidic acid-producing enzyme, as autotaxin, a multifunctional phosphodiesterase. J. Biol. Chem. 2002, 277, 39436–39442. [Google Scholar] [CrossRef]
- Brindley, D.N.; Lin, F.T.; Tigyi, G.J. Role of the autotaxin-lysophosphatidate axis in cancer resistance to chemotherapy and radiotherapy. Biochim. Biophys. Acta 2013, 1831, 74–85. [Google Scholar] [CrossRef]
- Samadi, N.; Bekele, R.; Capatos, D.; Venkatraman, G.; Sariahmetoglu, M.; Brindley, D.N. Regulation of lysophosphatidate signaling by autotaxin and lipid phosphate phosphatases with respect to tumor progression, angiogenesis, metastasis and chemo-resistance. Biochimie 2011, 93, 61–70. [Google Scholar] [CrossRef] [PubMed]
- Strub, G.M.; Paillard, M.; Liang, J.; Gomez, L.; Allegood, J.C.; Hait, N.C.; Maceyka, M.; Price, M.M.; Chen, Q.; Simpson, D.C.; et al. Sphingosine-1-phosphate produced by sphingosine kinase 2 in mitochondria interacts with prohibitin 2 to regulate complex IV assembly and respiration. FASEB J. 2011, 25, 600–612. [Google Scholar] [CrossRef] [PubMed]
- Hait, N.C.; Allegood, J.; Maceyka, M.; Strub, G.M.; Harikumar, K.B.; Singh, S.K.; Luo, C.; Marmorstein, R.; Kordula, T.; Milstien, S.; et al. Regulation of histone acetylation in the nucleus by sphingosine-1-phosphate. Science 2009, 325, 1254–1257. [Google Scholar] [CrossRef] [PubMed]
- Takabe, K.; Paugh, S.W.; Milstien, S.; Spiegel, S. “Inside-out” signaling of sphingosine-1-phosphate: Therapeutic targets. Pharmacol. Rev. 2008, 60, 181–195. [Google Scholar] [CrossRef]
- Sato, K.; Malchinkhuu, E.; Horiuchi, Y.; Mogi, C.; Tomura, H.; Tosaka, M.; Yoshimoto, Y.; Kuwabara, A.; Okajima, F. Critical role of ABCA1 transporter in sphingosine 1-phosphate release from astrocytes. J. Neurochem. 2007, 103, 2610–2619. [Google Scholar] [CrossRef]
- Kobayashi, N.; Nishi, T.; Hirata, T.; Kihara, A.; Sano, T.; Igarashi, Y.; Yamaguchi, A. Sphingosine 1-phosphate is released from the cytosol of rat platelets in a carrier-mediated manner. J. Lipid Res. 2006, 47, 614–621. [Google Scholar] [CrossRef]
- Ulrych, T.; Bohm, A.; Polzin, A.; Daum, G.; Nusing, R.M.; Geisslinger, G.; Hohlfeld, T.; Schror, K.; Rauch, B.H. Release of sphingosine-1-phosphate from human platelets is dependent on thromboxane formation. J. Thromb. Haemost. 2011, 9, 790–798. [Google Scholar] [CrossRef]
- Nagahashi, M.; Kim, E.Y.; Yamada, A.; Ramachandran, S.; Allegood, J.C.; Hait, N.C.; Maceyka, M.; Milstien, S.; Takabe, K.; Spiegel, S. Spns2, a transporter of phosphorylated sphingoid bases, regulates their blood and lymph levels, and the lymphatic network. FASEB J. 2013, 27, 1001–1011. [Google Scholar] [CrossRef]
- Vu, T.M.; Ishizu, A.N.; Foo, J.C.; Toh, X.R.; Zhang, F.; Whee, D.M.; Torta, F.; Cazenave-Gassiot, A.; Matsumura, T.; Kim, S.; et al. Mfsd2b is essential for the sphingosine-1-phosphate export in erythrocytes and platelets. Nature 2017, 550, 524–528. [Google Scholar] [CrossRef]
- English, D.; Welch, Z.; Kovala, A.T.; Harvey, K.; Volpert, O.V.; Brindley, D.N.; Garcia, J.G. Sphingosine 1-phosphate released from platelets during clotting accounts for the potent endothelial cell chemotactic activity of blood serum and provides a novel link between hemostasis and angiogenesis. FASEB J. 2000, 14, 2255–2265. [Google Scholar] [CrossRef]
- Yung, Y.C.; Stoddard, N.C.; Chun, J. LPA receptor signaling: Pharmacology, physiology, and pathophysiology. J. Lipid Res. 2014, 55, 1192–1214. [Google Scholar] [CrossRef] [PubMed]
- Tigyi, G.J.; Yue, J.; Norman, D.D.; Szabo, E.; Balogh, A.; Balazs, L.; Zhao, G.; Lee, S.C. Regulation of tumor cell-Microenvironment interaction by the autotaxin-lysophosphatidic acid receptor axis. Adv. Biol. Regul. 2019, 71, 183–193. [Google Scholar] [CrossRef] [PubMed]
- Long, J.S.; Yokoyama, K.; Tigyi, G.; Pyne, N.J.; Pyne, S. Lipid phosphate phosphatase-1 regulates lysophosphatidic acid- and platelet-derived-growth-factor-induced cell migration. Biochem. J. 2006, 394, 495–500. [Google Scholar] [CrossRef] [PubMed]
- Pilquil, C.; Dewald, J.; Cherney, A.; Gorshkova, I.; Tigyi, G.; English, D.; Natarajan, V.; Brindley, D.N. Lipid phosphate phosphatase-1 regulates lysophosphatidate-induced fibroblast migration by controlling phospholipase D2-dependent phosphatidate generation. J. Biol. Chem. 2006, 281, 38418–38429. [Google Scholar] [CrossRef]
- Alderton, F.; Darroch, P.; Sambi, B.; McKie, A.; Ahmed, I.S.; Pyne, N.; Pyne, S. G-protein-coupled receptor stimulation of the p42/p44 mitogen-activated protein kinase pathway is attenuated by lipid phosphate phosphatases 1, 1a, and 2 in human embryonic kidney 293 cells. J. Biol. Chem. 2001, 276, 13452–13460. [Google Scholar] [CrossRef]
- Bektas, M.; Payne, S.G.; Liu, H.; Goparaju, S.; Milstien, S.; Spiegel, S. A novel acylglycerol kinase that produces lysophosphatidic acid modulates cross talk with EGFR in prostate cancer cells. J. Cell Biol. 2005, 169, 801–811. [Google Scholar] [CrossRef]
- Gobeil, F., Jr.; Bernier, S.G.; Vazquez-Tello, A.; Brault, S.; Beauchamp, M.H.; Quiniou, C.; Marrache, A.M.; Checchin, D.; Sennlaub, F.; Hou, X.; et al. Modulation of pro-inflammatory gene expression by nuclear lysophosphatidic acid receptor type-1. J. Biol. Chem. 2003, 278, 38875–38883. [Google Scholar] [CrossRef]
- Zhang, C.; Baker, D.L.; Yasuda, S.; Makarova, N.; Balazs, L.; Johnson, L.R.; Marathe, G.K.; McIntyre, T.M.; Xu, Y.; Prestwich, G.D.; et al. Lysophosphatidic acid induces neointima formation through PPARgamma activation. J. Exp. Med. 2004, 199, 763–774. [Google Scholar] [CrossRef]
- Tang, X.; Zhao, Y.Y.; Dewald, J.; Curtis, J.M.; Brindley, D.N. Tetracyclines increase lipid phosphate phosphatase expression on plasma membranes and turnover of plasma lysophosphatidate. J. Lipid Res. 2016, 57, 597–606. [Google Scholar] [CrossRef]
- Nakayama, J.; Raines, T.A.; Lynch, K.R.; Slack-Davis, J.K. Decreased peritoneal ovarian cancer growth in mice lacking expression of lipid phosphate phosphohydrolase 1. PLoS ONE 2015, 10, e0120071. [Google Scholar] [CrossRef]
- Tomsig, J.L.; Snyder, A.H.; Berdyshev, E.V.; Skobeleva, A.; Mataya, C.; Natarajan, V.; Brindley, D.N.; Lynch, K.R. Lipid phosphate phosphohydrolase type 1 (LPP1) degrades extracellular lysophosphatidic acid in vivo. Biochem. J. 2009, 419, 611–618. [Google Scholar] [CrossRef] [PubMed]
- Yue, J.; Yokoyama, K.; Balazs, L.; Baker, D.L.; Smalley, D.; Pilquil, C.; Brindley, D.N.; Tigyi, G. Mice with transgenic overexpression of lipid phosphate phosphatase-1 display multiple organotypic deficits without alteration in circulating lysophosphatidate level. Cell. Signal. 2004, 16, 385–399. [Google Scholar] [CrossRef] [PubMed]
- Jesionowska, A.; Cecerska-Heryc, E.; Matoszka, N.; Dolegowska, B. Lysophosphatidic acid signaling in ovarian cancer. J. Recept. Signal. Transduct. Res. 2015, 35, 578–584. [Google Scholar] [CrossRef]
- Fang, X.; Schummer, M.; Mao, M.; Yu, S.; Tabassam, F.H.; Swaby, R.; Hasegawa, Y.; Tanyi, J.L.; LaPushin, R.; Eder, A.; et al. Lysophosphatidic acid is a bioactive mediator in ovarian cancer. Biochim. Biophys. Acta 2002, 1582, 257–264. [Google Scholar] [CrossRef]
- Baker, D.L.; Morrison, P.; Miller, B.; Riely, C.A.; Tolley, B.; Westermann, A.M.; Bonfrer, J.M.; Bais, E.; Moolenaar, W.H.; Tigyi, G. Plasma lysophosphatidic acid concentration and ovarian cancer. JAMA 2002, 287, 3081–3082. [Google Scholar] [CrossRef] [PubMed]
- Peest, U.; Sensken, S.C.; Andreani, P.; Hanel, P.; Van Veldhoven, P.P.; Graler, M.H. S1P-lyase independent clearance of extracellular sphingosine 1-phosphate after dephosphorylation and cellular uptake. J. Cell Biochem. 2008, 104, 756–772. [Google Scholar] [CrossRef] [PubMed]
- Mandala, S.M. Sphingosine-1-phosphate phosphatases. Prostaglandins Other Lipid Mediat. 2001, 64, 143–156. [Google Scholar] [CrossRef]
- Le Stunff, H.; Peterson, C.; Liu, H.; Milstien, S.; Spiegel, S. Sphingosine-1-phosphate and lipid phosphohydrolases. Biochim. Biophys. Acta 2002, 1582, 8–17. [Google Scholar] [CrossRef]
- Zhao, Y.; Kalari, S.K.; Usatyuk, P.V.; Gorshkova, I.; He, D.; Watkins, T.; Brindley, D.N.; Sun, C.; Bittman, R.; Garcia, J.G.; et al. Intracellular generation of sphingosine 1-phosphate in human lung endothelial cells: Role of lipid phosphate phosphatase-1 and sphingosine kinase 1. J. Biol. Chem. 2007, 282, 14165–14177. [Google Scholar] [CrossRef]
- Breart, B.; Ramos-Perez, W.D.; Mendoza, A.; Salous, A.K.; Gobert, M.; Huang, Y.; Adams, R.H.; Lafaille, J.J.; Escalante-Alcalde, D.; Morris, A.J.; et al. Lipid phosphate phosphatase 3 enables efficient thymic egress. J. Exp. Med. 2011, 208, 1267–1278. [Google Scholar] [CrossRef]
- Lopez-Juarez, A.; Morales-Lazaro, S.; Sanchez-Sanchez, R.; Sunkara, M.; Lomeli, H.; Velasco, I.; Morris, A.J.; Escalante-Alcalde, D. Expression of LPP3 in Bergmann glia is required for proper cerebellar sphingosine-1-phosphate metabolism/signaling and development. Glia 2011, 59, 577–589. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Yamanaka, M.; Anada, Y.; Igarashi, Y.; Kihara, A. A splicing isoform of LPP1, LPP1a, exhibits high phosphatase activity toward FTY720 phosphate. Biochem. Biophys. Res. Commun. 2008, 375, 675–679. [Google Scholar] [CrossRef] [PubMed]
- Mechtcheriakova, D.; Wlachos, A.; Sobanov, J.; Bornancin, F.; Zlabinger, G.; Baumruker, T.; Billich, A. FTY720-phosphate is dephosphorylated by lipid phosphate phosphatase 3. FEBS Lett. 2007, 581, 3063–3068. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.; Benesch, M.G.; Dewald, J.; Zhao, Y.Y.; Patwardhan, N.; Santos, W.L.; Curtis, J.M.; McMullen, T.P.; Brindley, D.N. Lipid phosphate phosphatase-1 expression in cancer cells attenuates tumor growth and metastasis in mice. J. Lipid Res. 2014, 55, 2389–2400. [Google Scholar] [CrossRef]
- Nishioka, T.; Frohman, M.A.; Matsuda, M.; Kiyokawa, E. Heterogeneity of phosphatidic acid levels and distribution at the plasma membrane in living cells as visualized by a Foster resonance energy transfer (FRET) biosensor. J. Biol. Chem. 2010, 285, 35979–35987. [Google Scholar] [CrossRef]
- Rizzo, M.A.; Shome, K.; Vasudevan, C.; Stolz, D.B.; Sung, T.C.; Frohman, M.A.; Watkins, S.C.; Romero, G. Phospholipase D and its product, phosphatidic acid, mediate agonist-dependent raf-1 translocation to the plasma membrane and the activation of the mitogen-activated protein kinase pathway. J. Biol. Chem. 1999, 274, 1131–1139. [Google Scholar] [CrossRef]
- Andresen, B.T.; Rizzo, M.A.; Shome, K.; Romero, G. The role of phosphatidic acid in the regulation of the Ras/MEK/Erk signaling cascade. FEBS Lett. 2002, 531, 65–68. [Google Scholar] [CrossRef]
- Brindley, D.N.; Pilquil, C. Lipid phosphate phosphatases and signaling. J. Lipid Res. 2009, 50, S225–S230. [Google Scholar] [CrossRef]
- Alderton, F.; Rakhit, S.; Kong, K.C.; Palmer, T.; Sambi, B.; Pyne, S.; Pyne, N.J. Tethering of the platelet-derived growth factor beta receptor to G-protein-coupled receptors. A novel platform for integrative signaling by these receptor classes in mammalian cells. J. Biol. Chem. 2001, 276, 28578–28585. [Google Scholar] [CrossRef]
- Gutierrez-Martinez, E.; Fernandez-Ulibarri, I.; Lazaro-Dieguez, F.; Johannes, L.; Pyne, S.; Sarri, E.; Egea, G. Lipid phosphate phosphatase 3 participates in transport carrier formation and protein trafficking in the early secretory pathway. J. Cell Sci. 2013, 126, 2641–2655. [Google Scholar] [CrossRef]
- Long, J.; Darroch, P.; Wan, K.F.; Kong, K.C.; Ktistakis, N.; Pyne, N.J.; Pyne, S. Regulation of cell survival by lipid phosphate phosphatases involves the modulation of intracellular phosphatidic acid and sphingosine 1-phosphate pools. Biochem. J. 2005, 391, 25–32. [Google Scholar] [CrossRef] [PubMed]
- Roberts, R.Z.; Morris, A.J. Role of phosphatidic acid phosphatase 2a in uptake of extracellular lipid phosphate mediators. Biochim. Biophys. Acta 2000, 1487, 33–49. [Google Scholar] [CrossRef]
- Brindley, D.N.; Pilquil, C.; Sariahmetoglu, M.; Reue, K. Phosphatidate degradation: Phosphatidate phosphatases (lipins) and lipid phosphate phosphatases. Biochim. Biophys. Acta 2009, 1791, 956–961. [Google Scholar] [CrossRef]
- Pettus, B.J.; Kitatani, K.; Chalfant, C.E.; Taha, T.A.; Kawamori, T.; Bielawski, J.; Obeid, L.M.; Hannun, Y.A. The coordination of prostaglandin E2 production by sphingosine-1-phosphate and ceramide-1-phosphate. Mol. Pharmacol. 2005, 68, 330–335. [Google Scholar] [CrossRef] [PubMed]
- Pyne, S.; Adams, D.R.; Pyne, N.J. Sphingosine 1-phosphate and sphingosine kinases in health and disease: Recent advances. Prog. Lipid Res. 2016, 62, 93–106. [Google Scholar] [CrossRef]
- Balazs, L.; Okolicany, J.; Ferrebee, M.; Tigyi, G. Topical application of LPA accelerates wound healing. Ann. N. Y. Acad. Sci. 2000, 905, 270–273. [Google Scholar] [CrossRef]
- van Corven, E.J.; van Rijswijk, A.; Jalink, K.; van der Bend, R.L.; van Blitterswijk, W.J.; Moolenaar, W.H. Mitogenic action of lysophosphatidic acid and phosphatidic acid on fibroblasts. Dependence on acyl-chain length and inhibition by suramin. Biochem. J. 1992, 281, 163–169. [Google Scholar] [CrossRef]
- Chabaud, S.; Marcoux, T.L.; Deschenes-Rompre, M.P.; Rousseau, A.; Morissette, A.; Bouhout, S.; Bernard, G.; Bolduc, S. Lysophosphatidic acid enhances collagen deposition and matrix thickening in engineered tissue. J. Tissue Eng. Regen. Med. 2015, 9, E65–E75. [Google Scholar] [CrossRef]
- Van Corven, E.J.; Groenink, A.; Jalink, K.; Eichholtz, T.; Moolenaar, W.H. Lysophosphatidate-induced cell proliferation: Identification and dissection of signaling pathways mediated by G proteins. Cell 1989, 59, 45–54. [Google Scholar] [CrossRef]
- Panetti, T.S.; Chen, H.; Misenheimer, T.M.; Getzler, S.B.; Mosher, D.F. Endothelial cell mitogenesis induced by LPA: Inhibition by thrombospondin-1 and thrombospondin-2. J. Lab. Clin. Med. 1997, 129, 208–216. [Google Scholar] [CrossRef]
- Dvorak, H.F. Tumors: Wounds that do not heal-redux. Cancer Immunol. Res. 2015, 3, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Benesch, M.G.; Ko, Y.M.; Tang, X.; Dewald, J.; Lopez-Campistrous, A.; Zhao, Y.Y.; Lai, R.; Curtis, J.M.; Brindley, D.N.; McMullen, T.P. Autotaxin is an inflammatory mediator and therapeutic target in thyroid cancer. Endocr. Relat. Cancer 2015, 22, 593–607. [Google Scholar] [CrossRef]
- Magkrioti, C.; Oikonomou, N.; Kaffe, E.; Mouratis, M.A.; Xylourgidis, N.; Barbayianni, I.; Megadoukas, P.; Harokopos, V.; Valavanis, C.; Chun, J.; et al. The Autotaxin-Lysophosphatidic Acid Axis Promotes Lung Carcinogenesis. Cancer Res. 2018, 78, 3634–3644. [Google Scholar] [CrossRef]
- Shao, Y.; Yu, Y.; He, Y.; Chen, Q.; Liu, H. Serum ATX as a novel biomarker for breast cancer. Medicine (Baltimore) 2019, 98, e14973. [Google Scholar] [CrossRef] [PubMed]
- Memet, I.; Tsalkidou, E.; Tsaroucha, A.K.; Lambropoulou, M.; Chatzaki, E.; Trypsianis, G.; Schizas, D.; Pitiakoudis, M.; Simopoulos, C. Autotaxin Expression in Hepatocellular Carcinoma. J. Investig. Surg. 2018, 31, 359–365. [Google Scholar] [CrossRef] [PubMed]
- Auciello, F.R.; Bulusu, V.; Oon, C.; Tait-Mulder, J.; Berry, M.; Bhattacharyya, S.; Tumanov, S.; Allen-Petersen, B.L.; Link, J.; Kendsersky, N.D.; et al. A Stromal Lysolipid-Autotaxin Signaling Axis Promotes Pancreatic Tumor Progression. Cancer Discov. 2019, 9, 617–627. [Google Scholar] [CrossRef] [PubMed]
- Nakai, Y.; Ikeda, H.; Nakamura, K.; Kume, Y.; Fujishiro, M.; Sasahira, N.; Hirano, K.; Isayama, H.; Tada, M.; Kawabe, T.; et al. Specific increase in serum autotaxin activity in patients with pancreatic cancer. Clin. Biochem. 2011, 44, 576–581. [Google Scholar] [CrossRef]
- Xu, A.; Ahsanul Kabir Khan, M.; Chen, F.; Zhong, Z.; Chen, H.C.; Song, Y. Overexpression of autotaxin is associated with human renal cell carcinoma and bladder carcinoma and their progression. Med. Oncol. 2016, 33, 131. [Google Scholar] [CrossRef]
- Nouh, M.A.; Wu, X.X.; Okazoe, H.; Tsunemori, H.; Haba, R.; Abou-Zeid, A.M.; Saleem, M.D.; Inui, M.; Sugimoto, M.; Aoki, J.; et al. Expression of autotaxin and acylglycerol kinase in prostate cancer: Association with cancer development and progression. Cancer Sci. 2009, 100, 1631–1638. [Google Scholar] [CrossRef]
- Patterson, A.D.; Maurhofer, O.; Beyoglu, D.; Lanz, C.; Krausz, K.W.; Pabst, T.; Gonzalez, F.J.; Dufour, J.F.; Idle, J.R. Aberrant lipid metabolism in hepatocellular carcinoma revealed by plasma metabolomics and lipid profiling. Cancer Res. 2011, 71, 6590–6600. [Google Scholar] [CrossRef]
- Masuda, A.; Nakamura, K.; Izutsu, K.; Igarashi, K.; Ohkawa, R.; Jona, M.; Higashi, K.; Yokota, H.; Okudaira, S.; Kishimoto, T.; et al. Serum autotaxin measurement in haematological malignancies: A promising marker for follicular lymphoma. Br. J. Haematol. 2008, 143, 60–70. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Shen, Z.; Wiper, D.W.; Wu, M.; Morton, R.E.; Elson, P.; Kennedy, A.W.; Belinson, J.; Markman, M.; Casey, G. Lysophosphatidic acid as a potential biomarker for ovarian and other gynecologic cancers. JAMA 1998, 280, 719–723. [Google Scholar] [CrossRef] [PubMed]
- Zeng, R.; Li, B.; Huang, J.; Zhong, M.; Li, L.; Duan, C.; Zeng, S.; Liu, W.; Lu, J.; Tang, Y.; et al. Lysophosphatidic Acid is a Biomarker for Peritoneal Carcinomatosis of Gastric Cancer and Correlates with Poor Prognosis. Genet. Test. Mol. Biomark. 2017, 21, 641–648. [Google Scholar] [CrossRef] [PubMed]
- Riaz, A.; Huang, Y.; Johansson, S. G-Protein-Coupled Lysophosphatidic Acid Receptors and Their Regulation of AKT Signaling. Int. J. Mol. Sci. 2016, 17, 215. [Google Scholar] [CrossRef]
- Zhang, G.; Cheng, Y.; Zhang, Q.; Li, X.; Zhou, J.; Wang, J.; Wei, L. ATXLPA axis facilitates estrogeninduced endometrial cancer cell proliferation via MAPK/ERK signaling pathway. Mol. Med. Rep. 2018, 17, 4245–4252. [Google Scholar]
- Herr, D.R. Potential use of g protein-coupled receptor-blocking monoclonal antibodies as therapeutic agents for cancers. Int. Rev. Cell Mol. Biol. 2012, 297, 45–81. [Google Scholar]
- Kishi, Y.; Okudaira, S.; Tanaka, M.; Hama, K.; Shida, D.; Kitayama, J.; Yamori, T.; Aoki, J.; Fujimaki, T.; Arai, H. Autotaxin is overexpressed in glioblastoma multiforme and contributes to cell motility of glioblastoma by converting lysophosphatidylcholine to lysophosphatidic acid. J. Biol. Chem. 2006, 281, 17492–17500. [Google Scholar] [CrossRef]
- Manning, T.J., Jr.; Parker, J.C.; Sontheimer, H. Role of lysophosphatidic acid and rho in glioma cell motility. Cell Motil. Cytoskelet. 2000, 45, 185–199. [Google Scholar] [CrossRef]
- Yamada, T.; Sato, K.; Komachi, M.; Malchinkhuu, E.; Tobo, M.; Kimura, T.; Kuwabara, A.; Yanagita, Y.; Ikeya, T.; Tanahashi, Y.; et al. Lysophosphatidic acid (LPA) in malignant ascites stimulates motility of human pancreatic cancer cells through LPA1. J. Biol. Chem. 2004, 279, 6595–6605. [Google Scholar] [CrossRef]
- Gong, Y.L.; Tao, C.J.; Hu, M.; Chen, J.F.; Cao, X.F.; Lv, G.M.; Li, P. Expression of lysophosphatidic acid receptors and local invasiveness and metastasis in Chinese pancreatic cancers. Curr. Oncol. 2012, 19, eS15–eS21. [Google Scholar] [CrossRef]
- Kato, K.; Yoshikawa, K.; Tanabe, E.; Kitayoshi, M.; Fukui, R.; Fukushima, N.; Tsujiuchi, T. Opposite roles of LPA1 and LPA3 on cell motile and invasive activities of pancreatic cancer cells. Tumour. Biol. 2012, 33, 1739–1744. [Google Scholar] [CrossRef] [PubMed]
- Shida, D.; Watanabe, T.; Aoki, J.; Hama, K.; Kitayama, J.; Sonoda, H.; Kishi, Y.; Yamaguchi, H.; Sasaki, S.; Sako, A.; et al. Aberrant expression of lysophosphatidic acid (LPA) receptors in human colorectal cancer. Lab. Investig. 2004, 84, 1352–1362. [Google Scholar] [CrossRef] [PubMed]
- Yun, C.C.; Sun, H.; Wang, D.; Rusovici, R.; Castleberry, A.; Hall, R.A.; Shim, H. LPA2 receptor mediates mitogenic signals in human colon cancer cells. Am. J. Physiol. Cell Physiol. 2005, 289, C2–C11. [Google Scholar] [CrossRef] [PubMed]
- Sun, K.; Cai, H.; Duan, X.; Yang, Y.; Li, M.; Qu, J.; Zhang, X.; Wang, J. Aberrant expression and potential therapeutic target of lysophosphatidic acid receptor 3 in triple-negative breast cancers. Clin. Exp. Med. 2015, 15, 371–380. [Google Scholar] [CrossRef] [PubMed]
- Li, T.T.; Alemayehu, M.; Aziziyeh, A.I.; Pape, C.; Pampillo, M.; Postovit, L.M.; Mills, G.B.; Babwah, A.V.; Bhattacharya, M. Beta-arrestin/Ral signaling regulates lysophosphatidic acid-mediated migration and invasion of human breast tumor cells. Mol. Cancer Res. 2009, 7, 1064–1077. [Google Scholar] [CrossRef] [PubMed]
- Burkhalter, R.J.; Westfall, S.D.; Liu, Y.; Stack, M.S. Lysophosphatidic Acid Initiates Epithelial to Mesenchymal Transition and Induces beta-Catenin-mediated Transcription in Epithelial Ovarian Carcinoma. J. Biol. Chem. 2015, 290, 22143–22154. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.; Liu, Z.; Wang, C.; Yao, B.; Zheng, X. EDG2 enhanced the progression of hepatocellular carcinoma by LPA/PI3K/AKT/mTOR signaling. Oncotarget 2017, 8, 66154–66168. [Google Scholar] [CrossRef]
- Shibue, T.; Weinberg, R.A. EMT, CSCs, and drug resistance: The mechanistic link and clinical implications. Nat. Rev. Clin. Oncol. 2017, 14, 611–629. [Google Scholar] [CrossRef]
- Seo, E.J.; Kwon, Y.W.; Jang, I.H.; Kim, D.K.; Lee, S.I.; Choi, E.J.; Kim, K.H.; Suh, D.S.; Lee, J.H.; Choi, K.U.; et al. Autotaxin Regulates Maintenance of Ovarian Cancer Stem Cells through Lysophosphatidic Acid-Mediated Autocrine Mechanism. Stem Cells 2016, 34, 551–564. [Google Scholar] [CrossRef]
- Banerjee, S.; Norman, D.D.; Lee, S.C.; Parrill, A.L.; Pham, T.C.; Baker, D.L.; Tigyi, G.J.; Miller, D.D. Highly Potent Non-Carboxylic Acid Autotaxin Inhibitors Reduce Melanoma Metastasis and Chemotherapeutic Resistance of Breast Cancer Stem Cells. J. Med. Chem. 2017, 60, 1309–1324. [Google Scholar] [CrossRef]
- Gupta, P.B.; Onder, T.T.; Jiang, G.; Tao, K.; Kuperwasser, C.; Weinberg, R.A.; Lander, E.S. Identification of selective inhibitors of cancer stem cells by high-throughput screening. Cell 2009, 138, 645–659. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Umezu-Goto, M.; Murph, M.; Lu, Y.; Liu, W.; Zhang, F.; Yu, S.; Stephens, L.C.; Cui, X.; Murrow, G.; et al. Expression of autotaxin and lysophosphatidic acid receptors increases mammary tumorigenesis, invasion, and metastases. Cancer Cell 2009, 15, 539–550. [Google Scholar] [CrossRef] [PubMed]
- Lee, Z.; Cheng, C.T.; Zhang, H.; Subler, M.A.; Wu, J.; Mukherjee, A.; Windle, J.J.; Chen, C.K.; Fang, X. Role of LPA4/p2y9/GPR23 in negative regulation of cell motility. Mol. Biol. Cell 2008, 19, 5435–5445. [Google Scholar] [CrossRef] [PubMed]
- Ishii, S.; Hirane, M.; Fukushima, K.; Tomimatsu, A.; Fukushima, N.; Tsujiuchi, T. Diverse effects of LPA4, LPA5 and LPA6 on the activation of tumor progression in pancreatic cancer cells. Biochem. Biophys. Res. Commun. 2015, 461, 59–64. [Google Scholar] [CrossRef]
- Jongsma, M.; Matas-Rico, E.; Rzadkowski, A.; Jalink, K.; Moolenaar, W.H. LPA is a chemorepellent for B16 melanoma cells: Action through the cAMP-elevating LPA5 receptor. PLoS ONE 2011, 6, e29260. [Google Scholar] [CrossRef]
- Lee, S.C.; Fujiwara, Y.; Liu, J.; Yue, J.; Shimizu, Y.; Norman, D.D.; Wang, Y.; Tsukahara, R.; Szabo, E.; Patil, R.; et al. Autotaxin and LPA1 and LPA5 receptors exert disparate functions in tumor cells versus the host tissue microenvironment in melanoma invasion and metastasis. Mol. Cancer Res. 2015, 13, 174–185. [Google Scholar] [CrossRef]
- Mathew, D.; Kremer, K.N.; Strauch, P.; Tigyi, G.; Pelanda, R.; Torres, R.M. LPA5 Is an Inhibitory Receptor That Suppresses CD8 T-Cell Cytotoxic Function via Disruption of Early TCR Signaling. Front. Immunol. 2019, 10, 1159. [Google Scholar] [CrossRef]
- Enooku, K.; Uranbileg, B.; Ikeda, H.; Kurano, M.; Sato, M.; Kudo, H.; Maki, H.; Koike, K.; Hasegawa, K.; Kokudo, N.; et al. Higher LPA2 and LPA6 mRNA Levels in Hepatocellular Carcinoma Are Associated with Poorer Differentiation, Microvascular Invasion and Earlier Recurrence with Higher Serum Autotaxin Levels. PLoS ONE 2016, 11, e0161825. [Google Scholar] [CrossRef]
- Takahashi, K.; Fukushima, K.; Otagaki, S.; Ishimoto, K.; Minami, K.; Fukushima, N.; Honoki, K.; Tsujiuchi, T. Effects of LPA1 and LPA6 on the regulation of colony formation activity in colon cancer cells treated with anticancer drugs. J. Recept. Signal. Transduct. Res. 2018, 38, 71–75. [Google Scholar] [CrossRef]
- Knowlden, S.; Georas, S.N. The autotaxin-LPA axis emerges as a novel regulator of lymphocyte homing and inflammation. J. Immunol. 2014, 192, 851–857. [Google Scholar] [CrossRef]
- Ray, R.; Rai, V. Lysophosphatidic acid converts monocytes into macrophages in both mice and humans. Blood 2017, 129, 1177–1183. [Google Scholar] [CrossRef] [PubMed]
- Benesch, M.G.; Ko, Y.M.; McMullen, T.P.; Brindley, D.N. Autotaxin in the crosshairs: Taking aim at cancer and other inflammatory conditions. FEBS Lett. 2014, 588, 2712–2727. [Google Scholar] [CrossRef] [PubMed]
- Benesch, M.G.K.; Yang, Z.; Tang, X.; Meng, G.; Brindley, D.N. Lysophosphatidate Signaling: The Tumor Microenvironment’s New Nemesis. Trends Cancer 2017, 3, 748–752. [Google Scholar] [CrossRef] [PubMed]
- Colotta, F.; Allavena, P.; Sica, A.; Garlanda, C.; Mantovani, A. Cancer-related inflammation, the seventh hallmark of cancer: Links to genetic instability. Carcinogenesis 2009, 30, 1073–1081. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.M.; Xu, Y.; Skill, N.J.; Sheng, H.; Zhao, Z.; Yu, M.; Saxena, R.; Maluccio, M.A. Autotaxin expression and its connection with the TNF-alpha-NF-kappaB axis in human hepatocellular carcinoma. Mol. Cancer 2010, 9, 71. [Google Scholar] [CrossRef] [PubMed]
- Cummings, R.; Zhao, Y.; Jacoby, D.; Spannhake, E.W.; Ohba, M.; Garcia, J.G.; Watkins, T.; He, D.; Saatian, B.; Natarajan, V. Protein kinase Cdelta mediates lysophosphatidic acid-induced NF-kappaB activation and interleukin-8 secretion in human bronchial epithelial cells. J. Biol. Chem. 2004, 279, 41085–41094. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; He, D.; Saatian, B.; Watkins, T.; Spannhake, E.W.; Pyne, N.J.; Natarajan, V. Regulation of lysophosphatidic acid-induced epidermal growth factor receptor transactivation and interleukin-8 secretion in human bronchial epithelial cells by protein kinase Cdelta, Lyn kinase, and matrix metalloproteinases. J. Biol. Chem. 2006, 281, 19501–19511. [Google Scholar] [CrossRef]
- Yu, S.; Murph, M.M.; Lu, Y.; Liu, S.; Hall, H.S.; Liu, J.; Stephens, C.; Fang, X.; Mills, G.B. Lysophosphatidic acid receptors determine tumorigenicity and aggressiveness of ovarian cancer cells. J. Natl. Cancer Inst. 2008, 100, 1630–1642. [Google Scholar] [CrossRef]
- Lin, S.; Wang, D.; Iyer, S.; Ghaleb, A.M.; Shim, H.; Yang, V.W.; Chun, J.; Yun, C.C. The absence of LPA2 attenuates tumor formation in an experimental model of colitis-associated cancer. Gastroenterology 2009, 136, 1711–1720. [Google Scholar] [CrossRef]
- Zhao, Y.; Usatyuk, P.V.; Cummings, R.; Saatian, B.; He, D.; Watkins, T.; Morris, A.; Spannhake, E.W.; Brindley, D.N.; Natarajan, V. Lipid phosphate phosphatase-1 regulates lysophosphatidic acid-induced calcium release, NF-kappaB activation and interleukin-8 secretion in human bronchial epithelial cells. Biochem. J. 2005, 385, 493–502. [Google Scholar] [CrossRef]
- Schmid, R.; Wolf, K.; Robering, J.W.; Strauss, S.; Strissel, P.L.; Strick, R.; Rubner, M.; Fasching, P.A.; Horch, R.E.; Kremer, A.E.; et al. ADSCs and adipocytes are the main producers in the autotaxin-lysophosphatidic acid axis of breast cancer and healthy mammary tissue in vitro. BMC Cancer 2018, 18, 1273. [Google Scholar] [CrossRef] [PubMed]
- Benesch, M.G.; Tang, X.; Dewald, J.; Dong, W.F.; Mackey, J.R.; Hemmings, D.G.; McMullen, T.P.; Brindley, D.N. Tumor-induced inflammation in mammary adipose tissue stimulates a vicious cycle of autotaxin expression and breast cancer progression. FASEB J. 2015, 29, 3990–4000. [Google Scholar] [CrossRef] [PubMed]
- Benesch, M.G.; Zhao, Y.Y.; Curtis, J.M.; McMullen, T.P.; Brindley, D.N. Regulation of autotaxin expression and secretion by lysophosphatidate and sphingosine 1-phosphate. J. Lipid Res. 2015, 56, 1134–1144. [Google Scholar] [CrossRef] [PubMed]
- Meng, G.; Tang, X.; Yang, Z.; Benesch, M.G.K.; Marshall, A.; Murray, D.; Hemmings, D.G.; Wuest, F.; McMullen, T.P.W.; Brindley, D.N. Implications for breast cancer treatment from increased autotaxin production in adipose tissue after radiotherapy. FASEB J. 2017, 31, 4064–4077. [Google Scholar] [CrossRef]
- Meng, G.; Wuest, M.; Tang, X.; Dufour, J.; Zhao, Y.; Curtis, J.M.; McMullen, T.P.W.; Murray, D.; Wuest, F.; Brindley, D.N. Repeated Fractions of X-Radiation to the Breast Fat Pads of Mice Augment Activation of the Autotaxin-Lysophosphatidate-Inflammatory Cycle. Cancers 2019, 11, 1816. [Google Scholar] [CrossRef]
- Meng, G.; Tang, X.; Yang, Z.; Zhao, Y.; Curtis, J.M.; McMullen, T.P.W.; Brindley, D.N. Dexamethasone decreases the autotaxin-lysophosphatidate-inflammatory axis in adipose tissue: Implications for the metabolic syndrome and breast cancer. FASEB J. 2019, 33, 1899–1910. [Google Scholar] [CrossRef]
- Meng, G.; Wuest, M.; Tang, X.; Dufour, J.; McMullen, T.P.W.; Wuest, F.; Murray, D.; Brindley, D.N. Dexamethasone Attenuates X-Ray-Induced Activation of the Autotaxin-Lysophosphatidate-Inflammatory Cycle in Breast Tissue and Subsequent Breast Fibrosis. Cancers 2020, 12, 999. [Google Scholar] [CrossRef]
- Shi, K.; Jiang, J.; Ma, T.; Xie, J.; Duan, L.; Chen, R.; Song, P.; Yu, Z.; Liu, C.; Zhu, Q.; et al. Dexamethasone attenuates bleomycin-induced lung fibrosis in mice through TGF-beta, Smad3 and JAK-STAT pathway. Int. J. Clin. Exp. Med. 2014, 7, 2645–2650. [Google Scholar]
- Chen, F.; Gong, L.; Zhang, L.; Wang, H.; Qi, X.; Wu, X.; Xiao, Y.; Cai, Y.; Liu, L.; Li, X.; et al. Short courses of low dose dexamethasone delay bleomycin-induced lung fibrosis in rats. Eur. J. Pharmacol. 2006, 536, 287–295. [Google Scholar] [CrossRef]
- Deng, W.; Wang, D.A.; Gosmanova, E.; Johnson, L.R.; Tigyi, G. LPA protects intestinal epithelial cells from apoptosis by inhibiting the mitochondrial pathway. Am. J. Physiol. Gastrointest. Liver Physiol. 2003, 284, G821–G829. [Google Scholar] [CrossRef]
- Meng, Y.; Kang, S.; Fishman, D.A. Lysophosphatidic acid inhibits anti-Fas-mediated apoptosis enhanced by actin depolymerization in epithelial ovarian cancer. FEBS Lett. 2005, 579, 1311–1319. [Google Scholar] [CrossRef] [PubMed]
- Sui, Y.; Yang, Y.; Wang, J.; Li, Y.; Ma, H.; Cai, H.; Liu, X.; Zhang, Y.; Wang, S.; Li, Z.; et al. Lysophosphatidic Acid Inhibits Apoptosis Induced by Cisplatin in Cervical Cancer Cells. Biomed. Res. Int. 2015, 2015, 598386. [Google Scholar] [CrossRef]
- Kang, Y.C.; Kim, K.M.; Lee, K.S.; Namkoong, S.; Lee, S.J.; Han, J.A.; Jeoung, D.; Ha, K.S.; Kwon, Y.G.; Kim, Y.M. Serum bioactive lysophospholipids prevent TRAIL-induced apoptosis via PI3K/Akt-dependent cFLIP expression and Bad phosphorylation. Cell Death Differ. 2004, 11, 1287–1298. [Google Scholar] [CrossRef] [PubMed]
- Rusovici, R.; Ghaleb, A.; Shim, H.; Yang, V.W.; Yun, C.C. Lysophosphatidic acid prevents apoptosis of Caco-2 colon cancer cells via activation of mitogen-activated protein kinase and phosphorylation of Bad. Biochim. Biophys. Acta 2007, 1770, 1194–1203. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Samadi, N.; Bekele, R.T.; Goping, I.S.; Schang, L.M.; Brindley, D.N. Lysophosphatidate induces chemo-resistance by releasing breast cancer cells from taxol-induced mitotic arrest. PLoS ONE 2011, 6, e20608. [Google Scholar] [CrossRef] [PubMed]
- Bekele, R.T.; Venkatraman, G.; Liu, R.Z.; Tang, X.; Mi, S.; Benesch, M.G.; Mackey, J.R.; Godbout, R.; Curtis, J.M.; McMullen, T.P.; et al. Oxidative stress contributes to the tamoxifen-induced killing of breast cancer cells: Implications for tamoxifen therapy and resistance. Sci. Rep. 2016, 6, 21164. [Google Scholar] [CrossRef]
- Venkatraman, G.; Benesch, M.G.; Tang, X.; Dewald, J.; McMullen, T.P.; Brindley, D.N. Lysophosphatidate signaling stabilizes Nrf2 and increases the expression of genes involved in drug resistance and oxidative stress responses: Implications for cancer treatment. FASEB J. 2015, 29, 772–785. [Google Scholar] [CrossRef]
- Deng, W.; Shuyu, E.; Tsukahara, R.; Valentine, W.J.; Durgam, G.; Gududuru, V.; Balazs, L.; Manickam, V.; Arsura, M.; VanMiddlesworth, L.; et al. The lysophosphatidic acid type 2 receptor is required for protection against radiation-induced intestinal injury. Gastroenterology 2007, 132, 1834–1851. [Google Scholar] [CrossRef]
- Tigyi, G.J.; Johnson, L.R.; Lee, S.C.; Norman, D.D.; Szabo, E.; Balogh, A.; Thompson, K.; Boler, A.; McCool, W.S. Lysophosphatidic acid type 2 receptor agonists in targeted drug development offer broad therapeutic potential. J. Lipid Res. 2019, 60, 464–474. [Google Scholar] [CrossRef]
- Hashimoto, S.; Mikami, S.; Sugino, H.; Yoshikawa, A.; Hashimoto, A.; Onodera, Y.; Furukawa, S.; Handa, H.; Oikawa, T.; Okada, Y.; et al. Lysophosphatidic acid activates Arf6 to promote the mesenchymal malignancy of renal cancer. Nat. Commun. 2016, 7, 10656. [Google Scholar] [CrossRef]
- Su, S.C.; Hu, X.; Kenney, P.A.; Merrill, M.M.; Babaian, K.N.; Zhang, X.Y.; Maity, T.; Yang, S.F.; Lin, X.; Wood, C.G. Autotaxin-lysophosphatidic acid signaling axis mediates tumorigenesis and development of acquired resistance to sunitinib in renal cell carcinoma. Clin. Cancer Res. 2013, 19, 6461–6472. [Google Scholar] [CrossRef] [PubMed]
- Fukushima, K.; Takahashi, K.; Yamasaki, E.; Onishi, Y.; Fukushima, N.; Honoki, K.; Tsujiuchi, T. Lysophosphatidic acid signaling via LPA1 and LPA3 regulates cellular functions during tumor progression in pancreatic cancer cells. Exp. Cell Res. 2017, 352, 139–145. [Google Scholar] [CrossRef]
- Nguyen, T.; Nioi, P.; Pickett, C.B. The Nrf2-antioxidant response element signaling pathway and its activation by oxidative stress. J. Biol. Chem. 2009, 284, 13291–13295. [Google Scholar] [CrossRef] [PubMed]
- Hemmings, D.G.; Brindley, D.N. Signalling by lysophosphatidate and its health implications. Essays Biochem. 2020. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.; Wuest, M.; Benesch, M.G.; Dufour, J.; Zhao, Y.; Curtis, J.M.; Monjardet, A.; Heckmann, B.; Murray, D.; Wuest, F.; et al. Inhibition of autotaxin with GLPG1690 increases the efficacy of radiotherapy and chemotherapy in a mouse model of breast cancer. Mol. Cancer Ther. 2019, 19, 63–74. [Google Scholar] [CrossRef]
- Kim, B.C.; Yoo, H.J.; Lee, H.C.; Kang, K.A.; Jung, S.H.; Lee, H.J.; Lee, M.; Park, S.; Ji, Y.H.; Lee, Y.S.; et al. Evaluation of premature senescence and senescence biomarkers in carcinoma cells and xenograft mice exposed to single or fractionated irradiation. Oncol. Rep. 2014, 31, 2229–2235. [Google Scholar] [CrossRef][Green Version]
- Jones, K.R.; Elmore, L.W.; Jackson-Cook, C.; Demasters, G.; Povirk, L.F.; Holt, S.E.; Gewirtz, D.A. p53-Dependent accelerated senescence induced by ionizing radiation in breast tumour cells. Int. J. Radiat. Biol. 2005, 81, 445–458. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Guo, X.; Gao, Y.; Lu, D.; Li, W. Tumor Cell-Accelerated Senescence Is Associated With DNA-PKcs Status and Telomere Dysfunction Induced by Radiation. Dose Response 2018, 16, 1559325818771527. [Google Scholar] [CrossRef]
- Pyne, N.J.; El Buri, A.; Adams, D.R.; Pyne, S. Sphingosine 1-phosphate and cancer. Adv. Biol. Regul. 2018, 68, 97–106. [Google Scholar] [CrossRef] [PubMed]
- Heffernan-Stroud, L.A.; Obeid, L.M. Sphingosine kinase 1 in cancer. Adv. Cancer Res. 2013, 117, 201–235. [Google Scholar]
- Hatoum, D.; Haddadi, N.; Lin, Y.; Nassif, N.T.; McGowan, E.M. Mammalian sphingosine kinase (SphK) isoenzymes and isoform expression: Challenges for SphK as an oncotarget. Oncotarget 2017, 8, 36898–36929. [Google Scholar] [CrossRef]
- Xia, P.; Gamble, J.R.; Wang, L.; Pitson, S.M.; Moretti, P.A.; Wattenberg, B.W.; D’Andrea, R.J.; Vadas, M.A. An oncogenic role of sphingosine kinase. Curr. Biol. 2000, 10, 1527–1530. [Google Scholar] [CrossRef]
- Chipuk, J.E.; McStay, G.P.; Bharti, A.; Kuwana, T.; Clarke, C.J.; Siskind, L.J.; Obeid, L.M.; Green, D.R. Sphingolipid metabolism cooperates with BAK and BAX to promote the mitochondrial pathway of apoptosis. Cell 2012, 148, 988–1000. [Google Scholar] [CrossRef] [PubMed]
- Hofmann, L.P.; Ren, S.; Schwalm, S.; Pfeilschifter, J.; Huwiler, A. Sphingosine kinase 1 and 2 regulate the capacity of mesangial cells to resist apoptotic stimuli in an opposing manner. Biol. Chem. 2008, 389, 1399–1407. [Google Scholar] [CrossRef] [PubMed]
- Maceyka, M.; Sankala, H.; Hait, N.C.; Le Stunff, H.; Liu, H.; Toman, R.; Collier, C.; Zhang, M.; Satin, L.S.; Merrill, A.H., Jr.; et al. SphK1 and SphK2, sphingosine kinase isoenzymes with opposing functions in sphingolipid metabolism. J. Biol. Chem. 2005, 280, 37118–37129. [Google Scholar] [CrossRef]
- Schnitzer, S.E.; Weigert, A.; Zhou, J.; Brune, B. Hypoxia enhances sphingosine kinase 2 activity and provokes sphingosine-1-phosphate-mediated chemoresistance in A549 lung cancer cells. Mol. Cancer Res. 2009, 7, 393–401. [Google Scholar] [CrossRef]
- Nemoto, S.; Nakamura, M.; Osawa, Y.; Kono, S.; Itoh, Y.; Okano, Y.; Murate, T.; Hara, A.; Ueda, H.; Nozawa, Y.; et al. Sphingosine kinase isoforms regulate oxaliplatin sensitivity of human colon cancer cells through ceramide accumulation and Akt activation. J. Biol. Chem. 2009, 284, 10422–10432. [Google Scholar] [CrossRef]
- Gao, P.; Smith, C.D. Ablation of sphingosine kinase-2 inhibits tumor cell proliferation and migration. Mol. Cancer Res. 2011, 9, 1509–1519. [Google Scholar] [CrossRef]
- Weigert, A.; Schiffmann, S.; Sekar, D.; Ley, S.; Menrad, H.; Werno, C.; Grosch, S.; Geisslinger, G.; Brune, B. Sphingosine kinase 2 deficient tumor xenografts show impaired growth and fail to polarize macrophages towards an anti-inflammatory phenotype. Int. J. Cancer 2009, 125, 2114–2121. [Google Scholar] [CrossRef]
- Wallington-Beddoe, C.T.; Powell, J.A.; Tong, D.; Pitson, S.M.; Bradstock, K.F.; Bendall, L.J. Sphingosine kinase 2 promotes acute lymphoblastic leukemia by enhancing MYC expression. Cancer Res. 2014, 74, 2803–2815. [Google Scholar] [CrossRef]
- Neubauer, H.A.; Pham, D.H.; Zebol, J.R.; Moretti, P.A.; Peterson, A.L.; Leclercq, T.M.; Chan, H.; Powell, J.A.; Pitman, M.R.; Samuel, M.S.; et al. An oncogenic role for sphingosine kinase 2. Oncotarget 2016, 7, 64886–64899. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Li, J.; Li, G.; Li, Y.; Xu, C.; Li, M.; Xu, G.; Fu, S. Prognostic significance of sphingosine kinase 2 expression in non-small cell lung cancer. Tumour Biol. 2014, 35, 363–368. [Google Scholar] [CrossRef] [PubMed]
- Nagahashi, M.; Ramachandran, S.; Kim, E.Y.; Allegood, J.C.; Rashid, O.M.; Yamada, A.; Zhao, R.; Milstien, S.; Zhou, H.; Spiegel, S.; et al. Sphingosine-1-phosphate produced by sphingosine kinase 1 promotes breast cancer progression by stimulating angiogenesis and lymphangiogenesis. Cancer Res. 2012, 72, 726–735. [Google Scholar] [CrossRef] [PubMed]
- Nagahashi, M.; Tsuchida, J.; Moro, K.; Hasegawa, M.; Tatsuda, K.; Woelfel, I.A.; Takabe, K.; Wakai, T. High levels of sphingolipids in human breast cancer. J. Surg. Res. 2016, 204, 435–444. [Google Scholar] [CrossRef]
- Lee, H.; Deng, J.; Kujawski, M.; Yang, C.; Liu, Y.; Herrmann, A.; Kortylewski, M.; Horne, D.; Somlo, G.; Forman, S.; et al. STAT3-induced S1PR1 expression is crucial for persistent STAT3 activation in tumors. Nat. Med. 2010, 16, 1421–1428. [Google Scholar] [CrossRef]
- Chang, Q.; Bournazou, E.; Sansone, P.; Berishaj, M.; Gao, S.P.; Daly, L.; Wels, J.; Theilen, T.; Granitto, S.; Zhang, X.; et al. The IL-6/JAK/Stat3 feed-forward loop drives tumorigenesis and metastasis. Neoplasia 2013, 15, 848–862. [Google Scholar] [CrossRef]
- Liang, J.; Nagahashi, M.; Kim, E.Y.; Harikumar, K.B.; Yamada, A.; Huang, W.C.; Hait, N.C.; Allegood, J.C.; Price, M.M.; Avni, D.; et al. Sphingosine-1-phosphate links persistent STAT3 activation, chronic intestinal inflammation, and development of colitis-associated cancer. Cancer Cell 2013, 23, 107–120. [Google Scholar] [CrossRef]
- Deng, J.; Liu, Y.; Lee, H.; Herrmann, A.; Zhang, W.; Zhang, C.; Shen, S.; Priceman, S.J.; Kujawski, M.; Pal, S.K.; et al. S1PR1-STAT3 signaling is crucial for myeloid cell colonization at future metastatic sites. Cancer Cell 2012, 21, 642–654. [Google Scholar] [CrossRef]
- Theiss, A.L. Sphingosine-1-phosphate: Driver of NFkappaB and STAT3 persistent activation in chronic intestinal inflammation and colitis-associated cancer. JAKSTAT 2013, 2, e24150. [Google Scholar] [CrossRef]
- Chae, S.S.; Paik, J.H.; Furneaux, H.; Hla, T. Requirement for sphingosine 1-phosphate receptor-1 in tumor angiogenesis demonstrated by in vivo RNA interference. J. Clin. Investig. 2004, 114, 1082–1089. [Google Scholar] [CrossRef]
- Wang, S.; Liang, Y.; Chang, W.; Hu, B.; Zhang, Y. Triple Negative Breast Cancer Depends on Sphingosine Kinase 1 (SphK1)/Sphingosine-1-Phosphate (S1P)/Sphingosine 1-Phosphate Receptor 3 (S1PR3)/Notch Signaling for Metastasis. Med. Sci. Monit. 2018, 24, 1912–1923. [Google Scholar] [CrossRef]
- Watson, C.; Long, J.S.; Orange, C.; Tannahill, C.L.; Mallon, E.; McGlynn, L.M.; Pyne, S.; Pyne, N.J.; Edwards, J. High expression of sphingosine 1-phosphate receptors, S1P1 and S1P3, sphingosine kinase 1, and extracellular signal-regulated kinase-1/2 is associated with development of tamoxifen resistance in estrogen receptor-positive breast cancer patients. Am. J. Pathol. 2010, 177, 2205–2215. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.J.; Thangada, S.; Claffey, K.P.; Ancellin, N.; Liu, C.H.; Kluk, M.; Volpi, M.; Sha’afi, R.I.; Hla, T. Vascular endothelial cell adherens junction assembly and morphogenesis induced by sphingosine-1-phosphate. Cell 1999, 99, 301–312. [Google Scholar] [CrossRef]
- Zhao, J.; Liu, J.; Lee, J.F.; Zhang, W.; Kandouz, M.; VanHecke, G.C.; Chen, S.; Ahn, Y.H.; Lonardo, F.; Lee, M.J. TGF-beta/SMAD3 Pathway Stimulates Sphingosine-1 Phosphate Receptor 3 Expression: IMPLICATION OF SPHINGOSINE-1 PHOSPHATE RECEPTOR 3 IN LUNG ADENOCARCINOMA PROGRESSION. J. Biol. Chem. 2016, 291, 27343–27353. [Google Scholar] [CrossRef] [PubMed]
- Goetzl, E.J.; Dolezalova, H.; Kong, Y.; Zeng, L. Dual mechanisms for lysophospholipid induction of proliferation of human breast carcinoma cells. Cancer Res. 1999, 59, 4732–4737. [Google Scholar]
- Asghar, M.Y.; Kemppainen, K.; Lassila, T.; Tornquist, K. Sphingosine 1-phosphate attenuates MMP2 and MMP9 in human anaplastic thyroid cancer C643 cells: Importance of S1P2. PLoS ONE 2018, 13, e0196992. [Google Scholar] [CrossRef]
- Liu, R.; Zhao, R.; Zhou, X.; Liang, X.; Campbell, D.J.; Zhang, X.; Zhang, L.; Shi, R.; Wang, G.; Pandak, W.M.; et al. Conjugated bile acids promote cholangiocarcinoma cell invasive growth through activation of sphingosine 1-phosphate receptor 2. Hepatology 2014, 60, 908–918. [Google Scholar] [CrossRef]
- Tang, X.; McMullen, T.P.W.; Brindley, D.N. Increasing the low lipid phosphate phosphatase 1 activity in breast cancer cells decreases transcription by AP-1 and expressions of matrix metalloproteinases and cyclin D1/D3. Theranostics 2019, 9, 6129–6142. [Google Scholar] [CrossRef]
- Leung, D.W.; Tompkins, C.K.; White, T. Molecular cloning of two alternatively spliced forms of human phosphatidic acid phosphatase cDNAs that are differentially expressed in normal and tumor cells. DNA Cell Biol. 1998, 17, 377–385. [Google Scholar] [CrossRef]
- Bhattacharjee, A.; Richards, W.G.; Staunton, J.; Li, C.; Monti, S.; Vasa, P.; Ladd, C.; Beheshti, J.; Bueno, R.; Gillette, M.; et al. Classification of human lung carcinomas by mRNA expression profiling reveals distinct adenocarcinoma subclasses. Proc. Natl. Acad. Sci. USA 2001, 98, 13790–13795. [Google Scholar] [CrossRef]
- Curtis, C.; Shah, S.P.; Chin, S.-F.; Turashvili, G.; Rueda, O.M.; Dunning, M.J.; Speed, D.; Lynch, A.G.; Samarajiwa, S.; Yuan, Y.; et al. The genomic and transcriptomic architecture of 2000 breast tumours reveals novel subgroups. Nature 2012, 486, 346–352. [Google Scholar] [CrossRef]
- Yoshihara, K.; Tajima, A.; Komata, D.; Yamamoto, T.; Kodama, S.; Fujiwara, H.; Suzuki, M.; Onishi, Y.; Hatae, M.; Sueyoshi, K.; et al. Gene expression profiling of advanced-stage serous ovarian cancers distinguishes novel subclasses and implicates ZEB2 in tumor progression and prognosis. Cancer Sci. 2009, 100, 1421–1428. [Google Scholar] [CrossRef]
- Gao, X.; Goggin, K.; Dowling, C.; Qian, J.; Hawdon, J.M. Two potential hookworm DAF-16 target genes, SNR-3 and LPP-1: Gene structure, expression profile, and implications of a cis-regulatory element in the regulation of gene expression. Parasit. Vectors 2015, 8, 14. [Google Scholar] [CrossRef][Green Version]
- Loziuk, P.; Meier, F.; Johnson, C.; Ghashghaei, H.T.; Muddiman, D.C. TransOmic analysis of forebrain sections in Sp2 conditional knockout embryonic mice using IR-MALDESI imaging of lipids and LC-MS/MS label-free proteomics. Anal. Bioanal. Chem. 2016, 408, 3453–3474. [Google Scholar] [CrossRef]
- Reschen, M.E.; Gaulton, K.J.; Lin, D.; Soilleux, E.J.; Morris, A.J.; Smyth, S.S.; O’Callaghan, C.A. Lipid-induced epigenomic changes in human macrophages identify a coronary artery disease-associated variant that regulates PPAP2B Expression through Altered C/EBP-beta binding. PLoS Genet. 2015, 11, e1005061. [Google Scholar] [CrossRef]
- Mao, G.; Smyth, S.S.; Morris, A.J. Regulation of PLPP3 gene expression by NF-kappaB family transcription factors. J. Biol. Chem. 2019, 294, 14009–14019. [Google Scholar] [CrossRef]
- Kotini, A.G.; Mpakali, A.; Agalioti, T. Dnmt3a1 upregulates transcription of distinct genes and targets chromosomal gene clusters for epigenetic silencing in mouse embryonic stem cells. Mol. Cell Biol. 2011, 31, 1577–1592. [Google Scholar] [CrossRef]
- Wary, K.K.; Thakker, G.D.; Humtsoe, J.O.; Yang, J. Analysis of VEGF-responsive genes involved in the activation of endothelial cells. Mol. Cancer 2003, 2, 25. [Google Scholar] [CrossRef][Green Version]
- Asirvatham, A.J.; Schmidt, M.; Gao, B.; Chaudhary, J. Androgens regulate the immune/inflammatory response and cell survival pathways in rat ventral prostate epithelial cells. Endocrinology 2006, 147, 257–271. [Google Scholar] [CrossRef]
- Imai, A.; Furui, T.; Tamaya, T.; Mills, G.B. A gonadotropin-releasing hormone-responsive phosphatase hydrolyses lysophosphatidic acid within the plasma membrane of ovarian cancer cells. J. Clin. Endocrinol. Metab. 2000, 85, 3370–3375. [Google Scholar] [CrossRef][Green Version]
- Harper, K.; Brochu-Gaudreau, K.; Saucier, C.; Dubois, C.M. Hypoxia Downregulates LPP3 and Promotes the Spatial Segregation of ATX and LPP1 During Cancer Cell Invasion. Cancers 2019, 11, 1403. [Google Scholar] [CrossRef]
- Tanyi, J.L.; Hasegawa, Y.; Lapushin, R.; Morris, A.J.; Wolf, J.K.; Berchuck, A.; Lu, K.; Smith, D.I.; Kalli, K.; Hartmann, L.C.; et al. Role of decreased levels of lipid phosphate phosphatase-1 in accumulation of lysophosphatidic acid in ovarian cancer. Clin. Cancer Res. 2003, 9, 3534–3545. [Google Scholar]
- Tanyi, J.L.; Morris, A.J.; Wolf, J.K.; Fang, X.; Hasegawa, Y.; Lapushin, R.; Auersperg, N.; Sigal, Y.J.; Newman, R.A.; Felix, E.A.; et al. The human lipid phosphate phosphatase-3 decreases the growth, survival, and tumorigenesis of ovarian cancer cells: Validation of the lysophosphatidic acid signaling cascade as a target for therapy in ovarian cancer. Cancer Res. 2003, 63, 1073–1082. [Google Scholar]
- Tang, X.; Wang, X.; Zhao, Y.Y.; Curtis, J.M.; Brindley, D.N. Doxycycline attenuates breast cancer related inflammation by decreasing plasma lysophosphatidate concentrations and inhibiting NF-kappaB activation. Mol. Cancer 2017, 16, 36. [Google Scholar] [CrossRef]
- Mao, X.Y.; Lee, M.J.; Zhu, J.; Zhu, C.; Law, S.M.; Snijders, A.M. Genome-wide screen identifies a novel prognostic signature for breast cancer survival. Oncotarget 2017, 8, 14003–14016. [Google Scholar] [CrossRef]
- Chatterjee, I.; Humtsoe, J.O.; Kohler, E.E.; Sorio, C.; Wary, K.K. Lipid phosphate phosphatase-3 regulates tumor growth via beta-catenin and Cyclin-D1 signaling. Mol. Cancer 2011, 10, 51. [Google Scholar] [CrossRef]
- Susanto, O.; Koh, Y.W.H.; Morrice, N.; Tumanov, S.; Thomason, P.A.; Nielson, M.; Tweedy, L.; Muinonen-Martin, A.J.; Kamphorst, J.J.; Mackay, G.M.; et al. LPP3 mediates self-generation of chemotactic LPA gradients by melanoma cells. J. Cell Sci. 2017, 130, 3455–3466. [Google Scholar] [CrossRef]
- Flanagan, J.M.; Funes, J.M.; Henderson, S.; Wild, L.; Carey, N.; Boshoff, C. Genomics screen in transformed stem cells reveals RNASEH2A, PPAP2C, and ADARB1 as putative anticancer drug targets. Mol. Cancer Ther. 2009, 8, 249–260. [Google Scholar] [CrossRef]
- Zhang, N.; Sundberg, J.P.; Gridley, T. Mice mutant for Ppap2c, a homolog of the germ cell migration regulator wunen, are viable and fertile. Genesis 2000, 27, 137–140. [Google Scholar] [CrossRef]
- Escalante-Alcalde, D.; Hernandez, L.; Le Stunff, H.; Maeda, R.; Lee, H.S.; Gang, C., Jr.; Sciorra, V.A.; Daar, I.; Spiegel, S.; Morris, A.J.; et al. The lipid phosphatase LPP3 regulates extra-embryonic vasculogenesis and axis patterning. Development 2003, 130, 4623–4637. [Google Scholar] [CrossRef]
- Ile, K.E.; Tripathy, R.; Goldfinger, V.; Renault, A.D. Wunen, a Drosophila lipid phosphate phosphatase, is required for septate junction-mediated barrier function. Development 2012, 139, 2535–2546. [Google Scholar] [CrossRef] [PubMed]
- Kai, M.; Sakane, F.; Jia, Y.J.; Imai, S.; Yasuda, S.; Kanoh, H. Lipid phosphate phosphatases 1 and 3 are localized in distinct lipid rafts. J. Biochem. 2006, 140, 677–686. [Google Scholar] [CrossRef] [PubMed]

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tang, X.; Brindley, D.N. Lipid Phosphate Phosphatases and Cancer. Biomolecules 2020, 10, 1263. https://doi.org/10.3390/biom10091263
Tang X, Brindley DN. Lipid Phosphate Phosphatases and Cancer. Biomolecules. 2020; 10(9):1263. https://doi.org/10.3390/biom10091263
Chicago/Turabian StyleTang, Xiaoyun, and David N. Brindley. 2020. "Lipid Phosphate Phosphatases and Cancer" Biomolecules 10, no. 9: 1263. https://doi.org/10.3390/biom10091263
APA StyleTang, X., & Brindley, D. N. (2020). Lipid Phosphate Phosphatases and Cancer. Biomolecules, 10(9), 1263. https://doi.org/10.3390/biom10091263





