EGFP Reporters for Direct and Sensitive Detection of Mutagenic Bypass of DNA Lesions
Abstract
1. Introduction
2. Materials and Methods
2.1. Reporter Constructs for Detection of TM
2.2. Reporter Constructs for Detection of Mutagenic TLS Templated by the AP Lesion
2.3. Phenotypic Screening of the EGFP Mutants
2.4. Phenotypic Validation of Newly Identified EGFP Mutants
2.5. Cell Culture, Transfection, and EGFP Expression Analysis
3. Results
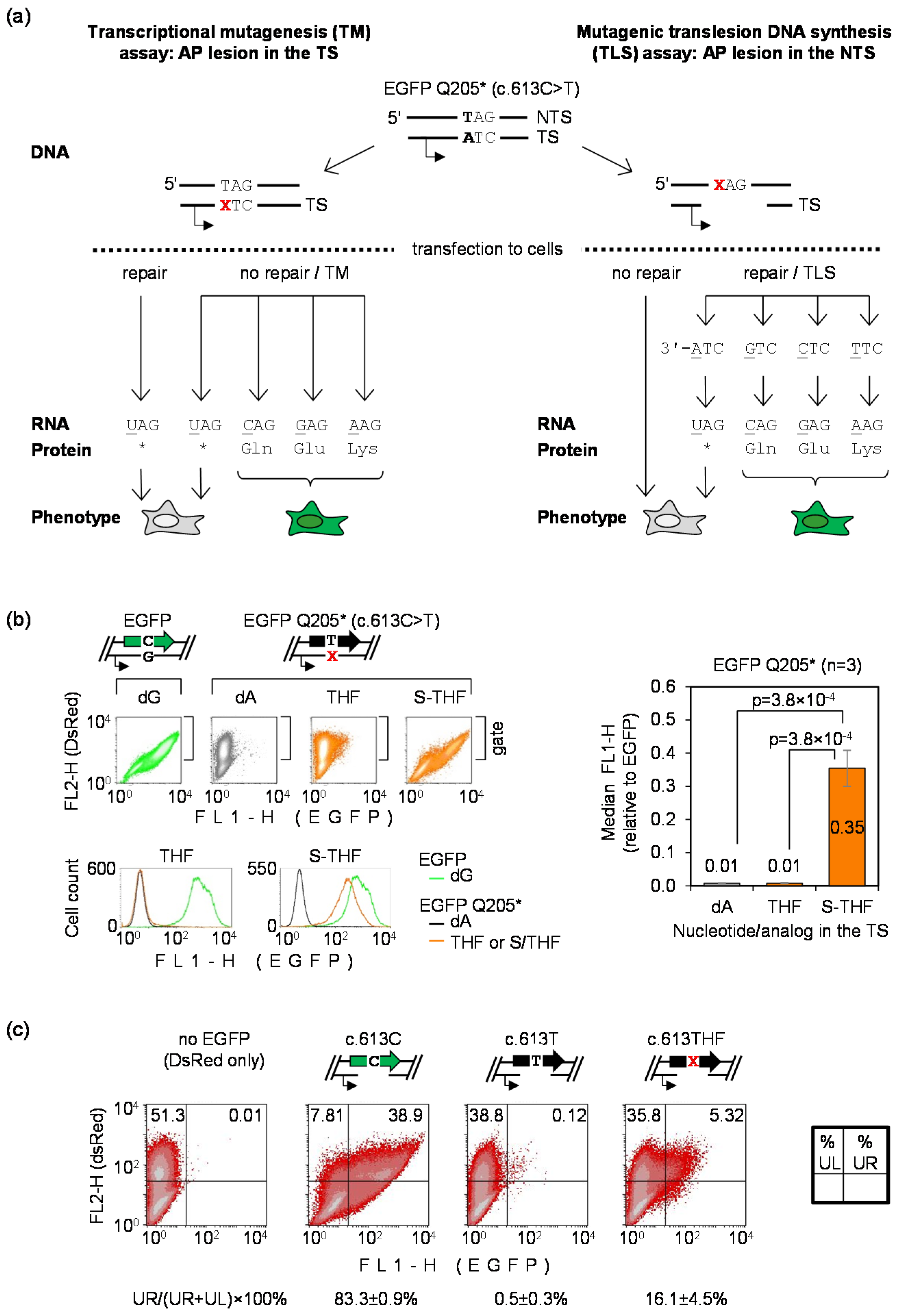
3.1. Detection of Mutagenic Bypass of Abasic Site During RNA and DNA Synthesis Using EGFP Q205* Reporter
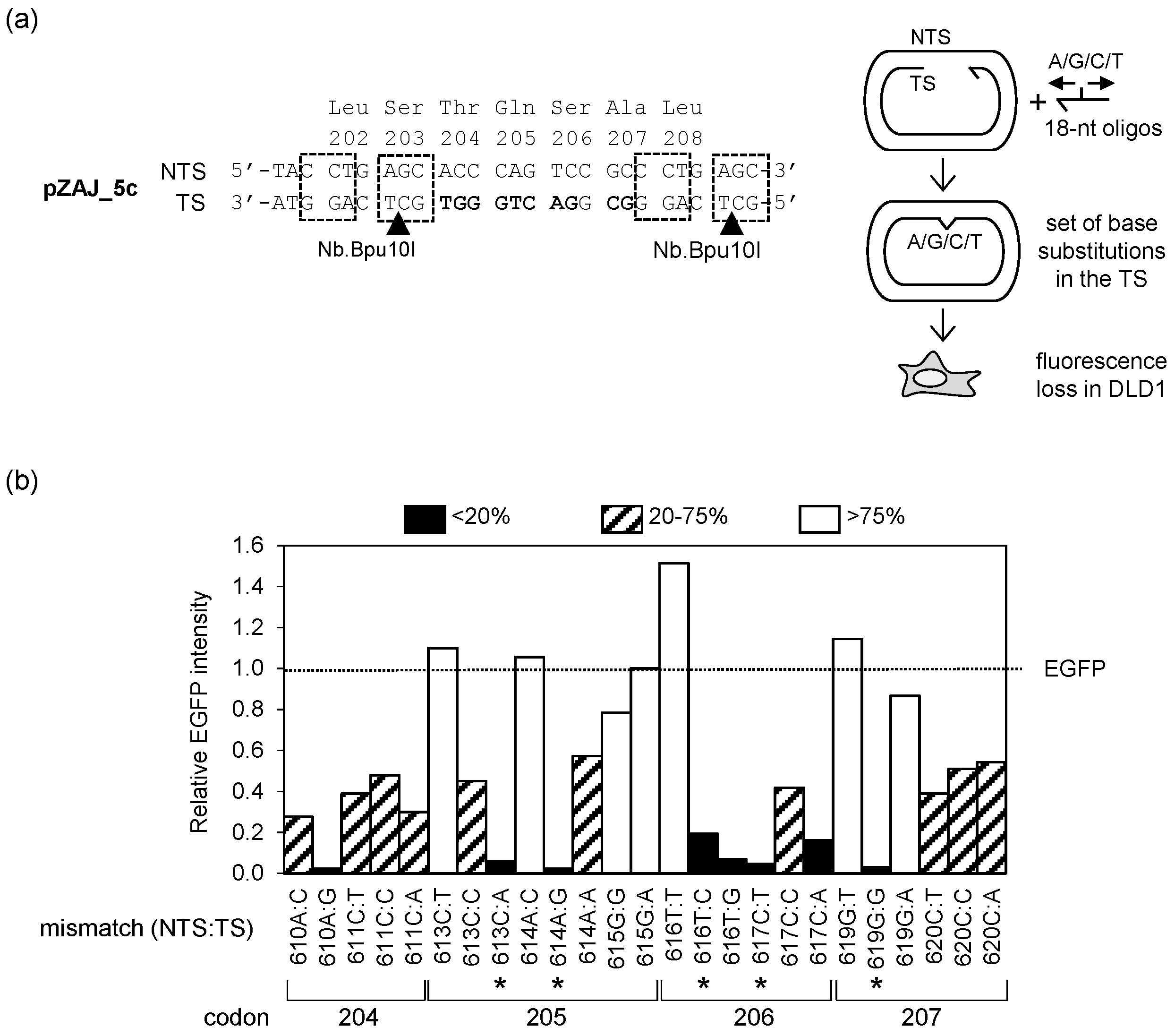
3.2. Screening the EGFP Sequence for Candidate Single Nucleotide Substitutions Leading to the Loss of Fluorescence
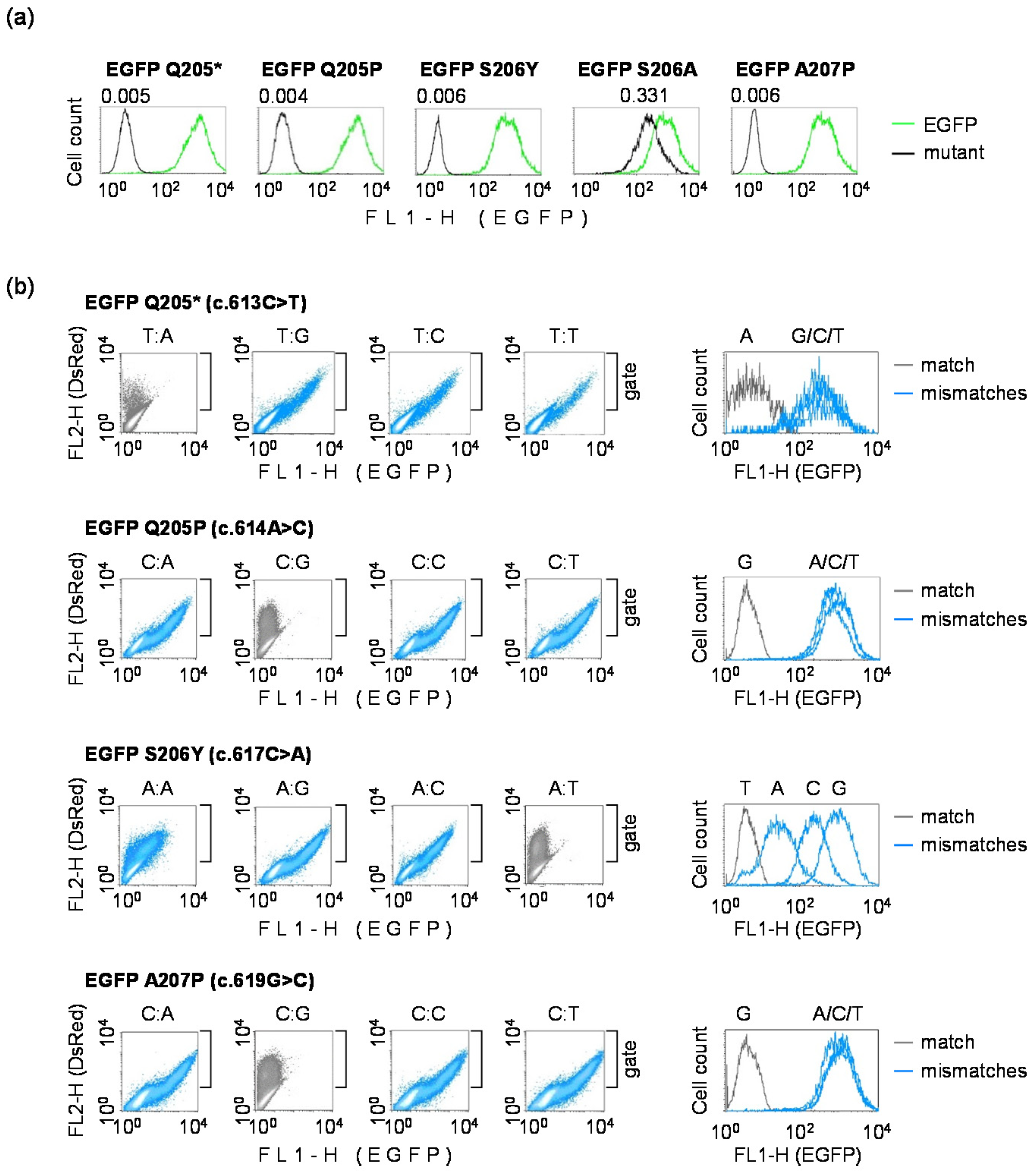
3.3. Validation of EGFP Mutants Carrying Inactivating Single Nucleotide Substitutions
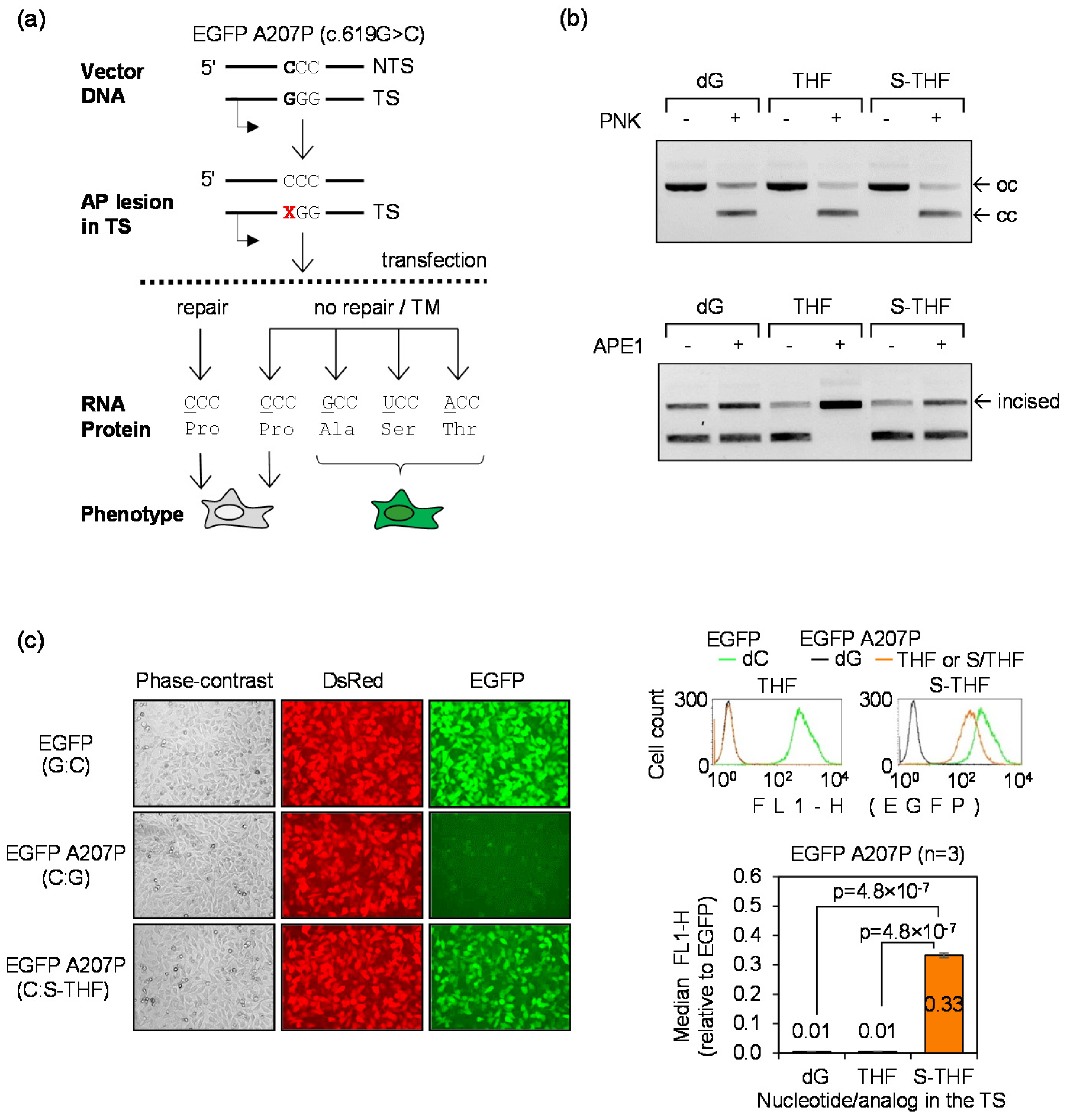
3.4. Detection of TM Using the A207P (c.619G>C) EGFP Mutant
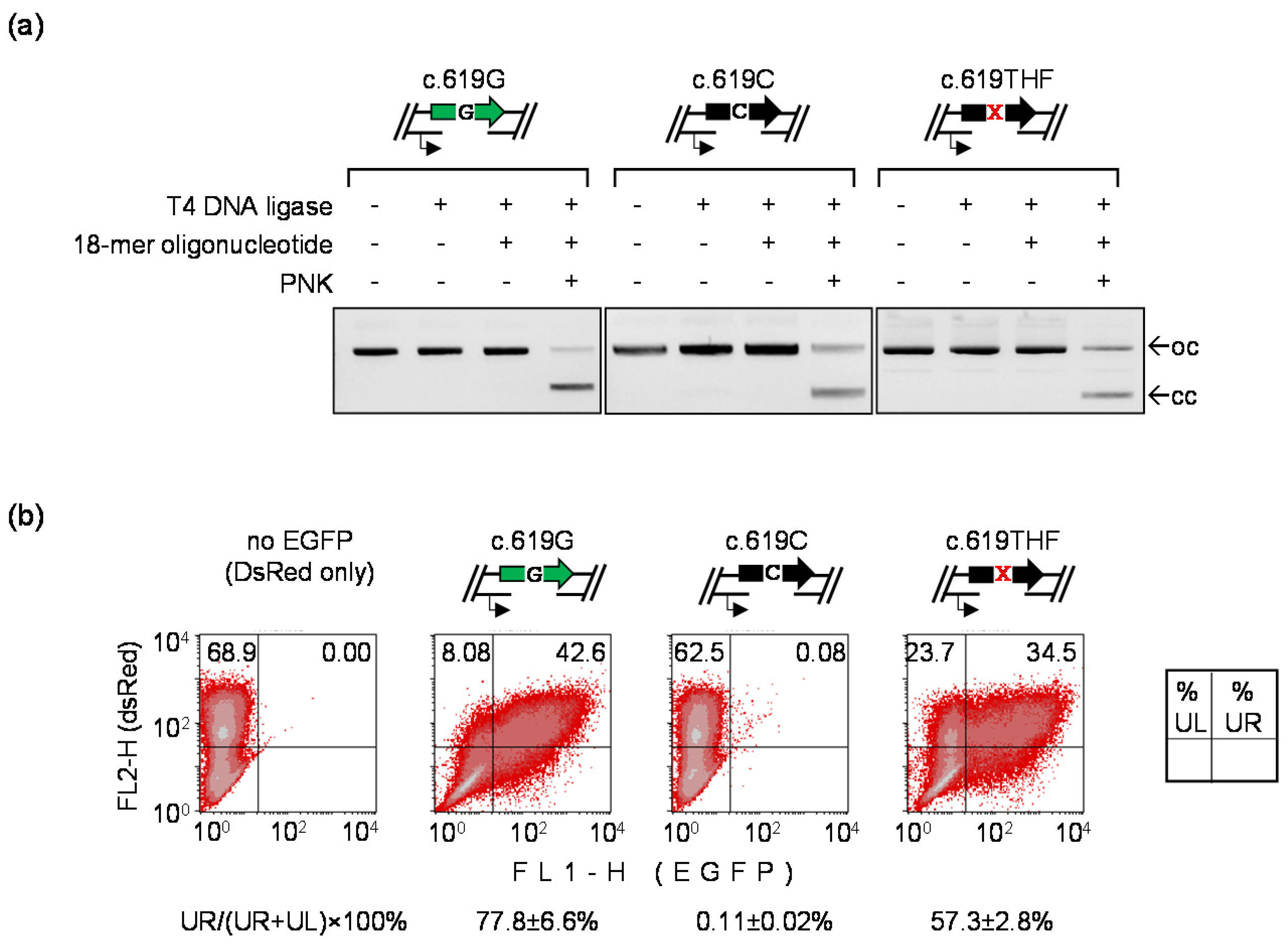
3.5. Detection of Mutagenic TLS Using the A207P (c.619G>C) EGFP Mutant
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Friedberg, E.C. Suffering in silence: The tolerance of DNA damage. Nat. Rev. Mol. Cell Biol. 2005, 6, 943–953. [Google Scholar] [CrossRef] [PubMed]
- Chatterjee, N.; Walker, G.C. Mechanisms of DNA damage, repair, and mutagenesis. Environ. Mol. Mutagen. 2017, 58, 235–263. [Google Scholar] [CrossRef] [PubMed]
- Izhar, L.; Ziv, O.; Cohen, I.S.; Geacintov, N.E.; Livneh, Z. Genomic assay reveals tolerance of DNA damage by both translesion DNA synthesis and homology-dependent repair in mammalian cells. Proc. Natl. Acad. Sci. USA 2013, 110, 1462–1469. [Google Scholar] [CrossRef] [PubMed]
- Lange, S.S.; Takata, K.I.; Wood, R.D. DNA polymerases and cancer. Nat. Rev. Cancer 2011, 11, 96–110. [Google Scholar] [CrossRef]
- Sale, J.E.; Lehmann, A.R.; Woodgate, R. Y-family DNA polymerases and their role in tolerance of cellular DNA damage. Nat. Rev. Mol. Cell Biol. 2012, 13, 141–152. [Google Scholar] [CrossRef]
- Bielas, J.H.; Loeb, K.R.; Rubin, B.P.; True, L.D.; Loeb, L.A. Human cancers express a mutator phenotype. Proc. Natl. Acad. Sci. USA 2006, 103, 18238–18242. [Google Scholar] [CrossRef]
- Hoeijmakers, J.H.J. Genome maintenance mechanisms for preventing cancer. Nature 2001, 411, 366–374. [Google Scholar] [CrossRef]
- Maga, G.; Villani, G.; Crespan, E.; Wimmer, U.; Ferrari, E.; Bertocci, B.; Hübscher, U. 8-oxo-guanine bypass by human DNA polymerases in the presence of auxiliary proteins. Nature 2007, 447, 606–608. [Google Scholar] [CrossRef]
- Choi, J.Y.; Lim, S.; Kim, E.J.; Jo, A.; Guengerich, F.P. Translesion Synthesis across Abasic Lesions by Human B-Family and Y-Family DNA Polymerases α, δ, η, ι, κ, and REV1. J. Mol. Biol. 2010, 404, 34–44. [Google Scholar] [CrossRef]
- Shibutani, S.; Suzuki, N.; Grollman, A.P. Mechanism of frameshift (deletion) generated by acetylaminofluorene-derived DNA adducts in vitro. Biochemistry 2004, 43, 15929–15935. [Google Scholar] [CrossRef]
- Schorr, S.; Schneider, S.; Lammens, K.; Hopfner, K.P.; Carell, T. Mechanism of replication blocking and bypass of Y-family polymerase η by bulky acetylaminofluorene DNA adducts. Proc. Natl. Acad. Sci. USA 2010, 107, 20720–20725. [Google Scholar] [CrossRef] [PubMed]
- Sherrer, S.M.; Taggart, D.J.; Pack, L.R.; Malik, C.K.; Basu, A.K.; Suo, Z. Quantitative analysis of the mutagenic potential of 1-aminopyrene-DNA adduct bypass catalyzed by Y-family DNA polymerases. Mutat. Res.-Fundam. Mol. Mech. Mutagen. 2012, 737, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Yasui, M.; Dong, H.; Bonala, R.R.; Suzuki, N.; Ohmori, H.; Hanaoka, F.; Johnson, F.; Grollman, A.P.; Shibutani, S. Mutagenic properties of 3-(deoxyguanosin-N2-yl)-2-acetylaminofluorene, a persistent acetylaminofluorene-derived DNA adduct in mammalian cells. Biochemistry 2004, 43, 15005–15013. [Google Scholar] [CrossRef]
- Avkin, S.; Adar, S.; Blander, G.; Livneh, Z. Quantitative measurement of translesion replication in human cells: Evidence for bypass of abasic sites by a replicative DNA polymerase. Proc. Natl. Acad. Sci. USA 2002, 99, 3764–3769. [Google Scholar] [CrossRef] [PubMed]
- Hashimoto, K.; Bonala, R.; Johnson, F.; Grollman, A.P.; Moriya, M. Y-family DNA polymerase-independent gap-filling translesion synthesis across aristolochic acid-derived adenine adducts in mouse cells. DNA Rep. 2016, 46, 55–60. [Google Scholar] [CrossRef]
- Basu, A.K.; Pande, P.; Bose, A. Translesion synthesis of 2-deoxyguanosine lesions by eukaryotic DNA polymerases. Chem. Res. Toxicol. 2017, 30, 61–72. [Google Scholar] [CrossRef]
- Ziv, O.; Zeisel, A.; Mirlas-Neisberg, N.; Swain, U.; Nevo, R.; Ben-Chetrit, N.; Martelli, M.P.; Rossi, R.; Schiesser, S.; Canman, C.E.; et al. Identification of novel DNA-damage tolerance genes reveals regulation of translesion DNA synthesis by nucleophosmin. Nat. Commun. 2014, 5, 5437. [Google Scholar] [CrossRef]
- Veaute, X.; Fuchs, R.P.P. Greater susceptibility to mutations in lagging strand of DNA replication in Escherichia coli than in leading strand. Science 1993, 261, 598–600. [Google Scholar] [CrossRef]
- Yagi, T.; Fujikawa, Y.; Sawai, T.; Takamura-Enya, T.; Ito-Harashima, S.; Kawanishi, M. Error-prone and error-free translesion DNA synthesis over site-specifically created DNA adducts of aryl hydrocarbons (3-nitrobenzanthrone and 4-aminobiphenyl). Toxicol. Res. 2017, 33, 265–272. [Google Scholar] [CrossRef][Green Version]
- Lans, H.; Hoeijmakers, J.H.J.; Vermeulen, W.; Marteijn, J.A. The DNA damage response to transcription stress. Nat. Rev. Mol. Cell Biol. 2019, 20, 766–784. [Google Scholar] [CrossRef]
- Brégeon, D.; Doetsch, P.W. Transcriptional mutagenesis: Causes and involvement in tumour development. Nat. Rev. Cancer 2011, 11, 218–227. [Google Scholar] [CrossRef] [PubMed]
- Saxowsky, T.T.; Meadows, K.L.; Klungland, A.; Doetsch, P.W. 8-Oxoguanine-mediated transcriptional mutagenesis causes Ras activation in mammalian cells. Proc. Natl. Acad. Sci. USA 2008, 105, 18877–18882. [Google Scholar] [CrossRef] [PubMed]
- Brégeon, D.; Peignon, P.A.; Sarasin, A. Transcriptional mutagenesis induced by 8-Oxoguanine in mammalian cells. PLoS Genet. 2009, 5, e1000577. [Google Scholar] [CrossRef] [PubMed]
- Ezerskyte, M.; Paredes, J.A.; Malvezzi, S.; Burns, J.A.; Margison, G.P.; Olsson, M.; Scicchitano, D.A.; Dreij, K. O6-methylguanine–induced transcriptional mutagenesis reduces p53 tumor-suppressor function. Proc. Natl. Acad. Sci. USA 2018, 115, 4731–4736. [Google Scholar] [CrossRef] [PubMed]
- Doetsch, P.W. Translesion synthesis by RNA polymerases: Occurrence and biological implications for transcriptional mutagenesis. Mutat. Res. 2002, 510, 131–140. [Google Scholar] [CrossRef]
- Holmquist, G.P. Cell-selfish modes of evolution and mutations directed after transcriptional bypass. Mutat. Res. Fundam. Mol. Mech. Mutagen. 2002, 510, 141–152. [Google Scholar] [CrossRef]
- Shaw, R.J.; Bonawitz, N.D.; Reines, D. Use of an in vivo reporter assay to test for transcriptional and translational fidelity in yeast. J. Biol. Chem. 2002, 277, 24420–24426. [Google Scholar] [CrossRef]
- Burns, J.A.; Dreij, K.; Cartularo, L.; Scicchitano, D.A. O6-Methylguanine induces altered proteins at the level of transcription in human cells. Nucleic Acids Res. 2010, 38, 8178–8187. [Google Scholar] [CrossRef]
- Nagel, Z.D.; Margulies, C.M.; Chaim, I.A.; McRee, S.K.; Mazzucato, P.; Ahmad, A.; Abo, R.P.; Butty, V.L.; Forget, A.L.; Samson, L.D. Multiplexed DNA repair assays for multiple lesions and multiple doses via transcription inhibition and transcriptional mutagenesis. Proc. Natl. Acad. Sci. USA 2014, 111, E1823–E1832. [Google Scholar] [CrossRef]
- Kitsera, N.; Rodriguez-Alvarez, M.; Emmert, S.; Carell, T.; Khobta, A. Nucleotide excision repair of abasic DNA lesions. Nucleic Acids Res. 2019, 47, 8537–8547. [Google Scholar] [CrossRef]
- Kitsera, N.; Stathis, D.; Lühnsdorf, B.; Müller, H.; Carell, T.; Epe, B.; Khobta, A. 8-Oxo-7,8-dihydroguanine in DNA does not constitute a barrier to transcription, but is converted into transcription-blocking damage by OGG1. Nucleic Acids Res. 2011, 39, 5926–5934. [Google Scholar] [CrossRef] [PubMed]
- Lühnsdorf, B.; Kitsera, N.; Warken, D.; Lingg, T.; Epe, B.; Khobta, A. Generation of reporter plasmids containing defined base modifications in the DNA strand of choice. Anal. Biochem. 2012, 425, 47–53. [Google Scholar] [CrossRef] [PubMed]
- Allgayer, J.; Kitsera, N.; Von Der Lippen, C.; Epe, B.; Khobta, A. Modulation of base excision repair of 8-oxoguanine by the nucleotide sequence. Nucleic Acids Res. 2013, 41, 8559–8571. [Google Scholar] [CrossRef] [PubMed]
- Lühnsdorf, B.; Epe, B.; Khobta, A. Excision of uracil from transcribed DNA negatively affects gene expression. J. Biol. Chem. 2014, 289, 22008–22018. [Google Scholar] [CrossRef] [PubMed]
- Khobta, A.; Anderhub, S.; Kitsera, N.; Epe, B. Gene silencing induced by oxidative DNA base damage: Association with local decrease of histone H4 acetylation in the promoter region. Nucleic Acids Res. 2010, 38, 4285–4295. [Google Scholar] [CrossRef]
- Allgayer, J.; Kitsera, N.; Bartelt, S.; Epe, B.; Khobta, A. Widespread transcriptional gene inactivation initiated by a repair intermediate of 8-oxoguanine. Nucleic Acids Res. 2016, 44, 7267–7280. [Google Scholar] [CrossRef]
- Brejc, K.; Sixma, T.K.; Kitts, P.A.; Kain, S.R.; Tsien, R.Y.; Ormö, M.; Remington, S.J. Structural basis for dual excitation and photoisomerization of the Aequorea victoria green fluorescent protein. Proc. Natl. Acad. Sci. USA 1997, 94, 2306–2311. [Google Scholar] [CrossRef]
- Gentil, A.; Cabral-Neto, J.B.; Mariage-Samson, R.; Margot, A.; Imbach, J.L.; Rayner, B.; Sarasin, A. Mutagenicity of a unique apurinic/apyrimidinic site in mammalian cells. J. Mol. Biol. 1992, 227, 981–984. [Google Scholar] [CrossRef]
- Takeshita, M.; Eisenberg, W. Mechanism of mutation on DNA templates containing synthetic abasic sites: Study with a double strand vector. Nucleic Acids Res. 1994, 22, 1897–1902. [Google Scholar] [CrossRef][Green Version]
- Lawrence, C.W.; Borden, A.; Banerjee, S.K.; Leclerc, J.E. Mutation frequency and spectrum resulting from a single abasic site in a single-stranded vector. Nucleic Acids Res. 1990, 18, 2153–2157. [Google Scholar] [CrossRef]
- Kucab, J.E.; Zou, X.; Morganella, S.; Joel, M.; Nanda, A.S.; Nagy, E.; Gomez, C.; Degasperi, A.; Harris, R.; Jackson, S.P.; et al. A Compendium of Mutational Signatures of Environmental Agents. Cell 2019, 177, 821–836. [Google Scholar] [CrossRef] [PubMed]
- Alexandrov, L.B.; Kim, J.; Haradhvala, N.J.; Huang, M.N.; Tian Ng, A.W.; Wu, Y.; Boot, A.; Covington, K.R.; Gordenin, D.A.; Bergstrom, E.N.; et al. The repertoire of mutational signatures in human cancer. Nature 2020, 578, 94–101. [Google Scholar] [CrossRef] [PubMed]
- Hollstein, M.; Alexandrov, L.B.; Wild, C.P.; Ardin, M.; Zavadil, J. Base changes in tumour DNA have the power to reveal the causes and evolution of cancer. Oncogene 2017, 36, 158–167. [Google Scholar] [CrossRef] [PubMed]
- Birnbaum, M.D.; Nemzow, L.; Kumar, A.; Gong, F.; Zhang, F. A Rapid and Precise Mutation-Activated Fluorescence Reporter for Analyzing Acute Mutagenesis Frequency. Cell Chem. Biol. 2018, 25, 1038–1049. [Google Scholar] [CrossRef]
- Fedeles, B.I.; Essigmann, J.M. Impact of DNA lesion repair, replication and formation on the mutational spectra of environmental carcinogens: Aflatoxin B 1 as a case study. DNA Repair 2018, 71, 12–22. [Google Scholar] [CrossRef]
- Koehl, P.; Burnouf, D.; Fuchs, R.P.P. Construction of plasmids containing a unique acetylaminofluorene adduct located within a mutation hot spot. A new probe for frameshift mutagenesis. J. Mol. Biol. 1989, 207, 355–364. [Google Scholar] [CrossRef]
- Levine, R.L.; Yang, I.Y.; Hossain, M.; Pandya, G.A.; Grollman, A.P.; Moriya, M. Mutagenesis induced by a single 1, N6-ethenodeoxyadenosine adduct in human cells. Cancer Res. 2000, 60, 4098–4104. [Google Scholar]
- Wee, H.A.; Brown, W.W.; Lippard, S.J. Preparation of mammalian expression vectors incorporating site-specifically platinated-DNA lesions. Bioconjug. Chem. 2009, 20, 1058–1063. [Google Scholar]
- Brégeon, D.; Doetsch, P.W. Reliable method for generating double-stranded DNA vectors containing site-specific base modifications. Biotechniques 2004, 37, 760–766. [Google Scholar] [CrossRef]
- Pagès, V.; Mazón, G.; Naiman, K.; Philippin, G.; Fuchs, R.P. Monitoring bypass of single replication-blocking lesions by damage avoidance in the Escherichia coli chromosome. Nucleic Acids Res. 2012, 40, 9036–9043. [Google Scholar] [CrossRef]
- Masłowska, K.H.; Laureti, L.; Pagès, V. iDamage: A method to integrate modified DNA into the yeast genome. Nucleic Acids Res. 2019, 47, e124. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Hays, J.B. Simple and rapid preparation of gapped plasmid DNA for incorporation of oligomers containing specific DNA lesions. Mol. Biotechnol. 2001, 19, 133–140. [Google Scholar] [CrossRef]
- Baker, D.J.; Wuenschell, G.; Xia, L.; Termini, J.; Bates, S.E.; Riggs, A.D.; O’Connor, T.R. Nucleotide excision repair eliminates unique DNA-protein cross-links from mammalian cells. J. Biol. Chem. 2007, 282, 22592–22604. [Google Scholar] [CrossRef] [PubMed]
- Kitsera, N.; Gasteiger, K.; Lühnsdorf, B.; Allgayer, J.; Epe, B.; Carell, T.; Khobta, A. Cockayne syndrome: Varied requirement of transcription-coupled nucleotide excision repair for the removal of three structurally different adducts from transcribed DNA. PLoS ONE 2014, 9, e94405. [Google Scholar] [CrossRef]
- Ji, S.; Park, D.; Kropachev, K.; Kolbanovskiy, M.; Fu, I.; Broyde, S.; Essawy, M.; Geacintov, N.E.; Tretyakova, N.Y. 5-Formylcytosine-induced DNA–peptide cross-links reduce transcription efficiency, but do not cause transcription errors in human cells. J. Biol. Chem. 2019, 294, 18387–18397. [Google Scholar] [CrossRef]
- Piekna-Przybylska, D. Reporter assays for BER pathway. In Methods in Molecular Biology; Balakrishnan, L., Stewart, J.A., Eds.; Springer: New York, NY, USA, 2019; pp. 145–160. ISBN 978-1-4939-9500-4. [Google Scholar]
- Jung, G.; Wiehler, J.; Zumbusch, A. The photophysics of green fluorescent protein: Influence of the key amino acids at positions 65, 203, and 222. Biophys. J. 2005, 88, 1932–1947. [Google Scholar] [CrossRef]
- Ehrig, T.; O’Kane, D.J.; Prendergast, F.G. Green-fluorescent protein mutants with altered fluorescence excitation spectra. FEBS Lett. 1995, 367, 163–166. [Google Scholar] [CrossRef]
- Stepanenko, O.V.; Kuznetsova, I.M.; Verkhusha, V.V.; Turoverov, K.K. Beta-Barrel Scaffold of Fluorescent Proteins. Folding, Stability and Role in Chromophore Formation. In International Review of Cell and Molecular Biology; Elsevier: Amsterdam, The Netherlands, 2013; Volume 302, pp. 221–278. [Google Scholar]
- Loeb, L.A.; Preston, B.D. Mutagenesis by Apurinic/Apyrimidinic Sites. Annu. Rev. Genet. 1986, 20, 201–230. [Google Scholar] [CrossRef]
- Weerasooriya, S.; Jasti, V.P.; Basu, A.K. Replicative bypass of abasic site in Escherichia coli and human cells: Similarities and differences. PLoS ONE 2014, 9, e107915. [Google Scholar] [CrossRef]
- Villani, G.; Shevelev, I.; Orlando, E.; Pospiech, H.; Syvaoja, J.E.; Markkanen, E.; Hubscher, U.; Le Gac, N.T. Gap-directed translesion DNA Synthesis of an abasic site on circular DNA templates by a human replication complex. PLoS ONE 2014, 9, e93908. [Google Scholar] [CrossRef]
- Mozzherin, D.J.; Shibutani, S.; Tan, C.K.; Downey, K.M.; Fisher, P.A. Proliferating cell nuclear antigen promotes DNA synthesis past template lesions by mammalian DNA polymerase δ. Proc. Natl. Acad. Sci. USA 1997, 94, 6126–6131. [Google Scholar] [CrossRef] [PubMed]
- Chaim, I.A.; Nagel, Z.D.; Jordan, J.J.; Mazzucato, P.; Ngo, L.P.; Samson, L.D. In vivo measurements of interindividual differences in DNA glycosylases and APE1 activities. Proc. Natl. Acad. Sci. USA 2017, 114, E10379–E10388. [Google Scholar] [CrossRef] [PubMed]
| Codon No. and Amino Acid. | Codon (5′→3′) | Nucleotide Position and the Nature of the Mismatch (NTS:TS) | Expected Amino Acid Change | % EGFP Expression |
|---|---|---|---|---|
| 204 Thr | ACC | 610A:C | T204A | 20–75% |
| 610A:G | T204P | <20% | ||
| 610A:A | T204S | no data 1 | ||
| 611C:T | T204N | 20–75% | ||
| 611C:C | T204S | 20–75% | ||
| 611C:A | T204I | 20–75% | ||
| 612C:T | = | not screened | ||
| 612C:C | = | not screened | ||
| 612C:A | = | not screened | ||
| 205 Gln | CAG | 613C:T | Q205K | >75% |
| 613C:C | Q205E | 20–75% | ||
| 613C:A | Q205 * | <20% | ||
| 614A:C | Q205R | >75% | ||
| 614A:G | Q205P | <20% | ||
| 614A:A | Q205L | 20–75% | ||
| 615G:T | = | not screened | ||
| 615G:G | Q205H | >75% | ||
| 615G:A | Q205H | >75% | ||
| 206 Ser | UCC | 616T:T | S206T | >75% 2 |
| 616T:C | S206A | <20% | ||
| 616T:G | S206P | <20% | ||
| 617C:T | S206Y | <20% | ||
| 617C:C | S206C | 20–75% | ||
| 617C:A | S206F | <20% | ||
| 618C:T | = | not screened | ||
| 618C:C | = | not screened | ||
| 618C:A | = | not screened | ||
| 207 Ala | GCC | 619G:T | A207T | >75% |
| 619G:G | A207P | <20% | ||
| 619G:A | A207S | >75% | ||
| 620C:T | A207D | 20–75% | ||
| 620C:C | A207G | 20–75% | ||
| 620C:A | A207V | 20–75% |
| Non-Fluorescent Mutants | Detectable Secondary Mutations | |||||
|---|---|---|---|---|---|---|
| Protein | DNA | Base Pair (NTS:TS) | DNA TS 3′→5′ (TLS) | RNA 5′→3′ (TM) | Amino Acid Change 1 | Brightness (% EGFP) 2 |
| Q205 * | c.613C>T | T:A | GTC | CAG | *205Q | 100 |
| CTC | GAG | *205E | ~70 | |||
| TTC | AAG | *205K | ~100 | |||
| Q205P | c.614A>C | C:G | GTC | CAG | P205Q | 100 |
| GAC | CUG | P205L | ~80 | |||
| GCC | CGG | P205R | ~100 | |||
| S206Y | c.617C>A | A:T | AGG | UCC | Y206S | 100 |
| AAG | UUC | Y206F | ~3 | |||
| ACG | UGC | Y206C | ~20 | |||
| A207P | c.619G>C | C:G | CGG | GCC | P207A | 100 |
| AGG | UCC | P207S | ~100 | |||
| TGG | ACC | P207T | ~100 | |||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rodriguez-Alvarez, M.; Kim, D.; Khobta, A. EGFP Reporters for Direct and Sensitive Detection of Mutagenic Bypass of DNA Lesions. Biomolecules 2020, 10, 902. https://doi.org/10.3390/biom10060902
Rodriguez-Alvarez M, Kim D, Khobta A. EGFP Reporters for Direct and Sensitive Detection of Mutagenic Bypass of DNA Lesions. Biomolecules. 2020; 10(6):902. https://doi.org/10.3390/biom10060902
Chicago/Turabian StyleRodriguez-Alvarez, Marta, Daria Kim, and Andriy Khobta. 2020. "EGFP Reporters for Direct and Sensitive Detection of Mutagenic Bypass of DNA Lesions" Biomolecules 10, no. 6: 902. https://doi.org/10.3390/biom10060902
APA StyleRodriguez-Alvarez, M., Kim, D., & Khobta, A. (2020). EGFP Reporters for Direct and Sensitive Detection of Mutagenic Bypass of DNA Lesions. Biomolecules, 10(6), 902. https://doi.org/10.3390/biom10060902





