Structural Studies of HNA Substrate Specificity in Mutants of an Archaeal DNA Polymerase Obtained by Directed Evolution
Abstract
1. Introduction
2. Materials and Methods
2.1. TgoT, TgoT_6G12, TgoT_RT521 Polymerases
2.1.1. TgoT DNA Polymerase
2.1.2. TgoT_6G12 DNA Polymerase
2.1.3. TgoT_RT521 Polymerase
2.2. Purification
2.3. Protein Crystallography
2.3.1. Duplex Preparation
2.3.2. TgoT DNA Polymerase
2.3.3. TgoT_RT521 DNA Polymerase
2.3.4. TgoT_6G12 DNA Polymerase
2.4. Mass Spectrometry
2.4.1. DNA Extraction
2.4.2. Mass Spectrometry
2.5. Molecular Dynamics (MD) Protocol
2.5.1. System Preparation
2.5.2. MD Simulations
2.5.3. HNA-DNA Duplex Conformation Analysis
3. Results
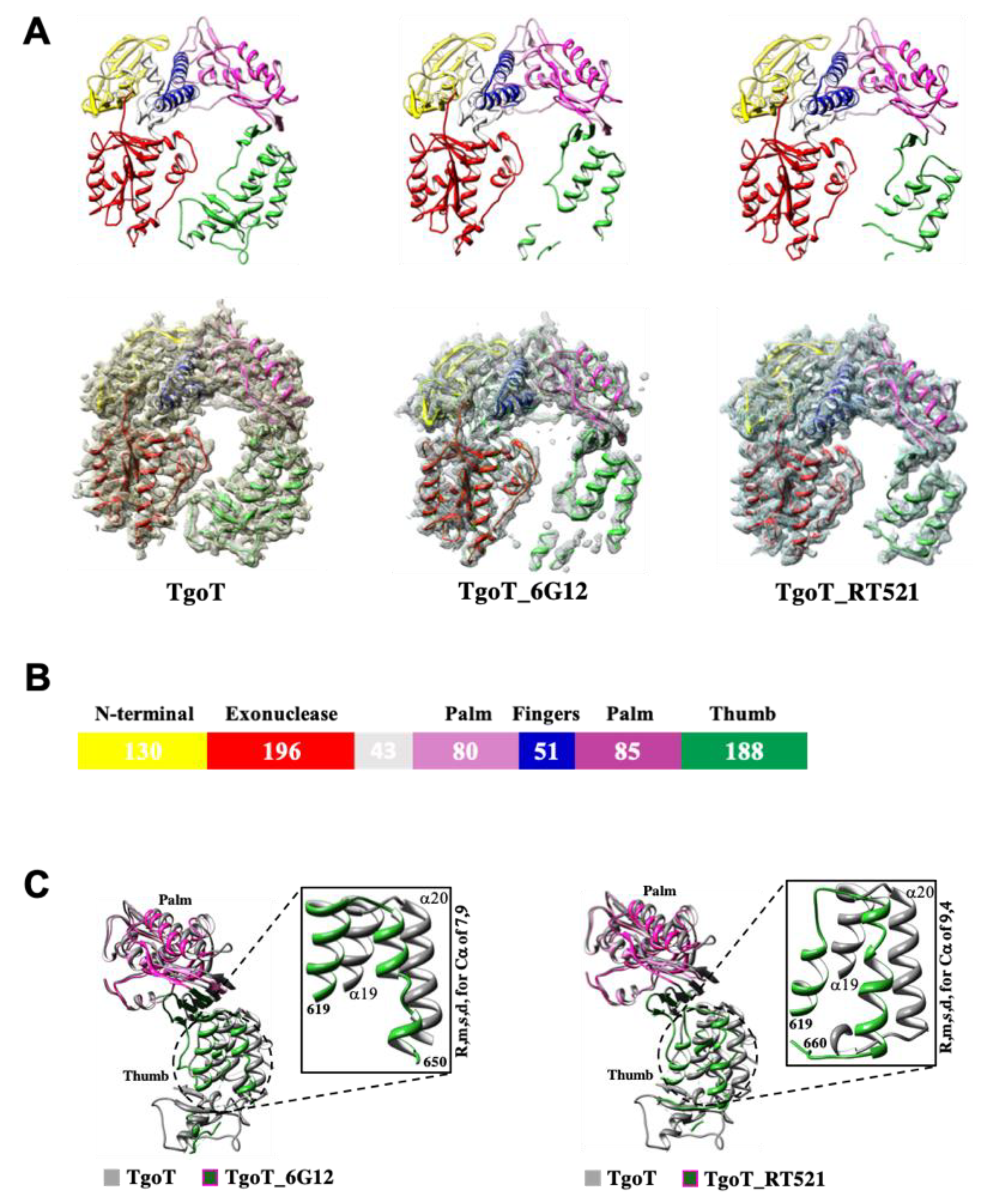
3.1. Structure Analysis of the Apo Form of Three Tgo Polymerase Variants
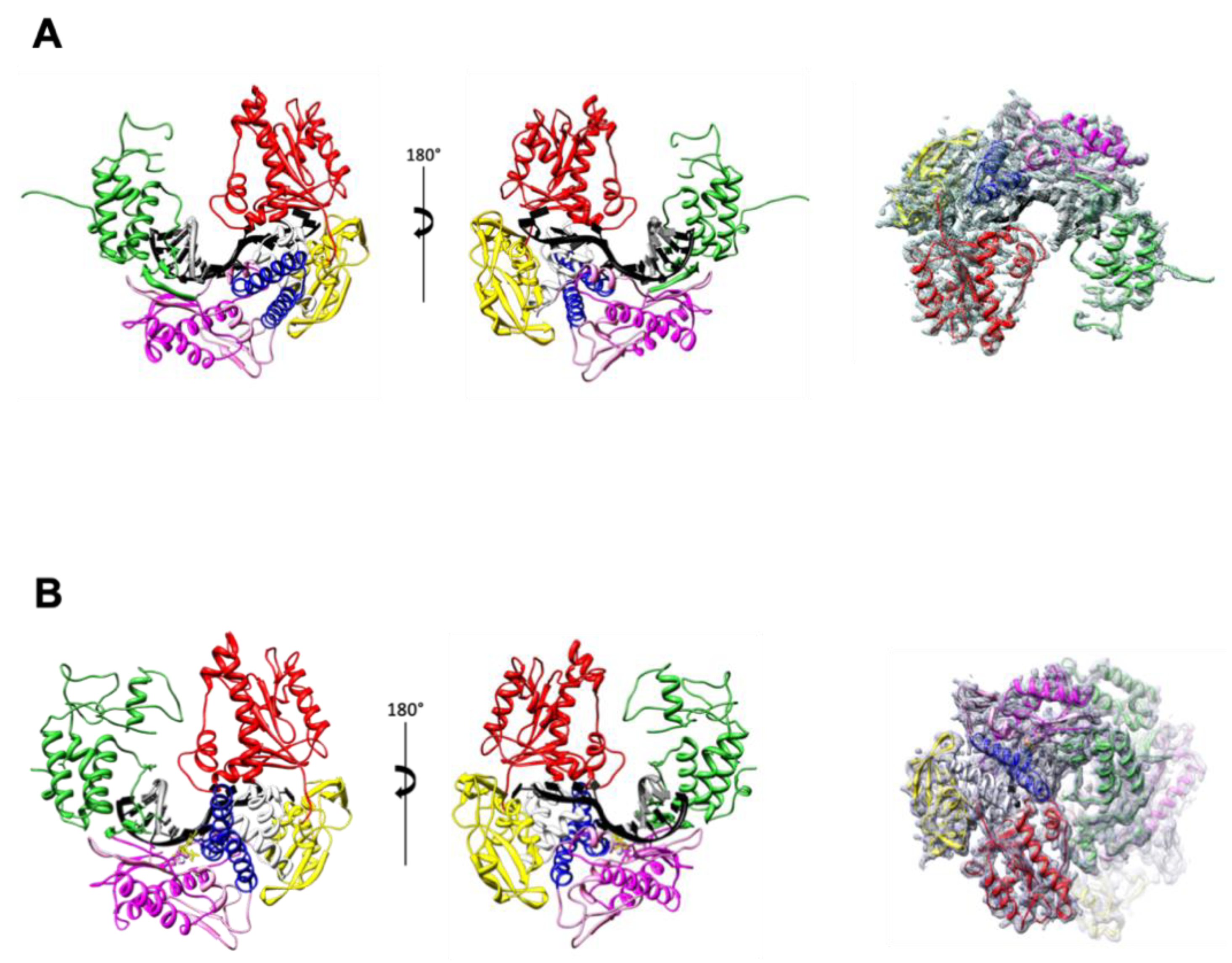
3.2. TgoT_6G12 in Binary and Ternary Complexes with DNA
3.2.1. Crystallization
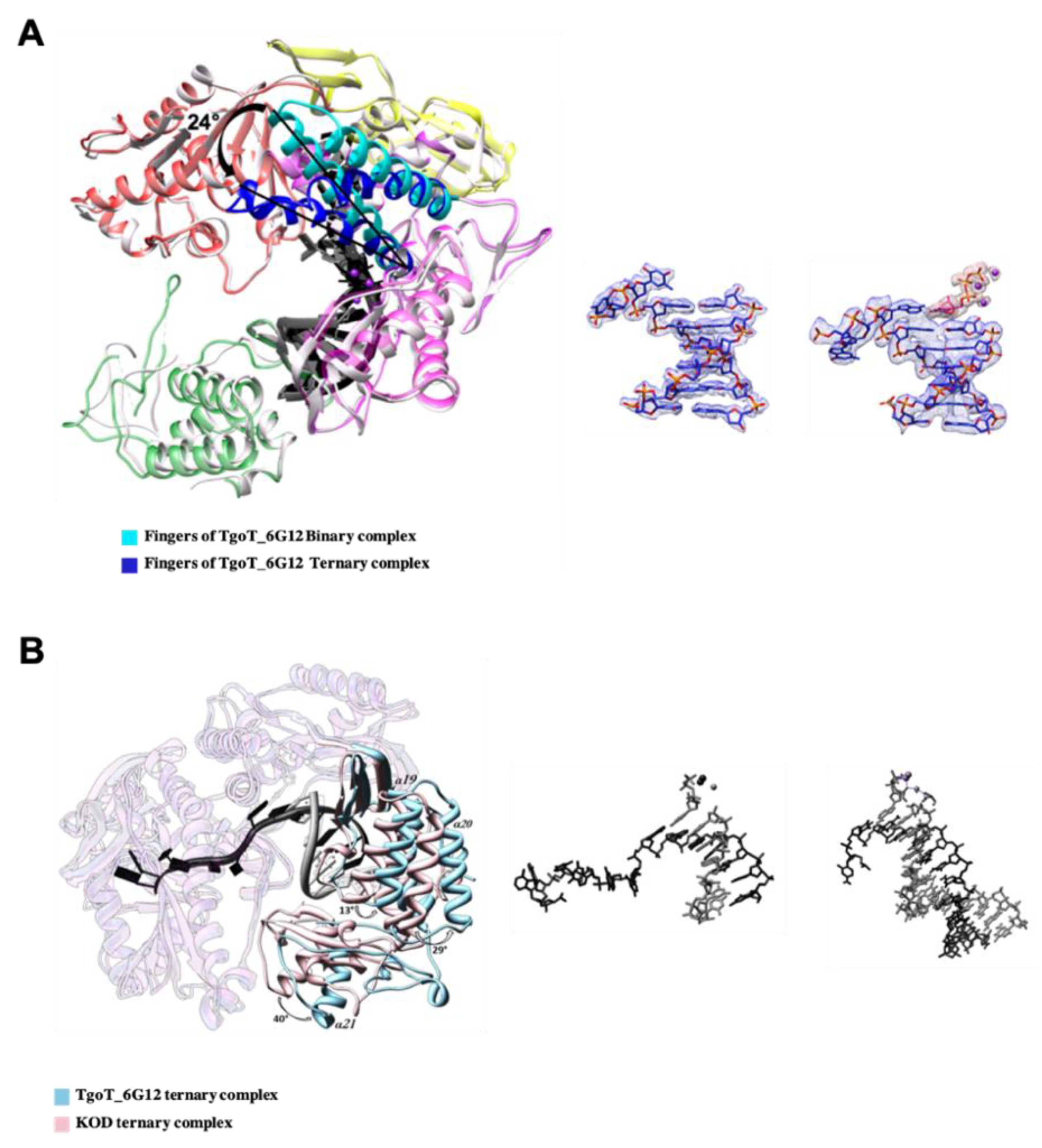
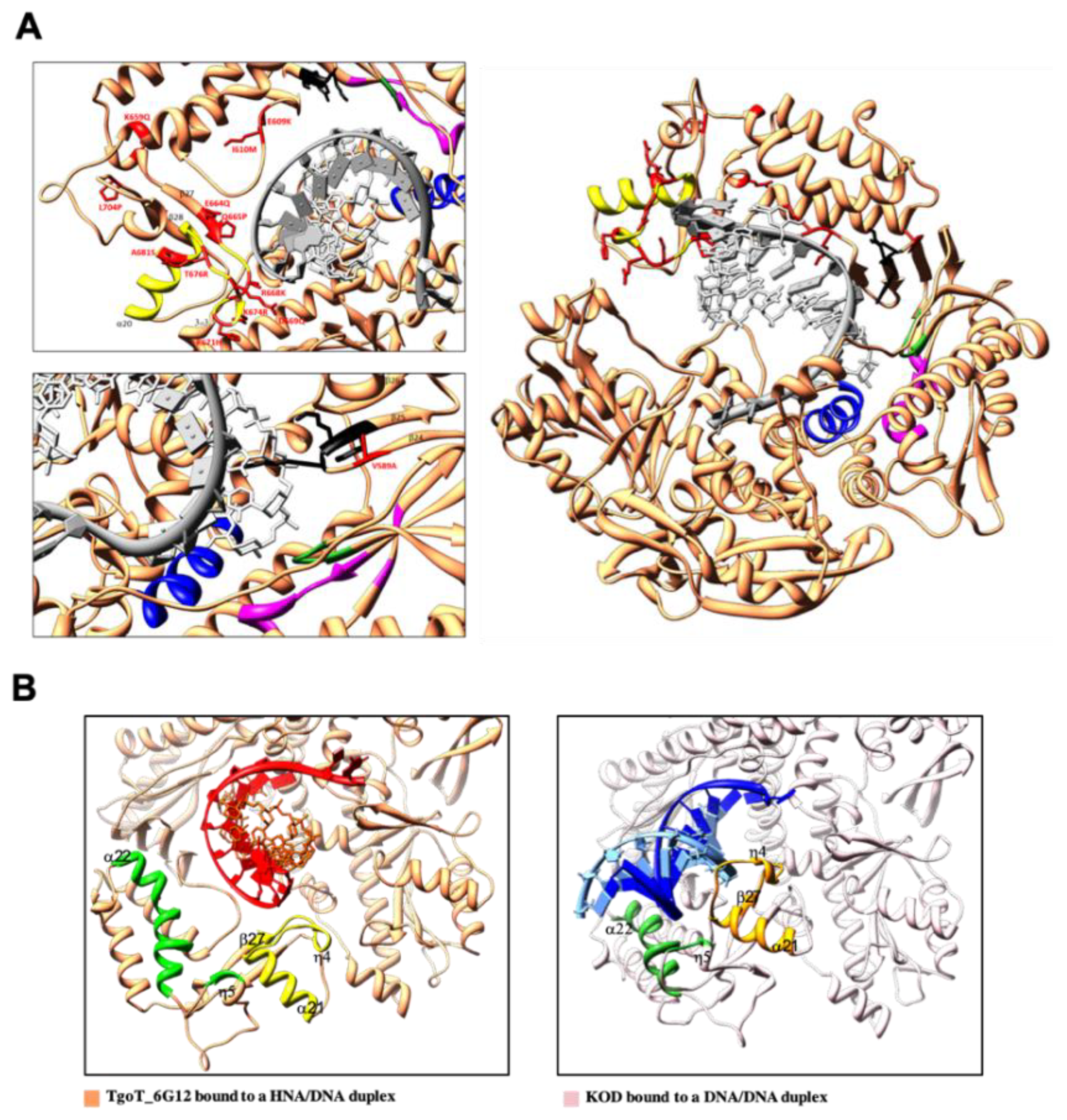
3.2.2. Conformational Changes
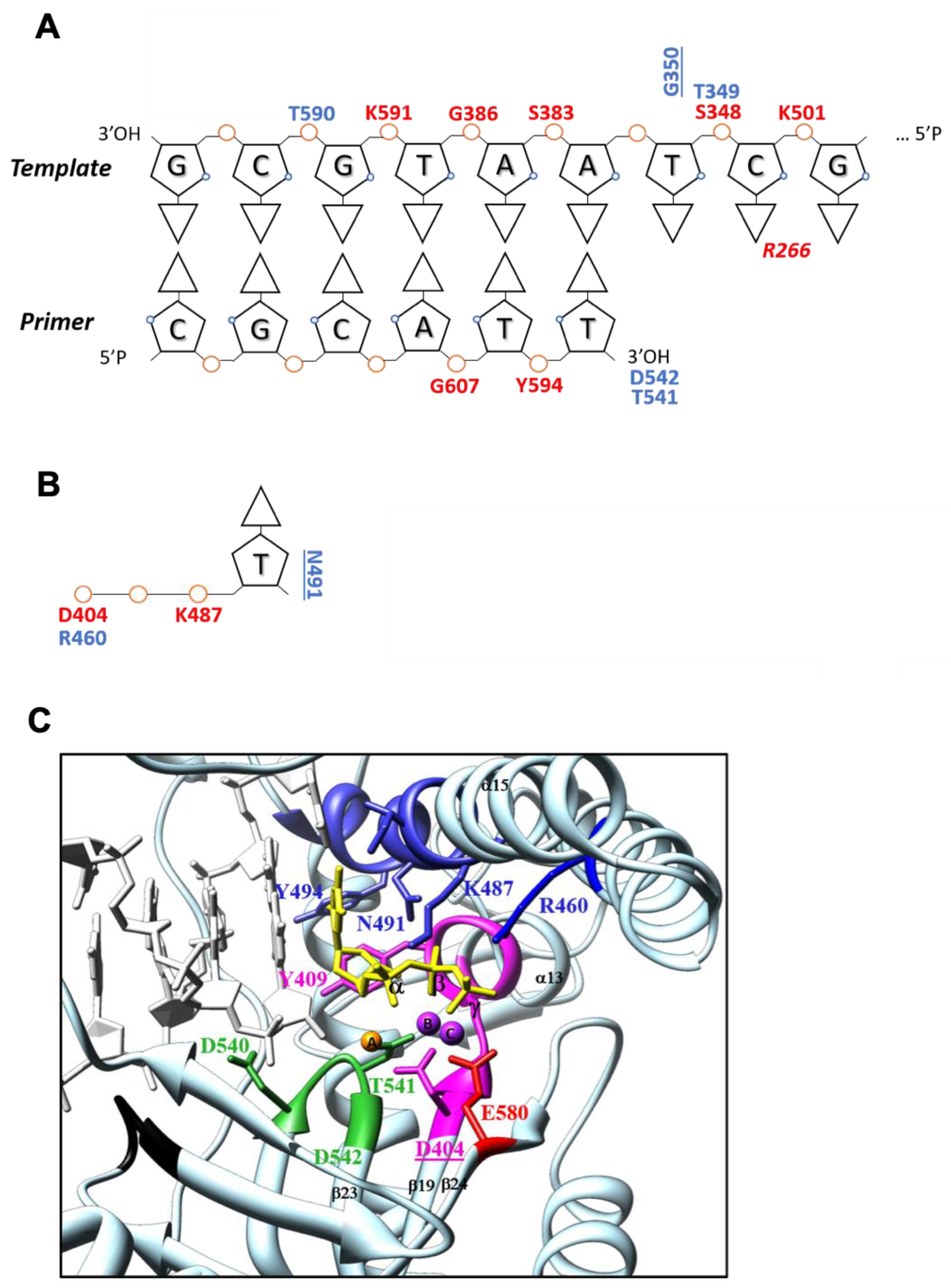
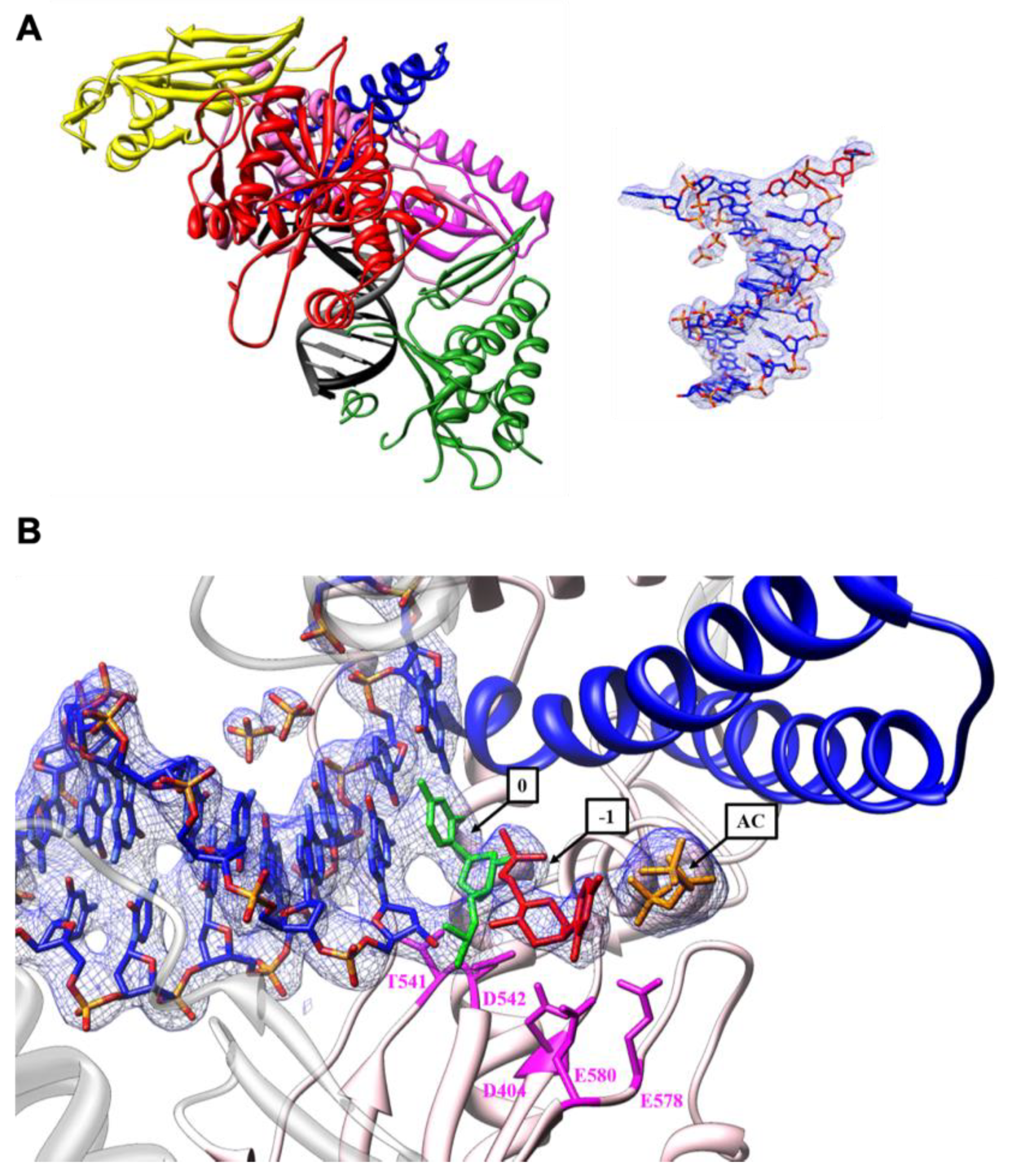
3.2.3. DNA Interactions
3.2.4. Active Site Analysis
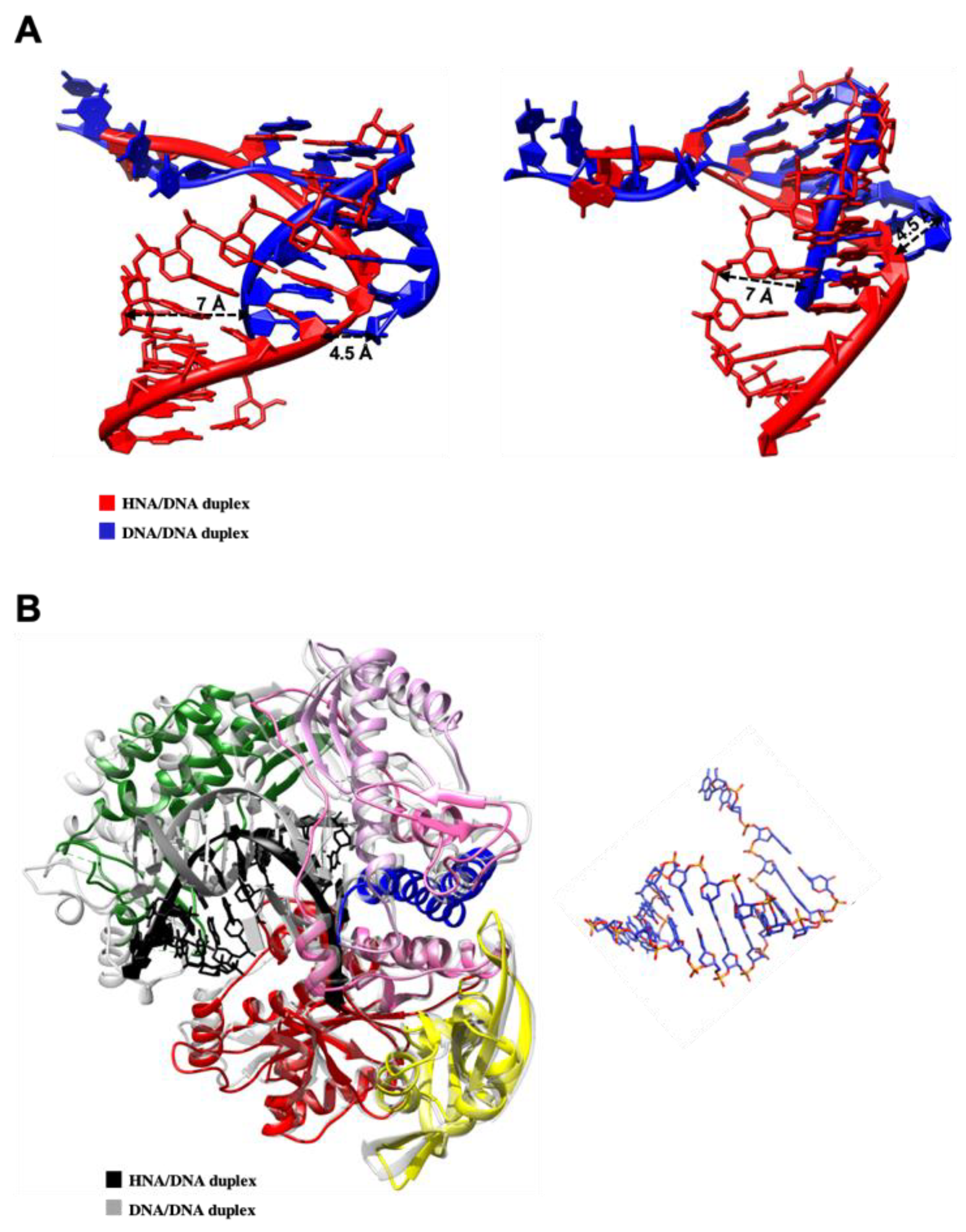
3.3. TgoT_6G12 Binding Mode to the HNA-DNA Duplex: In Silico Approach
3.3.1. HNA-DNA Hybrid Duplex
3.3.2. HNA-Specific TgoT Mutation Analysis in a Structural Context
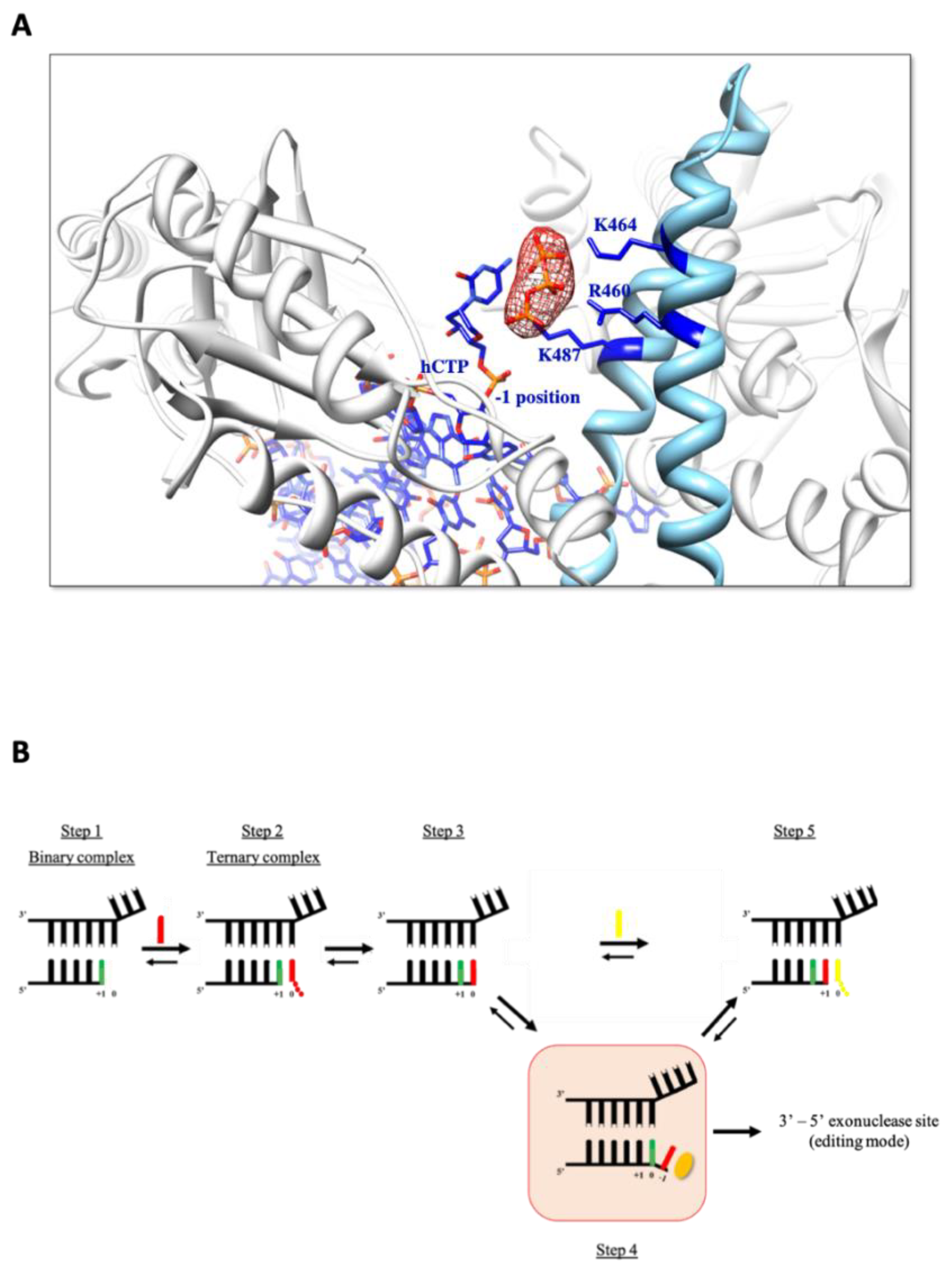
3.4. One Nucleotide-Backtracked State for TgoT_6G12 after Two Successive HNA Synthesis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Johansson, E.; Dixon, N. Replicative DNA Polymerases. Cold Spring Harb. Perspect. Biol. 2013, 5, a012799. [Google Scholar] [CrossRef] [PubMed]
- Lehman, I.R.; Bessman, M.J.; Simms, E.S.; Kornberg, A. Enzymatic synthesis of deoxyribonucleic acid. I. Preparation of substrates and partial purification of an enzyme from Escherichia coli. J. Biol. Chem. 1958, 233, 163–170. [Google Scholar] [PubMed]
- Kohlstaedt, A.L.; Wang, J.; Friedman, J.M.; Rice, A.P.; Steitz, A.T. Crystal structure at 3.5 A resolution of HIV-1 reverse transcriptase complexed with an inhibitor. Science 1992, 256, 1783–1790. [Google Scholar] [CrossRef] [PubMed]
- Delarue, M.; Poch, O.; Tordo, N.; Moras, D.; Argos, P. An attempt to unify the structure of polymerases. Protein Eng. Des. Sel. 1990, 3, 461–467. [Google Scholar] [CrossRef] [PubMed]
- Braithwaite, D.K.; Ito, J. Compilation, alignment, and phylogenetic relationships of DNA polymerases. Nucleic Acids Res. 1993, 21, 787–802. [Google Scholar] [CrossRef]
- Burgers, P.M.J.; Koonin, E.V.; Bruford, E.; Blanco, L.; Burtis, K.C.; Christman, M.F.; Copeland, W.C.; Friedberg, E.C.; Hanaoka, F.; Hinkle, D.C.; et al. Eukaryotic DNA Polymerases: Proposal for a Revised Nomenclature. J. Biol. Chem. 2001, 276, 43487–43490. [Google Scholar] [CrossRef]
- Wu, S.; Beard, W.A.; Pedersen, L.G.; Wilson, S.H. Structural Comparison of DNA Polymerase Architecture Suggests a Nucleotide Gateway to the Polymerase Active Site. Chem. Rev. 2014, 114, 2759–2774. [Google Scholar] [CrossRef][Green Version]
- Raia, P.; Carroni, M.; Henry, E.; Pehau-Arnaudet, G.; Brûlé, S.; Béguin, P.; Henneke, G.; Lindahl, E.; Delarue, M.; Sauguet, L. Structure of the DP1–DP2 PolD complex bound with DNA and its implications for the evolutionary history of DNA and RNA polymerases. PLoS Biol. 2019, 17, e3000122. [Google Scholar] [CrossRef]
- Batra, V.K.; Beard, W.A.; Shock, D.D.; Krahn, J.M.; Pedersen, L.C.; Wilson, S.H. Magnesium-Induced Assembly of a Complete DNA Polymerase Catalytic Complex. Structure 2006, 14, 757–766. [Google Scholar] [CrossRef]
- Swan, M.K.; Johnson, R.E.; Prakash, L.; Prakash, S.; Aggarwal, A.K. Structural basis of high-fidelity DNA synthesis by yeast DNA polymerase delta. Nat. Struct. Mol. Biol. 2009, 16, 979–986. [Google Scholar] [CrossRef]
- Kropp, H.M.; Betz, K.; Wirth, J.; Diederichs, K.; Marx, A. Crystal structures of ternary complexes of archaeal B-family DNA polymerases. PLoS ONE 2017, 12, e0188005. [Google Scholar] [CrossRef] [PubMed]
- Jamsen, J.A.; Beard, W.A.; Pedersen, L.C.; Shock, D.D.; Moon, A.F.; Krahn, J.M.; Bebenek, K.; Kunkel, T.A.; Wilson, S.H. Time-lapse crystallography snapshots of a double-strand break repair polymerase in action. Nat. Commun. 2017, 8, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Pelletier, H.; Sawaya, M.R.; Kumar, A.; Wilson, S.H.; Kraut, J. Structures of ternary complexes of rat DNA polymerase beta, a DNA template-primer, and ddCTP. Science 1994, 264, 1891–1903. [Google Scholar] [CrossRef] [PubMed]
- Gardner, A.F.; Kelman, Z. DNA polymerases in biotechnology. Front. Microbiol. 2014, 5, 659. [Google Scholar] [CrossRef] [PubMed]
- Aschenbrenner, J.; Marx, A. DNA polymerases and biotechnological applications. Curr. Opin. Biotechnol. 2017, 48, 187–195. [Google Scholar] [CrossRef]
- Herdewijn, P.; Marlière, P. Toward safe genetically modified organisms through the chemical diversification of nucleic acids. Chem. Biodivers. 2009, 6, 791–808. [Google Scholar] [CrossRef]
- Ruckman, J.; Green, L.S.; Beeson, J.; Waugh, S.; Gillette, W.L.; Henninger, D.D.; Claesson-Welsh, L.; Janjic, N. 2′-Fluoropyrimidine RNA-based Aptamers to the 165-Amino Acid Form of Vascular Endothelial Growth Factor (VEGF165). J. Biol. Chem. 1998, 273, 20556–20567. [Google Scholar] [CrossRef]
- Herdewijn, P. Nucleic Acids with a Six-Membered ‘Carbohydrate’ Mimic in the Backbone. Chem Biodivers. 2010, 7, 1–59. [Google Scholar] [CrossRef]
- Hasegawa, T.; Shoji, A.; Kuwahara, M.; Ozaki, H.; Sawai, H. Synthesis and property of DNA labeled with fluorescent acridone. Nucleic Acids Symp. Ser. 2006, 50, 145–146. [Google Scholar] [CrossRef]
- Kuwahara, M.; Suto, Y.; Minezaki, S.; Kitagata, R.; Nagashima, J.-I.; Sawai, H. Substrate property and incorporation accuracy of various dATP analogs during enzymatic polymerization using thermostable DNA polymerases. Nucleic Acids Symp. Ser. 2006, 50, 31–32. [Google Scholar] [CrossRef][Green Version]
- Lee, A.H.F.; Kool, E.T. A New Four-Base Genetic Helix, yDNA, Composed of Widened Benzopyrimidine−Purine Pairs. J. Am. Chem. Soc. 2005, 127, 3332–3338. [Google Scholar] [CrossRef]
- Liu, H.; Gao, J.; Lynch, S.R.; Saito, Y.D.; Maynard, L.; Kool, E.T. A four-base paired genetic helix with expanded size. Science 2003, 302, 868–871. [Google Scholar] [CrossRef]
- Hoshika, S.; Chen, F.; Leal, N.A.; Benner, S.A. Artificial Genetic Systems. Self-Avoiding DNA in PCR and Multiplexed PCR. Angew. Chem. Int. Ed. Engl. 2010, 49, 5554–5557. [Google Scholar] [CrossRef] [PubMed]
- Mahal, L. Faculty Opinions recommendation of Expanded genetic alphabets in the polymerase chain reaction. Angew. Chem. Int. Ed. Engl. 2010, 49, 177–180. [Google Scholar] [CrossRef]
- Pinheiro, V.B.; Holliger, P. The XNA world: Progress towards replication and evolution of synthetic genetic polymers. Curr. Opin. Chem. Biol. 2012, 16, 245–252. [Google Scholar] [CrossRef] [PubMed]
- Saenger, W. Principles of Nucleic Acid Structure; Springer: New York, NY, USA, 1984. [Google Scholar]
- Maier, T.; Przylas, I.; Strater, N.; Herdewijn, A.P.; Saenger, W. Reinforced HNA Backbone Hydration in the Crystal Structure of a Decameric HNA/RNA Hybrid. J. Am. Chem. Soc. 2005, 127, 2937–2943. [Google Scholar] [CrossRef] [PubMed]
- Chaput, J.C.; Herdewijn, P.; Hollenstein, M. Orthogonal Genetic Systems. ChemBioChem 2020, 21, 1408–1411. [Google Scholar] [CrossRef]
- Lescrinier, E.; Esnouf, R.; Schraml, J.; Busson, R.; Heus, H.; Hilbers, C.; Herdewijn, P. Solution structure of a HNA-RNA hybrid. Chem. Biol. 2000, 7, 719–731. [Google Scholar] [CrossRef]
- Boiziau, C.; Toulmé, J.-J. A Method to Select Chemically Modified Aptamers Directly. Antisense Nucleic Acid Drug Dev. 2001, 11, 379–385. [Google Scholar] [CrossRef]
- Pinheiro, V.B.; Taylor, A.I.; Cozens, C.; Abramov, M.; Renders, M.; Zhang, S.; Chaput, J.C.; Wengel, J.; Peak-Chew, S.-Y.; McLaughlin, S.H.; et al. Synthetic Genetic Polymers Capable of Heredity and Evolution. Science 2012, 336, 341–344. [Google Scholar] [CrossRef]
- Chim, N.; Shi, C.; Sau, S.P.; Nikoomanzar, A.; Chaput, J.C. Structural basis for TNA synthesis by an engineered TNA polymerase. Nat. Commun. 2017, 8, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Jackson, L.N.; Chim, N.; Shi, C.; Chaput, J.C. Crystal structures of a natural DNA polymerase that functions as an XNA reverse transcriptase. Nucleic Acids Res. 2019, 47, 6973–6983. [Google Scholar] [CrossRef] [PubMed]
- Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 2010, 66 Pt 2, 125–132. [Google Scholar] [CrossRef]
- Adams, P.D.; Afonine, P.V.; Bunkóczi, G.; Chen, V.B.; Davis, I.W.; Echols, N.; Headd, J.J.; Hung, L.-W.; Kapral, G.J.; Grosse-Kunstleve, R.W.; et al. PHENIX: A comprehensive Python-based system for macromolecular structure solution. Int. Tables Crystallogr. 2012, 66 Pt 2, 539–547. [Google Scholar] [CrossRef]
- Hopfner, K.-P.; Eichinger, A.; Engh, R.A.; Laue, F.; Ankenbauer, W.; Huber, R.; Angerer, B. Crystal structure of a thermostable type B DNA polymerase from Thermococcus gorgonarius. Proc. Natl. Acad. Sci. USA 1999, 96, 3600–3605. [Google Scholar] [CrossRef]
- Bricogne, G.; Blanc, E.; Brandl, M.; Flensburg, C.; Keller, P.; Paciorek, W.; Roversi, P.; Sharff, A.; Smart, O.S.; Vonrhein, C.; et al. BUSTER Version X.Y.Z.; Global Phasing Ltd.: Cambridge, UK, 2017. [Google Scholar]
- Emsley, P.; Cowtan, K. Coot: Model-building tools for molecular graphics. Acta Crystallogr. Sect. D Biol. Crystallogr. 2004, 60, 2126–2132. [Google Scholar] [CrossRef]
- Roberts, E.; Eargle, J.; Wright, D.; Luthey-Schulten, Z. MultiSeq: Unifying sequence and structure data for evolutionary analysis. BMC Bioinform. 2006, 7, 382. [Google Scholar] [CrossRef]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visual molecular dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef]
- Jorgensen, W.L.; Chandrasekhar, J.; Madura, J.D.; Impey, R.W.; Klein, M.L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 1983, 79, 926–935. [Google Scholar] [CrossRef]
- Maier, J.A.; Martinez, C.; Kasavajhala, K.; Wickstrom, L.; Hauser, K.E.; Simmerling, C. ff14SB: Improving the Accuracy of Protein Side Chain and Backbone Parameters from ff99SB. J. Chem. Theory Comput. 2015, 11, 3696–3713. [Google Scholar] [CrossRef]
- Ivani, I.; Dans, P.D.; Noy, A.; Pérez, A.; Faustino, I.; Hospital, A. Parmbsc1: A refined force field for DNA simulations. Nat. Methods 2016, 13, 55–58. [Google Scholar] [CrossRef] [PubMed]
- De Winter, H.; Lescrinier, E.; Van Aerschot, A.; Herdewijn, P. Molecular Dynamics Simulation To Investigate Differences in Minor Groove Hydration of HNA/RNA Hybrids As Compared to HNA/DNA Complexes. J. Am. Chem. Soc. 1998, 120, 5381–5394. [Google Scholar] [CrossRef]
- Darden, A.T.; York, D.M.; Pedersen, L. Particle mesh Ewald: AnN⋅log(N) method for Ewald sums in large systems. J. Chem. Phys. 1993, 98, 10089–10092. [Google Scholar] [CrossRef]
- Essmann, U.; Perera, L.; Berkowitz, M.L.; Darden, T.; Lee, H.; Pedersen, L.G. A smooth particle mesh Ewald method. J. Chem. Phys. 1995, 103, 8577–8593. [Google Scholar] [CrossRef]
- Hess, B.; Bekker, H.; Berendsen, H.J.; Fraaije, J.G. LINCS: A Linear Constraint Solver for molecular simulations. J. Comput. Chem. 1997, 18, 1463–1472. [Google Scholar] [CrossRef]
- Hess, B. P-LINCS: A Parallel Linear Constraint Solver for Molecular Simulation. J. Chem. Theory Comput. 2008, 4, 116–122. [Google Scholar] [CrossRef]
- Pronk, S.; Páll, S.; Schulz, R.; Larsson, P.; Bjelkmar, P.; Apostolov, R.; Shirts, M.R.; Smith, J.C.; Kasson, P.M.; Van Der Spoel, D.; et al. GROMACS 4.5: A high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 2013, 29, 845–854. [Google Scholar] [CrossRef]
- Van Der Spoel, D.; Lindahl, E.; Hess, B.; Groenhof, G.; Mark, A.E.; Berendsen, H.J.C. GROMACS: Fast, flexible, and free. J. Comput. Chem. 2005, 26, 1701–1718. [Google Scholar] [CrossRef]
- Berendsen, H.J.C.; Postma, J.P.M.; Van Gunsteren, W.F.; DiNola, A.; Haak, J.R. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 1984, 81, 3684–3690. [Google Scholar] [CrossRef]
- Bussi, G.; Donadio, D.; Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 2007, 126, 014101. [Google Scholar] [CrossRef]
- Westhof, E.; Romby, P.; Romaniuk, P.J.; Ebel, J.-P.; Ehresmann, C.; Ehresmann, B. Computer modeling from solution data of spinach chloroplast and of Xenopus laevis somatic and oocyte 5 S rRNAs. J. Mol. Biol. 1989, 207, 417–431. [Google Scholar] [CrossRef]
- Lu, X.-J.; Olson, W.K. 3DNA: A versatile, integrated software system for the analysis, rebuilding and visualization of three-dimensional nucleic-acid structures. Nat. Protoc. 2008, 3, 1213–1227. [Google Scholar] [CrossRef] [PubMed]
- Colasanti, A.V.; Lu, X.-J.; Olson, W.K. Analyzing and Building Nucleic Acid Structures with 3DNA. J. Vis. Exp. 2013, 74, e4401. [Google Scholar] [CrossRef] [PubMed]
- Bergen, K.; Betz, K.; Welte, W.; Diederichs, K.; Marx, A. Structures of KOD and 9° N DNA Polymerases Complexed with Primer Template Duplex. ChemBioChem 2013, 14, 1058–1062. [Google Scholar] [CrossRef] [PubMed]
- Rodrigueza, A.C.; Parka, H.-W.; Maoa, C.; Beese, L.S. Crystal structure of a pol α family DNA polymerase from the hyperthermophilic archaeon Thermococcus sp. 9° N-711Edited by D. Rees. J. Mol. Biol. 2000, 299, 447–462. [Google Scholar] [CrossRef] [PubMed]
- Brown, J.A.; Suo, Z. Unlocking the Sugar ‘Steric Gate’ of DNA Polymerases. Biochemistry 2011, 22, 1135–1142. [Google Scholar] [CrossRef]
- Pinheiro, V.B. Engineering-driven biological insights into DNA polymerase mechanism. Curr. Opin. Biotechnol. 2019, 60, 9–16. [Google Scholar] [CrossRef]
- Pavlov, A.R.; Pavlova, N.V.; Kozyavkin, S.A.; Slesarev, A.I. Recent developments in the optimization of thermostable DNA polymerases for efficient applications. Trends Biotechnol. 2004, 22, 253–260. [Google Scholar] [CrossRef]
- Houlihan, G.; Arangundy-Franklin, S.; Holliger, S.T.J.A.P. Exploring the Chemistry of Genetic Information Storage and Propagation through Polymerase Engineering. Accounts Chem. Res. 2017, 50, 1079–1087. [Google Scholar] [CrossRef]
- Anosova, I.; Kowal, E.A.; Sisco, N.J.; Sau, S.; Liao, J.-Y.; Bala, S. Structural Insights into Conformation Differences between DNA/TNA and RNA/TNA Chimeric Duplexes. Chembiochem. Eur. J. Chem. Biol. 2016, 15, 1705–1708. [Google Scholar] [CrossRef]
- Turtola, M.; Mäkinen, J.J.; Belogurov, A.G. Active site closure stabilizes the backtracked state of RNA polymerase. Nucleic Acids Res. 2018, 46, 10870–10887. [Google Scholar] [CrossRef] [PubMed]
| Type | Number of Residues | Number of Atoms |
|---|---|---|
| Protein | 755 | 12,435 |
| DNA | 13 | 414 |
| HNA | 11 | 378 |
| Mn2+ | 2 | 2 |
| Water | 31,407 | 94,221 |
| Na+ | 104 | 104 |
| Cl− | 85 | 85 |
| TgoT_Apo | TgoT_RT521_Apo | TgoT_6G12_Apo | TgoT_6G12_Binary | TgoT_6G12_Ternary | TgoT_6G12_Binary_hCTP | |
|---|---|---|---|---|---|---|
| Resolution range | 45.86–2.394 (2.48–2.394) | 43.69–2.35 (2.434–2.35) | 49.53–3.099 (3.21–3.099) | 46.1–2.797 (2.897–2.797) | 48.47–3.15 (3.263–3.15) | 49.74–3.0 (3.107–3.0) |
| Space group | P 21 21 21 | P 21 21 21 | P 21 21 21 | C 1 2 1 | P 21 21 21 | P 21 21 21 |
| Unit cell | 65.06 105.22 152.86 90 90 90 | 70.53 109.85 111.3 90 90 90 | 71.89 110.49 111.77 90 90 90 | 164.49 111.92 74.1 90 110.39 90 | 111.823 112.331 186.518 90 90 90 | 56.21 106.77 231.78 90 90 90 |
| Total reflections | 389,306 (33,918) | 158,299 (14,944) | 132,742 (13,197) | 134,380 (13,082) | 66,626 (3510) | 320,482 (30,357) |
| Unique reflections | 41,950 (3986) | 36,486 (3526) | 16,706 (1621) | 31,062 (3015) | 33,316 (1758) | 28,837 (2829) |
| Multiplicity | 9.3 (8.5) | 4.3 (4.2) | 7.9 (8.1) | 4.3 (4.3) | 2.0 (2.0) | 11.1 (10.7) |
| Completeness (%) | 99.28 (95.65) | 99.08 (97.67) | 99.37 (98.48) | 99.57 (98.11) | 80.49 (43.10) | 99.88 (99.79) |
| Mean I/sigma(I) | 11.45 (0.49) | 7.69 (0.94) | 8.72 (0.82) | 7.18 (0.78) | 17.89 (1.52) | 9.21 (0.53) |
| Wilson B-factor | 72.01 | 57.45 | 101.76 | 79.15 | 88.71 | 109.57 |
| R-merge | 0.1222 (4.034) | 0.09902 (1.565) | 0.193 (2.756) | 0.1749 (1.737) | 0.05693 (0.5029) | 0.2038 (3.171) |
| R-meas | 0.1294 (4.293) | 0.1126 (1.793) | 0.2067 (2.939) | 0.1997 (1.981) | 0.08052 (0.7112) | 0.2136 (3.332) |
| R-pim | 0.042 (1.442) | 0.05237 (0.8555) | 0.07307 (1.008) | 0.09487 (0.9382) | 0.05693 (0.5029) | 0.06285 (1.006) |
| CC1/2 | 0.999 (0.287) | 0.997 (0.486) | 0.998 (0.552) | 0.99 (0.319) | 0.997 (0.53) | 0.998 (0.292) |
| CC* | 1 (0.667) | 0.999 (0.809) | 0.999 (0.843) | 0.997 (0.696) | 0.999 (0.832) | 1 (0.672) |
| Reflections Used in Refinement | 41,845 (3979) | 36,401 (3516) | 16,647 (1619) | 31,027 (3011) | 33,305 (1758) | 28,816 (2825) |
| Reflections used for R-free | 2096 (199) | 1820 (177) | 834 (81) | 1550 (151) | 1648 (93) | 1442 (141) |
| R-work | 0.2410 (0.4986) | 0.225 (0.3986) | 0.2616 (0.4530) | 0.2308 (0.3985) | 0.2319 (0.3545) | 0.2253 (0.4123) |
| R-free | 0.2618 (0.5556) | 0.2469 (0.4188) | 0.2649 (0.5542) | 0.2422 (0.4173) | 0.2472 (0.3294) | 0.2429 (0.4564) |
| CC(work) | 0.939 (0.387) | 0.944 (0.675) | 0.933 (0.535) | 0.923 (0.517) | 0.951 (0.567) | 0.932 (0.418) |
| CC(free) | 0.939 (0.173) | 0.949 (0.713) | 0.841 (0.435) | 0.919 (0.566) | 0.923 (0.435) | 0.909 (0.530) |
| Number of Non-hydrogen atoms | 6174 | 5759 | 6154 | 6429 | 12,971 | 6535 |
| Macromolecules | 6094 | 5641 | 6150 | 6336 | 12,907 | 6472 |
| Ligands | 23 | 118 | 4 | 13 | 64 | 63 |
| Protein Residues | 742 | 685 | 749 | 721 | 1475 | 737 |
| RMS(bonds) | 0.009 | 0.009 | 0.008 | 0.008 | 0.009 | 0.009 |
| RMS(angles) | 1.29 | 1.28 | 1.15 | 1.23 | 1.37 | 1.24 |
| Ramachandran Favored (%) | 97.54 | 96.75 | 97.05 | 98.04 | 96.80 | 97.26 |
| Ramachandran Allowed (%) | 2.46 | 2.95 | 2.82 | 1.96 | 3.20 | 2.74 |
| Ramachandran Outliers (%) | 0.00 | 0.30 | 0.13 | 0.00 | 0.00 | 0.00 |
| Rotamer Outliers (%) | 3.58 | 2.85 | 1.54 | 3.02 | 0.86 | 3.29 |
| Clashscore | 2.77 | 2.48 | 3.49 | 3.99 | 3.24 | 5.54 |
| Average B-Factor | 100.24 | 95.01 | 140.31 | 90.08 | 104.56 | 128.22 |
| Macromolecules | 100.33 | 95.73 | 140.33 | 90.37 | 104.66 | 127.63 |
| Ligands | 156.24 | 60.15 | 109.55 | 152.13 | 84.25 | 188.89 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Samson, C.; Legrand, P.; Tekpinar, M.; Rozenski, J.; Abramov, M.; Holliger, P.; Pinheiro, V.B.; Herdewijn, P.; Delarue, M. Structural Studies of HNA Substrate Specificity in Mutants of an Archaeal DNA Polymerase Obtained by Directed Evolution. Biomolecules 2020, 10, 1647. https://doi.org/10.3390/biom10121647
Samson C, Legrand P, Tekpinar M, Rozenski J, Abramov M, Holliger P, Pinheiro VB, Herdewijn P, Delarue M. Structural Studies of HNA Substrate Specificity in Mutants of an Archaeal DNA Polymerase Obtained by Directed Evolution. Biomolecules. 2020; 10(12):1647. https://doi.org/10.3390/biom10121647
Chicago/Turabian StyleSamson, Camille, Pierre Legrand, Mustafa Tekpinar, Jef Rozenski, Mikhail Abramov, Philipp Holliger, Vitor B. Pinheiro, Piet Herdewijn, and Marc Delarue. 2020. "Structural Studies of HNA Substrate Specificity in Mutants of an Archaeal DNA Polymerase Obtained by Directed Evolution" Biomolecules 10, no. 12: 1647. https://doi.org/10.3390/biom10121647
APA StyleSamson, C., Legrand, P., Tekpinar, M., Rozenski, J., Abramov, M., Holliger, P., Pinheiro, V. B., Herdewijn, P., & Delarue, M. (2020). Structural Studies of HNA Substrate Specificity in Mutants of an Archaeal DNA Polymerase Obtained by Directed Evolution. Biomolecules, 10(12), 1647. https://doi.org/10.3390/biom10121647





