Euglena Central Metabolic Pathways and Their Subcellular Locations
Abstract
1. Introduction
2. Results
2.1. Pathway Localisation from Sequence Information
2.1.1. Metabolic Pathways in Euglena
2.1.2. Metabolic Pathways in the Euglena Plastid
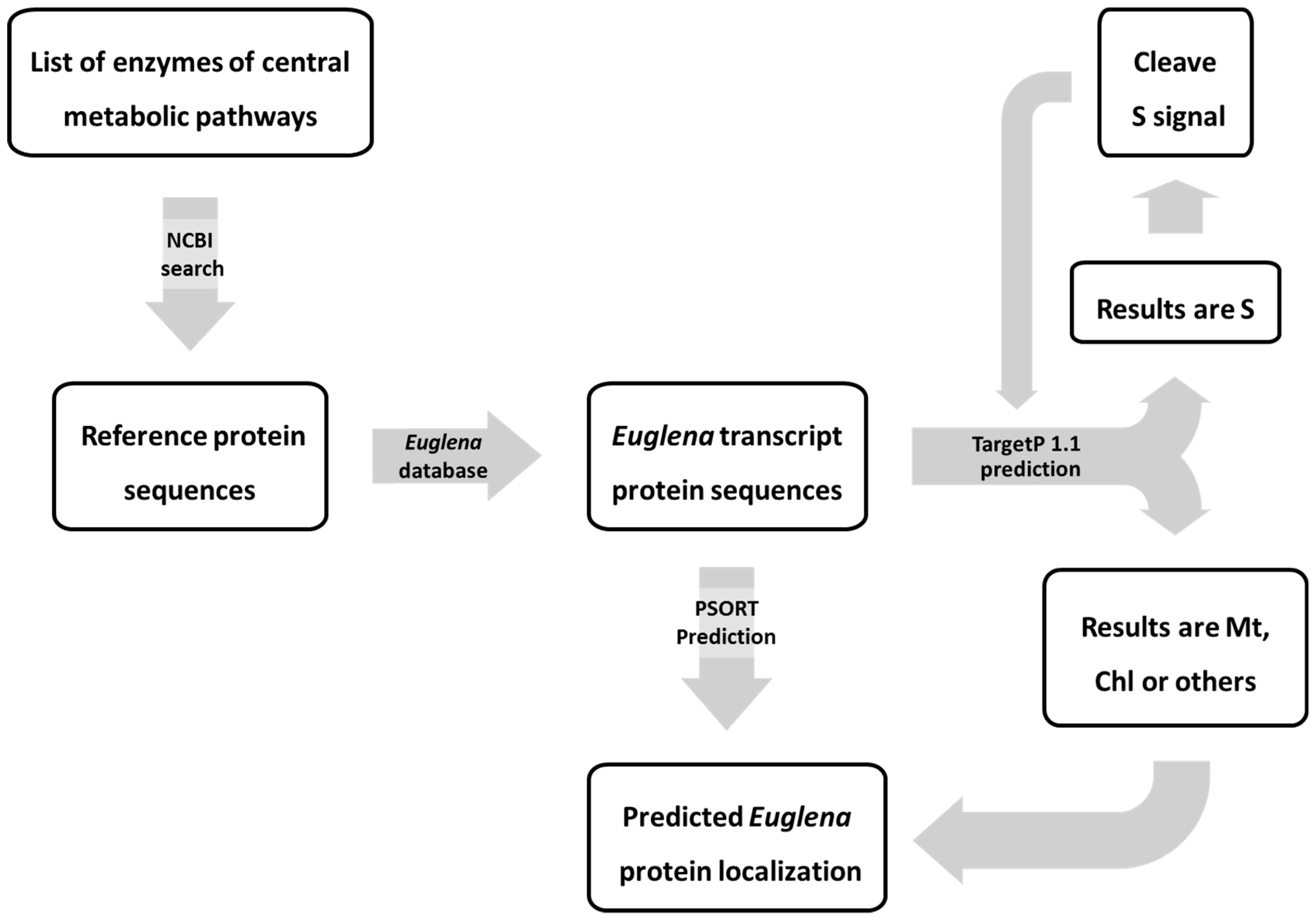
2.1.3. Predicting the Subcellular Location of Euglena Proteins
Mitochondrial Targeting
Proteins without Targeting Signals
Targeting for Secretion
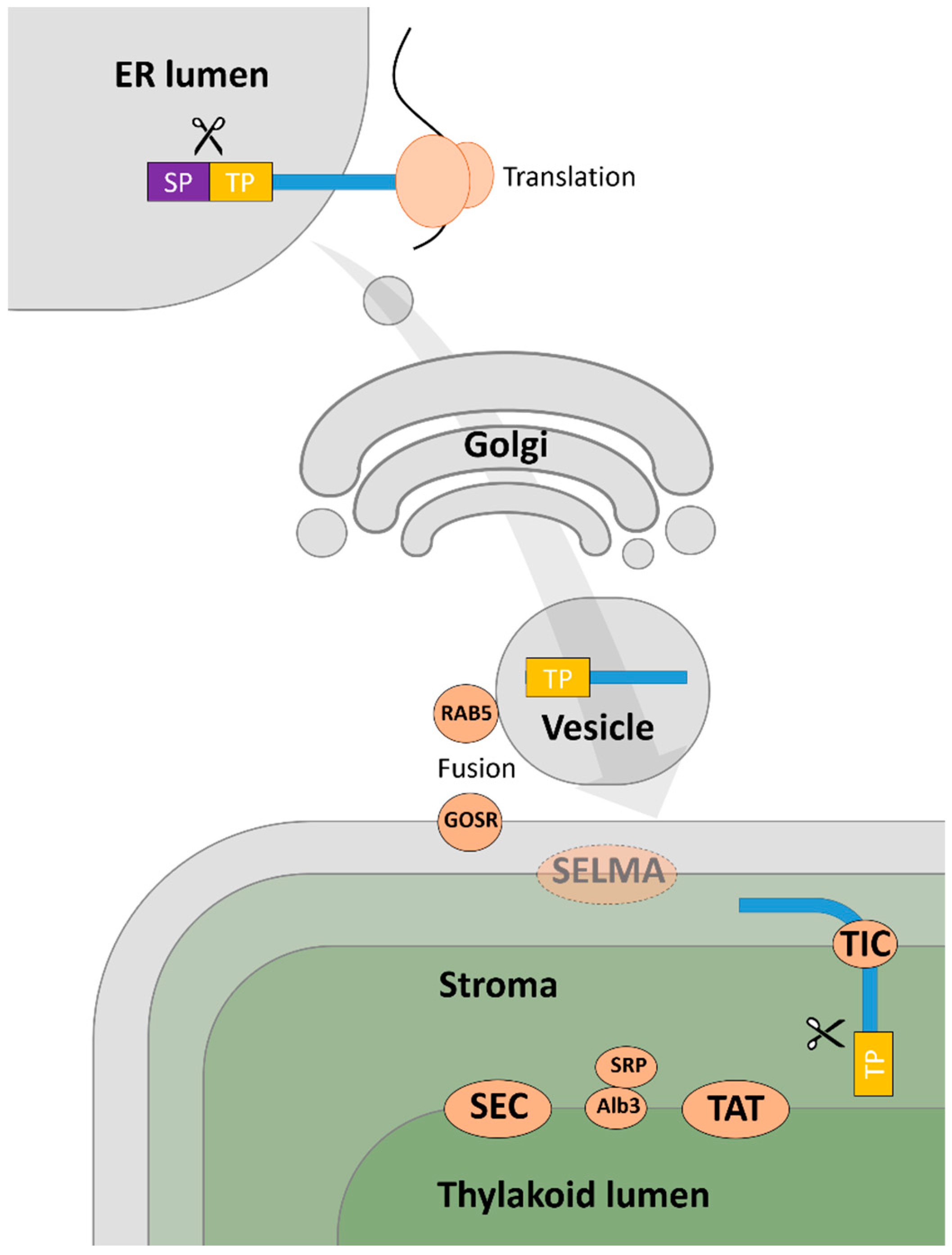
Chloroplast Targeting
2.2. Pathway Localisation from Biochemical/Proteomic Information
2.2.1. Central Metabolic Pathways of Euglena
Glycolysis and Gluconeogenesis
Pentose Phosphate Pathway
Anaplerotic Pathway: Dicarboxylic Acid Bypass
TCA Cycle
Glyoxylate Cycle
C2 Metabolism
2.2.2. Subcellular Locations of Biomass Production
Carbohydrate Biosynthesis
Amino Acid Biosynthesis
Lipid Biosynthesis
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Identification of Euglena Enzymes
5.2. ProteinTargeting Prediction for Euglena
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Einicker-Lamas, M.; Mezian, G.A.; Fernandes, T.B.; Silva, F.L.S.; Guerra, F.; Miranda, K.; Attias, M.; Oliveira, M.M. Euglena gracilis as a model for the study of Cu2+ and Zn2+ toxicity and accumulation in eukaryotic cells. Environ. Pollut. 2002, 120, 779–786. [Google Scholar] [CrossRef]
- Kitaoka, S.; Nakano, Y.; Miyatake, K.; Yokota, A. Enzymes and their functional location. In The Biology of Euglena; Buetow, D.E., Ed.; Academic Press Limited: London, UK, 1989; Volume 4, pp. 2–119. [Google Scholar]
- Wolken, J.J. Euglena: An Experimental Organism for Biochemical and Biophysical Studies; Institute of Microbiology, Rutgers: New Brunswick, NJ, USA, 1961; p. 173. [Google Scholar]
- Sato, S. The apicomplexan plastid and its evolution. Cell. Mol. Life Sci. 2011, 68, 1285–1296. [Google Scholar] [CrossRef] [PubMed]
- Ebenezer, T.E.; Zoltner, M.; Burrell, A.; Nenarokova, A.; Novák Vanclová, A.M.G.; Prasad, B.; Soukal, P.; Santana-Molina, C.; O’Neill, E.; Nankissoor, N.N.; et al. Transcriptome, proteome and draft genome of Euglena gracilis. BMC Biol. 2019, 17, 11. [Google Scholar] [CrossRef] [PubMed]
- Linton, E.W.; Karnkowska-Ishikawa, A.; Kim, J.I.; Shin, W.; Bennett, M.S.; Kwiatowski, J.; Zakryś, B.; Triemer, R.E. Reconstructing euglenoid evolutionary relationships using three genes: Nuclear SSU and LSU, and chloroplast SSU rDNA sequences and the description of Euglenaria gen. nov. (Euglenophyta). Protist 2010, 161, 603–619. [Google Scholar] [CrossRef] [PubMed]
- Ahmadinejad, N.; Dagan, T.; Martin, W. Genome history in the symbiotic hybrid Euglena gracilis. Gene 2007, 402, 35–39. [Google Scholar] [CrossRef] [PubMed]
- Parfrey, L.W.; Lahr, D.J.G.; Knoll, A.H.; Katz, L.A. Estimating the timing of early eukaryotic diversification with multigene molecular clocks. Proc. Natl. Acad. Sci. USA 2011, 108, 13624–13629. [Google Scholar] [CrossRef] [PubMed]
- Maruyama, S.; Suzaki, T.; Weber, A.P.; Archibald, J.M.; Nozaki, H. Eukaryote-to-eukaryote gene transfer gives rise to genome mosaicism in euglenids. BMC Evol. Biol. 2011, 11, 105. [Google Scholar] [CrossRef] [PubMed]
- Martin, W.; Somerville, C.C.; Loiseaux-de Goer, S. Molecular phylogenies of plastid origins and algal evolution. J. Mol. Evol. 1992, 35, 385–404. [Google Scholar] [CrossRef]
- O’Neill, E.C.; Trick, M.; Henrissat, B.; Field, R.A. Euglena in time: Evolution, control of central metabolic processes and multi-domain proteins in carbohydrate and natural product biochemistry. Perspect Sci. 2015, 6, 84–93. [Google Scholar] [CrossRef]
- Gissibl, A.; Sun, A.; Care, A.; Nevalainen, H.; Sunna, A. Bioproducts from Euglena gracilis: Synthesis and applications. Front. Bioeng. Biotechnol. 2019, 7, 108. [Google Scholar] [CrossRef]
- Mahapatra, D.M.; Chanakya, H.N.; Ramachandra, T.V. Euglena sp. as a suitable source of lipids for potential use as biofuel and sustainable wastewater treatment. J. Appl. Phycol. 2013, 25, 855–865. [Google Scholar] [CrossRef]
- Kruger, N.J.; Ratcliffe, R.G. Fluxes through plant metabolic networks: Measurements, predictions, insights and challenges. Biochem. J. 2015, 465, 27–38. [Google Scholar] [CrossRef] [PubMed]
- Jackson, C.; Knoll, A.H.; Chan, C.X.; Verbruggen, H. Plastid phylogenomics with broad taxon sampling further elucidates the distinct evolutionary origins and timing of secondary green plastids. Sci. Rep. 2018, 8, 1523. [Google Scholar] [CrossRef] [PubMed]
- O’Neill, E.C.; Trick, M.; Hill, L.; Rejzek, M.; Dusi, R.G.; Hamilton, C.J.; Zimba, P.V.; Henrissat, B.; Field, R.A. The transcriptome of Euglena gracilis reveals unexpected metabolic capabilities for carbohydrate and natural product biochemistry. Mol. Biosys. 2015, 11, 2808–2820. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, Y.; Tomiyama, T.; Maruta, T.; Tomita, M.; Ishikawa, T.; Arakawa, K. De novo assembly and comparative transcriptome analysis of Euglena gracilis in response to anaerobic conditions. BMC Genom. 2016, 17, 182. [Google Scholar] [CrossRef] [PubMed]
- O’Neill, E.C.; Saalbach, G.; Field, R.A. Chapter five - gene discovery for synthetic biology: Exploring the novel natural product biosynthetic capacity of eukaryotic microalgae. Methods Enzymol. 2016, 576, 99–120. [Google Scholar]
- Prigent, S.; Frioux, C.; Dittami, S.M.; Thiele, S.; Larhlimi, A.; Collet, G.; Gutknecht, F.; Got, J.; Eveillard, D.; Bourdon, J.; et al. Meneco, a topology-based gap-filling tool applicable to degraded genome-wide metabolic networks. PLoS Comput. Biol. 2017, 13, e1005276. [Google Scholar] [CrossRef] [PubMed]
- Hallick, R.B.; Hong, L.; Drager, R.G.; Favreau, M.R.; Monfort, A.; Orsat, B.; Spielmann, A.; Stutz, E. Complete sequence of Euglena gracilis chloroplast DNA. Nucleic Acids Res. 1993, 21, 3537–3544. [Google Scholar] [CrossRef] [PubMed]
- Turmel, M.; Gagnon, M.-C.; O’Kelly, C.J.; Otis, C.; Lemieux, C. The chloroplast genomes of the green algae Pyramimonas, Monomastix, and Pycnococcus shed new light on the evolutionary history of prasinophytes and the origin of the secondary chloroplasts of euglenids. Mol. Biol. Evol. 2009, 26, 631–648. [Google Scholar] [CrossRef]
- Martin, W.; Herrmann, R.G. Gene transfer from organelles to the nucleus: How much, what happens, and why? Plant Physiol. 1998, 118, 9–17. [Google Scholar] [CrossRef] [PubMed]
- Geimer, S.; Belicová, A.; Legen, J.; Sláviková, S.; Herrmann, R.G.; Krajčovič, J. Transcriptome analysis of the Euglena gracilis plastid chromosome. Curr. Genet. 2009, 55, 425–438. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.E.; Jurgenson, J.E.; Reardon, E.M.; Price, C.A. Plastid translation in organello and in vitro during light-induced development in Euglena. J. Biol. Chem. 1983, 258, 14478–14484. [Google Scholar] [PubMed]
- Bachvaroff, T.R.; Sanchez Puerta, M.V.; Delwiche, C.F. Chlorophyll c–containing plastid relationships based on analyses of a multigene data set with all four chromalveolate lineages. Mol. Biol. Evol. 2005, 22, 1772–1782. [Google Scholar] [CrossRef] [PubMed]
- Yoon, H.S.; Hackett, J.D.; Bhattacharya, D. A single origin of the peridinin- and fucoxanthin-containing plastids in dinoflagellates through tertiary endosymbiosis. Proc. Natl. Acad. Sci. USA 2002, 99, 11724. [Google Scholar] [CrossRef] [PubMed]
- Bolte, K.; Bullmann, L.; Hempel, F.; Bozarth, A.; Zauner, S.; Maier, U.-G. Protein targeting into secondary plastids. J. Eukaryot. Microbiol. 2009, 56, 9–15. [Google Scholar] [CrossRef] [PubMed]
- Horton, P.; Park, K.-J.; Obayashi, T.; Fujita, N.; Harada, H.; Adams-Collier, C.J.; Nakai, K. WoLF PSORT: protein localization predictor. Nucleic Acids Res. 2007, 35, W585–W587. [Google Scholar] [CrossRef]
- Emanuelsson, O.; Brunak, S.; von Heijne, G.; Nielsen, H. Locating proteins in the cell using TargetP, SignalP and related tools. Nat. Protoc. 2007, 2, 953–971. [Google Scholar] [CrossRef]
- Novák Vanclová, A.M.G.; Zoltner, M.; Kelly, S.; Soukal, P.; Záhonová, K.; Füssy, Z.; Ebenezer, T.E.; Lacová Dobáková, E.; Eliáš, M.; Lukeš, J.; et al. Proteome of the secondary plastid of Euglena gracilis reveals metabolic quirks and colourful history. bioRxiv 2019, 573709. [Google Scholar] [CrossRef]
- Durnford, D.G.; Gray, M.W. Analysis of Euglena gracilis plastid-targeted proteins reveals different classes of transit sequences. Eukaryot. Cell 2006, 5, 2079–2091. [Google Scholar] [CrossRef]
- Emanuelsson, O.; Nielsen, H.; Brunak, S.; von Heijne, G. Predicting subcellular localization of proteins based on their n-terminal amino acid sequence. J. Mol. Biol. 2000, 300, 1005–1016. [Google Scholar] [CrossRef]
- Gibbs, S.P. The chloroplasts of Euglena may have evolved from symbiotic green algae. Can. J. Bot. 1978, 56, 2883–2889. [Google Scholar] [CrossRef]
- Záhonová, K.; Füssy, Z.; Birčák, E.; Novák Vanclová, A.M.G.; Klimeš, V.; Vesteg, M.; Krajčovič, J.; Oborník, M.; Eliáš, M. Peculiar features of the plastids of the colourless alga Euglena longa and photosynthetic euglenophytes unveiled by transcriptome analyses. Sci. Rep. 2018, 8, 17012. [Google Scholar] [CrossRef] [PubMed]
- Sláviková, S.; Vacula, R.; Fang, Z.; Ehara, T.; Osafune, T.; Schwartzbach, S.D. Homologous and heterologous reconstitution of Golgi to chloroplast transport and protein import into the complex chloroplasts of Euglena. J. Cell Sci. 2005, 118, 1651–1661. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Maier, U.G.; Zauner, S.; Hempel, F. Protein import into complex plastids: Cellular organization of higher complexity. Eur. J. Cell Biol. 2015, 94, 340–348. [Google Scholar] [CrossRef] [PubMed]
- Smillie, R.M. Formation and function of soluble proteins in ghloroplasts. Can. J. Bot. 1963, 41, 123–154. [Google Scholar] [CrossRef]
- Bégin-Heick, N. The localization of enzymes of intermediary metabolism in Astasia and Euglena. Biochem. J. 1973, 134, 607. [Google Scholar] [CrossRef] [PubMed]
- Dockerty, A.; Merrett, M.J. Isolation and enzymic characterization of Euglena proplastids. Plant Physiol. 1979, 63, 468–473. [Google Scholar] [CrossRef]
- Graves, J.R.; Lynn, B. Effects of different substrates on glucose uptake and hexokinase activity in Euglena gracilis. J. Protozool. 1971, 18, 543–546. [Google Scholar] [CrossRef]
- Bäumer, D.; Preisfeld, A.; Ruppel, H.G. Isolation and characterization of paramylon synthase from Euglena gracilis (euglenophyceae). J. Phycol. 2001, 37, 38–46. [Google Scholar] [CrossRef]
- Lucchini, G. Control of glucose phosphorylation in Euglena gracilis I. Partial characterization of a glucokinase. Biochim. Biophys. Acta 1971, 242, 365–370. [Google Scholar] [CrossRef]
- Bégin-Heick, N. Oxygen toxicity and carbon deprivation in Astasia longa. Can. J. Biochem. 1970, 48, 251–258. [Google Scholar] [CrossRef] [PubMed]
- Miyatake, K.; Enomoto, T.; Kitaoka, S. Detection and subcellular distribution of pyrophosphate: D-fructose 6- phosphate phosphotransferase (PFP) in Euglena gracilis. Agric. Biol. Chem. 1984, 48, 2857–2859. [Google Scholar] [CrossRef][Green Version]
- Willard, J.M.; Gibbs, M. Purification and characterization of the fructose diphosphate aldolases from Anacystis is nidulans and Saprospira thermalis. Biochim. Biophys. Acta 1968, 151, 438–448. [Google Scholar] [CrossRef]
- Mo, Y.; Harris, B.G.; Gracy, R.W. Triosephosphate isomerases and aldolases from light- and dark-grown Euglena gracilis. Arch. Biochem. Biophys. 1973, 157, 580–587. [Google Scholar] [CrossRef]
- Latzko, E.; Gibbs, M. Enzyme activities of the carbon reduction cycle in some photosynthetic organisms. Plant Physiol. 1969, 44, 295. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Grissom, F.E.; Kahn, J.S. Glyceraldehyde-3-phosphate dehydrogenases from Euglena gracilis: Purification and physical and chemical characterization. Arch. Biochem. Biophys. 1975, 171, 444–458. [Google Scholar] [CrossRef]
- Bukowiecki, A.C.; Anderson, L.E. Multiple forms of aldolase and triose phosphate isomerase in diverse plant species. Plant Sci. Lett. 1974, 3, 381–386. [Google Scholar] [CrossRef]
- Forsee, W.T.; Kahn, J.S. Carbon dioxide fixation by isolated chloroplasts of Euglena gracilis: I. Isolation of functionally intact chloroplasts and their characterization. Arch. Biochem. Biophys. 1972, 150, 296–301. [Google Scholar] [CrossRef]
- Ammon, R.; Friedrich, G. The behavior of enzymes in Euglena gracilis. Acta Biol. Med. Ger. 1967, 19, 659–672. [Google Scholar]
- Hannaert, V.; Brinkmann, H.; Nowitzki, U.; Lee, J.A.; Albert, M.-A.; Sensen, C.W.; Gaasterland, T.; Miklós, M.; Michels, P.; Martin, W. Enolase from Trypanosoma brucei, from the amitochondriate protist Mastigamoeba balamuthi, and from the chloroplast and cytosol of Euglena gracilis: Pieces in the evolutionary puzzle of the eukaryotic glycolytic pathway. Mol. Biol. Evol. 2000, 17, 989–1000. [Google Scholar] [CrossRef]
- Ohmann, E. Die regulation der pyruvat-kinase in Euglena gracilis. Arch. Microbiol. 1969, 67, 273–292. [Google Scholar] [CrossRef]
- Ogawa, T.; Kimura, A.; Sakuyama, H.; Tamoi, M.; Ishikawa, T.; Shigeoka, S. Identification and characterization of cytosolic fructose-1,6-bisphosphatase in Euglena gracilis. Biosci. Biotechnol. Biochem. 2015, 79, 1957–1964. [Google Scholar] [CrossRef] [PubMed]
- Hovenkamp-Obbema, R. Effect of chloramphenicol on the development of proplastids in Euglena gracilis: II. The synthesis of carotenoids. Z. Pflanzenphysiol. 1974, 73, 439–447. [Google Scholar] [CrossRef]
- Kempner, E.S.; Miller, J.H. The molecular biology of Euglena gracilis: IV. Cellular stratification by centrifuging. Exp. Cell Res. 1968, 51, 141–149. [Google Scholar] [CrossRef]
- Ohmann, E.; Rindt, K.P.; Borriss, R. Glucose-6-phosphat-dehydrogenase in autotrophen mikroorganismen i. Die regulation der synthese der glucose-6-phosphat-dehydrogenase in Euglena gracilis und Rhodopseudomonas spheroides in abhängigkeit von den kulturbedingungen. Z. Allg. Mikrobiol. 1969, 9, 557–564. [Google Scholar] [CrossRef] [PubMed]
- Hovenkamp-Obbema, R.; Moorman, A.; Stegwee, D. Aminolaevulinate dehydratase in greening cells of Euglena gracilis. Z. Pflanzenphysiol. 1974, 72, 277–286. [Google Scholar] [CrossRef]
- Forsee, W.T.; Kahn, J.S. Carbon dioxide fixation by isolated chloroplasts of Euglena gracilis: II. Inhibition of CO2 fixation by AMP. Arch. Biochem. Biophys. 1972, 150, 302–309. [Google Scholar] [CrossRef]
- Ma, Y.A.N.; Jakowitsch, J.; Deusch, O.; Henze, K.; Martin, W.; Löffelhardt, W. Transketolase from Cyanophora paradoxa: In vitro import into cyanelles and pea chloroplasts and a complex history of a gene often, but not always, transferred in the context of secondary endosymbiosis. J. Eukaryot. Microbiol. 2009, 56, 568–576. [Google Scholar] [CrossRef]
- Stern, J.R.; Hegre, C.S. Inducible D-malic enzyme in Escherichia coli. Nature 1966, 212, 1611. [Google Scholar] [CrossRef]
- Peak, M.J.; Peak, J.G.; Ting, I.P. Isoenzymes of malate dehydrogenase and their regulation in Euglena gracilis Z. Biochim. Biophys. Acta 1972, 284, 1–15. [Google Scholar] [CrossRef]
- Peak, M.J.; Peak, J.G.; Ting, I.P. Light-induced reduction in specific activity of malate enzyme in Euglena gracilis Z. Biochem. Biophys. Res. Commun. 1972, 48, 1074–1078. [Google Scholar] [CrossRef]
- Codd, G.A.; Merrett, M.J. The regulation of glycolate metabolism in division synchronized cultures of Euglena. Plant Physiol. 1971, 47, 640–643. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Laval-Martin, D.; Farineau, J.; Pineau, B.; Calvayrac, R. Evolution of enzymes involved in carbon metabolism (phosphoenolpyruvate and ribulose-bisphosphate carboxylases, phosphoenolpyruvate carboxykinase) during the light-induced greening of Euglena gracilis strains Z and ZR. Planta 1981, 151, 157–167. [Google Scholar] [CrossRef] [PubMed]
- Briand, J.; Calvayrac, R.; Laval-Martin, D.; Farineau, J. Evolution of carboxylating enzymes involved in paramylon synthesis (phosphoenolpyruvate carboxylase and carboxykinase) in heterotrophically grown Euglena gracilis. Planta 1981, 151, 168–175. [Google Scholar] [CrossRef] [PubMed]
- Ohmann, E.; Plhák, F. Reinigung und eigenschaften von phosphoenolpyruvat-carboxylase aus Euglena gracilis. Eur. J. Biochem. 1969, 10, 43–55. [Google Scholar] [CrossRef] [PubMed]
- Miyatake, K.; Ito, T.; Kitaoka, S. Subcellular location and some properties of phosphoenolpyruvate carboxykinase (PEPCK) in Euglena gracilis. Agric. Biol. Chem. 1984, 48, 2139–2141. [Google Scholar] [CrossRef]
- Yokota, A.; Hosotani, K.; Kitaoka, S. Mechanism of metabolic regulation in photoassimilation of propionate in Euglena gracilis z. Arch. Biochem. Biophys. 1982, 213, 530–537. [Google Scholar] [CrossRef]
- Nakazawa, M.; Hayashi, R.; Takenaka, S.; Inui, H.; Ishikawa, T.; Ueda, M.; Sakamoto, T.; Nakano, Y.; Miyatake, K. Physiological functions of pyruvate:NADP+ oxidoreductase and 2-oxoglutarate decarboxylase in Euglena gracilis under aerobic and anaerobic conditions. Biosci. Biotechnol. Biochem. 2017, 81, 1386–1393. [Google Scholar] [CrossRef]
- Inui, H.; Ono, K.; Miyatake, K.; Nakano, Y.; Kitaoka, S. Purification and characterization of pyruvate:NADP+ oxidoreductase in Euglena gracilis. J. Biol. Chem. 1987, 262, 9130–9135. [Google Scholar]
- Rotte, C.; Stejskal, F.; Zhu, G.; Keithly, J.S.; Martin, W. Pyruvate:NADP oxidoreductase from the mitochondrion of Euglena gracilis and from the apicomplexan Cryptosporidium parvum: A biochemical relic linking pyruvate metabolism in mitochondriate and amitochondriate protists. Mol. Biol. Evol. 2001, 18, 710–720. [Google Scholar] [CrossRef][Green Version]
- Hoffmeister, M.; van der Klei, A.; Rotte, C.; van Grinsven, K.W.A.; van Hellemond, J.J.; Henze, K.; Tielens, A.G.M.; Martin, W. Euglena gracilis rhodoquinone:Ubiquinone ratio and mitochondrial proteome differ under aerobic and anaerobic conditions. J. Biol. Chem. 2004, 279, 22422–22429. [Google Scholar] [CrossRef] [PubMed]
- Collins, N.; Merrett, M.J. Microbody-marker enzymes during transition from phototrophic to organotrophic growth in Euglena. Plant Physiol. 1975, 55, 1018–1022. [Google Scholar] [CrossRef] [PubMed]
- Graves, L.B., Jr.; Trelease, R.N.; Grill, A.; Becker, W.M. Localization of glyoxylate cycle enzymes in glyoxysomes in Euglena. J. Protozool. 1972, 19, 527–532. [Google Scholar] [CrossRef]
- Cook, J.R.; Carver, M. Partial photo-repression of the glyoxylate by-pass in Euglena. Plant Cell Physiol. 1966, 7, 377–383. [Google Scholar]
- Cook, J.R.; Heinrich, B. Unbalanced respiratory growth of Euglena. Microbiology 1968, 53, 237–251. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Collins, N.; Merrett, M.J. The localization of glycollate-pathway enzymes in Euglena. Biochem. J. 1975, 148, 321–328. [Google Scholar] [CrossRef][Green Version]
- Oda, Y.; Miyatake, K.; Nakano, Y.; Kitaoka, S. Subcellular location and some properties of isocitrate dehydrogenase isozymes in Euglena gracilis. Agric. Biol. Chem. 1981, 45, 2619–2621. [Google Scholar] [CrossRef][Green Version]
- Shigeoka, S.; Onishi, T.; Maeda, K.; Nakano, Y.; Kitaoka, S. Occurrence of thiamin pyrophosphate-dependent 2-oxoglutarate decarboxylase in mitochondria of Euglena gracilis. FEBS Lett. 1986, 195, 43–47. [Google Scholar] [CrossRef]
- Shigeoka, S.; Nakano, Y. Characterization and molecular properties of 2-oxoglutarate decarboxylase from Euglena gracilis. Arch. Biochem. Biophys. 1991, 288, 22–28. [Google Scholar] [CrossRef]
- Tokunaga, M.; Nakano, Y.; Kitaoka, S. Separation and properties of the NAD-linked and NADP-linked isozymes of succinic semialdehyde dehydrogenase in Euglena gracilis. Biochim. Biophys. Acta 1976, 429, 55–62. [Google Scholar] [CrossRef]
- Barry, D.; Merrett, M.J. Malate dehydrogenase isoenzymes in division synchronized cultures of Euglena. Plant Physiol. 1973, 51, 1127–1132. [Google Scholar]
- Isegawa, Y.; Nakano, Y.; Kitaoka, S. Submitochondrial location and some properties of NAD+- and NADP+-linked malate dehydrogenase in Euglena. Agric. Biol. Chem. 1984, 48, 549–552. [Google Scholar] [CrossRef]
- Miyatake, K.; Washio, K.; Yokota, A.; Nakano, Y.; Kitaoka, S. Occurrence of two NAD+-linked malate dehydrogenase isozymes in Euglena gracilis. Agric. Biol. Chem. 1985, 49, 859–860. [Google Scholar] [CrossRef]
- Nakazawa, M. C2 metabolism in Euglena. Adv. Exp. Med. Biol. 2017, 979, 39–45. [Google Scholar] [PubMed]
- Graves, L.B.; Hanzely, L.; Trelease, R.N. The occurrence and fine structural characterization of microbodies in Euglena gracilis. Protoplasma 1971, 72, 141–152. [Google Scholar] [CrossRef] [PubMed]
- Ono, K.; Kondo, M.; Osafune, T.; Miyatake, K.; Inui, H.; Kitaoka, S.; Nishimura, M.; Nakano, Y. Presence of glyoxylate cycle enzymes in the mitochondria of Euglena gracilis. J. Eukaryot. Microbiol. 2003, 50, 92–96. [Google Scholar] [CrossRef]
- RodrÍGuez-Zavala, J.S.; Ortiz-Cruz, M.A.; Moreno-Sanchez, R. Characterization of an aldehyde dehydrogenase from Euglena gracilis. J. Eukaryot. Microbiol. 2006, 53, 36–42. [Google Scholar] [CrossRef]
- Yoval-Sánchez, B.; Jasso-Chávez, R.; Lira-Silva, E.; Moreno-Sánchez, R.; Rodríguez-Zavala, J.S. Novel mitochondrial alcohol metabolizing enzymes of Euglena gracilis. J. Bioenerg. Biomembr. 2011, 43, 519–530. [Google Scholar] [CrossRef]
- Ono, K.; Kawanaka, Y.; Izumi, Y.; Inui, H.; Miyatake, K.; Kitaoka, S.; Nakano, Y. Mitochondrial alcohol dehydrogenase from ethanol-grown Euglena gracilis. J. Biochem. 1995, 117, 1178–1182. [Google Scholar] [CrossRef]
- Graves, L.B., Jr.; Becker, W.M. Beta-oxidation in glyoxysomes from Euglena. J. Protozool. 1974, 21, 771–774. [Google Scholar] [CrossRef]
- Tucci, S.; Vacula, R.; Krajcovic, J.; Proksch, P.; Martin, W. Variability of wax ester fermentation in natural and bleached Euglena gracilis strains in response to oxygen and the elongase inhibitor flufenacet. J. Eukaryot. Microbiol. 2010, 57, 63–69. [Google Scholar] [CrossRef] [PubMed]
- Grimm, P.; Risse, J.M.; Cholewa, D.; Müller, J.M.; Beshay, U.; Friehs, K.; Flaschel, E. Applicability of Euglena gracilis for biorefineries demonstrated by the production of α-tocopherol and paramylon followed by anaerobic digestion. J. Biotechnol. 2015, 215, 72–79. [Google Scholar] [CrossRef] [PubMed]
- O’Neill, E.C.; Kuhaudomlarp, S.; Rejzek, M.; Fangel, J.U.; Alagesan, K.; Kolarich, D.; Willats, W.G.T.; Field, R.A. Exploring the glycans of Euglena gracilis. Biology 2017, 6, 45. [Google Scholar] [CrossRef] [PubMed]
- Marechal, L.R.; Goldemberg, S.H. Laminaribiose phosphorylase from Euglena gracilis. Biochem. Biophys. Res. Commun. 1963, 13, 106–107. [Google Scholar] [CrossRef]
- Tomos, A.D.; Northcote, D.H. A protein-glucan intermediate during paramylon synthesis. Biochem. J. 1978, 174, 283–290. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, Y.; Ogawa, T.; Maruta, T.; Yoshida, Y.; Arakawa, K.; Ishikawa, T. Glucan synthase-like 2 is indispensable for paramylon synthesis in Euglena gracilis. FEBS Lett. 2017, 591, 1360–1370. [Google Scholar] [CrossRef] [PubMed]
- Calvayrac, R.; Laval-Martin, D.; Briand, J.; Farineau, J. Paramylon synthesis by Euglena gracilis photoheterotrophically grown under low O2 pressure. Planta 1981, 153, 6–13. [Google Scholar] [CrossRef]
- Briand, J.; Calvayrac, R. Paramylon synthesis in heterotrophic and photoheterotrophic Euglena (Euglenophyceae). J. Phycol. 1980, 16, 234–239. [Google Scholar] [CrossRef]
- Takeda, T.; Nakano, Y.; Takahashi, M.; Konno, N.; Sakamoto, Y.; Arashida, R.; Marukawa, Y.; Yoshida, E.; Ishikawa, T.; Suzuki, K. Identification and enzymatic characterization of an endo-1,3-β-glucanase from Euglena gracilis. Phytochemistry 2015, 116, 21–27. [Google Scholar] [CrossRef]
- Barras, D.R.; Stone, B.A. β-1,3-glucan hydrolases from Euglena gracilis: II. Purification and properties of the β-1,3-glucan exo-hydrolase. Biochim. Biophys. Acta 1969, 191, 342–353. [Google Scholar] [CrossRef]
- Kuhaudomlarp, S.; Patron, N.J.; Henrissat, B.; Rejzek, M.; Saalbach, G.; Field, R.A. Identification of Euglena gracilis β-1,3-glucan phosphorylase and establishment of a new glycoside hydrolase (GH) family GH149. J. Biol. Chem. 2018, 293, 2865–2876. [Google Scholar] [CrossRef] [PubMed]
- Takenaka, S.; Kondo, T.; Nazeri, S.; Tamura, Y.; Tokunaga, M.; Tsuyama, S.; Miyatake, K.; Nakano, Y. Accumulation of trehalose as a compatible solute under osmotic stress in Euglena gracilis Z. J. Eukaryot. Microbiol. 1997, 44, 609–613. [Google Scholar] [CrossRef]
- Porchia, A.C.; Fiol, D.F.; Salerno, G.L. Differential synthesis of sucrose and trehalose in Euglena gracilis cells during growth and salt stress. Plant Sci. 1999, 149, 43–49. [Google Scholar] [CrossRef]
- Fiol, D.F.; Salerno, G.L. Trehalose synthesis in Euglena gracilis (euglenophyceae) occurs through an enzyme complex. J. Phycol. 2005, 41, 812–818. [Google Scholar] [CrossRef]
- Schluepmann, H.; Paul, M. Trehalose metabolites in Arabidopsis-elusive, active and central. Arabidopsis Book 2009, 7, e0122. [Google Scholar] [CrossRef]
- Vandesteene, L.; Ramon, M.; Le Roy, K.; Van Dijck, P.; Rolland, F. A single active trehalose-6-p synthase (TPS) and a family of putative regulatory TPS-like proteins in Arabidopsis. Mol. Plant 2010, 3, 406–419. [Google Scholar] [CrossRef] [PubMed]
- Murray, D.R.; Giovanelli, J.; Smillie, R.M. Photoassimilation of glycolate, glycine and serine by Euglena gracilis. J. Protozool. 1970, 17, 99–104. [Google Scholar] [CrossRef]
- Yokota, A.; Komura, H.; Kitaoka, S. Different metabolic fate of two carbons of glycolate in Euglena gracilis Z. In Advances in Photosynthesis Research: Proceedings of the VIth International Congress on Photosynthesis, Brussels, Belgium, August 1–6, 1983; Sybesma, C., Ed.; Springer Netherlands: Dordrecht, The Netherlands, 1984; pp. 407–410. [Google Scholar]
- Yokota, A.; Kawabata, A.; Kitaoka, S. Mechanism of glyoxylate decarboxylation in the glycolate pathway in Euglena gracilis Z. Plant Physiol. 1983, 71, 772–776. [Google Scholar] [CrossRef]
- Yokota, A.; Haga, S.; Kitaoka, S. Purification and some properties of glyoxylate reductase (NADP+) and its functional location in mitochondria in Euglena gracilis Z. Biochem. J. 1985, 227, 211–216. [Google Scholar] [CrossRef]
- Hasan, M.T.; Sun, A.; Mirzaei, M.; Te’o, J.; Hobba, G.; Sunna, A.; Nevalainen, H. A comprehensive assessment of the biosynthetic pathways of ascorbate, α-tocopherol and free amino acids in Euglena gracilis var. saccharophila. Algal Res. 2017, 27, 140–151. [Google Scholar] [CrossRef]
- Isegawa, Y.; Watanabe, F.; Kitaoka, S.; Nakano, Y. Subcellular distribution of cobalamin-dependent methionine synthase in Euglena gracilis Z. Phytochemistry 1993, 35, 59–61. [Google Scholar] [CrossRef]
- Sulzman, F.M.; Edmunds, L.N. Persisting circadian oscillations in enzyme activity in non-dividing cultures of Euglena. Biochem. Biophys. Res. Commun. 1972, 47, 1338–1344. [Google Scholar] [CrossRef]
- Edmunds, L.N.; Walther, W.G.; Sulzman, F.M. At cellular level. J. Interdiscip. Cycle Res. 1972, 3, 107–108. [Google Scholar] [CrossRef]
- Reinbothe, C.; Ortel, B.; Parthier, B.; Reinbothe, S. Cytosolic and plastid forms of 5-enolpyruvylshikimate-3-phosphate synthase in Euglena gracilis are differentially expressed during light-induced chloroplast development. Mol. Gen. Genet. 1994, 245, 616–622. [Google Scholar] [CrossRef] [PubMed]
- Richards, T.A.; Dacks, J.B.; Campbell, S.A.; Blanchard, J.L.; Foster, P.G.; McLeod, R.; Roberts, C.W. Evolutionary origins of the Eukaryotic shikimate pathway: gene fusions, horizontal gene transfer, and endosymbiotic replacements. Eukaryot. Cell 2006, 5, 1517–1531. [Google Scholar] [CrossRef] [PubMed]
- Oda, Y.; Nakano, Y.; Kitaoka, S. Utilization and toxicity of exogenous amino acids in Euglena gracilis. Microbiology 1982, 128, 853–858. [Google Scholar] [CrossRef]
- Oda, Y.; Nakano, Y.; Kitaoka, S. Occurrence and some properties of two threonine dehydratases in Euglena gracilis. Microbiology 1983, 129, 57–61. [Google Scholar] [CrossRef][Green Version]
- Park, B.-S.; Hirotani, A.; Nakano, Y.; Kitaoka, S. Purification and someproperties of arginine deiminase in Euglena gracilis Z. Agric. Biol. Chem. 1984, 48, 483–489. [Google Scholar]
- Koonin, E.V.; Galperin, M.Y. Evolution of central metabolic pathways: The playground of non-orthologous gene displacement. In Sequence—Evolution—Function: Computational Approaches in Comparative Genomics; Koonin, E.V., Galperin, M.Y., Eds.; Springer: Boston, MA, USA, 2003; pp. 295–355. [Google Scholar]
- Torruella, G.; Suga, H.; Riutort, M.; Pereto, J.; Ruiz-Trillo, I. The evolutionary history of lysine biosynthesis pathways within Eukaryotes. J. Mol. Evol. 2009, 69, 240–248. [Google Scholar] [CrossRef]
- Xu, H.; Andi, B.; Qian, J.; West, A.H.; Cook, P.F. The α-aminoadipate pathway for lysine biosynthesis in fungi. Cell Biochem. Biophys. 2006, 46, 43–64. [Google Scholar] [CrossRef]
- Creaser, E.H.; Varela-Torres, R. Immunological comparisons of histidinol dehydrogenases. Microbiology 1971, 67, 85–90. [Google Scholar] [CrossRef] [PubMed]
- Tokunaga, M.; Nakano, Y.; Kitaoka, S. Subcellular localization of the GABA-shunt enzymes in Euglena gracilis strain Z. J. Protozool. 1979, 26, 471–473. [Google Scholar] [CrossRef] [PubMed]
- Miyatake, K.; Kitaoka, S. NADH-dependent glutamat synthase in Euglena gracilis Z. Agric. Biol. Chem. 1981, 45, 1727–1729. [Google Scholar]
- Fayyaz-Chaudhary, M.; Javed, Q.; Merrett, M.J. Effect of growth conditions on NADPH-specific glutamate dehydrogenase activity of Euglena gracilis. New Phytol. 1985, 101, 367–376. [Google Scholar] [CrossRef]
- Shemin, D.; Rittenberg, D. The utilization of glycine for the synthesis of a porphyrin. J. Biol. Chem. 1945, 159, 567–568. [Google Scholar]
- Beale, S.I.; Castelfranco, P.A. Biosynthesis of delta-aminolevulinic-acid in higher-plants.1. Accumulation of delta-aminolevulinic-acid in greening plant-tissues. Plant Phys. 1974, 53, 291–296. [Google Scholar] [CrossRef] [PubMed]
- Iida, K.; Mimura, I.; Kajiwara, M. Evaluation of two biosynthetic pathways to δ-aminolevulinic acid in Euglena gracilis. Eur. J. Biochem. 2002, 269, 291–297. [Google Scholar] [CrossRef] [PubMed]
- Weinstein, J.D.; Beale, S.I. Separate physiological roles and subcellular compartments for two tetrapyrrole biosynthetic pathways in Euglena gracilis. J. Biol. Chem. 1983, 258, 6799–6807. [Google Scholar]
- Wolpert, J.S.; Ernst-Fonberg, M.L. Multienzyme complex for carbon dioxide fixation. Biochemistry 1975, 14, 1095–1102. [Google Scholar] [CrossRef]
- Inui, H.; Miyatake, K.; Nakano, Y.; Kitaoka, S. Wax ester fermentation in Euglena gracilis. FEBS Lett. 1982, 150, 89–93. [Google Scholar] [CrossRef]
- Kim, D.; Filtz, M.R.; Proteau, P.J. The methylerythritol phosphate pathway contributes to carotenoid but not phytol biosynthesis in Euglena gracilis. J. Nat. Prod. 2004, 67, 1067–1069. [Google Scholar] [CrossRef] [PubMed]
- Ferro, M.; Salvi, D.; Riviere-Rolland, H.; Vermat, T.; Seigneurin-Berny, D.; Grunwald, D.; Garin, J.; Joyard, J.; Rolland, N. Integral membrane proteins of the chloroplast envelope: identification and subcellular localization of new transporters. Proc. Natl. Acad. Sci. USA 2002, 99, 11487–11492. [Google Scholar] [CrossRef] [PubMed]
- O’Neill, E. An Exploration of Phosphorylases for the Synthesis of Carbohydrate Polymers; University of East Anglia: Norwich, UK, 2013. [Google Scholar]

| Marker Enzyme Name | EC Number | EG Transcript | WoLF PSORT Plant | WoLF PSORT Animal | WoLF PSORT Fungi | TargetP Plant | TargetP Nonplant | TargetP, Signal Cut | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Fumarase | 4.2.1.2 | 4764 | Chl: 8 | - | Mt: 27.5 | Cyt_Mt: 17 | - | Mt: 26 | - | - | Mt: 1 | Mt: 2 | - | |
| Succinic semialdehyde dehydrogenase | 1.2.1.16 | 7552 | Cyt: 11 | - | Cyt: 24 | - | - | Cyt: 22 | - | - | Cyt: 2 | Cyt: 1 | - | |
| 5575 | Mt: 10.5 | Chl_Mt: 7.5 | Mt: 28.5 | Cyt_Mt: 16 | - | Mt: 22.5 | Cyt_Mt: 14 | - | Mt: 1 | Mt | - | |||
| 3780 | Chl: 9 | - | Mt: 10.5 | Cyt_Mt: 10.3 | Cyt: 9 | Mt: 16 | Cyt: 7.5 | - | Mt: 5 | Mt: 4 | - | |||
| 5750 | Mt: 12.5 | Chl_Mt: 7.5 | Mt: 27.5 | Cyt_Mt: 16.5 | - | Mt: 25.5 | Cyt_Mt: 14 | - | Mt: 3 | Mt: 2 | - | |||
| Isocitrate dehydrogenase | 1.1.1.41, 1.1.1.42 | 6875 | Cyt: 7 | - | Cyt: 17 | Mt: 15 | - | Cyt: 22.5 | Cyt_Nu: 12 | - | Cyt: 3 | Cyt: 2 | - | |
| 10115 | Chl_Mt: 7.5 | Mt: 7 | Chl: 6 | Mt: 29 | Cyt_Mt: 16.5 | - | Mt: 22 | - | - | Mt: 1 | Mt: 1 | - | ||
| 10872 | Mt: 10 | - | Mt: 28 | - | - | Mt: 26.5 | Cyt_Mt: 14 | - | Mt: 3 | Mt: 5 | - | |||
| 10979 | - | - | Mt: 24 | Cyt_Mt: 15.3 | Mt_Per: 12.8 | Per: 15 | Cyt: 7.5 | - | Mt: 2 | Mt: 2 | - | |||
| NADPH:glyoxylate reductase | 1.1.1.79 | 5154 | Mt: 14 | - | Mt: 24 | Cyt: 6 | - | Mt: 14.5 | CytMt:14 | Cyt: 12.5 | Mt: 3 | Mt: 5 | - | |
| 13699 | Cyt: 7 | - | Mt: 27 | - | - | Cyt: 12.5 | Mt: 10 | Cyt_Nu: 7 | Cyt: 5 | Sec: 2 | - | |||
| Marker Enzyme Name | EC Number | EG Transcript | WoLF PSORT Plant | WoLF PSORT Animal | WoLF PSORT Fungi | TargetP Plant | TargetP Nonplant | TargetP, Signal Cut | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Glutamate dehydrogenase | 1.4.1.2 | 11196 | Cyt: 8 | Chl: 6 | Cyt_Nu: 15 | Nu: 8.5 | - | Cyt: 16 | Cyt_Nu: 9.5 | Cysk:7 | Cyt: 2 | Cyt: 2 | - |
| 10682 | Cyt: 8 | - | Cyt: 21 | Cyt_Nu: 16 | Nu: 9 | Cyt: 12.5 | Mt: 8 | Cyt_Nu: 8 | Cyt: 4 | Cyt: 3 | - | ||
| 7536 | Cyt: 11 | - | Cyt: 16.5 | Cyt_Mt: 13.6 | Cyt_Nu: 10.3 | Mt: 15 | Cyt: 7.5 | - | Mt: 5 | Cyt: 4 | - | ||
| 791 | Cyt: 6 | - | Cyt: 12 | Nu: 10 | - | Cyt: 11.5 | Cyt_Nu: 7.5 | Mt: 7 | Cyt: 4 | Cyt: 5 | - | ||
| NAD-lactate dehydrogenase | 1.1.1.27 | 11571 | - | - | Mt: 10 | Cyt: 7 | Cyt_Nu: 7 | Mt: 19 | Cyt: 8 | - | Cyt: 3 | Cyt: 5 | - |
| thiosulfate sulfurtransferase | 2.8.1.1 | 32721 | Cyt: 8.5 | - | Cyt: 17 | Cyt_Nu: 14 | Sec: 7 | Cyt: 9 | Cysk: 9 | Mt: 7 | Cyt | Cyt: 3 | - |
| 19233 | Chl: 13 | - | Sec: 14 | E.R.: 6 | - | Sec: 17 | - | - | Mt: 5 | Sec: 4 | - | ||
| 10931 | - | - | PM: 15 | Mt: 7 | - | Cyt: 8.5 | Mt: 7 | Sec: 7 | Cyt: 5 | Cyt: 3 | - | ||
| 27480 | - | - | Cyt_Nu: 15.5 | Nu: 13.5 | Cyt: 12.5 | Cyt: 10.5 | Cysk: 9 | Cyt_Nu: 8 | Mt: 5 | Mt: 5 | - | ||
| 18090 | Cyt: 6 | - | Sec: 28 | - | - | Sec: 18 | - | - | Cyt: 3 | Cyt: 2 | - | ||
| 46995 | Chl: 9 | - | Cyt: 18 | Cyt_Nu: 14.5 | Nu: 7 | Mt: 15.5 | Cyt_Mt: 10.3 | - | Cyt: 4 | Cyt: 5 | - | ||
| 1438 | Chl: 6 | - | Sec: 13 | E.R.: 11 | Mt: 6 | Sec: 9 | Mt: 7 | - | Mt: 5 | Sec: 4 | - | ||
| α-1,3-Man transferase | 2.4.1.258 | 11655 | PM: 6 | - | PM: 19 | - | - | Mt: 13 | PM: 9 | - | Sec: 4 | Sec: 1 | Cyt: 1 |
| α-1,2-Man transferase | 2.4.1.259/261 | 44449 | Chl: 12 | - | Sec: 27 | - | - | Mt: 10 | PM: 10 | - | Sec: 5 | Sec: 1 | Cyt: 1 |
| 15548 | PM: 12 | - | PM: 15.5 | Sec_PM: 10.5 | - | PM: 17 | - | - | Mt: 2 | Sec: 5 | - | ||
| α-1,6-Man transferase | 2.4.1.260 | 9095 | PM: 9 | - | PM: 13 | Sec: 11 | - | PM: 19 | - | - | Sec: 4 | Sec: 1 | Cyt: 1 |
| Dolichol phosphate mannose synthase | 2.4.1.83 | 21418 | - | - | PM: 11 | Cyt: 6 | Mt: 6 | Cyt: 8 | Mt: 7 | PM: 6 | Cyt: 1 | Cyt: 1 | - |
| Dolichyl-phosphate beta-glucosyltransferase | 2.4.1.117 | 15275 | PM: 7 | - | Sec: 20 | PM: 9 | - | PM: 12 | Sec: 12 | - | Sec: 5 | Sec: 2 | Cyt: 4 |
| α-1,3-Glc transferase | 2.4.1.267/265 | 22866 | - | - | Sec: 11 | PM: 9 | Mt: 7 | PM: 21 | - | - | Cyt: 5 | Sec: 5 | - |
| 24290 | PM: 7 | - | Sec: 16 | PM: 11 | - | PM: 18 | - | - | Mt: 3 | Sec: 1 | - | ||
| 22282 | PM: 6 | - | Sec: 9 | PM: 7 | Mt: 7 | PM: 17 | E.R.: 7 | Sec: 2 | Sec: 2 | Cyt: 2 | |||
| α-1,2-Glc transferase | 2.4.1.256 | 20467 | - | - | Sec: 15 | PM: 10 | - | PM: 17 | - | - | Mt: 1 | Mt: 3 | - |
| Dolichyldiphosphooligosaccharide-protein glycosyltransferase | 2.4.99.18 | 2741 | PM: 13 | - | PM: 22 | - | - | PM: 17 | E.R.: 6 | - | Mt: 5 | Sec: 2 | - |
| 10761 | PM: 11 | - | PM: 11 | Cyt: 9 | PM: 26 | - | - | Cyt: 2 | Cyt: 2 | - | |||
| 2733 | PM: 11 | - | PM: 24 | - | - | PM: 23 | - | - | Cyt: 5 | Cyt: 4 | - | ||
| Marker Enzyme Name | EC Number | EG Transcript | WoLF PSORT Plant | WoLF PSORT Animal | WoLF PSORT Fungi | TargetP Plant | TargetP Nonplant | TargetP, Signal Cut | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Ribulose-biphosphate carboxylase (small subunit) | 4.1.1.39 | 7123 | Chl: 13 | - | Cyt: 15.5 | Cyt_Nu: 14.5 | Nu: 8.5 | Nu: 7.5 | Cyt_Nu: 7.5 | Mt: 7 | Cyt: 2 | Cyt: 1 | - |
| 43244 | Chl: 8 | - | Sec: 30 | - | - | PM: 21 | - | - | Sec: 3 | Sec: 2 | Chl: 5 | ||
| 13866 | - | - | Mt: 12.5 | Cyt_Nu: 11 | Cyt: 10.5 | Cyt_Nu: 9.3 | Cyt_Mt: 9.3 | Cyt: 9 | Cyt: 4 | Cyt: 3 | - | ||
| 59484 | Chl: 7 | - | Sec: 29 | - | - | Nu: 10.5 | Cyt_Nu: 10 | Cyt: 8.5 | Sec: 3 | Sec: 5 | Cyt: 4 | ||
| NADP- glyceraldehyde-3-phosphate dehydrogenase | 1.2.1.9 | 4454 | Chl: 11 | - | Mt: 19.5 | Mt_Per: 11 | Cyt: 6.5 | Mt: 18 | Cyt: 6 | - | Mt: 4 | Mt: 4 | - |
| 5575 | Mt: 10.5 | Chl_Mt: 7.5 | Mt: 28.5 | Cyt_Mt: 16 | - | Mt: 22.5 | Cyt_Mt: 14 | - | Mt: 1 | Mt: 2 | - | ||
| 9413 | Chl: 12 | - | Mt: 22.5 | Mt_Per: 12 | - | Mt: 23 | - | - | Mt: 5 | Mt: 2 | - | ||
| 7552 | Cyt: 11 | - | Cyt: 24 | - | - | Cyt: 22 | - | - | Cyt: 2 | Cyt: 1 | - | ||
| 15045 | Mt: 8.5 | Chl_Mt: 7 | Mt: 24 | - | - | Mt: 26.5 | Cyt_Mt: 14 | - | Mt: 2 | Mt: 2 | - | ||
| Fructose-biphosphate aldolase | 4.1.2.13 | 15855 | Cyt: 8 | - | Nu: 12.5 | Cyt_Nu: 10.5 | Cyt: 7.5 | Cyt: 16 | Mt: 6 | - | Cyt: 4 | Cyt: 2 | - |
| 25832 | Mt: 8.5 | Chl_Mt: 7 | Mt: 16 | Cyt: 14 | - | Mt: 24.5 | Cyt_Mt: 14 | - | Mt: 2 | Mt: 4 | - | ||
| 7827 | Chl: 12 | - | E.R.: 16 | - | - | Sec: 15 | - | - | Chl: 5 | Sec: 4 | - | ||
| 5279 | Chl: 14 | - | Mt: 17 | E.R._Mt: 10.5 | Per: 6 | Mt: 15 | Cyt: 6 | - | Mt: 4 | Mt: 3 | - | ||
| Pyruvate Pi dikinase | 2.7.9.1 | 1635 | Chl: 6 | - | PM: 9 | Sec: 8 | - | Cyt: 11.5 | Mt: 9 | Cyt_Nu: 7.5 | Cyt: 1 | Cyt: 3 | - |
| Photosystem II D1 | - | 13288 | PM: 6 | - | PM: 12 | Lyso: 10 | Sec: 6 | PM: 21 | - | - | Sec: 1 | Sec: 1 | Cyt: 4 |
| - | 10715 | PM: 13 | - | PM: 26.5 | Sec_PM: 14 | - | PM: 27 | - | - | Cyt: 2 | Cyt: 4 | - | |
| Photosystem II D2 | - | 13288 | PM: 6 | - | PM: 12 | Lyso: 10 | Sec: 6 | PM: 21 | - | - | Sec: 1 | Sec: 1 | Cyt: 4 |
| - | 10715 | PM: 13 | - | PM: 26.5 | Sec_PM: 14 | - | PM: 27 | - | - | Cyt: 2 | Cyt: 4 | - | |
| Photosystem II CP47 | - | 899 | PM: 9.5 | - | PM: 31 | - | - | PM: 24 | - | - | Mt: 5 | Sec: 5 | Cyt: 2 |
| Photosystem II CP43 | - | 13640 | PM: 10 | - | PM: 20 | - | - | PM: 22 | - | - | Sec: 4 | Sec: 1 | Cyt: 4 |
| Cytochrome b6f | - | 5228 | PM: 6.5 | - | PM: 12 | Mt: 6 | - | PM: 13 | E.R.: 7 | - | Cyt: 4 | Cyt: 4 | - |
| - | 23844 | Chl: 13 | - | Mt: 16.5 | E.R._Mt: 13 | Mt_Per: 12.5 | Mt: 14 | - | - | Cyt: 5 | Mt: 5 | - | |
| Photosystem I A1 | - | 158 | PM: 10 | - | PM: 23 | - | - | PM: 25 | - | - | Mt: 5 | Mt: 4 | - |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Inwongwan, S.; Kruger, N.J.; Ratcliffe, R.G.; O’Neill, E.C. Euglena Central Metabolic Pathways and Their Subcellular Locations. Metabolites 2019, 9, 115. https://doi.org/10.3390/metabo9060115
Inwongwan S, Kruger NJ, Ratcliffe RG, O’Neill EC. Euglena Central Metabolic Pathways and Their Subcellular Locations. Metabolites. 2019; 9(6):115. https://doi.org/10.3390/metabo9060115
Chicago/Turabian StyleInwongwan, Sahutchai, Nicholas J. Kruger, R. George Ratcliffe, and Ellis C. O’Neill. 2019. "Euglena Central Metabolic Pathways and Their Subcellular Locations" Metabolites 9, no. 6: 115. https://doi.org/10.3390/metabo9060115
APA StyleInwongwan, S., Kruger, N. J., Ratcliffe, R. G., & O’Neill, E. C. (2019). Euglena Central Metabolic Pathways and Their Subcellular Locations. Metabolites, 9(6), 115. https://doi.org/10.3390/metabo9060115






