Postprandial Metabolic Effects of Fiber Mixes Revealed by in vivo Stable Isotope Labeling in Humans
Abstract
1. Introduction
2. Results and Discussion
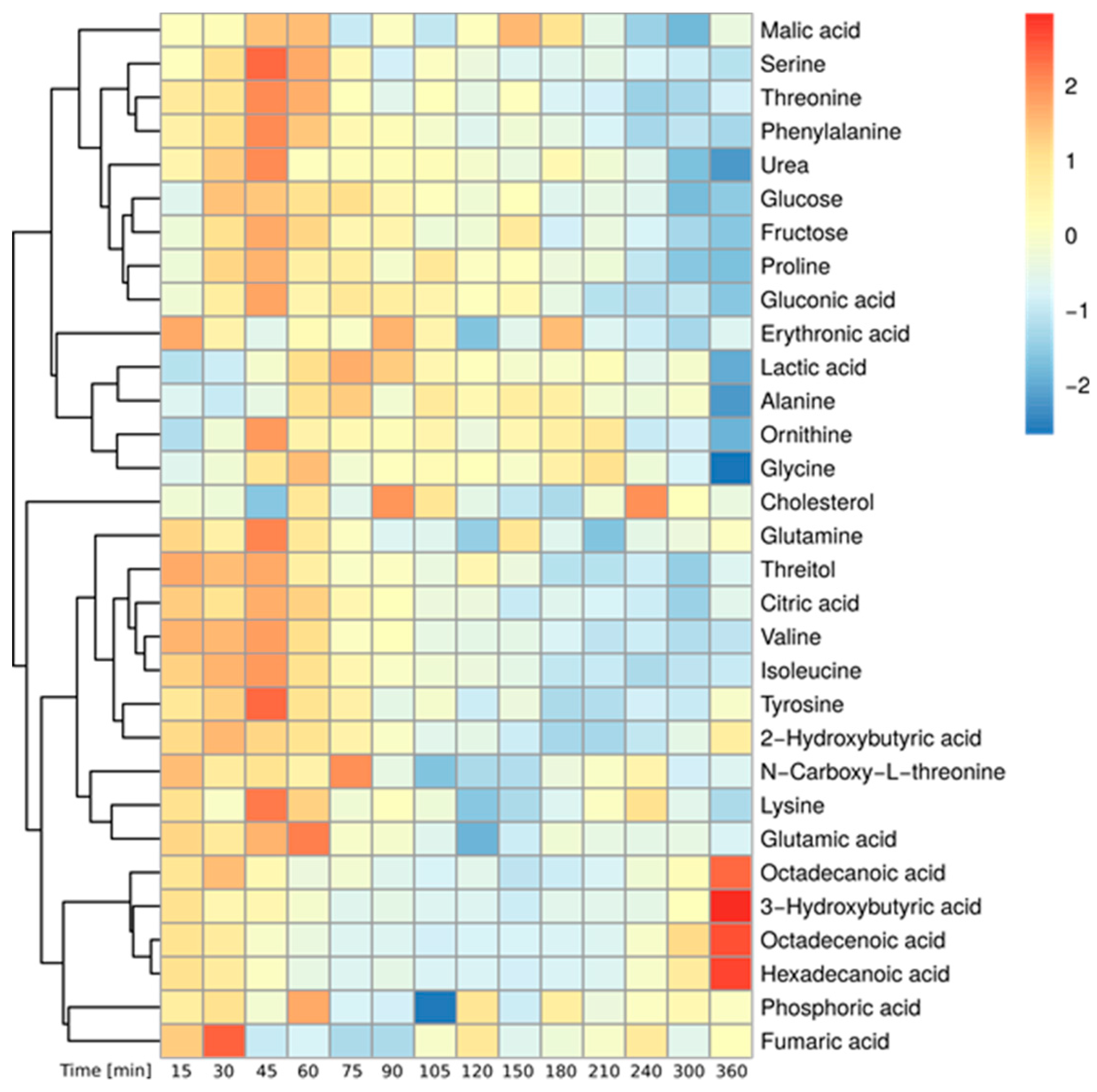
2.1. Postprandial Effect of Sharbati Flour Flatbreads on the Plasma Metabolome
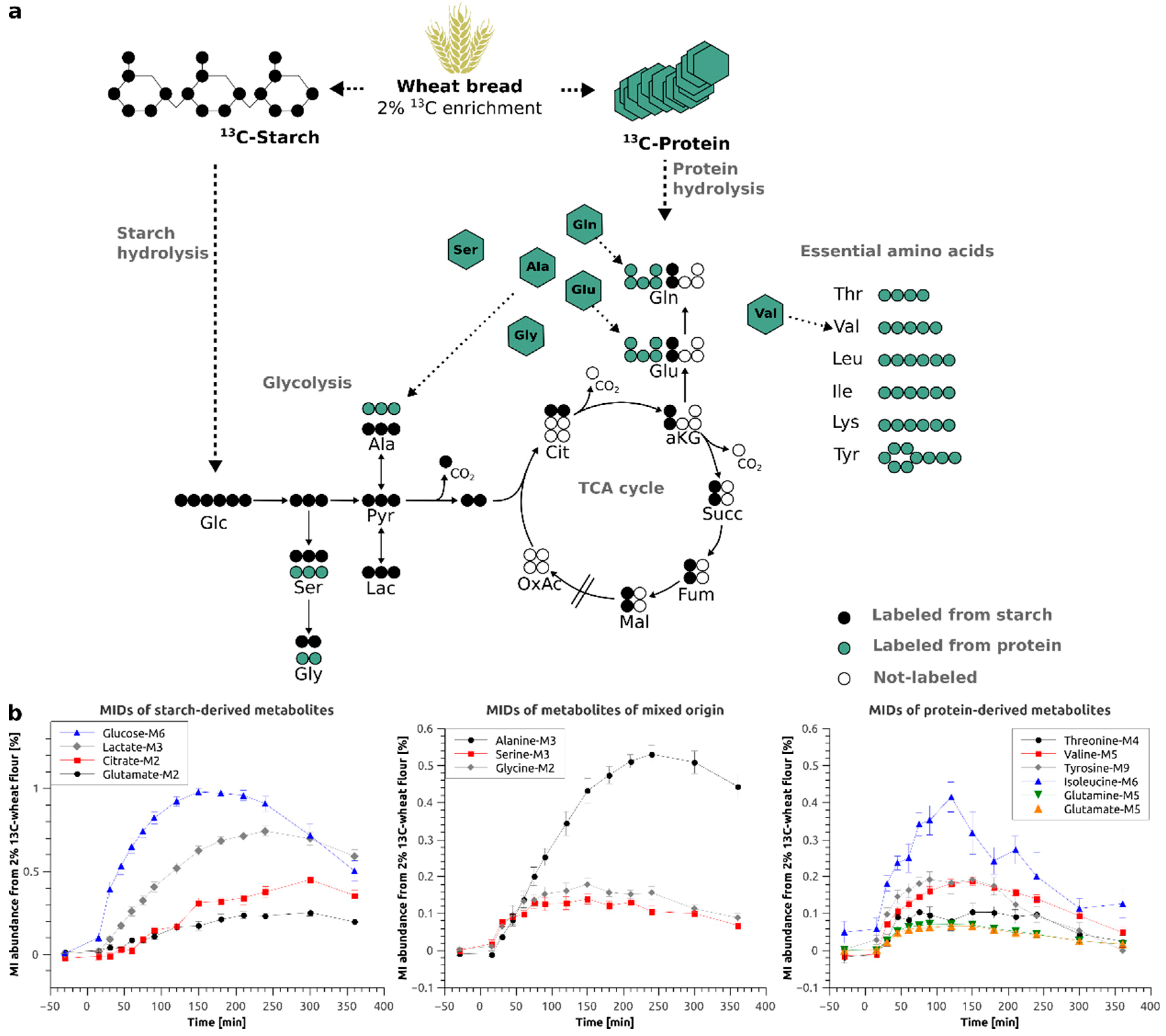
2.2. Incorporation of 13C Enrichment from 2% Labeled Wheat Flour in Central Carbon Metabolites
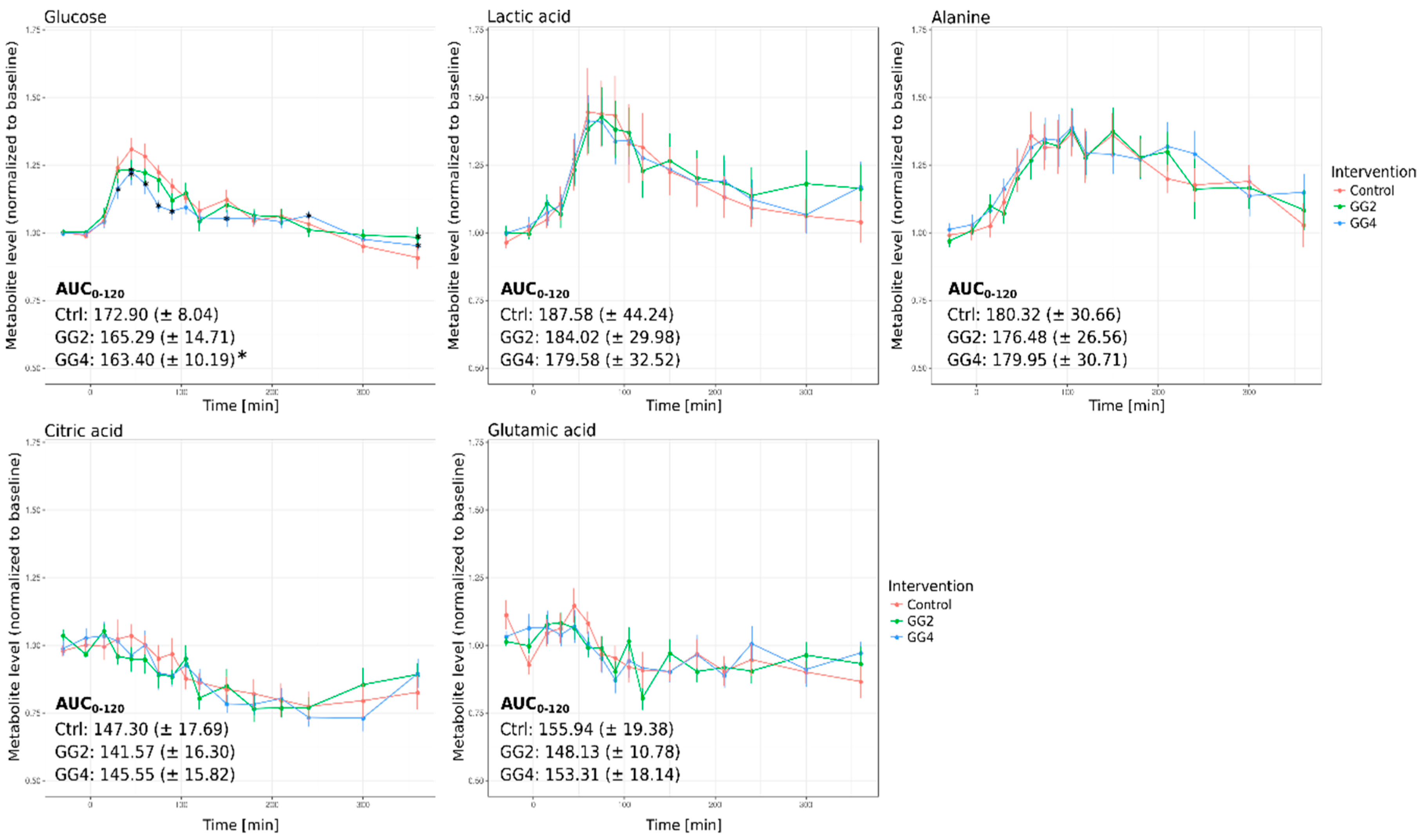
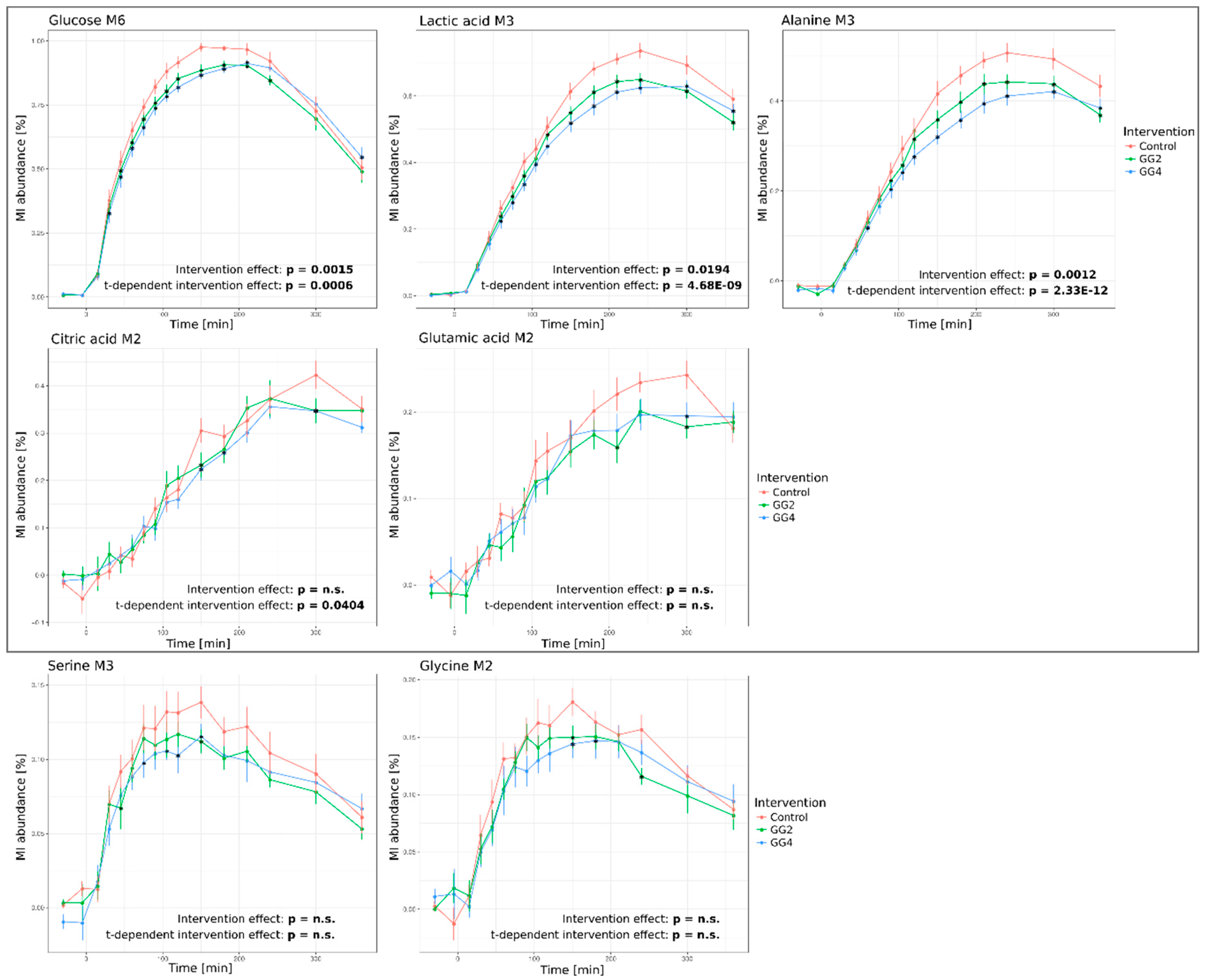
2.3. Effects of Fiber Supplementation on Postprandial Metabolite Levels and 13C-Enrichment
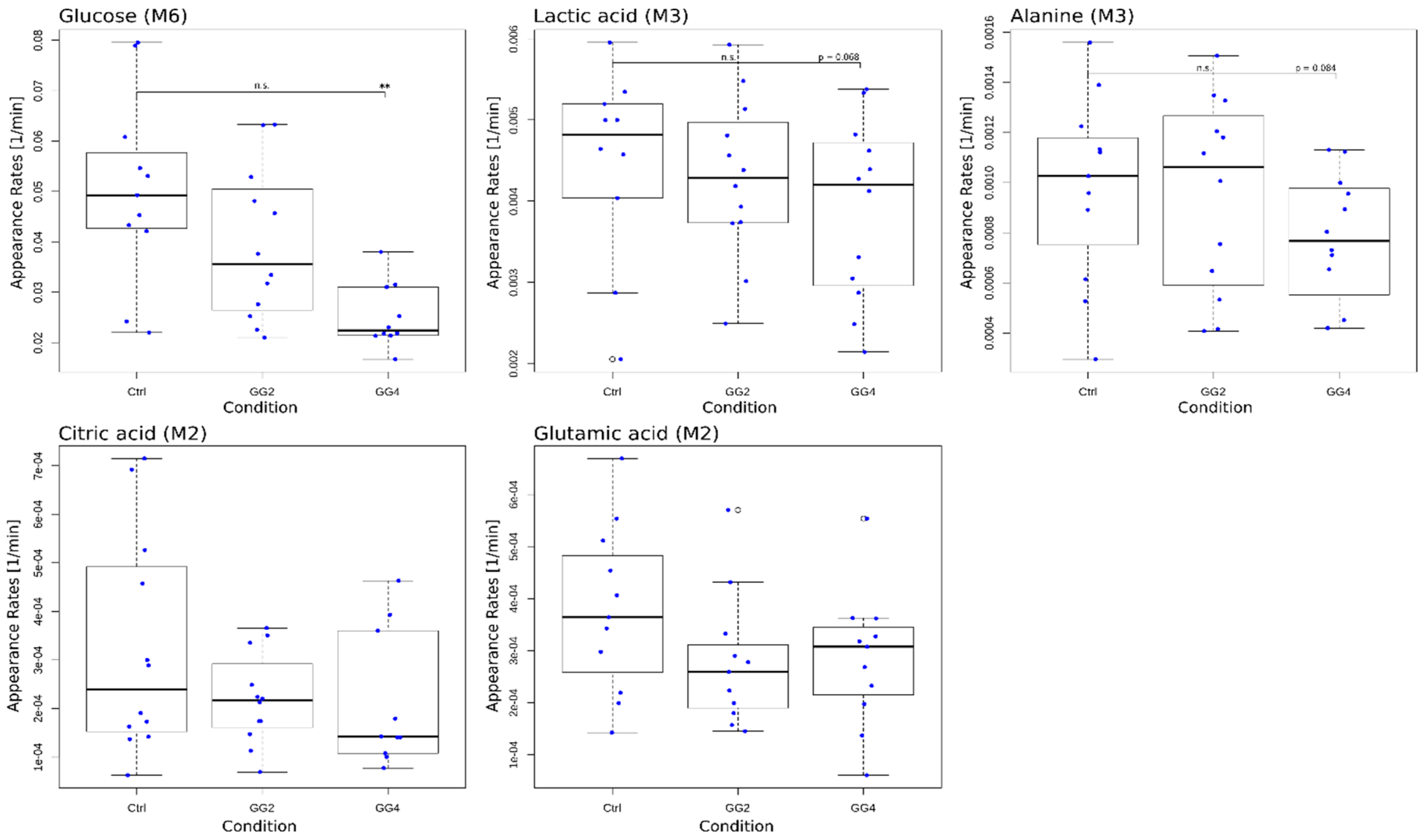
2.4. Postprandial Metabolic Turnover of Plasma Metabolites
2.5. Conclusions
3. Materials and Methods
3.1. Study Design
3.1.1. Subjects
3.1.2. Experimental Design
3.1.3. Test Product and Preparation
3.1.4. Sample Collection
3.2. Analytical Techniques
3.2.1. Metabolite Extraction from Plasma
3.2.2. Metabolite Derivatization and GC-MS Measurement
3.3. Data Analysis
3.3.1. MID Determination
3.3.2. Batch Quantification and Normalization
3.3.3. Contribution of Food-Derived Metabolites to the Total Metabolite Pool
3.3.4. AUC Determination
3.4. Statistical Analysis
3.4.1. Outlier Analysis
3.4.2. Repeated Measures (rm)ANOVA
3.4.3. Statistical Analysis of Single Time Points
3.5. Metabolic Modeling
Single Metabolites Model
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hu, F.B. Globalization of Diabetes: The Role of Diet, Lifestyle, and Genes. Diabetes Care 2011, 34, 1249–1257. [Google Scholar] [CrossRef] [PubMed]
- Tuomilehto, J.; Lindström, J.; Eriksson, J.G.; Valle, T.T.; Hämäläinen, H.; Ilanne-Parikka, P.; Kainänen-Kiukaanniemi, S.; Laakso, M.; Louheranta, A.; Rastas, M.; et al. Prevention of Type 2 Diabetes Mellitus By Changes In Lifestyle Among Subjects with Impaired Glucose Tolerance. N. Engl. J. Med. 2001, 344, 1343–1350. [Google Scholar] [CrossRef] [PubMed]
- Chiasson, J.-L.; Josse, R.G.; Gomis, R.; Hanefeld, M.; Karasik, A.; Laakso, M.; STOP-NIDDM Trail Research Group. Acarbose for Prevention of Type 2 Diabetes Mellitus: The STOP-NIDDM Randomised Trial. Lancet 2002, 359, 2072–2077. [Google Scholar] [CrossRef]
- Van de Laar, F.A.; Lucassen, P.L.; Akkermans, R.P.; Van de Lisdonk, E.H.; De Grauw, W.J. Alpha-Glucosidase Inhibitors for People with Impaired Glucose Tolerance or Impaired Fasting Blood Glucose. Cochrane Database Syst. Rev. 2006. [Google Scholar] [CrossRef] [PubMed]
- Schnell, O.; Mertes, G.; Standl, E. Acarbose and Metabolic Control in Patients with Type 2 Diabetes with Newly Initiated Insulin Therapy. Diabetes Obes. Metab. 2007, 9, 853–858. [Google Scholar] [CrossRef] [PubMed]
- Blaak, E.E.; Antoine, J.-M.; Benton, D.; Björck, I.; Bozzetto, L.; Brouns, F.; Diamant, M.; Dye, L.; Hulshof, T.; Holst, J.J.; et al. Impact of Postprandial Glycaemia on Health and Prevention of Disease. Obes. Rev. 2012, 13, 923–984. [Google Scholar] [CrossRef]
- Lafiandra, D.; Riccardi, G.; Shewry, P.R. Improving Cereal Grain Carbohydrates for Diet and Health. J. Cereal Sci. 2014, 59, 312–326. [Google Scholar] [CrossRef] [PubMed]
- Ray, K.S.; Singhania, P.R. Glycemic and Insulinemic Responses to Carbohydrate Rich Whole Foods. J. Food Sci. Technol. 2014, 51, 347–352. [Google Scholar] [CrossRef]
- Boers, H.M.; Macaulay, K.; Murray, P.; Seijen Ten Hoorn, J.; Hoogenraad, A.; Harry, P.F.; Vente-spreeuwenberg, M.A.M.; Mela, D.J. Efficacy of Different Fibres and Flour Mixes in South-Asian Flatbreads for Reducing Post-prandial Glucose Responses in Healthy Adults. Eur. J. Nutr. 2017, 56, 2049–2060. [Google Scholar] [CrossRef] [PubMed]
- Boers, H.M.; Macaulay, K.; Murray, P.; Dobriyal, R.; Mela, D.J.; Spreeuwenberg, M.A.M. Efficacy of Fibre Additions to Flatbread Flour Mixes for Reducing Post-Meal Glucose and Insulin Responses in Healthy Indian Subjects. Br. J. Nutr. 2017, 117, 386–394. [Google Scholar] [CrossRef]
- Ohashi, Y.; Sumitani, K.; Tokunaga, M.; Ishihara, N.; Okubo, T.; Fujisawa, T. Consumption of Partially Hydrolysed Guar Gum Stimulates Bifidobacteria and Butyrate-Producing Bacteria in the Human Large Intestine. Benef. Microbes 2015, 6, 451–455. [Google Scholar] [CrossRef] [PubMed]
- Papathanasopoulos, A.; Camilleri, M. Dietary Fiber Supplements: Effects in Obesity and Metabolic Syndrome and Relationship to Gastrointestinal Functions. Gastroenterology 2010, 138, 65–72. [Google Scholar] [CrossRef] [PubMed]
- Den Besten, G.; Gerding, A.; van Dijk, T.H.; Ciapaite, J.; Bleeker, A.; van Eunen, K.; Havinga, R.; Groen, A.K.; Reijngoud, D.-J.; Bakker, B.M. Protection against the Metabolic Syndrome by Guar Gum-Derived Short-Chain Fatty Acids Depends on Peroxisome Proliferator-Activated Receptor γ and Glucagon-Like Peptide-1. PLoS ONE 2015, 10, e0136364. [Google Scholar] [CrossRef] [PubMed]
- Osorio-Díaz, P.; Agama-Acevedo, E.; Mendoza-Vinalay, M.; Tovar, J.; Bello-Pérez, L.A. Pasta Added with Chickpea Flour: Chemical Composition, in Vitro Starch Digestibility and Predicted Glycemic Index. Cienc. Tecnol. Aliment. 2008, 6, 6–12. [Google Scholar] [CrossRef]
- Jukanti, A.K.; Gaur, P.M.; Gowda, C.L.L.; Chibbar, R.N. Nutritional Quality and Health Benefits of Chickpea (Cicer arietinum L.): A Review. Br. J. Nutr. 2012, 108, S11–S26. [Google Scholar] [CrossRef] [PubMed]
- Grundy, M.M.-L.; Edwards, C.H.; Mackie, A.R.; Gidley, M.J.; Butterworth, P.J.; Ellis, P.R. Re-Evaluation of the Mechanisms of Dietary Fibre and Implications for Macronutrient Bioaccessibility, Digestion and Postprandial Metabolism. Br. J. Nutr. 2016, 116, 816–833. [Google Scholar] [CrossRef] [PubMed]
- Ekström, L.M.N.K.; Björck, I.M.E.; Östman, E.M. On the Possibility to Affect the Course of Glycaemia, Insulinaemia, and Perceived Hunger/Satiety to Bread Meals in Healthy Volunteers. Food Funct. 2013, 4, 522–529. [Google Scholar] [CrossRef] [PubMed]
- Eelderink, C.; Moerdijk-Poortvliet, T.C.W.; Wang, H.; Schepers, M.; Preston, T.; Boer, T.; Vonk, R.J.; Schierbeek, H.; Priebe, M.G. The Glycemic Response Does Not Reflect the In Vivo Starch Digestibility of Fiber-Rich Wheat Products in Healthy Men. J. Nutr 2012, 142, 258–263. [Google Scholar] [CrossRef]
- Boers, H.M.; Van Dijk, T.H.; Hiemstra, H.; Hoogenraad, A.R.; Mela, D.J.; Peters, H.P.F.; Vonk, R.J.; Priebe, M.G. Effect of Fibre Additions to Flatbread Flour Mixes on Glucose Kinetics: A Randomised Controlled. Br. J. Nutr. 2017, 118, 777–787. [Google Scholar] [CrossRef] [PubMed]
- Brennan, L. Metabolomics in Nutrition Research: Current Status and Perspectives. Biochem. Soc. Trans. 2013, 41, 670–673. [Google Scholar] [CrossRef]
- Wishart, D.S. Metabolomics: Applications to Food Science and Nutrition Research. Trends Food Sci. Technol. 2008, 19, 482–493. [Google Scholar] [CrossRef]
- Cortassa, S.; Caceres, V.; Bell, L.N.; O’Rourke, B.; Paolocci, N.; Aon, M.A. From Metabolomics to Fluxomics: A Computational Procedure to Translate Metabolite Profiles into Metabolic Fluxes. Biophys. J. 2015, 108, 163–172. [Google Scholar] [CrossRef] [PubMed]
- Wasylenko, T.M.; Stephanopoulos, G. Kinetic Isotope Effects Significantly Influence Intracellular Metabolite 13C Labeling Patterns and Flux Determination. Biotechnol. J. 2013, 8, 1080–1089. [Google Scholar] [CrossRef] [PubMed]
- Shrestha, A.; Müllner, E.; Poutanen, K.; Mykkänen, H.; Moazzami, A.A. Metabolic Changes in Serum Metabolome in Response to a Meal. Eur. J. Nutr. 2017, 56, 671–681. [Google Scholar] [CrossRef] [PubMed]
- Shi, L.; Brunius, C.; Lindelöf, M.; Shameh, S.A.; Wu, H.; Lee, I.; Landberg, R.; Moazzami, A.A. Targeted Metabolomics Reveals Differences in the Extended Postprandial Plasma Metabolome of Healthy Subjects after Intake of Whole-Grain Rye Porridges versus Refined Wheat Bread. Mol. Nutr. Food Res. 2017, 61, 1600924. [Google Scholar] [CrossRef] [PubMed]
- Pantophlet, A.J.; Wopereis, S.; Eelderink, C.; Vonk, R.J.; Stroeve, J.H.; Bijlsma, S.; van Stee, L.; Bobeldijk, I.; Priebe, M.G. Metabolic Profiling Reveals Differences in Plasma Concentrations of Arabinose and Xylose after Consumption of Fiber-Rich Pasta and Wheat Bread with Differential Rates of Systemic Appearance of Exogenous Glucose in Healthy Men. J. Nutr. 2017, 147, 152–160. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Pellis, L.; van Erk, M.J.; van Ommen, B.; Bakker, G.C.M.; Hendriks, H.F.J.; Cnubben, N.H.P.; Kleemann, R.; van Someren, E.P.; Bobeldijk, I.; Rubingh, C.M.; et al. Plasma Metabolomics and Proteomics Profiling after a Postprandial Challenge Reveal Subtle Diet Effects on Human Metabolic Status. Metabolomics 2012, 8, 347–359. [Google Scholar] [CrossRef] [PubMed]
- Moazzami, A.A.; Shrestha, A.; Morrison, D.A.; Poutanen, K.; Mykkänen, H. Metabolomics Reveals Differences in Postprandial Responses to Breads and Fasting Metabolic Characteristics Associated with Postprandial Insulin Demand in Postmenopausal Women. J. Nutr. 2014, 144, 807–814. [Google Scholar] [CrossRef]
- Fiamoncini, J.; Rundle, M.; Gibbons, H.; Thomas, L.; Geillinger-Kästle, K.; Bunzel, D.; Trezzi, J.-P.; Kiselova-Kaneva, Y.; Wopereis, S.; Wahrheit, J.; et al. Plasma Metabolome Analysis Identifies Distinct Human Metabotypes in the Postprandial State with Different Susceptibility to Weight Loss–Mediated Metabolic Improvements. FASEB J. 2018, 32, 5447–5458. [Google Scholar] [CrossRef] [PubMed]
- Ferrannini, E.; Natali, A.; Camastra, S.; Nannipieri, M.; Mari, A.; Adam, K.P.; Milburn, M.V.; Kastenmüller, G.; Adamski, J.; Tuomi, T.; et al. Early Metabolic Markers of the Development of Dysglycemia and Type 2 Diabetes and Their Physiological Significance. Diabetes 2013, 62, 1730–1737. [Google Scholar] [CrossRef] [PubMed]
- Eelderink, C.; Schepers, M.; Preston, T.; Vonk, R.J.; Oudhuis, L.; Priebe, M.G. Slowly and Rapidly Digestible Starchy Foods Can Elicit a Similar Glycemic Response Because of Differential Tissue Glucose Uptake in Healthy Men. Am. J. Clin. Nutr. 2012, 96, 1017–1024. [Google Scholar] [CrossRef] [PubMed]
- Darmaun, D.; Matthews, D.E.; Bier, D.M. Glutamine and Glutamate Kinetics in Humans. Am. J. Physiol. 1986, 251, E117–E126. [Google Scholar] [CrossRef] [PubMed]
- Yang, R.D.; Matthews, D.E.; Bier, D.M.; Wen, Z.M.; Young, V.R. Response of Alanine Metabolism in Humans to Manipulation of Dietary Protein and Energy Intakes. Am. J. Physiol. 1986, 250, E39–E46. [Google Scholar] [CrossRef] [PubMed]
- Rizza, R.A.; Toffolo, G.; Cobelli, C. Accurate Measurement of Postprandial Glucose Turnover: Why Is It Difficult and How Can It Be Done (Relatively) Simply? Diabetes 2016, 65, 1133–1145. [Google Scholar] [CrossRef] [PubMed]
- Krämer, L.; Jäger, C.; Trezzi, J.-P.; Jacobs, D.M.; Hiller, K. Quantification of Stable Isotope Traces Close to Natural Enrichment in Human Plasma Metabolites Using Gas Chromatography-Mass Spectrometry. Metabolites 2018, 8, 15. [Google Scholar] [CrossRef] [PubMed]
- Grabacka, M.; Pierzchalska, M.; Dean, M.; Reiss, K. Regulation of Ketone Body Metabolism and the Role of PPARα. Int. J. Mol. Sci. 2016, 17, 2093. [Google Scholar] [CrossRef] [PubMed]
- Krug, S.; Kastenmuller, G.; Stuckler, F.; Rist, M.J.; Skurk, T.; Sailer, M.; Raffler, J.; Romisch-Margl, W.; Adamski, J.; Prehn, C.; et al. The Dynamic Range of the Human Metabolome Revealed by Challenges. FASEB J. 2012, 26, 2607–2619. [Google Scholar] [CrossRef] [PubMed]
- Van Der Borght, A.; Goesaert, H.; Veraverbeke, W.S.; Delcour, J.A. Fractionation of Wheat and Wheat Flour into Starch and Gluten: Overview of the Main Processes and the Factors Involved. J. Cereal Sci. 2005, 41, 221–237. [Google Scholar] [CrossRef]
- Rombouts, I.; Lamberts, L.; Celus, I.; Lagrain, B.; Brijs, K.; Delcour, J.A. Wheat Gluten Amino Acid Composition Analysis by High-Performance Anion-Exchange Chromatography with Integrated Pulsed Amperometric Detection. J. Chromatogr. A 2009, 1216, 5557–5562. [Google Scholar] [CrossRef]
- Adibi, S.A.; Gray, S.J. Intestinal Absorption of Essential Amino Acids in Man. Gastroenterology 1967, 52, 837–845. [Google Scholar] [CrossRef]
- Adibi, S.A.; Gray, S.J.; Menden, E. The Kinetics of Amino Acid Absorption and Alteration of Plasma Composition of Free Amino Acids After Intestinal Perfusion of Amino Acid Mixtures. Am. J. Clin. Nutr. 1967, 20, 24–33. [Google Scholar] [CrossRef] [PubMed]
- Webb, K.E. Intestinal Absorption of Protein Hydrolysis Products: A Review. J. Anim. Sci. 1990, 68, 3011–3022. [Google Scholar] [CrossRef]
- Levitt, D.G.; Levitt, M.D. A Model of Blood-Ammonia Homeostasis Based on a Quantitative Analysis of Nitrogen Metabolism in the Multiple Organs Involved in the Production, Catabolism, and Excretion of Ammonia in Humans. Clin. Exp. Gastroenterol. 2018, 11, 193–215. [Google Scholar] [CrossRef]
- Adeva, M.M.; Souto, G.; Blanco, N.; Donapetry, C. Ammonium Metabolism in Humans. Metabolism 2012, 61, 1495–1511. [Google Scholar] [CrossRef]
- Nie, C.; He, T.; Zhang, W.; Zhang, G.; Ma, X. Branched Chain Amino Acids: Beyond Nutrition Metabolism. Int. J. Mol. Sci. 2018, 19, 954. [Google Scholar] [CrossRef] [PubMed]
- Dario Frias, A.C.; Sgarbieri, V.C. Guar Gum Effects on Food Intake, Blood Serum Lipids and Glucose Levels of Wistar Rats. Plant Foods Hum. Nutr. 1998, 53, 15–28. [Google Scholar] [CrossRef]
- Mariotti, F.; Pueyo, M.E.; Tomé, D.; Benamouzig, R.; Mahé, S. Guar Gum Does Not Impair the Absorption and Utilization of Dietary Nitrogen but Affects Early Endogenous Urea Kinetics in Humans. Am. J. Clin. Nutr. 2001, 74, 487–493. [Google Scholar] [CrossRef] [PubMed]
- Rochus, K.; Janssens, G.P.J.; Van de Velde, H.; Verbrugghe, A.; Wuyts, B.; Vanhaecke, L.; Hesta, M. Highly Viscous Guar Gum Shifts Dietary Amino Acids from Metabolic Use to Fermentation Substrate in Domestic Cats. Br. J. Nutr. 2013, 109, 1022–1030. [Google Scholar] [CrossRef] [PubMed]
- Katz, J.; Lee, W.N.P.; Wals, P.A.; Bergner, E.A. Studies of Glycogen Synthesis and the Krebs Cycle by Mass Isotopomer Analysis with [U-13C]Glucose in Rats. J. Biol. Chem. 1989, 264, 12994–13001. [Google Scholar]
- Hiller, K.; Hangebrauk, J.; Jäger, C.; Spura, J.; Schreiber, K.; Schomburg, D. Metabolite Detector: Comprehensive Analysis Tool for Targeted and Nontargeted GC/MS Based Metabolome Analysis. Anal. Chem. 2009, 81, 3429–3439. [Google Scholar] [CrossRef] [PubMed]
- Jennings, M.E.; Matthews, D.E. Determination of Complex Isotopomer Patterns in Isotopically Labeled Compounds by Mass Spectrometry. Anal. Chem. 2005, 77, 6435–6444. [Google Scholar] [CrossRef] [PubMed]
- Trezzi, J.-P.; Galozzi, S.; Jaeger, C.; Barkovits, K.; Brockmann, K.; Maetzler, W.; Berg, D.; Marcus, K.; Betsou, F.; Hiller, K.; et al. Distinct Metabolomic Signature in Cerebrospinal Fluid in Early Parkinson’s Disease. Mov. Disord. 2017, 32, 1401–1408. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Y.; Hoon Yang, T.; Heinzle, E. 13C Metabolic Flux Analysis for Larger Scale Cultivation Using Gas Chromatography-Combustion-Isotope Ratio Mass Spectrometry. Metab. Eng. 2010, 12, 392–400. [Google Scholar] [CrossRef] [PubMed]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. Methodol. 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Marquardt, D.W. An Algorithm for Least-Squares Estimation of Nonlinear Parameters. J. Soc. Ind. Appl. Math. 1963, 11, 431–441. [Google Scholar] [CrossRef]
- Levenberg, K. A Method for the Solution of Certain Non-Linear Problems in Least Squares. Q. Appl. Math. 1944, 2, 164–168. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Schlicker, L.; Boers, H.M.; Dudek, C.-A.; Zhao, G.; Barua, A.; Trezzi, J.-P.; Meyer-Hermann, M.; Jacobs, D.M.; Hiller, K. Postprandial Metabolic Effects of Fiber Mixes Revealed by in vivo Stable Isotope Labeling in Humans. Metabolites 2019, 9, 91. https://doi.org/10.3390/metabo9050091
Schlicker L, Boers HM, Dudek C-A, Zhao G, Barua A, Trezzi J-P, Meyer-Hermann M, Jacobs DM, Hiller K. Postprandial Metabolic Effects of Fiber Mixes Revealed by in vivo Stable Isotope Labeling in Humans. Metabolites. 2019; 9(5):91. https://doi.org/10.3390/metabo9050091
Chicago/Turabian StyleSchlicker, Lisa, Hanny M. Boers, Christian-Alexander Dudek, Gang Zhao, Arnab Barua, Jean-Pierre Trezzi, Michael Meyer-Hermann, Doris M. Jacobs, and Karsten Hiller. 2019. "Postprandial Metabolic Effects of Fiber Mixes Revealed by in vivo Stable Isotope Labeling in Humans" Metabolites 9, no. 5: 91. https://doi.org/10.3390/metabo9050091
APA StyleSchlicker, L., Boers, H. M., Dudek, C.-A., Zhao, G., Barua, A., Trezzi, J.-P., Meyer-Hermann, M., Jacobs, D. M., & Hiller, K. (2019). Postprandial Metabolic Effects of Fiber Mixes Revealed by in vivo Stable Isotope Labeling in Humans. Metabolites, 9(5), 91. https://doi.org/10.3390/metabo9050091





