Degradation of Extracellular NAD+ Intermediates in Cultures of Human HEK293 Cells
Abstract
1. Introduction
2. Results
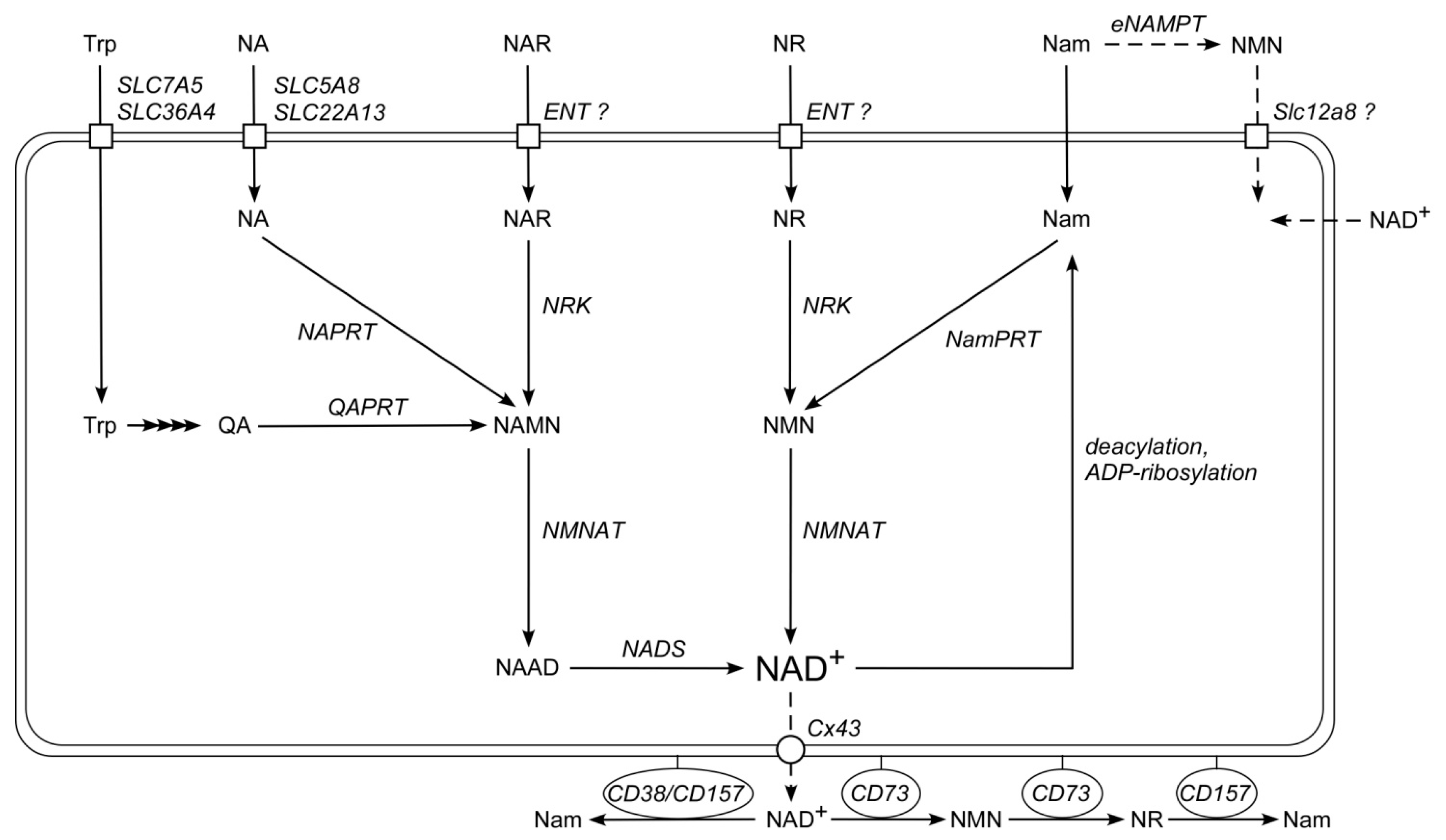
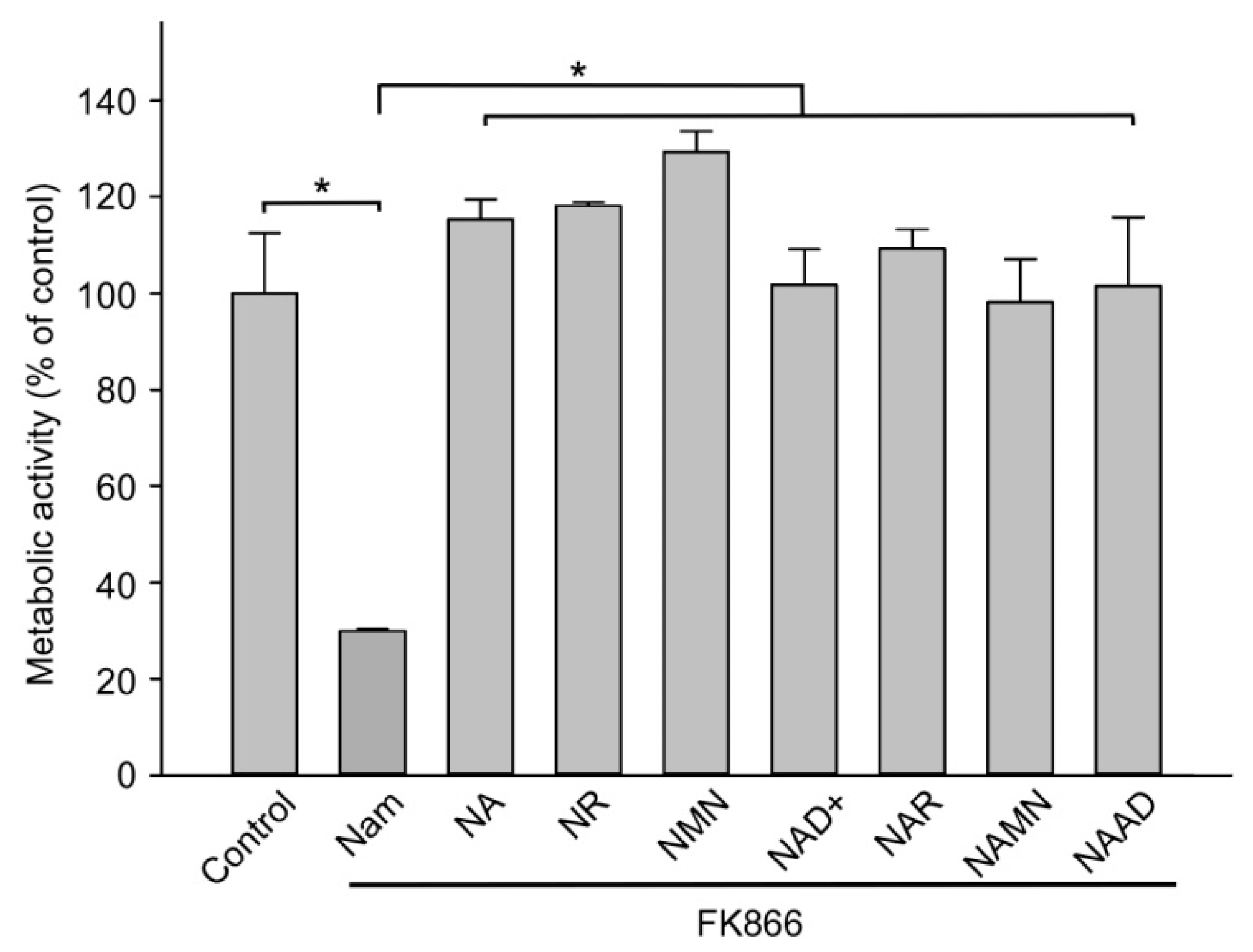
2.1. Major Extracellular NAD Intermediates Support NAD Generation in HEK293 Cells
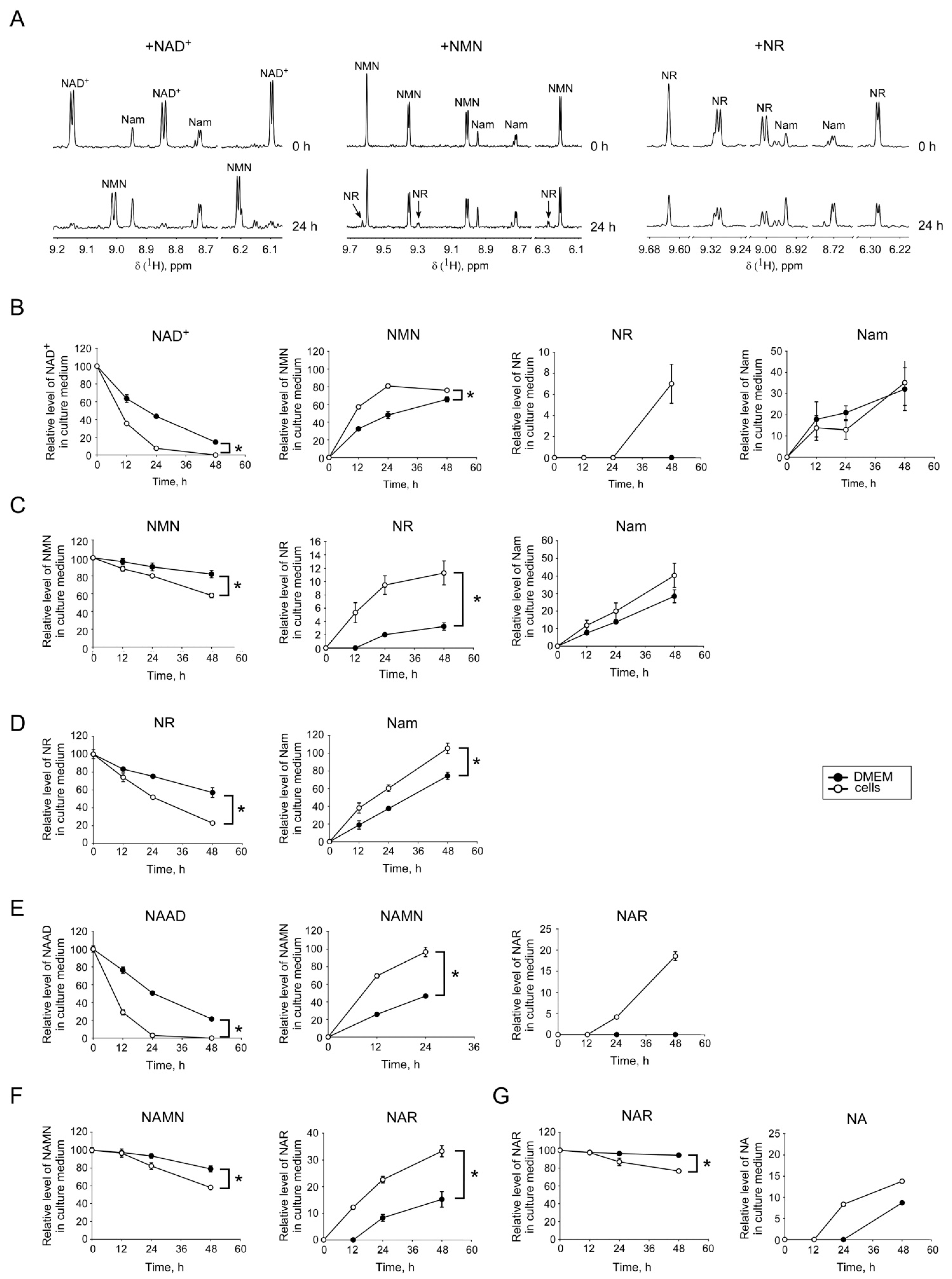
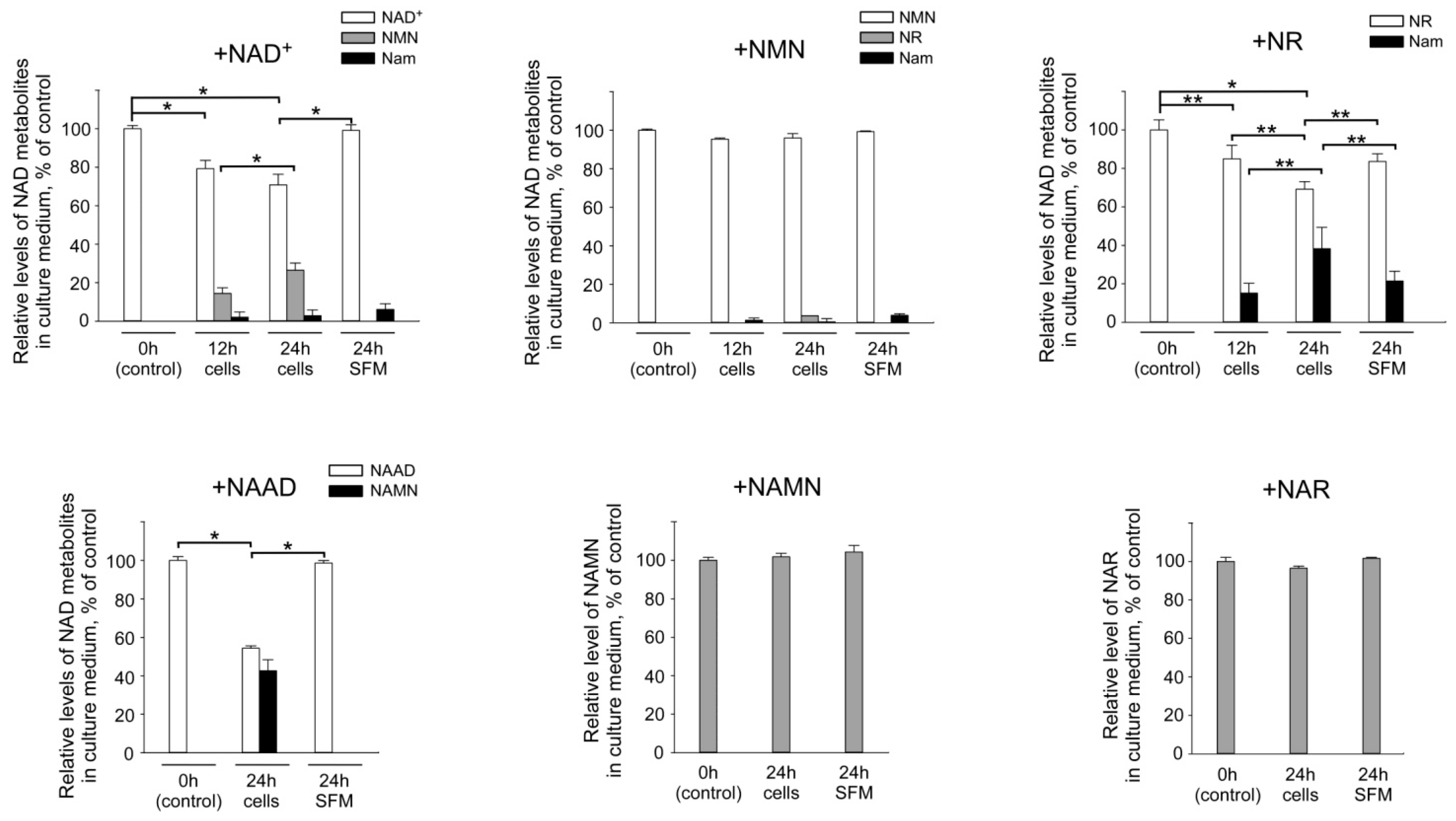
2.2. Interconversion and Degradation of Extracellular NAD Intermediates in Cultures of HEK293 Cells
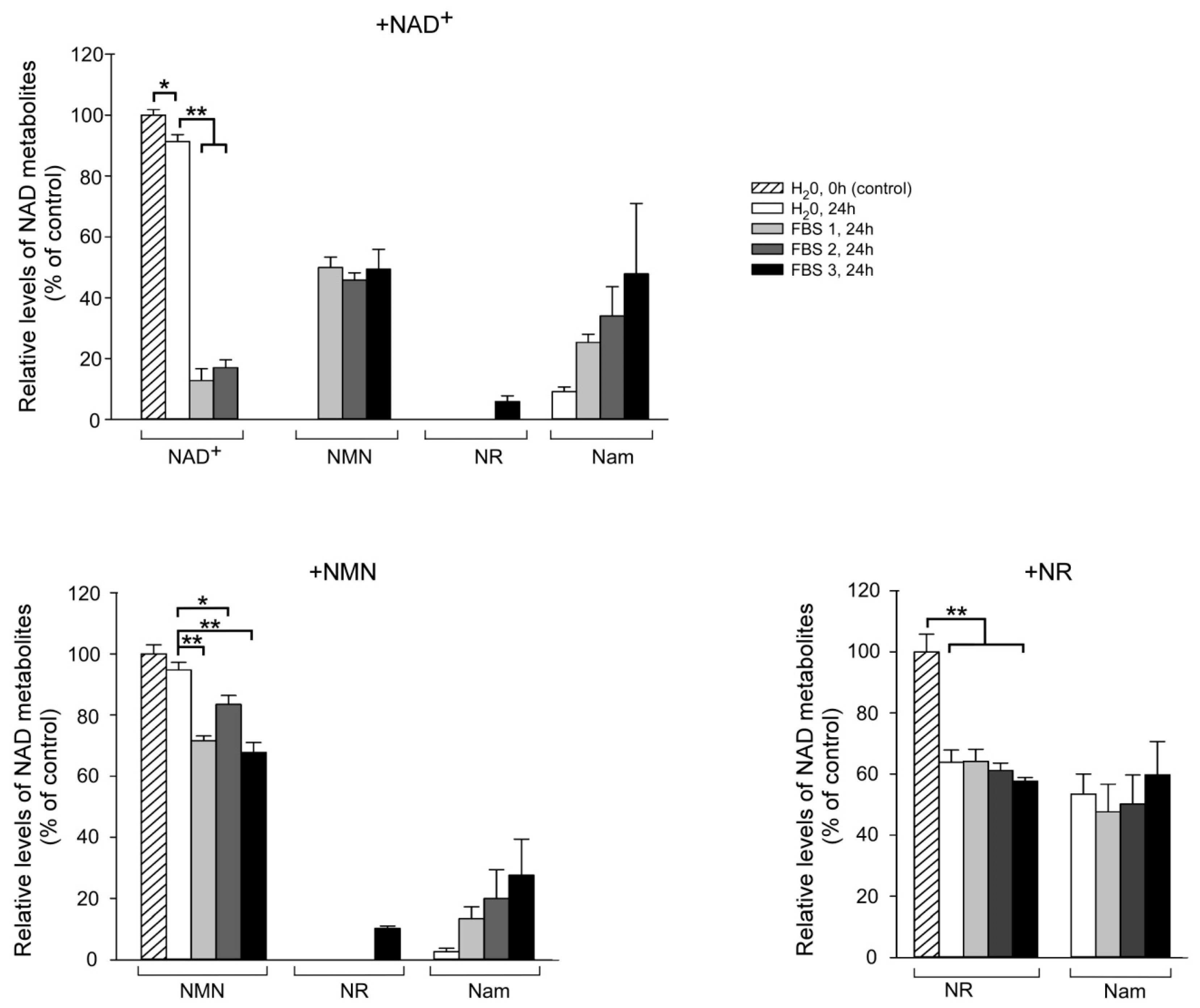
2.3. Fetal Bovine Serum (FBS) Has Pyridine Nucleotide-Degrading Activities
2.4. Cellular Ecto-Enzymes Metabolize Extracellular Pyridine Nucleotides
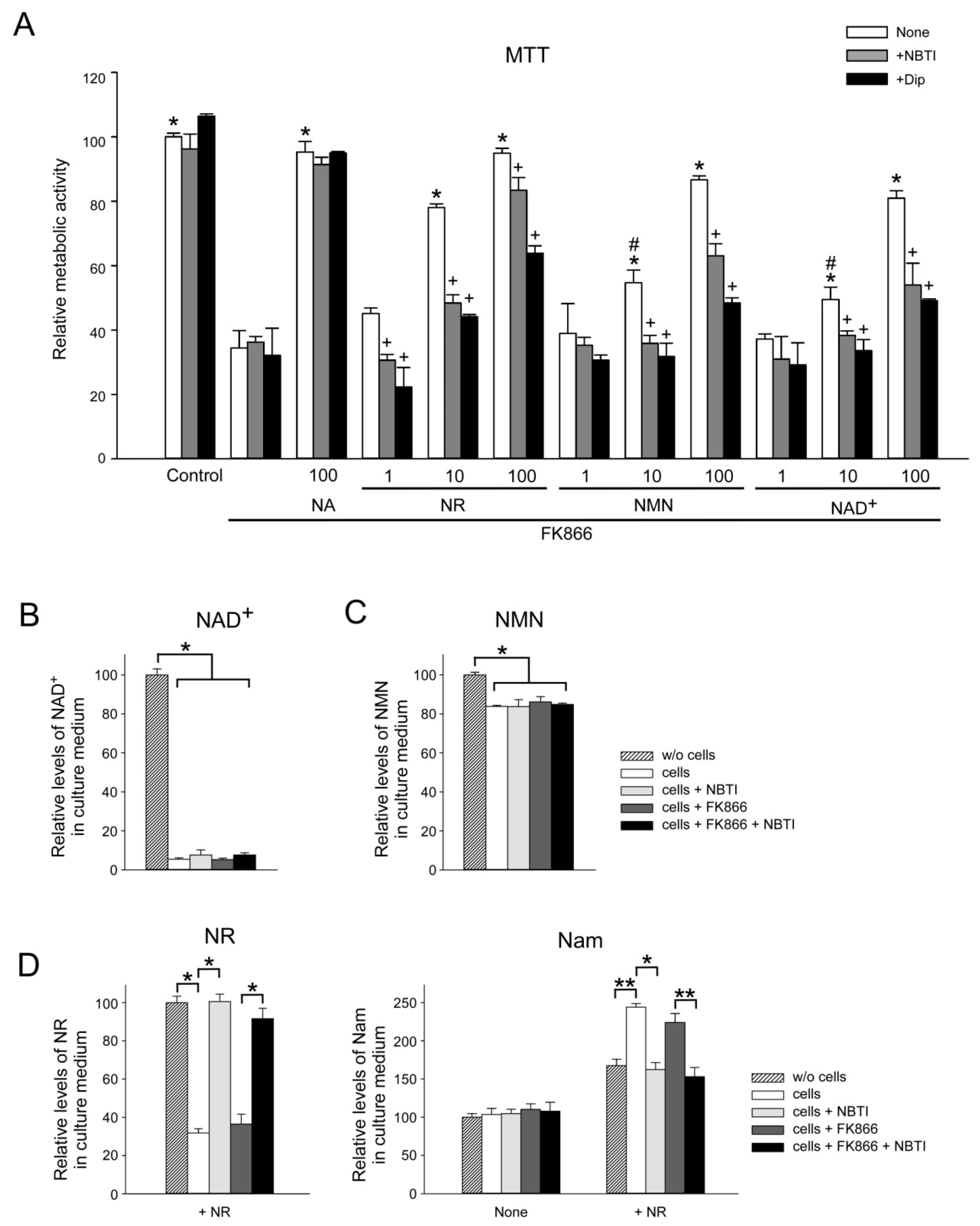
2.5. ENT Inhibition Suppresses NR, NMN, and NAD+ Utilization by HEK293 Cells
3. Discussion
4. Materials and Methods
4.1. Materials
4.2. Cell Culture
4.3. Pharmacological Treatments
4.4. NMR Sample Preparation
4.5. NMR Analysis
4.6. Statistical Analysis
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Houtkooper, R.H.; Canto, C.; Wanders, R.J.; Auwerx, J. The secret life of NAD+: An old metabolite controlling new metabolic signaling pathways. Endocr. Rev. 2010, 31, 194–223. [Google Scholar] [CrossRef] [PubMed]
- Kulikova, V.A.; Gromyko, D.V.; Nikiforov, A.A. The Regulatory Role of NAD in Human and Animal Cells. Biochemistry (Moscow) 2018, 83, 800–812. [Google Scholar] [CrossRef] [PubMed]
- Stromland, Ø.; Niere, M.; Nikiforov, A.A.; VanLinden, M.R.; Heiland, I.; Ziegler, M. Keeping the balance in NAD metabolism. Biochem. Soc. Transact. 2019, 47, 119–130. [Google Scholar] [CrossRef] [PubMed]
- Dölle, C.; Skoge, R.H.; Vanlinden, M.R.; Ziegler, M. NAD biosynthesis in humans–enzymes, metabolites and therapeutic aspects. Curr. Top. Med. Chem. 2013, 13, 2907–2917. [Google Scholar] [CrossRef] [PubMed]
- Nikiforov, A.; Kulikova, V.; Ziegler, M. The human NAD metabolome: Functions, metabolism and compartmentalization. Critical Rev. Biochem. Mol. Biol. 2015, 50, 284–297. [Google Scholar] [CrossRef]
- Bieganowski, P.; Brenner, C. Discoveries of nicotinamide riboside as a nutrient and conserved NRK genes establish a Preiss-Handler independent route to NAD+ in fungi and humans. Cell 2004, 117, 495–502. [Google Scholar] [CrossRef]
- Kulikova, V.; Shabalin, K.; Nerinovski, K.; Dölle, C.; Niere, M.; Yakimov, A.; Redpath, P.; Khodorkovskiy, M.; Migaud, M.E.; Ziegler, M.; et al. Generation, Release, and Uptake of the NAD Precursor Nicotinic Acid Riboside by Human Cells. J. Biol. Chem. 2015, 290, 27124–27137. [Google Scholar] [CrossRef]
- Bahn, A.; Hagos, Y.; Reuter, S.; Balen, D.; Brzica, H.; Krick, W.; Burckhardt, B.C.; Sabolic, I.; Burckhardt, G. Identification of a new urate and high affinity nicotinate transporter, hOAT10 (SLC22A13). J. Biol. Chem. 2008, 283, 16332–16341. [Google Scholar] [CrossRef]
- Gopal, E.; Miyauchi, S.; Martin, P.M.; Ananth, S.; Roon, P.; Smith, S.B.; Ganapathy, V. Transport of nicotinate and structurally related compounds by human SMCT1 (SLC5A8) and its relevance to drug transport in the mammalian intestinal tract. Pharm. Res. 2007, 24, 575–584. [Google Scholar] [CrossRef]
- Kanai, Y.; Segawa, H.; Miyamoto, K.; Uchino, H.; Takeda, E.; Endou, H. Expression cloning and characterization of a transporter for large neutral amino acids activated by the heavy chain of 4F2 antigen (CD98). J. Biol. Chem. 1998, 273, 23629–23632. [Google Scholar] [CrossRef]
- Pillai, S.M.; Meredith, D. SLC36A4 (hPAT4) is a high affinity amino acid transporter when expressed in Xenopus laevis oocytes. J. Biol. Chem. 2011, 286, 2455–2640. [Google Scholar] [CrossRef] [PubMed]
- Nikiforov, A.; Dölle, C.; Niere, M.; Ziegler, M. Pathways and subcellular compartmentation of NAD biosynthesis in human cells: From entry of extracellular precursors to mitochondrial NAD generation. J. Biol. Chem. 2011, 286, 21767–21778. [Google Scholar] [CrossRef] [PubMed]
- Pittelli, M.; Felici, R.; Pitozzi, V.; Giovannelli, L.; Bigagli, E.; Cialdai, F.; Romano, G.; Moroni, F.; Chiarugi, A. Pharmacological effects of exogenous NAD on mitochondrial bioenergetics, DNA repair, and apoptosis. Mol. Pharmacol. 2011, 80, 1136–1146. [Google Scholar] [CrossRef] [PubMed]
- Zheng, T.; Xu, S.Y.; Zhou, S.Q.; Lai, L.Y.; Li, L. Nicotinamide adenine dinucleotide (NAD+) repletion attenuates bupivacaine-induced neurotoxicity. Neurochem. Res. 2013, 38, 1880–19884. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Wang, P.; Liu, X.; He, D.; Liang, C.; Yu, Y. Exogenous NAD(+) supplementation protects H9c2 cardiac myoblasts against hypoxia/reoxygenation injury via Sirt1-p53 pathway. Fundam. Clin. Pharmacol. 2014, 28, 180–189. [Google Scholar] [CrossRef]
- Yoshino, J.; Baur, J.A.; Imai, S.I. NAD(+) Intermediates: The Biology and Therapeutic Potential of NMN and NR. Cell Metabol. 2018, 27, 513–528. [Google Scholar] [CrossRef]
- Rajman, L.; Chwalek, K.; Sinclair, D.A. Therapeutic Potential of NAD-Boosting Molecules: The In Vivo Evidence. Cell Metabol. 2018, 27, 529–547. [Google Scholar] [CrossRef]
- Ratajczak, J.; Joffraud, M.; Trammell, S.A.; Ras, R.; Canela, N.; Boutant, M.; Kulkarni, S.S.; Rodrigues, M.; Redpath, P.; Migaud, M.E.; et al. NRK1 controls nicotinamide mononucleotide and nicotinamide riboside metabolism in mammalian cells. Nat. Commun. 2016, 7, 13103. [Google Scholar] [CrossRef]
- Grozio, A.; Sociali, G.; Sturla, L.; Caffa, I.; Soncini, D.; Salis, A.; Raffaelli, N.; De Flora, A.; Nencioni, A.; Bruzzone, S. CD73 protein as a source of extracellular precursors for sustained NAD+ biosynthesis in FK866-treated tumor cells. J. Biol. Chem. 2013, 288, 25938–25949. [Google Scholar] [CrossRef]
- Garavaglia, S.; Bruzzone, S.; Cassani, C.; Canella, L.; Allegrone, G.; Sturla, L.; Mannino, E.; Millo, E.; De Flora, A.; Rizzi, M. The high-resolution crystal structure of periplasmic Haemophilus influenzae NAD nucleotidase reveals a novel enzymatic function of human CD73 related to NAD metabolism. Biochem. J. 2012, 441, 131–141. [Google Scholar] [CrossRef]
- Malavasi, F.; Deaglio, S.; Funaro, A.; Ferrero, E.; Horenstein, A.L.; Ortolan, E.; Vaisitti, T.; Aydin, S. Evolution and function of the ADP ribosyl cyclase/CD38 gene family in physiology and pathology. Physiol. Rev. 2008, 88, 841–886. [Google Scholar] [CrossRef] [PubMed]
- Preugschat, F.; Carter, L.H.; Boros, E.E.; Porter, D.J.; Stewart, E.L.; Shewchuk, L.M. A pre-steady state and steady state kinetic analysis of the N-ribosyl hydrolase activity of hCD157. Arch. Biochem. Biophys. 2014, 564, 156–163. [Google Scholar] [CrossRef] [PubMed]
- Billington, R.A.; Travelli, C.; Ercolano, E.; Galli, U.; Roman, C.B.; Grolla, A.A.; Canonico, P.L.; Condorelli, F.; Genazzani, A.A. Characterization of NAD uptake in mammalian cells. J. Biol. Chem. 2008, 283, 6367–6374. [Google Scholar] [CrossRef] [PubMed]
- Formentini, L.; Moroni, F.; Chiarugi, A. Detection and pharmacological modulation of nicotinamide mononucleotide (NMN) in vitro and in vivo. Biochem. Pharmacol. 2009, 77, 1612–1620. [Google Scholar] [CrossRef] [PubMed]
- Grozio, A.; Mills, K.F.; Yoshino, J.; Bruzzone, S.; Sociali, G.; Tokizane, K.; Lei, H.C.; Cunningham, R.; Sasaki, Y.; Migaud, M.E.; et al. Slc12a8 is a nicotinamide mononucleotide transporter. Nat. Metabol. 2019, 1, 47–57. [Google Scholar] [CrossRef]
- Schmidt, M.; Brenner, C. Absence of evidence that Slc12a8 encodes a nicotinamide mononucleotide transporter. Nat. Metabol. 2019, 1, 660–661. [Google Scholar] [CrossRef]
- Shabalin, K.; Nerinovski, K.; Yakimov, A.; Kulikova, V.; Svetlova, M.; Solovjeva, L.; Khodorkovskiy, M.; Gambaryan, S.; Cunningham, R.; Migaud, M.E.; et al. NAD Metabolome Analysis in Human Cells Using (1)H NMR Spectroscopy. Int. J. Mol. Sci. 2018, 19, 3906. [Google Scholar] [CrossRef]
- Hasmann, M.; Schemainda, I. FK866, a highly specific noncompetitive inhibitor of nicotinamide phosphoribosyltransferase, represents a novel mechanism for induction of tumor cell apoptosis. Cancer Res. 2003, 63, 7436–7442. [Google Scholar]
- Koziolkiewicz, M.; Gendaszewska, E.; Maszewska, M.; Stein, C.A.; Stec, W.J. The mononucleotide-dependent, nonantisense mechanism of action of phosphodiester and phosphorothioate oligonucleotides depends upon the activity of an ecto-5′-nucleotidase. Blood 2001, 98, 995–1002. [Google Scholar] [CrossRef]
- Yegutkin, G.G.; Samburski, S.S.; Jalkanen, S. Soluble purine-converting enzymes circulate in human blood and regulate extracellular ATP level via counteracting pyrophosphatase and phosphotransfer reactions. FASEB J. Off. Public. Fed. Am. Soc. Experiment. Biol. 2003, 17, 1328–1330. [Google Scholar] [CrossRef]
- Gendaszewska-Darmach, E.; Maszewska, M.; Zaklos, M.; Koziolkiewicz, M. Degradation of extracellular nucleotides and their analogs in HeLa and HUVEC cell cultures. Acta Biochim. Polonica 2003, 50, 973–984. [Google Scholar] [PubMed]
- Mills, K.F.; Yoshida, S.; Stein, L.R.; Grozio, A.; Kubota, S.; Sasaki, Y.; Redpath, P.; Migaud, M.E.; Apte, R.S.; Uchida, K.; et al. Long-Term Administration of Nicotinamide Mononucleotide Mitigates Age-Associated Physiological Decline in Mice. Cell Metabol. 2016, 24, 795–806. [Google Scholar] [CrossRef] [PubMed]
- Trammell, S.A.; Yu, L.; Redpath, P.; Migaud, M.E.; Brenner, C. Nicotinamide Riboside Is a Major NAD+ Precursor Vitamin in Cow Milk. J. Nutr. 2016, 146, 957–963. [Google Scholar] [CrossRef] [PubMed]
- Revollo, J.R.; Korner, A.; Mills, K.F.; Satoh, A.; Wang, T.; Garten, A.; Dasgupta, B.; Sasaki, Y.; Wolberger, C.; Townsend, R.R.; et al. Nampt/PBEF/Visfatin regulates insulin secretion in beta cells as a systemic NAD biosynthetic enzyme. Cell Metabol. 2007, 6, 363–375. [Google Scholar] [CrossRef] [PubMed]
- Romacho, T.; Azcutia, V.; Vazquez-Bella, M.; Matesanz, N.; Cercas, E.; Nevado, J.; Carraro, R.; Rodriguez-Manas, L.; Sanchez-Ferrer, C.F.; Peiro, C. Extracellular PBEF/NAMPT/visfatin activates pro-inflammatory signalling in human vascular smooth muscle cells through nicotinamide phosphoribosyltransferase activity. Diabetologia 2009, 52, 2455–2463. [Google Scholar] [CrossRef] [PubMed]
- Kim, U.H.; Kim, M.K.; Kim, J.S.; Han, M.K.; Park, B.H.; Kim, H.R. Purification and characterization of NAD glycohydrolase from rabbit erythrocytes. Arch. Biochem. Biophys. 1993, 305, 147–152. [Google Scholar] [CrossRef] [PubMed]
- O’Reilly, T.; Niven, D.F. Levels of nicotinamide adenine dinucleotide in extracellular body fluids of pigs may be growth-limiting for Actinobacillus pleuropneumoniae and Haemophilus parasuis. Canad. J. Vet. Res. Revue Canad. Recherche Vet. 2003, 67, 229–231. [Google Scholar]
- Adriouch, S.; Hubert, S.; Pechberty, S.; Koch-Nolte, F.; Haag, F.; Seman, M. NAD+ released during inflammation participates in T cell homeostasis by inducing ART2-mediated death of naive T cells in vivo. J. Immunol. 2007, 179, 186–194. [Google Scholar] [CrossRef]
- Bruzzone, S.; Guida, L.; Zocchi, E.; Franco, L.; De Flora, A. Connexin 43 hemi channels mediate Ca2+-regulated transmembrane NAD+ fluxes in intact cells. FASEB J. Off. Public. Fed. Am. Soc. Exp. Biol. 2001, 15, 10–12. [Google Scholar] [CrossRef]
- Bruzzone, S.; Franco, L.; Guida, L.; Zocchi, E.; Contini, P.; Bisso, A.; Usai, C.; De Flora, A. A self-restricted CD38-connexin 43 cross-talk affects NAD+ and cyclic ADP-ribose metabolism and regulates intracellular calcium in 3T3 fibroblasts. J. Biol. Chem. 2001, 276, 48300–48308. [Google Scholar] [CrossRef]
- Smyth, L.M.; Bobalova, J.; Mendoza, M.G.; Lew, C.; Mutafova-Yambolieva, V.N. Release of beta-nicotinamide adenine dinucleotide upon stimulation of postganglionic nerve terminals in blood vessels and urinary bladder. J. Biol. Chem. 2004, 279, 48893–48903. [Google Scholar] [CrossRef] [PubMed]
- Breen, L.T.; Smyth, L.M.; Yamboliev, I.A.; Mutafova-Yambolieva, V.N. beta-NAD is a novel nucleotide released on stimulation of nerve terminals in human urinary bladder detrusor muscle. Am. J. Physiol. Renal Physiol. 2006, 290, F486–F495. [Google Scholar] [CrossRef] [PubMed]
- Mutafova-Yambolieva, V.N.; Hwang, S.J.; Hao, X.; Chen, H.; Zhu, M.X.; Wood, J.D.; Ward, S.M.; Sanders, K.M. Beta-nicotinamide adenine dinucleotide is an inhibitory neurotransmitter in visceral smooth muscle. Proc. Natl. Acad. Sci. USA 2007, 104, 16359–16364. [Google Scholar] [CrossRef] [PubMed]
- Makarov, M.V.; Harris, N.W.; Rodrigues, M.; Migaud, M.E. Scalable syntheses of traceable ribosylated NAD(+) precursors. Organic Biomol. Chem. 2019, 17, 8716–8720. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kulikova, V.; Shabalin, K.; Nerinovski, K.; Yakimov, A.; Svetlova, M.; Solovjeva, L.; Kropotov, A.; Khodorkovskiy, M.; Migaud, M.E.; Ziegler, M.; et al. Degradation of Extracellular NAD+ Intermediates in Cultures of Human HEK293 Cells. Metabolites 2019, 9, 293. https://doi.org/10.3390/metabo9120293
Kulikova V, Shabalin K, Nerinovski K, Yakimov A, Svetlova M, Solovjeva L, Kropotov A, Khodorkovskiy M, Migaud ME, Ziegler M, et al. Degradation of Extracellular NAD+ Intermediates in Cultures of Human HEK293 Cells. Metabolites. 2019; 9(12):293. https://doi.org/10.3390/metabo9120293
Chicago/Turabian StyleKulikova, Veronika, Konstantin Shabalin, Kirill Nerinovski, Alexander Yakimov, Maria Svetlova, Ljudmila Solovjeva, Andrey Kropotov, Mikhail Khodorkovskiy, Marie E. Migaud, Mathias Ziegler, and et al. 2019. "Degradation of Extracellular NAD+ Intermediates in Cultures of Human HEK293 Cells" Metabolites 9, no. 12: 293. https://doi.org/10.3390/metabo9120293
APA StyleKulikova, V., Shabalin, K., Nerinovski, K., Yakimov, A., Svetlova, M., Solovjeva, L., Kropotov, A., Khodorkovskiy, M., Migaud, M. E., Ziegler, M., & Nikiforov, A. (2019). Degradation of Extracellular NAD+ Intermediates in Cultures of Human HEK293 Cells. Metabolites, 9(12), 293. https://doi.org/10.3390/metabo9120293





