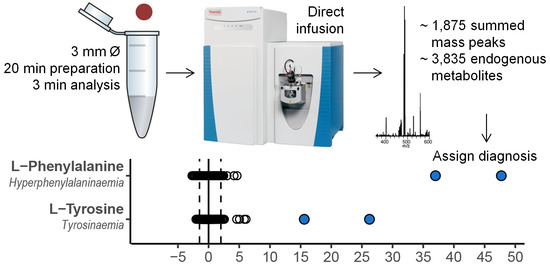
Direct Infusion Based Metabolomics Identifies Metabolic Disease in Patients’ Dried Blood Spots and Plasma
Abstract
1. Introduction
2. Results
2.1. Reproducibility Assessment
2.2. Evaluation of the Clinical Value of the Method
2.3. Direct-Infusion Based Metabolomics in Metabolic Diagnostics
3. Discussion
4. Methods
4.1. Sample Collection
4.2. Patient Inclusion and Sample Selection
4.3. Sample Preparation
4.4. DI-HRMS Analysis
4.5. Data Processing
4.6. Data Analysis
4.7. Evaluation of the Clinical Value of the Method
4.8. Reproducibility Assessment
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Coene, K.L.M.; Kluijtmans, L.A.J.; van der Heeft, E. Next-generation metabolic screening: Targeted and untargeted metabolomics for the diagnosis of inborn errors of metabolism in individual patients. J. Inherit. Metab. Dis. 2018, 41, 337–353. [Google Scholar] [CrossRef] [PubMed]
- Janeckova, H.; Hron, K.; Wojtowicz, P.; Hlídková, E.; Barešová, A.; Friedecký, D.; Zídková, L.; Hornik, P.; Behúlová, D.; Procházková, D.; et al. Targeted metabolomic analysis of plasma samples for the diagnosis of inherited metabolic disorders. J. Chromatogr. A 2012, 1226, 13–17. [Google Scholar] [CrossRef] [PubMed]
- Pitt, J.J.; Eggington, M.; Kahler, S.G. Comprehensive screening of urine samples for inborn errors of metabolism by electrospray tandem mass spectrometry. Clin. Chem. 2002, 48, 1970–1980. [Google Scholar] [PubMed]
- Jacob, M.; Malkawi, A.; Albast, N.; Al Bougha, S.; Lopata, A.; Dasouki, M.; Abdel Rahman, A.M. A targeted metabolomics approach for clinical diagnosis of inborn errors of metabolism. Analyt. Chim. Acta 2018, 1025, 141–153. [Google Scholar] [CrossRef] [PubMed]
- Kurko, J.; Tringham, M.; Tanner, L.; Näntö-Salonen, K.; Vähä-Mäkilä, M.; Nygren, H.; Pöhö, P.; Lietzen, N.; Mattila, I.; Olkku, A.; et al. Imbalance of plasma amino acids, metabolites and lipids in patients with lysinuric protein intolerance. Metabolism 2016, 65, 1361–1375. [Google Scholar] [CrossRef] [PubMed]
- Dercksen, M.; Koekemoer, G.; Duran, M.; Wanders, R.J.A.; Mienie, L.J.; Reinecke, C.J. Organic acid profile of isovaleric acidemia: A comprehensive metabolomics approach. Metabolomics 2013, 9, 765–777. [Google Scholar] [CrossRef]
- Smuts, I.; van der Westhuizen, F.H.; Louw, R.; Mienie, L.J.; Engelke, U.F.H.; Wevers, R.A.; Mason, S.; Koekemoer, G.; Reinecke, C.J. Disclosure of a putative biosignature for respiratory chain disorders through a metabolomics approach. Metabolomics 2013, 9, 379–391. [Google Scholar] [CrossRef]
- Najdekr, L.; Gardlo, A.; Madrova, L.; Friedecký, D.; Janečková, H.; Correa, E.S.; Goodacre, R.; Adam, T. Oxidized phosphatidylcholines suggest oxidative stress in patients with medium-chain acyl-CoA dehydrogenase deficiency. Talanta 2015, 139, 62–66. [Google Scholar] [CrossRef]
- Tebani, A.; Schmitz-Afonso, I.; Abily-Donval, L.; Héron, B.; Piraud, M.; Ausseil, J.; Brassier, A.; De Lonlay, P.; Zerimech, F.; Vaz, F.M.; et al. Urinary metabolic phenotyping of mucopolysaccharidosis type I combining untargeted and targeted strategies with data modeling. Clin. Chim. Acta 2017, 475, 7–14. [Google Scholar] [CrossRef]
- Wangler, M.F.; Huber, L.; Donti, T.R.; Ventura, M.J.; Miller, M.J.; Braverman, N.; Gawron, K.; Bose, M.; Moser, A.B.; Jones, R.O.; et al. A metabolomics map of Zellweger spectrum disorders reveals novel disease biomarkers. Genet. Med. 2018, 20, 1274–1283. [Google Scholar] [CrossRef]
- Wikoff, W.R.; Gangoiti, J.A.; Barshop, B.A.; Siuzdak, G. Metabolomics identifies perturbations in human disorders of propionate metabolism. Clin. Chem. 2007, 53, 2169–2176. [Google Scholar] [CrossRef] [PubMed]
- Denes, J.; Szabo, E.; Robinette, S.L.; Szatmári, I.; Szőnyi, L.; Kreuder, J.G.; Rauterberg, E.W.; Takáts, Z. Metabonomics of newborn screening dried blood spot samples: A novel approach in the screening and diagnostics of inborn errors of metabolism. Anal. Chem. 2012, 84, 10113–10120. [Google Scholar] [CrossRef] [PubMed]
- Peretz, H.; Watson, D.G.; Blackburn, G.; Zhang, T.; Lagziel, A.; Shtauber-Naamati, M.; Morad, T.; Keren-Tardai, E.; Greenshpun, V.; Usher, S.; et al. Urine metabolomics reveals novel physiologic functions of human aldehyde oxidase and provides biomarkers for typing xanthinuria. Metabolomics 2012, 8, 951–959. [Google Scholar] [CrossRef]
- Atwal, P.S.; Donti, T.R.; Cardon, A.L.; Bacino, C.A.; Sun, Q.; Emrick, L.; Reid Sutton, V.; Elsea, S.H. Aromatic L-amino acid decarboxylase deficiency diagnosed by clinical metabolomics profiling of plasma. Mol. Genet. Metab. 2015, 115, 91–94. [Google Scholar] [CrossRef] [PubMed]
- Venter, L.; Lindeque, Z.; Jansen van Rensburg, P.; van der Westhuizen, F.; Smuts, I.; Louw, R. Untargeted urine metabolomics reveals a biosignature for muscle respiratory chain deficiencies. Metabolomics 2015, 11, 111–121. [Google Scholar] [CrossRef]
- Abela, L.; Simmons, L.; Steindl, K.; Schmitt, B.; Mastrangelo, M.; Joset, P.; Papuc, M.; Sticht, H.; Baumer, A.; Crowther, L.M.; et al. N(8)-acetylspermidine as a potential plasma biomarker for Snyder-Robinson syndrome identified by clinical metabolomics. J. Inherit. Metab. Dis. 2016, 39, 131–137. [Google Scholar] [CrossRef] [PubMed]
- Donti, T.R.; Cappuccio, G.; Hubert, L.; Neira, J.; Atwal, P.S.; Miller, M.J.; Cardon, A.L.; Sutton, V.R.; Porter, B.E.; Baumer, F.M.; et al. Diagnosis of adenylosuccinate lyase deficiency by metabolomic profiling in plasma reveals a phenotypic spectrum. Mol. Genet. Metab. Rep. 2016, 8, 61–66. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.J.; Bostwick, B.L.; Kennedy, A.D.; Donti, T.R.; Sun, Q.; Sutton, V.R.; Elsea, S.H. Chronic oral L-carnitine supplementation drives marked plasma TMAO elevations in patients with organic acidemias despite dietary meat restrictions. JIMD Rep. 2016, 30, 39–44. [Google Scholar]
- Abela, L.; Spiegel, R.; Crowther, L.M.; Klein, A.; Steindl, K.; Papuc, S.M.; Joset, P.; Zehavi, Y.; Rauch, A.; Plecko, B.; et al. Plasma metabolomics reveals a diagnostic metabolic fingerprint for mitochondrial aconitase (ACO2) deficiency. PLoS ONE 2017, 12. [Google Scholar] [CrossRef]
- Glinton, K.E.; Benke, P.J.; Lines, M.A.; Geraghty, M.T.; Chakraborty, P.; Al-Dirbashi, O.Y.; Jiang, Y.; Kennedy, A.D.; Grotewiel, M.S.; Suttona, V.R.; et al. Disturbed phospholipid metabolism in serine biosynthesis defects revealed by metabolomic profiling. Mol. Genet. Metab. 2018, 123, 309–316. [Google Scholar] [CrossRef]
- Miller, M.J.; Kennedy, A.D.; Eckhart, A.D.; Burrage, L.C.; Wulff, J.E.; Miller, L.A.; Milburn, M.V.; Ryals, J.A.; Beaudet, A.L.; Sun, Q.; et al. Untargeted metabolomic analysis for the clinical screening of inborn errors of metabolism. J. Inherit. Metab. Dis. 2015, 38, 1029–1039. [Google Scholar] [CrossRef] [PubMed]
- de Sain-van der Velden, M.G.M.; van der Ham, M.; Gerrits, J.; Prinsen, H.C.M.T.; Willemsen, M.; Pras-Raves, M.L.; Jans, J.J.; Verhoeven-Duif, N.M. Quantification of metabolites in dried blood spots by direct infusion high resolution mass spectrometry. Anal. Chim. Acta 2017, 979, 45–50. [Google Scholar] [CrossRef] [PubMed]
- Kertesz, V.; Van Berkel, G.J. Fully automated liquid extraction-based surface sampling and ionization using a chip-based robotic nanoelectrospray platform. J. Mass Spectrom. 2010, 45, 252–260. [Google Scholar] [CrossRef] [PubMed]
- Kirwan, J.A.; Broadhurst, D.I.; Davidson, R.L.; Viant, M.R. Characterising and correcting batch variation in an automated direct infusion mass spectrometry (DIMS) metabolomics workflow. Anal. Bioanal. Chem. 2013, 405, 5147–5157. [Google Scholar] [CrossRef] [PubMed]
- González-Domínguez, R.; García-Barrera, T.; Gómez-Ariza, J.L. Using direct infusion mass spectrometry for serum metabolomics in Alzheimer’s disease. Anal. Bioanal. Chem. 2014, 406, 7137–7148. [Google Scholar] [CrossRef] [PubMed]
- Lokhov, P.G.; Trifonova, O.P.; Maslov, D.L.; Balashova, E.E.; Archakov, A.I.; Shestakova, E.A.; Shestakova, M.V.; Dedov, I.I. Diagnosing impaired glucose intolerance using direct infusion mass spectrometry of blood plasma. PLoS ONE 2014, 9, e105343. [Google Scholar] [CrossRef] [PubMed]
- Koulman, A.; Prentice, P.; Wong, M.C.Y.; Matthews, L.; Bond, N.J.; Eiden, M.; Griffin, J.L.; Dunger, D.B. The development and validation of a fast and robust dried blood spot based lipid profiling method to study infant metabolism. Metabolomics 2014, 10, 1018–1025. [Google Scholar] [CrossRef]
- Anand, S.; Barnes, J.M.; Young, S.A.; Garcia, D.M.; Tolley, H.D.; Kauwe, J.S.K.; Graves, S.W. Discovery and confirmation of diagnostic serum lipid biomarkers for Alzheimer’s disease using direct infusion mass spectrometry. J. Alzheimers Dis. 2017, 59, 277–290. [Google Scholar] [CrossRef]
- Anand, S.; Young, S.; Esplin, M.S.; Peaden, B.; Tolley, H.D.; Porter, T.F.; Varner, M.W.; D’Alton, M.E.; Jackson, B.J.; Graves, S.W. Detection and confirmation of serum lipid biomarkers for preeclampsia using direct infusion mass spectrometry. J. Lipid Res. 2017, 57, 687–696. [Google Scholar] [CrossRef]
- Ramos, R.J.; Pras-Raves, M.L.; Gerrits, J.; van der Ham, M.; Willemsen, M.; Prinsen, H.; Burgering, B.; Jans, J.J.; Verhoeven-Duif, N.M. Vitamin B6 is essential for serine de novo biosynthesis. J. Inherit. Metab. Dis. 2017, 40, 883–891. [Google Scholar] [CrossRef]
- Sain-van der Velden, M.G.M.; Diekman, E.F.; Jans, J.J.; van der Ham, M.; Prinsen, B.H.; Visser, G.; Verhoeven-Duif, N.M. Differences between acylcarnitine profiles in plasma and bloodspots. Mol. Genet. Metabol. 2013, 110, 116–121. [Google Scholar] [CrossRef] [PubMed]
- Primassin, S.; Spiekerkoetter, U. ESI-MS/MS measurement of free carnitine and its precursor γ-butyrobetaine in plasma and dried blood spots from patients with organic acidurias and fatty acid oxidation disorders. Mol. Genet. Metabol. 2010, 101, 145–150. [Google Scholar] [CrossRef] [PubMed]
- Heiner-Fokkema, M.R.; Vaz, F.M.; Maatman, R.; Kluijtmans, L.A.J.; van Spronsen, F.J.; Reijngoud, D.J. Reliable diagnosis of carnitine palmitoyltransferase type IA deficiency by analysis of plasma acylcarnitine profiles. JIMD Rep. 2016, 32, 33–39. [Google Scholar] [CrossRef] [PubMed]
- Wishart, D.S.; Jewison, T.; Guo, A.C.; Wilson, M.; Knox, C.; Liu, Y.; Djoumbou, Y.; Mandal, R.; Aziat, F.; Dong, E.; et al. HMDB 3.0—The Human Metabolome Database in 2013. Nucleic Acids Res. 2013, 41, 801–807. [Google Scholar] [CrossRef] [PubMed]
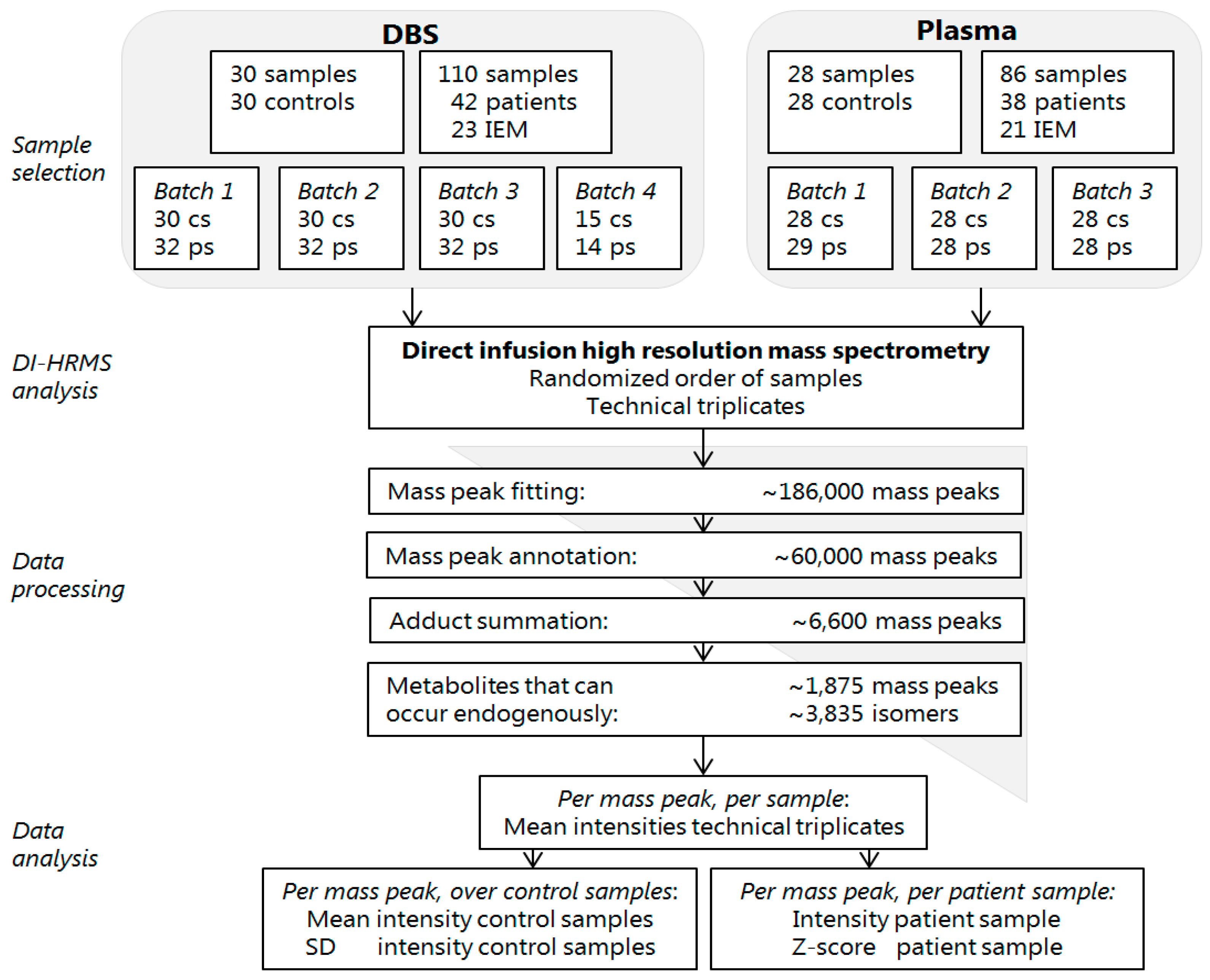

| DBS | Plasma | ||||||
|---|---|---|---|---|---|---|---|
| Batch 1 | Batch 2 | Batch 3 | Batch 4 | Batch 1 | Batch 2 | Batch 3 | |
| Mass peak fitting | 185,661 | 176,934 | 197,681 | 190,172 | 192,198 | 177,879 | 185,642 |
| Mass peak annotation | 59,543 | 56,250 | 63,360 | 60,979 | 62,503 | 58,212 | 60,450 |
| Adduct summation | 6580 | 6625 | 6598 | 6611 | |||
| Endogenous mass peaks * | 1874 | 1885 | 1874 | 1875 | 1875 | 1867 | 1868 |
| Endogenous metabolite annotations * | 3822 | 3863 | 3826 | 3839 | 3832 | 3847 | 3817 |
| DBS | Plasma | ||||||
|---|---|---|---|---|---|---|---|
| Batch 1 | Batch 2 | Batch 3 | Batch 4 | Batch 1 | Batch 2 | Batch 3 | |
| 15N;2−13C-glycine | 0.23 | 0.16 | 0.18 | 0.24 | 0.22 | 0.21 | 0.79 |
| 2H4-alanine | 0.20 | 0.14 | 0.16 | 0.20 | 0.20 | 0.21 | 0.19 |
| 2H3-leucine | 0.18 | 0.14 | 0.15 | 0.18 | 0.60 | 0.55 | 0.50 |
| 2H3-methionine | 0.31 | 0.30 | 0.36 | 0.39 | 1.70 | 0.22 | 0.20 |
| 13C6-phenylalanine | 0.19 | 0.16 | 0.14 | 0.18 | 0.21 | 0.20 | 0.19 |
| 13C6-tyrosine | 0.19 | 0.17 | 0.16 | 0.20 | 0.22 | 0.21 | 0.18 |
| 2H3-aspartate | 0.24 | 0.22 | 0.22 | 0.25 | 0.23 | 0.24 | 0.26 |
| 2H3-glutamate | 0.17 | 0.15 | 0.14 | 0.18 | 0.20 | 0.21 | 0.15 |
| 2H2-ornithine | 0.21 | 0.19 | 0.17 | 0.21 | 0.14 | 0.17 | 0.12 |
| 2H2-citrulline | 0.16 | 0.16 | 0.14 | 0.18 | 0.18 | 0.19 | 0.14 |
| 2H4;13C-arginine | 0.21 | 0.17 | 0.16 | 0.20 | 0.17 | 0.18 | 0.16 |
| 2H8-valine | 0.18 | 0.14 | 0.15 | 0.18 | 0.20 | 0.19 | 0.18 |
| 2H9-carnitine | 0.27 | 0.21 | 0.22 | 0.30 | 0.22 | 0.24 | 0.21 |
| 2H3-acetylcarnitine | 0.89 | 0.21 | 0.82 | 0.92 | 0.46 | 0.46 | 0.74 |
| 2H3-propionylcarnitine | 0.21 | 0.16 | 0.16 | 0.20 | 0.19 | 0.20 | 0.20 |
| 2H3-butyrylcarnitine | 3.39 | 0.63 | 1.34 | 1.53 | 0.77 | 0.92 | 1.08 |
| 2H9-isovalerylcarnitine | 0.20 | 0.13 | 0.15 | 0.17 | 0.19 | 0.20 | 0.19 |
| 2H3-octanoylcarnitine | 0.18 | 0.12 | 0.14 | 0.17 | 0.16 | 0.21 | 0.20 |
| 2H9-myristoylcarnitine | 0.20 | 0.14 | 0.14 | 0.17 | 0.20 | 0.22 | 0.20 |
| 2H3-palmitoylcarnitne | 0.19 | 0.16 | 0.15 | 0.18 | 0.23 | 0.23 | 0.21 |
| 5th percentile | 0.16 | 0.13 | 0.14 | 0.17 | 0.16 | 0.18 | 0.14 |
| Median | 0.20 | 0.16 | 0.16 | 0.20 | 0.21 | 0.21 | 0.20 |
| 95th percentile | 2.96 | 0.30 | 0.79 | 0.89 | 0.76 | 0.55 | 0.79 |
| Batch 1 | Batch 2 | Batch 3 | Batch 4 | Batch 5 | Batch 6 | Batch 7 | RSD | |
|---|---|---|---|---|---|---|---|---|
| Propionic aciduria | ||||||||
| Propionylcarnitine | 40.23 | 66.57 | 47.17 | 70.07 | 61.18 | 52.29 | 52.66 | 0.19 |
| Glycine | 12.99 | 20.75 | 17.28 | 16.42 | 23.48 | 24.54 | 10.12 | 0.30 |
| Propionylglycine | 7.89 | 7.69 | 9.26 | 7.54 | 12.69 | 9.09 | 6.10 | 0.24 |
| Lysinuric protein intolerance | ||||||||
| Citrulline | 23.86 | 26.55 | 18.98 | 29.23 | 24.10 | 30.03 | 22.51 | 0.15 |
| Glutamine | 3.32 | 3.40 | 3.56 | 4.68 | 3.39 | 4.81 | 2.07 | 0.26 |
| Lysine | −2.07 | −2.13 | −1.89 | −2.07 | −2.25 | −1.97 | −1.71 | 0.09 |
| Phenylketonuria | ||||||||
| Phenylalanine | 34.29 | 17.93 | 16.79 | 23.13 | 21.74 | 16.21 | 14.19 | 0.33 |
| DBS #1 | DBS #2 | Plasma #1 | Plasma #2 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Patient Diagnosis | Metabolite * | Z-sc. | Rank | Correct Diagn. | Z-sc. | Rank | Correct Diagn. | Z-sc. | Rank | Correct Diagn. | Z-sc. | Rank | Correct Diagn. | |
| Urea cycle | OTC deficiency | Orotic acid | 5.7 | 1 | Yes (n = 2) | −0.5 | No (n = 3) | 11.7 | 2 | Yes (n = 2) | 2.3 | Yes (n = 1) | ||
| Uridine | 1.6 | −0.5 | 7.1 | 39.2 | 6 | |||||||||
| 5-Oxoproline | −0.7 | 0.2 | 0.4 | 9.0 | ||||||||||
| Uracil | −0.8 | −0.7 | 4.0 | 4.0 | ||||||||||
| Orotidine | 0.1 | −0.5 | −1.1 | 4.0 | ||||||||||
| L-Lysine | 0.0 | −0.2 | 0.3 | 3.3 | ||||||||||
| Citrulline | −0.3 | −1.8 | −7 | −0.6 | −2.8 | −7 | ||||||||
| Branched-chain amino acid metabolism | MSUD | Ketoleucine | 23.3 | 2 | Yes (n = 4) | 3.0 | 20 | Yes (n = 3) | 65.0 | 7 | Yes (n = 4) | 13.3 | 17 | Yes (n = 3) |
| (n = Iso) leucine | 12.4 | 6 | 0.4 | 37.4 | 10 | 24.7 | 7 | |||||||
| 2-Hydroxy-3-methylbutyr. acid | 9.4 | 8 | −0.4 | 579.1 | 1 | 234.8 | 1 | |||||||
| Alpha-ketoisovaleric acid | 4.8 | 2.2 | 39.5 | 9 | 21.2 | 9 | ||||||||
| IVA | Isovalerylcarnitine | 137.9 | 1 | Yes (n = 2) | 42.5 | 2 | Yes (n = 2) | 84.7 | 1 | Yes (n = 1) | 92.5 | 1 | Yes (n = 3) | |
| 3-Hydroxyisovaleric acid | 0.0 | −0.1 | 0.4 | 0.1 | ||||||||||
| 3-MCC | 3-Hydroxyisovaleric acid | 5.4 | 1 | Yes (n = 2) | 33.1 | 1 | Yes (n = 2) | 17.8 | 4 | No (n = 1) | 825.8 | 1 | Yes (n = 2) | |
| 3-Methylcrotonylglycine | 0.7 | 22.8 | 2 | −0.1 | 2.7 | |||||||||
| Isovalerylcarnitine | 0.6 | −1.2 | 7.0 | 11 | 0.7 | |||||||||
| 3-Hydroxyisovalerylcarnitine | 0.6 | −1.6 | −0.2 | 0.0 | ||||||||||
| MMA | Propionylcarnitine | 13.3 | 2 | Yes (n = 3) | 75.4 | 1 | Yes (n = 1) | |||||||
| Methylcitric acid | 7.3 | 4 | 4.3 | |||||||||||
| Methylmalonic acid | 0.2 | 16.6 | 4 | |||||||||||
| Methylmalonylcarnitine | 1.1 | 0.7 | ||||||||||||
| Lysine metabolism | GA-1 | Glutarylcarnitine | 18.6 | 2 | Yes (n = 4) | 4.9 | 10 | Yes (n = 2) | 26.3 | 1 | Yes (n = 3) | 27.9 | 5 | Yes (n = 3) |
| Glutaric acid | 7.9 | 3 | −0.9 | 6.4 | 17 | 71.6 | 3 | |||||||
| 3-hydroxyglutaric acid | −0.3 | 0.2 | 10.5 | 8 | 8.2 | 11 | ||||||||
| Glutaconic acid | −1.64 | 27.9 | 2 | |||||||||||
| Phenylalanine and tyrosine metabolism | PKU | Phenylalanine | 47.7 | 1 | Yes (n = 1) | 37.0 | 3 | Yes (n = 1) | ||||||
| Hydroxyphenylacetic acid | 10.9 | 4 | 1.9 | |||||||||||
| N-acetylphenylalanine | 6.3 | 9 | 7.0 | 22 | ||||||||||
| Tyrosine | −1.0 | −0.1 | ||||||||||||
| Tyrosinaemia | 4-Hydroxyphenyllactic acid | 150.7 | 1 | Yes (n = 1) | 125.6 | 2 | Yes (n = 1) | 206.5 | 1 | Yes (n = 3) | 263.5 | 13 | Yes (n = 3) | |
| Tyrosine | 26.2 | 3 | 15.6 | 6 | 35.0 | 3 | 33.7 | |||||||
| 4-Hydroxyphenylacetic acid | 4.6 | 6.3 | 9 | 2.2 | 2.0 | |||||||||
| 4-Hydroxyphenylpyruvic acid | 0.2 | 2.0 | 10.4 | 8 | 6.8 | |||||||||
| Succinylacetone | −1.5 | −1.2 | 0.2 | 1.1 | ||||||||||
| Sulphur amino acid metabolism | MAT1A deficiency | Methionine sulfoxide | 72.2 | 1 | Yes (n = 5) | 53.4 | 2 | Yes (n = 5) | 1106.7 | 1 | Yes (n = 1) | 632.2 | 1 | Yes (n = 3) |
| Methionine | 57.1 | 2 | 96.4 | 1 | 118.8 | 4 | 47.4 | 6 | ||||||
| S-adenosylmethionine | 0.5 | −0.3 | 0.1 | 0.3 | ||||||||||
| S-adenosylhomocysteine | −0.8 | 0.4 | 0.4 | 0.1 | ||||||||||
| CBS deficiency | Methionine sulfoxide | 22.4 | 2 | Yes (n = 4) | 778.9 | 1 | Yes (n = 2) | |||||||
| Methionine | 31.1 | 3 | 2.6 | |||||||||||
| Homocystine | 3.2 | 7 | 1.3 | |||||||||||
| Homocysteine | 2.6 | 12 | 2.2 | |||||||||||
| MTHFR deficiency | Homocysteine thiolactone | 28.0 | 1 | Yes (n = 6) | 7.5 | 3 | Yes | −0.3 | No (n = 1) | 0.1 | No (n = 3) | |||
| Homocystine | 1.1 | 4.8 | 9 | (n = 3) | −0.2 | 0.3 | ||||||||
| Methionine | 0.2 | 0.0 | −2.4 | −20 | −2.3 | −12 | ||||||||
| Molybdenum cofactor deficiency | Xanthine | 59.3 | 1 | Yes (n = 1) | 40.7 | 3 | Yes | 55.5 | 7 | Yes (n = 1) | ||||
| Alpha amino adipic semialdeh. | 3.4 | 1.5 | (n = 1) | 6.9 | ||||||||||
| Cysteine-S-sulfate | −0.9 | 0.6 | 11.8 | 22 | ||||||||||
| Cysteine | −1.0 | −2.6 | −2.1 | −14 | ||||||||||
| Uric acid | −1.4 | −0.8 | −2.6 | −5 | ||||||||||
| Serine and glycine metabolism | NKH | Glycine | 3.7 | 18 | Yes (n = 2) | 2.0 | No (n = 3) | 3.4 | Yes (n = 3) | 2.2 | No (n = 3) | |||
| 3-PGDH deficiency | Serine | 5.1 | 1 | No (n = 3) | 0.8 | No | −2.5 | −4 | Yes (n = 2) | −2.4 | −6 | Yes (n = 2) | ||
| Glycine | 2.1 | −0.1 | (n = 3) | −1.6 | −1.8 | |||||||||
| Proline metabolism | OAT deficiency | Proline | 4.0 | 11 | Yes (n = 6) | 4.0 | Yes (n = 5) | |||||||
| Ornithine | 2.8 | 18 | −0.8 | |||||||||||
| Amino acid transport | LPI | Citrulline | 8.5 | 2 | Yes (n = 3) | 16.1 | 13 | Yes (n = 3) | ||||||
| Serine | 6.2 | 3 | 2.4 | |||||||||||
| Proline | 6.4 | 4 | 0.2 | |||||||||||
| Threonine | 5.7 | 7 | 0.6 | |||||||||||
| Lysine | −2.0 | −7 | −1.3 | |||||||||||
| Ornithine | −1.5 | 0.6 | ||||||||||||
| Arginine | −1.0 | −1.0 | ||||||||||||
| Fatty acid oxidation | VLCAD deficiency | C14:1 carnitine | 28.9 | 1 | Yes (n = 1) | 0.6 | No (n = 3) | 7.3 | 34 | Yes (n = 1) | 5.8 | Yes (n = 1) | ||
| C14:2 carnitine | 15.7 | 2 | 1.4 | 7.6 | 33 | 2.8 | ||||||||
| C14-carnitine | 3.7 | 1.5 | 1.4 | 2.4 | ||||||||||
| LCHAD deficiency | C14-OH carnitine | 3.1 | 35 | Yes (n = 1) | 8.3 | 14 | Yes (n = 2) | 8.2 | Yes (n = 2) | |||||
| C16-OH carnitine | 3.0 | 37 | 22.7 | 2 | 37.3 | 12 | ||||||||
| C16-OH:1 carnitine | 1.5 | 23.8 | 1 | 41.6 | 11 | |||||||||
| C18-OH carnitine | 0.7 | 21.9 | 3 | 29.8 | 17 | |||||||||
| MCAD deficiency | C8-carnitine | 56.5 | 1 | Yes (n = 2) | 111.5 | 1 | Yes (n = 3) | 189.3 | 1 | Yes (n = 3) | 143.4 | 1 | Yes (n = 2) | |
| C6-carnitine | 7.3 | 6 | 16.0 | 3 | 51.7 | 2 | 55.7 | 2 | ||||||
| C10:1-carnitine | 1.7 | 8.1 | 7 | 24.9 | 4 | 11.6 | 5 | |||||||
| C10-carnitine | 1.1 | 2.6 | 7.3 | 12 | 3.2 | |||||||||
| OCTN2 deficiency | L-Carnitine | −2.0 | Yes (n = 1) | −1.3 | Yes (n = 4) | −2.4 | −3 | Yes (n = 2) | −2.3 | −6 | Yes (n = 1) | |||
| Acetylcarnitine | −1.9 | −0.9 | −2.5 | −1 | −2.5 | −9 | ||||||||
| C16-carnitine | −1.7 | −1.3 | −1.1 | −0.3 | ||||||||||
| C16:1-carnitineC18-carnitine | −2.6–1.7 | −5 | −1.1–1.7 | −2 | −1.3–0.6 | −1.8–0.9 | ||||||||
| C18:1-carnitine | −2.3 | −12 | −1.8 | −1 | −1.1 | −1.0 | ||||||||
| CPT1 deficiency | L-Carnitine | 19.0 | 1 | Yes (n = 2) | 19.0 | 1 | Yes (n = 6) | −2.7 | −84 | No (n = 2) | 1.8 | No (n = 2) | ||
| C0/(n = C16 + C18) ratio | 10.3 | 3 | 8.4 | 3 | −1.6 | −0.3 | ||||||||
| C16-carnitine | −3.1 | −1 | −1.8 | −5 | −2.7 | −82 | −0.2 | |||||||
| C18-carnitine | −2.6 | −3 | −2.2 | −2 | −1.1 | −0.6 | ||||||||
| C18:1-carnitine | −2.6 | −4 | −2.5 | −1 | 0.0 | 1.1 | ||||||||
| CPT2 deficiency | C16+C18:1/C2 ratio | 2.2 | 25 | Yes (n = 2) | 4.8 | 1 | Yes (n = 3) | −1.4 | Yes (n = 4) | 0.1 | Yes (n = 2) | |||
| Acetylcarnitine | −1.7 | −8 | −2.4 | −1 | 8.8 | 9 | 6.5 | 6 | ||||||
| C16-carnitine | −0.6 | −1.4 | 9.3 | 8 | 6.7 | 5 | ||||||||
| C18-carnitine | −0.6 | −1.7 | 4.1 | 3.1 | ||||||||||
| C18:1-carnitine | −0.7 | −1.8 | ||||||||||||
| Creatine biosynthesis | GAMT deficiency | Guanidoacetic acid | 20.9 | 1 | Yes (n = 2) | 39.2 | 2 | Yes (n = 1) | 25.1 | 1 | Yes (n = 3) | 35.9 | 1 | Yes (n = 1) |
| Creatine | −1.4 | −1.2 | 1.8 | −1.7 | ||||||||||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Haijes, H.A.; Willemsen, M.; Van der Ham, M.; Gerrits, J.; Pras-Raves, M.L.; Prinsen, H.C.M.T.; Van Hasselt, P.M.; De Sain-van der Velden, M.G.M.; Verhoeven-Duif, N.M.; Jans, J.J.M. Direct Infusion Based Metabolomics Identifies Metabolic Disease in Patients’ Dried Blood Spots and Plasma. Metabolites 2019, 9, 12. https://doi.org/10.3390/metabo9010012
Haijes HA, Willemsen M, Van der Ham M, Gerrits J, Pras-Raves ML, Prinsen HCMT, Van Hasselt PM, De Sain-van der Velden MGM, Verhoeven-Duif NM, Jans JJM. Direct Infusion Based Metabolomics Identifies Metabolic Disease in Patients’ Dried Blood Spots and Plasma. Metabolites. 2019; 9(1):12. https://doi.org/10.3390/metabo9010012
Chicago/Turabian StyleHaijes, Hanneke A., Marcel Willemsen, Maria Van der Ham, Johan Gerrits, Mia L. Pras-Raves, Hubertus C. M. T. Prinsen, Peter M. Van Hasselt, Monique G. M. De Sain-van der Velden, Nanda M. Verhoeven-Duif, and Judith J. M. Jans. 2019. "Direct Infusion Based Metabolomics Identifies Metabolic Disease in Patients’ Dried Blood Spots and Plasma" Metabolites 9, no. 1: 12. https://doi.org/10.3390/metabo9010012
APA StyleHaijes, H. A., Willemsen, M., Van der Ham, M., Gerrits, J., Pras-Raves, M. L., Prinsen, H. C. M. T., Van Hasselt, P. M., De Sain-van der Velden, M. G. M., Verhoeven-Duif, N. M., & Jans, J. J. M. (2019). Direct Infusion Based Metabolomics Identifies Metabolic Disease in Patients’ Dried Blood Spots and Plasma. Metabolites, 9(1), 12. https://doi.org/10.3390/metabo9010012




_Verhoeven-Duif.png)