Exometabolomic Analysis of Cross-Feeding Metabolites
Abstract
1. Introduction
2. Materials and Methods
2.1. Chemicals
2.2. Culture Methods
2.2.1. Sugar Substrate Preference of C. Fimi and Y. Lipolytica
2.2.2. Substrate Preference of C. Fimi in a Mixture of Sugars
2.2.3. Co-culture of C. Fimi and Y. Lipolytica Cultured with Cellulose and Galactomannan
2.2.4. Co-Culture of C. Fimi and Y. Lipolytica Cultured with Ionic-Liquids Treated Switchgrass
2.3. GC-MS Analysis
2.3.1. Sample Preparation
2.3.2. Data Acquisition
2.3.3. Targeted Sugar Quantitation
2.3.4. GC-MS Untargeted Metabolite Analysis
3. Results and Discussion
3.1. Sugar Substrate Preference of C. Fimi and Y. Lipolytica
3.2. Substrate Preference of C. Fimi in a Mixture of Sugars
3.3. Co-Culture of C. Fimi and Y. Lipolytica Cultured with Cellulose and Galactomannan
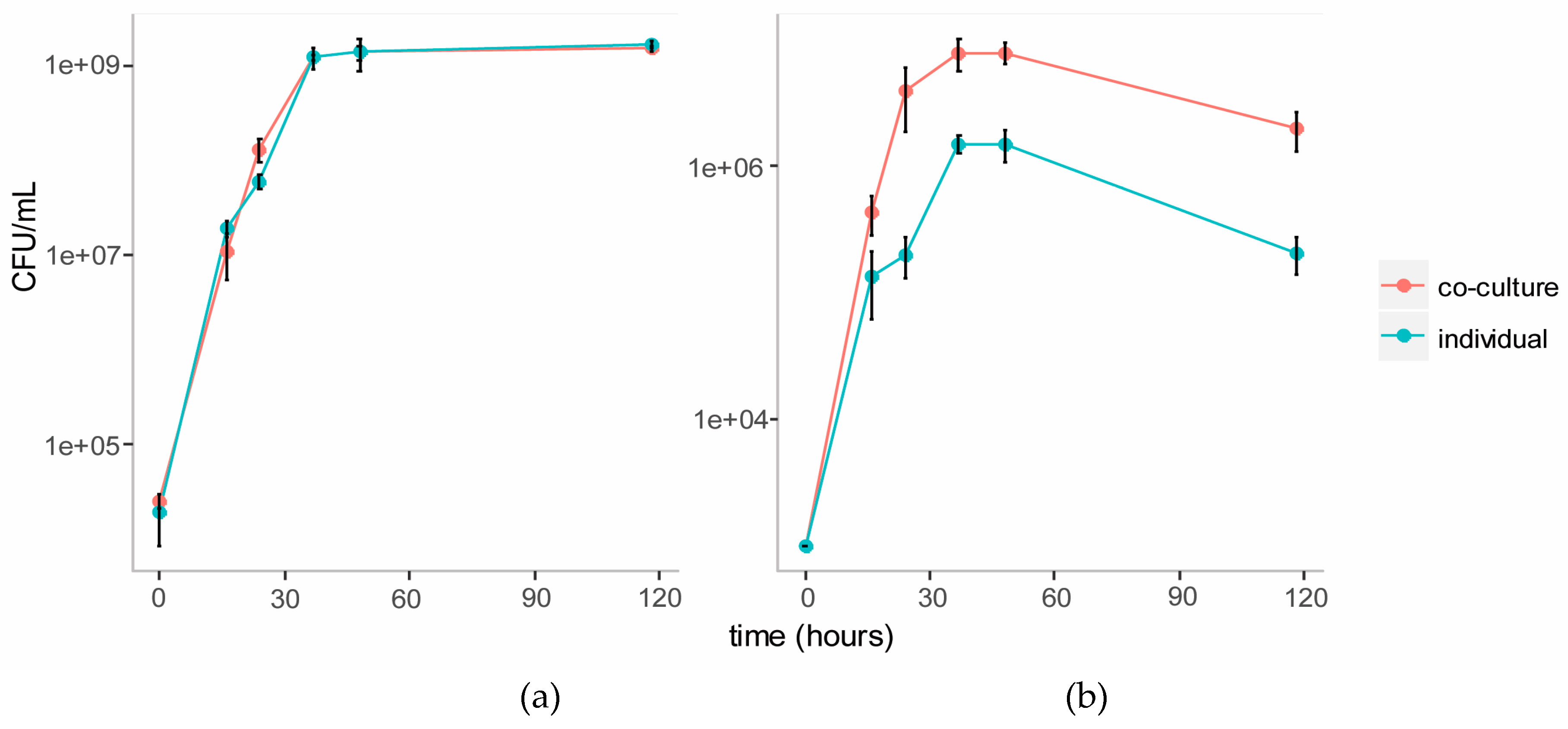
3.3.1. Growth Curves on Cellulose Glucomannan Substrates
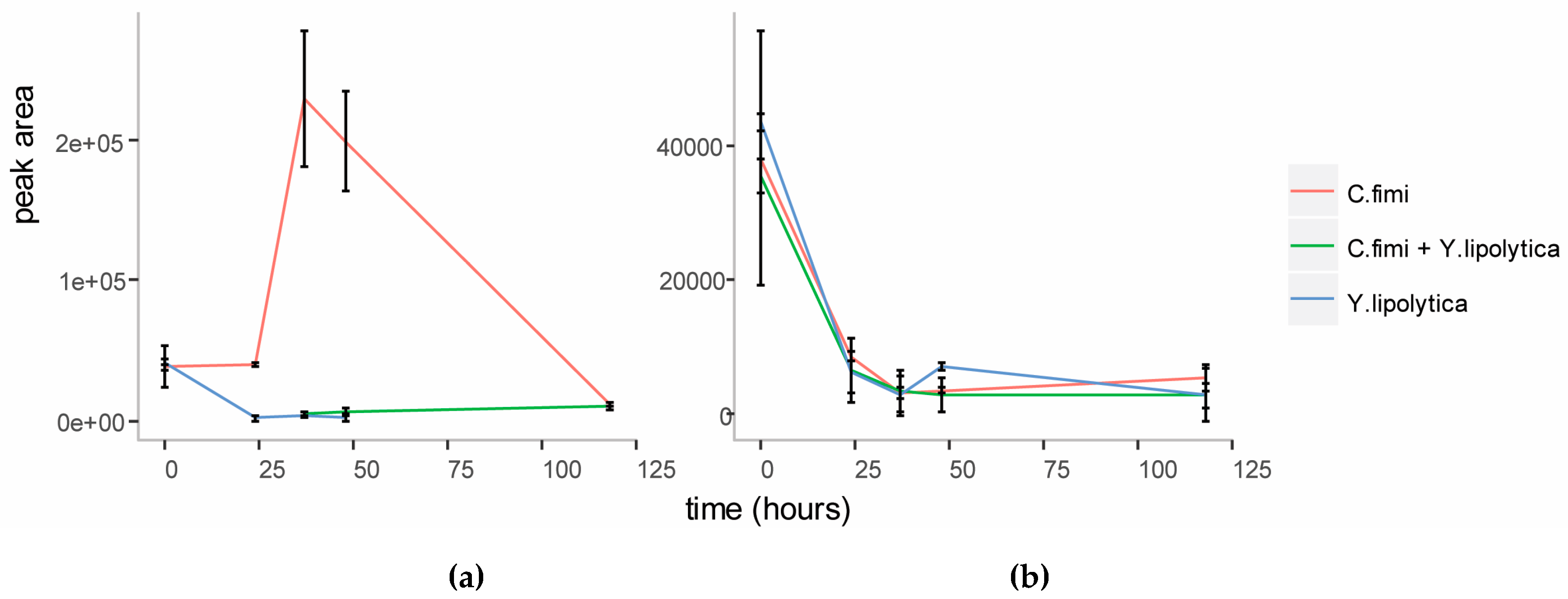
3.3.2. Targeted Sugar Analysis from Growth on Cellulose Glucomannan Substrates
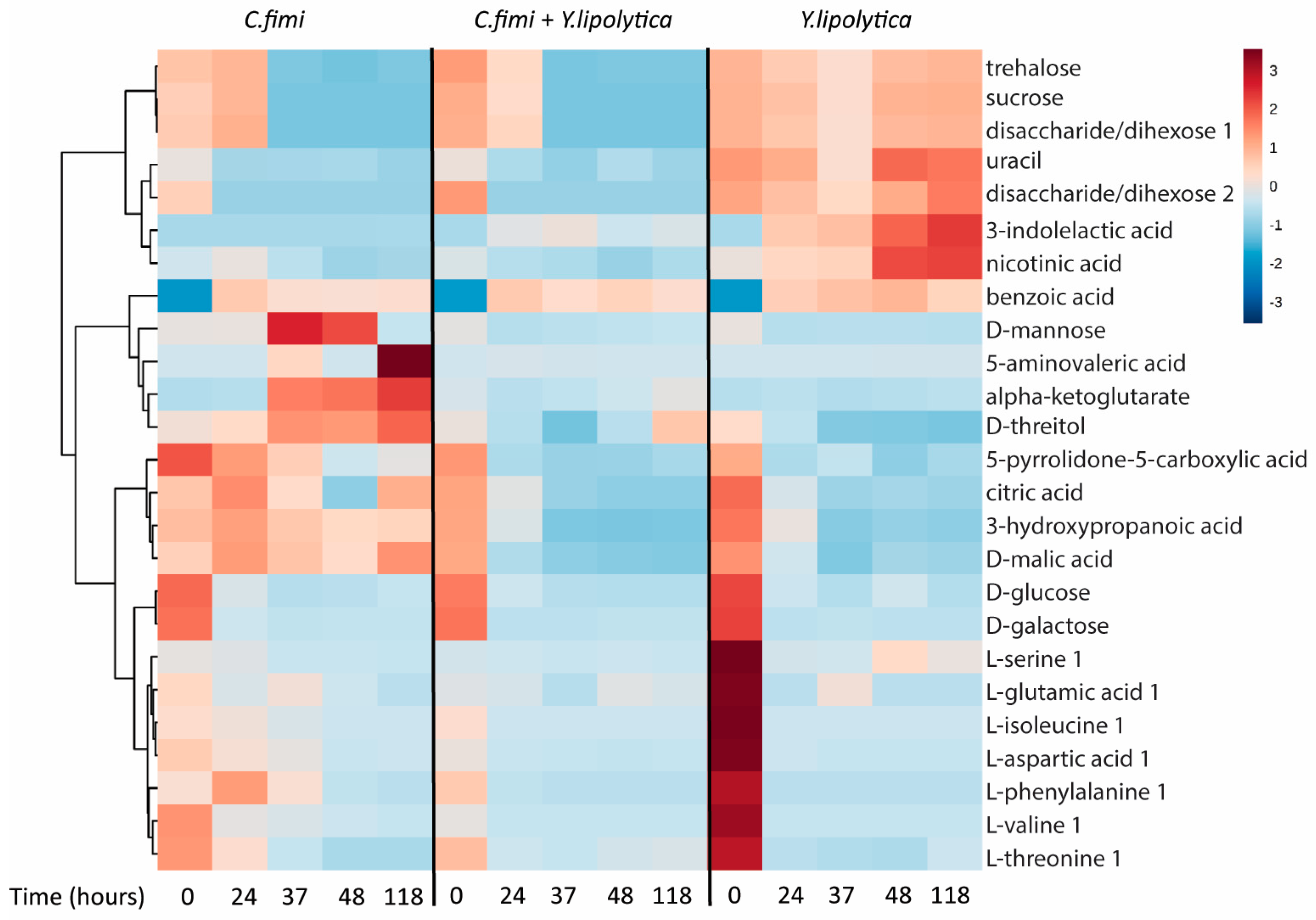
3.3.3. Untargeted Metabolite Analysis on Cellulose Galactomannan Substrates
3.4. Co-Culture of C. Fimi and Y. Lipolytica Cultured With Ionic Liquid Treated Switchgrass
3.4.1. Growth Curves on Ionic Liquid Treated Switchgrass
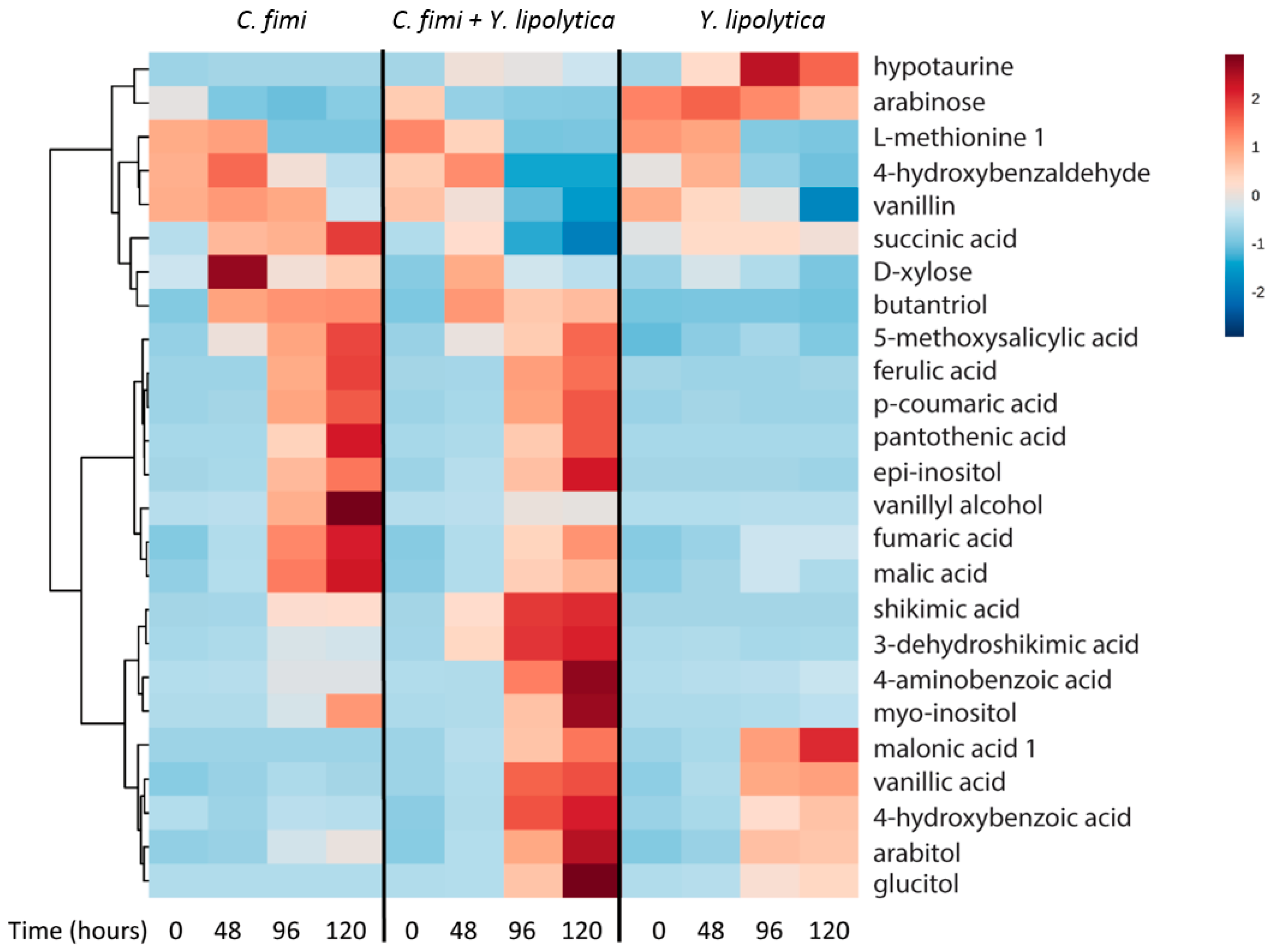
3.4.2. Untargeted Metabolite Analysis from Ionic Liquid-Treated Switchgrass
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Johns, N.I.; Blazejewski, T.; Gomes, A.L.; Wang, H.H. Principles for designing synthetic microbial communities. Curr. Opin. Microbiol. 2016, 31, 146–153. [Google Scholar] [CrossRef] [PubMed]
- Zuroff, T.R.; Curtis, W.R. Developing symbiotic consortia for lignocellulosic biofuel production. Appl. Microbiol. Biotechnol. 2012, 93, 1423–1435. [Google Scholar] [CrossRef] [PubMed]
- Hahn-Hägerdal, B.; Häggström, M. Production of ethanol from cellulose, Solka Floc BW 200, in a fedbatch mixed culture of Trichoderma reesei, C 30, and Saccharomyces cerevisiae. Appl. Microbiol. Biotechnol. 1985, 22, 187–189. [Google Scholar] [CrossRef]
- Minty, J.J.; Singer, M.E.; Scholz, S.A.; Bae, C.-H.; Ahn, J.-H.; Foster, C.E.; Liao, J.C.; Lin, X.N. Design and characterization of synthetic fungal-bacterial consortia for direct production of isobutanol from cellulosic biomass. Proc. Natl. Acad. Sci. 2013, 110, 14592–14597. [Google Scholar] [CrossRef] [PubMed]
- Bayer, T.S.; Widmaier, D.M.; Temme, K.; Mirsky, E.A.; Santi, D.V.; Voigt, C.A. Synthesis of methyl halides from biomass using engineered microbes. J. Am. Chem. Soc. 2009, 131, 6508–6515. [Google Scholar] [CrossRef] [PubMed]
- Zelezniak, A.; Andrejev, S.; Ponomarova, O.; Mende, D.R.; Bork, P.; Patil, K.R. Metabolic dependencies drive species co-occurrence in diverse microbial communities. Proc. Natl. Acad. Sci. 2015, 112, 6449–6454. [Google Scholar] [CrossRef] [PubMed]
- Silva, L.P.; Northen, T.R. Exometabolomics and MSI: Deconstructing how cells interact to transform their small molecule environment. Curr. Opin. Biotechnol. 2015, 34, 209–216. [Google Scholar] [CrossRef] [PubMed]
- Allen, J.; Davey, H.M.; Broadhurst, D.; Heald, J.K.; Rowland, J.J.; Oliver, S.G.; Kell, D.B. High-throughput classification of yeast mutants for functional genomics using metabolic footprinting. Nat. Biotechnol. 2003, 21, 692–696. [Google Scholar] [CrossRef] [PubMed]
- Kell, D.B.; Brown, M.; Davey, H.M.; Dunn, W.B.; Spasic, I.; Oliver, S.G. Metabolic footprinting and systems biology: The medium is the message. Nat. Rev. Microbiol. 2005, 3, 557–565. [Google Scholar] [CrossRef] [PubMed]
- Baran, R.; Brodie, E.L.; Mayberry-Lewis, J.; Hummel, E.; Da Rocha, U.N.; Chakraborty, R.; Bowen, B.P.; Karaoz, U.; Cadillo-Quiroz, H.; Garcia-Pichel, F.; et al. Exometabolite niche partitioning among sympatric soil bacteria. Nat. Commun. 2015, 6, 8289. [Google Scholar] [CrossRef] [PubMed]
- Kosina, S.M.; Danielewicz, M.A.; Mohammed, M.; Ray, J.; Suh, Y.; Yilmaz, S.; Singh, A.K.; Arkin, A.P.; Deutschbauer, A.M.; Northen, T.R. Exometabolomics assisted design and validation of synthetic obligate mutualism. ACS Synth. Biol. 2015, 5, 569–576. [Google Scholar] [CrossRef] [PubMed]
- Görke, B.; Stülke, J. Carbon catabolite repression in bacteria: Many ways to make the most out of nutrients. Nat. Rev. Microbiol. 2008, 6, 613–624. [Google Scholar] [CrossRef] [PubMed]
- Christopherson, M.R.; Suen, G.; Bramhacharya, S.; Jewell, K.A.; Aylward, F.O.; Mead, D.; Brumm, P.J. The genome sequences of cellulomonas fimi and “cellvibrio gilvus” reveal the cellulolytic strategies of two facultative anaerobes, transfer of “cellvibrio gilvus” to the genus cellulomonas, and proposal of cellulomonas gilvus sp. nov. PLoS ONE 2013, 8, e53954. [Google Scholar] [CrossRef] [PubMed]
- Beopoulos, A.; Chardot, T.; Nicaud, J.-M. Yarrowia lipolytica: A model and a tool to understand the mechanisms implicated in lipid accumulation. Biochimie 2009, 91, 692–696. [Google Scholar] [CrossRef] [PubMed]
- Whittle, D.J.; Kilburn, D.G.; Warren, R.A.J.; Miller, R.C. Molecular cloning of a cellulomonas fimi cellulase gene in escherichia coli: Recombinant DNA; plasmid pBR322; immunoassay. Gene 1982, 17, 139–145. [Google Scholar] [CrossRef]
- Kind, T.; Wohlgemuth, G.; Lee, D.Y.; Lu, Y.; Palazoglu, M.; Shahbaz, S.; Fiehn, O. FiehnLib: Mass spectral and retention index libraries for metabolomics based on quadrupole and time-of-flight gas chromatography/mass spectrometry. Anal. Chem. 2009, 81, 10038–10048. [Google Scholar] [CrossRef] [PubMed]
- Stackebrandt, E.; Schumann, P.; Prauser, H. The family cellulomonadaceae. In The Prokaryotes; Springer: New York, NY, USA, 2006; pp. 938–1001. [Google Scholar]
- Stoppok, W.; Rapp, P.; Wagner, F. Formation, location, and regulation of endo-1,4-β-glucanases and β-glucosidases from cellulomonas uda. Appl. Environ. Microbiol. 1982, 44, 44–53. [Google Scholar] [PubMed]
- Xia, J.; Wishart, D.S. Using metaboanalyst 3.0 for comprehensive metabolomics data analysis. Curr. Protoc. Bioinforma. 2016, 55. Available online: http://europepmc.org/abstract/med/27603023 (accessed on 1 October 2017).
- Harzevili, F.D. Yarrowia lipolytica: An overview. In Biotechnological Applications of the Yeast Yarrowia Lipolytica; Springer: New York, NY, USA, 2014; pp. 1–16. [Google Scholar]
- David, K.; Ragauskas, A.J. Switchgrass as an energy crop for biofuel production: A review of its ligno-cellulosic chemical properties. Energy Environ. Sci. 2010, 3, 1182–1190. [Google Scholar] [CrossRef]
- Varanasi, P.; Singh, P.; Auer, M.; Adams, P.D.; Simmons, B.A.; Singh, S. Survey of renewable chemicals produced from lignocellulosic biomass during ionic liquid pretreatment. Biotechnol. Biofuels 2013, 6, 14. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.W. Nutritional requirements and growth of a cellulomonas species on cellulosic substrates. J. Ferment. Technol. 1982, 60, 99–104. [Google Scholar]
- Ledesma-Amaro, R.; Nicaud, J.-M. Metabolic engineering for expanding the substrate range of yarrowia lipolytica. Trends Biotechnol. 2016, 34, 798–809. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, S.; Chisti, Y.; Banerjee, U.C. Production of shikimic acid. Biotechnol. Adv. 2011, 30, 1425–1431. [Google Scholar] [CrossRef] [PubMed]



© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lubbe, A.; Bowen, B.P.; Northen, T. Exometabolomic Analysis of Cross-Feeding Metabolites. Metabolites 2017, 7, 50. https://doi.org/10.3390/metabo7040050
Lubbe A, Bowen BP, Northen T. Exometabolomic Analysis of Cross-Feeding Metabolites. Metabolites. 2017; 7(4):50. https://doi.org/10.3390/metabo7040050
Chicago/Turabian StyleLubbe, Andrea, Benjamin P. Bowen, and Trent Northen. 2017. "Exometabolomic Analysis of Cross-Feeding Metabolites" Metabolites 7, no. 4: 50. https://doi.org/10.3390/metabo7040050
APA StyleLubbe, A., Bowen, B. P., & Northen, T. (2017). Exometabolomic Analysis of Cross-Feeding Metabolites. Metabolites, 7(4), 50. https://doi.org/10.3390/metabo7040050



