Genome-Scale Metabolic Modeling Reveals Sequential Dysregulation of Glutathione Metabolism in Livers from Patients with Alcoholic Hepatitis
Abstract
1. Introduction
2. Materials and Methods
2.1. The Patient Samples and RNA-Seq Data
2.2. Generating Genome-Scale Metabolic Models
2.3. Calculating the Objective Function and Performing Flux Balance Analysis
2.4. Functional Annotation
2.5. Protein–Protein Interaction Network Analysis
2.6. Dimensionality Reduction and Trajectory Analysis
2.7. Statistical Testing
2.8. Data Manipulation and Plotting
2.9. Data Availability and Model Reproducibility
3. Results
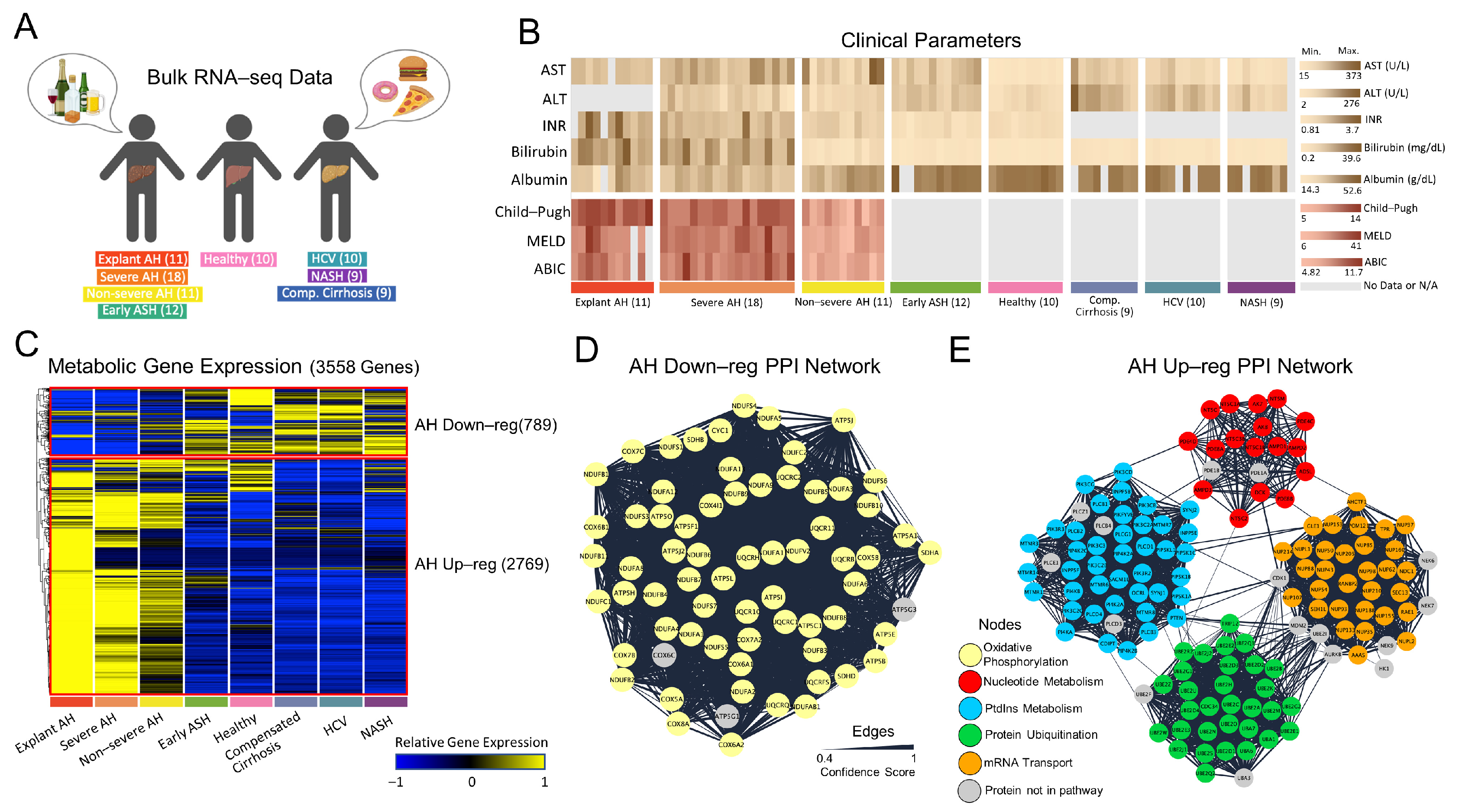
3.1. Regulatory Analysis of Patients with AH
3.2. Structural Analysis of Generated Metabolic Models
3.3. Metabolic Functional Analysis of Patients with AH
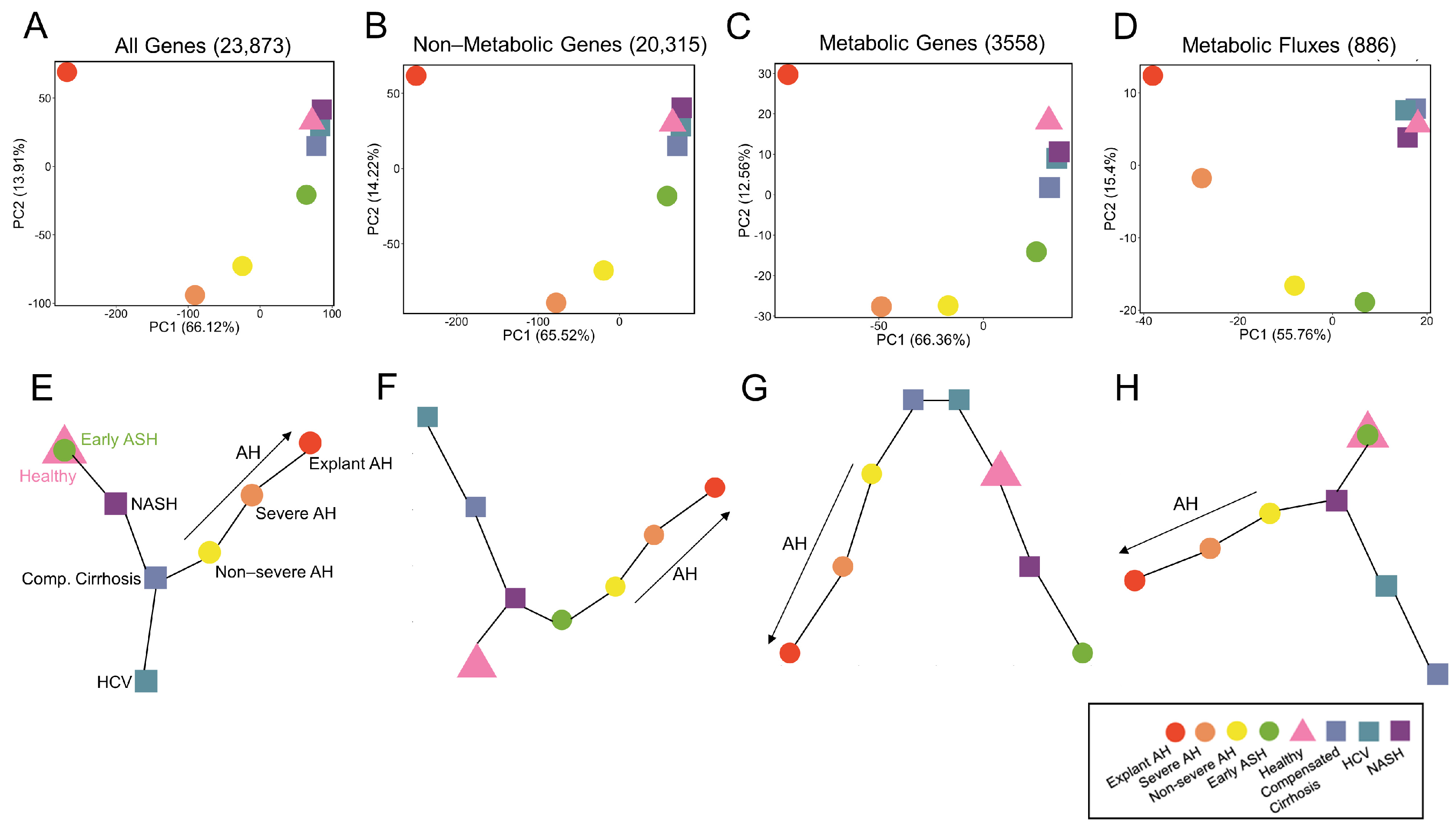
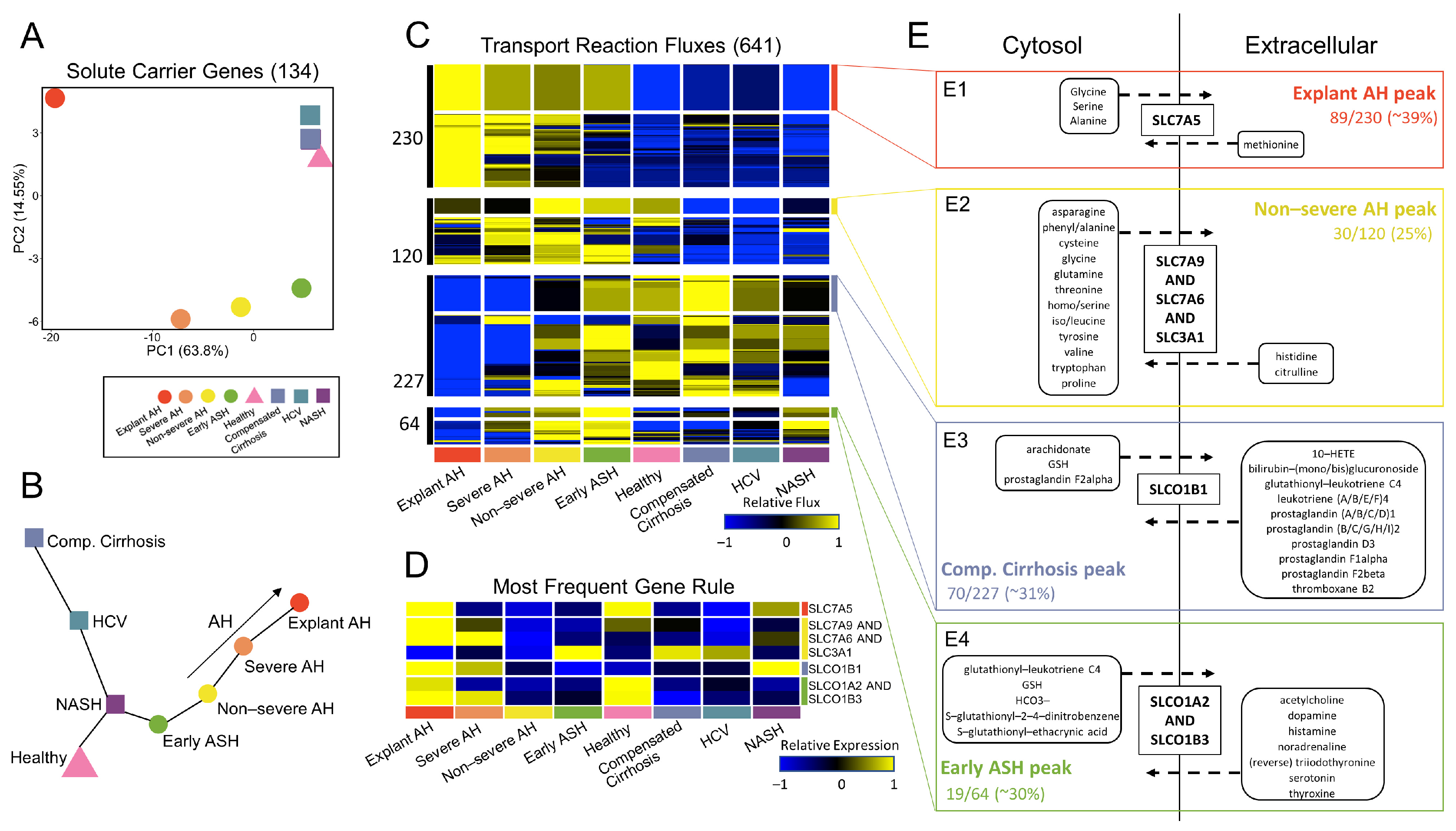
3.4. Systems-Level Analysis of Metabolic Regulatory and Functional Variability
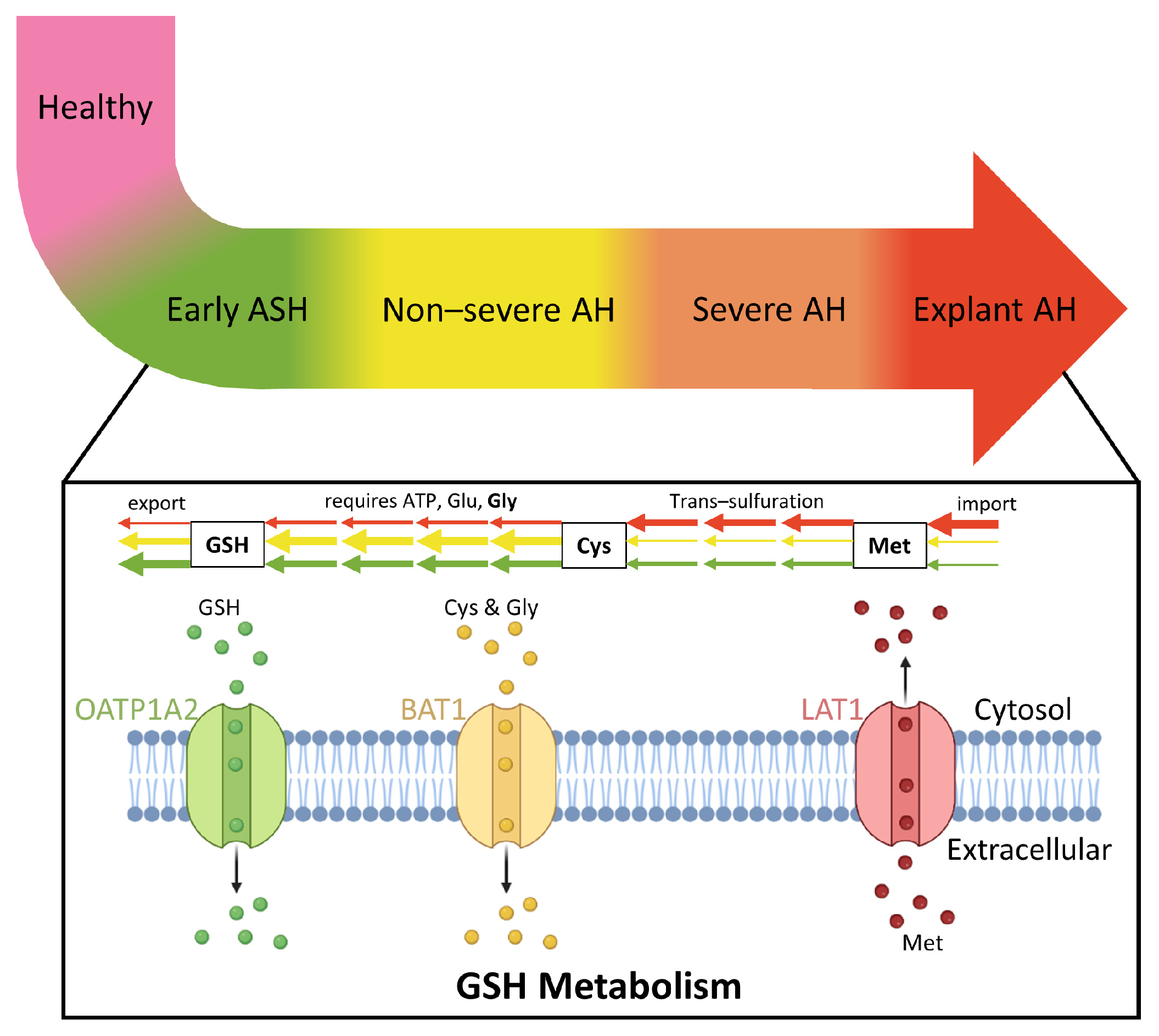
3.5. Glutathione Metabolism Dysregulated in Patients with AH
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lucey, M.R.; Mathurin, P.; Morgan, T.R. Alcoholic Hepatitis. N. Engl. J. Med. 2009, 360, 2758–2769. [Google Scholar] [CrossRef] [PubMed]
- Bertha, M.; Choi, G.; Mellinger, J. Diagnosis and Treatment of Alcohol-Associated Liver Disease: A Patient-Friendly Summary of the 2019 AASLD Guidelines. Clin. Liver Dis. 2021, 17, 418–423. [Google Scholar] [CrossRef] [PubMed]
- Singal, A.K.; Walia, I.; Singal, A.; Soloway, R.D. Corticosteroids and Pentoxifylline for the Treatment of Alcoholic Hepatitis: Current Status. World J. Hepatol. 2011, 3, 205–210. [Google Scholar] [CrossRef] [PubMed]
- Patel, R.; Mueller, M. Alcoholic Liver Disease. In StatPearls; StatPearls Publishing LLC: Treasure Island, FL, USA, 2022. [Google Scholar]
- Dang, K.; Hirode, G.; Singal, A.K.; Sundaram, V.; Wong, R.J. Alcoholic Liver Disease Epidemiology in the United States: A Retrospective Analysis of 3 US Databases. Am. J. Gastroenterol. 2020, 115, 96–104. [Google Scholar] [CrossRef]
- Cholankeril, G.; Ahmed, A. Alcoholic Liver Disease Replaces Hepatitis C Virus Infection as the Leading Indication for Liver Transplantation in the United States. Clin. Gastroenterol. Hepatol. Off. Clin. Pract. J. Am. Gastroenterol. Assoc. 2018, 16, 1356–1358. [Google Scholar] [CrossRef]
- Gao, B.; Bataller, R. Alcoholic Liver Disease: Pathogenesis and New Therapeutic Targets. Gastroenterology 2011, 141, 1572–1585. [Google Scholar] [CrossRef]
- Toshikuni, N.; Tsutsumi, M.; Arisawa, T. Clinical Differences between Alcoholic Liver Disease and Nonalcoholic Fatty Liver Disease. World J. Gastroenterol. 2014, 20, 8393–8406. [Google Scholar] [CrossRef]
- Ikejima, K.; Kon, K.; Yamashina, S. Nonalcoholic Fatty Liver Disease and Alcohol-Related Liver Disease: From Clinical Aspects to Pathophysiological Insights. Clin. Mol. Hepatol. 2020, 26, 728–735. [Google Scholar] [CrossRef]
- Rasineni, K.; Penrice, D.D.; Natarajan, S.K.; McNiven, M.A.; McVicker, B.L.; Kharbanda, K.K.; Casey, C.A.; Harris, E.N. Alcoholic vs Non-Alcoholic Fatty Liver in Rats: Distinct Differences in Endocytosis and Vesicle Trafficking despite Similar Pathology. BMC Gastroenterol. 2016, 16, 27. [Google Scholar] [CrossRef]
- Greuter, T.; Malhi, H.; Gores, G.J.; Shah, V.H. Therapeutic Opportunities for Alcoholic Steatohepatitis and Nonalcoholic Steatohepatitis: Exploiting Similarities and Differences in Pathogenesis. JCI Insight 2017, 2, e95354. [Google Scholar] [CrossRef] [PubMed]
- Wen, C.-S.; Ho, C.-M. Alcohol or Not: A Review Comparing Initial Mechanisms, Contributing Factors, and Liver Transplantation Outcomes Between Alcoholic and Nonalcoholic Steatohepatitis. EMJ 2018, 3, 40–48. [Google Scholar]
- Mitra, S.; De, A.; Chowdhury, A. Epidemiology of Non-Alcoholic and Alcoholic Fatty Liver Diseases. Transl. Gastroenterol. Hepatol. 2020, 5, 16. [Google Scholar] [CrossRef] [PubMed]
- Ceni, E.; Mello, T.; Galli, A. Pathogenesis of Alcoholic Liver Disease: Role of Oxidative Metabolism. World J. Gastroenterol. 2014, 20, 17756–17772. [Google Scholar] [CrossRef] [PubMed]
- Robinson, K.E.; Shah, V.H. Pathogenesis and Pathways: Nonalcoholic Fatty Liver Disease & Alcoholic Liver Disease. Transl. Gastroenterol. Hepatol. 2020, 5, 49. [Google Scholar] [CrossRef] [PubMed]
- Teschke, R. Alcoholic Liver Disease: Alcohol Metabolism, Cascade of Molecular Mechanisms, Cellular Targets, and Clinical Aspects. Biomedicines 2018, 6, 106. [Google Scholar] [CrossRef]
- Massey, V.; Parrish, A.; Argemi, J.; Moreno, M.; Mello, A.; García-Rocha, M.; Altamirano, J.; Odena, G.; Dubuquoy, L.; Louvet, A.; et al. Integrated Multiomics Reveals Glucose Use Reprogramming and Identifies a Novel Hexokinase in Alcoholic Hepatitis. Gastroenterology 2021, 160, 1725–1740.e2. [Google Scholar] [CrossRef]
- Argemi, J.; Latasa, M.U.; Atkinson, S.R.; Blokhin, I.O.; Massey, V.; Gue, J.P.; Cabezas, J.; Lozano, J.J.; Van Booven, D.; Bell, A.; et al. Defective HNF4alpha-Dependent Gene Expression as a Driver of Hepatocellular Failure in Alcoholic Hepatitis. Nat. Commun. 2019, 10, 3126. [Google Scholar] [CrossRef]
- Mardinoglu, A.; Agren, R.; Kampf, C.; Asplund, A.; Uhlen, M.; Nielsen, J. Genome-Scale Metabolic Modelling of Hepatocytes Reveals Serine Deficiency in Patients with Non-Alcoholic Fatty Liver Disease. Nat. Commun. 2014, 5, 3083. [Google Scholar] [CrossRef]
- Agren, R.; Mardinoglu, A.; Asplund, A.; Kampf, C.; Uhlen, M.; Nielsen, J. Identification of Anticancer Drugs for Hepatocellular Carcinoma through Personalized Genome-scale Metabolic Modeling. Mol. Syst. Biol. 2014, 10, 721. [Google Scholar] [CrossRef]
- Naik, A.; Rozman, D.; Belič, A. SteatoNet: The First Integrated Human Metabolic Model with Multi-Layered Regulation to Investigate Liver-Associated Pathologies. PLoS Comput. Biol. 2014, 10, e1003993. [Google Scholar] [CrossRef]
- Gille, C.; Bölling, C.; Hoppe, A.; Bulik, S.; Hoffmann, S.; Hübner, K.; Karlstädt, A.; Ganeshan, R.; König, M.; Rother, K.; et al. HepatoNet1: A Comprehensive Metabolic Reconstruction of the Human Hepatocyte for the Analysis of Liver Physiology. Mol. Syst. Biol. 2010, 6, 411. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Eddy, J.A.; Price, N.D. Reconstruction of Genome-Scale Metabolic Models for 126 Human Tissues Using MCADRE. BMC Syst. Biol. 2012, 6, 153. [Google Scholar] [CrossRef] [PubMed]
- Hyötyläinen, T.; Jerby, L.; Petäjä, E.M.; Mattila, I.; Jäntti, S.; Auvinen, P.; Gastaldelli, A.; Yki-Järvinen, H.; Ruppin, E.; Orešič, M. Genome-Scale Study Reveals Reduced Metabolic Adaptability in Patients with Non-Alcoholic Fatty Liver Disease. Nat. Commun. 2016, 7, 8994. [Google Scholar] [CrossRef] [PubMed]
- Sen, P.; Govaere, O.; Sinioja, T.; McGlinchey, A.; Geng, D.; Ratziu, V.; Bugianesi, E.; Schattenberg, J.M.; Vidal-Puig, A.; Allison, M.; et al. Quantitative Modeling of Human Liver Reveals Dysregulation of Glycosphingolipid Pathways in Nonalcoholic Fatty Liver Disease. iScience 2022, 25, 104949. [Google Scholar] [CrossRef]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast Universal RNA-Seq Aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef]
- Li, B.; Dewey, C.N. RSEM: Accurate Transcript Quantification from RNA-Seq Data with or without a Reference Genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef]
- Robinson, J.L.; Kocabaş, P.; Wang, H.; Cholley, P.-E.; Cook, D.; Nilsson, A.; Anton, M.; Ferreira, R.; Domenzain, I.; Billa, V.; et al. An Atlas of Human Metabolism. Sci. Signal. 2020, 13, eaaz1482. [Google Scholar] [CrossRef]
- Agren, R.; Liu, L.; Shoaie, S.; Vongsangnak, W.; Nookaew, I.; Nielsen, J. The RAVEN Toolbox and Its Use for Generating a Genome-Scale Metabolic Model for Penicillium Chrysogenum. PLoS Comput. Biol. 2013, 9, e1002980. [Google Scholar] [CrossRef]
- Nilsson, A.; Nielsen, J. Genome Scale Metabolic Modeling of Cancer. Metab. Eng. 2017, 43, 103–112. [Google Scholar] [CrossRef]
- Kim, M.K.; Lane, A.; Kelley, J.J.; Lun, D.S. E-Flux2 and SPOT: Validated Methods for Inferring Intracellular Metabolic Flux Distributions from Transcriptomic Data. PLoS ONE 2016, 11, e0157101. [Google Scholar] [CrossRef]
- Sherman, B.T.; Hao, M.; Qiu, J.; Jiao, X.; Baseler, M.W.; Lane, H.C.; Imamichi, T.; Chang, W. DAVID: A Web Server for Functional Enrichment Analysis and Functional Annotation of Gene Lists (2021 Update). Nucleic Acids Res. 2022, 50, W216–W221. [Google Scholar] [CrossRef] [PubMed]
- Thomas, P.D.; Campbell, M.J.; Kejariwal, A.; Mi, H.; Karlak, B.; Daverman, R.; Diemer, K.; Muruganujan, A.; Narechania, A. PANTHER: A Library of Protein Families and Subfamilies Indexed by Function. Genome Res. 2003, 13, 2129–2141. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Gable, A.L.; Nastou, K.C.; Lyon, D.; Kirsch, R.; Pyysalo, S.; Doncheva, N.T.; Legeay, M.; Fang, T.; Bork, P.; et al. The STRING Database in 2021: Customizable Protein–Protein Networks, and Functional Characterization of User-Uploaded Gene/Measurement Sets. Nucleic Acids Res. 2021, 49, D605–D612. [Google Scholar] [CrossRef] [PubMed]
- Doncheva, N.T.; Morris, J.H.; Gorodkin, J.; Jensen, L.J. Cytoscape StringApp: Network Analysis and Visualization of Proteomics Data. J. Proteome Res. 2019, 18, 623–632. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A Software Environment for Integrated Models of Biomolecular Interaction Networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Bader, G.D.; Hogue, C.W. An Automated Method for Finding Molecular Complexes in Large Protein Interaction Networks. BMC Bioinform. 2003, 4, 2. [Google Scholar] [CrossRef]
- Erdemir, A.; Mulugeta, L.; Ku, J.P.; Drach, A.; Horner, M.; Morrison, T.M.; Peng, G.C.Y.; Vadigepalli, R.; Lytton, W.W.; Myers, J.G. Credible Practice of Modeling and Simulation in Healthcare: Ten Rules from a Multidisciplinary Perspective. J. Transl. Med. 2020, 18, 369. [Google Scholar] [CrossRef]
- García-Ruiz, C.; Kaplowitz, N.; Fernandez-Checa, J.C. Role of Mitochondria in Alcoholic Liver Disease. Curr. Pathobiol. Rep. 2013, 1, 159–168. [Google Scholar] [CrossRef]
- Middleton, P.; Vergis, N. Mitochondrial Dysfunction and Liver Disease: Role, Relevance, and Potential for Therapeutic Modulation. Ther. Adv. Gastroenterol. 2021, 14, 175628482110313. [Google Scholar] [CrossRef]
- Meikle, P.J.; Mundra, P.A.; Wong, G.; Rahman, K.; Huynh, K.; Barlow, C.K.; Duly, A.M.P.; Haber, P.S.; Whitfield, J.B.; Seth, D. Circulating Lipids Are Associated with Alcoholic Liver Cirrhosis and Represent Potential Biomarkers for Risk Assessment. PLoS ONE 2015, 10, e0130346. [Google Scholar] [CrossRef]
- French, S.W.; Bardag-Gorce, F. Ubiquitin-Proteasome Pathway in the Pathogenesis of Liver Disease. In Signaling Pathways in Liver Diseases; Dufour, J.-F., Clavien, P.-A., Trautwein, C., Graf, R., Eds.; Springer: Berlin/Heidelberg, Germany, 2005; pp. 377–389. ISBN 978-3-540-22934-6. [Google Scholar]
- Donohue, T.M. The Ubiquitin-Proteasome System and Its Role in Ethanol-Induced Disorders. Addict. Biol. 2002, 7, 15–28. [Google Scholar] [CrossRef] [PubMed]
- Park, J.-S.; Ma, H.; Roh, Y.-S. Ubiquitin Pathways Regulate the Pathogenesis of Chronic Liver Disease. Biochem. Pharmacol. 2021, 193, 114764. [Google Scholar] [CrossRef] [PubMed]
- Régnier, M.; Polizzi, A.; Guillou, H.; Loiseau, N. Sphingolipid Metabolism in Non-Alcoholic Fatty Liver Diseases. Biochimie 2019, 159, 9–22. [Google Scholar] [CrossRef] [PubMed]
- Häfliger, P.; Charles, R.-P. The L-Type Amino Acid Transporter LAT1—An Emerging Target in Cancer. Int. J. Mol. Sci. 2019, 20, 2428. [Google Scholar] [CrossRef]
- Ricciotti, E.; FitzGerald, G.A. Prostaglandins and Inflammation. Arterioscler. Thromb. Vasc. Biol. 2011, 31, 986–1000. [Google Scholar] [CrossRef] [PubMed]
- Schoen Smith, J.M.; Lautt, W.W. The Role of Prostaglandins in Triggering the Liver Regeneration Cascade. Nitric Oxide 2005, 13, 111–117. [Google Scholar] [CrossRef] [PubMed]
- Lu, S.C. Dysregulation of Glutathione Synthesis in Liver Disease. Liver Res. 2020, 4, 64–73. [Google Scholar] [CrossRef]
- Rachakonda, V.; Gabbert, C.; Raina, A.; Bell, L.N.; Cooper, S.; Malik, S.; Behari, J. Serum Metabolomic Profiling in Acute Alcoholic Hepatitis Identifies Multiple Dysregulated Pathways. PLoS ONE 2014, 9, e113860. [Google Scholar] [CrossRef]
- Cree, M.G.; Newcomer, B.R.; Katsanos, C.S.; Sheffield-Moore, M.; Chinkes, D.; Aarsland, A.; Urban, R.; Wolfe, R.R. Intramuscular and Liver Triglycerides Are Increased in the Elderly. J. Clin. Endocrinol. Metab. 2004, 89, 3864–3871. [Google Scholar] [CrossRef]
- Basu, R.; Dalla Man, C.; Campioni, M.; Basu, A.; Klee, G.; Toffolo, G.; Cobelli, C.; Rizza, R.A. Effects of Age and Sex on Postprandial Glucose Metabolism. Diabetes 2006, 55, 2001–2014. [Google Scholar] [CrossRef]
- Kim, I.H.; Kisseleva, T.; Brenner, D.A. Aging and Liver Disease. Curr. Opin. Gastroenterol. 2015, 31, 184–191. [Google Scholar] [CrossRef] [PubMed]
- Lutchmansingh, F.K.; Hsu, J.W.; Bennett, F.I.; Badaloo, A.V.; McFarlane-Anderson, N.; Gordon-Strachan, G.M.; Wright-Pascoe, R.A.; Jahoor, F.; Boyne, M.S. Glutathione Metabolism in Type 2 Diabetes and Its Relationship with Microvascular Complications and Glycemia. PLoS ONE 2018, 13, e0198626. [Google Scholar] [CrossRef] [PubMed]
- CDC. National Diabetes Statistics Report | Diabetes. Available online: https://www.cdc.gov/diabetes/data/statistics-report/index.html (accessed on 15 November 2022).
- Lischner, M.W.; Alexander, J.F.; Galambos, J.T. Natural History of Alcoholic Hepatitis: I. The Acute Disease. Am. J. Dig. Dis. 1971, 16, 481–494. [Google Scholar] [CrossRef] [PubMed]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Manchel, A.; Mahadevan, R.; Bataller, R.; Hoek, J.B.; Vadigepalli, R. Genome-Scale Metabolic Modeling Reveals Sequential Dysregulation of Glutathione Metabolism in Livers from Patients with Alcoholic Hepatitis. Metabolites 2022, 12, 1157. https://doi.org/10.3390/metabo12121157
Manchel A, Mahadevan R, Bataller R, Hoek JB, Vadigepalli R. Genome-Scale Metabolic Modeling Reveals Sequential Dysregulation of Glutathione Metabolism in Livers from Patients with Alcoholic Hepatitis. Metabolites. 2022; 12(12):1157. https://doi.org/10.3390/metabo12121157
Chicago/Turabian StyleManchel, Alexandra, Radhakrishnan Mahadevan, Ramon Bataller, Jan B. Hoek, and Rajanikanth Vadigepalli. 2022. "Genome-Scale Metabolic Modeling Reveals Sequential Dysregulation of Glutathione Metabolism in Livers from Patients with Alcoholic Hepatitis" Metabolites 12, no. 12: 1157. https://doi.org/10.3390/metabo12121157
APA StyleManchel, A., Mahadevan, R., Bataller, R., Hoek, J. B., & Vadigepalli, R. (2022). Genome-Scale Metabolic Modeling Reveals Sequential Dysregulation of Glutathione Metabolism in Livers from Patients with Alcoholic Hepatitis. Metabolites, 12(12), 1157. https://doi.org/10.3390/metabo12121157






