The Role of AKR1B10 in Physiology and Pathophysiology
Abstract
1. Introduction
2. Gene Regulation of AKR1B10
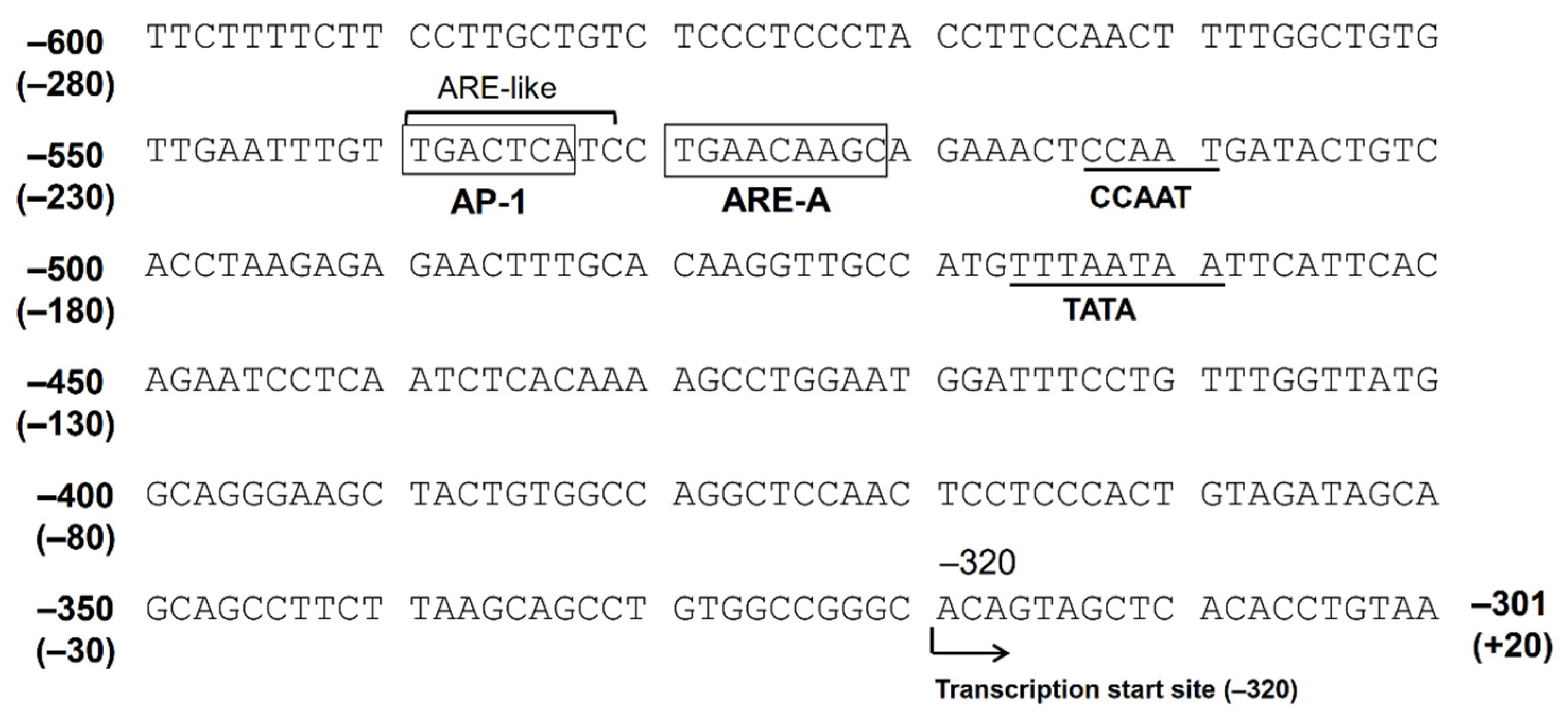
2.1. Factors Regulating AKR1B10 Expression
2.2. Contribution of Nrf2 to AKR1B10 Induction
2.3. The Function of AP-1 Protein in AKR1B10 Gene Regulation
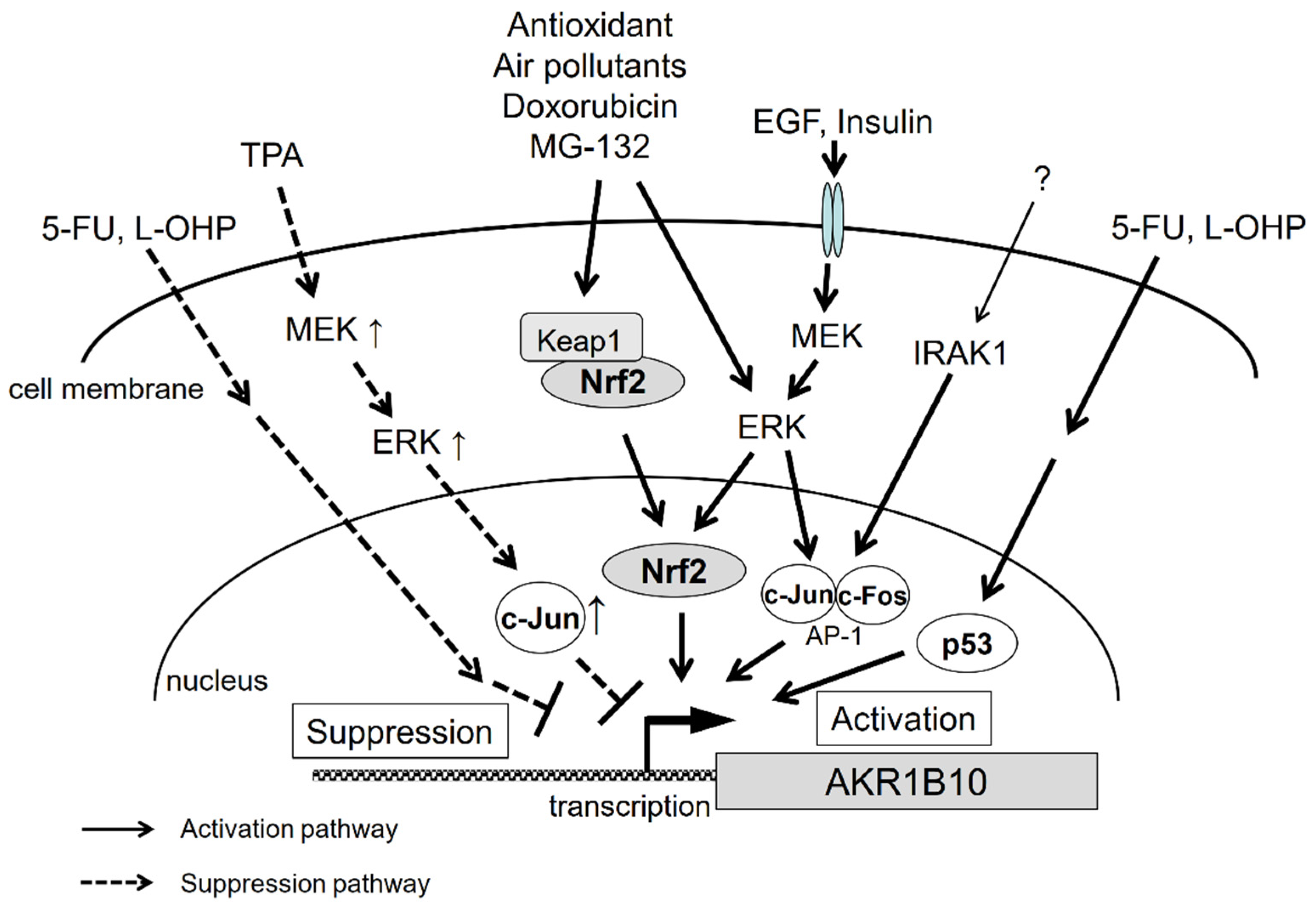
2.4. Signal Transduction
3. AKR1B10 as a Multifunctional NADPH-Dependent Reductase
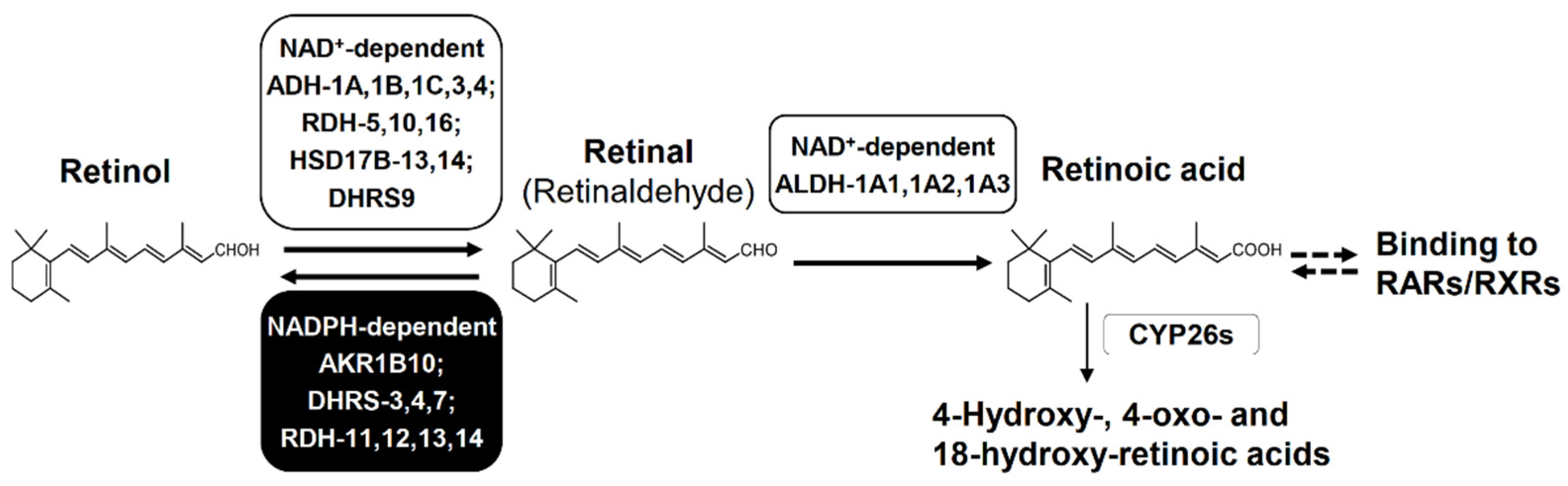
3.1. Retinoid Metabolism
3.2. Detoxification of Reactive Carbonyl Species (RCS)
3.3. Isoprenoid Metabolism
3.4. Xenobiotic Metabolism
4. Moonlighting Functions of AKR1B10
- (1)
- Fatty acid/lipid synthesis. In breast cancer RAO-3 cells, AKR1B10 interacts with acetyl-CoA carboxylase-α (a rate-limiting enzyme of de novo fatty acid synthesis), preventing acetyl-CoA carboxylase-α ubiquitination and proteolysis, and thereby promoting fatty acid/lipid synthesis [64].
- (2)
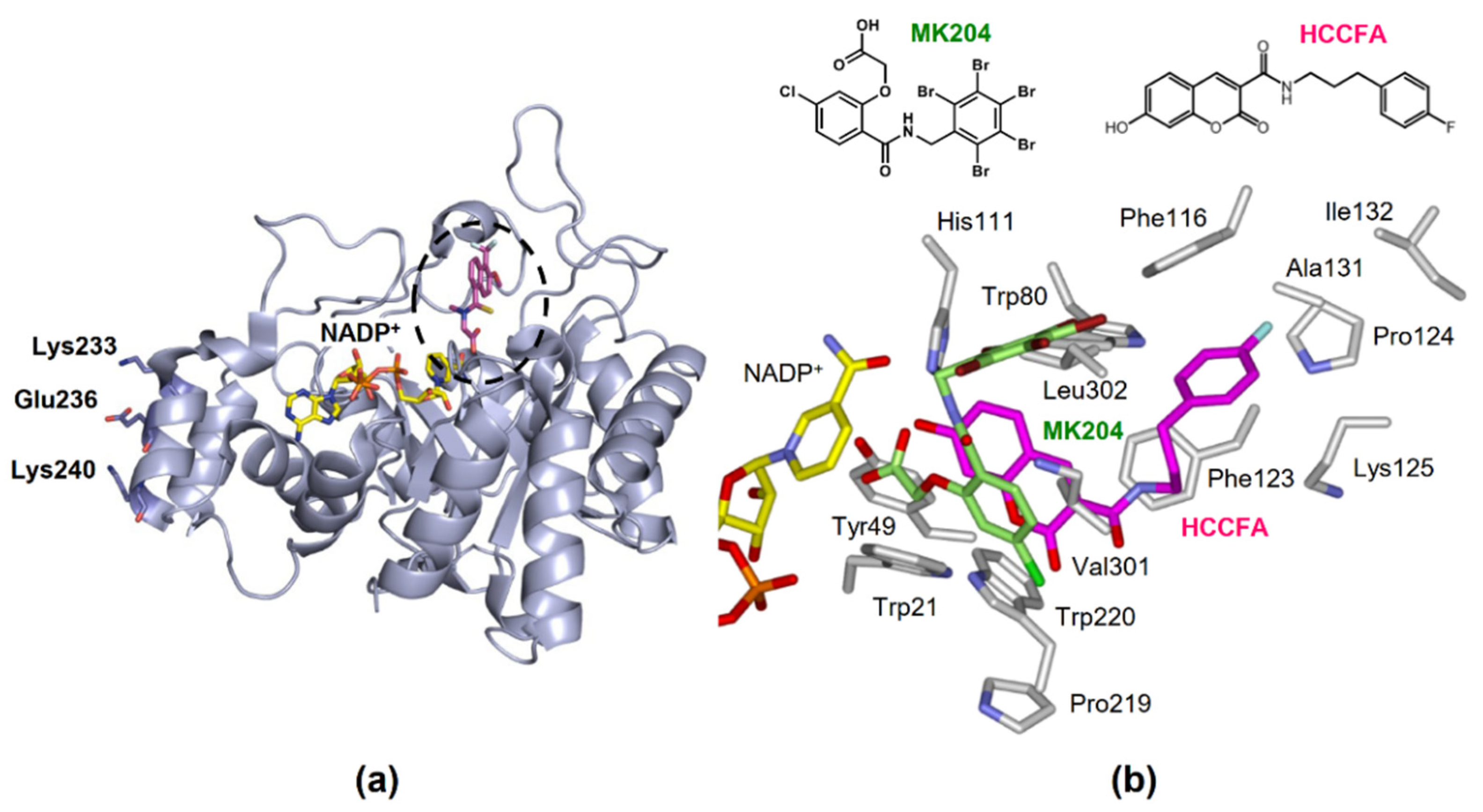
- Interaction with heat shock protein (HSP) 90α in AKR1B10 secretion. Cytosolic AKR1B10 is secreted from cells through a lysosome-mediated nonclassical pathway, increasing its presence in breast cancer patients’ serum [65]. The AKR1B10 secretion is mediated by interaction with HSP90α, which binds to Lys-233, Glu-236, and Lys-240 in AKR1B10 [66] (Figure 4a).
- (3)
- Interaction with glyceraldehyde-3-phosphate dehydrogenase (GAPDH). A recent report shows that AKR1B10 interacts with GAPDH in CRC HT29 cells [67]. The interaction inhibits the nuclear import of GAPDH, and subsequently results in autophagy repression, for which AKR1B10 reductase activity is likely to be important.
5. AKR1B10 in the Gastrointestinal Tract and Cancer
6. Diseases Associated with AKR1B10 Elevation
6.1. Hepatocellular Carcinoma (HCC)
6.2. Nonalcoholic Fatty Liver Disease (NAFLD)
6.3. Lung Cancer
6.4. Breast Cancer
6.5. Pancreatic Cancer
6.6. Oral Cancer
6.7. Other Cancers
6.8. Non-Neoplastic Skin Diseases
7. AKR1B10 in Anti-Cancer Drug Resistance
8. AKR1B10 Inhibitors
9. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Penning, T.M. The aldo-keto reductases (AKRs): Overview. Chem. Biol. Interact. 2015, 234, 236–246. [Google Scholar] [CrossRef]
- Cao, D.; Fan, S.T.; Chung, S.S. Identification and characterization of a novel human aldose reductase-like gene. J. Biol. Chem. 1998, 273, 11429–11435. [Google Scholar] [CrossRef] [PubMed]
- Weber, S.; Salabei, J.K.; Moller, G.; Kremmer, E.; Bhatnagar, A.; Adamski, J.; Barski, O.A. Aldo-keto Reductase 1B15 (AKR1B15): A mitochondrial human aldo-keto reductase with activity toward steroids and 3-keto-acyl-CoA conjugates. J. Biol. Chem. 2015, 290, 6531–6545. [Google Scholar] [CrossRef] [PubMed]
- Endo, S.; Matsunaga, T.; Mamiya, H.; Ohta, C.; Soda, M.; Kitade, Y.; Tajima, K.; Zhao, H.T.; El-Kabbani, O.; Hara, A. Kinetic studies of AKR1B10, human aldose reductase-like protein: Endogenous substrates and inhibition by steroids. Arch. Biochem. Biophys. 2009, 487, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Gimenez-Dejoz, J.; Kolar, M.H.; Ruiz, F.X.; Crespo, I.; Cousido-Siah, A.; Podjarny, A.; Barski, O.A.; Fanfrlik, J.; Pares, X.; Farres, J.; et al. Substrate Specificity, Inhibitor Selectivity and Structure-Function Relationships of Aldo-Keto Reductase 1B15: A Novel Human Retinaldehyde Reductase. PLoS ONE 2015, 10, e0134506. [Google Scholar] [CrossRef]
- Gimenez-Dejoz, J.; Weber, S.; Fernandez-Pardo, A.; Moller, G.; Adamski, J.; Porte, S.; Pares, X.; Farres, J. Engineering aldo-keto reductase 1B10 to mimic the distinct 1B15 topology and specificity towards inhibitors and substrates, including retinoids and steroids. Chem. Biol. Interact. 2019, 307, 186–194. [Google Scholar] [CrossRef]
- Kabututu, Z.; Manin, M.; Pointud, J.C.; Maruyama, T.; Nagata, N.; Lambert, S.; Lefrancois-Martinez, A.M.; Martinez, A.; Urade, Y. Prostaglandin F2{alpha} Synthase Activities of Aldo-Keto Reductase 1B1, 1B3 and 1B7. J. Biochem. 2009, 145, 161–168. [Google Scholar] [CrossRef]
- Fukumoto, S.; Yamauchi, N.; Moriguchi, H.; Hippo, Y.; Watanabe, A.; Shibahara, J.; Taniguchi, H.; Ishikawa, S.; Ito, H.; Yamamoto, S.; et al. Overexpression of the aldo-keto reductase family protein AKR1B10 is highly correlated with smokers’ non-small cell lung carcinomas. Clin. Cancer Res. 2005, 11, 1776–1785. [Google Scholar] [CrossRef]
- Hyndman, D.J.; Flynn, T.G. Sequence and expression levels in human tissues of a new member of the aldo-keto reductase family. Biochim. Biophys. Acta 1998, 1399, 198–202. [Google Scholar] [CrossRef]
- Oates, P.J. Aldose reductase, still a compelling target for diabetic neuropathy. Curr. Drug Targets 2008, 9, 14–36. [Google Scholar] [CrossRef] [PubMed]
- Chang, K.C.; Petrash, J.M. Aldo-Keto Reductases: Multifunctional Proteins as Therapeutic Targets in Diabetes and Inflammatory Disease. Adv. Exp. Med. Biol. 2018, 1032, 173–202. [Google Scholar] [CrossRef] [PubMed]
- Scuric, Z.; Stain, S.C.; Anderson, W.F.; Hwang, J.J. New member of aldose reductase family proteins overexpressed in human hepatocellular carcinoma. Hepatology 1998, 27, 943–950. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Zhong, L.; Krishack, P.A.; Robbins, S.; Cao, J.X.; Zhao, Y.; Chung, S.; Cao, D. Structure and promoter characterization of aldo-keto reductase family 1 B10 gene. Gene 2009, 437, 39–44. [Google Scholar] [CrossRef] [PubMed]
- Nishinaka, T.; Miura, T.; Okumura, M.; Nakao, F.; Nakamura, H.; Terada, T. Regulation of aldo-keto reductase AKR1B10 gene expression: Involvement of transcription factor Nrf2. Chem. Biol. Interact. 2011, 191, 185–191. [Google Scholar] [CrossRef] [PubMed]
- Nishinaka, T.; Miura, T.; Sakou, M.; Hidaka, C.; Sasaoka, C.; Okamura, A.; Okamoto, A.; Terada, T. Down-regulation of aldo-keto reductase AKR1B10 gene expression by a phorbol ester via the ERK/c-Jun signaling pathway. Chem. Biol. Interact. 2015, 234, 274–281. [Google Scholar] [CrossRef]
- Zinovieva, O.L.; Grineva, E.N.; Krasnov, G.S.; Karpov, D.S.; Zheltukhin, A.O.; Snezhkina, A.V.; Kudryavtseva, A.V.; Mashkova, T.D.; Lisitsyn, N.A. Treatment of cancer cells with chemotherapeutic drugs results in profound changes in expression of genes encoding aldehyde-metabolizing enzymes. J. Cancer 2019, 10, 4256–4263. [Google Scholar] [CrossRef]
- Ebert, B.; Kisiela, M.; Wsol, V.; Maser, E. Proteasome inhibitors MG-132 and bortezomib induce AKR1C1, AKR1C3, AKR1B1, and AKR1B10 in human colon cancer cell lines SW-480 and HT-29. Chem. Biol. Interact. 2011, 191, 239–249. [Google Scholar] [CrossRef]
- Morikawa, Y.; Kezuka, C.; Endo, S.; Ikari, A.; Soda, M.; Yamamura, K.; Toyooka, N.; El-Kabbani, O.; Hara, A.; Matsunaga, T. Acquisition of doxorubicin resistance facilitates migrating and invasive potentials of gastric cancer MKN45 cells through up-regulating aldo-keto reductase 1B10. Chem. Biol. Interact. 2015, 230, 30–39. [Google Scholar] [CrossRef]
- Liu, Z.; Yan, R.; Al-Salman, A.; Shen, Y.; Bu, Y.; Ma, J.; Luo, D.X.; Huang, C.; Jiang, Y.; Wilber, A.; et al. Epidermal growth factor induces tumour marker AKR1B10 expression through activator protein-1 signalling in hepatocellular carcinoma cells. Biochem. J. 2012, 442, 273–282. [Google Scholar] [CrossRef] [PubMed]
- Shaw, N.; Yang, B.; Millward, A.; Demaine, A.; Hodgkinson, A. AKR1B10 is induced by hyperglycaemia and lipopolysaccharide in patients with diabetic nephropathy. Cell Stress Chaperones 2014, 19, 281–287. [Google Scholar] [CrossRef]
- Van Zoelen, E.J.; Duarte, I.; Hendriks, J.M.; van der Woning, S.P. TGFbeta-induced switch from adipogenic to osteogenic differentiation of human mesenchymal stem cells: Identification of drug targets for prevention of fat cell differentiation. Stem Cell Res. Ther. 2016, 7, 123. [Google Scholar] [CrossRef] [PubMed]
- Matsunaga, T.; Morikawa, Y.; Haga, M.; Endo, S.; Soda, M.; Yamamura, K.; El-Kabbani, O.; Tajima, K.; Ikari, A.; Hara, A. Exposure to 9,10-phenanthrenequinone accelerates malignant progression of lung cancer cells through up-regulation of aldo-keto reductase 1B10. Toxicol. Appl. Pharmacol. 2014, 278, 180–189. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Wang, G.; Ricard, M.J.; Ferris, B.; Strulovici-Barel, Y.; Salit, J.; Hackett, N.R.; Gudas, L.J.; Crystal, R.G. Smoking-induced upregulation of AKR1B10 expression in the airway epithelium of healthy individuals. Chest 2010, 138, 1402–1410. [Google Scholar] [CrossRef]
- Itoh, K.; Igarashi, K.; Hayashi, N.; Nishizawa, M.; Yamamoto, M. Cloning and characterization of a novel erythroid cell-derived CNC family transcription factor heterodimerizing with the small Maf family proteins. Mol. Cell. Biol. 1995, 15, 4184–4193. [Google Scholar] [CrossRef] [PubMed]
- Rushmore, T.H.; Morton, M.R.; Pickett, C.B. The antioxidant responsive element. Activation by oxidative stress and identification of the DNA consensus sequence required for functional activity. J. Biol. Chem. 1991, 266, 11632–11639. [Google Scholar] [CrossRef]
- Kobayashi, A.; Kang, M.I.; Okawa, H.; Ohtsuji, M.; Zenke, Y.; Chiba, T.; Igarashi, K.; Yamamoto, M. Oxidative stress sensor Keap1 functions as an adaptor for Cul3-based E3 ligase to regulate proteasomal degradation of Nrf2. Mol. Cell. Biol. 2004, 24, 7130–7139. [Google Scholar] [CrossRef]
- Tong, K.I.; Padmanabhan, B.; Kobayashi, A.; Shang, C.; Hirotsu, Y.; Yokoyama, S.; Yamamoto, M. Different electrostatic potentials define ETGE and DLG motifs as hinge and latch in oxidative stress response. Mol. Cell. Biol. 2007, 27, 7511–7521. [Google Scholar] [CrossRef]
- Mimura, J.; Inose-Maruyama, A.; Taniuchi, S.; Kosaka, K.; Yoshida, H.; Yamazaki, H.; Kasai, S.; Harada, N.; Kaufman, R.J.; Oyadomari, S.; et al. Concomitant Nrf2- and ATF4-activation by Carnosic Acid Cooperatively Induces Expression of Cytoprotective Genes. Int. J. Mol. Sci. 2019, 20, 1706. [Google Scholar] [CrossRef]
- MacLeod, A.K.; McMahon, M.; Plummer, S.M.; Higgins, L.G.; Penning, T.M.; Igarashi, K.; Hayes, J.D. Characterization of the cancer chemopreventive NRF2-dependent gene battery in human keratinocytes: Demonstration that the KEAP1-NRF2 pathway, and not the BACH1-NRF2 pathway, controls cytoprotection against electrophiles as well as redox-cycling compounds. Carcinogenesis 2009, 30, 1571–1580. [Google Scholar] [CrossRef]
- Rooney, J.P.; Chorley, B.; Hiemstra, S.; Wink, S.; Wang, X.; Bell, D.A.; van de Water, B.; Corton, J.C. Mining a human transcriptome database for chemical modulators of NRF2. PLoS ONE 2020, 15, e0239367. [Google Scholar] [CrossRef]
- Nishinaka, T.; Miura, T.; Shimizu, K.; Terada, T. Identification and characterization of functional antioxidant response elements in the promoter of the aldo-keto reductase AKR1B10 gene. Chem. Biol. Interact. 2017. [Google Scholar] [CrossRef] [PubMed]
- Chiu, R.; Boyle, W.J.; Meek, J.; Smeal, T.; Hunter, T.; Karin, M. The c-Fos protein interacts with c-Jun/AP-1 to stimulate transcription of AP-1 responsive genes. Cell 1988, 54, 541–552. [Google Scholar] [CrossRef]
- Cheng, B.Y.; Lau, E.Y.; Leung, H.W.; Leung, C.O.; Ho, N.P.; Gurung, S.; Cheng, L.K.; Lin, C.H.; Lo, R.C.; Ma, S.; et al. IRAK1 Augments Cancer Stemness and Drug Resistance via the AP-1/AKR1B10 Signaling Cascade in Hepatocellular Carcinoma. Cancer Res. 2018, 78, 2332–2342. [Google Scholar] [CrossRef] [PubMed]
- Ruiz, F.X.; Porte, S.; Pares, X.; Farres, J. Biological role of aldo-keto reductases in retinoic Acid biosynthesis and signaling. Front. Pharmacol. 2012, 3, 58. [Google Scholar] [CrossRef] [PubMed]
- Kedishvili, N.Y. Retinoic Acid Synthesis and Degradation. Subcell. Biochem. 2016, 81, 127–161. [Google Scholar] [CrossRef] [PubMed]
- Matsunaga, T.; Endo, S.; Maeda, S.; Ishikura, S.; Tajima, K.; Tanaka, N.; Nakamura, K.T.; Imamura, Y.; Hara, A. Characterization of human DHRS4: An inducible short-chain dehydrogenase/reductase enzyme with 3beta-hydroxysteroid dehydrogenase activity. Arch. Biochem. Biophys. 2008, 477, 339–347. [Google Scholar] [CrossRef]
- Stambergova, H.; Zemanova, L.; Lundova, T.; Malcekova, B.; Skarka, A.; Safr, M.; Wsol, V. Human DHRS7, promising enzyme in metabolism of steroids and retinoids? J. Steroid Biochem. Mol. Biol. 2016, 155, 112–119. [Google Scholar] [CrossRef]
- Belyaeva, O.V.; Adams, M.K.; Popov, K.M.; Kedishvili, N.Y. Generation of Retinaldehyde for Retinoic Acid Biosynthesis. Biomolecules 2019, 10, 5. [Google Scholar] [CrossRef]
- Adams, M.K.; Belyaeva, O.V.; Wu, L.; Kedishvili, N.Y. The retinaldehyde reductase activity of DHRS3 is reciprocally activated by retinol dehydrogenase 10 to control retinoid homeostasis. J. Biol. Chem. 2014, 289, 14868–14880. [Google Scholar] [CrossRef]
- Pequerul, R.; Vera, J.; Gimenez-Dejoz, J.; Crespo, I.; Coines, J.; Porte, S.; Rovira, C.; Pares, X.; Farres, J. Structural and kinetic features of aldehyde dehydrogenase 1A (ALDH1A) subfamily members, cancer stem cell markers active in retinoic acid biosynthesis. Arch. Biochem. Biophys. 2020, 681, 108256. [Google Scholar] [CrossRef]
- Conklin, D.; Prough, R.; Bhatanagar, A. Aldehyde metabolism in the cardiovascular system. Mol. Biosyst. 2007, 3, 136–150. [Google Scholar] [CrossRef] [PubMed]
- Singh, M.; Kapoor, A.; Bhatnagar, A. Oxidative and reductive metabolism of lipid-peroxidation derived carbonyls. Chem. Biol. Interact. 2015, 234, 261–273. [Google Scholar] [CrossRef] [PubMed]
- Spite, M.; Baba, S.P.; Ahmed, Y.; Barski, O.A.; Nijhawan, K.; Petrash, J.M.; Bhatnagar, A.; Srivastava, S. Substrate specificity and catalytic efficiency of aldo-keto reductases with phospholipid aldehydes. Biochem. J. 2007, 405, 95–105. [Google Scholar] [CrossRef] [PubMed]
- Baba, S.P.; Barski, O.A.; Ahmed, Y.; O’Toole, T.E.; Conklin, D.J.; Bhatnagar, A.; Srivastava, S. Reductive metabolism of AGE precursors: A metabolic route for preventing AGE accumulation in cardiovascular tissue. Diabetes 2009, 58, 2486–2497. [Google Scholar] [CrossRef] [PubMed]
- Zhong, L.; Liu, Z.; Yan, R.; Johnson, S.; Fang, X.; Cao, D. Aldo-keto reductase family 1 B10 protein detoxifies dietary and lipid-derived alpha, beta-unsaturated carbonyls at physiological levels. Biochem. Biophys. Res. Commun. 2009, 387, 245–250. [Google Scholar] [CrossRef] [PubMed]
- Martin, H.J.; Maser, E. Role of human aldo-keto-reductase AKR1B10 in the protection against toxic aldehydes. Chem. Biol. Interact. 2009, 178, 145–150. [Google Scholar] [CrossRef] [PubMed]
- Shen, Y.; Zhong, L.; Johnson, S.; Cao, D. Human aldo-keto reductases 1B1 and 1B10: A comparative study on their enzyme activity toward electrophilic carbonyl compounds. Chem. Biol. Interact. 2011, 191, 192–198. [Google Scholar] [CrossRef]
- Matsunaga, T.; Yamaji, Y.; Tomokuni, T.; Morita, H.; Morikawa, Y.; Suzuki, A.; Yonezawa, A.; Endo, S.; Ikari, A.; Iguchi, K.; et al. Nitric oxide confers cisplatin resistance in human lung cancer cells through upregulation of aldo-keto reductase 1B10 and proteasome. Free Radic. Res. 2014, 48, 1371–1385. [Google Scholar] [CrossRef]
- Vander Jagt, D.L.; Kolb, N.S.; Vander Jagt, T.J.; Chino, J.; Martinez, F.J.; Hunsaker, L.A.; Royer, R.E. Substrate specificity of human aldose reductase: Identification of 4-hydroxynonenal as an endogenous substrate. Biochim. Biophys. Acta 1995, 1249, 117–126. [Google Scholar] [CrossRef]
- Ramana, K.V.; Dixit, B.L.; Srivastava, S.; Balendiran, G.K.; Srivastava, S.K.; Bhatnagar, A. Selective recognition of glutathiolated aldehydes by aldose reductase. Biochemistry 2000, 39, 12172–12180. [Google Scholar] [CrossRef]
- Burczynski, M.E.; Sridhar, G.R.; Palackal, N.T.; Penning, T.M. The reactive oxygen species- and Michael acceptor-inducible human aldo-keto reductase AKR1C1 reduces the alpha,beta-unsaturated aldehyde 4-hydroxy-2-nonenal to 1,4-dihydroxy-2-nonene. J. Biol. Chem. 2001, 276, 2890–2897. [Google Scholar] [CrossRef] [PubMed]
- Endo, S.; Matsunaga, T.; Ohta, C.; Soda, M.; Kanamori, A.; Kitade, Y.; Ohno, S.; Tajima, K.; El-Kabbani, O.; Hara, A. Roles of rat and human aldo-keto reductases in metabolism of farnesol and geranylgeraniol. Chem. Biol. Interact. 2011, 191, 261–268. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Li, H.; Yang, Y.; Liao, J.; Yang, G.Y. Knockdown or inhibition of aldo-keto reductase 1B10 inhibits pancreatic carcinoma growth via modulating Kras-E-cadherin pathway. Cancer Lett. 2014, 355, 273–280. [Google Scholar] [CrossRef]
- Chung, Y.T.; Matkowskyj, K.A.; Li, H.; Bai, H.; Zhang, W.; Tsao, M.S.; Liao, J.; Yang, G.Y. Overexpression and oncogenic function of aldo-keto reductase family 1B10 (AKR1B10) in pancreatic carcinoma. Mod. Pathol. 2012. [Google Scholar] [CrossRef] [PubMed]
- Matsunaga, T.; Wada, Y.; Endo, S.; Soda, M.; El-Kabbani, O.; Hara, A. Aldo-Keto Reductase 1B10 and Its Role in Proliferation Capacity of Drug-Resistant Cancers. Front. Pharmacol. 2012, 3, 5. [Google Scholar] [CrossRef]
- Matsunaga, T.; Endo, S.; Takemura, M.; Soda, M.; Yamamura, K.; Tajima, K.; Miura, T.; Terada, T.; El-Kabbani, O.; Hara, A. Reduction of cytotoxic p-quinone metabolites of tert-butylhydroquinone by human aldo-keto reductase (AKR) 1B10. Drug Metab. Pharmacokinet. 2012, 27, 553–558. [Google Scholar] [CrossRef]
- Martin, H.J.; Breyer-Pfaff, U.; Wsol, V.; Venz, S.; Block, S.; Maser, E. Purification and characterization of akr1b10 from human liver: Role in carbonyl reduction of xenobiotics. Drug Metab. Dispos. 2006, 34, 464–470. [Google Scholar] [CrossRef] [PubMed]
- Bains, O.S.; Grigliatti, T.A.; Reid, R.E.; Riggs, K.W. Naturally occurring variants of human aldo-keto reductases with reduced in vitro metabolism of daunorubicin and doxorubicin. J. Pharmacol. Exp. Ther. 2010, 335, 533–545. [Google Scholar] [CrossRef]
- Skarydova, L.; Nobilis, M.; Wsol, V. Role of carbonyl reducing enzymes in the phase I biotransformation of the non-steroidal anti-inflammatory drug nabumetone in vitro. Xenobiotica 2013, 43, 346–354. [Google Scholar] [CrossRef] [PubMed]
- Malatkova, P.; Skarka, A.; Musilova, K.; Wsol, V. Reductive metabolism of tiaprofenic acid by the human liver and recombinant carbonyl reducing enzymes. Chem. Biol. Interact. 2017. [Google Scholar] [CrossRef]
- Breyer-Pfaff, U.; Nill, K. Carbonyl reduction of naltrexone and dolasetron by oxidoreductases isolated from human liver cytosol. J. Pharm. Pharmacol. 2004, 56, 1601–1606. [Google Scholar] [CrossRef] [PubMed]
- Quinn, A.M.; Harvey, R.G.; Penning, T.M. Oxidation of PAH trans-Dihydrodiols by Human Aldo-Keto Reductase AKR1B10. Chem. Res. Toxicol. 2008, 21, 2207–2215. [Google Scholar] [CrossRef]
- Conway, L.P.; Garg, N.; Lin, W.; Vujasinovic, M.; Lohr, J.M.; Globisch, D. Chemoselective probe for detailed analysis of ketones and aldehydes produced by gut microbiota in human samples. Chem. Commun. (Camb.) 2019, 55, 9080–9083. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Yan, R.; Zu, X.; Cheng, J.M.; Rao, K.; Liao, D.F.; Cao, D. Aldo-keto Reductase Family 1 B10 Affects Fatty Acid Synthesis by Regulating the Stability of Acetyl-CoA Carboxylase-{alpha} in Breast Cancer Cells. J. Biol. Chem. 2008, 283, 3418–3423. [Google Scholar] [CrossRef] [PubMed]
- Luo, D.X.; Huang, M.C.; Ma, J.; Gao, Z.; Liao, D.F.; Cao, D. Aldo-keto reductase family 1, member B10 is secreted through a lysosome-mediated non-classical pathway. Biochem. J. 2011, 438, 71–80. [Google Scholar] [CrossRef]
- Luo, D.; Bu, Y.; Ma, J.; Rajput, S.; He, Y.; Cai, G.; Liao, D.F.; Cao, D. Heat shock protein 90-alpha mediates aldo-keto reductase 1B10 (AKR1B10) protein secretion through secretory lysosomes. J. Biol. Chem. 2013, 288, 36733–36740. [Google Scholar] [CrossRef]
- Li, W.; Liu, C.; Huang, Z.; Shi, L.; Zhong, C.; Zhou, W.; Meng, P.; Li, Z.; Wang, S.; Luo, F.; et al. AKR1B10 negatively regulates autophagy through reducing GAPDH upon glucose starvation in colon cancer. J. Cell Sci. 2021. [Google Scholar] [CrossRef]
- Shen, Y.; Ma, J.; Yan, R.; Ling, H.; Li, X.; Yang, W.; Gao, J.; Huang, C.; Bu, Y.; Cao, Y.; et al. Impaired self-renewal and increased colitis and dysplastic lesions in colonic mucosa of AKR1B8-deficient mice. Clin. Cancer Res. 2015, 21, 1466–1476. [Google Scholar] [CrossRef]
- Lee, H.J.; Nam, K.T.; Park, H.S.; Kim, M.A.; Lafleur, B.J.; Aburatani, H.; Yang, H.K.; Kim, W.H.; Goldenring, J.R. Gene expression profiling of metaplastic lineages identifies CDH17 as a prognostic marker in early stage gastric cancer. Gastroenterology 2010, 139, 213–225.e3. [Google Scholar] [CrossRef]
- Kropotova, E.S.; Tychko, R.A.; Zinov’eva, O.L.; Zyrianova, A.F.; Khankin, S.L.; Cherkes, V.L.; Aliev, V.A.; Beresten, S.F.; Oparina, N.; Mashkova, T.D. [Downregulation of AKR1B10 gene expression in colorectal cancer]. Mol. Biol. (Mosk.) 2010, 44, 216–222. [Google Scholar] [CrossRef]
- Ohashi, T.; Idogawa, M.; Sasaki, Y.; Suzuki, H.; Tokino, T. AKR1B10, a transcriptional target of p53, is downregulated in colorectal cancers associated with poor prognosis. Mol. Cancer Res. 2013, 11, 1554–1563. [Google Scholar] [CrossRef] [PubMed]
- Kropotova, E.S.; Zinovieva, O.L.; Zyryanova, A.F.; Dybovaya, V.I.; Prasolov, V.S.; Beresten, S.F.; Oparina, N.Y.; Mashkova, T.D. Altered expression of multiple genes involved in retinoic acid biosynthesis in human colorectal cancer. Pathol. Oncol. Res. 2014, 20, 707–717. [Google Scholar] [CrossRef] [PubMed]
- Zu, X.; Yan, R.; Pan, J.; Zhong, L.; Cao, Y.; Ma, J.; Cai, C.; Huang, D.; Liu, J.; Chung, F.L.; et al. Aldo-keto reductase 1B10 protects human colon cells from DNA damage induced by electrophilic carbonyl compounds. Mol. Carcinog. 2017, 56, 118–129. [Google Scholar] [CrossRef]
- Taskoparan, B.; Seza, E.G.; Demirkol, S.; Tuncer, S.; Stefek, M.; Gure, A.O.; Banerjee, S. Opposing roles of the aldo-keto reductases AKR1B1 and AKR1B10 in colorectal cancer. Cell. Oncol. (Dordr.) 2017, 40, 563–578. [Google Scholar] [CrossRef] [PubMed]
- Yao, Y.; Wang, X.; Zhou, D.; Li, H.; Qian, H.; Zhang, J.; Jiang, L.; Wang, B.; Lin, Q.; Zhu, X. Loss of AKR1B10 promotes colorectal cancer cells proliferation and migration via regulating FGF1-dependent pathway. Aging (Albany N. Y.) 2020, 12, 13059–13075. [Google Scholar] [CrossRef]
- Dai, G.P.; Wang, L.P.; Wen, Y.Q.; Ren, X.Q.; Zuo, S.G. Identification of key genes for predicting colorectal cancer prognosis by integrated bioinformatics analysis. Oncol. Lett. 2020, 19, 388–398. [Google Scholar] [CrossRef]
- Kropotova, E.S.; Zinov’eva, O.L.; Zyrianova, A.F.; Choinzonov, E.L.; Afanas’ev, S.G.; Cherdyntseva, N.V.; Beresten, S.F.; Oparina, N.; Mashkova, T.D. [Expression of genes involved in retinoic acid biosynthesis in human gastric cancer]. Mol. Biol. (Mosk.) 2013, 47, 317–330. [Google Scholar] [CrossRef]
- Yao, H.B.; Xu, Y.; Chen, L.G.; Guan, T.P.; Ma, Y.Y.; He, X.J.; Xia, Y.J.; Tao, H.Q.; Shao, Q.S. AKR1B10, a good prognostic indicator in gastric cancer. Eur. J. Surg. Oncol. 2014, 40, 318–324. [Google Scholar] [CrossRef]
- Ahmed, S.M.U.; Jiang, Z.N.; Zheng, Z.H.; Li, Y.; Wang, X.J.; Tang, X. AKR1B10 expression predicts response of gastric cancer to neoadjuvant chemotherapy. Oncol. Lett. 2019, 17, 773–780. [Google Scholar] [CrossRef]
- Osaki, L.H.; Gama, P. MAPKs and signal transduction in the control of gastrointestinal epithelial cell proliferation and differentiation. Int. J. Mol. Sci. 2013, 14, 10143–10161. [Google Scholar] [CrossRef]
- Mashkova, T.D.; Oparina, N.; Zinov’eva, O.L.; Kropotova, E.S.; Dubovaia, V.I.; Poltaraus, A.B.; Fridman, M.V.; Kopantsev, E.P.; Vinogradova, T.V.; Zinov’eva, M.V.; et al. [Transcription TIMP3, DAPk1 and AKR1B10 genes in squamous cell lung cancer]. Mol. Biol. (Mosk.) 2006, 40, 1047–1054. [Google Scholar] [CrossRef]
- Kang, M.W.; Lee, E.S.; Yoon, S.Y.; Jo, J.; Lee, J.; Kim, H.K.; Choi, Y.S.; Kim, K.; Shim, Y.M.; Kim, J.; et al. AKR1B10 is associated with smoking and smoking-related non-small-cell lung cancer. J. Int. Med. Res. 2011, 39, 78–85. [Google Scholar] [CrossRef]
- Szymanowska-Narloch, A.; Jassem, E.; Skrzypski, M.; Muley, T.; Meister, M.; Dienemann, H.; Taron, M.; Rosell, R.; Rzepko, R.; Jarzab, M.; et al. Molecular profiles of non-small cell lung cancers in cigarette smoking and never-smoking patients. Adv. Med. Sci. 2013, 58, 196–206. [Google Scholar] [CrossRef]
- MacLeod, A.K.; Acosta-Jimenez, L.; Coates, P.J.; McMahon, M.; Carey, F.A.; Honda, T.; Henderson, C.J.; Wolf, C.R. Aldo-keto reductases are biomarkers of NRF2 activity and are co-ordinately overexpressed in non-small cell lung cancer. Br. J. Cancer 2016, 115, 1530–1539. [Google Scholar] [CrossRef]
- Kuznetsova, E.S.; Zinovieva, O.L.; Oparina, N.Y.; Prokofjeva, M.M.; Spirin, P.V.; Favorskaya, I.A.; Zborovskaya, I.B.; Lisitsyn, N.A.; Prassolov, V.S.; Mashkova, T.D. [Abnormal expression of genes that regulate retinoid metabolism and signaling in non-small-cell lung cancer]. Mol. Biol. (Mosk.) 2016, 50, 255–265. [Google Scholar] [CrossRef]
- Ludovini, V.; Bianconi, F.; Siggillino, A.; Piobbico, D.; Vannucci, J.; Metro, G.; Chiari, R.; Bellezza, G.; Puma, F.; Della Fazia, M.A.; et al. Gene identification for risk of relapse in stage I lung adenocarcinoma patients: A combined methodology of gene expression profiling and computational gene network analysis. Oncotarget 2016, 7, 30561–30574. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Zhao, Y.; Gu, L.; Niu, X.; Lu, S. Inhibiting proliferation and migration of lung cancer using small interfering RNA targeting on Aldo-keto reductase family 1 member B10. Mol. Med. Rep. 2018, 17, 2153–2160. [Google Scholar] [CrossRef]
- Hung, J.J.; Yeh, Y.C.; Hsu, W.H. Prognostic significance of AKR1B10 in patients with resected lung adenocarcinoma. Thorac. Cancer 2018, 9, 1492–1499. [Google Scholar] [CrossRef]
- Cong, Z.; Diao, Y.; Xu, Y.; Li, X.; Jiang, Z.; Shao, C.; Ji, S.; Shen, Y.; De, W.; Qiang, Y. Long non-coding RNA linc00665 promotes lung adenocarcinoma progression and functions as ceRNA to regulate AKR1B10-ERK signaling by sponging miR-98. Cell Death Dis. 2019, 10, 84. [Google Scholar] [CrossRef]
- Liu, W.; Song, J.; Du, X.; Zhou, Y.; Li, Y.; Li, R.; Lyu, L.; He, Y.; Hao, J.; Ben, J.; et al. AKR1B10 (Aldo-keto reductase family 1 B10) promotes brain metastasis of lung cancer cells in a multi-organ microfluidic chip model. Acta Biomater. 2019, 91, 195–208. [Google Scholar] [CrossRef]
- Ma, J.; Luo, D.X.; Huang, C.; Shen, Y.; Bu, Y.; Markwell, S.; Gao, J.; Liu, J.; Zu, X.; Cao, Z.; et al. AKR1B10 overexpression in breast cancer: Association with tumor size, lymph node metastasis and patient survival and its potential as a novel serum marker. Int. J. Cancer 2012, 131, E862–E871. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; He, X.; Xing, S.; Ni, J.; Zhang, W.; Xu, X.; Gao, F.; Gai, J.; Zhao, Z.; Li, J.; et al. Overexpression of Aldo-keto reductase family 1 B10 protein in ductal carcinoma in situ of the breast correlates with HER2 positivity. Cancer Biomark. 2013, 13, 181–192. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Verhulst, S.; Shen, Y.; Bu, Y.; Cao, Y.; He, Y.; Wang, Y.; Huang, D.; Cai, C.; Rao, K.; et al. AKR1B10 promotes breast cancer metastasis through integrin alpha5/delta-catenin mediated FAK/Src/Rac1 signaling pathway. Oncotarget 2016. [Google Scholar] [CrossRef]
- Reddy, K.A.; Kumar, P.U.; Srinivasulu, M.; Triveni, B.; Sharada, K.; Ismail, A.; Reddy, G.B. Overexpression and enhanced specific activity of aldoketo reductases (AKR1B1 & AKR1B10) in human breast cancers. Breast 2017, 31, 137–143. [Google Scholar] [CrossRef]
- Li, J.; Guo, Y.; Duan, L.; Hu, X.; Zhang, X.; Hu, J.; Huang, L.; He, R.; Hu, Z.; Luo, W.; et al. AKR1B10 promotes breast cancer cell migration and invasion via activation of ERK signaling. Oncotarget 2017, 8, 33694–33703. [Google Scholar] [CrossRef]
- Van Weverwijk, A.; Koundouros, N.; Iravani, M.; Ashenden, M.; Gao, Q.; Poulogiannis, G.; Jungwirth, U.; Isacke, C.M. Metabolic adaptability in metastatic breast cancer by AKR1B10-dependent balancing of glycolysis and fatty acid oxidation. Nat. Commun. 2019, 10, 2698. [Google Scholar] [CrossRef]
- Connor, J.P.; Esbona, K.; Matkowskyj, K.A. AKR1B10 expression by immunohistochemistry in surgical resections and fine needle aspiration cytology material in patients with cystic pancreatic lesions; potential for improved nonoperative diagnosis. Hum. Pathol. 2017, 70, 77–83. [Google Scholar] [CrossRef]
- Fang, C.Y.; Lin, Y.H.; Chen, C.L. Overexpression of AKR1B10 predicts tumor recurrence and short survival in oral squamous cell carcinoma patients. J. Oral Pathol. Med. 2019, 48, 712–719. [Google Scholar] [CrossRef]
- Ko, H.H.; Cheng, S.L.; Lee, J.J.; Chen, H.M.; Kuo, M.Y.; Cheng, S.J. Expression of AKR1B10 as an independent marker for poor prognosis in human oral squamous cell carcinoma. Head Neck 2017, 39, 1327–1332. [Google Scholar] [CrossRef]
- El-Hariry, I.; Pignatelli, M.; Lemoine, N. Fibroblast growth factor 1 and fibroblast growth factor 2 immunoreactivity in gastrointestinal tumours. J. Pathol. 1997, 181, 39–45. [Google Scholar] [CrossRef]
- Henriksson, M.L.; Edin, S.; Dahlin, A.M.; Oldenborg, P.A.; Oberg, A.; Van Guelpen, B.; Rutegard, J.; Stenling, R.; Palmqvist, R. Colorectal cancer cells activate adjacent fibroblasts resulting in FGF1/FGFR3 signaling and increased invasion. Am. J. Pathol. 2011, 178, 1387–1394. [Google Scholar] [CrossRef]
- Liu, N.; Zhang, J.; Sun, S.; Yang, L.; Zhou, Z.; Sun, Q.; Niu, J. Expression and clinical significance of fibroblast growth factor 1 in gastric adenocarcinoma. Onco Targets Ther. 2015, 8, 615–621. [Google Scholar] [CrossRef]
- Matsunaga, T.; Suzuki, A.; Kezuka, C.; Okumura, N.; Iguchi, K.; Inoue, I.; Soda, M.; Endo, S.; El-Kabbani, O.; Hara, A.; et al. Aldo-keto reductase 1B10 promotes development of cisplatin resistance in gastrointestinal cancer cells through down-regulating peroxisome proliferator-activated receptor-gamma-dependent mechanism. Chem. Biol. Interact. 2016, 256, 142–153. [Google Scholar] [CrossRef]
- DiStefano, J.K.; Davis, B. Diagnostic and Prognostic Potential of AKR1B10 in Human Hepatocellular Carcinoma. Cancers 2019, 11, 486. [Google Scholar] [CrossRef]
- Liu, Y.; Zhang, J.; Liu, H.; Guan, G.; Zhang, T.; Wang, L.; Qi, X.; Zheng, H.; Chen, C.C.; Liu, J.; et al. Compensatory upregulation of aldo-keto reductase 1B10 to protect hepatocytes against oxidative stress during hepatocarcinogenesis. Am. J. Cancer Res. 2019, 9, 2730–2748. [Google Scholar]
- Wang, T.; Zhang, K.H. New Blood Biomarkers for the Diagnosis of AFP-Negative Hepatocellular Carcinoma. Front. Oncol. 2020, 10, 1316. [Google Scholar] [CrossRef]
- Raza, S.; Rajak, S.; Anjum, B.; Sinha, R.A. Molecular links between non-alcoholic fatty liver disease and hepatocellular carcinoma. Hepatoma Res. 2019, 5, 42. [Google Scholar] [CrossRef]
- Starmann, J.; Falth, M.; Spindelbock, W.; Lanz, K.L.; Lackner, C.; Zatloukal, K.; Trauner, M.; Sultmann, H. Gene expression profiling unravels cancer-related hepatic molecular signatures in steatohepatitis but not in steatosis. PLoS ONE 2012, 7, e46584. [Google Scholar] [CrossRef]
- Tsuzura, H.; Genda, T.; Sato, S.; Murata, A.; Kanemitsu, Y.; Narita, Y.; Ishikawa, S.; Kikuchi, T.; Mori, M.; Hirano, K.; et al. Expression of aldo-keto reductase family 1 member b10 in the early stages of human hepatocarcinogenesis. Int. J. Mol. Sci. 2014, 15, 6556–6568. [Google Scholar] [CrossRef]
- Bitter, A.; Rummele, P.; Klein, K.; Kandel, B.A.; Rieger, J.K.; Nussler, A.K.; Zanger, U.M.; Trauner, M.; Schwab, M.; Burk, O. Pregnane X receptor activation and silencing promote steatosis of human hepatic cells by distinct lipogenic mechanisms. Arch. Toxicol. 2015, 89, 2089–2103. [Google Scholar] [CrossRef]
- Pettinelli, P.; Arendt, B.M.; Teterina, A.; McGilvray, I.; Comelli, E.M.; Fung, S.K.; Fischer, S.E.; Allard, J.P. Altered hepatic genes related to retinol metabolism and plasma retinol in patients with non-alcoholic fatty liver disease. PLoS ONE 2018, 13, e0205747. [Google Scholar] [CrossRef]
- Arendt, B.M.; Teterina, A.; Pettinelli, P.; Comelli, E.M.; Ma, D.W.L.; Fung, S.K.; McGilvray, I.D.; Fischer, S.E.; Allard, J.P. Cancer-related gene expression is associated with disease severity and modifiable lifestyle factors in non-alcoholic fatty liver disease. Nutrition 2019, 62, 100–107. [Google Scholar] [CrossRef]
- Kanno, M.; Kawaguchi, K.; Honda, M.; Horii, R.; Takatori, H.; Shimakami, T.; Kitamura, K.; Arai, K.; Yamashita, T.; Sakai, Y.; et al. Serum aldo-keto reductase family 1 member B10 predicts advanced liver fibrosis and fatal complications of nonalcoholic steatohepatitis. J. Gastroenterol. 2019, 54, 549–557. [Google Scholar] [CrossRef] [PubMed]
- Feng, G.; Li, X.P.; Niu, C.Y.; Liu, M.L.; Yan, Q.Q.; Fan, L.P.; Li, Y.; Zhang, K.L.; Gao, J.; Qian, M.R.; et al. Bioinformatics analysis reveals novel core genes associated with nonalcoholic fatty liver disease and nonalcoholic steatohepatitis. Gene 2020, 742, 144549. [Google Scholar] [CrossRef]
- Govaere, O.; Cockell, S.; Tiniakos, D.; Queen, R.; Younes, R.; Vacca, M.; Alexander, L.; Ravaioli, F.; Palmer, J.; Petta, S.; et al. Transcriptomic profiling across the nonalcoholic fatty liver disease spectrum reveals gene signatures for steatohepatitis and fibrosis. Sci. Transl. Med. 2020, 12. [Google Scholar] [CrossRef] [PubMed]
- Zeng, F.; Zhang, Y.; Han, X.; Zeng, M.; Gao, Y.; Weng, J. Predicting Non-Alcoholic Fatty Liver Disease Progression and Immune Deregulations by Specific Gene Expression Patterns. Front. Immunol. 2020, 11, 609900. [Google Scholar] [CrossRef] [PubMed]
- Tebay, L.E.; Robertson, H.; Durant, S.T.; Vitale, S.R.; Penning, T.M.; Dinkova-Kostova, A.T.; Hayes, J.D. Mechanisms of activation of the transcription factor Nrf2 by redox stressors, nutrient cues, and energy status and the pathways through which it attenuates degenerative disease. Free Radic. Biol. Med. 2015, 88, 108–146. [Google Scholar] [CrossRef]
- Cullinan, S.B.; Zhang, D.; Hannink, M.; Arvisais, E.; Kaufman, R.J.; Diehl, J.A. Nrf2 is a direct PERK substrate and effector of PERK-dependent cell survival. Mol. Cell. Biol. 2003, 23, 7198–7209. [Google Scholar] [CrossRef]
- Cubillos-Angulo, J.M.; Fukutani, E.R.; Cruz, L.A.B.; Arriaga, M.B.; Lima, J.V.; Andrade, B.B.; Queiroz, A.T.L.; Fukutani, K.F. Systems biology analysis of publicly available transcriptomic data reveals a critical link between AKR1B10 gene expression, smoking and occurrence of lung cancer. PLoS ONE 2020, 15, e0222552. [Google Scholar] [CrossRef]
- Li, C.P.; Goto, A.; Watanabe, A.; Murata, K.; Ota, S.; Niki, T.; Aburatani, H.; Fukayama, M. AKR1B10 in usual interstitial pneumonia: Expression in squamous metaplasia in association with smoking and lung cancer. Pathol. Res. Pract. 2008, 204, 295–304. [Google Scholar] [CrossRef]
- Huang, C.; Cao, Z.; Ma, J.; Shen, Y.; Bu, Y.; Khoshaba, R.; Shi, G.; Huang, D.; Liao, D.F.; Ji, H.; et al. AKR1B10 activates diacylglycerol (DAG) second messenger in breast cancer cells. Mol. Carcinog. 2018, 57, 1300–1310. [Google Scholar] [CrossRef]
- Hayes, A.J.; Skouras, C.; Haugk, B.; Charnley, R.M. Keap1-Nrf2 signalling in pancreatic cancer. Int. J. Biochem. Cell Biol. 2015, 65, 288–299. [Google Scholar] [CrossRef] [PubMed]
- Ko, H.H.; Peng, H.H.; Cheng, S.J.; Kuo, M.Y. Increased salivary AKR1B10 level: Association with progression and poor prognosis of oral squamous cell carcinoma. Head Neck 2018, 40, 2642–2647. [Google Scholar] [CrossRef] [PubMed]
- Nagaraj, N.S.; Beckers, S.; Mensah, J.K.; Waigel, S.; Vigneswaran, N.; Zacharias, W. Cigarette smoke condensate induces cytochromes P450 and aldo-keto reductases in oral cancer cells. Toxicol. Lett. 2006, 165, 182–194. [Google Scholar] [CrossRef]
- Yoshitake, H.; Takahashi, M.; Ishikawa, H.; Nojima, M.; Iwanari, H.; Watanabe, A.; Aburatani, H.; Yoshida, K.; Ishi, K.; Takamori, K.; et al. Aldo-keto reductase family 1, member B10 in uterine carcinomas: A potential risk factor of recurrence after surgical therapy in cervical cancer. Int. J. Gynecol. Cancer 2007, 17, 1300–1306. [Google Scholar] [CrossRef]
- Hevir, N.; Sinkovec, J.; Lanisnik Rizner, T. Decreased levels of AKR1B1 and AKR1B10 in cancerous endometrium compared to adjacent non-cancerous tissue. Chem. Biol. Interact. 2013, 202, 226–233. [Google Scholar] [CrossRef]
- Sinreih, M.; Stupar, S.; Cemazar, L.; Verdenik, I.; Frkovic Grazio, S.; Smrkolj, S.; Rizner, T.L. STAR and AKR1B10 are down-regulated in high-grade endometrial cancer. J. Steroid Biochem. Mol. Biol. 2017, 171, 43–53. [Google Scholar] [CrossRef]
- Ooi, A.; Wong, J.C.; Petillo, D.; Roossien, D.; Perrier-Trudova, V.; Whitten, D.; Min, B.W.; Tan, M.H.; Zhang, Z.; Yang, X.J.; et al. An antioxidant response phenotype shared between hereditary and sporadic type 2 papillary renal cell carcinoma. Cancer Cell 2011, 20, 511–523. [Google Scholar] [CrossRef]
- He, Y.C.; Shen, Y.; Cao, Y.; Tang, F.Q.; Tian, D.F.; Huang, C.F.; Tao, H.; Zhou, F.L.; Zhang, B.; Song, L.; et al. Overexpression of AKR1B10 in nasopharyngeal carcinoma as a potential biomarker. Cancer Biomark. 2016, 16, 127–135. [Google Scholar] [CrossRef]
- Guo, Y.; Luo, W.; Hu, Z.; Li, J.; Li, X.; Cao, H.; Li, J.; Wen, B.; Zhang, J.; Cheng, H.; et al. Low expression of Aldo-keto reductase 1B10 is a novel independent prognostic indicator for nasopharyngeal carcinoma. Cell Biosci. 2016, 6, 18. [Google Scholar] [CrossRef]
- Liu, X.; Hu, Z.; Qu, J.; Li, J.; Gong, K.; Wang, L.; Jiang, J.; Li, X.; He, R.; Duan, L.; et al. AKR1B10 confers resistance to radiotherapy via FFA/TLR4/NF-κB axis in nasopharyngeal carcinoma. Int. J. Biol. Sci. 2021, 17, 756–767. [Google Scholar] [CrossRef] [PubMed]
- Breton, J.; Gage, M.C.; Hay, A.W.; Keen, J.N.; Wild, C.P.; Donnellan, C.; Findlay, J.B.; Hardie, L.J. Proteomic screening of a cell line model of esophageal carcinogenesis identifies cathepsin D and aldo-keto reductase 1C2 and 1B10 dysregulation in Barrett’s esophagus and esophageal adenocarcinoma. J. Proteome Res. 2008, 7, 1953–1962. [Google Scholar] [CrossRef] [PubMed]
- Nancarrow, D.J.; Clouston, A.D.; Smithers, B.M.; Gotley, D.C.; Drew, P.A.; Watson, D.I.; Tyagi, S.; Hayward, N.K.; Whiteman, D.C.; Australian Cancer, S.; et al. Whole genome expression array profiling highlights differences in mucosal defense genes in Barrett’s esophagus and esophageal adenocarcinoma. PLoS ONE 2011, 6, e22513. [Google Scholar] [CrossRef]
- Sumantran, V.N.; Mishra, P.; Bera, R.; Sudhakar, N. Microarray Analysis of Differentially-Expressed Genes Encoding CYP450 and Phase II Drug Metabolizing Enzymes in Psoriasis and Melanoma. Pharmaceutics 2016, 8, 4. [Google Scholar] [CrossRef]
- Gao, Y.; Yi, X.; Ding, Y. Combined Transcriptomic Analysis Revealed AKR1B10 Played an Important Role in Psoriasis through the Dysregulated Lipid Pathway and Overproliferation of Keratinocyte. Biomed. Res. Int. 2017, 2017, 8717369. [Google Scholar] [CrossRef]
- Jumper, N.; Hodgkinson, T.; Arscott, G.; Har-Shai, Y.; Paus, R.; Bayat, A. The Aldo-Keto Reductase AKR1B10 Is Up-Regulated in Keloid Epidermis, Implicating Retinoic Acid Pathway Dysregulation in the Pathogenesis of Keloid Disease. J. Investig. Dermatol. 2016, 136, 1500–1512. [Google Scholar] [CrossRef]
- Jumper, N.; Hodgkinson, T.; Paus, R.; Bayat, A. Site-specific gene expression profiling as a novel strategy for unravelling keloid disease pathobiology. PLoS ONE 2017, 12, e0172955. [Google Scholar] [CrossRef]
- Ghosh, D.; Ding, L.; Sivaprasad, U.; Geh, E.; Biagini Myers, J.; Bernstein, J.A.; Khurana Hershey, G.K.; Mersha, T.B. Multiple Transcriptome Data Analysis Reveals Biologically Relevant Atopic Dermatitis Signature Genes and Pathways. PLoS ONE 2015, 10, e0144316. [Google Scholar] [CrossRef]
- Soares, C.T.; Fachin, L.R.V.; Trombone, A.P.F.; Rosa, P.S.; Ghidella, C.C.; Belone, A.F.F. Potential of AKR1B10 as a Biomarker and Therapeutic Target in Type 2 Leprosy Reaction. Front. Med. (Lausanne) 2018, 5, 263. [Google Scholar] [CrossRef]
- Matsunaga, T.; El-Kabbani, O.; Hara, A. Aldo-keto reductases as new therapeutic targets for colon cancer chemoresistance. In Molecular Mechanisms of Tumor Cell Resistance to Chemotherapy: Targeted Therapies to Reverse Resistance; Bonavida, B., Ed.; Springer: New York, NY, USA, 2013; pp. 109–133. [Google Scholar]
- Penning, T.M. Aldo-Keto Reductase Regulation by the Nrf2 System: Implications for Stress Response, Chemotherapy Drug Resistance, and Carcinogenesis. Chem. Res. Toxicol. 2017, 30, 162–176. [Google Scholar] [CrossRef]
- Hashimoto, Y.; Imanishi, K.; Tokui, N.; Okamoto, T.; Okamoto, A.; Hatakeyama, S.; Yoneyama, T.; Koie, T.; Kamimura, N.; Ohyama, C. Carboplatin-gemcitabine combination chemotherapy upregulates AKR1B10 expression in bladder cancer. Int. J. Clin. Oncol. 2013, 18, 177–182. [Google Scholar] [CrossRef] [PubMed]
- Matsunaga, T.; Yamane, Y.; Iida, K.; Endo, S.; Banno, Y.; El-Kabbani, O.; Hara, A. Involvement of the aldo-keto reductase, AKR1B10, in mitomycin-c resistance through reactive oxygen species-dependent mechanisms. Anticancer Drugs 2011, 22, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Matsunaga, T.; Kawabata, S.; Yanagihara, Y.; Kezuka, C.; Kato, M.; Morikawa, Y.; Endo, S.; Chen, H.; Iguchi, K.; Ikari, A. Pathophysiological roles of autophagy and aldo-keto reductases in development of doxorubicin resistance in gastrointestinal cancer cells. Chem. Biol. Interact. 2019, 314, 108839. [Google Scholar] [CrossRef] [PubMed]
- Matsunaga, T.; Saito, H.; Endo, S.; Iguchi, K.; Soda, M.; El-Kabbani, O.; Hara, A.; Ikari, A. Roles of aldo-keto reductases 1B10 and 1C3 and ATP-binding cassette transporter in docetaxel tolerance. Free Radic. Res. 2016, 50, 1296–1308. [Google Scholar] [CrossRef]
- Heibein, A.D.; Guo, B.; Sprowl, J.A.; Maclean, D.A.; Parissenti, A.M. Role of aldo-keto reductases and other doxorubicin pharmacokinetic genes in doxorubicin resistance, DNA binding, and subcellular localization. BMC Cancer 2012, 12, 381. [Google Scholar] [CrossRef]
- Bacolod, M.D.; Lin, S.M.; Johnson, S.P.; Bullock, N.S.; Colvin, M.; Bigner, D.D.; Friedman, H.S. The gene expression profiles of medulloblastoma cell lines resistant to preactivated cyclophosphamide. Curr. Cancer Drug Targets 2008, 8, 172–179. [Google Scholar] [CrossRef][Green Version]
- Singh, A.; Misra, V.; Thimmulappa, R.K.; Lee, H.; Ames, S.; Hoque, M.O.; Herman, J.G.; Baylin, S.B.; Sidransky, D.; Gabrielson, E.; et al. Dysfunctional KEAP1-NRF2 interaction in non-small-cell lung cancer. PLoS Med. 2006, 3, e420. [Google Scholar] [CrossRef]
- Shibata, T.; Kokubu, A.; Gotoh, M.; Ojima, H.; Ohta, T.; Yamamoto, M.; Hirohashi, S. Genetic alteration of Keap1 confers constitutive Nrf2 activation and resistance to chemotherapy in gallbladder cancer. Gastroenterology 2008, 135, 1358–1368. [Google Scholar] [CrossRef]
- Kang, J.S.; Nam, L.B.; Yoo, O.K.; Keum, Y.S. Molecular mechanisms and systemic targeting of NRF2 dysregulation in cancer. Biochem. Pharmacol. 2020, 177, 114002. [Google Scholar] [CrossRef]
- Zhong, L.; Shen, H.; Huang, C.; Jing, H.; Cao, D. AKR1B10 induces cell resistance to daunorubicin and idarubicin by reducing C13 ketonic group. Toxicol. Appl. Pharmacol. 2011, 255, 40–47. [Google Scholar] [CrossRef]
- Yan, R.; Zu, X.; Ma, J.; Liu, Z.; Adeyanju, M.; Cao, D. Aldo-keto reductase family 1 B10 gene silencing results in growth inhibition of colorectal cancer cells: Implication for cancer intervention. Int. J. Cancer 2007, 121, 2301–2306. [Google Scholar] [CrossRef]
- Huang, L.; He, R.; Luo, W.; Zhu, Y.S.; Li, J.; Tan, T.; Zhang, X.; Hu, Z.; Luo, D. Aldo-Keto Reductase Family 1 Member B10 Inhibitors: Potential Drugs for Cancer Treatment. Recent Pat. Anticancer Drug Discov. 2016, 11, 184–196. [Google Scholar] [CrossRef]
- Soda, M.; Hu, D.; Endo, S.; Takemura, M.; Li, J.; Wada, R.; Ifuku, S.; Zhao, H.T.; El-Kabbani, O.; Ohta, S.; et al. Design, synthesis and evaluation of caffeic acid phenethyl ester-based inhibitors targeting a selectivity pocket in the active site of human aldo-keto reductase 1B10. Eur. J. Med. Chem. 2012, 48, 321–329. [Google Scholar] [CrossRef]
- Takemura, M.; Endo, S.; Matsunaga, T.; Soda, M.; Zhao, H.T.; El-Kabbani, O.; Tajima, K.; Iinuma, M.; Hara, A. Selective inhibition of the tumor marker aldo-keto reductase family member 1B10 by oleanolic acid. J. Nat. Prod. 2011, 74, 1201–1206. [Google Scholar] [CrossRef]
- Endo, S.; Matsunaga, T.; Soda, M.; Tajima, K.; Zhao, H.T.; El-Kabbani, O.; Hara, A. Selective inhibition of the tumor marker AKR1B10 by antiinflammatory N-phenylanthranilic acids and glycyrrhetic acid. Biol. Pharm. Bull. 2010, 33, 886–890. [Google Scholar] [CrossRef]
- Hintzpeter, J.; Seliger, J.M.; Hofman, J.; Martin, H.J.; Wsol, V.; Maser, E. Inhibition of human anthracycline reductases by emodin—A possible remedy for anthracycline resistance. Toxicol. Appl. Pharmacol. 2016, 293, 21–29. [Google Scholar] [CrossRef]
- Hara, A.; Endo, S.; Matsunaga, T.; Soda, M.; El-Kabbani, O.; Yashiro, K. Inhibition of aldo-keto reductase family 1 member B10 by unsaturated fatty acids. Arch. Biochem. Biophys. 2016, 609, 69–76. [Google Scholar] [CrossRef]
- Seliger, J.M.; Cicek, S.S.; Witt, L.T.; Martin, H.J.; Maser, E.; Hintzpeter, J. Selective Inhibition of Human AKR1B10 by n-Humulone, Adhumulone and Cohumulone Isolated from Humulus lupulus Extract. Molecules 2018, 23, 3041. [Google Scholar] [CrossRef]
- Seliger, J.M.; Misuri, L.; Maser, E.; Hintzpeter, J. The hop-derived compounds xanthohumol, isoxanthohumol and 8-prenylnaringenin are tight-binding inhibitors of human aldo-keto reductases 1B1 and 1B10. J. Enzyme Inhib. Med. Chem. 2018, 33, 607–614. [Google Scholar] [CrossRef]
- Bateman, R.L.; Rauh, D.; Tavshanjian, B.; Shokat, K.M. Human carbonyl reductase 1 is an S-nitrosoglutathione reductase. J. Biol. Chem. 2008, 283, 35756–35762. [Google Scholar] [CrossRef]
- Hara, A.; Endo, S.; Matsunaga, T.; El-Kabbani, O.; Miura, T.; Nishinaka, T.; Terada, T. Human carbonyl reductase 1 participating in intestinal first-pass drug metabolism is inhibited by fatty acids and acyl-CoAs. Biochem. Pharmacol. 2017, 138, 185–192. [Google Scholar] [CrossRef] [PubMed]
- Hara, A.; Endo, S.; Matsunaga, T.; Soda, M.; Yashiro, K.; El-Kabbani, O. Long-chain fatty acids inhibit human members of the aldo-keto reductase 1C subfamily. J. Biochem. 2017, 162, 371–379. [Google Scholar] [CrossRef]
- Endo, S.; Xia, S.; Suyama, M.; Morikawa, Y.; Oguri, H.; Hu, D.; Ao, Y.; Takahara, S.; Horino, Y.; Hayakawa, Y.; et al. Synthesis of Potent and Selective Inhibitors of Aldo-Keto Reductase 1B10 and Their Efficacy against Proliferation, Metastasis, and Cisplatin Resistance of Lung Cancer Cells. J. Med. Chem. 2017, 60, 8441–8455. [Google Scholar] [CrossRef]
- Cousido-Siah, A.; Ruiz, F.X.; Fanfrlik, J.; Gimenez-Dejoz, J.; Mitschler, A.; Kamlar, M.; Vesely, J.; Ajani, H.; Pares, X.; Farres, J.; et al. IDD388 Polyhalogenated Derivatives as Probes for an Improved Structure-Based Selectivity of AKR1B10 Inhibitors. ACS Chem. Biol. 2016, 11, 2693–2705. [Google Scholar] [CrossRef]
- Chen, W.; Chen, X.; Zhou, S.; Zhang, H.; Wang, L.; Xu, J.; Hu, X.; Yin, W.; Yan, G.; Zhang, J. Design and synthesis of polyhydroxy steroids as selective inhibitors against AKR1B10 and molecular docking. Steroids 2016, 110, 1–8. [Google Scholar] [CrossRef]
- Geng, N.; Jin, Y.Y.; Zhu, S.X.; Li, Y.R.; Zheng, L.Y.; Zhu, W.J.; Li, Y.W.; Han, C.; Dou, X.G.; Bai, H. [Aldo-keto reductase family 1 B10 participates in the regulation of hepatoma cell cycle through p27/p-Rb signaling pathway]. Zhonghua Gan Zang Bing Za Zhi 2020, 28, 861–867. [Google Scholar] [CrossRef]
- Shi, L.; Guo, S.; Zhang, S.; Gao, X.; Liu, A.; Wang, Q.; Zhang, T.; Zhang, Y.; Wen, A. Glycyrrhetinic acid attenuates disturbed vitamin a metabolism in non-alcoholic fatty liver disease through AKR1B10. Eur. J. Pharmacol. 2020, 883, 173167. [Google Scholar] [CrossRef]
- Gimenez-Dejoz, J.; Weber, S.; Barski, O.A.; Moller, G.; Adamski, J.; Pares, X.; Porte, S.; Farres, J. Characterization of AKR1B16, a novel mouse aldo-keto reductase. Chem. Biol. Interact. 2017, 276, 182–193. [Google Scholar] [CrossRef]
- Pastel, E.; Pointud, J.C.; Volat, F.; Martinez, A.; Lefrancois-Martinez, A.M. Aldo-Keto Reductases 1B in Endocrinology and Metabolism. Front. Pharmacol. 2012, 3, 148. [Google Scholar] [CrossRef] [PubMed]

| Agent * | Signal Molecule | Cell ** | References |
|---|---|---|---|
| Up-regulation | |||
| Ethoxyquin | Nrf2 | Lung cancer A549, H23 | [14] |
| MG-132, bortezomib | Nrf2 | CRC SW-480, HT29 | [17] |
| Doxorubicin | Nrf2 | Gastric cancer MKN45 | [18] |
| EGF, insulin | AP-1, ERK | HCC HepG2, Hep3B | [19] |
| Lipopolysaccharide | Blood mononuclear cells | [20] | |
| BMP, IBMX | Mesenchymal stem cells | [21] | |
| 9,10-Phenanthrenequinone | Nrf2, ERK | Lung cancer A549 | [22] |
| Cigarette smoke extract | Airway epithelium | [23] | |
| Carnosic acid, t-BHQ | Nrf2 | Astrocytoma U373MG | [15] |
| 5-FU, L-OHP | p53 | CRC HT116 | [16] |
| Down-regulation | |||
| TPA | c-Jun, ERK | Lung cancer A549 | [15] |
| 5-FU, L-OHP | CRC HT29 | [16] |
| Enzyme | Km (μM) | kcat (min−1) | kcat/Km (min−1 μM−1) | Assay Condition (pH and °C) | References |
|---|---|---|---|---|---|
| AKR1B10 | 30.9 | 121 | 3.9 | pH 7.0, 35 °C | [45] |
| 31 | 119 | 3.8 | pH 7.0, 35 °C | [47] | |
| 4.7 | 27 | 6.1 | pH 7.4, 25 °C | [48] | |
| AKR1B1 | 22 | 102 | 4.6 | pH 7.0, 25 °C | [49] |
| 31 | 16.6 | 1.2 | pH 7.0, 25 °C | [50] | |
| 716 | 50 | 0.07 | pH 6.4, 0.3 M Li2SO4, 35 °C | [47] | |
| AKR1B15 | 2.2 | 5.2 | 2.5 | pH 7.0, 25 °C | [5] |
| AKR1C1 | 34 | 8.8 | 0.27 | pH 7.0, 37 °C | [51] |
| ALDH1A1 | 27 | 10 | 0.38 | pH 8.0, 25 °C | [40] |
| ALDH1A2 | 7.5 | 31 | 4.1 | pH 8.0, 25 °C | [40] |
| ALDH1A3 | 40 | 7.4 | 0.185 | pH 8.0, 25 °C | [40] |
| Organ | Cancer Type * | Assay | AKR1B10 Level ** | Prognosis | References | |
|---|---|---|---|---|---|---|
| Lesion | Serum | |||||
| Colorectum | CRC, ADC | mRNA, protein | Low | Poor | [68,70,71,72,73,74,75,76] | |
| mRNA | Low | Poor | [24] | |||
| Stomach | ADC | mRNA, protein | Low | Poor | [69,77,78,79] | |
| Lung | NSCLC, SCC, ADC | mRNA, protein | High | Poor | [8,81,82,83,84,85,86,87,88,89,90] | |
| protein | High | [90] | ||||
| Breast | ADC, ductal carcinoma | mRNA, protein | High | Poor | [91,92,93,94,95,96] | |
| protein | High | Poor | [91] | |||
| Pancreas | ADC, MCT | protein | High | [55,97] | ||
| Oral cavity | SCC | protein | High | Poor | [98,99] | |
| Cell | Drug * | Suggested Role of AKR1B10 | References | |
|---|---|---|---|---|
| CRC | HT29 cell | L-OHP | Promotion of cell proliferation by modulating isoprenoid metabolism. | [55] |
| HT29 cell | MMC | Detoxification of RCS and drug metabolism | [143] | |
| Lovo cell | CDDP | Detoxification of RCS and down-regulation of PPARγ | [103] | |
| Lovo cell | DOX | Autophagy suppression by detoxification of RCS | [144] | |
| Gastric cancer MKN45 cell | CDDP | Detoxification of RCS and down-regulation of PPARγ | [103] | |
| DOX | Detoxification of RCS and elevation of migrating and invasive potentials through MMP2 induction | [18] | ||
| Lung cancer A549 cell | CDDP | NO production by detoxification of RCS | [48] | |
| DTX | Detoxification of RCS | [145] | ||
| Breast cancer MCF-7 cell | DOX | Drug metabolism | [146] | |
| Prostate cancer Du145 cell | DTX | Detoxification of RCS | [145] | |
| Medulloblastoma D341 MED cell | CPA | Metabolism of a reactive metabolite, aldophosphamide | [147] | |
| Inhibitor * | IC50 (µM) ** | IC50 Ratio (1B10/1B1) | References | |
|---|---|---|---|---|
| AKR1B10 | AKR1B1 | |||
| HCCFA | 0.0035 | 0.277 | 79 | [164] |
| HAHE | 0.0062 | 4.9 | 790 | [154] |
| MK204 | 0.080 | 21.7 | 271 | [165] |
| Oleanolic acid | 0.090 | 124 | 1370 | [155] |
| Epalrestat | 0.33 | 0.021 | 0.06 | [156] |
| Androst-3β,5α,6β,19-tetrol | 0.83 | >100 | 120 | [166] |
| Emodin | 0.99 | 12 | 12 | [157] |
| Arachidonic acid | 1.1 | 24 | 22 | [158] |
| Cohumulone | 1.35 | >125 | >93 | [159] |
| 8-Prenylnaringenin | 3.96 | 1.87 | 0.47 | [160] |
| Glycyrrhetinic acid | 4.9 | 280 | 57 | [156] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Endo, S.; Matsunaga, T.; Nishinaka, T. The Role of AKR1B10 in Physiology and Pathophysiology. Metabolites 2021, 11, 332. https://doi.org/10.3390/metabo11060332
Endo S, Matsunaga T, Nishinaka T. The Role of AKR1B10 in Physiology and Pathophysiology. Metabolites. 2021; 11(6):332. https://doi.org/10.3390/metabo11060332
Chicago/Turabian StyleEndo, Satoshi, Toshiyuki Matsunaga, and Toru Nishinaka. 2021. "The Role of AKR1B10 in Physiology and Pathophysiology" Metabolites 11, no. 6: 332. https://doi.org/10.3390/metabo11060332
APA StyleEndo, S., Matsunaga, T., & Nishinaka, T. (2021). The Role of AKR1B10 in Physiology and Pathophysiology. Metabolites, 11(6), 332. https://doi.org/10.3390/metabo11060332






