Serum Metabolomic and Lipoprotein Profiling of Pancreatic Ductal Adenocarcinoma Patients of African Ancestry
Abstract
:1. Introduction
2. Results
2.1. Patients’ Demographic and Clinicopathological Characteristics
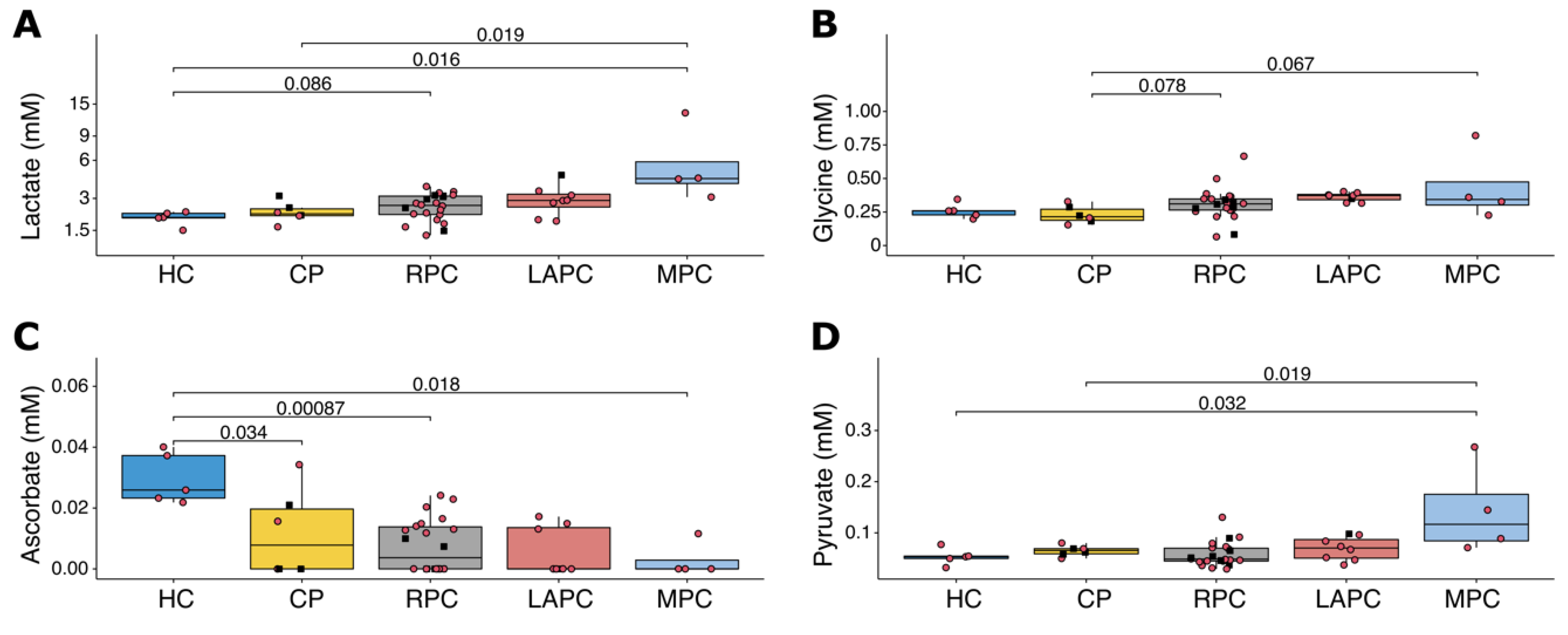
2.2. Metabolic and Lipoprotein Signatures in the Different Tumour Stages
2.3. Dysregulated Metabolites in Patient Survival
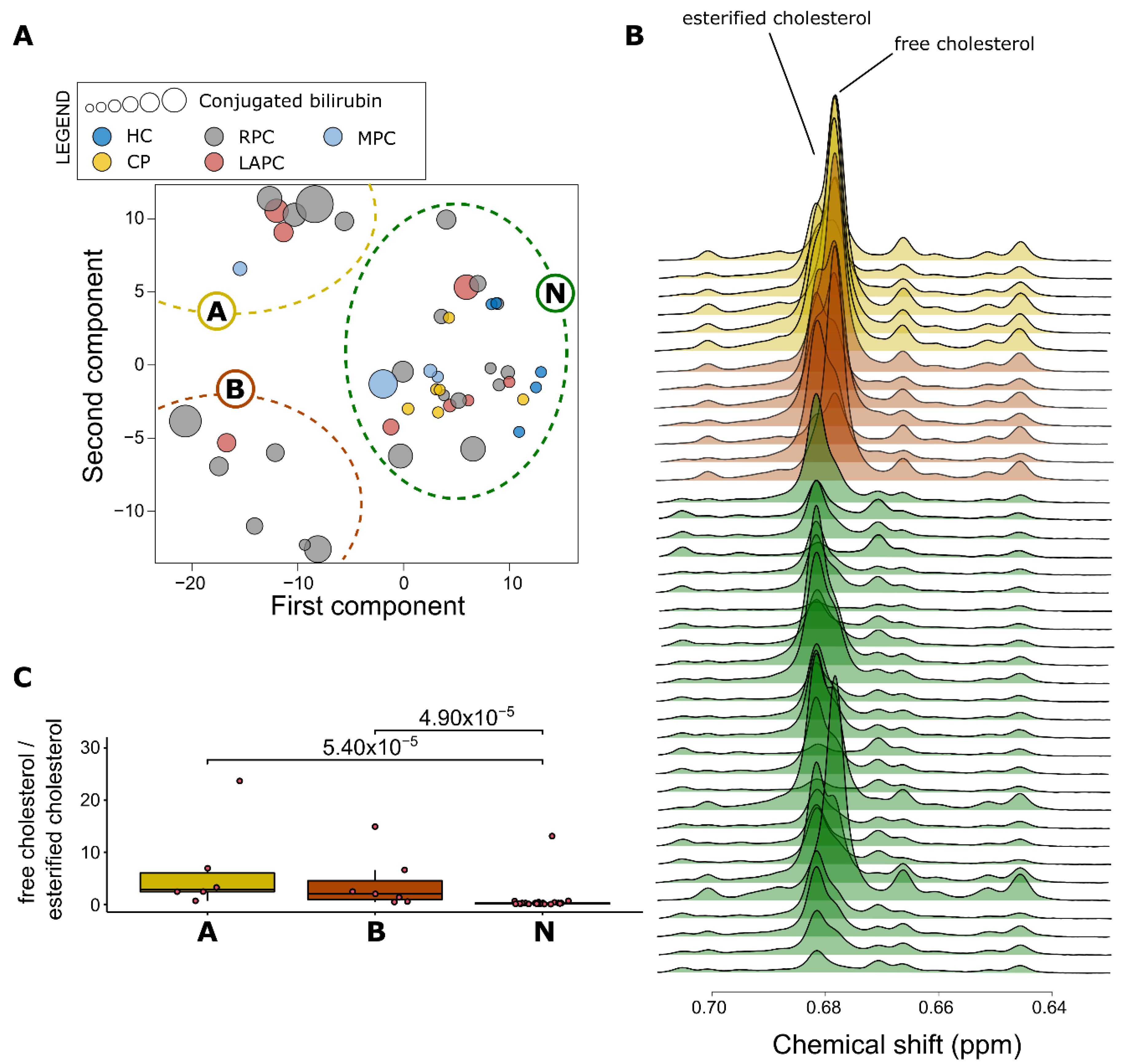
2.4. Impact of Raised Bilirubin Levels on Metabolites and Lipoproteins in PDAC
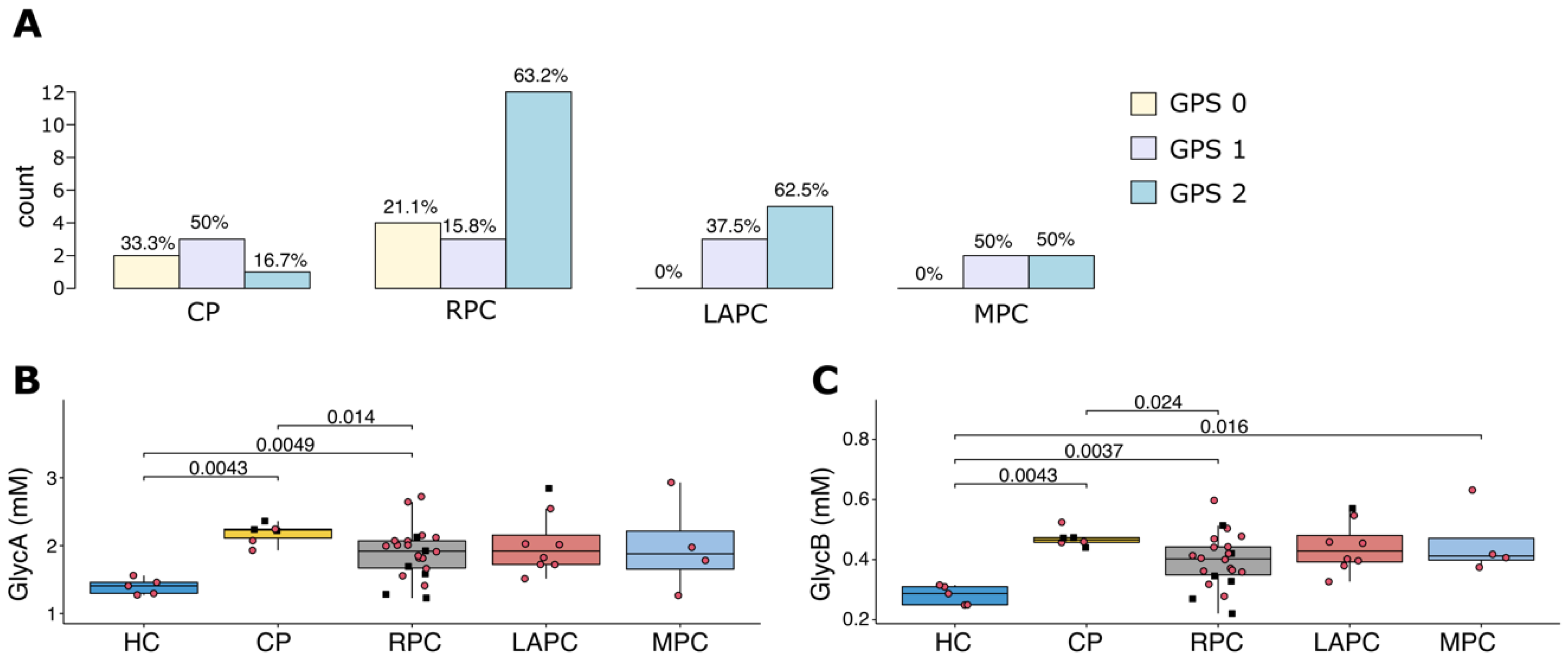
2.5. Impact of Diabetes and Inflammation on Metabolites and Lipoproteins Levels
3. Discussion
4. Materials and Methods
4.1. Sample Collection and Processing
4.2. Serum Sample Preparation
4.3. Lipid Extracts Preparation
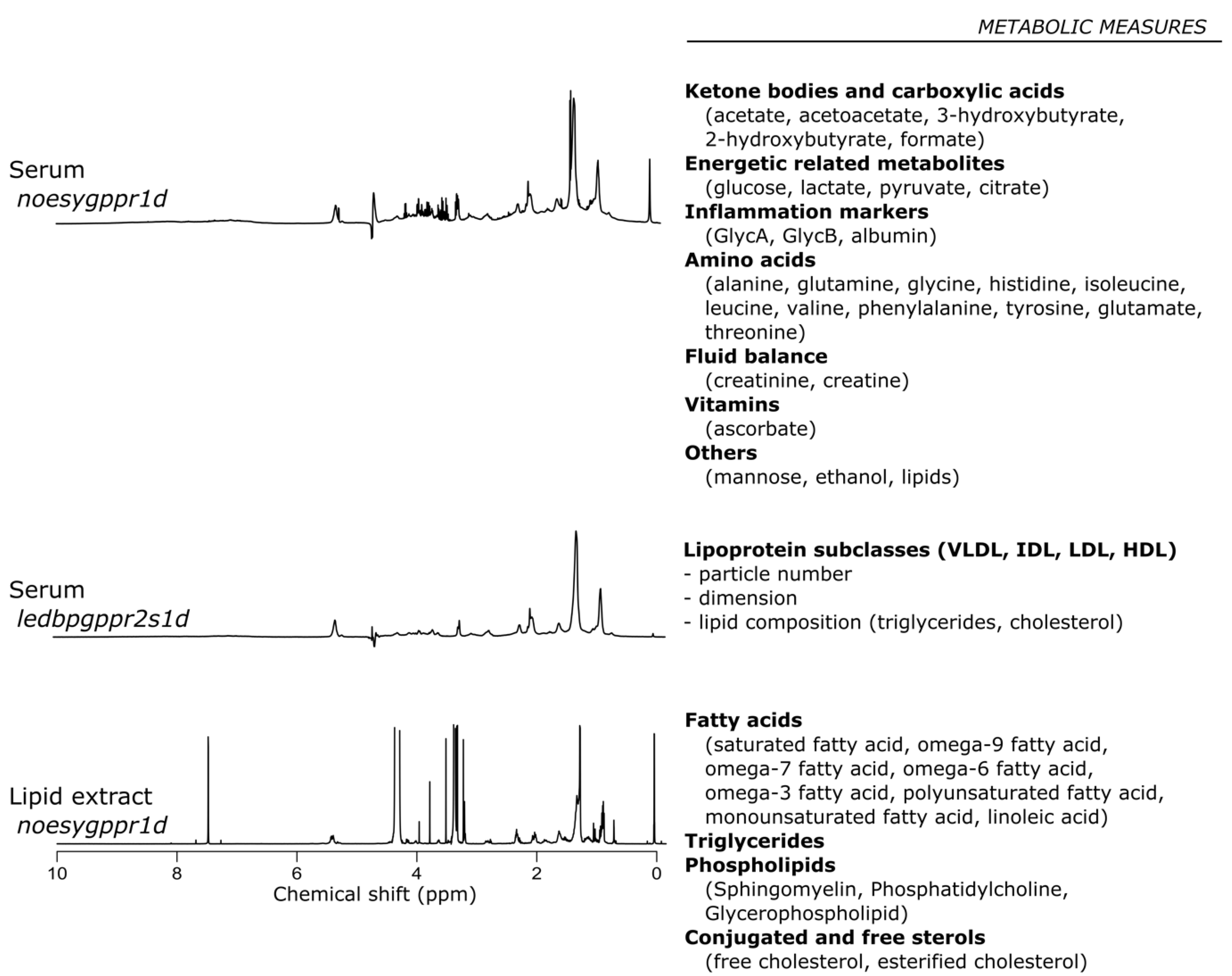
4.4. Nuclear Magnetic Resonance Spectroscopic Analysis
4.5. Nuclear Magnetic Resonance Profiling
4.6. Statistic and Data Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Yao, W.; Maitra, A.; Ying, H. Recent insights into the biology of pancreatic cancer. EBioMedicine 2020, 53, 102655. [Google Scholar] [CrossRef] [PubMed]
- Pourshams, A.; Sepanlou, S.G.; Ikuta, K.S.; Bisignano, C.; Safiri, S.; Roshandel, G.; Sharif, M.; Khatibian, M.; Fitzmaurice, C.; Nixon, M.R.; et al. The global, regional, and national burden of pancreatic cancer and its attributable risk factors in 195 countries and territories, 1990–2017: A systematic analysis for the Global Burden of Disease Study 2017. Lancet Gastroenterol. Hepatol. 2019, 4, 934–947. [Google Scholar] [CrossRef] [Green Version]
- Satyananda, V.; Gupta, R.; Hari, D.M.; Yeh, J.; Chen, K.T. Advances in Translational Research and Clinical Care in Pancreatic Cancer: Where Are We Headed? Gastroenterol. Res. Pract. 2019, 2019, 7690528. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer Statistics, 2021. CA Cancer J. Clin. 2021, 71, 7–33. [Google Scholar] [CrossRef] [PubMed]
- Sarantis, P.; Koustas, E.; Papadimitropoulou, A.; Papavassiliou, A.G.; Karamouzis, M.V. Pancreatic ductal adenocarcinoma: Treatment hurdles, tumor microenvironment and immunotherapy. World J. Gastrointest. Oncol. 2020, 12, 173–181. [Google Scholar] [CrossRef]
- Porta, M.; Fabregat, X.; Malats, N.; Guarner, L.; Carrato, A.; de Miguel, A.; Ruiz, L.; Jariod, M.; Costafreda, S.; Coll, S.; et al. Exocrine pancreatic cancer: Symptoms at presentation and their relation to tumour site and stage. Clin. Transl. Oncol. 2005, 7, 189–197. [Google Scholar] [CrossRef] [PubMed]
- McWilliams, R.R.; Maisonneuve, P.; Bamlet, W.R.; Petersen, G.M.; Li, D.; Risch, H.A.; Yu, H.; Fontham, E.T.H.; Luckett, B.; Bosetti, C.; et al. Risk Factors for Early-Onset and Very-Early-Onset Pancreatic Adenocarcinoma: A Pancreatic Cancer Case-Control Consortium (PanC4) Analysis. Pancreas 2016, 45, 311–316. [Google Scholar] [CrossRef] [Green Version]
- Silverman, D.T.; Hoover, R.N.; Brown, L.M.; Swanson, G.M.; Schiffman, M.; Greenberg, R.S.; Hayes, R.B.; Lillemoe, K.D.; Schoenberg, J.B.; Schwartz, A.G.; et al. Why Do Black Americans Have a Higher Risk of Pancreatic Cancer than White Americans? Epidemiology 2003, 14, 45–54. [Google Scholar] [CrossRef]
- Yadav, D.; Lowenfels, A.B. The Epidemiology of Pancreatitis and Pancreatic Cancer. Gastroenterology 2013, 144, 1252–1261. [Google Scholar] [CrossRef] [Green Version]
- Khawja, S.N.; Mohammed, S.; Silberfein, E.J.; Musher, B.L.; Fisher, W.E.; Van Buren, G.I. Pancreatic Cancer Disparities in African Americans. Pancreas 2015, 44, 522–527. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D.; Weinberg, R.A. Hallmarks of Cancer: The Next Generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef] [Green Version]
- Tennant, D.A.; Durán, R.V.; Gottlieb, E. Targeting metabolic transformation for cancer therapy. Nat. Rev. Cancer 2010, 10, 267–277. [Google Scholar] [CrossRef]
- Warburg, O.; Wind, F.; Negelein, E. The Metabolism of Tumors in the Body. J. Gen. Physiol. 1927, 8, 519–530. [Google Scholar] [CrossRef] [Green Version]
- Cacciatore, S.; Zadra, G.; Bango, C.; Penney, K.L.; Tyekucheva, S.; Yanes, O.; Loda, M. Metabolic Profiling in Formalin-Fixed and Paraffin-Embedded Prostate Cancer Tissues. Mol. Cancer Res. 2017, 15, 439–447. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Labbé, D.P.; Zadra, G.; Yang, M.; Reyes, J.M.; Lin, C.Y.; Cacciatore, S.; Ebot, E.M.; Creech, A.L.; Giunchi, F.; Fiorentino, M.; et al. High-fat diet fuels prostate cancer progression by rewiring the metabolome and amplifying the MYC program. Nat. Commun. 2019, 10, 4358. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Elebo, N.; Fru, P.; Omoshoro-Jones, J.; Candy, G.P.; Nweke, E.E. Role of different immune cells and metabolic pathways in modulating the immune response in pancreatic cancer. Mol. Med. Rep. 2020, 22, 4981–4991. [Google Scholar] [CrossRef] [PubMed]
- Lu, C.; Thompson, C.B. Metabolic regulation of epigenetics. Cell Metab. 2012, 16, 9–17. [Google Scholar] [CrossRef] [Green Version]
- Moore, H.B.; Culp-Hill, R.; Reisz, J.A.; Lawson, P.J.; Sauaia, A.; Schulick, R.D.; Del Chiaro, M.; Nydam, T.L.; Moore, E.E.; Hansen, K.C.; et al. The metabolic time line of pancreatic cancer: Opportunities to improve early detection of adenocarcinoma. Am. J. Surg. 2019, 218, 1206–1212. [Google Scholar] [CrossRef]
- Bertini, I.; Cacciatore, S.; Jensen, B.V.; Schou, J.V.; Johansen, J.S.; Kruhøffer, M.; Luchinat, C.; Nielsen, D.L.; Turano, P. Metabolomic NMR Fingerprinting to Identify and Predict Survival of Patients with Metastatic Colorectal Cancer. Cancer Res. 2012, 72, 356–364. [Google Scholar] [CrossRef] [Green Version]
- Cacciatore, S.; Wium, M.; Licari, C.; Ajayi-Smith, A.; Masieri, L.; Anderson, C.; Salukazana, A.S.; Kaestner, L.; Carini, M.; Carbone, G.M.J.C.; et al. Inflammatory metabolic profile of South African patients with prostate cancer. Cancer Metab. 2021, 9, 29. [Google Scholar] [CrossRef]
- Michálková, L.; Horník, Š.; Sykora, J.; Habartová, L.; Setnicka, V. Diagnosis of pancreatic cancer via 1H NMR metabolomics of human plasma. Analyst 2018, 143, 5974–5978. [Google Scholar] [CrossRef]
- Michálková, L.; Horník, Š.; Sýkora, J.; Habartová, L.; Setnička, V.; Bunganič, B. Early Detection of Pancreatic Cancer in Type 2 Diabetes Mellitus Patients Based on 1H NMR Metabolomics. J. Proteome Res. 2021, 20, 1744–1753. [Google Scholar] [CrossRef]
- Bathe, O.F.; Shaykhutdinov, R.; Kopciuk, K.; Weljie, A.M.; McKay, A.; Sutherland, F.R.; Dixon, E.; Dunse, N.; Sotiropoulos, D.; Vogel, H.J. Feasibility of Identifying Pancreatic Cancer Based on Serum Metabolomics. Cancer Epidemiol. Biomark. Prev. 2011, 20, 140–147. [Google Scholar] [CrossRef] [Green Version]
- OuYang, D.; Xu, J.; Huang, H.; Chen, Z. Metabolomic Profiling of Serum from Human Pancreatic Cancer Patients Using 1H NMR Spectroscopy and Principal Component Analysis. Appl. Biochem. Biotechnol. 2011, 165, 148–154. [Google Scholar] [CrossRef]
- Fest, J.; Vijfhuizen, L.S.; Goeman, J.J.; Veth, O.; Joensuu, A.; Perola, M.; Männistö, S.; Ness-Jensen, E.; Hveem, K.; Haller, T.; et al. Search for Early Pancreatic Cancer Blood Biomarkers in Five European Prospective Population Biobanks Using Metabolomics. Endocrinology 2019, 160, 1731–1742. [Google Scholar] [CrossRef] [Green Version]
- Mehta, K.Y.; Wu, H.-J.; Menon, S.S.; Fallah, Y.; Zhong, X.; Rizk, N.; Unger, K.; Mapstone, M.; Fiandaca, M.S.; Federoff, H.J.; et al. Metabolomic biomarkers of pancreatic cancer: A meta-analysis study. Oncotarget 2017, 8, 68899–68915. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Donelli, G.; Guaglianone, E.; Di Rosa, R.; Fiocca, F.; Basoli, A. Plastic biliary stent occlusion: Factors involved and possible preventive approaches. Clin. Med. Res. 2007, 5, 53–60. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kruse, E.J. Palliation in Pancreatic Cancer. Surg. Clin. N. Am. 2010, 90, 355–364. [Google Scholar] [CrossRef]
- Ćwiklińska, A.; Mickiewicz, A.; Kowalski, R.; Kortas-Stempak, B.; Kuchta, A.; Mucha, K.; Makowiecki, M.; Gliwińska, A.; Lewandowski, K.; Pączek, L.; et al. Detection of Lipoprotein X (LpX): A challenge in patients with severe hypercholesterolaemia. J. Med. Biochem. 2020, 39, 283–289. [Google Scholar] [CrossRef] [PubMed]
- Yan, L.; Tonack, S.; Smith, R.; Dodd, S.; Jenkins, R.E.; Kitteringham, N.; Greenhalf, W.; Ghaneh, P.; Neoptolemos, J.P.; Costello, E. Confounding Effect of Obstructive Jaundice in the Interpretation of Proteomic Plasma Profiling Data for Pancreatic Cancer. J. Proteome Res. 2009, 8, 142–148. [Google Scholar] [CrossRef] [PubMed]
- Velazquez-Torres, G.; Fuentes-Mattei, E.; Choi, H.H.; Yeung, S.-C.J.; Meng, X.; Lee, M.-H. Diabetes mellitus type 2 drives metabolic reprogramming to promote pancreatic cancer growth. Gastroenterol. Rep. 2020, 8, 261–276. [Google Scholar] [CrossRef]
- Zechner, D.; Radecke, T.; Amme, J.; Bürtin, F.; Albert, A.-C.; Partecke, L.I.; Vollmar, B. Impact of diabetes type II and chronic inflammation on pancreatic cancer. BMC Cancer 2015, 15, 51. [Google Scholar] [CrossRef] [Green Version]
- Correig, E.; Ibarretxe, D.; Rodriguez, D.; Pardo, M.; Rodriguez, C.; Plana, N.; Masana, L.; Amigo, N.J.A. Exploring lipoprotein patterns in general population with advanced 1H-NMR testing by using an unsupervised clustering approach. Atherosclerosis 2018, 275, e97. [Google Scholar]
- Lamiquiz-Moneo, I.; Civeira, F.; Gómez-Coronado, D.; Blanco-Vaca, F.; Villafuerte-Ledesma, H.M.; Gil, M.; Amigó, N.; Mateo-Gallego, R.; Cenarro, A. Lipid Profile Rather Than the LCAT Mutation Explains Renal Disease in Familial LCAT Deficiency. J. Clin. Med. 2019, 8, 1860. [Google Scholar] [CrossRef] [Green Version]
- McMillan, D.C. The systemic inflammation-based Glasgow Prognostic Score: A decade of experience in patients with cancer. Cancer Treat. Rev. 2013, 39, 534–540. [Google Scholar] [CrossRef]
- Gilliam, A.D.; Lobo, D.N.; Rowlands, B.J.; Beckingham, I.J. The ‘two-week’ target for the diagnosis of pancreatic carcinoma: An achievable aim? Eur. J. Surg. Oncol. (EJSO) 2003, 29, 575–579. [Google Scholar] [CrossRef]
- Cao, L.; Wu, J.; Qu, X.; Sheng, J.; Cui, M.; Liu, S.; Huang, X.; Xiang, Y.; Li, B.; Zhang, X.; et al. Glycometabolic rearrangements–aerobic glycolysis in pancreatic cancer: Causes, characteristics and clinical applications. J. Exp. Clin. Cancer Res. 2020, 39, 267. [Google Scholar] [CrossRef]
- McDonald, O.G.; Li, X.; Saunders, T.; Tryggvadottir, R.; Mentch, S.J.; Warmoes, M.O.; Word, A.E.; Carrer, A.; Salz, T.H.; Natsume, S.; et al. Epigenomic reprogramming during pancreatic cancer progression links anabolic glucose metabolism to distant metastasis. Nat. Genet. 2017, 49, 367–376. [Google Scholar] [CrossRef] [PubMed]
- Mentch, S.; Locasale, J. One-carbon metabolism and epigenetics: Understanding the specificity. Ann. N. Y. Acad. Sci. 2016, 1363, 91–98. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, S.; Xu, H.; Wang, W.; Li, S.; Li, H.; Li, T.; Zhang, W.; Yu, X.; Liu, L. The role of collagen in cancer: From bench to bedside. J. Transl. Med. 2019, 17, 309. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Amelio, I.; Cutruzzolá, F.; Antonov, A.; Agostini, M.; Melino, G. Serine and glycine metabolism in cancer. Trends Biochem. Sci. 2014, 39, 191–198. [Google Scholar] [CrossRef]
- Pan, S.; Fan, M.; Liu, Z.; Li, X.; Wang, H. Serine, glycine and one-carbon metabolism in cancer. Int. J. Oncol. 2021, 58, 158–170. [Google Scholar] [CrossRef]
- Park, S.; Ahn, S.; Shin, Y.; Yang, Y.; Yeom, C.H. Vitamin C in Cancer: A Metabolomics Perspective. Front. Physiol. 2018, 9, 762. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yun, J.; Mullarky, E.; Lu, C.; Bosch, K.N.; Kavalier, A.; Rivera, K.; Roper, J.; Chio, I.I.C.; Giannopoulou, E.G.; Rago, C.; et al. Vitamin C selectively kills KRAS and BRAF mutant colorectal cancer cells by targeting GAPDH. Science 2015, 350, 1391–1396. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Polireddy, K.; Dong, R.; Reed, G.; Yu, J.; Chen, P.; Williamson, S.; Violet, P.-C.; Pessetto, Z.; Godwin, A.K.; Fan, F.; et al. High Dose Parenteral Ascorbate Inhibited Pancreatic Cancer Growth and Metastasis: Mechanisms and a Phase I/IIa study. Sci. Rep. 2017, 7, 17188. [Google Scholar] [CrossRef] [Green Version]
- Chen, Q.; Espey, M.G.; Sun, A.Y.; Pooput, C.; Kirk, K.L.; Krishna, M.C.; Khosh, D.B.; Drisko, J.; Levine, M. Pharmacologic doses of ascorbate act as a prooxidant and decrease growth of aggressive tumor xenografts in mice. Proc. Natl. Acad. Sci. USA 2008, 105, 11105–11109. [Google Scholar] [CrossRef] [Green Version]
- Drisko, J.A.; Serrano, O.K.; Spruce, L.R.; Chen, Q.; Levine, M. Treatment of pancreatic cancer with intravenous vitamin C: A case report. Anticancer. Drugs 2018, 29, 373–379. [Google Scholar] [CrossRef]
- Sousa, C.M.; Kimmelman, A.C.J.C. The complex landscape of pancreatic cancer metabolism. Carcinogenesis 2014, 35, 1441–1450. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Laffel, L. Ketone bodies: A review of physiology, pathophysiology and application of monitoring to diabetes. Diabetes/Metab. Res. Rev. 1999, 15, 412–426. [Google Scholar] [CrossRef]
- Halperin, M.L.; Hammeke, M.; Josse, R.G.; Jungas, R.L. Metabolic acidosis in the alcoholic: A pathophysiologic approach. Metabolism 1983, 32, 308–315. [Google Scholar] [CrossRef]
- Shakery, A.; Pourvali, K.; Ghorbani, A.; Fereidani, S.S.; Zand, H. Beta-Hydroxybutyrate Promotes Proliferation, Migration and Stemness in a Subpopulation of 5FU Treated SW480 Cells: Evidence for Metabolic Plasticity in Colon Cancer. Asian Pac. J. Cancer Prev. 2018, 19, 3287–3294. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Martinez-Outschoorn, U.E.; Prisco, M.; Ertel, A.; Tsirigos, A.; Lin, Z.; Pavlides, S.; Wang, C.; Flomenberg, N.; Knudsen, E.S.; Howell, A.; et al. Ketones and lactate increase cancer cell “stemness”, driving recurrence, metastasis and poor clinical outcome in breast cancer. Cell Cycle 2011, 10, 1271–1286. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lim, D.J.; Sharma, Y.; Thompson, C.H. Vitamin C and alcohol: A call to action. BMJ Nutr. Prev. Health 2018, 1, 17–22. [Google Scholar] [CrossRef] [Green Version]
- Tripathi, N.; Jialal, I. Conjugated Hyperbilirubinemia; StatPearls Publishing: Treasure Island, FL, USA, 2020. [Google Scholar]
- Méndez-Sánchez, N.; Vítek, L.; Aguilar-Olivos, N.E.; Uribe, M. Bilirubin as a Biomarker in Liver Disease. In Biomarkers in Liver Disease; Patel, V.B., Preedy, V.R., Eds.; Springer: Dordrecht, The Netherlands, 2017; pp. 281–304. [Google Scholar] [CrossRef]
- Di Gangi, I.M.; Mazza, T.; Fontana, A.; Copetti, M.; Fusilli, C.; Ippolito, A.; Mattivi, F.; Latiano, A.; Andriulli, A.; Vrhovsek, U.J.O. Metabolomic profile in pancreatic cancer patients: A consensus-based approach to identify highly discriminating metabolites. Oncotarget 2016, 7, 5815. [Google Scholar] [CrossRef] [PubMed]
- Shadhu, K.; Xi, C. Inflammation and pancreatic cancer: An updated review. Saudi J. Gastroenterol. 2019, 25, 3–13. [Google Scholar] [CrossRef]
- Nordmann, T.M.; Dror, E.; Schulze, F.; Traub, S.; Berishvili, E.; Barbieux, C.; Böni-Schnetzler, M.; Donath, M.Y. The Role of Inflammation in β-cell Dedifferentiation. Sci. Rep. 2017, 7, 6285. [Google Scholar] [CrossRef]
- Harris, P.A.; Taylor, R.; Thielke, R.; Payne, J.; Gonzalez, N.; Conde, J.G. Research electronic data capture (REDCap)—A metadata-driven methodology and workflow process for providing translational research informatics support. J. Biomed. Inform. 2009, 42, 377–381. [Google Scholar] [CrossRef] [Green Version]
- Tempero, M.A.; Malafa, M.P.; Chiorean, E.G.; Czito, B.; Scaife, C.; Narang, A.K.; Fountzilas, C.; Wolpin, B.M.; Al-Hawary, M.; Asbun, H.; et al. Pancreatic Adenocarcinoma, Version 1.2019. J. Natl. Compr. Cancer Netw. 2019, 17, 202–210. [Google Scholar] [CrossRef] [Green Version]
- Soweid, A.M. The borderline resectable and locally advanced pancreatic ductal adenocarcinoma: Definition. Endosc. Ultrasound 2017, 6, S76–S78. [Google Scholar] [CrossRef]
- Marshall, I.; Higinbotham, J.; Bruce, S.; Freise, A. Use of voigt lineshape for quantification of in vivo 1H spectra. Magn. Reson. Med. 1997, 37, 651–657. [Google Scholar] [CrossRef]
- Serkova, N.; Florian Fuller, T.; Klawitter, J.; Freise, C.E.; Niemann, C.U. 1H-NMR–based metabolic signatures of mild and severe ischemia/reperfusion injury in rat kidney transplants. Kidney Int. 2005, 67, 1142–1151. [Google Scholar] [CrossRef] [Green Version]
- Mallol, R.; Amigó, N.; Rodríguez, M.A.; Heras, M.; Vinaixa, M.; Plana, N.; Rock, E.; Ribalta, J.; Yanes, O.; Masana, L.; et al. Liposcale: A novel advanced lipoprotein test based on 2D diffusion-ordered 1H NMR spectroscopy. J. Lipid Res. 2015, 56, 737–746. [Google Scholar] [CrossRef] [Green Version]
- Cacciatore, S.; Tenori, L.; Luchinat, C.; Bennett, P.R.; MacIntyre, D.A. KODAMA: An R package for knowledge discovery and data mining. Bioinformatics 2017, 33, 621–623. [Google Scholar] [CrossRef] [Green Version]
- Bray, R.; Cacciatore, S.; Jiménez, B.; Cartwright, R.; Digesu, A.; Fernando, R.; Holmes, E.; Nicholson, J.K.; Bennett, P.R.; MacIntyre, D.A.; et al. Urinary Metabolic Phenotyping of Women with Lower Urinary Tract Symptoms. J. Proteome Res. 2017, 16, 4208–4216. [Google Scholar] [CrossRef]
- Reynolds, A.P.; Richards, G.; de la Iglesia, B.; Rayward-smith, V.J. Clustering Rules: A Comparison of Partitioning and Hierarchical Clustering Algorithms. J. Math. Model. Algorithms 2006, 5, 475–504. [Google Scholar] [CrossRef]
- Rousseeuw, P.J. Silhouettes: A graphical aid to the interpretation and validation of cluster analysis. J. Comput. Appl. Math. 1987, 20, 53–65. [Google Scholar] [CrossRef] [Green Version]
- Eriksson, L.; Jaworska, J.; Worth, A.P.; Cronin, M.T.D.; McDowell, R.M.; Gramatica, P. Methods for reliability and uncertainty assessment and for applicability evaluations of classification- and regression-based QSARs. Environ. Health Perspect. 2003, 111, 1361–1375. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Szymańska, E.; Saccenti, E.; Smilde, A.K.; Westerhuis, J.A. Double-check: Validation of diagnostic statistics for PLS-DA models in metabolomics studies. Metabolomics 2012, 8, 3–16. [Google Scholar] [CrossRef] [PubMed] [Green Version]

| Feature | HC (n = 6) | CP (n = 6) | RPC (n = 22) | LAPC (n = 8) | MPC (n = 4) | p-Value |
|---|---|---|---|---|---|---|
| HIV status | 0.831 | |||||
| Negative, n (%) | 6 (100.0) | 5 (83.3) | 19 (86.4) | 6 (75.0) | 4 (100.0) | |
| Positive, n (%) | 0 (0.0) | 1 (16.7) | 3 (13.6) | 2 (25.0) | 0 (0.0) | |
| Gender | 0.286 | |||||
| female, n (%) | 3 (50.0) | 0 (0.0) | 8 (36.4) | 2 (25.0) | 2 (50.0) | |
| male, n (%) | 3 (50.0) | 6 (100.0) | 14 (63.6) | 6 (75.0) | 2 (50.0) | |
| Smoking | 0.450 | |||||
| no, n (%) | 6 (100.0) | 1 (16.7) | 12 (54.5) | 4 (50.0) | 2 (50.0) | |
| yes, n (%) | 0 (0.0) | 5 (83.3) | 10 (45.5) | 4 (50.0) | 2 (50.0) | |
| Alcohol | 0.962 | |||||
| no, n (%) | 4 (66.67) | 3 (50.0) | 13 (59.1) | 4 (50.0) | 2 (50.0) | |
| yes, n (%) | 2 (33.33) | 3 (50.0) | 9 (40.9) | 4 (50.0) | 2 (50.0) | |
| Age, median (IQR) | 37 (24–54) | 51 (46–57) | 63 (50–67) | 56 (48–62) | 56 (46–70) | 0.439 |
| Obstructive jaundice | 0.013 | |||||
| no, n (%) | 6 (100.0) | 6 (100.0) | 8 (36.4) | 2 (25.0) | 1 (25.0) | |
| yes, n (%) | 0 (0.0) | 0 (0.0) | 14 (63.6) | 6 (75.0) | 3 (75.0) | |
| Cholangitis | 0.145 | |||||
| no, n (%) | 6 (100.0) | 6 (100.0) | 20 (90.9) | 7 (87.5) | 2 (50.0) | |
| yes, n (%) | 0 (0.0) | 0 (0.0) | 2 (9.1) | 1 (12.5) | 2 (50.0) | |
| T2DM | 0.322 | |||||
| no, n (%) | 6 (100.0) | 3 (50.0) | 16 (72.7) | 7 (87.5) | 4 (100.0) | |
| yes, n (%) | 0 (0.0) | 3 (50.0) | 6 (27.3) | 1 (12.5) | 0 (0.0) | |
| Hypertension | 0.560 | |||||
| no, n (%) | 6 (100.0) | 6 (100.0) | 17 (77.3) | 6 (75.0) | 4 (100.0) | |
| yes, n (%) | 0 (0.0) | 0 (0.0) | 5 (22.7) | 2 (25.0) | 0 (0.0) | |
| Feature | * Physiological Range | CP Median | RPC Median | LAPC Median | MPC Median | p-Value | FDR |
|---|---|---|---|---|---|---|---|
| Total Protein (g/L) | 60–78 | 66.0 | 59.0 | 66.0 | 69.0 | 0.289 | 0.330 |
| Albumin (g/L) | 35–52 | 36.5 | 30.0 | 27.0 | 32.5 | 0.361 | 0.361 |
| Total Bilirubin (µmol/L) | 5–21 | 5.0 | 154.0 | 120.0 | 58.0 | 0.006 | 0.030 |
| Conjugated Bilirubin (µmol/L) | 0–3 | 2.0 | 141.0 | 112.5 | 45.0 | 0.008 | 0.030 |
| Alanine transaminase (U/L) | 10–40 | 18.0 | 88.0 | 29.5 | 38.0 | 0.051 | 0.082 |
| Aspartate transaminase (U/L) | 15–40 | 28.5 | 104.0 | 55.0 | 75.0 | 0.019 | 0.052 |
| Alkaline phosphatase (U/L) | 53–128 | 74.0 | 615.0 | 337.0 | 314.5 | 0.025 | 0.052 |
| Gamma glutamyl transferase (U/L) | <68 | 61.5 | 751.0 | 301.0 | 483.0 | 0.151 | 0.201 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Elebo, N.; Omoshoro-Jones, J.; Fru, P.N.; Devar, J.; De Wet van Zyl, C.; Vorster, B.C.; Smith, M.; Cacciatore, S.; Zerbini, L.F.; Candy, G.; et al. Serum Metabolomic and Lipoprotein Profiling of Pancreatic Ductal Adenocarcinoma Patients of African Ancestry. Metabolites 2021, 11, 663. https://doi.org/10.3390/metabo11100663
Elebo N, Omoshoro-Jones J, Fru PN, Devar J, De Wet van Zyl C, Vorster BC, Smith M, Cacciatore S, Zerbini LF, Candy G, et al. Serum Metabolomic and Lipoprotein Profiling of Pancreatic Ductal Adenocarcinoma Patients of African Ancestry. Metabolites. 2021; 11(10):663. https://doi.org/10.3390/metabo11100663
Chicago/Turabian StyleElebo, Nnenna, Jones Omoshoro-Jones, Pascaline N. Fru, John Devar, Christiaan De Wet van Zyl, Barend Christiaan Vorster, Martin Smith, Stefano Cacciatore, Luiz F. Zerbini, Geoffrey Candy, and et al. 2021. "Serum Metabolomic and Lipoprotein Profiling of Pancreatic Ductal Adenocarcinoma Patients of African Ancestry" Metabolites 11, no. 10: 663. https://doi.org/10.3390/metabo11100663
APA StyleElebo, N., Omoshoro-Jones, J., Fru, P. N., Devar, J., De Wet van Zyl, C., Vorster, B. C., Smith, M., Cacciatore, S., Zerbini, L. F., Candy, G., & Nweke, E. E. (2021). Serum Metabolomic and Lipoprotein Profiling of Pancreatic Ductal Adenocarcinoma Patients of African Ancestry. Metabolites, 11(10), 663. https://doi.org/10.3390/metabo11100663







