An Integrated Multi-Disciplinary Perspective for Addressing Challenges of the Human Gut Microbiome
Abstract
1. Introduction
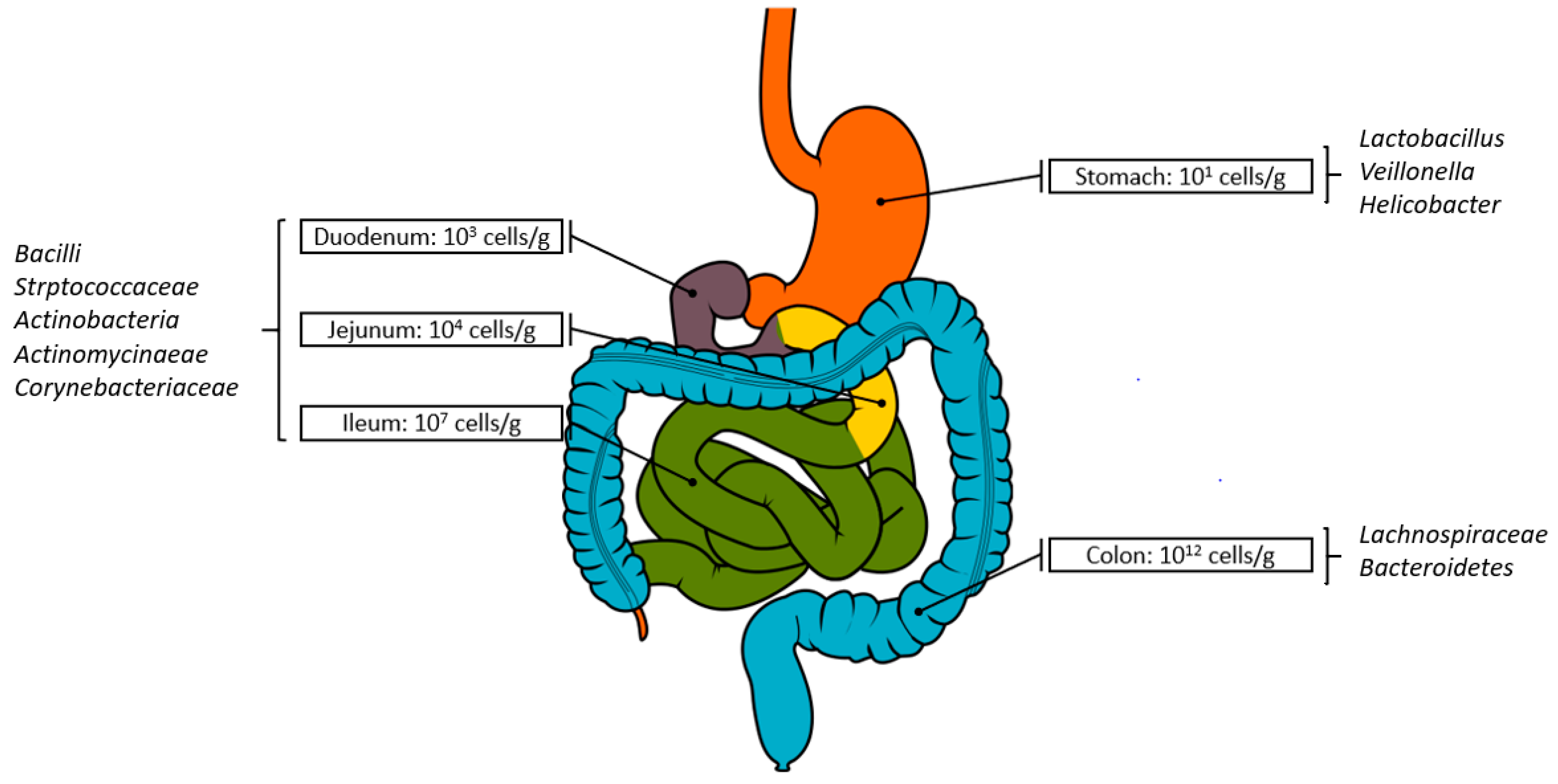
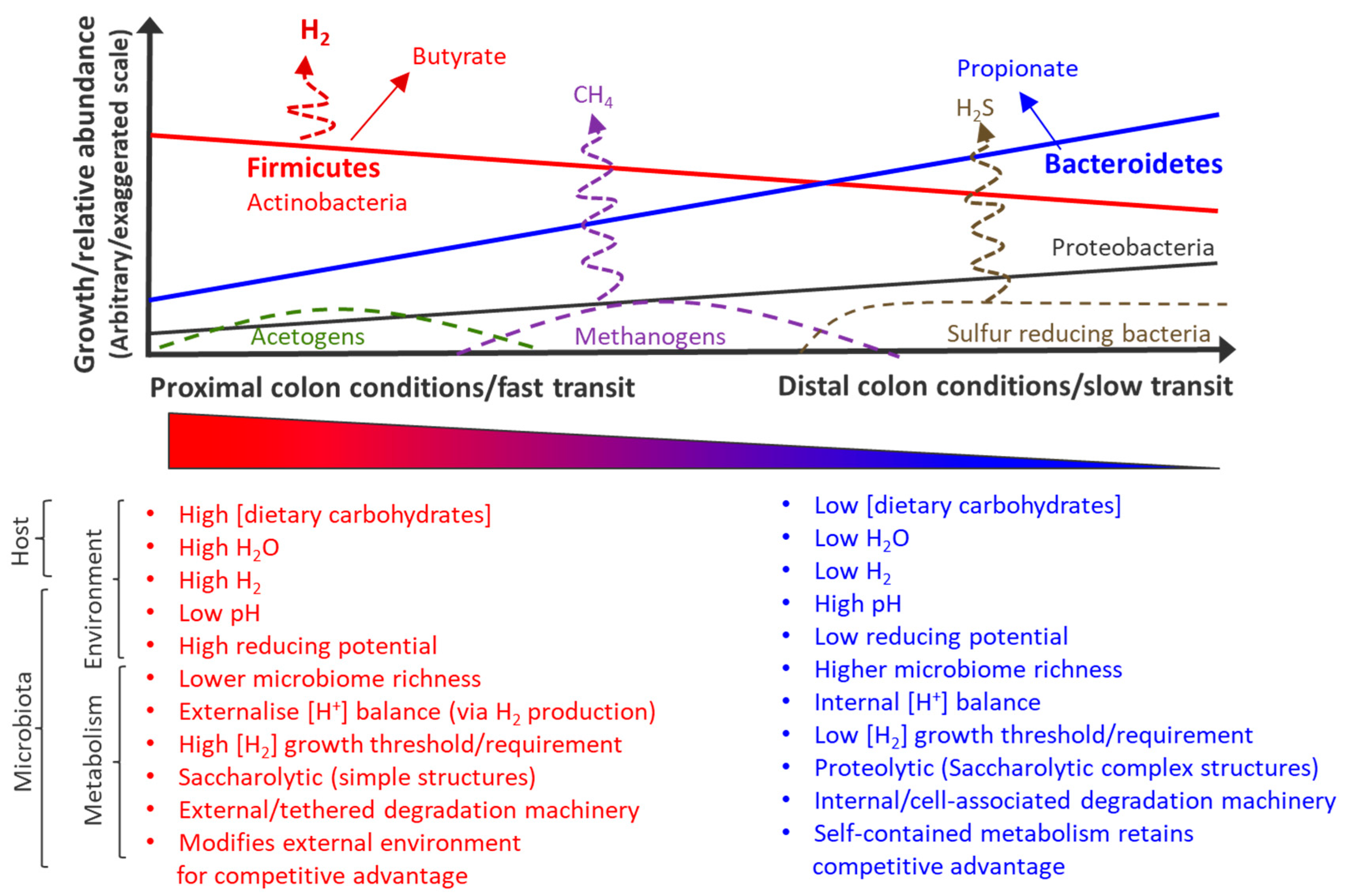
2. Biogeography of the Microbiome and Metabolome: Implications for Faecal Samples as Proxies
3. The Role of Microbiome Structure and Function in Human Health and Disease
4. Analysing Microbiome Structure and Function with Non-Metabolomics Approaches
5. Analysing Microbiome Function with Metabolomics
6. Integrating Multi-Omics Datasets
7. Future Directions that Would Accelerate an Integrated Approach
7.1. New Sampling Techniques
7.2. New and Emerging Techniques and Disciplines: Culturomics, the Interactome, Foodomics, and the Exposome
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Robinson, C.J.; Bohannan, B.J.; Young, V.B. From structure to function: The ecology of host-associated microbial communities. Microbiol. Mol. Biol. Rev. 2010, 74, 453–476. [Google Scholar] [CrossRef] [PubMed]
- Shreiner, A.B.; Kao, J.Y.; Young, V.B. The gut microbiome in health and in disease. Curr. Opin. Gastroenterol. 2015, 31, 69. [Google Scholar] [CrossRef] [PubMed]
- Nash, A.K.; Auchtung, T.A.; Wong, M.C.; Smith, D.P.; Gesell, J.R.; Ross, M.C.; Stewart, C.J.; Metcalf, G.A.; Muzny, D.M.; Gibbs, R.A. The gut mycobiome of the Human Microbiome Project healthy cohort. Microbiome 2017, 5, 153. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, T.S.; Raes, J.; Bork, P. The human gut microbiome: From association to modulation. Cell 2018, 172, 1198–1215. [Google Scholar] [CrossRef] [PubMed]
- Heintz-Buschart, A.; Wilmes, P. Human gut microbiome: Function matters. Trends Microbiol. 2018, 26, 563–574. [Google Scholar] [CrossRef] [PubMed]
- Tropini, C.; Earle, K.A.; Huang, K.C.; Sonnenburg, J.L. The gut microbiome: Connecting spatial organization to function. Cell Host Microbe 2017, 21, 433–442. [Google Scholar] [CrossRef]
- Lynch, S.V.; Pedersen, O. The human intestinal microbiome in health and disease. N. Engl. J. Med. 2016, 375, 2369–2379. [Google Scholar] [CrossRef]
- Couch, R.D.; Navarro, K.; Sikaroodi, M.; Gillevet, P.; Forsyth, C.B.; Mutlu, E.; Engen, P.A.; Keshavarzian, A. The approach to sample acquisition and its impact on the derived human fecal microbiome and VOC metabolome. PLoS ONE 2013, 8, e81163. [Google Scholar] [CrossRef]
- Zierer, J.; Jackson, M.A.; Kastenmüller, G.; Mangino, M.; Long, T.; Telenti, A.; Mohney, R.P.; Small, K.S.; Bell, J.T.; Steves, C.J.; et al. The fecal metabolome as a functional readout of the gut microbiome. Nat. Genet. 2018, 50, 790–795. [Google Scholar] [CrossRef]
- Marchesi, J.R. Prokaryotic and eukaryotic diversity of the human gut. Adv. Appl. Microbiol. 2010, 72, 43–62. [Google Scholar]
- Huseyin, C.E.; O’Toole, P.W.; Cotter, P.D.; Scanlan, P.D. Forgotten fungi-the gut mycobiome in human health and disease. FEMS Microbiol. Rev. 2017, 41, 479–511. [Google Scholar] [CrossRef] [PubMed]
- Koh, A.Y. Gastrointestinal colonization of fungi. Curr. Fungal Infect. Rep. 2013, 7, 144–151. [Google Scholar] [CrossRef]
- Underhill, D.M.; Iliev, I.D. The mycobiota: Interactions between commensal fungi and the host immune system. Nat. Rev. Immunol. 2014, 14, 405–416. [Google Scholar] [CrossRef] [PubMed]
- Seed, P.C. The human mycobiome. Cold Spring Harb. Perspect. Med. 2015, 5, a019810. [Google Scholar] [CrossRef] [PubMed]
- Huffnagle, G.B.; Noverr, M.C. The emerging world of the fungal microbiome. Trends Microbiol. 2013, 21, 334–341. [Google Scholar] [CrossRef] [PubMed]
- Wegener Parfrey, L.; Walters, W.A.; Knight, R. Microbial eukaryotes in the human microbiome: Ecology, evolution, and future directions. Front. Microbiol. 2011, 2, 153. [Google Scholar] [CrossRef]
- Ley, R.E.; Peterson, D.A.; Gordon, J.I. Ecological and evolutionary forces shaping microbial diversity in the human intestine. Cell 2006, 124, 837–848. [Google Scholar] [CrossRef]
- Garrett, W.S.; Gordon, J.I.; Glimcher, L.H. Homeostasis and inflammation in the intestine. Cell 2010, 140, 859–870. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Lauber, C.L.; Walters, W.A.; Berg-Lyons, D.; Lozupone, C.A.; Turnbaugh, P.J.; Fierer, N.; Knight, R. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl. Acad. Sci. USA 2011, 108, 4516–4522. [Google Scholar] [CrossRef]
- Bolhuis, H.; Cretoiu, M.S.; Stal, L.J. Molecular ecology of microbial mats. FEMS Microbiol. Ecol. 2014, 90, 335–350. [Google Scholar]
- Huttenhower, C.; Knight, R.; Brown, C.T.; Caporaso, J.G.; Clemente, J.C.; Gevers, D.; Franzosa, E.A.; Kelley, S.T.; Knights, D.; Ley, R.E. Advancing the microbiome research community. Cell 2014, 159, 227–230. [Google Scholar] [CrossRef] [PubMed]
- Norman, J.M.; Handley, S.A.; Virgin, H.W. Kingdom-agnostic metagenomics and the importance of complete characterization of enteric microbial communities. Gastroenterology 2014, 146, 1459–1469. [Google Scholar] [CrossRef] [PubMed]
- Yoon, S.S.; Kim, E.-K.; Lee, W.-J. Functional genomic and metagenomic approaches to understanding gut microbiota–animal mutualism. Curr. Opin. Microbiol. 2015, 24, 38–46. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, C.; Dollive, S.; Grunberg, S.; Chen, J.; Li, H.; Wu, G.D.; Lewis, J.D.; Bushman, F.D. Archaea and fungi of the human gut microbiome: Correlations with diet and bacterial residents. PLoS ONE 2013, 8, e66019. [Google Scholar] [CrossRef] [PubMed]
- Schulze, J.; Sonnenborn, U. Yeasts in the gut: From commensals to infectious agents. Dtsch. Arztebl. Int. 2009, 106, 837. [Google Scholar] [CrossRef]
- Suhr, M.J.; Hallen-Adams, H.E. The human gut mycobiome: Pitfalls and potentials—A mycologist’s perspective. Mycologia 2015, 107, 1057–1073. [Google Scholar] [CrossRef]
- Sender, R.; Fuchs, S.; Milo, R. Revised estimates for the number of human and bacteria cells in the body. PLoS Biol. 2016, 14, e1002533. [Google Scholar] [CrossRef]
- Lauber, C.L.; Zhou, N.; Gordon, J.I.; Knight, R.; Fierer, N. Effect of storage conditions on the assessment of bacterial community structure in soil and human-associated samples. FEMS Microbiol. Lett. 2010, 307, 80–86. [Google Scholar] [CrossRef]
- Wu, G.D.; Lewis, J.D.; Hoffmann, C.; Chen, Y.-Y.; Knight, R.; Bittinger, K.; Hwang, J.; Chen, J.; Berkowsky, R.; Nessel, L.; et al. Sampling and pyrosequencing methods for characterizing bacterial communities in the human gut using 16S sequence tags. BMC Microbiol. 2010, 10, 206. [Google Scholar] [CrossRef]
- Roesch, L.F.; Casella, G.; Simell, O.; Krischer, J.; Wasserfall, C.H.; Schatz, D.; Atkinson, M.A.; Neu, J.; Triplett, E.W. Influence of fecal sample storage on bacterial community diversity. Open Microbiol. J. 2009, 3, 40. [Google Scholar] [CrossRef]
- Panek, M.; Čipčić Paljetak, H.; Barešić, A.; Perić, M.; Matijašić, M.; Lojkić, I.; Vranešić Bender, D.; Krznarić, Ž.; Verbanac, D. Methodology challenges in studying human gut microbiota—Effects of collection, storage, DNA extraction and next generation sequencing technologies. Sci. Rep. 2018, 8, 5143. [Google Scholar] [CrossRef] [PubMed]
- Vandeputte, D.; Tito, R.Y.; Vanleeuwen, R.; Falony, G.; Raes, J. Practical considerations for large-scale gut microbiome studies. FEMS Microbiol. Rev. 2017, 41, S154–S167. [Google Scholar] [CrossRef] [PubMed]
- Falony, G.; Vieira-Silva, S.; Raes, J. Richness and ecosystem development across faecal snapshots of the gut microbiota. Nat. Microbiol. 2018, 3, 526–528. [Google Scholar] [CrossRef] [PubMed]
- Wolfe, A.J. The acetate switch. Microbiol. Mol. Biol. Rev. 2005, 69, 12–50. [Google Scholar] [CrossRef] [PubMed]
- Flamholz, A.; Noor, E.; Bar-Even, A.; Liebermeister, W.; Milo, R. Glycolytic strategy as a tradeoff between energy yield and protein cost. Proc. Natl. Acad. Sci. USA 2013, 110, 10039–10044. [Google Scholar] [CrossRef] [PubMed]
- Herrmann, E.; Young, W.; Rosendale, D.; Reichert-Grimm, V.; Riedel, C.U.; Conrad, R.; Egert, M. RNA-based stable isotope probing suggests Allobaculum spp. as particularly active glucose assimilators in a complex murine microbiota cultured in vitro. BioMed Res. Int. 2017. [Google Scholar] [CrossRef]
- Mahowald, M.A.; Rey, F.E.; Seedorf, H.; Turnbaugh, P.J.; Fulton, R.S.; Wollam, A.; Shah, N.; Wang, C.; Magrini, V.; Wilson, R.K.; et al. Characterizing a model human gut microbiota composed of members of its two dominant bacterial phyla. Proc. Natl. Acad. Sci. USA 2009, 106, 5859–5864. [Google Scholar] [CrossRef]
- Tuncil, Y.E.; Xiao, Y.; Porter, N.T.; Reuhs, B.L.; Martens, E.C.; Hamaker, B.R. Reciprocal Prioritization to Dietary Glycans by Gut Bacteria in a Competitive Environment Promotes Stable Coexistence. mBio 2017, 8. [Google Scholar] [CrossRef]
- Kim, C. Identification of Rumen Methanolgens, Characterization of Substrate Requirements and Measurement of Hydrogen Thresholds. Master’s Thesis, Massey University, Palmerston North, New Zealand, 2012. [Google Scholar]
- Donohoe, D.R.; Garge, N.; Zhang, X.; Sun, W.; O’Connell, T.M.; Bunger, M.K.; Bultman, S.J. The microbiome and butyrate regulate energy metabolism and autophagy in the mammalian colon. Cell Metab. 2011, 13, 517–526. [Google Scholar] [CrossRef]
- den Besten, G.; Havinga, R.; Bleeker, A.; Rao, S.; Gerding, A.; van Eunen, K.; Groen, A.K.; Reijngoud, D.J.; Bakker, B.M. The short-chain fatty acid uptake fluxes by mice on a guar gum supplemented diet associate with amelioration of major biomarkers of the metabolic syndrome. PLoS ONE 2014, 9, e107392. [Google Scholar] [CrossRef]
- Kinross, J.M.; Darzi, A.W.; Nicholson, J.K. Gut microbiome-host interactions in health and disease. Genome Med. 2011, 3, 14. [Google Scholar] [CrossRef] [PubMed]
- Walsh, C.J.; Guinane, C.M.; O’Toole, P.W.; Cotter, P.D. Beneficial modulation of the gut microbiota. FEBS Lett. 2014, 588, 4120–4130. [Google Scholar] [CrossRef] [PubMed]
- Hennessy, A.A.; Ross, R.P.; Fitzgerald, G.F.; Caplice, N.; Stanton, C. Role of the gut in modulating lipoprotein metabolism. Curr. Cardiol. Rep. 2014, 16, 515. [Google Scholar] [CrossRef] [PubMed]
- O’Mahony, S.M.; Clarke, G.; Borre, Y.; Dinan, T.; Cryan, J. Serotonin, tryptophan metabolism and the brain-gut-microbiome axis. Behav. Brain Res. 2015, 277, 32–48. [Google Scholar] [CrossRef]
- Ramakrishna, B.S. Role of the gut microbiota in human nutrition and metabolism. J. Gastroenterol. Hepatol. 2013, 28, 9–17. [Google Scholar] [CrossRef]
- Fuller, M. Determination of protein and amino acid digestibility in foods including implications of gut microbial amino acid synthesis. Br. J. Nutr. 2012, 108, S238–S246. [Google Scholar] [CrossRef]
- Dutton, R.J.; Turnbaugh, P.J. Taking a metagenomic view of human nutrition. Curr. Opin. Clin. Nutr. Metab. Care 2012, 15, 448–454. [Google Scholar] [CrossRef]
- Kau, A.L.; Ahern, P.P.; Griffin, N.W.; Goodman, A.L.; Gordon, J.I. Human nutrition, the gut microbiome and the immune system. Nature 2011, 474, 327. [Google Scholar] [CrossRef]
- Brestoff, J.R.; Artis, D. Commensal bacteria at the interface of host metabolism and the immune system. Nat. Immunol. 2013, 14, 676. [Google Scholar] [CrossRef]
- Hooper, L.V.; Littman, D.R.; Macpherson, A.J. Interactions between the microbiota and the immune system. Science 2012, 336, 1268–1273. [Google Scholar] [CrossRef]
- Lee, Y.K.; Mazmanian, S.K. Has the microbiota played a critical role in the evolution of the adaptive immune system? Science 2010, 330, 1768–1773. [Google Scholar] [CrossRef] [PubMed]
- Maslowski, K.M.; Mackay, C.R. Diet, gut microbiota and immune responses. Nat. Immunol. 2010, 12, 5. [Google Scholar] [CrossRef] [PubMed]
- Thaiss, C.A.; Zmora, N.; Levy, M.; Elinav, E. The microbiome and innate immunity. Nature 2016, 535, 65. [Google Scholar] [CrossRef] [PubMed]
- Zackular, J.P.; Rogers, M.A.; Ruffin, M.T.; Schloss, P.D. The human gut microbiome as a screening tool for colorectal cancer. Cancer Prev. Res. 2014, 7, 1112–1121. [Google Scholar] [CrossRef]
- Feng, Q.; Liang, S.; Jia, H.; Stadlmayr, A.; Tang, L.; Lan, Z.; Zhang, D.; Xia, H.; Xu, X.; Jie, Z. Gut microbiome development along the colorectal adenoma–carcinoma sequence. Nat. Commun. 2015, 6, 6528. [Google Scholar] [CrossRef] [PubMed]
- Bajaj, J.S.; Heuman, D.M.; Hylemon, P.B.; Sanyal, A.J.; White, M.B.; Monteith, P.; Noble, N.A.; Unser, A.B.; Daita, K.; Fisher, A.R. Altered profile of human gut microbiome is associated with cirrhosis and its complications. J. Hepatol. 2014, 60, 940–947. [Google Scholar] [CrossRef]
- Del Chierico, F.; Nobili, V.; Vernocchi, P.; Russo, A.; Stefanis, C.D.; Gnani, D.; Furlanello, C.; Zandonà, A.; Paci, P.; Capuani, G. Gut microbiota profiling of pediatric nonalcoholic fatty liver disease and obese patients unveiled by an integrated meta-omics-based approach. Hepatology 2017, 65, 451–464. [Google Scholar] [CrossRef]
- Schippa, S.; Iebba, V.; Barbato, M.; Di Nardo, G.; Totino, V.; Checchi, M.P.; Longhi, C.; Maiella, G.; Cucchiara, S.; Conte, M.P. A distinctive ‘microbial signature’ in celiac pediatric patients. BMC Microbiol. 2010, 10, 175. [Google Scholar] [CrossRef]
- Plummer, M.; Franceschi, S.; Vignat, J.; Forman, D.; de Martel, C. Global burden of gastric cancer attributable to Helicobacter pylori. Int. J. Cancer 2015, 136, 487–490. [Google Scholar] [CrossRef]
- Finegold, S.M.; Dowd, S.E.; Gontcharova, V.; Liu, C.; Henley, K.E.; Wolcott, R.D.; Youn, E.; Summanen, P.H.; Granpeesheh, D.; Dixon, D. Pyrosequencing study of fecal microflora of autistic and control children. Anaerobe 2010, 16, 444–453. [Google Scholar] [CrossRef]
- Finegold, S.M.; Downes, J.; Summanen, P.H. Microbiology of regressive autism. Anaerobe 2012, 18, 260–262. [Google Scholar] [CrossRef] [PubMed]
- Mezzelani, A.; Landini, M.; Facchiano, F.; Raggi, M.E.; Villa, L.; Molteni, M.; De Santis, B.; Brera, C.; Caroli, A.M.; Milanesi, L. Environment, dysbiosis, immunity and sex-specific susceptibility: A translational hypothesis for regressive autism pathogenesis. Nutr. Neurosci. 2015, 18, 145–161. [Google Scholar] [CrossRef] [PubMed]
- Scheperjans, F.; Aho, V.; Pereira, P.A.B.; Koskinen, K.; Paulin, L.; Pekkonen, E.; Haapaniemi, E.; Kaakkola, S.; Eerola-Rautio, J.; Pohja, M.; et al. Gut microbiota are related to Parkinson’s disease and clinical phenotype. Mov. Disord. 2015, 30, 350–358. [Google Scholar] [CrossRef] [PubMed]
- Larsen, N.; Vogensen, F.K.; Van Den Berg, F.W.J.; Nielsen, D.S.; Andreasen, A.S.; Pedersen, B.K.; Al-Soud, W.A.; Sørensen, S.J.; Hansen, L.H.; Jakobsen, M. Gut Microbiota in Human Adults with Type 2 Diabetes Differs from Non-Diabetic Adults. PLoS ONE 2010, 5, e9085. [Google Scholar] [CrossRef]
- Qin, J.; Li, Y.; Cai, Z.; Li, S.; Zhu, J.; Zhang, F.; Liang, S.; Zhang, W.; Guan, Y.; Shen, D.; et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 2012, 490, 55–60. [Google Scholar] [CrossRef] [PubMed]
- Dicksved, J.; Halfvarson, J.; Rosenquist, M.; Järnerot, G.; Tysk, C.; Apajalahti, J.; Engstrand, L.; Jansson, J.K. Molecular analysis of the gut microbiota of identical twins with Crohn’s disease. ISME J. 2008, 2, 716. [Google Scholar] [CrossRef] [PubMed]
- Sokol, H.; Lepage, P.; Seksik, P.; Doré, J.; Marteau, P. Molecular comparison of dominant microbiota associated with injured versus healthy mucosa in ulcerative colitis. Gut 2007, 56, 152–154. [Google Scholar] [CrossRef][Green Version]
- Sokol, H.; Seksik, P.; Furet, J.; Firmesse, O.; Nion-Larmurier, I.; Beaugerie, L.; Cosnes, J.; Corthier, G.; Marteau, P.; Doré, J. Low counts of Faecalibacterium prausnitzii in colitis microbiota. Inflamm. Bowel Dis. 2009, 15, 1183–1189. [Google Scholar] [CrossRef]
- Lepage, P.; Häsler, R.; Spehlmann, M.E.; Rehman, A.; Zvirbliene, A.; Begun, A.; Ott, S.; Kupcinskas, L.; Doré, J.; Raedler, A. Twin study indicates loss of interaction between microbiota and mucosa of patients with ulcerative colitis. Gastroenterology 2011, 141, 227–236. [Google Scholar] [CrossRef]
- Drossman, D.A. Functional gastrointestinal disorders: History, pathophysiology, clinical features, and Rome IV. Gastroenterology 2016, 150, 1262–1279.e1262. [Google Scholar] [CrossRef]
- Nafarin, A.R.; Hegar, B.; Sjakti, H.A.; Vandenplas, Y. Gut microbiome pattern in adolescents with functional gastrointestinal disease. Int. J. Pediatr. Adolesc. Med. 2019, 6, 12–15. [Google Scholar] [CrossRef] [PubMed]
- Shin, A.; Preidis, G.A.; Shulman, R.; Kashyap, P. The gut microbiome in adult and pediatric functional gastrointestinal disorders. Clin. Gastroenterol. Hepatol. 2019, 17, 256–274. [Google Scholar] [CrossRef] [PubMed]
- Foster, J.A.; Rinaman, L.; Cryan, J.F. Stress & the gut-brain axis: Regulation by the microbiome. Neurobiol Stress 2017, 7, 124–136. [Google Scholar] [CrossRef] [PubMed]
- Grenham, S.; Clarke, G.; Cryan, J.F.; Dinan, T.G. Brain–gut–microbe communication in health and disease. Front. Physiol. 2011, 2, 94. [Google Scholar] [CrossRef] [PubMed]
- Sampson, T.R.; Mazmanian, S.K. Control of brain development, function, and behavior by the microbiome. Cell Host Microbe 2015, 17, 565–576. [Google Scholar] [CrossRef] [PubMed]
- Sharon, G.; Sampson, T.R.; Geschwind, D.H.; Mazmanian, S.K. The central nervous system and the gut microbiome. Cell 2016, 167, 915–932. [Google Scholar] [CrossRef]
- Rieder, R.; Wisniewski, P.J.; Alderman, B.L.; Campbell, S.C. Microbes and mental health: A review. Brain Behav. Immun. 2017, 66, 9–17. [Google Scholar] [CrossRef]
- Bruce-Keller, A.J.; Salbaum, J.M.; Berthoud, H.-R. Harnessing gut microbes for mental health: Getting from here to there. Biol. Psychiatry 2018, 83, 214–223. [Google Scholar] [CrossRef]
- Archie, E.A.; Tung, J. Social behavior and the microbiome. Curr. Opin. Behav. Sci. 2015, 6, 28–34. [Google Scholar] [CrossRef]
- Gomez, A.; Luckey, D.; Taneja, V. The gut microbiome in autoimmunity: Sex matters. Clin. Immunol. 2015, 159, 154–162. [Google Scholar] [CrossRef]
- Clemente, J.C.; Manasson, J.; Scher, J.U. The role of the gut microbiome in systemic inflammatory disease. BMJ 2018, 360, j5145. [Google Scholar] [CrossRef] [PubMed]
- Mielcarz, D.W.; Kasper, L.H. The gut microbiome in multiple sclerosis. Curr. Treat. Options Neurol. 2015, 17, 18. [Google Scholar] [CrossRef] [PubMed]
- Weis, M. Impact of the gut microbiome in cardiovascular and autoimmune diseases. Clin. Sci. 2018, 132, 2387–2389. [Google Scholar] [CrossRef] [PubMed]
- Zitvogel, L.; Ayyoub, M.; Routy, B.; Kroemer, G. Microbiome and anticancer immunosurveillance. Cell 2016, 165, 276–287. [Google Scholar] [CrossRef] [PubMed]
- Gopalakrishnan, V.; Helmink, B.A.; Spencer, C.N.; Reuben, A.; Wargo, J.A. The influence of the gut microbiome on cancer, immunity, and cancer immunotherapy. Cancer Cell 2018, 33, 570–580. [Google Scholar] [CrossRef] [PubMed]
- Wong, S.H.; Kwong, T.N.; Wu, C.-Y.; Yu, J. Clinical applications of gut microbiota in cancer biology. Semin. Cancer Biol. 2019, 55, 28–36. [Google Scholar] [CrossRef]
- Bhutia, Y.D.; Ogura, J.; Sivaprakasam, S.; Ganapathy, V. Gut microbiome and colon cancer: Role of bacterial metabolites and their molecular targets in the host. Curr. Colorectal Cancer Rep. 2017, 13, 111–118. [Google Scholar] [CrossRef]
- Gagnière, J.; Raisch, J.; Veziant, J.; Barnich, N.; Bonnet, R.; Buc, E.; Bringer, M.-A.; Pezet, D.; Bonnet, M. Gut microbiota imbalance and colorectal cancer. World J. Gastroenterol. 2016, 22, 501. [Google Scholar] [CrossRef]
- Goubet, A.-G.; Daillère, R.; Routy, B.; Derosa, L.; Roberti, P.M.; Zitvogel, L. The impact of the intestinal microbiota in therapeutic responses against cancer. Comptes Rendus Boil. 2018, 341, 284–289. [Google Scholar] [CrossRef]
- West, N.R.; Powrie, F. Immunotherapy not working? Check your microbiota. Cancer Cell 2015, 28, 687–689. [Google Scholar] [CrossRef]
- Singh, V.; San Yeoh, B.; Vijay-Kumar, M. Gut microbiome as a novel cardiovascular therapeutic target. Curr. Opin. Pharmacol. 2016, 27, 8–12. [Google Scholar] [CrossRef] [PubMed]
- Kitai, T.; Tang, W.W. The Role and Impact of gut microbiota in cardiovascular disease. Revista Española de Cardiología 2017, 70, 799–800. [Google Scholar] [CrossRef] [PubMed]
- Antza, C.; Stabouli, S.; Kotsis, V. Gut microbiota in kidney disease and hypertension. Pharmacol. Res. 2018, 130, 198–203. [Google Scholar] [CrossRef] [PubMed]
- Mazidi, M.; Rezaie, P.; Kengne, A.P.; Mobarhan, M.G.; Ferns, G.A. Gut microbiome and metabolic syndrome. Diabetes Metab. Syndr. Clin. Res. Rev. 2016, 10, S150–S157. [Google Scholar] [CrossRef]
- John, G.K.; Mullin, G.E. The Gut Microbiome and Obesity. Curr. Oncol. Rep. 2016, 18, 45. [Google Scholar] [CrossRef]
- Mondot, S.; Lepage, P. The human gut microbiome and its dysfunctions through the meta-omics prism. Ann. N. Y. Acad. Sci. 2016, 1372, 9–19. [Google Scholar] [CrossRef]
- Segata, N.; Boernigen, D.; Tickle, T.L.; Morgan, X.C.; Garrett, W.S.; Huttenhower, C. Computational meta’omics for microbial community studies. Mol. Syst. Biol. 2013, 9, 666. [Google Scholar] [CrossRef]
- Iwai, S.; Weinmaier, T.; Schmidt, B.L.; Albertson, D.G.; Poloso, N.J.; Dabbagh, K.; DeSantis, T.Z. Piphillin: Improved prediction of metagenomic content by direct inference from human microbiomes. PLoS ONE 2016, 11, e0166104. [Google Scholar] [CrossRef]
- Chen, I.-M.A.; Markowitz, V.M.; Chu, K.; Palaniappan, K.; Szeto, E.; Pillay, M.; Ratner, A.; Huang, J.; Andersen, E.; Huntemann, M.; et al. IMG/M: Integrated genome and metagenome comparative data analysis system. Nucleic Acids Res. 2016, 45, D507–D516. [Google Scholar] [CrossRef]
- Dehal, P.S.; Joachimiak, M.P.; Price, M.N.; Bates, J.T.; Baumohl, J.K.; Chivian, D.; Friedland, G.D.; Huang, K.H.; Keller, K.; Novichkov, P.S.; et al. MicrobesOnline: An integrated portal for comparative and functional genomics. Nucleic Acids Res. 2009, 38, D396–D400. [Google Scholar] [CrossRef]
- Uchiyama, I. MBGD: Microbial genome database for comparative analysis. Nucleic Acids Res. 2003, 31, 58–62. [Google Scholar] [CrossRef] [PubMed]
- Page, A.J.; Cummins, C.A.; Hunt, M.; Wong, V.K.; Reuter, S.; Holden, M.T.G.; Fookes, M.; Falush, D.; Keane, J.A.; Parkhill, J. Roary: Rapid large-scale prokaryote pan genome analysis. Bioinformatics 2015, 31, 3691–3693. [Google Scholar] [CrossRef] [PubMed]
- Yoon, S.-H.; Ha, S.-M.; Kwon, S.; Lim, J.; Kim, Y.; Seo, H.; Chun, J. Introducing EzBioCloud: A taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int. J. Syst. Evol. Microbiol. 2017, 67, 1613. [Google Scholar] [CrossRef] [PubMed]
- Whiteside, M.D.; Winsor, G.L.; Laird, M.R.; Brinkman, F.S.L. OrtholugeDB: A bacterial and archaeal orthology resource for improved comparative genomic analysis. Nucleic Acids Res. 2012, 41, D366–D376. [Google Scholar] [CrossRef] [PubMed]
- Blom, J.; Kreis, J.; Spänig, S.; Juhre, T.; Bertelli, C.; Ernst, C.; Goesmann, A. EDGAR 2.0: An enhanced software platform for comparative gene content analyses. Nucleic Acids Res. 2016, 44, W22–W28. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Blom, J.; Glaeser, S.P.; Jaenicke, S.; Juhre, T.; Rupp, O.; Schwengers, O.; Spänig, S.; Goesmann, A. A review of bioinformatics platforms for comparative genomics. Recent developments of the EDGAR 2.0 platform and its utility for taxonomic and phylogenetic studies. J. Biotechnol. 2017, 261, 2–9. [Google Scholar] [CrossRef] [PubMed]
- Wilkinson, T.J.; Huws, S.A.; Edwards, J.E.; Kingston-Smith, A.H.; Siu-Ting, K.; Hughes, M.; Rubino, F.; Friedersdorff, M.; Creevey, C.J. CowPI: A Rumen Microbiome Focussed Version of the PICRUSt Functional Inference Software. Front. Microbiol. 2018, 9. [Google Scholar] [CrossRef]
- Langille, M.G. Exploring Linkages between Taxonomic and Functional Profiles of the Human Microbiome. MSystems 2018, 3, e00163-17. [Google Scholar] [CrossRef]
- Odamaki, T.; Kato, K.; Sugahara, H.; Hashikura, N.; Takahashi, S.; Xiao, J.-Z.; Abe, F.; Osawa, R. Age-related changes in gut microbiota composition from newborn to centenarian: A cross-sectional study. BMC Microbiol. 2016, 16, 90. [Google Scholar] [CrossRef]
- Aßhauer, K.P.; Wemheuer, B.; Daniel, R.; Meinicke, P. Tax4Fun: Predicting functional profiles from metagenomic 16S rRNA data. Bioinformatics 2015, 31, 2882–2884. [Google Scholar] [CrossRef]
- Lurie-Weinberger, M.N.; Gophna, U. Archaea in and on the Human Body: Health Implications and Future Directions. PLOS Pathog. 2015, 11, e1004833. [Google Scholar] [CrossRef] [PubMed]
- Koskinen, K.; Pausan, M.R.; Perras, A.K.; Beck, M.; Bang, C.; Mora, M.; Schilhabel, A.; Schmitz, R.; Moissl-Eichinger, C. First insights into the diverse human archaeome: Specific detection of archaea in the gastrointestinal tract, lung, and nose and on skin. mBio 2017, 8, e00824-17. [Google Scholar] [CrossRef] [PubMed]
- Miller, T.L.; Wolin, M.; De Macario, E.C.; Macario, A. Isolation of Methanobrevibacter smithii from human feces. Appl. Environ. Microbiol. 1982, 43, 227–232. [Google Scholar] [CrossRef] [PubMed]
- Miller, T.L.; Wolin, M.J. Methanosphaera stadtmaniae gen. nov., sp. nov.: A species that forms methane by reducing methanol with hydrogen. Arch. Microbiol. 1985, 141, 116–122. [Google Scholar] [CrossRef] [PubMed]
- Dridi, B.; Fardeau, M.-L.; Ollivier, B.; Raoult, D.; Drancourt, M. Methanomassiliicoccus luminyensis gen. nov., sp. nov., a methanogenic archaeon isolated from human faeces. Int. J. Syst. Evol. Microbiol. 2012, 62, 1902–1907. [Google Scholar] [CrossRef] [PubMed]
- Bang, C.; Weidenbach, K.; Gutsmann, T.; Heine, H.; Schmitz, R.A. The intestinal archaea Methanosphaera stadtmanae and Methanobrevibacter smithii activate human dendritic cells. PLoS ONE 2014, 9, e99411. [Google Scholar] [CrossRef]
- Lecours, P.B.; Marsolais, D.; Cormier, Y.; Berberi, M.; Haché, C.; Bourdages, R.; Duchaine, C. Increased prevalence of Methanosphaera stadtmanae in inflammatory bowel diseases. PLoS ONE 2014, 9, e87734. [Google Scholar] [CrossRef]
- Million, M.; Angelakis, E.; Maraninchi, M.; Henry, M.; Giorgi, R.; Valero, R.; Vialettes, B.; Raoult, D. Correlation between body mass index and gut concentrations of Lactobacillus reuteri, Bifidobacterium animalis, Methanobrevibacter smithii and Escherichia coli. Int. J. Obes. 2013, 37, 1460. [Google Scholar] [CrossRef]
- Le Chatelier, E.; Nielsen, T.; Qin, J.; Prifti, E.; Hildebrand, F.; Falony, G.; Almeida, M.; Arumugam, M.; Batto, J.-M.; Kennedy, S. Richness of human gut microbiome correlates with metabolic markers. Nature 2013, 500, 541. [Google Scholar] [CrossRef]
- Schwiertz, A.; Taras, D.; Schäfer, K.; Beijer, S.; Bos, N.A.; Donus, C.; Hardt, P.D. Microbiota and SCFA in Lean and Overweight Healthy Subjects. Obesity 2010, 18, 190–195. [Google Scholar] [CrossRef]
- Pimentel, M.; Gunsalus, R.P.; Rao, S.S.; Zhang, H. Methanogens in human health and disease. Am. J. Gastroenterol. Suppl. 2012, 1, 28. [Google Scholar] [CrossRef]
- McKay, L.F.; Eastwood, M.; Brydon, W. Methane excretion in man--a study of breath, flatus, and faeces. Gut 1985, 26, 69–74. [Google Scholar] [CrossRef] [PubMed]
- Hallen-Adams, H.E.; Suhr, M.J. Fungi in the healthy human gastrointestinal tract. Virulence 2017, 8, 352–358. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.K.; Yang, Y.S.; Stefka, A.T.; Sun, G.; Peng, L.H. Review article: Fungal microbiota and digestive diseases. Aliment. Pharmacol. Ther. 2014, 39, 751–766. [Google Scholar] [CrossRef] [PubMed]
- Gouba, N.; Drancourt, M. Digestive tract mycobiota: A source of infection. Médecine et Maladies Infectieuses 2015, 45, 9–16. [Google Scholar] [CrossRef]
- Dollive, S.; Peterfreund, G.L.; Sherrill-Mix, S.; Bittinger, K.; Sinha, R.; Hoffmann, C.; Nabel, C.S.; Hill, D.A.; Artis, D.; Bachman, M.A. A tool kit for quantifying eukaryotic rRNA gene sequences from human microbiome samples. Genome Biol. 2012, 13, R60. [Google Scholar] [CrossRef]
- Ott, S.J.; Kühbacher, T.; Musfeldt, M.; Rosenstiel, P.; Hellmig, S.; Rehman, A.; Drews, O.; Weichert, W.; Timmis, K.N.; Schreiber, S. Fungi and inflammatory bowel diseases: Alterations of composition and diversity. Scand. J. Gastroenterol. 2008, 43, 831–841. [Google Scholar] [CrossRef]
- Richard, M.L.; Lamas, B.; Liguori, G.; Hoffmann, T.W.; Sokol, H. Gut Fungal Microbiota: The Yin and Yang of Inflammatory Bowel Disease. Inflamm. Bowel Dis. 2014, 21, 656–665. [Google Scholar] [CrossRef]
- Ramaswamy, K.; Correa, M.; Koshy, A. Non-healing gastric ulcer associated with Candida infection. Indian J. Med Microbiol. 2007, 25, 57. [Google Scholar]
- Santelmann, H.; Howard, J.M. Yeast metabolic products, yeast antigens and yeasts as possible triggers for irritable bowel syndrome. Eur. J. Gastroenterol. Hepatol. 2005, 17, 21–26. [Google Scholar] [CrossRef]
- Krause, R.; Reisinger, E. Candida and antibiotic-associated diarrhoea. Clin. Microbiol. Infect. 2005, 11, 1–2. [Google Scholar] [CrossRef] [PubMed]
- Stringer, A.M.; Gibson, R.J.; Logan, R.M.; Bowen, J.M.; Yeoh, A.S.; Hamilton, J.; Keefe, D.M. Gastrointestinal microflora and mucins may play a critical role in the development of 5-fluorouracil-induced gastrointestinal mucositis. Exp. Biol. Med. 2009, 234, 430–441. [Google Scholar] [CrossRef] [PubMed]
- Cadwell, K. Expanding the Role of the Virome: Commensalism in the Gut. J. Virol. 2015, 89, 1951–1953. [Google Scholar] [CrossRef] [PubMed]
- Duerkop, B.A.; Hooper, L.V. Resident viruses and their interactions with the immune system. Nat. Immunol. 2013, 14, 654. [Google Scholar] [CrossRef] [PubMed]
- Foca, A.; Liberto, M.C.; Quirino, A.; Marascio, N.; Zicca, E.; Pavia, G. Gut Inflammation and Immunity: What Is the Role of the Human Gut Virome? Mediat. Inflamm. 2015, 2015, 7. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Nava, G.M.; Stappenbeck, T.S. Host genetic susceptibility, dysbiosis and viral triggers in IBD. Curr. Opin. Gastroenterol. 2011, 27, 321. [Google Scholar] [CrossRef] [PubMed]
- Cario, E. Microbiota and innate immunity in intestinal inflammation and neoplasia. Curr. Opin. Gastroenterol. 2013, 29, 85–91. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Wang, W.-L.; Xu, S.-Y.; Ren, Z.-G.; Tao, L.; Jiang, J.-W.; Zheng, S.-S. Application of metagenomics in the human gut microbiome. World J. Gastroenterol. WJG 2015, 21, 803. [Google Scholar] [CrossRef]
- Zou, Y.; Xue, W.; Luo, G.; Deng, Z.; Qin, P.; Guo, R.; Sun, H.; Xia, Y.; Liang, S.; Dai, Y.; et al. 1,520 reference genomes from cultivated human gut bacteria enable functional microbiome analyses. Nat. Biotechnol. 2019, 37, 179–185. [Google Scholar] [CrossRef]
- Gosalbes, M.J.; Durbán, A.; Pignatelli, M.; Abellan, J.J.; Jiménez-Hernández, N.; Pérez-Cobas, A.E.; Latorre, A.; Moya, A. Metatranscriptomic Approach to Analyze the Functional Human Gut Microbiota. PLoS ONE 2011, 6, e17447. [Google Scholar] [CrossRef] [PubMed]
- Franzosa, E.A.; Morgan, X.C.; Segata, N.; Waldron, L.; Reyes, J.; Earl, A.M.; Giannoukos, G.; Boylan, M.R.; Ciulla, D.; Gevers, D.; et al. Relating the metatranscriptome and metagenome of the human gut. Proc. Natl. Acad. Sci. USA 2014, 111, E2329–E2338. [Google Scholar] [CrossRef] [PubMed]
- Verberkmoes, N.C.; Russell, A.L.; Shah, M.; Godzik, A.; Rosenquist, M.; Halfvarson, J.; Lefsrud, M.G.; Apajalahti, J.; Tysk, C.; Hettich, R.L. Shotgun metaproteomics of the human distal gut microbiota. ISME J. 2009, 3, 179. [Google Scholar] [CrossRef] [PubMed]
- Erickson, A.R.; Cantarel, B.L.; Lamendella, R.; Darzi, Y.; Mongodin, E.F.; Pan, C.; Shah, M.; Halfvarson, J.; Tysk, C.; Henrissat, B. Integrated metagenomics/metaproteomics reveals human host-microbiota signatures of Crohn’s disease. PLoS ONE 2012, 7, e49138. [Google Scholar] [CrossRef] [PubMed]
- Kolmeder, C.A.; De Been, M.; Nikkilä, J.; Ritamo, I.; Mättö, J.; Valmu, L.; Salojärvi, J.; Palva, A.; Salonen, A.; de Vos, W.M. Comparative metaproteomics and diversity analysis of human intestinal microbiota testifies for its temporal stability and expression of core functions. PLoS ONE 2012, 7, e29913. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.; Yu, N.; Ding, X.; Wang, J.; Pan, Y. Dime: A novel framework for de novo metagenomic sequence assembly. J. Comput. Biol. 2015, 22, 159–177. [Google Scholar] [CrossRef]
- Laserson, J.; Jojic, V.; Koller, D. Genovo: De novo assembly for metagenomes. J. Comput. Biol. 2011, 18, 429–443. [Google Scholar] [CrossRef]
- Pell, J.; Hintze, A.; Canino-Koning, R.; Howe, A.; Tiedje, J.M.; Brown, C.T. Scaling metagenome sequence assembly with probabilistic de Bruijn graphs. Proc. Natl. Acad. Sci. USA 2012, 109, 13272–13277. [Google Scholar] [CrossRef]
- Lai, B.; Ding, R.; Li, Y.; Duan, L.; Zhu, H. A de novo metagenomic assembly program for shotgun DNA reads. Bioinformatics 2012, 28, 1455–1462. [Google Scholar] [CrossRef]
- Peng, Y.; Leung, H.C.M.; Yiu, S.M.; Chin, F.Y.L. Meta-IDBA: A de Novo assembler for metagenomic data. Bioinformatics 2011, 27, i94–i101. [Google Scholar] [CrossRef]
- Treangen, T.J.; Koren, S.; Astrovskaya, I.; Sommer, D.; Liu, B.; Pop, M. MetAMOS: A metagenomic assembly and analysis pipeline for AMOS. Genome Biol. 2011, 12, P25. [Google Scholar] [CrossRef]
- Namiki, T.; Hachiya, T.; Tanaka, H.; Sakakibara, Y. MetaVelvet: An extension of Velvet assembler to de novo metagenome assembly from short sequence reads. Nucleic Acids Res. 2012, 40, e155. [Google Scholar] [CrossRef] [PubMed]
- Kultima, J.R.; Sunagawa, S.; Li, J.; Chen, W.; Chen, H.; Mende, D.R.; Arumugam, M.; Pan, Q.; Liu, B.; Qin, J. MOCAT: A metagenomics assembly and gene prediction toolkit. PLoS ONE 2012, 7, e47656. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Zhu, H.; Ruan, J.; Qian, W.; Fang, X.; Shi, Z.; Li, Y.; Li, S.; Shan, G.; Kristiansen, K.; et al. De novo assembly of human genomes with massively parallel short read sequencing. Genome Res. 2010, 20, 265–272. [Google Scholar] [CrossRef] [PubMed]
- Ye, Y.; Tang, H. An ORFome assembly approach to metagenomics sequences analysis. J. Bioinform. Comput. Biol. 2009, 7, 455–471. [Google Scholar] [CrossRef]
- Kim, M.; Ligo, J.G.; Emad, A.; Farnoud, F.; Milenkovic, O.; Veeravalli, V.V. MetaPar: Metagenomic sequence assembly via iterative reclassification. In Proceedings of the 2013 IEEE Global Conference on Signal and Information Processing, Austin, TX, USA, 3–5 December 2013; pp. 43–46. [Google Scholar]
- Afiahayati; Sato, K.; Sakakibara, Y. An extended genovo metagenomic assembler by incorporating paired-end information. PeerJ 2013, 1, e196. [Google Scholar] [CrossRef]
- Wu, M.; Eisen, J.A. A simple, fast, and accurate method of phylogenomic inference. Genome Biol. 2008, 9, R151. [Google Scholar] [CrossRef]
- Wu, M.; Scott, A.J. Phylogenomic analysis of bacterial and archaeal sequences with AMPHORA2. Bioinformatics 2012, 28, 1033–1034. [Google Scholar] [CrossRef]
- Kerepesi, C.; Banky, D.; Grolmusz, V. AmphoraNet: The webserver implementation of the AMPHORA2 metagenomic workflow suite. Gene 2014, 533, 538–540. [Google Scholar] [CrossRef]
- Gerlach, W.; Stoye, J. Taxonomic classification of metagenomic shotgun sequences with CARMA3. Nucleic Acids Res. 2011, 39, e91. [Google Scholar] [CrossRef]
- Gerlach, W.; Stoye, J. Taxonomic classification of metagenomic shotgun sequences with CARMA3. In Encyclopedia of Metagenomics: Genes, Genomes and Metagenomes: Basics, Methods, Databases and Tools; Nelson, K.E., Ed.; Springer: New York, NY, USA, 2015; pp. 653–660. [Google Scholar]
- Patil, K.R.; Haider, P.; Pope, P.B.; Turnbaugh, P.J.; Morrison, M.; Scheffer, T.; McHardy, A.C. Taxonomic metagenome sequence assignment with structured output models. Nat. Methods 2011, 8, 191. [Google Scholar] [CrossRef] [PubMed]
- Ounit, R.; Wanamaker, S.; Close, T.J.; Lonardi, S. CLARK: Fast and accurate classification of metagenomic and genomic sequences using discriminative k-mers. BMC Genom. 2015, 16, 236. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, T.S.; Haque, M.; Mande, S.S. DiScRIBinATE: A rapid method for accurate taxonomic classification of metagenomic sequences. BMC Bioinform. 2010, 11, S14. [Google Scholar] [CrossRef] [PubMed]
- Silva, G.G.Z.; Cuevas, D.A.; Dutilh, B.E.; Edwards, R.A. FOCUS: An alignment-free model to identify organisms in metagenomes using non-negative least squares. PeerJ 2014, 2, e425. [Google Scholar] [CrossRef] [PubMed]
- Mohammed, M.H.; Ghosh, T.S.; Reddy, R.M.; Reddy, C.V.S.K.; Singh, N.K.; Mande, S.S. INDUS-a composition-based approach for rapid and accurate taxonomic classification of metagenomic sequences. BMC Genom. 2011, 12, S4. [Google Scholar] [CrossRef]
- Horton, M.; Bodenhausen, N.; Bergelson, J. MARTA: A suite of Java-based tools for assigning taxonomic status to DNA sequences. Bioinformatics 2009, 26, 568–569. [Google Scholar] [CrossRef]
- Wang, Y.; Leung, H.C.; Yiu, S.-M.; Chin, F.Y. MetaCluster 5.0: A two-round binning approach for metagenomic data for low-abundance species in a noisy sample. Bioinformatics 2012, 28, i356–i362. [Google Scholar] [CrossRef]
- Segata, N.; Waldron, L.; Ballarini, A.; Narasimhan, V.; Jousson, O.; Huttenhower, C. Metagenomic microbial community profiling using unique clade-specific marker genes. Nat. Methods 2012, 9, 811. [Google Scholar] [CrossRef]
- Truong, D.T.; Franzosa, E.A.; Tickle, T.L.; Scholz, M.; Weingart, G.; Pasolli, E.; Tett, A.; Huttenhower, C.; Segata, N. MetaPhlAn2 for enhanced metagenomic taxonomic profiling. Nat. Methods 2015, 12, 902. [Google Scholar] [CrossRef]
- Liu, B.; Gibbons, T.; Ghodsi, M.; Pop, M. MetaPhyler: Taxonomic profiling for metagenomic sequences. In Proceedings of the 2010 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), Hong Kong, China, 18–21 December 2010; pp. 95–100. [Google Scholar]
- Kultima, J.R.; Coelho, L.P.; Forslund, K.; Huerta-Cepas, J.; Li, S.S.; Driessen, M.; Voigt, A.Y.; Zeller, G.; Sunagawa, S.; Bork, P. MOCAT2: A metagenomic assembly, annotation and profiling framework. Bioinformatics 2016, 32, 2520–2523. [Google Scholar] [CrossRef]
- Gori, F.; Folino, G.; Jetten, M.S.; Marchiori, E. MTR: Taxonomic annotation of short metagenomic reads using clustering at multiple taxonomic ranks. Bioinformatics 2010, 27, 196–203. [Google Scholar] [CrossRef] [PubMed]
- Rosen, G.L.; Reichenberger, E.R.; Rosenfeld, A.M. NBC: The Naive Bayes Classification tool webserver for taxonomic classification of metagenomic reads. Bioinformatics 2010, 27, 127–129. [Google Scholar] [CrossRef] [PubMed]
- Berger, S.A.; Stamatakis, A. PaPaRa 2.0: A vectorized algorithm for probabilistic phylogeny-aware alignment extension. 2012. Available online: https://pdfs.semanticscholar.org/2b04/11608d4b9fe622ea7aa4df57a1913c625530.pdf (accessed on 1 March 2020).
- Darling, A.E.; Jospin, G.; Lowe, E.; Matsen, F.A.I.V.; Bik, H.M.; Eisen, J.A. PhyloSift: Phylogenetic analysis of genomes and metagenomes. PeerJ 2014, 2, e243. [Google Scholar] [CrossRef] [PubMed]
- Brady, A.; Salzberg, S.L. Phymm and PhymmBL: Metagenomic phylogenetic classification with interpolated Markov models. Nat. Methods 2009, 6, 673. [Google Scholar] [CrossRef]
- Nalbantoglu, O.U.; Way, S.F.; Hinrichs, S.H.; Sayood, K. RAIphy: Phylogenetic classification of metagenomics samples using iterative refinement of relative abundance index profiles. BMC Bioinform. 2011, 12, 41. [Google Scholar] [CrossRef] [PubMed]
- MacDonald, N.J.; Parks, D.H.; Beiko, R.G. RITA: Rapid identification of high-confidence taxonomic assignments for metagenomic data. Nucleic Acids Res. 2012, 40, e111. [Google Scholar] [CrossRef] [PubMed]
- Monzoorul Haque, M.; Ghosh, T.S.; Komanduri, D.; Mande, S.S. SOrt-ITEMS: Sequence orthology based approach for improved taxonomic estimation of metagenomic sequences. Bioinformatics 2009, 25, 1722–1730. [Google Scholar] [CrossRef]
- Mohammed, M.H.; Ghosh, T.S.; Singh, N.K.; Mande, S.S. SPHINX—An algorithm for taxonomic binning of metagenomic sequences. Bioinformatics 2010, 27, 22–30. [Google Scholar] [CrossRef]
- Diaz, N.N.; Krause, L.; Goesmann, A.; Niehaus, K.; Nattkemper, T.W. TACOA–Taxonomic classification of environmental genomic fragments using a kernelized nearest neighbor approach. BMC Bioinform. 2009, 10, 56. [Google Scholar] [CrossRef]
- Schreiber, F.; Gumrich, P.; Daniel, R.; Meinicke, P. Treephyler: Fast taxonomic profiling of metagenomes. Bioinformatics 2010, 26, 960–961. [Google Scholar] [CrossRef]
- Abubucker, S.; Segata, N.; Goll, J.; Schubert, A.M.; Izard, J.; Cantarel, B.L.; Rodriguez-Mueller, B.; Zucker, J.; Thiagarajan, M.; Henrissat, B. Metabolic reconstruction for metagenomic data and its application to the human microbiome. PLoS Comput. Biol. 2012, 8, e1002358. [Google Scholar] [CrossRef] [PubMed]
- Hyland, C.; Pinney, J.W.; McConkey, G.A.; Westhead, D.R. metaSHARK: A WWW platform for interactive exploration of metabolic networks. Nucleic Acids Res. 2006, 34, W725–W728. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Larsen, P.E.; Collart, F.R.; Field, D.; Meyer, F.; Keegan, K.P.; Henry, C.S.; McGrath, J.; Quinn, J.; Gilbert, J.A. Predicted Relative Metabolomic Turnover (PRMT): Determining metabolic turnover from a coastal marine metagenomic dataset. Microb. Inform. Exp. 2011, 1, 4. [Google Scholar] [CrossRef] [PubMed]
- Li, W. Analysis and comparison of very large metagenomes with fast clustering and functional annotation. BMC Bioinform. 2009, 10, 359. [Google Scholar] [CrossRef]
- Friedman, J.; Alm, E.J. Inferring correlation networks from genomic survey data. PLoS Comput. Biol. 2012, 8, e1002687. [Google Scholar] [CrossRef]
- Schwager, E.; Weingart, G.; Bielski, C.; Huttenhower, C. CCREPE: Compositionality Corrected by Permutation and Renormalization. 2014. Available online: http://citeseerx.ist.psu.edu/viewdoc/download?doi=10.1.1.683.4029&rep=rep1&type=pdf (accessed on 1 March 2020).
- Peng, Y.; Leung, H.C.M.; Yiu, S.M.; Chin, F.Y.L. IDBA-UD: A de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth. Bioinformatics 2012, 28, 1420–1428. [Google Scholar] [CrossRef]
- Harrington, E.D.; Arumugam, M.; Raes, J.; Bork, P.; Relman, D.A. SmashCell: A software framework for the analysis of single-cell amplified genome sequences. Bioinformatics 2010, 26, 2979–2980. [Google Scholar] [CrossRef][Green Version]
- Khosrovian, K.; Pfahl, D.; Garousi, V. GENSIM 2.0: A customizable process simulation model for software process evaluation. In Proceedings of the International Conference on Software Process, Leipzig, Germany, 10–11 May 2008; pp. 294–306. [Google Scholar]
- Richter, D.C.; Ott, F.; Auch, A.F.; Schmid, R.; Huson, D.H. MetaSim—A Sequencing Simulator for Genomics and Metagenomics. PLoS ONE 2008, 3, e3373. [Google Scholar] [CrossRef]
- White, J.R.; Nagarajan, N.; Pop, M. Statistical methods for detecting differentially abundant features in clinical metagenomic samples. PLoS Comput. Biol. 2009, 5, e1000352. [Google Scholar] [CrossRef]
- Segata, N.; Izard, J.; Waldron, L.; Gevers, D.; Miropolsky, L.; Garrett, W.S.; Huttenhower, C. Metagenomic biomarker discovery and explanation. Genome Biol. 2011, 12, R60. [Google Scholar] [CrossRef]
- Kristiansson, E.; Hugenholtz, P.; Dalevi, D. ShotgunFunctionalizeR: An R-package for functional comparison of metagenomes. Bioinformatics 2009, 25, 2737–2738. [Google Scholar] [CrossRef] [PubMed]
- Knights, D.; Kuczynski, J.; Charlson, E.S.; Zaneveld, J.; Mozer, M.C.; Collman, R.G.; Bushman, F.D.; Knight, R.; Kelley, S.T. Bayesian community-wide culture-independent microbial source tracking. Nat. Methods 2011, 8, 761. [Google Scholar] [CrossRef] [PubMed]
- Seshadri, R.; Kravitz, S.A.; Smarr, L.; Gilna, P.; Frazier, M. CAMERA: A community resource for metagenomics. PLoS Biol. 2007, 5, e75. [Google Scholar] [CrossRef] [PubMed]
- Riehle, K.; Coarfa, C.; Jackson, A.; Ma, J.; Tandon, A.; Paithankar, S.; Raghuraman, S.; Mistretta, T.-A.; Saulnier, D.; Raza, S. The Genboree Microbiome Toolset and the analysis of 16S rRNA microbial sequences. BMC Bioinform. 2012, 13, S11. [Google Scholar] [CrossRef] [PubMed]
- Asnicar, F.; Weingart, G.; Tickle, T.L.; Huttenhower, C.; Segata, N. Compact graphical representation of phylogenetic data and metadata with GraPhlAn. PeerJ 2015, 3, e1029. [Google Scholar] [CrossRef] [PubMed]
- Huson, D.H.; Auch, A.F.; Qi, J.; Schuster, S.C. MEGAN analysis of metagenomic data. Genome Res. 2007, 17, 377–386. [Google Scholar] [CrossRef]
- Goll, J.; Rusch, D.B.; Tanenbaum, D.M.; Thiagarajan, M.; Li, K.; Methé, B.A.; Yooseph, S. METAREP: JCVI metagenomics reports—an open source tool for high-performance comparative metagenomics. Bioinformatics 2010, 26, 2631–2632. [Google Scholar] [CrossRef]
- Meyer, F.; Paarmann, D.; D’Souza, M.; Olson, R.; Glass, E.M.; Kubal, M.; Paczian, T.; Rodriguez, A.; Stevens, R.; Wilke, A. The metagenomics RAST server—A public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinform. 2008, 9, 386. [Google Scholar] [CrossRef]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Pena, A.G.; Goodrich, J.K.; Gordon, J.I. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335. [Google Scholar] [CrossRef]
- Arumugam, M.; Harrington, E.D.; Foerstner, K.U.; Raes, J.; Bork, P. SmashCommunity: A metagenomic annotation and analysis tool. Bioinformatics 2010, 26, 2977–2978. [Google Scholar] [CrossRef] [PubMed]
- Parks, D.H.; Beiko, R.G. STAMP: Statistical analysis of metagenomic profiles. In Encyclopedia of Metagenomics: Genes, Genomes and Metagenomes. Basics, Methods, Databases and Tools; Springer: Berlin, Germany, 2015; pp. 641–645. [Google Scholar]
- Stocker, S.; Snajder, R.; Rainer, J.; Trajanoski, S.; Gorkiewicz, G.; Trajanoski, Z.; Thallinger, G.G. SnoWMAn: High-throughput phylotyping, analysis and comparison of microbial communities. 2011; Under Revision. [Google Scholar]
- Huse, S.M.; Welch, D.B.M.; Voorhis, A.; Shipunova, A.; Morrison, H.G.; Eren, A.M.; Sogin, M.L. VAMPS: A website for visualization and analysis of microbial population structures. BMC Bioinform. 2014, 15, 41. [Google Scholar] [CrossRef] [PubMed]
- Rinschen, M.M.; Ivanisevic, J.; Giera, M.; Siuzdak, G. Identification of bioactive metabolites using activity metabolomics. Nat. Rev. Mol. Cell Biol. 2019. [Google Scholar] [CrossRef]
- Piazza, I.; Kochanowski, K.; Cappelletti, V.; Fuhrer, T.; Noor, E.; Sauer, U.; Picotti, P. A Map of Protein-Metabolite Interactions Reveals Principles of Chemical Communication. Cell 2018, 172, 358–372.e323. [Google Scholar] [CrossRef]
- Larsen, P.E.; Dai, Y. Metabolome of human gut microbiome is predictive of host dysbiosis. GigaScience 2015, 4, 42. [Google Scholar] [CrossRef]
- Holmes, E.; Li, J.V.; Marchesi, J.R.; Nicholson, J.K. Gut microbiota composition and activity in relation to host metabolic phenotype and disease risk. Cell Metab. 2012, 16, 559–564. [Google Scholar] [CrossRef]
- Rooks, M.G.; Garrett, W.S. Gut microbiota, metabolites and host immunity. Nat. Rev. Immunol. 2016, 16, 341. [Google Scholar] [CrossRef]
- Morrison, D.J.; Preston, T. Formation of short chain fatty acids by the gut microbiota and their impact on human metabolism. Gut Microbes 2016, 7, 189–200. [Google Scholar] [CrossRef]
- Psichas, A.; Sleeth, M.; Murphy, K.; Brooks, L.; Bewick, G.; Hanyaloglu, A.; Ghatei, M.; Bloom, S.; Frost, G. The short chain fatty acid propionate stimulates GLP-1 and PYY secretion via free fatty acid receptor 2 in rodents. Int. J. Obes. 2015, 39, 424. [Google Scholar] [CrossRef]
- Chambers, E.S.; Morrison, D.J.; Frost, G. Control of appetite and energy intake by SCFA: What are the potential underlying mechanisms? Proc. Nutr. Soc. 2015, 74, 328–336. [Google Scholar] [CrossRef] [PubMed]
- Corrêa-Oliveira, R.; Fachi, J.L.; Vieira, A.; Sato, F.T.; Vinolo, M.A.R. Regulation of immune cell function by short-chain fatty acids. Clin. Transl. Immunol. 2016, 5, e73. [Google Scholar] [CrossRef] [PubMed]
- Levy, M.; Blacher, E.; Elinav, E. Microbiome, metabolites and host immunity. Curr. Opin. Microbiol. 2017, 35, 8–15. [Google Scholar] [CrossRef] [PubMed]
- Claus, S.P.; Guillou, H.; Ellero-Simatos, S. The gut microbiota: A major player in the toxicity of environmental pollutants? NPJ Biofilms Microbiomes 2016, 2, 16003. [Google Scholar] [CrossRef] [PubMed]
- LeBlanc, J.G.; Milani, C.; De Giori, G.S.; Sesma, F.; Van Sinderen, D.; Ventura, M. Bacteria as vitamin suppliers to their host: A gut microbiota perspective. Curr. Opin. Biotechnol. 2013, 24, 160–168. [Google Scholar] [CrossRef] [PubMed]
- Laparra, J.M.; Sanz, Y. Interactions of gut microbiota with functional food components and nutraceuticals. Pharmacol. Res. 2010, 61, 219–225. [Google Scholar] [CrossRef] [PubMed]
- Carding, S.; Verbeke, K.; Vipond, D.T.; Corfe, B.M.; Owen, L.J. Dysbiosis of the gut microbiota in disease. Microb. Ecol. Health Dis. 2015, 26, 26191. [Google Scholar] [CrossRef]
- Yau, Y.; Leong, R.W.; Zeng, M.; Wasinger, V.C. Proteomics and metabolomics in inflammatory bowel disease. J. Gastroenterol. Hepatol. 2013, 28, 1076–1086. [Google Scholar] [CrossRef]
- Lin, H.M.; Helsby, N.A.; Rowan, D.D.; Ferguson, L.R. Using metabolomic analysis to understand inflammatory bowel diseases. Inflamm. Bowel Dis. 2011, 17, 1021–1029. [Google Scholar] [CrossRef]
- Le Gall, G.; Noor, S.O.; Ridgway, K.; Scovell, L.; Jamieson, C.; Johnson, I.T.; Colquhoun, I.J.; Kemsley, E.K.; Narbad, A. Metabolomics of fecal extracts detects altered metabolic activity of gut microbiota in ulcerative colitis and irritable bowel syndrome. J. Proteome Res. 2011, 10, 4208–4218. [Google Scholar] [CrossRef]
- Phua, L.C.; Chue, X.P.; Koh, P.K.; Cheah, P.Y.; Ho, H.K.; Chan, E.C.Y. Non-invasive fecal metabonomic detection of colorectal cancer. Cancer Biol. Ther. 2014, 15, 389–397. [Google Scholar] [CrossRef] [PubMed]
- Nicholson, J.K.; Holmes, E.; Kinross, J.; Burcelin, R.; Gibson, G.; Jia, W.; Pettersson, S. Host-gut microbiota metabolic interactions. Science 2012, 336, 1262–1267. [Google Scholar] [CrossRef] [PubMed]
- Vernocchi, P.; Del Chierico, F.; Putignani, L. Gut Microbiota Profiling: Metabolomics Based Approach to Unravel Compounds Affecting Human Health. Front. Microbiol. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Binder, H.J. Role of colonic short-chain fatty acid transport in diarrhea. Annu. Rev. Physiol. 2010, 72, 297–313. [Google Scholar] [CrossRef] [PubMed]
- Chambers, E.S.; Viardot, A.; Psichas, A.; Morrison, D.J.; Murphy, K.G.; Zac-Varghese, S.E.; MacDougall, K.; Preston, T.; Tedford, C.; Finlayson, G.S. Effects of targeted delivery of propionate to the human colon on appetite regulation, body weight maintenance and adiposity in overweight adults. Gut 2015, 64, 1744–1754. [Google Scholar] [CrossRef]
- Qiu, Y.; Cai, G.; Su, M.; Chen, T.; Liu, Y.; Xu, Y.; Ni, Y.; Zhao, A.; Cai, S.; Xu, L.X. Urinary metabonomic study on colorectal cancer. J. Proteome Res. 2010, 9, 1627–1634. [Google Scholar] [CrossRef]
- Calvani, R.; Miccheli, A.; Capuani, G.; Miccheli, A.T.; Puccetti, C.; Delfini, M.; Iaconelli, A.; Nanni, G.; Mingrone, G. Gut microbiome-derived metabolites characterize a peculiar obese urinary metabotype. Int. J. Obes. 2010, 34, 1095. [Google Scholar] [CrossRef]
- Zhao, L.; Liu, X.; Xie, L.; Gao, H.; Lin, D. 1H NMR-based metabonomic analysis of metabolic changes in streptozotocin-induced diabetic rats. Anal. Sci. 2010, 26, 1277–1282. [Google Scholar] [CrossRef]
- Zheng, X.; Xie, G.; Zhao, A.; Zhao, L.; Yao, C.; Chiu, N.H.; Zhou, Z.; Bao, Y.; Jia, W.; Nicholson, J.K. The footprints of gut microbial–mammalian co-metabolism. J. Proteome Res. 2011, 10, 5512–5522. [Google Scholar] [CrossRef]
- Amaretti, A.; Raimondi, S.; Leonardi, A.; Quartieri, A.; Rossi, M. Hydrolysis of the rutinose-conjugates flavonoids rutin and hesperidin by the gut microbiota and bifidobacteria. Nutrients 2015, 7, 2788–2800. [Google Scholar] [CrossRef]
- Taverniti, V.; Guglielmetti, S. Health-promoting properties of Lactobacillus helveticus. Front. Microbiol. 2012, 3, 392. [Google Scholar] [CrossRef] [PubMed]
- Marín, L.; Miguélez, E.M.; Villar, C.J.; Lombó, F. Bioavailability of dietary polyphenols and gut microbiota metabolism: Antimicrobial properties. BioMed Res. Int. 2015, 2015. [Google Scholar] [CrossRef] [PubMed]
- Dawson, P.A.; Lan, T.; Rao, A. Bile acid transporters. J. Lipid Res. 2009, 50, 2340–2357. [Google Scholar] [CrossRef] [PubMed]
- Suhre, K.; Meisinger, C.; Döring, A.; Altmaier, E.; Belcredi, P.; Gieger, C.; Chang, D.; Milburn, M.V.; Gall, W.E.; Weinberger, K.M. Metabolic footprint of diabetes: A multiplatform metabolomics study in an epidemiological setting. PLoS ONE 2010, 5, e13953. [Google Scholar] [CrossRef]
- Swann, J.R.; Want, E.J.; Geier, F.M.; Spagou, K.; Wilson, I.D.; Sidaway, J.E.; Nicholson, J.K.; Holmes, E. Systemic gut microbial modulation of bile acid metabolism in host tissue compartments. Proc. Natl. Acad. Sci. USA 2011, 108, 4523–4530. [Google Scholar] [CrossRef]
- Wang, Z.; Klipfell, E.; Bennett, B.J.; Koeth, R.; Levison, B.S.; DuGar, B.; Feldstein, A.E.; Britt, E.B.; Fu, X.; Chung, Y.-M. Gut flora metabolism of phosphatidylcholine promotes cardiovascular disease. Nature 2011, 472, 57. [Google Scholar] [CrossRef]
- Martin, F.-P.J.; Sprenger, N.; Montoliu, I.; Rezzi, S.; Kochhar, S.; Nicholson, J.K. Dietary modulation of gut functional ecology studied by fecal metabonomics. J. Proteome Res. 2010, 9, 5284–5295. [Google Scholar] [CrossRef]
- Bercik, P.; Denou, E.; Collins, J.; Jackson, W.; Lu, J.; Jury, J.; Deng, Y.; Blennerhassett, P.; Macri, J.; McCoy, K.D. The intestinal microbiota affect central levels of brain-derived neurotropic factor and behavior in mice. Gastroenterology 2011, 141, 599–609.e593. [Google Scholar] [CrossRef]
- Keszthelyi, D.; Troost, F.; Masclee, A. Understanding the role of tryptophan and serotonin metabolism in gastrointestinal function. Neurogastroenterol. Motil. 2009, 21, 1239–1249. [Google Scholar] [CrossRef]
- Koenig, J.E.; Spor, A.; Scalfone, N.; Fricker, A.D.; Stombaugh, J.; Knight, R.; Angenent, L.T.; Ley, R.E. Succession of microbial consortia in the developing infant gut microbiome. Proc. Natl. Acad. Sci. USA 2011, 108, 4578–4585. [Google Scholar] [CrossRef]
- Said, H.M. Intestinal absorption of water-soluble vitamins in health and disease. Biochem. J. 2011, 437, 357–372. [Google Scholar] [CrossRef] [PubMed]
- Rossi, M.; Amaretti, A. Probiotic Properties of Bifidobacteria; Caister Academic Press: Norfolk, UK, 2010. [Google Scholar]
- Magnúsdóttir, S.; Ravcheev, D.; De Crécy-Lagard, V.; Thiele, I. Systematic genome assessment of B-vitamin biosynthesis suggests co-operation among gut microbes. Front. Genet. 2015, 6, 148. [Google Scholar] [CrossRef] [PubMed]
- Hanfrey, C.C.; Pearson, B.M.; Hazeldine, S.; Lee, J.; Gaskin, D.J.; Woster, P.M.; Phillips, M.A.; Michael, A.J. Alternative spermidine biosynthetic route is critical for growth of Campylobacter jejuni and is the dominant polyamine pathway in human gut microbiota. J. Biol. Chem. 2011, 286, 43301–43312. [Google Scholar] [CrossRef] [PubMed]
- Serino, M.; Luche, E.; Gres, S.; Baylac, A.; Bergé, M.; Cenac, C.; Waget, A.; Klopp, P.; Iacovoni, J.; Klopp, C. Metabolic adaptation to a high-fat diet is associated with a change in the gut microbiota. Gut 2012, 61, 543–553. [Google Scholar] [CrossRef] [PubMed]
- Muccioli, G.G.; Naslain, D.; Bäckhed, F.; Reigstad, C.S.; Lambert, D.M.; Delzenne, N.M.; Cani, P.D. The endocannabinoid system links gut microbiota to adipogenesis. Mol. Syst. Biol. 2010, 6, 392. [Google Scholar] [CrossRef] [PubMed]
- Fiehn, O. Metabolomics—the link between genotypes and phenotypes. Plant Mol. Biol. 2002, 48, 155–171. [Google Scholar] [CrossRef]
- Weckwerth, W.; Morgenthal, K. Metabolomics: From pattern recognition to biological interpretation. Drug Discov. Today 2005, 10, 1551–1558. [Google Scholar] [CrossRef]
- De Preter, V.; Van Staeyen, G.; Esser, D.; Rutgeerts, P.; Verbeke, K. Development of a screening method to determine the pattern of fermentation metabolites in faecal samples using on-line purge-and-trap gas chromatographic–mass spectrometric analysis. J. Chromatogr. A 2009, 1216, 1476–1483. [Google Scholar] [CrossRef]
- Dudley, E.; Yousef, M.; Wang, Y.; Griffiths, W.J. Targeted metabolomics and mass spectrometry. Adv. Protein Chem. Struct. Biol. 2010, 80, 45–83. [Google Scholar] [CrossRef]
- Nicholson, J.K.; Connelly, J.; Lindon, J.C.; Holmes, E. Metabonomics: A platform for studying drug toxicity and gene function. Nat. Rev. Drug Discov. 2002, 1, 153. [Google Scholar] [CrossRef]
- Wikoff, W.R.; Anfora, A.T.; Liu, J.; Schultz, P.G.; Lesley, S.A.; Peters, E.C.; Siuzdak, G. Metabolomics analysis reveals large effects of gut microflora on mammalian blood metabolites. Proc. Natl. Acad. Sci. USA 2009, 106, 3698–3703. [Google Scholar] [CrossRef] [PubMed]
- Panopoulos, A.D.; Yanes, O.; Ruiz, S.; Kida, Y.S.; Diep, D.; Tautenhahn, R.; Herrerías, A.; Batchelder, E.M.; Plongthongkum, N.; Lutz, M.; et al. The metabolome of induced pluripotent stem cells reveals metabolic changes occurring in somatic cell reprogramming. Cell Res. 2011, 22, 168. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Gao, Y. Physiological conditions can be reflected in human urine proteome and metabolome. Expert Rev. Proteom. 2015, 12, 623–636. [Google Scholar] [CrossRef] [PubMed]
- Smith, C.A.; Want, E.J.; O’Maille, G.; Abagyan, R.; Siuzdak, G. XCMS: Processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal. Chem. 2006, 78, 779–787. [Google Scholar] [CrossRef]
- Baran, R.; Kochi, H.; Saito, N.; Suematsu, M.; Soga, T.; Nishioka, T.; Robert, M.; Tomita, M. MathDAMP: A package for differential analysis of metabolite profiles. BMC Bioinform. 2006, 7, 530. [Google Scholar] [CrossRef]
- Lommen, A. MetAlign: Interface-driven, versatile metabolomics tool for hyphenated full-scan mass spectrometry data preprocessing. Anal. Chem. 2009, 81, 3079–3086. [Google Scholar] [CrossRef]
- Katajamaa, M.; Miettinen, J.; Orešič, M. MZmine: Toolbox for processing and visualization of mass spectrometry based molecular profile data. Bioinformatics 2006, 22, 634–636. [Google Scholar] [CrossRef]
- De Preter, V.; Verbeke, K. Metabolomics as a diagnostic tool in gastroenterology. World J. Gastrointest. Pharmacol. Ther. 2013, 4, 97. [Google Scholar] [CrossRef]
- Kopka, J.; Schauer, N.; Krueger, S.; Birkemeyer, C.; Usadel, B.; Bergmüller, E.; Dörmann, P.; Weckwerth, W.; Gibon, Y.; Stitt, M.; et al. GMD@CSB.DB: The Golm Metabolome Database. Bioinformatics 2004, 21, 1635–1638. [Google Scholar] [CrossRef]
- Smith, C.A.; O’Maille, G.; Want, E.J.; Qin, C.; Trauger, S.A.; Brandon, T.R.; Custodio, D.E.; Abagyan, R.; Siuzdak, G. METLIN: A metabolite mass spectral database. Ther. Drug Monit. 2005, 27, 747–751. [Google Scholar] [CrossRef]
- Kanehisa, M.; Araki, M.; Goto, S.; Hattori, M.; Hirakawa, M.; Itoh, M.; Katayama, T.; Kawashima, S.; Okuda, S.; Tokimatsu, T. KEGG for linking genomes to life and the environment. Nucleic Acids Res. 2007, 36, D480–D484. [Google Scholar] [CrossRef] [PubMed]
- Degtyarenko, K.; De Matos, P.; Ennis, M.; Hastings, J.; Zbinden, M.; McNaught, A.; Alcántara, R.; Darsow, M.; Guedj, M.; Ashburner, M. ChEBI: A database and ontology for chemical entities of biological interest. Nucleic Acids Res. 2007, 36, D344–D350. [Google Scholar] [CrossRef] [PubMed]
- Wishart, D.S.; Tzur, D.; Knox, C.; Eisner, R.; Guo, A.C.; Young, N.; Cheng, D.; Jewell, K.; Arndt, D.; Sawhney, S.; et al. HMDB: The Human Metabolome Database. Nucleic Acids Res. 2007, 35, D521–D526. [Google Scholar] [CrossRef] [PubMed]
- Wishart, D.S.; Feunang, Y.D.; Marcu, A.; Guo, A.C.; Liang, K.; Vázquez-Fresno, R.; Sajed, T.; Johnson, D.; Li, C.; Karu, N.; et al. HMDB 4.0: The human metabolome database for 2018. Nucleic Acids Res. 2017, 46, D608–D617. [Google Scholar] [CrossRef] [PubMed]
- Ulrich, E.L.; Akutsu, H.; Doreleijers, J.F.; Harano, Y.; Ioannidis, Y.E.; Lin, J.; Livny, M.; Mading, S.; Maziuk, D.; Miller, Z. BioMagResBank. Nucleic Acids Res. 2007, 36, D402–D408. [Google Scholar] [CrossRef] [PubMed]
- Cui, Q.; Lewis, I.A.; Hegeman, A.D.; Anderson, M.E.; Li, J.; Schulte, C.F.; Westler, W.M.; Eghbalnia, H.R.; Sussman, M.R.; Markley, J.L. Metabolite identification via the Madison Metabolomics Consortium Database. Nat. Biotechnol. 2008, 26, 162–164. [Google Scholar] [CrossRef]
- Schellenberger, J.; Park, J.O.; Conrad, T.M.; Palsson, B.Ø. BiGG: A Biochemical Genetic and Genomic knowledgebase of large scale metabolic reconstructions. BMC Bioinform. 2010, 11, 213. [Google Scholar] [CrossRef]
- Horai, H.; Arita, M.; Kanaya, S.; Nihei, Y.; Ikeda, T.; Suwa, K.; Ojima, Y.; Tanaka, K.; Tanaka, S.; Aoshima, K. MassBank: A public repository for sharing mass spectral data for life sciences. J. Mass Spectrom. 2010, 45, 703–714. [Google Scholar] [CrossRef]
- Skogerson, K.; Wohlgemuth, G.; Barupal, D.K.; Fiehn, O. The volatile compound BinBase mass spectral database. BMC Bioinform. 2011, 12, 321. [Google Scholar] [CrossRef]
- May, J.C.; McLean, J.A. Advanced Multidimensional Separations in Mass Spectrometry: Navigating the Big Data Deluge. Annu. Rev. Anal. Chem. 2016, 9, 387–409. [Google Scholar] [CrossRef]
- Schmidt, R.; Etalo, D.W.; De Jager, V.; Gerards, S.; Zweers, H.; De Boer, W.; Garbeva, P. Microbial small talk: Volatiles in fungal–bacterial interactions. Front. Microbiol. 2016, 6, 1495. [Google Scholar] [CrossRef] [PubMed]
- Young, G. Fungal pathogenesis: Fungal communication gets volatile. Nat. Rev. Microbiol. 2009, 7, 6. [Google Scholar] [CrossRef]
- Baranska, A.; Mujagic, Z.; Smolinska, A.; Dallinga, J.; Jonkers, D.; Tigchelaar, E.; Dekens, J.; Zhernakova, A.; Ludwig, T.; Masclee, A. Volatile organic compounds in breath as markers for irritable bowel syndrome: A metabolomic approach. Aliment. Pharmacol. Ther. 2016, 44, 45–56. [Google Scholar] [CrossRef] [PubMed]
- Chong, J.; Xia, J. Computational approaches for integrative analysis of the metabolome and microbiome. Metabolites 2017, 7, 62. [Google Scholar] [CrossRef] [PubMed]
- Kowalski, B.R. Chemometrics: Theory and Application; ACS Publications: Washington, DC, USA, 1977. [Google Scholar]
- Tauler, R.; Parastar, H. Big (bio)chemical data mining using Chemometric methods: A need for chemists. Angew. Chem. Int. Ed. 2018. [Google Scholar] [CrossRef]
- Noecker, C.; Eng, A.; Srinivasan, S.; Theriot, C.M.; Young, V.B.; Jansson, J.K.; Fredricks, D.N.; Borenstein, E. Metabolic model-based integration of microbiome taxonomic and metabolomic profiles elucidates mechanistic links between ecological and metabolic variation. MSystems 2016, 1, e00013–e00015. [Google Scholar] [CrossRef]
- Noronha, A.; Modamio, J.; Jarosz, Y.; Guerard, E.; Sompairac, N.; Preciat, G.; Daníelsdóttir, A.D.; Krecke, M.; Merten, D.; Haraldsdóttir, H.S. The Virtual Metabolic Human database: Integrating human and gut microbiome metabolism with nutrition and disease. Nucleic Acids Res. 2018, 47, D614–D624. [Google Scholar]
- Aguiar-Pulido, V.; Huang, W.; Suarez-Ulloa, V.; Cickovski, T.; Mathee, K.; Narasimhan, G. Metagenomics, Metatranscriptomics, and Metabolomics Approaches for Microbiome Analysis:Supplementary Issue: Bioinformatics Methods and Applications for Big Metagenomics Data. Evol. Bioinform. 2016, 12s1, EBO.S36436. [Google Scholar] [CrossRef]
- Kaput, J.; Van Ommen, B.; Kremer, B.; Priami, C.; Monteiro, J.P.; Morine, M.; Pepping, F.; Diaz, Z.; Fenech, M.; He, Y. Consensus statement understanding health and malnutrition through a systems approach: The ENOUGH program for early life. Genes Nutr. 2014, 9, 378. [Google Scholar] [CrossRef][Green Version]
- Shoaie, S.; Nielsen, J. Elucidating the interactions between the human gut microbiota and its host through metabolic modeling. Front. Genet. 2014, 5, 86. [Google Scholar] [CrossRef]
- Jacobsen, U.P.; Nielsen, H.B.; Hildebrand, F.; Raes, J.; Sicheritz-Ponten, T.; Kouskoumvekaki, I.; Panagiotou, G. The chemical interactome space between the human host and the genetically defined gut metabotypes. ISME J. 2012, 7, 730. [Google Scholar] [CrossRef] [PubMed]
- McHardy, I.H.; Goudarzi, M.; Tong, M.; Ruegger, P.M.; Schwager, E.; Weger, J.R.; Graeber, T.G.; Sonnenburg, J.L.; Horvath, S.; Huttenhower, C.; et al. Integrative analysis of the microbiome and metabolome of the human intestinal mucosal surface reveals exquisite inter-relationships. Microbiome 2013, 1, 17. [Google Scholar] [CrossRef] [PubMed]
- Thaiss, C.A.; Elinav, E. The remedy within: Will the microbiome fulfill its therapeutic promise? J. Mol. Med. 2017, 95, 1021–1027. [Google Scholar] [CrossRef] [PubMed]
- Young, V.B.; Kahn, S.A.; Schmidt, T.M.; Chang, E.B. Studying the Enteric Microbiome in Inflammatory Bowel Diseases: Getting through the Growing Pains and Moving Forward. Front. Microbiol. 2011, 2, 144. [Google Scholar] [CrossRef] [PubMed]
- Uhr, G.T.; Dohnalová, L.; Thaiss, C.A. The dimension of time in host-microbiome interactions. MSystems 2019, 4, e00216-18. [Google Scholar] [CrossRef] [PubMed]
- Hacılar, H.; Nalbantoğlu, O.U.; Bakir-Güngör, B. Machine Learning Analysis of Inflammatory Bowel Disease-Associated Metagenomics Dataset. In Proceedings of the 2018 3rd International Conference on Computer Science and Engineering (UBMK), Sarajevo, Bosnia-Herzegovina, 20–23 September 2018; pp. 434–438. [Google Scholar]
- Dave, M.; Higgins, P.D.; Middha, S.; Rioux, K.P. The human gut microbiome: Current knowledge, challenges, and future directions. Transl. Res. 2012, 160, 246–257. [Google Scholar] [CrossRef] [PubMed]
- Tyanova, S.; Temu, T.; Sinitcyn, P.; Carlson, A.; Hein, M.Y.; Geiger, T.; Mann, M.; Cox, J. The Perseus computational platform for comprehensive analysis of (prote) omics data. Nat. Methods 2016, 13, 731. [Google Scholar] [CrossRef] [PubMed]
- Lin, E.; Lane, H.-Y. Machine learning and systems genomics approaches for multi-omics data. Biomark. Res. 2017, 5, 2. [Google Scholar] [CrossRef]
- Swan, A.L.; Stekel, D.J.; Hodgman, C.; Allaway, D.; Alqahtani, M.H.; Mobasheri, A.; Bacardit, J. A machine learning heuristic to identify biologically relevant and minimal biomarker panels from omics data. BMC Genom. 2015, 16, S2. [Google Scholar] [CrossRef]
- Huang, S.; Chaudhary, K.; Garmire, L.X. More is better: Recent progress in multi-omics data integration methods. Front. Genet. 2017, 8, 84. [Google Scholar] [CrossRef]
- Dubourg-Felonneau, G.; Cannings, T.; Cotter, F.; Thompson, H.; Patel, N.; Cassidy, J.W.; Clifford, H.W. A Framework for Implementing Machine Learning on Omics Data. arXiv 2018, arXiv:1811.10455. [Google Scholar]
- Saulnier, D.M.; Riehle, K.; Mistretta, T.A.; Diaz, M.A.; Mandal, D.; Raza, S.; Weidler, E.M.; Qin, X.; Coarfa, C.; Milosavljevic, A.; et al. Gastrointestinal Microbiome Signatures of Pediatric Patients With Irritable Bowel Syndrome. Gastroenterology 2011, 141, 1782–1791. [Google Scholar] [CrossRef] [PubMed]
- Christley, S.; Cockrell, C.; An, G. Computational Studies of the Intestinal Host-Microbiota Interactome. Computation 2015, 3, 2–28. [Google Scholar] [CrossRef]
- Costello, Z.; Martin, H.G. A machine learning approach to predict metabolic pathway dynamics from time-series multiomics data. NPJ Syst. Biol. Appl. 2018, 4, 19. [Google Scholar] [CrossRef] [PubMed]
- Acharjee, A.; Ament, Z.; West, J.A.; Stanley, E.; Griffin, J.L. Integration of metabolomics, lipidomics and clinical data using a machine learning method. BMC Bioinform. 2016, 17, 440. [Google Scholar] [CrossRef]
- Sung, J.; Kim, S.; Cabatbat, J.J.T.; Jang, S.; Jin, Y.-S.; Jung, G.Y.; Chia, N.; Kim, P.-J. Global metabolic interaction network of the human gut microbiota for context-specific community-scale analysis. Nat. Commun. 2017, 8, 15393. [Google Scholar] [CrossRef]
- Hsu, Y.-H.H.; Churchhouse, C.; Pers, T.H.; Mercader, J.M.; Metspalu, A.; Fischer, K.; Fortney, K.; Morgen, E.K.; Gonzalez, C.; Gonzalez, M.E.; et al. PAIRUP-MS: Pathway analysis and imputation to relate unknowns in profiles from mass spectrometry-based metabolite data. PLoS Comput. Biol. 2019, 15, e1006734. [Google Scholar] [CrossRef]
- Menon, R.; Ramanan, V.; Korolev, K.S. Interactions between species introduce spurious associations in microbiome studies. PLoS Comput. Biol. 2018, 14, e1005939. [Google Scholar] [CrossRef]
- Sankaran, K.; Holmes, S. Interactive Visualization of Hierarchically Structured Data. J. Comput. Graph. Stat. 2018, 27, 553–563. [Google Scholar] [CrossRef]
- Rahnavard, G.; Franzosa, E.A.; McIver, L.J.; Schwager, E.; Weingart, G.; Moon, Y.S.; Morgan, X.C.; Waldron, L.; Huttenhower, C. High-Sensitivity Pattern Discovery in Large Multi’omic Datasets. 2017. Available online: https://huttenhower.sph.harvard.edu/halla (accessed on 1 March 2020).
- Wang, J.; Tang, L.; Zhou, H.; Zhou, J.; Glenn, T.C.; Shen, C.-L.; Wang, J.-S. Long-term treatment with green tea polyphenols modifies the gut microbiome of female sprague-dawley rats. J. Nutr. Biochem. 2018, 56, 55–64. [Google Scholar] [CrossRef]
- Kalantar-zadeh, K.; Yao, C.K.; Berean, K.J.; Ha, N.; Ou, J.Z.; Ward, S.A.; Pillai, N.; Hill, J.; Cottrell, J.J.; Dunshea, F.R.; et al. Intestinal Gas Capsules: A Proof-of-Concept Demonstration. Gastroenterology 2016, 150, 37–39. [Google Scholar] [CrossRef] [PubMed]
- Amrane, S.; Raoult, D.; Lagier, J.-C. Metagenomics, culturomics, and the human gut microbiota. Expert Rev. Anti-Infect. Ther. 2018, 16, 373–375. [Google Scholar] [CrossRef] [PubMed]
- Bilen, M.; Dufour, J.-C.; Lagier, J.-C.; Cadoret, F.; Daoud, Z.; Dubourg, G.; Raoult, D. The contribution of culturomics to the repertoire of isolated human bacterial and archaeal species. Microbiome 2018, 6, 94. [Google Scholar] [CrossRef] [PubMed]
- Lagier, J.-C.; Hugon, P.; Khelaifia, S.; Fournier, P.-E.; La Scola, B.; Raoult, D. The Rebirth of Culture in Microbiology through the Example of Culturomics To Study Human Gut Microbiota. Clin. Microbiol. Rev. 2015, 28, 237–264. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, C.; Lachaize, C.; Janody, F.; Bellon, B.; Röder, L.; Euzenat, J.; Rechenmann, F.; Jacq, B. Grasping at molecular interactions and genetic networks in Drosophila melanogaster using FlyNets, an Internet database. Nucleic Acids Res. 1999, 27, 89–94. [Google Scholar] [CrossRef]
- Gundogdu, A.; Nalbantoglu, U. Human genome-microbiome interaction: Metagenomics frontiers for the aetiopathology of autoimmune diseases. Microb. Genom. 2017, 3, e000112. [Google Scholar] [CrossRef]
- Guven-Maiorov, E.; Tsai, C.-J.; Nussinov, R. Structural host-microbiota interaction networks. PLoS Comput. Biol. 2017, 13, e1005579. [Google Scholar] [CrossRef]
- Yuan, C.; Burns, M.; Subramanian, S.; Blekhman, R. Interaction between Host MicroRNAs and the Gut Microbiota in Colorectal Cancer. bioRxiv 2017, 192401. [Google Scholar] [CrossRef]
- Tomás-Barberán, F.A.; Espín, J.C. Effect of Food Structure and Processing on (Poly)phenol–Gut Microbiota Interactions and the Effects on Human Health. Annu. Rev. Food Sci. Technol. 2019. [Google Scholar] [CrossRef]
| Disease/Disorder | Positively Implicated Members of Microbiota (↑) | Negatively Implicated Members of Microbiota (↓) | Reference |
|---|---|---|---|
| Colorectal cancer | Fusobacterium Porphyromonas | Clostridium Bacteroides Lachnospiraceae | [55] |
| Colitis-associated colorectal cancer | Bifidobacterium E. coli | [56] | |
| Cirrhosis | Enterococcaeae Staphylococcaceae Enterobacteriaceae | Clostridiales XIV Ruminococcaceae Lachnospiraceae Veillonellaceae Porphyromonadaceae | [57] |
| Non-alcoholic fatty liver disease and steatohepatitis | Ruminococcus Dorea | Oscillospira | [58] |
| Celiac’s disease | Bacteroides vulgatus Escherichia coli | Clostridium coccoides | [59] |
| Gastric cancer | Helicobacter pylori | [60] | |
| Autism | Bacteroidetes Proteobacteria | Actinobacteria Firmicutes | [61,62,63] |
| Parkinson’s Disease | Enterobacteriaceae | Prevotellaceae | [64] |
| Type 2 diabetes | Betaproteobacteria | Firmicutes Clostridia | [65,66] |
| IBD – Crohn’s Disease | Bacteroides ovatus Bacteroides vulgatus | Bacteroides uniformis | [67] |
| IBD – Ulcerative colitis | Gammaproteobacteria Deltaproteobacteria Actinobacteria Proteobacteria | Firmicutes | [68,69,70] |
| Method | Tool | Description | Reference |
|---|---|---|---|
| Assembly | DIME | Combines the DIvide, conquer, and MErge strategies | [147] |
| Genovo | Generative probabilistic model of reads | [148] | |
| Khmer | Probabilistic de Bruijn graphs | [149] | |
| MAP | OLC (Overlap/Layout/Consensus) strategy for longer reads | [150] | |
| Meta-IDBA | De Bruijn graph approach | [151] | |
| metAMOS | A Modular Open-Source Assembler component for metagenomes | [152] | |
| MetaVelvet | De Bruijn graph approach | [153] | |
| MOCAT | a metagenomics assembly and gene prediction toolkit | [154] | |
| SOAPdenovo | Single-genome assembler commonly tuned for metagenomes | [155] | |
| MetaORFA | Gene-targeted assembly approach | [156] | |
| MetaPAR | Metagenomic sequence assembly via iterative reclassification | [157] | |
| XGenovo | An extended Genovo assembler by incorporating paired-end information | [158] | |
| Taxonomic profiling | Amphora | Automated pipeline for Phylogenomic Analysis | [159,160,161] |
| CARMA3 | Taxonomic classification of metagenomic shotgun sequences | [162,163] | |
| ClaMS | Classifier for Metagenomic Sequences | [164] | |
| CLARK | Fast and accurate classification of metagenomic and genomic sequences using discriminative k-mers | [165] | |
| DiScRIBinATE | Distance Score Ratio for Improved Binning and Taxonomic Estimation | [166] | |
| FOCUS | An agile composition-based approach using non-negative least squares | [167] | |
| INDUS | Composition-based approach for rapid and accurate taxonomic classification of metagenomic sequences | [168] | |
| MARTA | Suite of Java-based tools for assigning taxonomic status to DNA sequences | [169] | |
| MetaCluster | Binning algorithm for high-throughput sequencing reads | [170] | |
| MetaPhlAn | Profiles the composition of microbial communities from metagenomic shotgun sequencing data | [171,172] | |
| MetaPhyler | Taxonomic classifier for metagenomic shotgun reads using phylogenetic marker reference genes | [173] | |
| MOCAT2 | A metagenomic assembly, annotation and profiling framework | [174] | |
| MTR | Taxonomic annotation of short metagenomic reads using clustering at multiple taxonomic ranks | [175] | |
| NBC | Naive Bayes Classification tool for taxonomic assignment | [176] | |
| PaPaRa | Aligning short reads to reference alignments and trees | [177] | |
| PhyloPythia | Accurate phylogenetic classification of variable-length DNA fragments | [164] | |
| PhyloSift | Phylogenetic analysis of metagenomic samples | [178] | |
| Phymm | Classification system designed for metagenomics experiments that assigns taxonomic labels to short DNA Reads | [179] | |
| RAIphy | Phylogenetic classification of metagenomics samples using iterative refinement of relative abundance index Profiles | [180] | |
| RITA | Classifying short genomic fragments from novel lineages using composition and homology | [181] | |
| SOrt-ITEMS | Sequence orthology-based approach for improved taxonomic estimation of metagenomic sequences | [182] | |
| SPHINX | Algorithm for taxonomic binning of metagenomic sequences | [183] | |
| TACOA | Taxonomic classification of environmental genomic fragments using a kernelized nearest neighbour approach | [184] | |
| Treephyler | Fast taxonomic profiling of metagenomes | [185] | |
| Functional profiling | HUMAnN | Determines the presence/absence and abundance of microbial pathways in meta-omic data | [186] |
| metaSHARK | web platform for interactive exploration of metabolic networks | [187] | |
| MOCAT2 | A metagenomic assembly, annotation and profiling framework | [174] | |
| PRMT | Predicted Relative Metabolomic Turnover: determining metabolic turnover from a coastal marine metagenomic dataset | [188] | |
| RAMMCAP | Rapid analysis of Multiple Metagenomes with Clustering and Annotation Pipeline | [189] | |
| Interaction networks | SparCC | Estimates correlation values from compositional data for network inference | [190] |
| CCREPE | Predicts microbial relationships within and between microbial habitats for network inference | [191] | |
| Single-cell sequencing | IDBA-UD | De Bruijn graph approach for uneven sequencing depths | [192] |
| SmashCell | Software framework for the analysis of single-cell amplified genome sequences | [193] | |
| Simulators | GenSIM | Error-model based simulator of next-generation sequencing data | [194] |
| Metasim | A sequencing simulator for genomics and metagenomics | [195] | |
| Statistical tests | Metastats | Statistical analysis software for comparing metagenomic samples | [196] |
| LefSe | Nonparametric test for biomarker discovery in proportional microbial community data | [197] | |
| ShotgunFunctionalizeR | A statistical test based on a Poisson model for metagenomic functional comparisons | [198] | |
| SourceTracker | A Bayesian approach to identify and quantify contaminants in a given community | [199] | |
| General toolkit | CAMERA | Dashboard for environmental metagenomic and genomic data, metadata, and comparative analysis tools | [200] |
| GenBoree | A web-based platform for multi-omic research and data analysis using the latest bioinformatics tools | [201] | |
| GraPhlAn | Compact graphical representation of phylogenetic data and metadata | [202] | |
| IMG/M | Integrated metagenome data management and comparative analysis system | [100] | |
| MEGAN | Software for metagenomic, metatranscriptomic, metaproteomic, and rRNA analysis | [203] | |
| METAREP | Online storage and analysis environment for meta-omic data | [204] | |
| MG-RAST | Storage, quality control, annotation and comparison of meta-omic samples | [205] | |
| Mothur | An open-source software for microbial ecology community analysis | [206] | |
| QIIME | An open-source bioinformatics pipeline for performing microbiome analysis from raw DNA sequencing data | [207] | |
| SmashCommunity | Stand-alone annotation and analysis pipeline suitable for meta-omic data | [208] | |
| STAMP | Comparative meta-omics software package | [209] | |
| SnoWMan | Pipeline for analysis of microbiome data | [210] | |
| VAMPS | Visualization and analysis of microbial population structure | [211] |
| Metabolite Class | Metabolites | Related Bacteria | Biological Functions | Disease/Disorder | Reference |
|---|---|---|---|---|---|
| Short-chain fatty acids |
|
|
|
| [40,232,233] |
| Phenolic, Benzoyl and phenyl derivatives |
|
|
|
| [230,234,235,236,237,238,239,240] |
| Bile salts |
|
|
|
| [241,242,243] |
| Choline metabolites |
|
|
|
| [244,245] |
| Indole derivatives |
|
|
|
| [246,247] |
| Vitamins |
|
|
| [248,249,250,251] | |
| Polyamines |
|
|
| [252] | |
| Lipids |
|
|
| [253] | |
| Others |
|
|
| [237,254] |
| Database | Web Address/URL | Available Since/Reference |
|---|---|---|
| Global Metabolome Database (GMD) | http://gmd.mpimp-golm.mpg.de/ | 2004, Kopka, et al. [268] |
| METLIN | https://metlin.scripps.edu/ | 2005, Smith, et al. [269] |
| Kyoto Encyclopedia of Genes and Genomes (KEGG) | http://www.genome.jp/kegg/ | 1995, Kanehisa, et al. [270] |
| Chemicals Entities of Biological Interest (ChEBI) | http://www.ebi.ac.uk/chebi/ | 2004, Degtyarenko, et al. [271] |
| Human Metabolome Database (HMDB) | http://www.hmdb.ca/ | 2007, Wishart et al. [272,273] |
| Biological Magnetic Resonance Data Bank (BMRB) | http://www.bmrb.wisc.edu/ | 2007, Ulrich, et al. [274] |
| Madison Metabolomics Consortium (MMC) Database | http://mmcd.nmrfam.wisc.edu/ | 2008, Cui, et al. [275] |
| BiGG (a knowledgebase of Biochemically, Genetically and Genomically structured genome-scale metabolic network reconstructions) | http://bigg.ucsd.edu/ | 2010, Schellenberger, et al. [276] |
| MassBank | http://www.massbank.jp/ | 2010, Horai, et al. [277] |
| SetupX and BinBase | https://fiehnlab.ucdavis.edu/ | 2011, Skogerson, et al. [278] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shah, R.M.; McKenzie, E.J.; Rosin, M.T.; Jadhav, S.R.; Gondalia, S.V.; Rosendale, D.; Beale, D.J. An Integrated Multi-Disciplinary Perspective for Addressing Challenges of the Human Gut Microbiome. Metabolites 2020, 10, 94. https://doi.org/10.3390/metabo10030094
Shah RM, McKenzie EJ, Rosin MT, Jadhav SR, Gondalia SV, Rosendale D, Beale DJ. An Integrated Multi-Disciplinary Perspective for Addressing Challenges of the Human Gut Microbiome. Metabolites. 2020; 10(3):94. https://doi.org/10.3390/metabo10030094
Chicago/Turabian StyleShah, Rohan M., Elizabeth J. McKenzie, Magda T. Rosin, Snehal R. Jadhav, Shakuntla V. Gondalia, Douglas Rosendale, and David J. Beale. 2020. "An Integrated Multi-Disciplinary Perspective for Addressing Challenges of the Human Gut Microbiome" Metabolites 10, no. 3: 94. https://doi.org/10.3390/metabo10030094
APA StyleShah, R. M., McKenzie, E. J., Rosin, M. T., Jadhav, S. R., Gondalia, S. V., Rosendale, D., & Beale, D. J. (2020). An Integrated Multi-Disciplinary Perspective for Addressing Challenges of the Human Gut Microbiome. Metabolites, 10(3), 94. https://doi.org/10.3390/metabo10030094








