Abstract
Zingiberaceae is commonly known as the ginger family and has been extensively studied in the last decades for its pharmacological purposes. The study of ginger includes microorganisms known as endophytes, which raise interest for the research community because they can produce a wide range of secondary metabolites. This review discusses the secondary metabolites of endophytes from the Zingiberaceae family and their pharmacological activities. We detail the group of secondary metabolites, updated for its absolute structures, source and part origins, and, especially, pharmacological divided properties. Zingiberaceae endophytes have 106 volatile compounds and 52 isolated constituents, including 17 polyketides, five nonribosomal peptides, five aromatic compounds, three alkaloids, and 21 terpene-alkaloids. They have antimicrobial, anticancer, antioxidant, and anti-inflammatory activities. Secondary metabolites from plant endophytes of the Zingiberaceae family have the potential to be therapeutic drugs in the future. Research on endophytic bacteria or fungi has been little performed. Therefore, this study supports a new drug discovery from Zingiberaceae endophytes and compares them for future drug development.
1. Introduction
Zingiberaceae is commonly known as the ginger family. It is a flowering plant of more than 1300 species divided into 52 genera. The plant is distributed throughout tropical Africa, Asia, and America [1,2,3,4]. Some plant genera of this family are beneficial for use in medicinal products, dyes, condiments, and spices [5,6,7,8]. Over the last three decades, Zingiberaceae has been studied extensively for substantial pharmacological and clinical investigations [9,10,11]. Interestingly, the phytochemical study of Zingiberaceae continues to explore their endophytes [12,13]. Endophytes are living microorganisms inside internal plant tissue and they are a beneficial symbiont to their host. These microorganisms have emerged as potential pharmacological metabolites producers [14,15,16,17].
In the last 16 years, endophytes from Zingiberaceae have been reported as producing several metabolites. There are volatile compounds, polyketides, peptides, aromatics, alkaloids, and hybrid terpene alkaloids, and they also have antimicrobial, anticancer, antioxidant, and anti-inflammatory properties [18,19,20,21,22,23,24,25,26,27].
The study of Zingiberaceae endophytes so far includes (1) species identification of endophytes microorganisms using molecular and genomic tools, (2) pharmacological potential screening, and (3) compound isolation of the natural product chemistry field area. A comprehensive review of these studies has not been conducted.
For this reason, we discuss the secondary metabolites of Zingiberaceae-associated endophytes with special emphasis on pharmacological purposes. This review highlights the chemical content of different strains classified by their biosynthesis chemical origins and their same activities. Furthermore, we add data on their parts, origin, function, biological roles, and absolute structure updates. All databases were concluded, including isolated secondary metabolites, species name, their host, absolute structures, volatile compounds, and their pharmacological activity from Scopus and SciFinder search engines. The keywords are “endophytes”, “Zingiberaceae genus member”, and “name of compound” collected from May 1975 to August 2022 [18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56]. This review might be helpful in future drug development, especially from Zingiberaceae endophytes.
2. Botany
Zingiberaceae is a large family, usually classified into four subfamilies: Hedychieae (leaves parallel to the rhizome, staminodes laterally petaloid, not fused with the labellum); Zingibereae (out of the anthers and covered by elongated anthers); Alpinieae (leaves perpendicular to the rhizome, lateral staminode absent or small and fused to the labellum); and Globbeae (elongated and curved filaments, l-locular gynoecium) [56,57,58,59]. Members of this family have distribution in tropical Asia, especially Indo-Malaysia. This family is an economic asset, an essential source of spice plants, such as Curcuma domestica (turmeric), Elettaria cardamomum (cardamom), and Zingiber officinale (ginger). Some species are grown as cultivated ornamental plants, for example, Alpinia and Hedychium [60,61,62].
Zingiberaceae grows from zero to more than 2000 m above sea level. It generally grows in areas with high rainfall and in humid places. Several other species are found in secondary forests, open forests, riverbanks, and swamps, and sometimes grow in open areas with full sun. Some species of Etlingera grow in secondary forests or open forest sites [63,64,65,66].
Zingiberaceae consists of several parts: rhizome, leaves, flowers, and fruit. These parts are used for several purposes by local communities. For example, ginger has been widely used over time because it has significant economic potential, such as traditional medicines for herbs, spices, cooking spices, hair toners, beverage ingredients, vegetables, and food seasonings. Others, such as Etlingera elatior, is used to make chili sauce or add seasoning to grilled rice. More than 60 species of Zingiberaceae are used from both cultivated and wild species from the forest [66,67,68,69,70].
3. Natural Product Diversity
Based on literature collected from May 1975 to August 2022, a total of 52 phytochemicals have been isolated from endophytic microorganisms living inside the Zingiberaceae family, including 17 polyketides, five nonribosomal peptides, five aromatics, three alkaloids, and 21 terpene-alkaloids. The isolated and identified phytochemicals are summarized in Table 1. Furthermore, the endophytic microorganism isolated from the Zingiberaceae family is rich in essential oil and volatile constituent. One hundred six volatile compounds, including 42% terpenes, 20% aromatic compounds, 12% organic acids, 5% esters, and 21 others, have been detected using gas chromatography–mass spectroscopy analysis. The volatile constituents are summarized in Table 2.

Table 1.
Compounds isolated from Zingiberaceae endophytes.

Table 2.
Volatile constituents from Zingiberaceae endophytes.
3.1. Volatile Constituents
Volatile compounds are often used in many pharmaceuticals and for other products such as soaps, cosmetics, toiletry products, and perfumes [71,72,73,74,75,76]. Volatile oils are extracted from some plant organs, including flowers, bark, leaves, roots, seeds, and others [77,78,79,80]. Interestingly, a microorganism that lives inside a plant, known as an endophyte, also produces volatile compounds [81,82].
Some reported Zingiberaceae endophytes produce volatile constituents, including terpene, steroid, alkaloid, aromatic, flavonoid, and others. There are Arthrinium sp. from Z. cassumunar, T. harzianum from C. longa, B. specifera from Z. nimmonii, A. flavus from K. rotunda, etc. [12,30,31,32,33,34]. All the reported volatile compounds from the Zingiberaceae endophyte are summarized in Table 2.
Pharmacological activities of some endophyte genera from Z. officinale show antibacterial activities against S. aureus B. subtilis S. typhimurium identified as Fusarium oxysporum and Fungal sp., GFV1. Some major compounds from fungal crude extract using GC-MS analysis includes tyrosol, benzene acetic acid, ergone, dehydromevalonic lactone, N-aminopyrrolidine, and many bioactive fatty acids and their derivatives, which include linoleic acid, oleic acid, myristic acid, hexadecenoic acid, palmitic acid methyl ester, and methyl linoleate [12].
Anisha and her co-workers [34] performed research on endophyte genera from Z. officinale and found danthron, an anthraquinone derivate from Paraconiothyrium sp., which has broad-spectrum antimicrobial activity. The PCR analysis of this endophyte showed the existence of a non-reducing polyketide synthase gene and was responsible for synthesizing anthraquinone.
The ethyl extract from Aspergillus terreus and B. specifera has shown antibacterial activity against six pathogenic bacterial strains viz., B. subtilis (MTCC 121), Staphylococcus aureus (MTCC 7443), Pseudomonas aeruginosa (MTCC 7093), Escherichia coli (MTCC 729), Enterobacter aerogenes (MTCC 111), and Klebsiella pneumoniae (MTCC 661), with minimum inhibitory concentrations (MIC) of 0.04–0.14 mg/mL. Seven major compounds with antibacterial activity were detected with GC–MS: (1) bicyclo[3.2.0]heptan-2-one, 6-hydroxy-5-methyl-6-vinyl; (2) adipic acid divinyl ester; (3) 1,4-naphthoquinone, 6-acetyl-2,5-dihydroxy; (4) decanedioic acid, 3,7-dimethyl ester; (5) (Z)-4-hexenoic acid 2-acetyl- 2-methyl-ethyl ester, and (6) butanoic acid 2-acetyl-3-methyl-methyl ester. These compounds are volatile esters and phenolic and adipic acid [32].
The Arthrinium sp. MFLUCC16-1053, an endophyte from Z. cassumunar, has shown antibacterial activity against S. aureus and E. coli with MIC 31.25 and 7.81 µg/mL, respectively. This fungus also has antioxidant activity with DPPH, scavenging at an IC50 value of 28.47 µg/mL. The major volatile compound was tested using gas chromatography–mass spectrometry and analysis revealed β-cyclocitral, 3E-cembrene A, laurenan-2-one, sclareol, 2Z,6E-farnesol, cembrene, β-isocomene, and γ-curcumene [30]. The volatile constituent from the Zingiberaceae endophytes has shown potential uses as an antibacterial and antioxidant. This information can be helpful for further development in the future in the field of drug discovery.
3.2. Polyketides
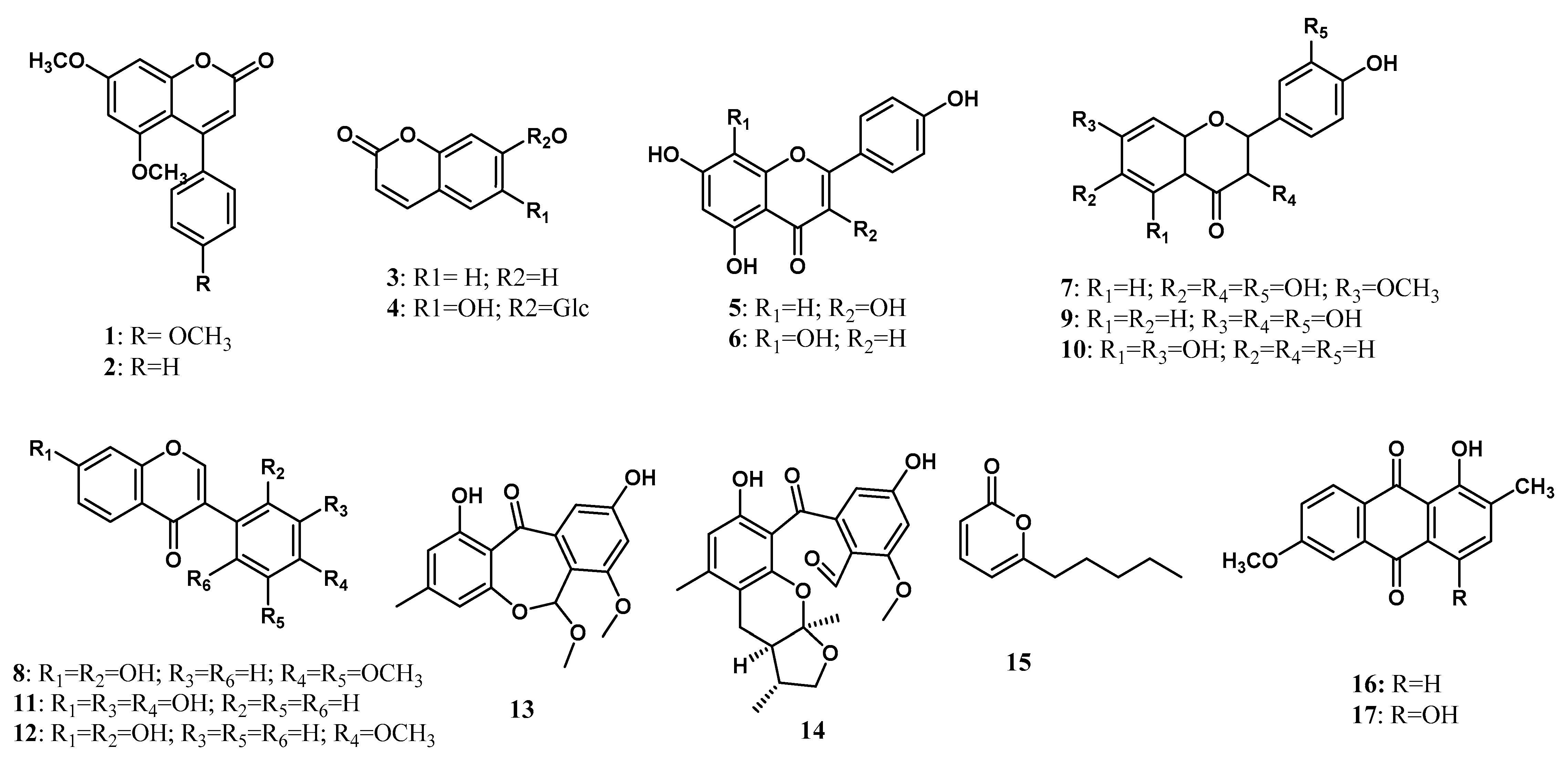
A total of 17 polyketides have been isolated from endophytic microorganisms living inside the Zingiberaceae plant. Polyketides are a large family of natural products that are known to exhibit a high degree of structural diversity with fascinating activities. Plants, bacteria, and fungi can produce polyketides through polyketide synthases enzymes [83,84,85,86]. Polyketides of fungal origin have diverse structures, ranging from antibiotics to toxins [87], which are widely used in pharmacological aspects. Taechowisan et al. [18] isolated two coumarins from the ethyl acetate extract of Streptomyces aureofaciens living inside the root tissue of Z. officinale, namely 5,7-Dimethoxy-4-p-methoxylphenylcoumarin (1) and 5,7-Dimethoxy-4-phenylcoumarin (2). Three years later, Taechowisan et al. [19] isolated another two coumarin, umbelliferone (3) and chicorii (4), together with two flavonoids, kaempferol (5) and isoscutellarin (6), from endophytic Streptomyces sp. Tc052 on the roots of A. galanga. Afterward, Taechowisan et al. [28] isolated two new flavonoids, 7-methoxy-3, 3′,4′,6-tetrahydroxyflavone (7) and 2′,7-dihydroxy-4′,5′-dimethoxyisoflavone (8) along with four known flavonoids, fisetin (9), naringenin (10), 3′-hydroxydaidzein (11), and xenognosin B (12) from endophytic Streptomyces sp. BT01 living inside the root tissue of B. rotunda (L.). These new compound structures were characterized using infrared, mass spectra, NMR spectroscopic data, and a polarimeter. In addition, the absolute configuration of 10 has been determined using VCD and DFT calculations [35]. Flavonoids were mentioned to prevent injury caused by free radicals and direct scavenging of free radicals [88].
Hammerschidt et al. [24] isolated two new polyketides from the endophytic fungus Xylaria sp. (healthy leaves of C. xanthorrhiza), that are rugosin J (13) and xylarugosin (14). These two new compounds were characterized using NMR data and mass spectral analysis, and the ECD spectrum confirmed the absolute structures. Five years later, Suebrasri and his co-workers [23] isolated 6-n-pentyl-2H-pyran-2-one (15) for the first time from Trichoderma erinaceum ST-KKU2 living inside Z. officinale. More recently, Taechowisan et al. [22] isolated 1-hydroxy-2-methyl-6-methoxyanthraquinone (16) and 6-methoxy-2-methylquinizarin (17) from endophytic Streptomyces sp. W08 from the pseudostem tissue of Amomum krevanh. The chemical structures of polyketides isolated from the endophytic microorganism of the Zingiberaceae family are shown in Figure 1.
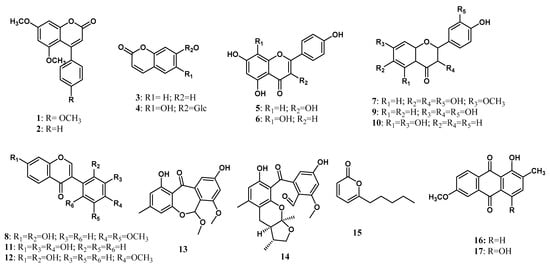
Figure 1.
Polyketides from Zingiberaceae endophytes.
3.3. Nonribosomal Peptides
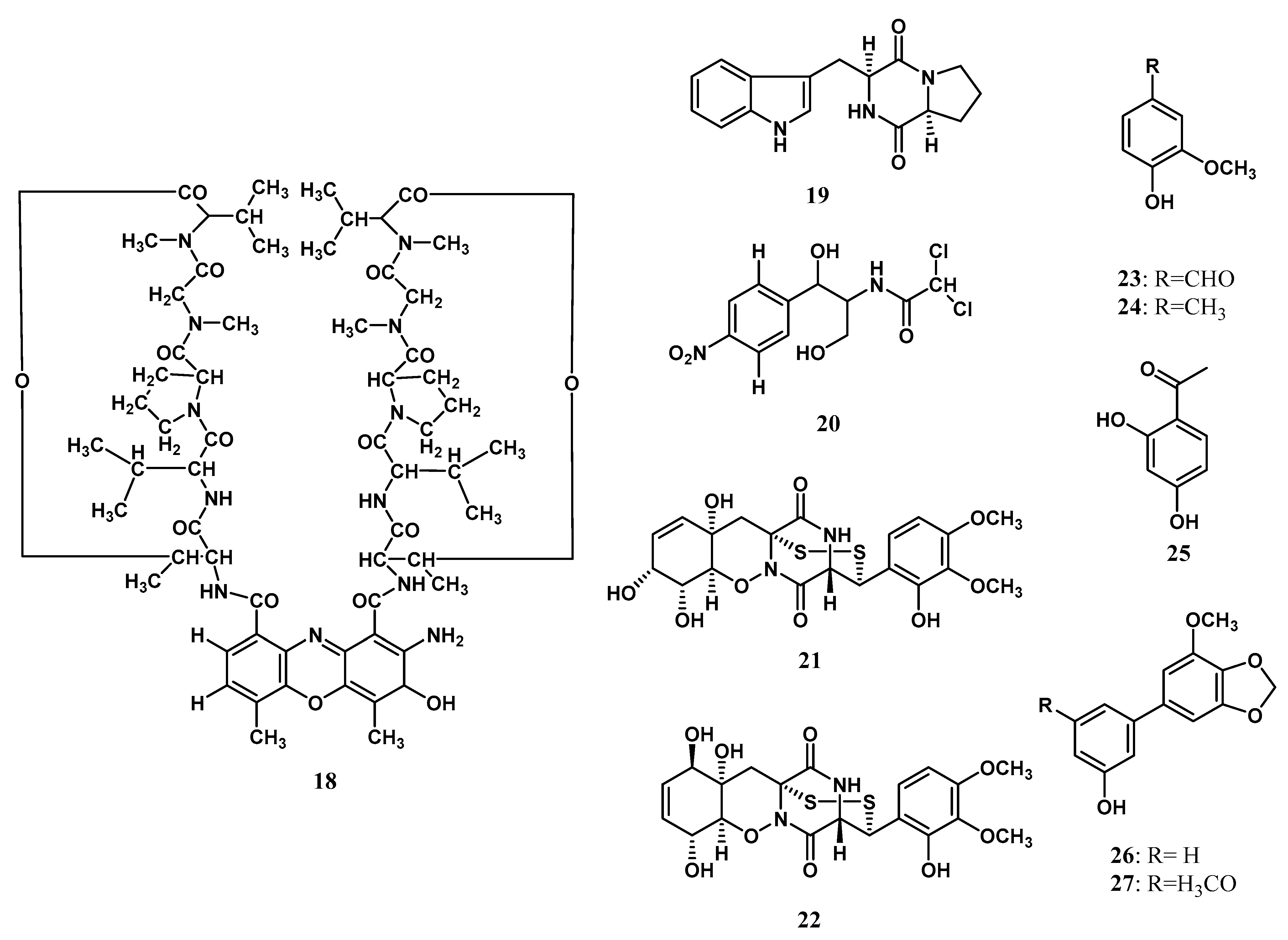
Ribosomes are molecular tools that translate mRNA into protein. the independent ribosomal tools of amide bond formation are called nonribosomal peptide synthesis [87,89,90,91]. The nonribosomal peptide consists of amino acid proteinogenic or non-proteinogenic linked with a peptide bond [92]. Each domain of NRPs provides a specific function, such as recognizing amino acids, activation with amino acids, bonding in between, and releasing the peptide [93]. The nonribosomal peptide was first isolated from the endophytic microorganism Zingiberaceae family by Taechowisan et al. [19]. Actinomycin D (18) was isolated from Streptomyces sp. Tc022 from the plant A. galanga. Over the next ten years, Alshaibani [25] isolated cyclic peptides that are brevianamide F (19) and 2,2-dichloro-N-[(1r,2r)-2-hydroxy-1-(hydroxymethyl)-2-(4-nitrophenyl) ethyl]-acetamide (20) from Streptomyces omiyaensis NBRC 11449T living inside Z. spectabile. Harwoko et al. [26] isolated two dithiodiketopiperazine derivatives from endophytic fungi T. harzianum associated with medicinal plants Z. officinale, namely pretrichodermamide G (21) and pretrichodermamide A (22). The absolute structure of 21 was determined to be the same as pretrichodermamide D, according to the common biosynthetic origin and its optical rotation value [26]. Meanwhile, the absolute stereostructure of 22 was previously determined using X-ray crystallographic analysis [36]. Compound 18 was first isolated and characterized by mass spectra analysis, 1D, and 2D NMR data. The chemical structures of nonribosomal peptides isolated from the endophytic microorganism Zingiberaceae family are shown in Figure 2.
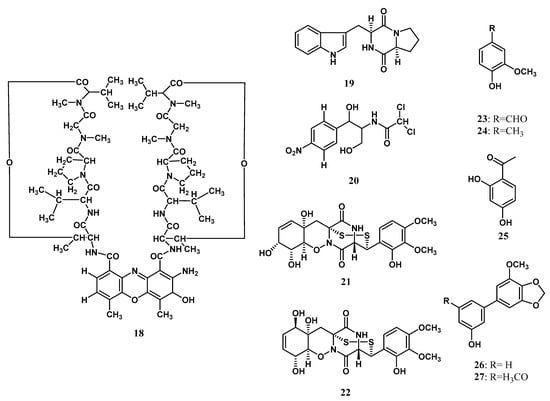
Figure 2.
Nonribosomal peptides and aromatic compounds from Zingiberaceae endophytes.
3.4. Aromatic Compounds
Aromatic compounds are found as essential oils that are volatile at room temperature [94]. In plants, aromatics are responsible for altering the odor and flavoring [95], attracting pollinators and seed dispersers and providing defense against pathogens [96,97]. Aromatic fragrance compounds are of great commercial interest in research, health, food, cosmetic, and health industries [96]. The first aromatic derivatives isolated from endophytic microorganisms living inside Zingibereacae were vanillin (23) and 3-methoxy-4-hydroxytoluene (24) by Taechowisan et al. [18] from Streptomyces aureofaciens SMUAc130 of the roots tissue of Z. officinale Rosc. The research followed the finding of resacetophenone (25) that was isolated by Hammerschmidt et al. [24] from Xylaria sp. living inside C. xanthorrhiza. Two years later, Taechowisan et al. [21] isolated 3′-hydroxy-5-methoxy-3,4-methylenedioxybiphenyl (26) and 3′-hydroxy-5,5′-dimethoxy-3,4-methylenedioxybiphenyl (27) from Streptomyces sp. BO-07 associated with the plant B. rotunda (L.). The structure of aromatic compounds is shown in Figure 2.
3.5. Alkaloids
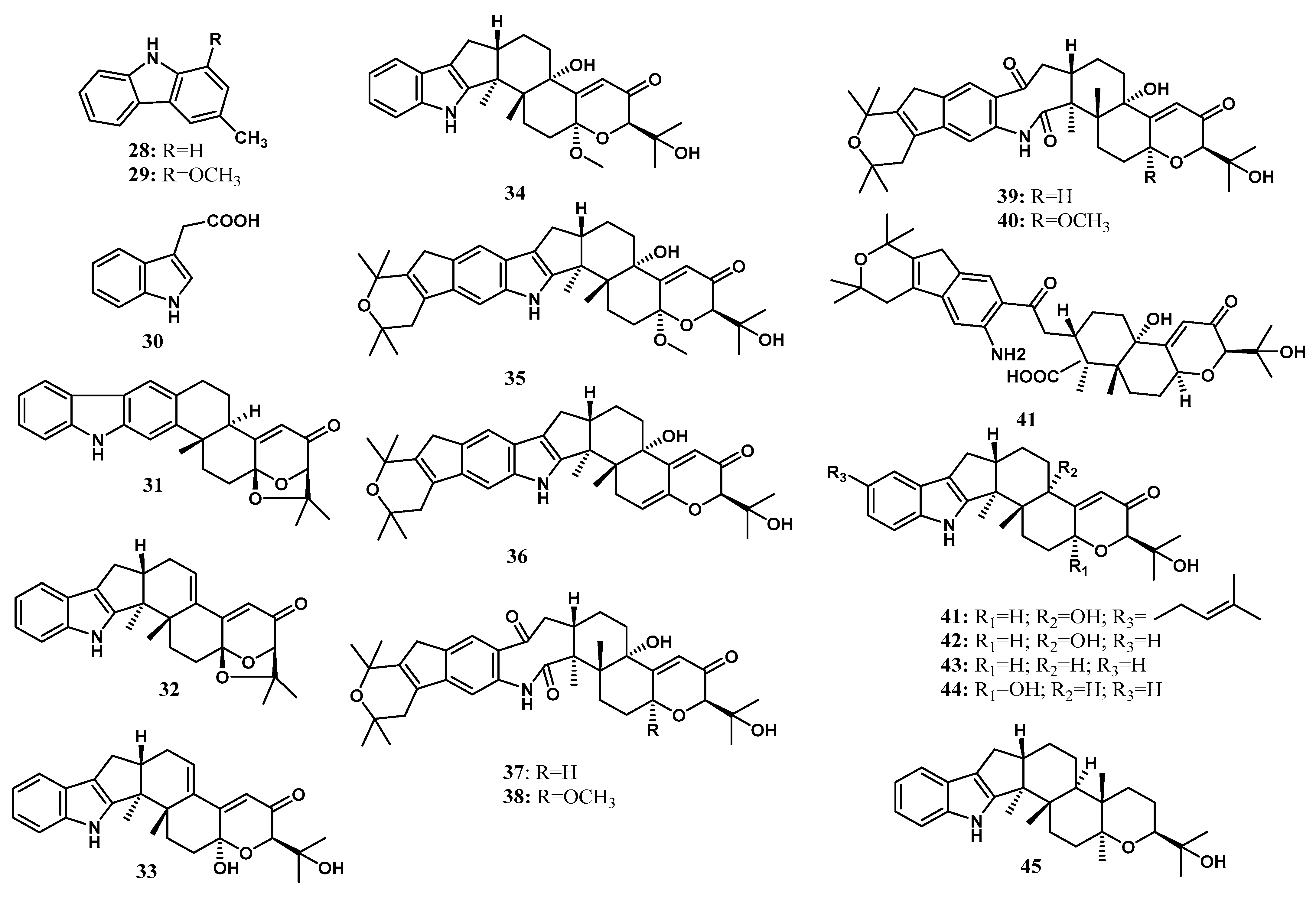
Alkaloids are mainly characterized by nitrogen atoms on the structure [98,99,100,101]. Alkaloids are familiar additive drugs to pharmacological relevance. The commercial value of alkaloids is primarily as medicines, flavorings, and poisons [102]. Alkaloids are produced by various organisms, including plants, animals, bacteria, and fungi. They are responsible as protective agents for biotic and abiotic stress [98]. Two alkaloids were isolated by Taechowisan et al. [29] from the endophytic Streptomyces sp. LJK109 from the roots of A. galanga (L.) Wild, which are 3-methylcarbazole (28) and 1-methoxy-3-methylcarbazole (29). Indole acetic acid (30) was characterized to be produced in various endophytic microorganisms, including B. subtilis CL1, Bacillus sp. CL3, Burkholderia thailandensis CL4, Agrobacterium tumefaciens CL5, Klebsiella sp. CL6, Bacillus cereus CL7, Pseudomonas putida CL9, Pseudomonas fluorescens CLI2, and Azotobacter chroococcum CL13; living inside the medicinal plant C. longa L. [37], Paenibacillus favisporus, and Paenibacillus sp. associated with the rhizome of C. longa L. [38], Pseudomonas sp.; living inside Z. officinale [39], Pseudomonas, Pantoea agglomerans, Aeromonas, Serratia, Enterobacter asburiae, and Rhizobium inside the roots, stem, tubers, and leaves of Z. officinale Roscoe [40], Ochrobactrum, Agrobacterium, Acinetobacter, Stenotrophomonas, Serratia and Bacillus associated inside Z. officinale Roscoe [41], Bacillus cereus (ECL1), Bacillus thuringiensis (ECL2), Bacillus sp. (ECL3), Bacillus pumilis (ECL4), Pseudomonas putida (ECL5), and Clavibacter michiganensis (ECL6); living inside C. longa L. [37], T. harzianum, T. asperellum, T. atroviride; and associated with the plant C. longa L. [31], A. flavus inside Alpinia sp. [42]. The structure of the isolated alkaloids is shown in Figure 3.
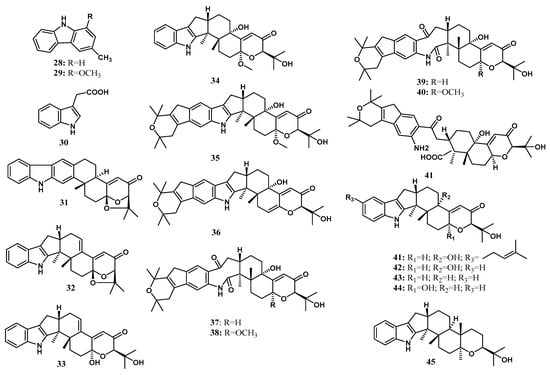
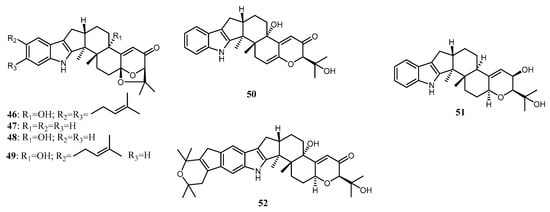
Figure 3.
Alkaloid and indole diterpenoids from Zingiberaceae endophytes.
3.6. Indole Diterpenoids
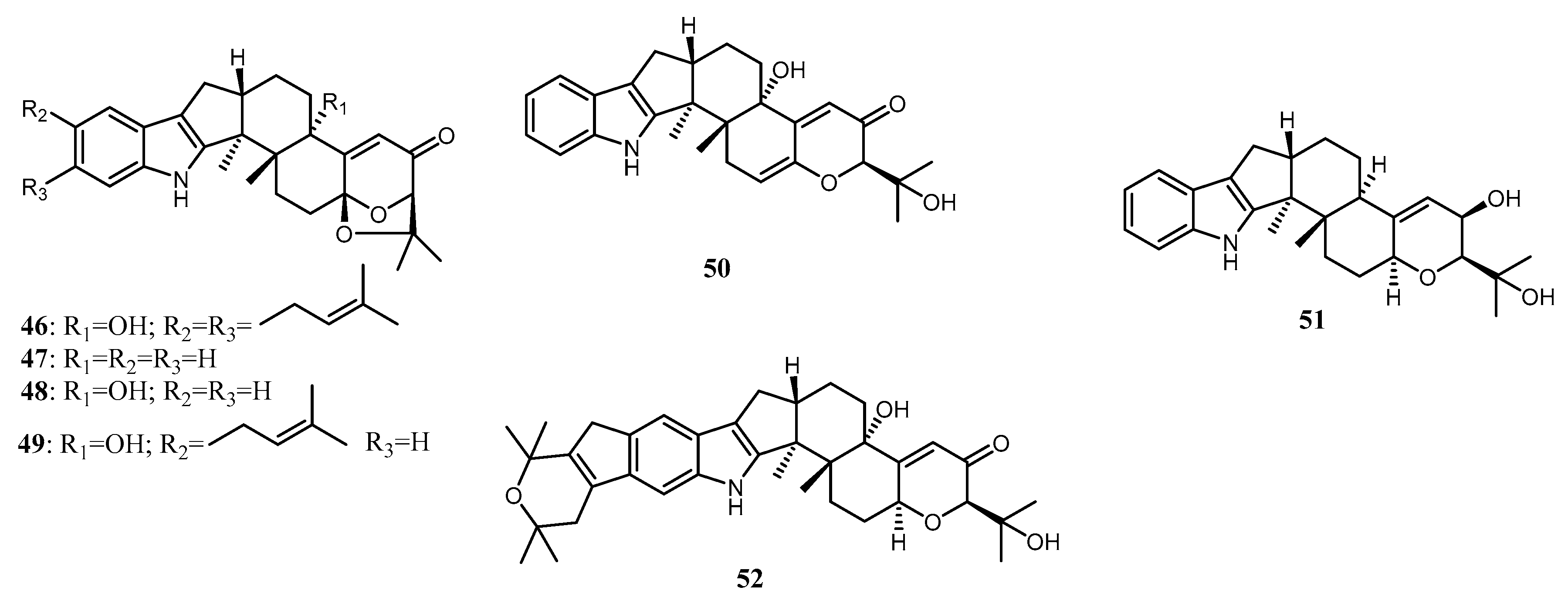
Indole terpenoids are fungal secondary metabolites with diverse structures and a broad range of biological activities [40,103,104]. Indole diterpenoids, a subclass of the indole terpenoids, have recently caught interest due to their promising biological properties [105]. Arianti and her co-workers [23] isolated nine indole diterpenoids along with 13 known congeners from the endophytic fungi Penicillium sp ZO-R1-1 associated with the roots of the medicinal plant Z. officinale. Nine new indole diterpenoids, namely shearilicine (31), paspalinine-13-ene (32), 7-hydroxypacilline-13-ene (33), 7-methoxypaxilline (34), shearinine N (35), shearinine O (36), shearinine P (37), 7-methoxyshearinine P (38), and shearinine Q (39), along with known indole diterpenoids, including emindole SB (40), 21-isopentenylpaxilline (41), paxilline (42), dehydroxypaxilline (43), 7-hydroxy-13-dehydroxypaxilline (44), paspaline (45), shearinine F (46), paspalicine (47), paspalinine (48), paspalitrem A (49), 6,7-dehydropaxilline (50), 10β-hydroxy-13-desoxypaxilline (51), and pyrapaxilline (52), were characterized and identified using mass spectra analysis and 1D and 2D NMR data. Meanwhile, the absolute configuration of the new natural products was determined using the TDDFT-ECD approach and confirmed by single-crystal X-ray determination through anomalous dispersion. The total synthesis of 41 has confirmed its structural assignment and absolute stereochemistry [43]. The absolute configuration of 42 was first reported by Springer et al. [44], using X-ray crystallography analysis. Based on that, 43 was determined to have the same absolute configuration as 42 [45]. Recently, the absolute configuration of 44 was determined by ECD calculation [46]. In addition, the absolute structures of 45 and 46 [47], as well as 50 [48], have been determined using comprehensive spectroscopic data analysis and circular dichroism (CD) calculation. The structure of indole diterpenoids isolated from Penicillium sp. ZO-R1-11 is shown in Figure 3.
4. Pharmacological Activities
4.1. Antimicrobial Activity
Compounds 1, 2, 23, and 24 were evaluated for their antifungal activity by 14 using a paper-disc assay method against C. musae and F. oxysporum. Compounds 1 and 2 give the same percentage for inhibiting the growth of C. musae and F. oxysporum, namely 66% and 72%, respectively. The MIC values of 1 and 2 for inhibition of C. musae were 120 and 150 µg/mL, respectively. Meanwhile, 23 and 24 showed weak inhibition, i.e., 32% and 35% for 23 and 26% and 30% for 24 [18]. Ten years later, El-Gendy & El-Bondkly [49] conducted the antimycotic activity of compound 1 against several dermatophytes and other pathogenic fungi, namely T. rubrum, T. mentagrophytes, M. gypseum, E. floccosum, A. niger, A. fumigatus, F. oxysporum, C. albicans, and C. humicolus. The results give the MIC values of 7.5, 90, 100, 50, 20, 10, 22, 15, and 10 µg/mL, respectively and give the MFC values of 100, 90, 150, 66, 50, 35, 49, 20, and 32 µg/mL, respectively [49].
Compound 4 was evaluated for its antibacterial effectiveness against S. aureus, E. coli, and P. aeruginosea, and antifungal activity against C. albicans with the observation by measuring the inhibited zones (IZ) in mm with DMF as a control. The results showed the IZ values of 4 are 14, 20, 14, and 20 mm, respectively, compared with the IZ values of DMF being 14, 18, 12, and 17, respectively. However, the MIC and MBC values of 4 against S. aureus and E. coli showed an inactive result; meanwhile, the activity of 4 against P. aeroginosea gives the same MIC and MBC values of 62.5 µg/mL, respectively. In addition, the antifungal activity of 4 against C. albicans gives a MIC value of 62.5 µg/mL, but a weak MBC value of >500 µg/mL [50]. In 2008, Taechowisan et al. [20] evaluated the antibacterial and antifungal activities of compounds 3, 4, 5, and 6 against S. aureus, E. coli, P. aeruginosa, B. subtilis, C. albicans, and C. musae. The MIC values lower or equal to 128 µg/mL were obtained with compounds 3, 5, and 6 on all tested microbial species; meanwhile, with compound 4, they were only obtained on S. aureus. The lowest MIC value of 16 µg/mL was obtained by compound 5 against S. aureus and C. albicans, and also on compound 6 against C. albicans. The MMC determinations of compounds 3, 5, and 6 against the tester microorganisms were 40%, 100%, and 80%, respectively, within the tested interval (0.50–256 µg/mL) [20]. On the previous research, compound 18 was also tested for its antifungal activity against C. albicans and C. musae, with MIC values of 20 and 10 mg/mL, respectively [19].
The antibacterial activity of 7, 8, 9, 10, 11, and 12 were evaluated against S. aureus ATCC25932, B. cereus ATCC7064, B. subtilis ATCC6633, E. coli ATCC10536, and P. aeruginosa ATCC27853. Compound 8 demonstrated strong activity with MIC values of 32 µg/mL against S. aureus, B. cereus, and B. subtilis. Compounds 7 and 9 gave MIC values of 32, 64, and 64 µg/mL against S. aureus, B. cereus, and B. subtilis, respectively. However, compounds 7 and 8 had weak activity against E. coli, with MIC values of 128 µg/mL. MIC values of 64 µg/mL were also obtained with compound 10 against S. aureus and compound 11 against S. aureus, B. cereus, and B. subtilis. MIC values of 128 µg/mL were obtained with compound 10 against B. cereus & B. subtilis and compound 12 against S. aureus, B. cereus, and B. subtilis. Compounds 9, 10, and 11 had weak activity against E. coli, with MIC values of 256 µg/mL. Meanwhile, 12 had the weakest activity against E. coli, with MIC values of 512 µg/mL. All of the tested compounds showed weak activity against P. aureginosa, with MIC values of 256–512 µg/mL [28].
The in vitro antibacterial activity of 19 and 20 were evaluated by Alshaibani [25] against MRSA with MIC and MBC calculation. Compound 19 gives MIC and MBC values of 16 µg/mL and 32 µg/mL; meanwhile, compound 20 gives MIC and MBC values of 8 and 64 µg/mL (21). Compound 22 was evaluated for its antifungal activity against the pathogenic fungus U. maydis with MIC values of 1 mg/mL (2 mM). In addition, 22 displayed antibacterial activity against the human pathogenic bacterium M. tuberculosis with MIC values of 25 µg/mL (50 µM) [26].
Compounds 26 and 27 were evaluated for their antibacterial activity against S. aureus ATCC25932, B. cereus ATCC7064, B. subtilis ATCC6633, E. coli ATCC10536, Salmonella typhi ATCC19430, P. aeruginosa ATCC27853, and Serratia marcescens ATCC8100 compared with ampicillin and chloramphenicol as a positive control. Based on the IZ calculation, compounds 26 and 27 showed the highest activity against S. aureus, B. cereus, and B. subtilis (35.0 mm, 34.0 mm & 35.5 mm, and 33.0 mm, 32.5 mm & 33.5 mm, respectively). These results showed more potent activity than the positive control. However, compounds 26 and 27 showed moderate activity against E. coli, S. typhi, and S. marcescens (14.8 mm, 17.5 mm & 15.5 ± 1.65 mm, and 13.0 mm, 15.5 mm & 14.5 mm, respectively) and had weak activity against P. aeruginosa (12.0 mm and 10.0 mm, respectively). These results were also confirmed with the calculation of high MIC values of 26 and 27 against E. coli, S. typhi, S. marcescens, and P. aeruginosa (256 to 512 µg/mL), indicating 26 and 27 as moderate inhibitors against Gram-negative bacteria and the MBC values >512 µg/mL indicating no bactericidal activity against Gram-negative bacteria. On the other hand, MIC values of 0.5 µg/mL were obtained from both 26 and 27 against S. aureus, B. cereus, and B. subtilis. In addition, compound 26 showed the lowest MBC values of 2 µg/mL against Gram-positive bacteria, whereas compound 27 showed greater MBC varies in 4–16 µg/mL [21].
In 2012, Taechowisan et al. [29] also evaluated the antifungal activity of 28 and 29 against nine phytopathogenic fungi, namely A. porri, C. gloeosporioides, C. musae, Curvularia sp., Drechsler sp., Exserohilum sp., F. oxysporum, Verticillium sp., and S. rolfsii using the paper disk method. Based on the percentage of growth inhibition calculation, compound 28 showed growth inhibition of all tested fungi in 20.5%, 16.4%, 35.6%, 28.9%, 27.2%, 22.9%, 40.0%, 30.6%, and 28.6%, respectively; meanwhile, compound 29 showed the growth inhibition of all tested fungi in 24.1%, 20.2%, 33.8%, 31.2%, 36.1%, 34.3%, 36.2%, 29.4%, and 26.7%, respectively. Furthermore, the MIC values of 60 µg/mL were obtained from both 28 and 29 against A. porri, Curvularia sp., Drechsler sp., and Verticilllium sp., and from 29 against C. gloeosporioides and S. rolfsii. Meanwhile, MIC values of 120 µg/mL were obtained from both 28 and 29 against C. musae & Exserohilum sp. and from 28 against S. rolfsii. The weak MIC values of 240 were obtained from both 28 and 29 against F. oxysporum. However, compound 28 displayed the strongest activity against C. gloeosporioides with MIC values of 30 µg/mL [29].
Hu et al. [51] evaluated the antibacterial activity of 40 and 45. Compound 40 showed potent activity against the aquatic pathogens P. aeruginosa, V. parahaemolyticus, and V. alginolyticus, with MIC values of 1.0, 2.0, and 1,0 µg/mL, respectively, compared with the positive control chloromycetin (0.5 µg/mL on the tested microbial). In addition, compounds 40 and 45 displayed activity against the human pathogen E. coli, with MIC values of 4.0 and 0.5 µg/mL, respectively, compared with the positive control chloromycetin, with MIC values of 1.0 µg/mL. The result show that compound 45 has more potent activity than the positive control against E. coli [51].
4.2. Anticancer Activity
Compounds 31, 32, 33, 36, 37, 40, 41, 44, and 52 were evaluated for their cytotoxic activity toward the murine L5178Y cell lines with the IC50 values of 3.6, 5.3, 5.3, 8.1, 7.6, 18.3, 12.9, 6.2, and 10.9 µM, respectively. Compound 31 exhibited more potent activity than the positive control kahalalide F (IC50 4.3 µM) [23]. In addition, compounds 31, 32, 34, 35, 36, 37, 38, 39, 40, 43, 45, 49, 51, and 52 were evaluated for their cytotoxic activity against the A2780 human ovarian cancer cell line with the IC50 values of 9.7, 12.2, 12.2, 32.2, 7.8, 11.9, 19.4, 51.5, 8.2, 17.1, 5.3, 19.8, 28.5, and 12.8 µM, respectively [23]. The results indicated strong potential activity from 31, 36, 40, and 45. Compounds 31, 32, 34, 35, 36, 37, and 38 were also evaluated for their cytotoxicity against human urothelial bladder cancer cell line J82. The compounds active in bladder cancer cell lines are of high scientific interest regarding the rapid chemoresistance development of the cell. The tested compounds gave IC50 values of 40.6, 42.1, 55.3, 96.7, 31.7, 29.4, and 73.0 µM, respectively. These results indicate a lower potency than their activity against L5178Y and/or A2780 cell lines [23]. Furthermore, compounds 31, 32, 33, 36, 37, 40, 44, and 45 were tested toward the human embryonic kidney cell line HEK-293 with the IC50 values of 28.5, 21.7, 27.9, 37.4, 28.3, 44.6, 39.8, and 43.0 µM, respectively [23]. In addition, compound 44 gave inhibitory activity against protein tyrosine phosphatases with IC50 values of 13 and 17 µg/mL against PTP1B and TCPTP, respectively [52].
The cytotoxic and anticancer activities of 26 and 27 were evaluated against the three tumor cell lines: HepG2, HeLa, and Huh7, and one murine fibroblast cell line, L929, using the MTT assay. The results show that compounds 26 and 27 exhibited significant anticancer activity against HeLa cells with IC50 values of 3.04 and 3.96 µg/mL, respectively. The anticancer activity of 26 and 27 against HepG2 and Huh7 cells also showed high potential, with IC50 values of 15.42 & 17.52 µg/mL and 18.73 & 20.30 µg/mL, respectively. However, these compounds showed the weakest cytotoxic activity toward the L929 cell line, with IC50 values of 182.28 and 216.33 µg/mL, respectively [21].
4.3. Antioxidant Activity
In 2009, compounds 5 and 6 were evaluated for their protective effects on glutamate-induced cytotoxicity in mouse hippocampal HT22 cells. This cell line lacks ionotropic glutamate receptors, resulting in a high concentration of glutamate that inhibits cysteine uptake and depletes intracellular glutathione, which leads to the accumulation of reactive oxygen species (ROS). The effective protection ratios of 5 and 6 at a concentration of 100 µM are 62.4 ± 2.8% and 55.3 ± 3.4%, respectively, compared to the positive control, Trolox, with a protection ratio of 92.5 ± 2.5% at the same concentration [53]. Furthermore, the free radical scavenging activity of 5 and 6 was also measured by the interaction with stable free radical DPPH. The results exhibited potent scavenging effects on DPPH radical with the IC50 value of 60.74 and 75.65 µM, respectively, compared with L-Ascorbic acid as a positive control, with an IC50 value of 72.35 µM [53].
Compounds 26 and 27 were evaluated for their antioxidant activity using the decoloration of the ethanolic solution of DPPH. The absorption disappears in the presence of an active radical scavenger, and the subsequent decolorization is stoichiometric at a chosen range about the degree of reduction. The antioxidant activity of 26 and 27 gave the SC50 values of 85.84 and 88.26 µg/mL, respectively, compared with the positive control L-ascorbic acid, with an SC50 value of 50.25 µg/mL [21].
4.4. Anti-Inflammatory Activity
Compound 12 was evaluated for its anti-inflammatory activity against the release of β-glucuronidase and lysozyme from rat neutrophils. The result showed the IC50 value of 80.9 and >100 µM, respectively, compared to the positive control trifluoperazine against the release of β-glucuronidase and lysozyme, with IC50 values of 16.9 and 12.8 µM, respectively [54]. In 2014, the anti-inflammatory activity of compounds 42 and 52 was reported with the inhibition of LPS-induced NO production of mouse macrophage cell line RAW264.7. As a result, 52 inhibited the LPS-induced NO production at 10–30 µg/mL. On the other hand, compound 42 inhibited the NO production at 10 µg/mL. In addition, compound 52 inhibited with lower toxicity than 42 [55].
5. Future Perspectives
Some compounds from Zingiberaceae endophytes have the potency for further development. For example, for antimicrobials, there are 2′,7-dihydroxy-4′,5′-dimethoxyisoflavone demonstrating strong activity against S. aureus, B. cereus, and B. subtilis, and pretrichodermamide A show activity as an antimycobacterial with MIC values of 25 µg/mL. Paspaline shows more potent activity than the positive control (chloromycetin) against human pathogenic E. coli. Meanwhile, 3-methylcarbazole and 1-methoxy-3-methylcarbazole have activity against nine phytopathogenic fungi, namely A. porri, C. gloeosporioides, C. musae, Curvularia sp., Drechsler sp., Exserohilum sp., F. oxysporum, Verticillium sp., and S. rolfsii. A mechanism of antimicrobials can further examine these compounds: cell wall interference, protein, and nucleic acid synthesis, as well as inhibition of the metabolic pathway, membrane function, and ATP synthase [106]. In vivo studies of antimicrobial compounds in antiseptic efficacy are needed for antiseptic development. These studies of in vivo models are necessary at more advanced development stages, while the in vitro models are essential at the discovery stages [107].
Some cancer lines have been tested from a Zingiberaceae endophytes compound: murine L5178Y cell lines, A2780 human ovarian cancer cell line, human urothelial bladder cancer cell line J82, human embryonic kidney cell line HEK-293, HepG2, HeLa, Huh7, and one murine fibroblast cell line. The potent anticancer compound includes 3′-hydroxy-5-methoxy-3,4-methylenedioxybiphenyl, 3′-hydroxy-5,5′-dimethoxy-3,4 methylenedioxyphenyl, shearilicine, shearinine O, emindole SB, 7-hydroxy-13-dehydroxypaxilline, and paspaline. However, these compounds are needed for mechanism action studies, such as mitochondria-dependent cytochrome C, caspase activation, and extrinsic mechanisms of apoptosis (activation of tumor necrotic factor) [108]. Such studies could lead to new perspectives on how these compounds can induce apoptosis.
The antioxidant and anti-inflammatory activity have also been tested. For antioxidants, there is reactive oxygen species protection from glutamate-induced cytotoxicity in mouse hippocampal HT22 cells and DPPH. Two methylenedioxyphenyl from Streptomyces sp. BO-07 has potent as an antioxidant. The anti-inflammatory activities were reported to inhibit LPS-induced NO production of the mouse macrophage cell line RAW264.7 with the potent compound pyrapaxilline. However, further studies are needed on this compound, such as the production of nitrite, iNOS (inducible nitric oxide synthase) mRNA, and protein expression [109].
There are several methods to enhance the chemical diversity of bacterial and fungal endophyte metabolites. According to Betrand et al. [110], the co-culture of endophytes with other microorganisms can stimulate the activation of the silent gene. The genome approach strategy can be used, such as mutasynthesis, metabolic engineering, and heterologous expression. Also, fermentation media modification with the principle of one strain of many compounds can be performed [111]. Because of the complexity of microbial extraction, the analytical method (e.g., mass spectrometry methods and metabolomics) is the key to the successful detection and identification of new compounds [110]. The discoveries of new compounds from endophytes, such as Zingiberaceae endophytes, are interesting for pharmacological studies and natural product diversity.
6. Conclusions
In summary, Zingiberaceae plant endophytes have been studied thus far for secondary metabolites in Alipinia, Amomoum, Boesenbergia, Curcuma, and Zingiber genus. There are different strains of bacteria and fungi endophytes, including Streptomyces sp., Bacillus sp., Agrobacterium sp., Xylaria sp., Trichoderma sp., Aspergillus sp., Penicillium sp., etc. This endophyte produced many secondary metabolites such as polyketide, peptide, aromatic, alkaloid, and hybrid terpene alkaloid. The volatile constituent has been identified from GC–MS Spectra, produced by endophyte genera of Curcuma, Kaempferia, Zingiber, and other plants, such as terpene, steroid, alkaloid, aromatic, flavonoid, and others. These compounds have pharmacological activities for antibacterial, antifungal, anticancer, antioxidant, and anti-inflammatory. A comprehensive review of secondary metabolites from Zingiberaceae endophytes and their pharmacology activities has filled the gaps in information between the studies: species identification of endophytes compounds isolation and pharmacological potency. Hopefully, this review will increase knowledge for the development of endophytic research, especially in plants of the Zingiberaceae family.
Author Contributions
L.N. and A.A. wrote the manuscript, and U.S. provided critical input. U.S. supervised work development, helped in data interpretation and manuscript evaluation, and acted as the corresponding author. Both authors discussed the results and contributed to the final manuscript. All authors have read and agreed to the published version of the manuscript.
Funding
The Ministry of Education and Culture, Indonesia, for providing the funds for this research under the Doctoral Dissertation Research Grant, no: 1318/UN6.3.1/PT.00/2022 by Unang Supratman.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Conflicts of Interest
The authors declare no conflict of interest.
Abbreviations
| DFT | Density Functional Theory |
| DMF | Dimethylformamide |
| DPPH | 2,2-diphenyl-1-picrylhydrazyl |
| ECD | Electronic Circular Dichroism |
| GC–MS | Gas Chromatography–Mass Spectroscopy |
| IC50 | Inhibitory Concentration 50 |
| IZ | Inhibition Zones |
| LPS-induced NO | Lipopolysaccharide-Induced Nitric Oxide |
| MBC | Minimum Bactericidal Concentration |
| MFC | Minimum Fungicidal Concentration |
| MIC | Minimum Inhibitory Concentration |
| MMC | Minimum Microbicidal Concentration |
| MRSA | Methicillin-Resistant Staphylococcus aureus |
| MTT | (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) tetrazolium |
| NMR | Nuclear Magnetic Resonance |
| PCR | Polymerase Chain Reaction |
| PTP1B | Protein Tyrosine Phosphatase 1B |
| ROS | Reactive Oxygen Species |
| SC50 | Scavenging Concentration 50 |
| TCPTP | T-Cell Protein Tyrosine Phosphatase |
| TDDFT | Time-Dependent Density-Functional Theory |
| VCD | Vibrational Circular Dichroism |
References
- López, E.I.C.; Balcázar, M.F.H.; Mendoza, J.M.R.; Ortiz, A.D.R.; Melo, M.T.O.; Parrales, R.S.; Delgado, T.H. Antimicrobial activity of essential oil of Zingiber officinale Roscoe (Zingiberaceae). Am. J. Plant Sci. 2017, 8, 1511–1524. [Google Scholar] [CrossRef]
- Ewon, K.; Bhagya, A.S. A review on golden species of Zingiberaceae family around the world: Genus Curcuma. Afr. J. Agric. Res. 2019, 14, 519–531. [Google Scholar] [CrossRef]
- Tamokou, J.D.D.; Mbaveng, A.T.; Kuete, V. Antimicrobial activities of African medicinal spices and vegetables. In Medicinal Spices and Vegetables from Africa; Academic Press: Cambridge, MA, USA, 2017; pp. 207–237. [Google Scholar]
- Preetha, T.S.; Hemanthakumar, A.S.; Krishnan, P.N. A comprehensive review of Kaempferia galanga L. (Zingiberaceae): A high sought medicinal plant in Tropical Asia. J. Med. Plants Stud. 2016, 4, 270–276. [Google Scholar]
- Devi, N.B.; Singh, P.K.; Das, A.K. Ethnomedicinal utilization of Zingiberaceae in the valley districts of Manipur. J. Environ. Sci. Toxicol. Food Technol. 2014, 8, 21–23. [Google Scholar]
- Furmuly, A.M.; Azemi, N. A Review on Golden Species of Zingiberaceae Family Genus Curcuma. Sci. Proc. Ser. 2020, 2, 133–138. [Google Scholar] [CrossRef]
- Akinola, A.A.; Ahmad, S.; Maziah, M. Total antioxidant capacity, flavonoid, phenolic acid and polyphenol content in ten selected species of Zingiberaceae rhizomes. Afr. J. Tradit. Complementary Altern. Med. 2014, 11, 7–13. [Google Scholar] [CrossRef]
- Kumar, K.M.P.; Asish, G.R.; Sabu, M.; Balachandran, I. Significance of gingers (Zingiberaceae) in Indian system of medicine-Ayurveda: An overview. Anc. Sci. Life 2013, 32, 253. [Google Scholar] [CrossRef]
- Wohlmuth, H. Phytochemistry and Pharmacology of Plants from the Ginger family, Zingiberaceae. Ph.D. Thesis, Southern Cross University, East Lismore, Australia, 2008. [Google Scholar]
- Victório, C.P. Therapeutic value of the genus Alpinia, Zingiberaceae. Rev. Bras. Farmacogn. 2011, 21, 194–201. [Google Scholar] [CrossRef]
- Zahara, M.; Hasanah, M.; Zalianda, R. Identification of Zingiberaceae as medicinal plants in Gunung Cut Village, Aceh Barat Daya, Indonesia. J. Trop. Hortic. 2018, 1, 24–28. [Google Scholar] [CrossRef]
- Anisha, C.; Radhakrishnan, E.K. Metabolite analysis of endophytic fungi from cultivars of Zingiber officinale Rosc. identifies myriad of bioactive compounds including tyrosol. 3 Biotech 2017, 7, 1–10. [Google Scholar] [CrossRef]
- Krishnapura, P.R.; Belur, P.D. Isolation and screening of endophytes from the rhizomes of some Zingiberaceae plants for L-asparaginase production. Prep. Biochem. Biotechnol. 2016, 46, 281–287. [Google Scholar] [CrossRef] [PubMed]
- Shiono, Y.; Sasaki, T.; Shibuya, F.; Yasuda, Y.; Koseki, T.; Supratman, U. Isolation of a phomoxanthone A derivative, a new metabolite of tetrahydroxanthone, from a Phomopsis sp. isolated from the mangrove, Rhizhopora mucronata. Nat. Prod. Commun. 2013, 8, 1934578X1300801220. [Google Scholar] [CrossRef]
- Supratman, U.; Hirai, N.; Sato, S.; Watanabe, K.; Malik, A.; Annas, S.; Harneti, D.; Maharani, R.; Koseki, T.; Shiono, Y. New naphthoquinone derivatives from Fusarium napiforme of a mangrove plant. Nat. Prod. Res. 2021, 35, 1406–1412. [Google Scholar] [CrossRef] [PubMed]
- Supratman, U.; Suzuki, T.; Nakamura, T.; Yokoyama, Y.; Harneti, D.; Maharani, R.; Salam, S.; Abdullah, F.F.; Koseki, T.; Shiono, Y. New metabolites produced by endophyte Clonostachys rosea B5−2. Nat. Prod. Res. 2021, 35, 1525–1531. [Google Scholar] [CrossRef] [PubMed]
- Azhari, A.; Supratman, U. The Chemistry and Pharmacology of Fungal Genus Periconia: A Review. Sci. Pharm. 2021, 89, 34. [Google Scholar] [CrossRef]
- Taechowisan, T.; Lu, C.; Shen, Y.; Lumyong, S. Secondary metabolites from endophytic Streptomyces aureofaciens CMUAc130 and their antifungal activity. Microbiology 2005, 151, 1691–1695. [Google Scholar] [CrossRef] [PubMed]
- Taechowisan, T.; Wanbanjob, A.; Tuntiwachwuttikul, P.; Taylor, W.C. Identification of Streptomyces sp. Tc022, an endophyte in Alpinia galanga, and the isolation of actinomycin D. Ann. Microbiol. 2006, 56, 113–117. [Google Scholar] [CrossRef]
- Taechowisan, T.; Chuaychot, N.; Chanaphat, S.; Wanbanjob, A.; Shen, Y. Biological activity of chemical constituents isolated from Streptomyces sp. Tc052, an endophyte in Alpinia galanga. Int. J. Pharm. 2008, 4, 95–101. [Google Scholar] [CrossRef]
- Taechowisan, T.; Chaisaeng, S.; Phutdhawong, W.S. Antibacterial, antioxidant and anticancer activities of biphenyls from Streptomyces sp. BO-07: An endophyte in Boesenbergia rotunda (L.) Mansf A. Food Agric. Immunol. 2017, 28, 1330–1346. [Google Scholar] [CrossRef]
- Taechowisan, T.; Samsawat, T.; Puckdee, W.; Phutdhawong, W.S. Cytotoxicity and antibacterial activities of crude extract of Streptomyces sp. W08, an endophyte of Amomum krervanh Pierre. J. Appl. Pharm. Sci. 2021, 11, 134–138. [Google Scholar]
- Suebrasri, T.; Somteds, A.; Harada, H.; Kanokmedhakul, S.; Jogloy, S.; Ekprasert, J.; Lumyong, S.; Boonlue, S. Novel endophytic fungi with fungicidal metabolites suppress sclerotium disease. Rhizosphere 2020, 16, 100250. [Google Scholar] [CrossRef]
- Hammerschmidt, L.; Ola, A.; Mueller, W.E.; Lin, W.; Mandi, A.; Kurtan, T.; Proksch, P.; Aly, A.H. Two new metabolites from the endophytic fungus Xylaria sp. isolated from the medicinal plant Curcuma xanthorrhiza. Tetrahedron Lett. 2015, 56, 1193–1197. [Google Scholar] [CrossRef]
- Alshaibani, M.M.; Jalil, J.; Sidik, N.M.; Edrada-Ebel, R.; Zin, N.M. Isolation and characterization of cyclo-(tryptophanyl-prolyl) and chloramphenicol from Streptomyces sp. SUK 25 with antimethicillin-resistant Staphylococcus aureus activity. Drug Des. Dev. Ther. 2016, 10, 1817. [Google Scholar]
- Harwoko, H.; Daletos, G.; Stuhldreier, F.; Lee, J.; Wesselborg, S.; Feldbrügge, M.; Müller, W.E.; Kalscheuer, R.; Ancheeva, E.; Proksch, P. Dithiodiketopiperazine derivatives from endophytic fungi Trichoderma harzianum and Epicoccum nigrum. Nat. Prod. Res. 2021, 35, 257–265. [Google Scholar] [CrossRef] [PubMed]
- Ariantari, N.P.; Ancheeva, E.; Wang, C.; Mándi, A.; Knedel, T.O.; Kurtán, T.; Chaidir, C.; Müller, W.E.; Kassack, M.U.; Janiak, C.; et al. Indole diterpenoids from an endophytic Penicillium sp. J. Nat. Prod. 2019, 82, 1412–1423. [Google Scholar] [CrossRef]
- Taechowisan, T.; Chanaphat, S.; Ruensamran, W.; Phutdhawong, W.S. Antibacterial activity of new flavonoids from Streptomyces sp. BT01; an endophyte in Boesenbergia rotunda (L.) Mansf. J. Appl. Pharm. Sci. 2014, 4, 8–13. [Google Scholar]
- Taechowisan, T.; Chanaphat, S.; Ruensamran, W.; Phutdhawong, W.S. Antifungal activity of 3-methylcarbazoles from Streptomyces sp. LJK109; an endophyte in Alpinia galangal. J. Appl. Pharm. Sci. 2012, 2, 124–128. [Google Scholar]
- Pansanit, A.; Pripdeevech, P. Antibacterial secondary metabolites from an endophytic fungus, Arthrinium sp. MFLUCC16-1053 isolated from Zingiber cassumunar. Mycology 2018, 9, 264–272. [Google Scholar] [CrossRef]
- Vinayarani, G.; Prakash, H.S. Fungal endophytes of turmeric (Curcuma longa L.) and their biocontrol potential against pathogens Pythium aphanidermatum and Rhizoctonia solani. World J. Microbiol. Biotechnol. 2018, 34, 1–17. [Google Scholar] [CrossRef]
- Das, M.; Prakash, H.S.; Nalini, M.S. Antibacterial metabolites from Bipolaris specifera, an endophytic fungus from the endemic medicinal plant, Zingiber nimmonii (J. Graham) Dalzell. 3 Biotech 2020, 10, 1–8. [Google Scholar] [CrossRef]
- Krishnakumar, P.; Varghese, M.; Joe, M.G.; Rajagopal, A.; Varghese, L. Identification and bioactivities of endophytic fungi from Lagenandra toxicaria Dalz. and Kaempferia rotunda L. J. Appl. Biol. Biotechnol. 2021, 9, 1–2. [Google Scholar]
- Anisha, C.; Sachidanandan, P.; Radhakrishnan, E.K. Endophytic Paraconiothyrium sp. from Zingiber officinale Rosc. displays broad-spectrum antimicrobial activity by production of danthron. Curr. Microbiol. 2018, 75, 343–352. [Google Scholar] [CrossRef] [PubMed]
- Abbate, S.; Burgi, L.F.; Castiglioni, E.; Lebon, F.; Longhi, G.; Toscano, E.; Caccamese, S. Assessment of configurational and conformational properties of naringenin by vibrational circular dichroism. Chirality Pharmacol. Biol. Chem. Conseq. Mol. Asymmetry 2009, 21, 436–441. [Google Scholar] [CrossRef] [PubMed]
- Seephonkai, P.; Kongsaeree, P.; Prabpai, S.; Isaka, M.; Thebtaranonth, Y. Transformation of an irregularly bridged epidithiodiketopiperazine to trichodermamide A. Org. Lett. 2006, 8, 3073–3075. [Google Scholar] [CrossRef]
- Kumar, A.; Singh, M.; Singh, P.P.; Singh, S.K.; Singh, P.K.; Pandey, K.D. Isolation of plant growth promoting rhizobacteria and their impact on growth and curcumin content in Curcuma longa L. Biocatal. Agric. Biotechnol. 2016, 8, 1–7. [Google Scholar] [CrossRef]
- Aswathy, A.J.; Jasim, B.; Jyothis, M.; Radhakrishnan, E.K. Identification of two strains of Paenibacillus sp. as indole 3 acetic acid-producing rhizome-associated endophytic bacteria from Curcuma longa. 3 Biotech 2013, 3, 219–224. [Google Scholar] [CrossRef]
- Jasim, B.; Joseph, A.A.; John, C.J.; Mathew, J.; Radhakrishnan, E.K. Isolation and characterization of plant growth promoting endophytic bacteria from the rhizome of Zingiber officinale. 3 Biotech 2014, 4, 197–204. [Google Scholar] [CrossRef]
- Chen, J.; Zhang, P.; Ye, X.; Wei, B.; Emam, M.; Zhang, H.; Wang, H. The structural diversity of marine microbial secondary metabolites based on co-culture strategy: 2009–2019. Mar. Drugs 2020, 18, 449. [Google Scholar] [CrossRef]
- Zhang, Y.; Kang, X.; Liu, H.; Liu, Y.; Li, Y.; Yu, X.; Zhao, K.; Gu, Y.; Xu, K.; Chen, C.; et al. Endophytes isolated from ginger rhizome exhibit growth promoting potential for Zea mays. Arch. Agron. Soil Sci. 2018, 64, 1302–1314. [Google Scholar] [CrossRef]
- Hartanto, A.; Lutfia, A.; Munir, E. Identification of a Potential IAA-Producing Fungus Isolated from Alpinia sp. Rhizome in Hutan Sibayak, North Sumatera. In Journal of Physics: Conference Series; IOP Publishing: Bristol, UK, 2019; Volume 1351, p. 012024. [Google Scholar]
- Smith, A.B.; Cui, H. Total synthesis of (−)-21-isopentenylpaxilline. Org. Lett. 2003, 5, 587–590. [Google Scholar] [CrossRef]
- Springer, J.P.; Clardy, J.; Wells, J.M.; Cole, R.J.; Kirksey, J.W. The structure of paxilline, a tremorgenic metabolite of Penicillium paxilli Bainier. Tetrahedron Lett. 1975, 16, 2531–2534. [Google Scholar] [CrossRef]
- Nozawa, K.; Nakajima, S.; Kawai, K.I.; Udagawa, S.I. Isolation and structures of indoloditerpenes, possible biosynthetic intermediates to the tremorgenic mycotoxin, paxilline, from Emericella striata. J. Chem. Soc. Perkin Trans. 1988, 1, 2607–2610. [Google Scholar] [CrossRef]
- Yu, J.; Wang, J.P.; Liu, S.F.; Yin, C.Y.; Tang, D.Y.; Li, Y.H.; Zhang, L.X. 7-Methoxy-13-dehydroxypaxilline: New indole diterpenoid from an endophytic fungus Penicillium sp. Nb 19. Nat. Prod. Res. 2022, 36, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.; Gessner, G.; Groth, I.; Lange, C.; Christner, A.; Bruhn, T.; Deng, Z.; Li, X.; Heinemann, S.H.; Grabley, S.; et al. Shearinines D–K, new indole triterpenoids from an endophytic Penicillium sp. (strain HKI0459) with blocking activity on large-conductance calcium-activated potassium channels. Tetrahedron 2007, 63, 435–444. [Google Scholar] [CrossRef]
- Gao, N.; Shang, Z.C.; Yu, P.; Luo, J.; Jian, K.L.; Kong, L.Y.; Yang, M.H. Alkaloids from the endophytic fungus Penicillium brefeldianum and their cytotoxic activities. Chin. Chem. Lett. 2017, 28, 1194–1199. [Google Scholar] [CrossRef]
- El-Gendy, M.M.; El-Bondkly, A.M. Production and genetic improvement of a novel antimycotic agent, saadamycin, against dermatophytes and other clinical fungi from endophytic Streptomyces sp. Hedaya48. J. Ind. Microbiol. Biotechnol. 2010, 37, 831–841. [Google Scholar] [CrossRef]
- El-Lakany, A.M.; Aboul-Ela, M.A.; Abdul-Ghani, M.M.; Mekky, H. Chemical constituents and biological activities of Cichorium intybus L. Nat. Prod. Sci. 2004, 10, 69–73. [Google Scholar]
- Hu, X.Y.; Meng, L.H.; Li, X.; Yang, S.Q.; Li, X.M.; Wang, B.G. Three new indole diterpenoids from the sea-anemone-derived fungus Penicillium sp. AS-79. Mar. Drugs 2017, 15, 137. [Google Scholar] [CrossRef]
- Zhou, L.M.; Kong, F.D.; Fan, P.; Ma, Q.Y.; Xie, Q.Y.; Li, J.H.; Zheng, H.Z.; Zheng, Z.H.; Yuan, J.Z.; Dai, H.F.; et al. Indole-diterpenoids with protein tyrosine phosphatase inhibitory activities from the marine-derived fungus Penicillium sp. KFD28. J. Nat. Prod. 2019, 82, 2638–2644. [Google Scholar] [CrossRef]
- Taechowisan, T.; Chuaychot, N.; Chanaphat, S.; Wanbanjob, A.; Shen, Y. Cytoprotective activity of chemical constituents isolated from Streptomyces sp. Int. J. Biol. Chem. 2009, 3, 11–17. [Google Scholar] [CrossRef]
- Chan, S.C.; Chang, Y.S.; Wang, J.P.; Chen, S.C.; Kuo, S.C. Three new flavonoids and antiallergic, anti-inflammatory constituents from the heartwood of Dalbergia odorifera. Planta Med. 1998, 64, 153–158. [Google Scholar] [CrossRef] [PubMed]
- Matsui, C.; Ikeda, Y.; Iinuma, H.; Kushida, N.; Kunisada, T.; Simizu, S.; Umezawa, K. Isolation of a novel paxilline analog pyrapaxilline from fungus that inhibits LPS-induced NO production. J. Antibiot. 2014, 67, 787–790. [Google Scholar] [CrossRef] [PubMed]
- Salasiah, M.; Meekiong, K. Preliminary anatomical study on leaf surfaces of bornean Zingiberaceae (tribe Alpinieae) from north east sarawak. Malays. Appl. Biol. 2018, 47, 289–293. [Google Scholar]
- Pedersen, L.B. Phylogenetic analysis of the subfamily Alpinioideae (Zingiberaceae), particularly Etlingera Giseke, based on nuclear and plastid DNA. Plant Syst. Evol. 2004, 245, 239–258. [Google Scholar] [CrossRef]
- Gevú, K.V.; Da Cunha, M.; Barros, C.F.; Pereira, S.M.; Lima, H.R.P. Structural analysis of subterranean organs in Zingiberaceae. Plant Syst. Evol. 2014, 300, 1089–1098. [Google Scholar] [CrossRef]
- Zhao, H.; Xiao, M.H.; Zhong, Y.; Wang, Y.Q. Leaf epidermal micromorphology of Zingiber (Zingiberaceae) from China and its systematic significance. PhytoKeys 2022, 190, 131. [Google Scholar] [CrossRef]
- Eswani, N.; Abd Kudus, K.; Nazre, M.; Noor, A.A.; Ali, M. Medicinal plant diversity and vegetation analysis of logged over hill forest of Tekai Tembeling Forest Reserve, Jerantut, Pahang. J. Agric. Sci. 2010, 2, 189. [Google Scholar] [CrossRef]
- Kress, W.J.; Prince, L.M.; Williams, K.J. The phylogeny and a new classification of the gingers (Zingiberaceae): Evidence from molecular data. Am. J. Bot. 2002, 89, 1682–1696. [Google Scholar] [CrossRef]
- Saensouk, P.; Saensouk, S. Diversity, traditional uses and conservation status of Zingiberaceae in Udorn Thani Province, Thailand. Biodiversitas J. Biol. Divers. 2021, 22, 3083–3097. [Google Scholar] [CrossRef]
- Larsen, K.; Lock, J.M.; Maas, H.; Maas, P.J.M. Zingiberaceae. In Flowering Plants· Monocotyledons; Springer: Berlin/Heidelberg, Germany, 1998; pp. 474–495. [Google Scholar]
- Newman, M.; Lhuillier, A.; Poulsen, A.D. Checklist of the Zingiberaceae of Malesia. Blumea. Suppl. 2004, 16, 1–166. [Google Scholar]
- Kuehny, J.S.; Sarmiento, M.; Paz, M.P.; Branch, P.C. Effect of light intensity, photoperiod and plant growth retardants on production of Zingiberaceae as pot plants. Acta Hortic. 2005, 683, 145. [Google Scholar] [CrossRef]
- Thomas, S.; Britto, S.J.; Mani, B. A comprehensive account on the genus Hedychium J. konig (Zingiberaceae) in south India. Plant Sci. Today 2022, 9, 12–18. [Google Scholar] [CrossRef]
- Basak, S.; Sarma, G.C.; Rangan, L. Ethnomedical uses of Zingiberaceous plants of Northeast India. J. Ethnopharmacol. 2010, 132, 286–296. [Google Scholar]
- Navia, Z.I.; Audira, D.; Afifah, N.; Turnip, K.; NURAINI, N.; Suwardi, A.B. Ethnobotanical investigation of spice and condiment plants used by the Taming tribe in Aceh, Indonesia. Biodiversitas J. Biol. Divers. 2020, 21, 4467–4473. [Google Scholar] [CrossRef]
- Rachkeeree, A.; Kantadoung, K.; Suksathan, R.; Puangpradab, R.; Page, P.A.; Sommano, S.R. Nutritional compositions and phytochemical properties of the edible flowers from selected Zingiberaceae found in Thailand. Front. Nutr. 2018, 5, 3. [Google Scholar] [CrossRef] [PubMed]
- Sharma, S.; Ghataury, S.K.; Sarathe, A.; Dubey, G.; Parkhe, G. Curcuma angustifolia Roxb, (Zingiberaceae): Ethnobotany, phytochemistry and pharmacology: A review. J. Pharmacogn. Phytochem. 2019, 8, 1535–1540. [Google Scholar]
- Handa, S.S.; Khanuja, S.P.S.; Longo, G.; Rakesh, D.D. Extraction Technologies for Medicinal and Aromatic Plants; UNIDO: Trieste, Italy, 2008. [Google Scholar]
- Sarkic, A.; Stappen, I. Essential oils and their single compounds in cosmetics—A critical review. Cosmetics 2018, 5, 11. [Google Scholar] [CrossRef]
- Mamusa, M.; Resta, C.; Sofroniou, C.; Baglioni, P. Encapsulation of volatile compounds in liquid media: Fragrances, flavors, and essential oils in commercial formulations. Adv. Colloid Interface Sci. 2021, 298, 102544. [Google Scholar] [CrossRef]
- Aburjai, T.; Natsheh, F.M. Plants used in cosmetics. Phytother. Res. Int. J. Devoted Pharmacol. Toxicol. Eval. Nat. Prod. Deriv. 2003, 17, 987–1000. [Google Scholar] [CrossRef]
- Sharmeen, J.B.; Mahomoodally, F.M.; Zengin, G.; Maggi, F. Essential oils as natural sources of fragrance compounds for cosmetics and cosmeceuticals. Molecules 2021, 26, 666. [Google Scholar] [CrossRef]
- Abate, L.; Bachheti, A.; Bachheti, R.K.; Husen, A.; Getachew, M.; Pandey, D.P. Potential role of forest-based plants in essential oil production: An approach to cosmetic and personal health care applications. In Non-Timber Forest Products; Springer: Cham, Switzerland, 2021; pp. 1–18. [Google Scholar]
- Tongnuanchan, P.; Benjakul, S. Essential oils: Extraction, bioactivities, and their uses for food preservation. J. Food Sci. 2014, 79, R1231–R1249. [Google Scholar] [CrossRef] [PubMed]
- Figueiredo, A.C.; Barroso, J.G.; Pedro, L.G.; Scheffer, J.J. Factors affecting secondary metabolite production in plants: Volatile components and essential oils. Flavour Fragr. J. 2008, 23, 213–226. [Google Scholar] [CrossRef]
- Okigbo, R.N.; Anuagasi, C.L.; Amadi, J.E. Advances in selected medicinal and aromatic plants indigenous to Africa. J. Med. Plants Res. 2009, 3, 86–95. [Google Scholar]
- Paibon, W.; Yinmoi, C.A.; Tembab, N.; Boonlue, W.; Jampachaisri, K.; Nuengchamnong, N.; Waranuch, N.; Ingkaninan, K. Comparison and evaluation of volatile oils from three different extraction methods for some Thai fragrant flowers. Int. J. Cosmet. Sci. 2011, 33, 150–156. [Google Scholar] [CrossRef] [PubMed]
- Kaddes, A.; Fauconnier, M.L.; Sassi, K.; Nasraoui, B.; Jijakli, M.H. Endophytic fungal volatile compounds as solution for sustainable agriculture. Molecules 2019, 24, 1065. [Google Scholar] [CrossRef] [PubMed]
- Plaszkó, T.; Szűcs, Z.; Kállai, Z.; Csoma, H.; Vasas, G.; Gonda, S. Volatile organic compounds (VOCs) of endophytic fungi growing on extracts of the host, horseradish (Armoracia rusticana). Metabolites 2020, 10, 451. [Google Scholar] [CrossRef] [PubMed]
- Yu, D.; Xu, F.; Zeng, J.; Zhan, J. Type III polyketide synthases in natural product biosynthesis. IUBMB Life 2012, 64, 285–295. [Google Scholar] [CrossRef]
- Hutchinson, C.R.; Kennedy, J.; Park, C.; Kendrew, S.; Auclair, K.; Vederas, J. Aspects of the biosynthesis of non-aromatic fungal polyketides by iterative polyketide synthases. Antonie Van Leeuwenhoek 2000, 78, 287–295. [Google Scholar] [CrossRef]
- Wang, H.; Fewer, D.P.; Holm, L.; Rouhiainen, L.; Sivonen, K. Atlas of nonribosomal peptide and polyketide biosynthetic pathways reveals common occurrence of nonmodular enzymes. Proc. Natl. Acad. Sci. USA 2014, 111, 9259–9264. [Google Scholar] [CrossRef]
- Miyanaga, A. Structure and function of polyketide biosynthetic enzymes: Various strategies for production of structurally diverse polyketides. Biosci. Biotechnol. Biochem. 2017, 81, 2227–2236. [Google Scholar] [CrossRef]
- Bhattarai, K.; Kabir, M.E.; Bastola, R.; Baral, B. Fungal natural products galaxy: Biochemistry and molecular genetics toward blockbuster drugs discovery. Adv. Genet. 2021, 107, 193–284. [Google Scholar] [PubMed]
- Sun, D.J.; Zhu, L.J.; Zhao, Y.Q.; Zhen, Y.Q.; Zhang, L.; Lin, C.C.; Chen, L.X. Diarylheptanoid: A privileged structure in drug discovery. Fitoterapia 2020, 142, 104490. [Google Scholar] [CrossRef] [PubMed]
- Dell, M.; Dunbar, K.L.; Hertweck, C. Ribosome-independent peptide biosynthesis: The challenge of a unifying nomenclature. Nat. Prod. Rep. 2022, 39, 453–459. [Google Scholar] [CrossRef] [PubMed]
- Hur, G.H.; Vickery, C.R.; Burkart, M.D. Explorations of catalytic domains in nonribosomal peptide synthetase enzymology. Nat. Prod. Rep. 2012, 29, 1074–1098. [Google Scholar] [CrossRef] [PubMed]
- Little, R.F.; Hertweck, C. Chain release mechanisms in polyketide and nonribosomal peptide biosynthesis. Nat. Prod. Rep. 2022, 39, 163–205. [Google Scholar] [CrossRef] [PubMed]
- Finking, R.; Marahiel, M.A. Biosynthesis of Nonribosomal Peptides. Annu. Rev. Microbiol. 2004, 58, 453–488. [Google Scholar] [CrossRef] [PubMed]
- Keller, N.P.; Turner, G.; Bennett, J.W. Fungal secondary metabolism—From biochemistry to genomics. Nat. Rev. Microbiol. 2005, 3, 937–947. [Google Scholar] [CrossRef]
- Wang, Y.T.; Zhu, L.; Zeng, D.; Long, W.; Zhu, S.M. Chemical composition and anti-inflammatory activities of essential oil from Trachydium roylei. J. Food Drug Anal. 2016, 24, 602–609. [Google Scholar] [CrossRef]
- Samarth, R.M.; Samarth, M.; Matsumoto, Y. Medicinally important aromatic plants with radioprotective activity. Future Sci. OA 2017, 3, FSO247. [Google Scholar] [CrossRef]
- Kumar, Y.; Prakash, O.; Tripathi, H.; Tandon, S.; Gupta, M.M.; Rahman, L.U.; Khan, F. AromaDb: A database of medicinal and aromatic plant’s aroma molecules with phytochemistry and therapeutic potentials. Front. Plant Sci. 2018, 9, 1081. [Google Scholar] [CrossRef]
- Schwab, W.; Davidovich-Rikanati, R.; Lewinsohn, E. Biosynthesis of plant-derived flavor compounds. Plant J. 2008, 54, 712–732. [Google Scholar] [CrossRef] [PubMed]
- Roy, A. A review on the alkaloids an important therapeutic compound from plants. IJPB 2017, 3, 1–9. [Google Scholar]
- Ziegler, J.; Facchini, P.J. Alkaloid biosynthesis: Metabolism and trafficking. Annu. Rev. Plant Biol. 2008, 59, 735. [Google Scholar] [CrossRef] [PubMed]
- Aniszewski, T. Alkaloids-Secrets of Life: Alkaloid Chemistry, Biological Significance, Applications and Ecological Role; Elsevier: Amsterdam, The Netherlands, 2007. [Google Scholar]
- Yang, L.; Stöckigt, J. Trends for diverse production strategies of plant medicinal alkaloids. Nat. Prod. Rep. 2010, 27, 1469–1479. [Google Scholar] [CrossRef] [PubMed]
- Bribi, N. Pharmacological activity of alkaloids: A review. Asian J. Bot. 2018, 1, 1–6. [Google Scholar]
- Jiang, M.; Wu, Z.; Liu, L.; Chen, S. The chemistry and biology of fungal meroterpenoids (2009–2019). Org. Biomol. Chem. 2021, 19, 1644–1704. [Google Scholar] [CrossRef]
- Zhang, P.; Li, X.; Wang, B.G. Secondary metabolites from the marine algal-derived endophytic fungi: Chemical diversity and biological activity. Planta Med. 2016, 82, 832–842. [Google Scholar] [CrossRef]
- Corsello, M.A.; Kim, J.; Garg, N.K. Indole diterpenoid natural products as the inspiration for new synthetic methods and strategies. Chem. Sci. 2017, 8, 5836–5844. [Google Scholar] [CrossRef]
- Radulovic, N.S.; Blagojevic, P.D.; Stojanovic-Radic, Z.Z.; Stojanovic, N.M. Antimicrobial plant metabolites: Structural diversity and mechanism of action. Curr. Med. Chem. 2013, 20, 932–952. [Google Scholar]
- Agafonova, M.N.; Kazakova, R.R.; Lubina, A.P.; Zeldi, M.I.; Nikitina, E.V.; Balakin, K.V.; Shtyrlin, Y.G. Antibacterial activity profile of miramistin in in vitro and in vivo models. Microb. Pathog. 2020, 142, 104072. [Google Scholar] [CrossRef]
- Thushara, R.M.; Hemshekhar, M.; Santhosh, M.S.; Devaraja, S.; Kemparaju, K.; Girish, K.S. Differential action of phytochemicals on platelet apoptosis: A biological overview. Curr. Med. Chem. 2013, 20, 1018–1027. [Google Scholar] [PubMed]
- Bhaskaran, N.; Shukla, S.; Srivastava, J.K.; Gupta, S. Chamomile: An anti-inflammatory agent inhibits inducible nitric oxide synthase expression by blocking RelA/p65 activity. Int. J. Mol. Med. 2010, 26, 935–940. [Google Scholar] [PubMed]
- Bertrand, S.; Bohni, N.; Schnee, S.; Schumpp, O.; Gindro, K.; Wolfender, J.L. Metabolite induction via microorganism co-culture: A potential way to enhance chemical diversity for drug discovery. Biotechnol. Adv. 2014, 32, 1180–1204. [Google Scholar] [CrossRef] [PubMed]
- Pan, R.; Bai, X.; Chen, J.; Zhang, H.; Wang, H. Exploring structural diversity of microbe secondary metabolites using OSMAC strategy: A literature review. Front. Microbiol. 2019, 10, 294. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).