New Quinazolin-4(3H)-one Derivatives Incorporating Hydrazone and Pyrazole Scaffolds as Antimicrobial Agents Targeting DNA Gyraze Enzyme
Abstract
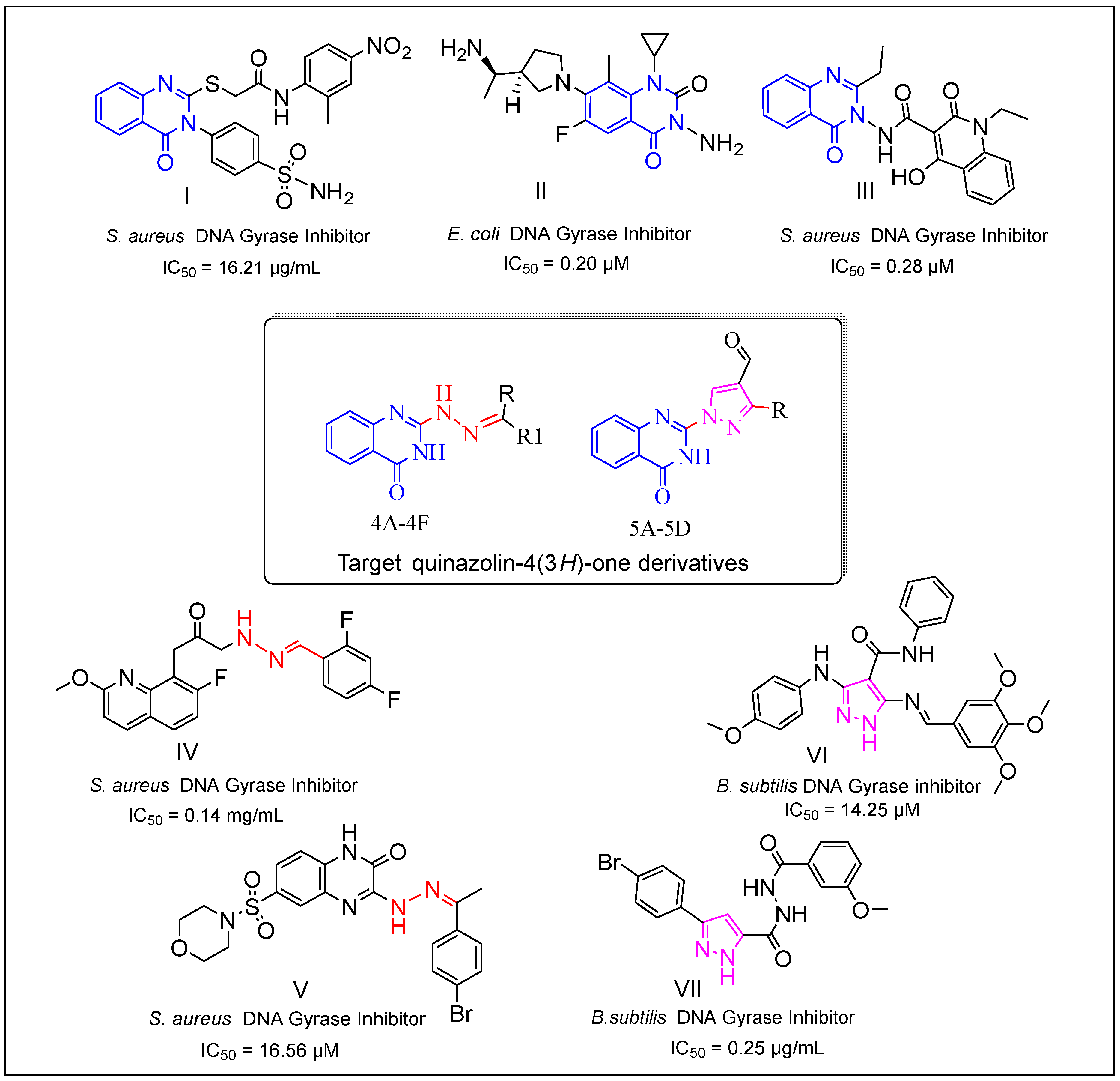
1. Introduction
2. Materials and Methods
2.1. Chemistry
2.1.1. General Information
2.1.2. Synthesis of 2-hydrazinylquinazolin-4(3H)-one (2)
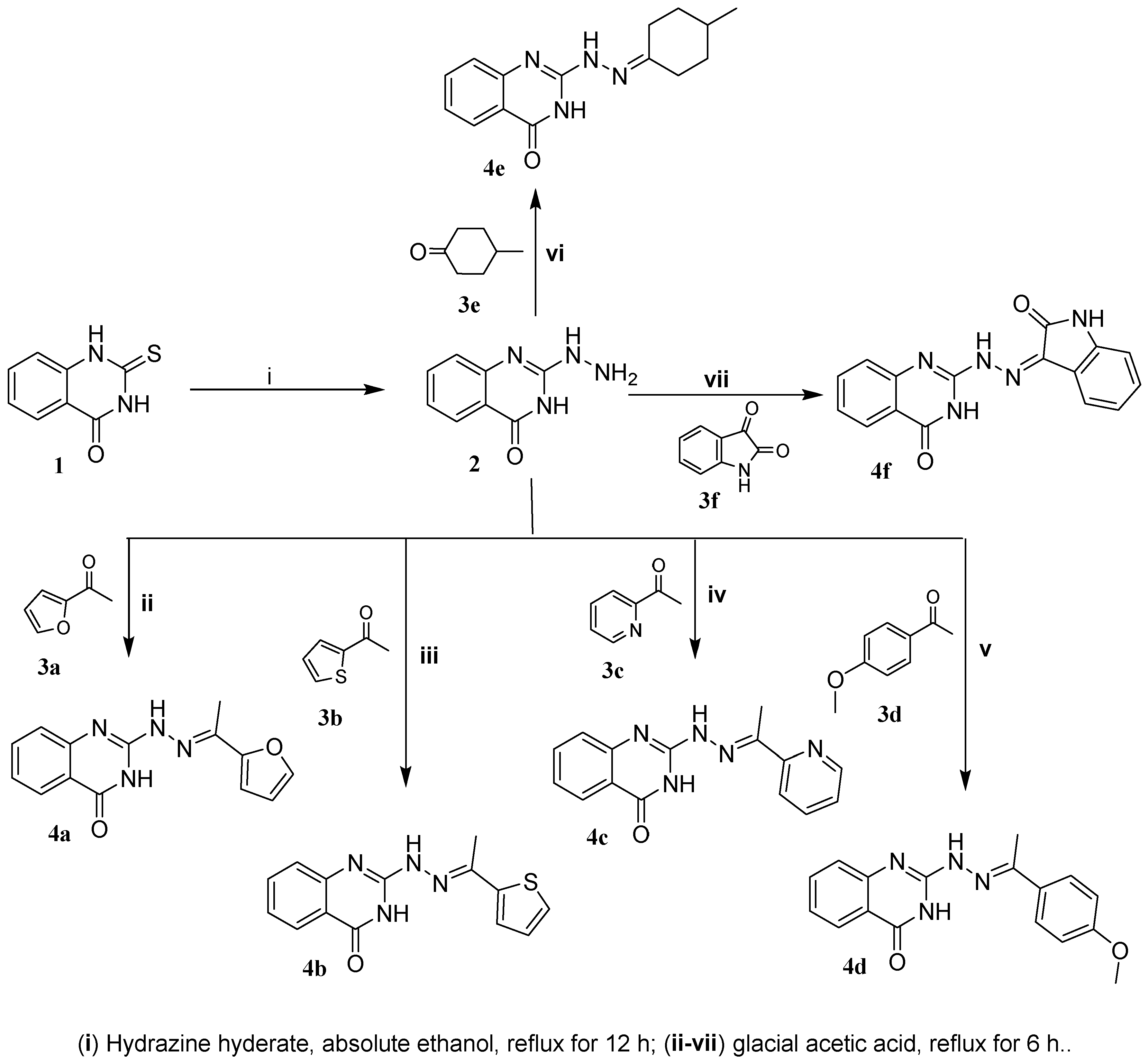
2.1.3. Synthesis of 2-(2-ethylidene-hydrazinyl)quinazolin-4(3H)-one Derivatives 4a–f
2-(2-(1-(Furan-2-yl)ethylidene)hydrazinyl)quinazolin-4(3H)-one (4a)
2-(2-(1-(Thiophen-2-yl)ethylidene)hydrazinyl)quinazolin-4(3H)-one (4b)
2-(2-(1-(Pyridin-2-yl)ethylidene)hydrazinyl)quinazolin-4(3H)-one (4c)
2-(2-(1-(4-Methoxyphenyl)ethylidene)hydrazinyl)quinazolin-4(3H)-one (4d)
2-(2-(4-Methylcyclohexylidene)hydrazinyl)quinazolin-4(3H)-one (4e)
2-(2-(2-Oxoindolin-3-ylidene)hydrazinyl)quinazolin-4(3H)-one (4f)
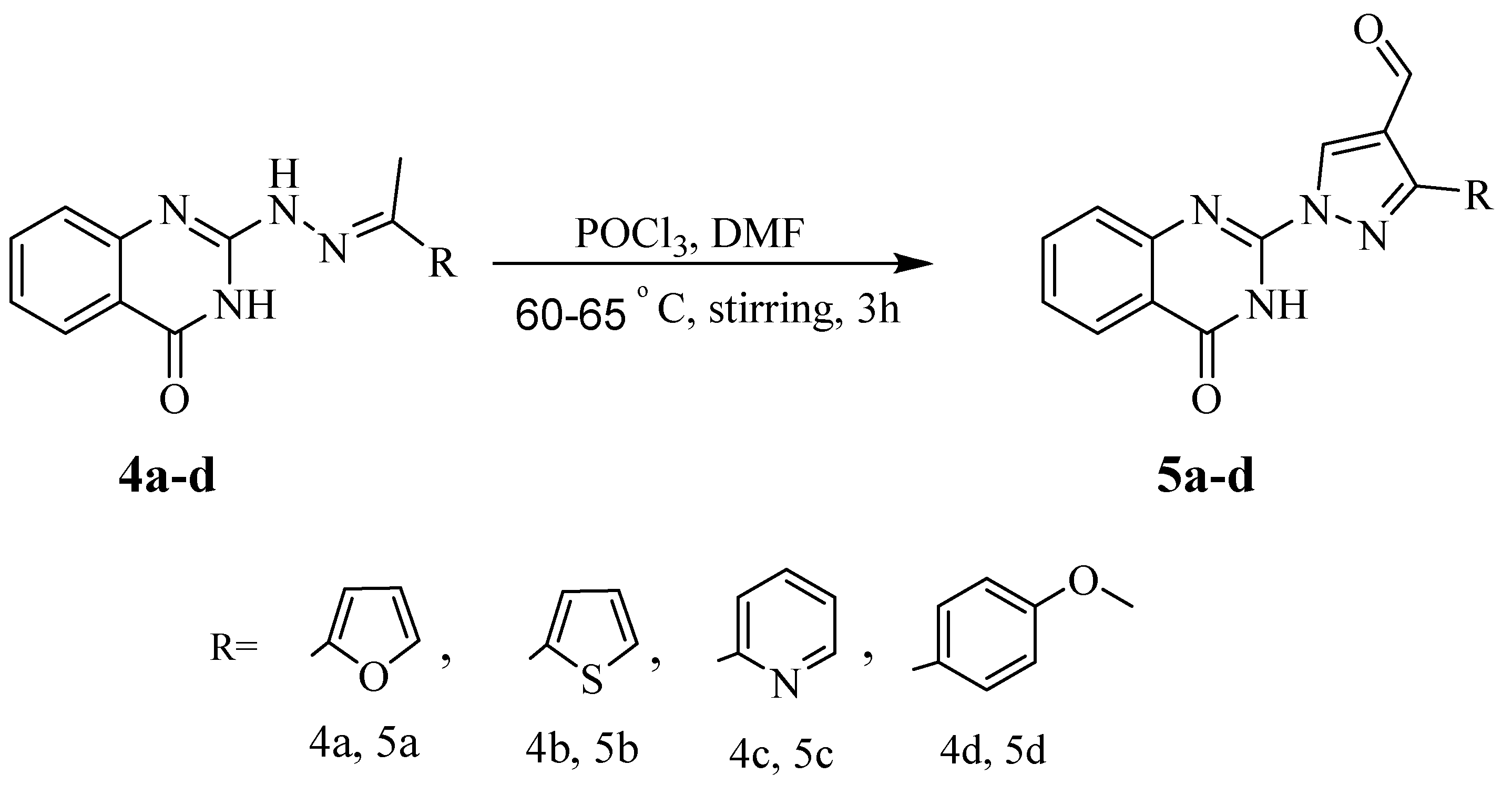
2.1.4. Synthesis of 1-(4-oxo-3,4-dihydroquinazolin-2-yl)-1H-pyrazole-4-carbaldehyde Derivatives (5a–d)
3-(Furan-2-yl)-1-(4-oxo-3,4-dihydroquinazolin-2-yl)-1H-pyrazole-4-carbaldehyde (5a)
1-(4-Oxo-3,4-dihydroquinazolin-2-yl)-3-(thiophen-2-yl)-1H-pyrazole-4-carbaldehyde (5b)
1-(4-Oxo-3,4-dihydroquinazolin-2-yl)-3-(pyridin-2-yl)-1H-pyrazole-4-carbaldehyde (5c)
3-(4-Methoxyphenyl)-1-(4-oxo-3,4-dihydroquinazolin-2-yl)-1H-pyrazole-4-carbaldehyde (5d)
2.2. Antimicrobial Screening
2.3. DNA Gyrase Supercoiling Inhibition Assay
2.4. Molecular Modeling Studies
2.5. In Silico ADME Prediction Study
3. Results and Discussion
3.1. Chemistry
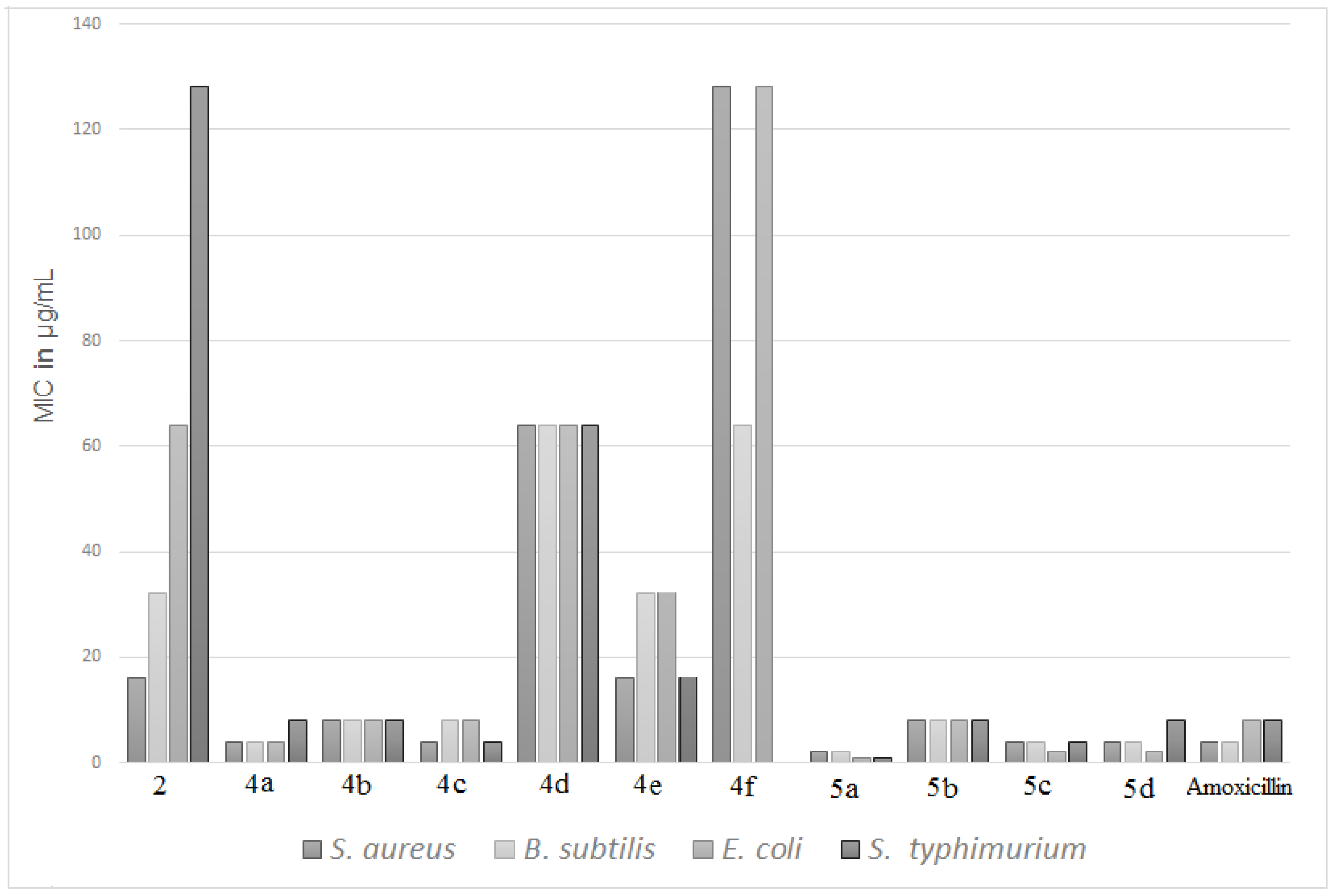
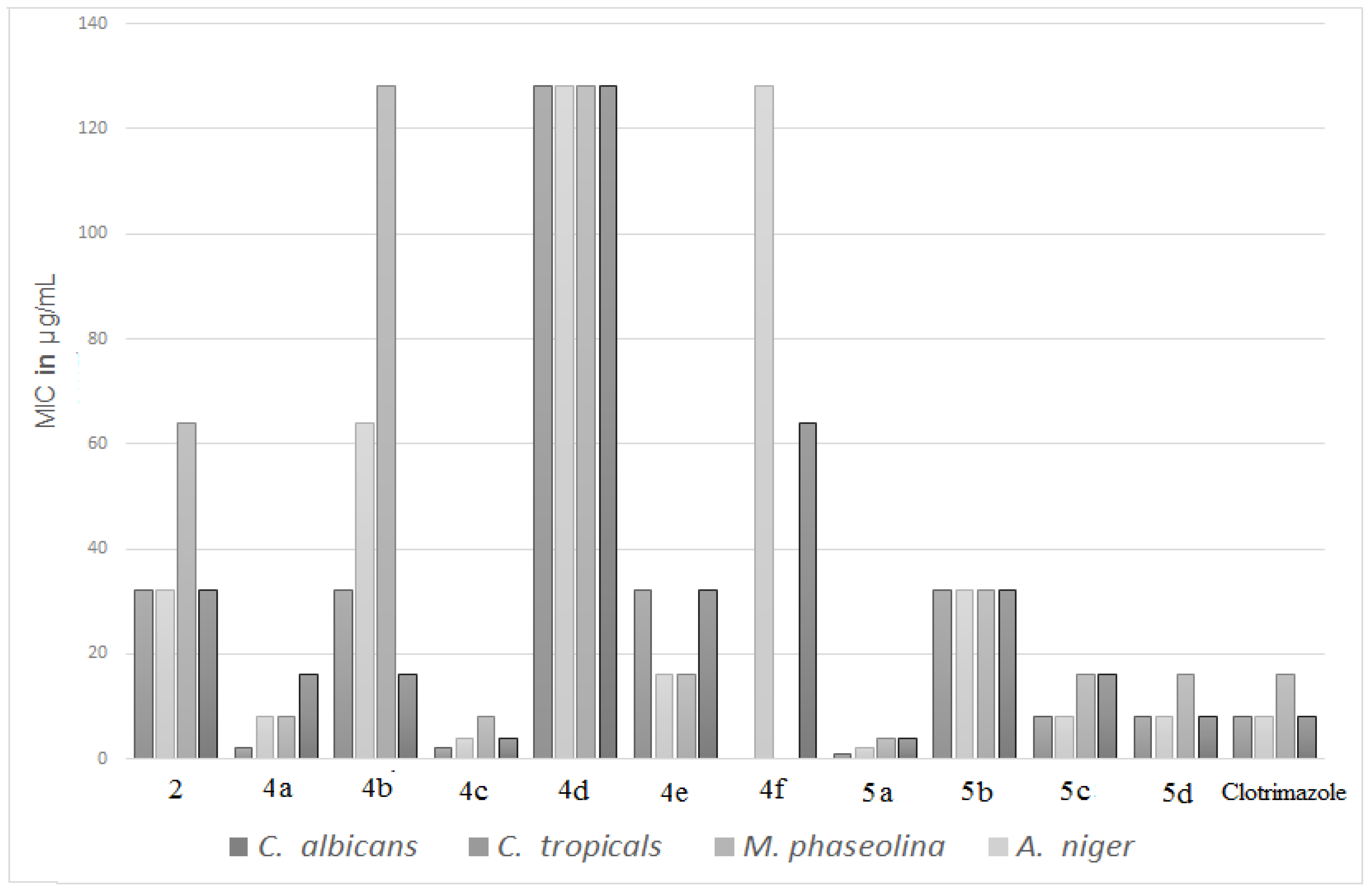
3.2. Antimicrobial Activity
3.3. In Vitro DNA Gyrase Inhibitory Activity
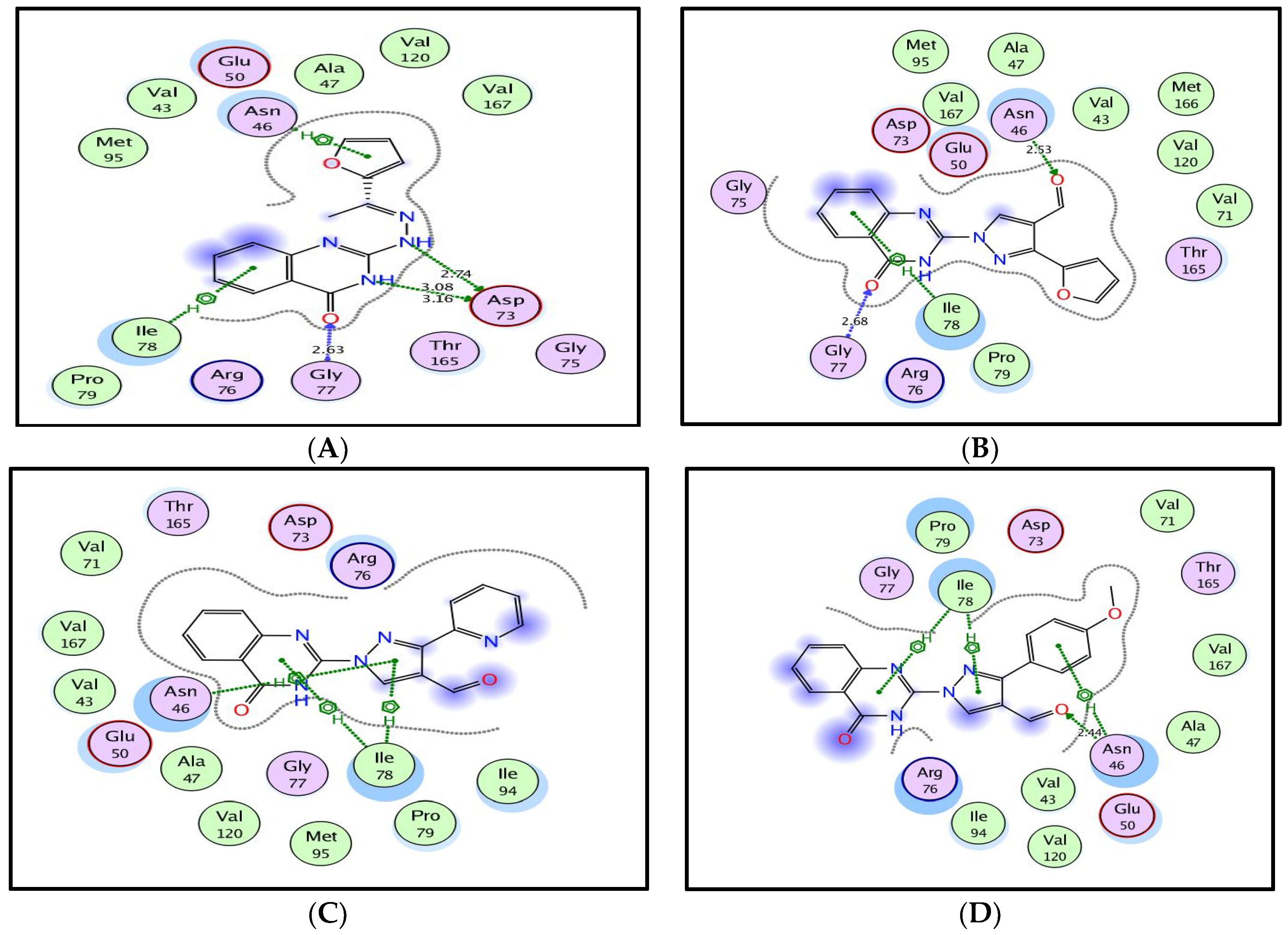
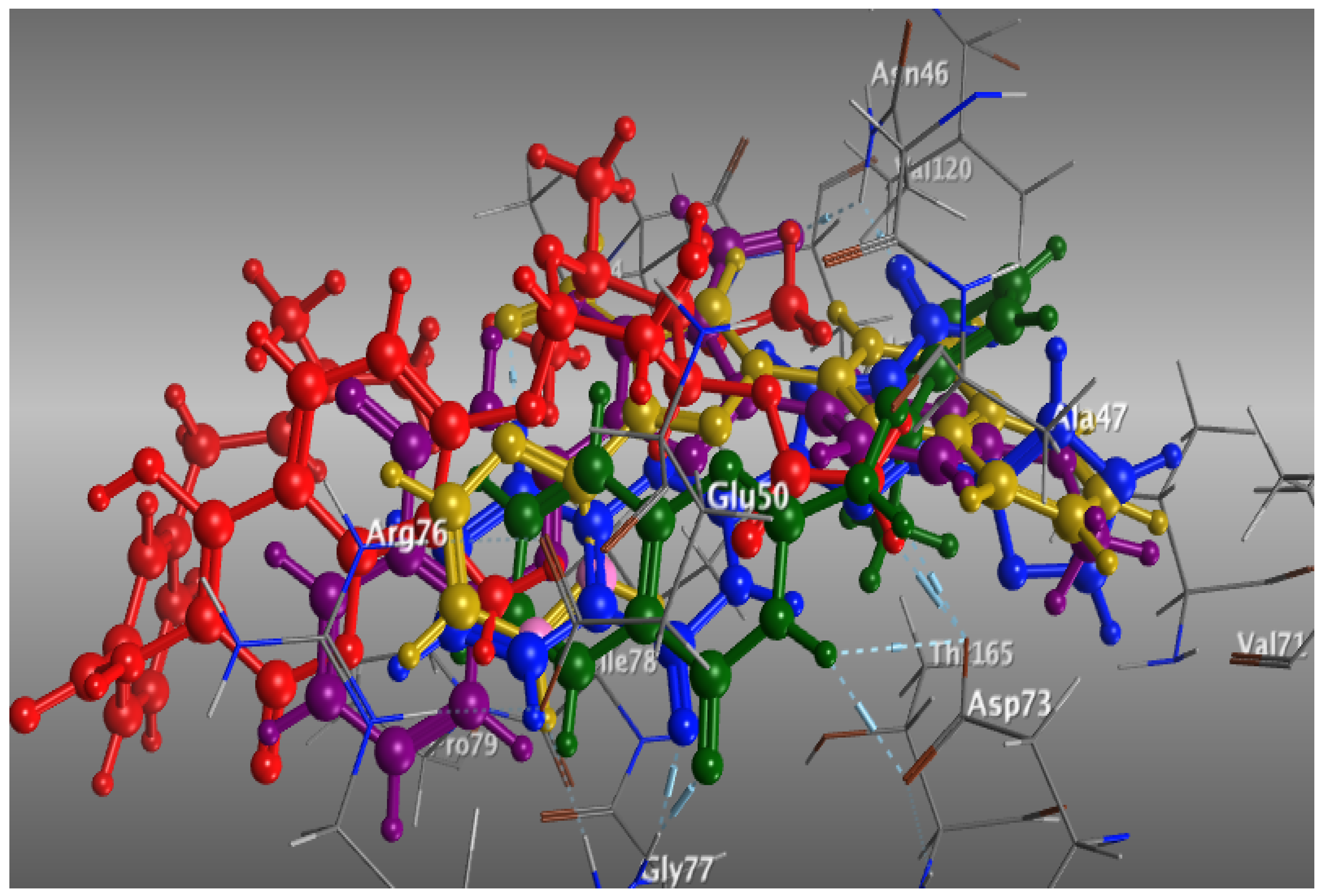
3.4. Molecular Docking Study on E. coli DNA Gyrase B Kinase
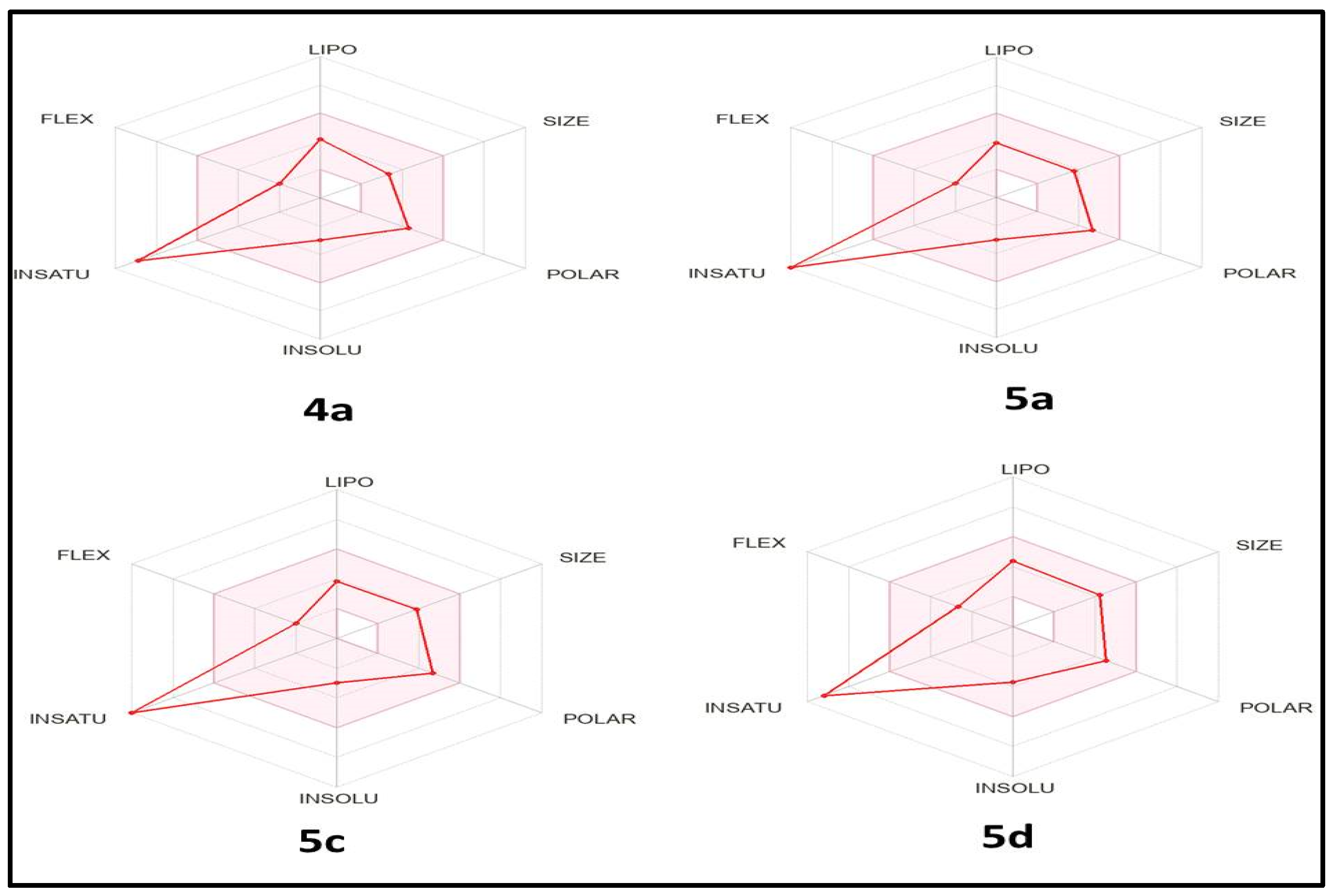
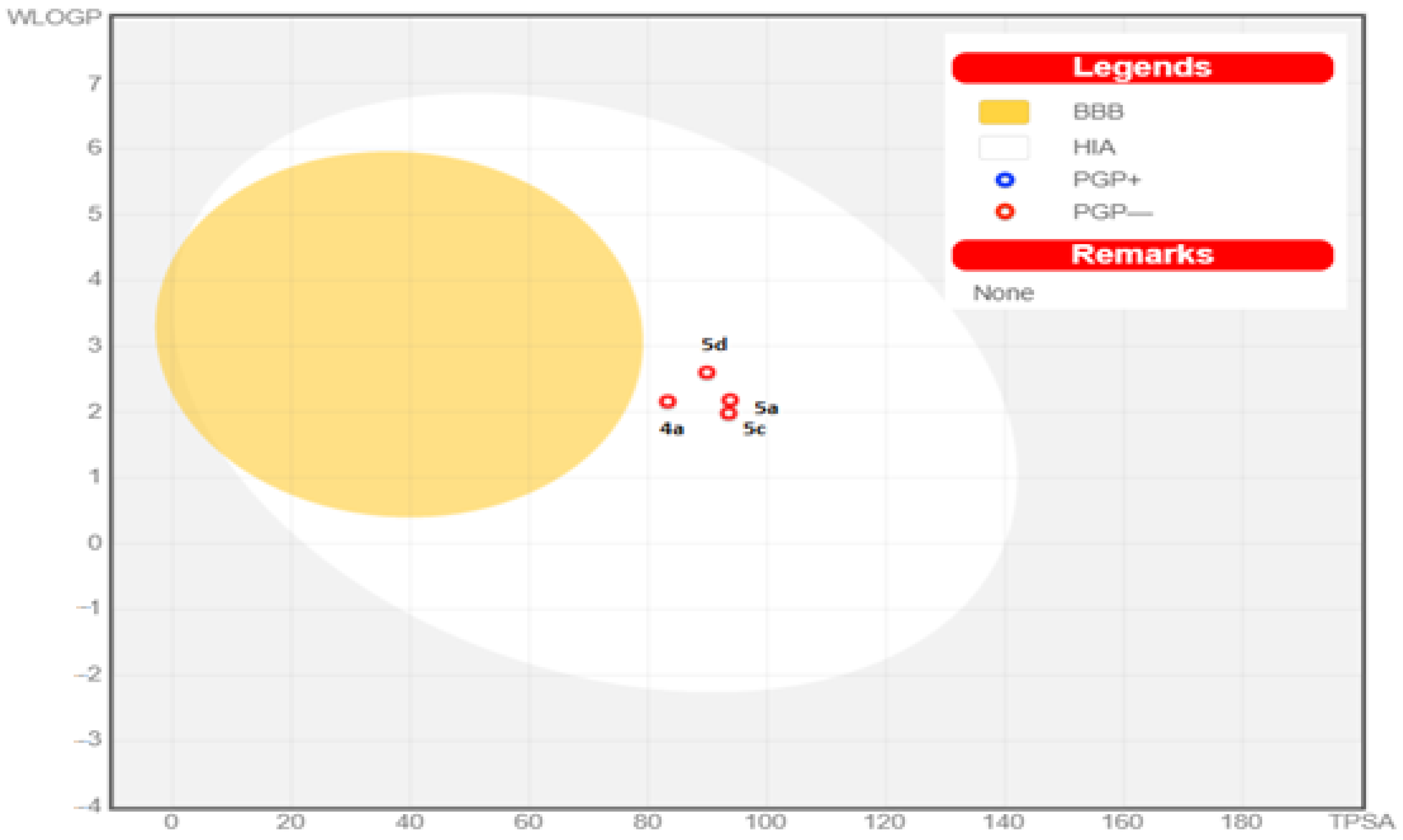
3.5. In Silico ADME Study
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sifri, Z.C.; Chokshi, A.; Cennimo, D.; Horng, H. Global Contributors to Antibiotic Resistance. J. Glob. Infect. Dis. 2019, 11, 36–42. [Google Scholar] [CrossRef] [PubMed]
- Lomazzi, M.; Moore, M.; Johnson, A.; Balasegaram, M.; Borisch, B. Antimicrobial resistance—Moving forward? BMC Public Health 2019, 19, 858. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Webste, T.J. Bacteria antibiotic resistance: New challenges and opportunities for implant-associated orthopedic infections. J. Orthop. Res. 2018, 36, 22–32. [Google Scholar] [CrossRef] [PubMed]
- Teitzel, G. Responding to Antimicrobial Resistance with Novel Therapeutics. Trends Microbiol. 2019, 27, 285–286. [Google Scholar] [CrossRef] [PubMed]
- Egorov, A.M.; Ulyashova, M.M.; Rubtsova, M.Y. Bacterial Enzymes and Antibiotic Resistance. Acta Naturae 2018, 10, 33–48. [Google Scholar] [CrossRef] [PubMed]
- Dubey, K.K.; Sharma, M. Reprogramming of antibiotics to combat antimicrobial resistance. Arch. Pharm. 2020, 353, e2000168. [Google Scholar] [CrossRef]
- Klostermeier, D. Why Two? On the Role of (A-)Symmetry in Negative Supercoiling of DNA by Gyrase. Int. J. Mol. Sci. 2018, 19, 1489. [Google Scholar] [CrossRef]
- Ma, Z.; Mduli, K. Mycobacterium tuberculosis DNA Gyrase as a Target for Drug Discovery. Infect. Disord. Drug Targets 2007, 7, 159–168. [Google Scholar] [CrossRef]
- Annunziato, G. Strategies to Overcome Antimicrobial Resistance (AMR) Making Use of Non-Essential Target Inhibitors: A Review. Int. J. Mol. Sci. 2019, 20, 5844. [Google Scholar] [CrossRef]
- Yi, L.; Lu, X. New Strategy on Antimicrobial-resistance: Inhibitors of DNA Replication Enzymes. Curr. Med. Chem. 2019, 26, 1761–1787. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Li, Q.; Su, M.; Luo, F.; Liu, Y.; Wang, D.; Fan, Y. Design, synthesis, and biological evaluation of novel 6-(pyridin-3-yl) quinazolin-4(3H)-one derivatives as potential anticancer agents via PI3K inhibition. Bioorg. Med. Chem. 2021, 46, 116346. [Google Scholar] [CrossRef] [PubMed]
- Mirgany, T.O.; Abdalla, A.N.; Arifuzzaman, M.; Motiur Rahman, A.F.M.; Al-Salem, H.S. Quinazolin-4(3H)-one based potential multiple tyrosine kinase inhibitors with excellent cytotoxicity. J. Enzyme Inhib Med. Chem. 2021, 36, 2055–2067. [Google Scholar] [CrossRef] [PubMed]
- Ramadan, S.K.; Elrazaz, E.Z.; Abouzid, K.A.M.; El-Naggar, A.M. Design, synthesis and in silico studies of new quinazolinone derivatives as antitumor PARP-1 inhibitors. RSC Adv. 2020, 10, 29475–29492. [Google Scholar] [CrossRef] [PubMed]
- Masri, A.; Anwar, A.; Khan, N.A.; Shahbaz, M.S.; Khan, K.M.; Shahabuddin, S.; Siddiqui, R. Antibacterial Effects of Quinazolin-4(3H)-One Functionalized-Conjugated Silver Nanoparticles. Antibiotics 2019, 8, 179. [Google Scholar] [CrossRef] [PubMed]
- Kavitha, K.; Mohan, S.; Srinivasan, N.; Suresh, R. Design, Synthesis, Characterization of Antimicrobial activity of Schiff bases of novel Quinazolin-4-one Derivatives. Res. J. Pharm. Tech. 2021, 14, 466–470. [Google Scholar] [CrossRef]
- Demirel, U.U.; Yilmaz, A.; Türkdağı, H.; Öztürk, B.; Arslan, U. Investigation of Some 3H-Quinazolin-4-One Derivatives in Vitro Antimicrobial Effect and Cytotoxicity on Human Gingival Fibroblasts. Polycycl. Aromat. Compd. 2021, 41, 1623–1630. [Google Scholar] [CrossRef]
- Thorat, D.B.; Shivkumar, B.; Rao, N.; Mohankumar, K.M. Synthesis and Evaluation of new 4(3H)-quinazolinones derivatives as potential Anti-Inflammatory agents. Asian J. Pharm. Sci. 2021, 11, 213–218. [Google Scholar] [CrossRef]
- Kumar, S.; Aghara, J.C.; Alex, A.T.; Aranjani, J.M.; Joesph, A. Novel Quinolone Substituted Quinazolin-4(3H)-Ones as Anti-Inflammatory, Anticancer Agents: Synthesis and Biological Screening. Indian J. Pharm. Educ. Res. 2018, 52, S268–S276. [Google Scholar] [CrossRef]
- Ayyad, R.A.; Sakr, H.M.; El-Gamal, K.M. Design, Synthesis, Computer Modeling and Analgesic Activity of Some New Disubstituted Quinazolin-4(3H)-ones. Med. Chem. 2016, 6, 299–305. [Google Scholar] [CrossRef]
- Osarumwense, P.; Edema, M.; Usifoh, C. Synthesis and Anagesic activities of Quinazolin-4(3H)-One, 2-Methyl-4(3H)-Quinazolinone and 2–Phenyl-4(3H)-quinazolin-4(3H)–one. J. Drug Deliv. Ther. 2020, 10, 87–91. [Google Scholar] [CrossRef]
- Birhan, Y.S.; Bekhit, A.A.; Hymete, A. In vivo antimalarial evaluation of some 2,3-disubstituted-4(3H)-quinazolinone derivatives. BMC Res. Notes 2015, 8, 589. [Google Scholar] [CrossRef]
- Bule, M.H.; Haymete, A.; Kefale, B. Synthesis and In-Vivo Pharmacological Evaluation of Some Novel 4(3H)- Quinazolinone Derivatives as Potential Anti-malarial Agents. Drug. Des. 2015, 4, 121. [Google Scholar] [CrossRef]
- Chen, L.; Wang, X.; Tang, X.; Xia, R.; Guo, T.; Zhang, C.; Li, X.; Xue, W. Design, synthesis, antiviral bioactivities and interaction mechanisms of penta-1,4-diene-3-one oxime ether derivatives containing a quinazolin-4(3H)-one scaffold. BMC Chem. 2019, 13, 34. [Google Scholar] [CrossRef]
- Tran, T.P.; Ellsworth, E.L.; Sanchez, J.P.; Watson, B.M.; Stier, M.A.; Showalter, H.D.H.; Domagala, J.M.; Shapiro, M.A.; Joannides, E.T.; Gracheck, S.J.; et al. Structure–activity relationships of 3-aminoquinazolinediones, a new class of bacterial type-2 topoisomerase (DNA gyrase and topo IV) inhibitors. Bioorg. Med. Chem. Lett. 2007, 17, 1312. [Google Scholar] [CrossRef]
- Ghorab, M.M.; Alqahtani, A.S.; Soliman, A.M.; Askar, A.A. Novel N-(Substituted) Thioacetamide Quinazolinone Benzenesulfonamides as Antimicrobial Agents. Int. J. Nanomed. 2020, 15, 3161–3180. [Google Scholar] [CrossRef]
- Xue, W.; Wang, Y.; Lian, X.; Li, X.; Pang, J.; Kirchmair, J.; Wu, K.; Han, Z.; You, X.; Zhang, H.; et al. Discovery of N-quinazolinone-4-hydroxy-2-quinolone-3-carboxamides as DNA gyrase B-targeted antibacterial agents. J. Enzyme Inhib. Med. Chem. 2022, 37, 1620–1631. [Google Scholar] [CrossRef]
- Verma, G.; Marella, A.; Shaquiquzzaman, M.; Akhtar, M.; Ali, M.R.; Alam, M.M. A review exploring biological activities of hydrazones. J. Pharm. Bioallied Sci. 2014, 6, 69–80. [Google Scholar] [CrossRef]
- Rollas, S.; Küçükgüzel, S.G. Biological Activities of Hydrazone Derivatives. Molecules 2007, 12, 1910–1939. [Google Scholar] [CrossRef]
- Ansari, A.; Ali, A.; Asif, A. Shamsuzzman Review: Biologically active pyrazole derivatives. New J. Chem. 2017, 41, 16–41. [Google Scholar] [CrossRef]
- Karrouchi, K.; Radi, S.; Ramli, Y.; Taoufik, J.; Mabkhot, Y.N.; Al-aizari, F.A.; Ansar, M. Synthesis and Pharmacological Activities of Pyrazole Derivatives: A Review. Molecules 2018, 23, 134. [Google Scholar] [CrossRef]
- Sridhar, P.; Alagumuthu, M.; Arumugam, S.; Reddy, S.R. Synthesis of quinoline acetohydrazide-hydrazone derivatives evaluated as DNA gyrase inhibitors and potent antimicrobial agents. RSC Adv. 2016, 6, 64460–64468. [Google Scholar] [CrossRef]
- Ragab, A.; Elsisi, D.M.; Abu Ali, O.A.; Abusaif, M.S.; Askar, A.A.; Farag, A.A.; Ammar, Y.A. Design, synthesis of new novel quinoxalin-2(1H)-one derivatives incorporating hydrazone, hydrazine, and pyrazole moieties as antimicrobial potential with in-silico ADME and molecular docking simulation. Arab. J. Chem. 2022, 15, 103497. [Google Scholar] [CrossRef]
- Hassan, A.S.; Askar, A.A.; Naglah, A.M.; Almehizia, A.A.; Ragab, A. Discovery of New Schiff Bases Tethered Pyrazole Moiety: Design, Synthesis, Biological Evaluation, and Molecular Docking Study as Dual Targeting DHFR/DNA Gyrase Inhibitors with Immunomodulatory Activity. Molecules 2020, 25, 2593. [Google Scholar] [CrossRef]
- Sun, J.; Lv, P.C.; Yin, Y.; Yuan, R.J.; Ma, J.; Zhu, H.L. Synthesis, structure and antibacterial activity of potent DNA gyrase inhibitors: N’-benzoyl-3-(4-bromophenyl)-1H-pyrazole-5-carbohydrazide derivatives. PLoS ONE 2013, 8, e69751. [Google Scholar] [CrossRef]
- Viegas-Junior, C.; Danuello, A.; da Silva Bolzani, V.; Barreiro, E.J.; Fraga, C.A. Molecular hybridization: A useful tool in the design of new drug prototypes. Curr. Med. Chem. 2007, 14, 1829–1852. [Google Scholar] [CrossRef] [PubMed]
- Sree, B.S.; Rao, A.L.; Babu, P.S.; Kiran, K.S.; Vandana, K. Synthesis and antimicrobial screening of some novel quinazolinones and its derivatives. Indian J. Pharm. Pharmacol. 2017, 4, 130–133. [Google Scholar] [CrossRef]
- Gruner, M.; Rehwald, M.; Katrin Eckert, K.; Gewald, K. New Syntheses of 2-Alkylthio-4-Oxo-3,4-Dihydroquinazolines, 2-Alkylthioquinazolines, as Well as Their Hetero Analogues. Heterocycles 2000, 53, 2363–2377. [Google Scholar] [CrossRef]
- Penna, C.A.; Marino, S.G.; Gutkind, G.O.; Clavin, M.; Ferraro, G.; Martino, V. Antimicrobial activity of Eupatorium species growing in Argentina. J. Herbs Spices Med. Plants 1988, 5, 21–28. [Google Scholar] [CrossRef]
- Wiegand, I.; Hilpert, K.; Hancock, R.E. Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat. Protoc. 2008, 3, 163–175. [Google Scholar] [CrossRef]
- Phillips, J.W.; Goetz, M.A.; Smith, S.K.; Zink, D.L.; Polishook, J.; Onishi, R.; Salowe, S.; Wiltsie, J.; Allocco, J.; Sigmund, J.; et al. Discovery of Kibdelomycin, A Potent New Class of Bacterial Type II Topoisomerase Inhibitor by Chemical-Genetic Profiling in Staphylococcus aureus. Chem. Biol. 2011, 18, 955–965. [Google Scholar] [CrossRef]
- Maxwell, A.; Burton, N.P.; O’Hagan, N. High-throughput assays for DNA gyrase and other topoisomerases. Nucleic Acids Res. 2006, 34, e104. [Google Scholar] [CrossRef]
- Mohi El-Deen, E.M.; Abd El-Meguid, E.A.; Karam, E.A.; Nossier, E.S.; Ahmed, M.F. Synthesis and Biological Evaluation of New Pyridothienopyrimidine Derivatives as Antibacterial Agents and Escherichia coli Topoisomerase II Inhibitors. Antibiotics 2020, 9, 695. [Google Scholar] [CrossRef]
- Hashem, H.E.; Amr, A.E.G.E.; Nossier, E.S.; Elsayed, E.A.; Azmy, E.M. Synthesis, antimicrobial activity and molecular docking of novel thiourea derivatives tagged with thiadiazole, imidazole and triazine moieties as potential DNA gyrase and topoisomerase IV inhibitors. Molecules 2020, 25, 2766. [Google Scholar] [CrossRef]
- Mohi El-Deen, E.M.; El-Meguid, E.A.A.; Hasabelnaby, S.; Karam, E.A.; Nossier, E.S.; El-Deen, E.M.; El-Meguid, E.A.A. Synthesis, Docking Studies, and In Vitro Evaluation of Some Novel Thienopyridines and Fused Thienopyridine-Quinolines as Antibacterial Agents and DNA Gyrase Inhibitors. Molecules 2019, 24, 3650. [Google Scholar] [CrossRef]
- Daina, A.; Michielin, O.; Zoete, V. SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 2017, 7, 42717. [Google Scholar] [CrossRef]
- Othman, I.M.; Alamshany, Z.M.; Tashkandi, N.Y.; Gad-Elkareem, M.A.; Anwar, M.M.; Nossier, E.S. New pyrimidine and pyrazole-based compounds as potential EGFR inhibitors: Synthesis, anticancer, antimicrobial evaluation and computational studies. Bioorg. Chem. 2021, 114, 105078. [Google Scholar] [CrossRef]
- Mohi El-Deen, E.M.; Anwar, M.M.; Abd El-Gwaad, A.A.; Karam, E.A.; El-Ashrey, M.K.; Kassab, R.R. Design and synthesis of some novel pyridothienopyrimidine derivatives and their biological evaluation as antimicrobial and anticancer agents targeting EGFR enzyme. Arab. J. Chem. 2022, 15, 103751. [Google Scholar] [CrossRef]
- El-Tombary, A.A.; Ismail, K.A.; Aboulwafa, O.M.; Omar, A.M.M.; El-Azzouni, M.Z.; El-Mansoury, S.T. Novel triazolo [4, 3-a] quinazolinone and bis-triazolo [4, 3-a: 4, 3′-c] quinazolines: Synthesis and antitoxoplasmosis effect. Farmaco 1999, 54, 486–495. [Google Scholar] [CrossRef]
- Rafeeq, M.; Reddy, B.S.; Reddy, C.V.R.; Naidu, A.; Dubey, P.K. Green and efficient synthesis of 2-(4-oxo-3, 4-dihydroquinazolin-2-yl)-2, 3-dihydropthalazine-1, 4-dione. Indian J. Chem. 2015, 54B, 412–417. [Google Scholar] [CrossRef]
- Singh, K.; Ralhan, S.; Sharma, P.K.; Dhawan, S.N. Vilsmeier–Haack Reaction on Hydrazones: A Convenient Synthesis of 4-formylpyrazoles. J. Chem. Res. 2005, 2005, 316–318. [Google Scholar] [CrossRef]
| Antibacterial Activity | Antifungal Activity | |||||||
|---|---|---|---|---|---|---|---|---|
| Compd. | S. aureus | B. subtilis | E. coli | S. typhimurium | C. albicans | C. tropicals | M. phaseolina | A. niger |
| 2 | 25 | 22 | 20 | 14 | 20 | 20 | 19 | 19 |
| 4a | 30 | 29 | 30 | 27 | 32 | 25 | 27 | 25 |
| 4b | 26 | 24 | 26 | 26 | 20 | 18 | 16 | 26 |
| 4c | 29 | 28 | 28 | 29 | 30 | 32 | 29 | 30 |
| 4d | 17 | 18 | 20 | 16 | 15 | 14 | 15 | 18 |
| 4e | 24 | 22 | 21 | 22 | 20 | 22 | 22 | 20 |
| 4f | 15 | 17 | 16 | - | - | 14 | - | 17 |
| 5a | 35 | 34 | 35 | 38 | 39 | 35 | 30 | 29 |
| 5b | 26 | 24 | 26 | 26 | 16 | 21 | 20 | 19 |
| 5c | 30 | 30 | 33 | 31 | 27 | 28 | 24 | 22 |
| 5d | 31 | 29 | 34 | 26 | 28 | 29 | 25 | 27 |
| Amoxicillin | 29 | 28 | 28 | 27 | - | - | - | - |
| Clotrimazole | - | - | - | - | 26 | 25 | 24 | 26 |
| Antibacterial Activity | Antifungal Activity | |||||||
|---|---|---|---|---|---|---|---|---|
| Compd. | S. aureus | B. subtilis | E. coli | S. typhimurium | C. albicans | C. tropicals | M. phaseolina | A. niger |
| 2 | 16 | 32 | 64 | 128 | 32 | 32 | 64 | 32 |
| 4a | 4 | 4 | 4 | 8 | 2 | 8 | 8 | 16 |
| 4b | 8 | 8 | 8 | 8 | 32 | 64 | 128 | 16 |
| 4c | 4 | 8 | 8 | 4 | 2 | 4 | 8 | 4 |
| 4d | 64 | 64 | 64 | 64 | 128 | 128 | 128 | 128 |
| 4e | 16 | 32 | 32 | 16 | 32 | 16 | 16 | 32 |
| 4f | 128 | 64 | 128 | - | - | 128 | - | 64 |
| 5a | 2 | 2 | 1 | 1 | 1 | 2 | 4 | 4 |
| 5b | 4 | 8 | 8 | 8 | 32 | 32 | 32 | 32 |
| 5c | 4 | 4 | 2 | 4 | 8 | 8 | 16 | 16 |
| 5d | 4 | 4 | 2 | 8 | 8 | 8 | 16 | 8 |
| Amoxicillin | 4 | 4 | 8 | 8 | - | - | - | - |
| Clotrimazole | - | - | - | - | 8 | 8 | 16 | 8 |
| Compound No. | DNA Gyrase Supercoiling Inhibition IC50 (µM) |
|---|---|
| 4a | 4.17 ± 0.07 |
| 4b | 16.91 ± 0.36 |
| 4c | 15.08 ± 0.34 |
| 5a | 3.19 ± 0.06 |
| 5b | 10.82 ± 0.24 |
| 5c | 4.09 ± 0.13 |
| 5d | 3.51 ± 0.10 |
| Novobiocin | 4.12 ± 0.11 |
| Compd. | Docking Score (Kcal/mol) | Amino Acid Residues (Bond Length A°) | Atoms of Compound | Type of Bond |
|---|---|---|---|---|
| Novobiocin | −10.25 | Asn46(3.27); | H(OH)(oxan-4-yl); | H-don |
| Asp73(1.91); | H(OCONH2); | H-don | ||
| Arg76 | C6H2(coumarin) | Arene–cation | ||
| 4a | −10.48 | Asn46; | furan; | Arene–cation |
| Asp73(3.08,3.16); | NH(quinazolinone) | H-don | ||
| Asp73(2.74); | NH(hydrazinyl) | H-don | ||
| Gly77(2.63); | O(quinazolinone) | H-acc | ||
| Ile78 | Quinazolinone | Arene–cation | ||
| 4b | −8.70 | Gly77(2.66); | O(quinazolinone) | H-acc |
| Ile78 | Quinazolinone | Arene–cation | ||
| 4c | −8.66 | Gly77(2.83); | O(quinazolinone) | H-acc |
| Ile78 | Quinazolinone | Arene–cation | ||
| 5a | −10.97 | Asn46(2.53); | O(CHO) | H-acc |
| Gly77(2.68); | O(quinazolinone) | H-acc | ||
| Ile78 | Quinazolinone | Arene–cation | ||
| 5b | −8.80 | Gly77(2.51); | O(quinazolinone) | H-acc |
| Ile78 | Quinazolinone | Arene–cation | ||
| 5c | −9.89 | Asn46; | Pyrazole | Arene–cation |
| Ile78; | Pyrazole | Arene–cation | ||
| Ile78 | Quinazolinone | Arene–cation | ||
| 5d | −10.74 | Asn46(2.44); | O(CHO) | H-acc |
| Asn46; | 4-methoxyphenyl | Arene–cation | ||
| Ile78; | Pyrazole | Arene–cation | ||
| Ile78 | Quinazolinone | Arene–cation |
| Compd. | MW 1 | nHBD 2 | nHBA 3 | nRB 4 | MLogP 5 | TPSA (Å2) 6 | Violations 7 |
|---|---|---|---|---|---|---|---|
| 4a | 268.27 | 2 | 4 | 3 | 1.58 | 83.28 | 0 |
| 5a | 306.28 | 1 | 5 | 3 | 1.38 | 93.78 | 0 |
| 5c | 317.30 | 1 | 5 | 3 | 1.58 | 93.53 | 0 |
| 5d | 346.34 | 1 | 5 | 4 | 2.05 | 89.87 | 0 |
| Comp. No. | GIT Absorption | BBB Permeability | P-gp Substrate | Bioavailability Score | PAINS Alert |
|---|---|---|---|---|---|
| 4a | High | NO | NO | 0.55 | 0 |
| 5a | High | NO | NO | 0.55 | 0 |
| 5c | High | NO | NO | 0.55 | 0 |
| 5d | High | NO | NO | 0.55 | 0 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mohi El-Deen, E.M.; Nossier, E.S.; Karam, E.A. New Quinazolin-4(3H)-one Derivatives Incorporating Hydrazone and Pyrazole Scaffolds as Antimicrobial Agents Targeting DNA Gyraze Enzyme. Sci. Pharm. 2022, 90, 52. https://doi.org/10.3390/scipharm90030052
Mohi El-Deen EM, Nossier ES, Karam EA. New Quinazolin-4(3H)-one Derivatives Incorporating Hydrazone and Pyrazole Scaffolds as Antimicrobial Agents Targeting DNA Gyraze Enzyme. Scientia Pharmaceutica. 2022; 90(3):52. https://doi.org/10.3390/scipharm90030052
Chicago/Turabian StyleMohi El-Deen, Eman M., Eman S. Nossier, and Eman A. Karam. 2022. "New Quinazolin-4(3H)-one Derivatives Incorporating Hydrazone and Pyrazole Scaffolds as Antimicrobial Agents Targeting DNA Gyraze Enzyme" Scientia Pharmaceutica 90, no. 3: 52. https://doi.org/10.3390/scipharm90030052
APA StyleMohi El-Deen, E. M., Nossier, E. S., & Karam, E. A. (2022). New Quinazolin-4(3H)-one Derivatives Incorporating Hydrazone and Pyrazole Scaffolds as Antimicrobial Agents Targeting DNA Gyraze Enzyme. Scientia Pharmaceutica, 90(3), 52. https://doi.org/10.3390/scipharm90030052







