Synthesis and Evaluation of (1,4-Disubstituted)-1,2,3-triazoles as Estrogen Receptor Beta Agonists
Abstract
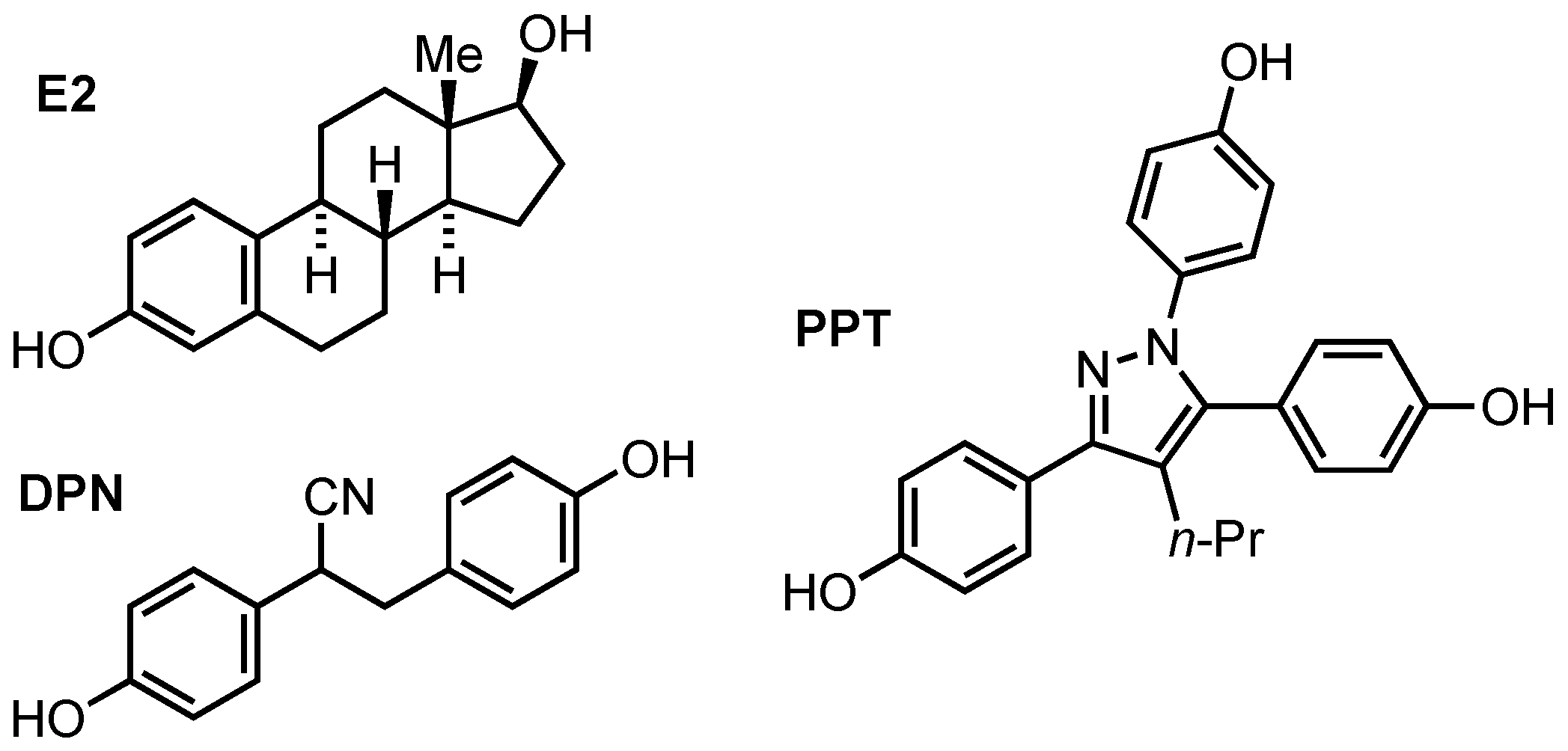
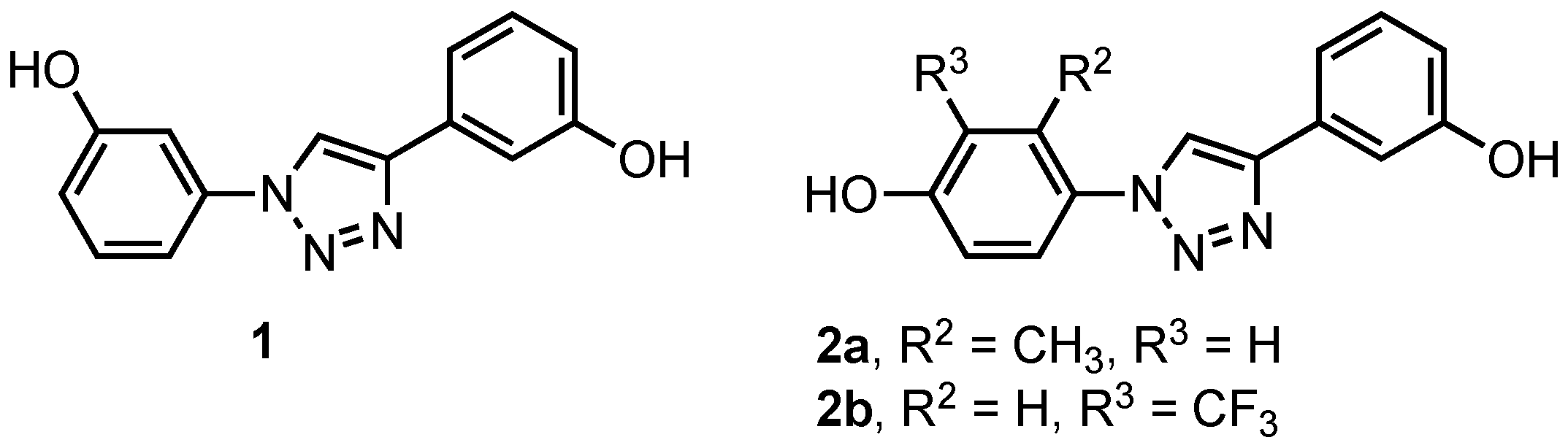
1. Introduction
2. Materials and Methods
2.1. Chemistry
2.1.1. General Experimental Outline
2.1.2. General Procedure for the Preparation of Azidophenols
2.1.3. General Procedure for Triazole Synthesis
2.2. ERβ Assays
2.2.1. TR-FRET Assay
2.2.2. Cell-Based Functional Assay
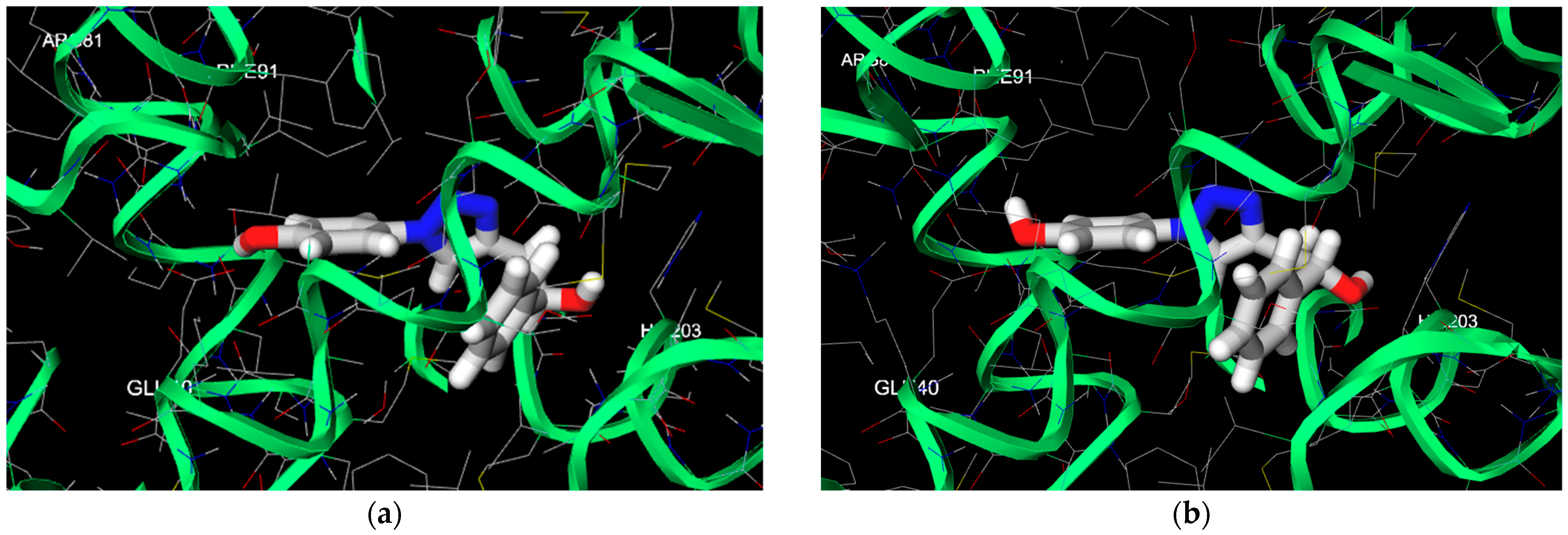
2.3. Computational
3. Results
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Dahlman-Wright, K.; Cavailles, V.; Fuqua, S.A.; Jordan, V.C.; Katzenellenbogen, J.A.; Korach, K.S.; Maggi, A.; Muramatsu, M.; Parker, M.G.; Gustafsson, J.A. International Union of Pharmacology, LXIV. Estrogen receptors. Pharmacol. Rev. 2006, 58, 773–781. [Google Scholar] [CrossRef] [PubMed]
- Lane, P.H. Estrogen receptors in the kidney: Lessons from genetically altered mice. Gend. Med. 2008, 5, S11–S18. [Google Scholar] [CrossRef]
- Weiser, M.J.; Foradori, C.D.; Handa, R.J. Estrogen receptor beta in the brain: From form to function. Brain Res. Rev. 2008, 5, 309–320. [Google Scholar] [CrossRef] [PubMed]
- Sengupta, S.; Jordan, V.C. Novel selective estrogen receptor modulators. In Estrogen Action, Selective Estrogen Receptor Modulators and Women’s Health; Imperial College Press: London, UK, 2013; pp. 325–359. [Google Scholar]
- Farzaheh, S.; Zarghi, A. Estrogen Receptor Ligands: A Review (2013–2015). Sci. Pharm. 2016, 84, 409–427. [Google Scholar] [CrossRef]
- Stauffer, S.R.; Coletta, C.J.; Tedesco, R.; Nishiguchi, G.; Carlson, K.; Sun, J.; Katzenellenbogen, B.S.; Katzenellenbogen, J.A. Pyrazole Ligands: Structure-Affinity/Activity Relationships and Estrogen Receptor-α-Selective Agonists. J. Med. Chem. 2000, 43, 4934–4947. [Google Scholar] [CrossRef] [PubMed]
- Meyers, M.J.; Sun, J.; Carlson, K.E.; Marriner, G.A.; Katzenellenbogen, B.S.; Katzenellenbogen, J.A. Estrogen Receptor-β Potency-Selective Ligands: Structure-Activity Relationship Studies of Diarylpropionitriles and Their Acetylene and Polar Analogues. J. Med. Chem. 2001, 44, 4230–4251. [Google Scholar] [CrossRef] [PubMed]
- Minutolo, F.; Macchia, M.; Katzenellenbogen, B.S.; Katzenellenbogen, J.A. Estrogen receptor β ligands: Recent advances and biomedical applications. Med. Res. Rev. 2011, 31, 364–442. [Google Scholar] [CrossRef] [PubMed]
- Tornoe, C.W.; Christensen, C.; Meldal, M. Peptidotriazoles on solid phase; [1,2,3]-Triazole by regiospecific copper (I)-catalyzed 1,3-dipolar cycloaddition of terminal alkynes to azide. J. Org. Chem. 2002, 67, 3057–3064. [Google Scholar] [CrossRef]
- Rostovtsev, V.; Green, L.G.; Fokin, V.V.; Sharpless, K.B. A stepwise Huisgen cycloaddition process: Copper (I) catalyzed regioselective “ligation” off azide and terminal alkynes. Angew. Chem. Int. Ed. 2002, 41, 2596–2599. [Google Scholar] [CrossRef]
- Agalave, S.G.; Maujan, S.R.; Pore, V.S. Click Chemistry: 1,2,3-Triazoles as Pharmacophores. Chem. Asian J. 2011, 6, 2696–2718. [Google Scholar] [CrossRef]
- Kharb, R.; Sharma, P.C.; Yar, M.S. Pharmacological significance of triazole scaffold. J. Enzyme Inhib. Med. Chem. 2011, 26, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Massarotti, A.; Aprile, S.; Mercalli, V.; Del Grosso, E.; Grosa, G.; Sorba, G.; Tron, G.C. Are 1,4- and 1,5-Disubstituted 1,2,3-Triazoles Good Pharmacophoric Groups? ChemMedChem 2014, 9, 2497–2508. [Google Scholar] [CrossRef] [PubMed]
- Dheer, D.; Singh, V.; Shankar, R. Medicinal attributes of 1,2,3-triazoles: Current developments. Bioorg. Chem. 2017, 71, 30–54. [Google Scholar] [CrossRef] [PubMed]
- Bozorov, K.; Zhao, J.; Aisa, H.A. 1,2,3-Triazole-containing hybrids as leads in medicinal chemistry: A recent overview. Bioorg. Med. Chem. 2019, 27, 3511–3531. [Google Scholar] [CrossRef] [PubMed]
- Rani, A.; Singh, G.; Singh, A.; Maqbool, U.; Kaur, G.; Singh, J. CuAAC-ensembled 1,2,3-triazole-linked isosteres as pharmacophores in drug discovery: Review. RSC Adv. 2020, 10, 5610–5635. [Google Scholar] [CrossRef]
- Dalvie, D.K.; Kalgutkar, A.S.; Khojasteh-Bakht, S.C.; Obach, R.S.; O’Donnell, J.P. Biotransformation Reactions of Five-Membered Aromatic Heterocyclic Rings. Chem. Res. Toxicol. 2002, 15, 269–299. [Google Scholar] [CrossRef]
- Pirali, T.; Gatti, S.; Di Brisco, R.; Tacchi, S.; Zaninetti, R.; Brunelli, E.; Massarotti, A.; Sorba, G.; Canonico, P.L.; Moro, L.; et al. Estrogenic Analogs Synthesized by Click Chemistry. ChemMedChem 2007, 2, 437–440. [Google Scholar] [CrossRef]
- Demkowicz, S.; Filipiak, K.; Maslyk, M.; Ciepielski, J.; de Pascual-Teresa, S.; Martin-Santamaria, S.; de Pascual-Teresa, B.; Ramos, A. New clicked full agonists of the estrogen receptor β. RSC Adv. 2013, 3, 3697–3706. [Google Scholar] [CrossRef]
- McCullough, C.M.; Neumann, T.S.; Gone, J.R.; He, Z.; Herrild, C.; Wondergem, J.; Pandey, R.K.; Donaldson, W.A.; Sem, D.S. Probing the human estrogen receptor-a binding requirements for phenolic mono- and di-hydroxyl compounds: A combined synthesis, binding and docking study. Bioorg. Med. Chem. 2014, 22, 303–310. [Google Scholar] [CrossRef][Green Version]
- Hanson, A.M.; Perera, K.L.I.S.; Kim, J.; Pandey, R.K.; Sweeney, N.; Lu, X.; Imhoff, A.; Mackinnon, A.C.; Wargolet, A.J.; Van Hart, R.M.; et al. A–C Estrogens as Potent and Selective Estrogen Receptor-Beta Agonists (SERBAs) to Enhance Memory Consolidation under Low-Estrogen Conditions. J. Med. Chem. 2018, 61, 4720–4738. [Google Scholar] [CrossRef]
- Perera, K.L.I.S.; Hanson, A.M.; Lindeman, S.; Imhoff, A.; Lu, X.; Sem, D.S.; Donaldson, W.A. Synthesis and evaluation of 4-cycloheptylphenols as selective Estrogen receptor-b agonists (SERBAs). Eur. J. Med. Chem. 2018, 157, 791–804. [Google Scholar] [CrossRef] [PubMed]
- Wetzel, E.A.; Hanson, A.M.; Troutfetter, C.L.; Burkett, D.J.; Sem, D.S.; Donaldson, W.A. Synthesis and evaluation of 17a-triazolyl and 9a-cyano derivatives of estradiol. Bioorg. Med. Chem. 2020, 28, 115670. [Google Scholar]
- Fleischer, A.W.; Schalk, J.C.; Wetzel, E.A.; Hanson, A.M.; Sem, D.S.; Donaldson, W.A.; Frick, K.M. Long-term oral administration of a novel estrogen receptor beta agonists enhances memory and alleviates drug-induced vasodilation in young ovariectomized mice. Horm. Behav. 2021, 130, 104948. [Google Scholar] [CrossRef] [PubMed]
- Mohammed, S.; Padala, A.K.; Dar, B.A.; Sing, B.; Sreedhar, B.; Vishwakarma, R.A.; Bharate, S.B. Recyclable clay supported Cu (II) catalyzed tandem one-pot synthesis of 1-aryl-1,2,3-traizoles. Tetrahedron 2012, 68, 8156–8162. [Google Scholar] [CrossRef]
- Shang, J.-Q.; Fu, H.; Li, Y.; Yang, T.; Gao, C.; Li, Y.-M. Copper-catalyzed decarboxylation/cycloaddition cascade of alkynyl carboxylic acids with azide. Tetrahedron 2019, 75, 253–259. [Google Scholar] [CrossRef]
- Xiao, Z.; Fokkens, M.; Chen, D.; Kok, T.; Proietti, G.; van Merkerk, R.; Poelarends, G.J.; Dekker, F.J. Structure-activity relationships for binding of 4-substituted triazole-phenols to macrophage migration inhibitory factor (MIF). Eur. J. Med. Chem. 2020, 186, 111849. [Google Scholar] [CrossRef] [PubMed]
- Shiau, A.K.; Barstad, D.; Radek, J.T.; Meyers, M.J.; Nettles, K.W.; Katzenellenbogen, B.S.; Katzenellenbogen, J.A.; Agard, D.A.; Greene, G.L. Structural characterization of a subtype-selective ligand reveals a novel mode of estrogen receptor antagonism. Nat. Struct. Biol. 2002, 9, 359–364. [Google Scholar]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization and multithreading. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef]
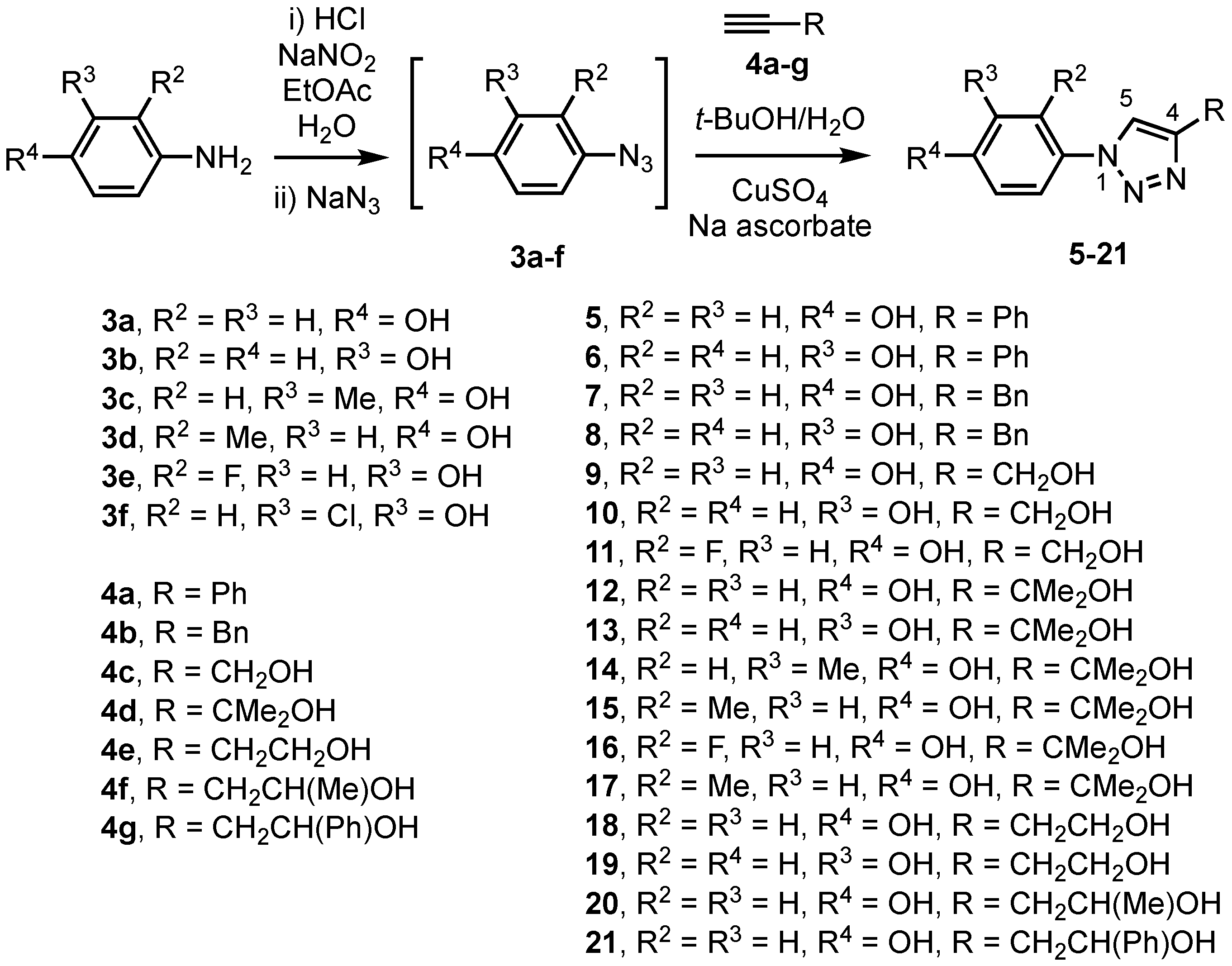
| Compound | Yield | ERβ EC50 (μM) | Volume (calc. Å3) 1 | O–O Distance 2 | Docking Score 3 | |
|---|---|---|---|---|---|---|
| 5 | 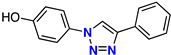 | 70% | 5.53 | 212 | – | −7.6 |
| 6 | 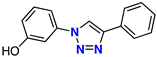 | 68% | ND | 212 | – | −7.9 |
| 7 | 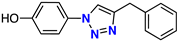 | 62% | 9.04 | 229 | – | −7.9 |
| 8 | 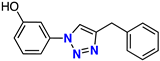 | 48% | 25.8 | 229 | – | −8.0 |
| 9 | 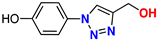 | 90% | 44.6 | 165 | 10.1 Å | −6.6 |
| 10 | 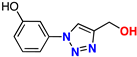 | 78% | 4.28 | 165 | 9.1 Å | −7.9 |
| 11 | 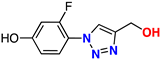 | 67% | 9.69 | 170 | 10.2 Å | −7.4 |
| 12 | 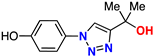 | 74% | >50 | 198 | 9.3 Å | −6.6 |
| 13 | 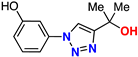 | 69% | 9.15 | 198 | 9.2 Å | −7.3 |
| 14 | 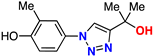 | 80% | 48.1 | 215 | 9.3 Å | −6.6 |
| 15 | 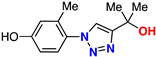 | 81% | ND | 215 | 9.4 Å | −7.0 |
| 16 | 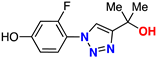 | 55% | 18.2 | 203 | 9.3 Å | −7.2 |
| 17 | 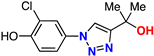 | 76% | >50 | 212 | 9.4 Å | −6.8 |
| 18 | 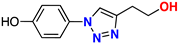 | 78% | 18.2 | 182 | 10.8 Å | −7.2 |
| 19 | 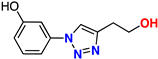 | 82% | >50 | 179 | 10.6 Å | −7.0 |
| (±)-20 | 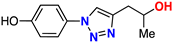 | 62% | 13.6 | 199 | 11.1 Å | −7.7 |
| (±)-21 | 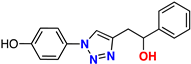 | 69% | 1.59 | 254 | 9.2–10.9 Å | −9.3 (R) −9.3 (S) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wetzel, E.A.; Corriero, G.C.; Brown-Ford, S.; Sem, D.S.; Donaldson, W.A. Synthesis and Evaluation of (1,4-Disubstituted)-1,2,3-triazoles as Estrogen Receptor Beta Agonists. Sci. Pharm. 2022, 90, 46. https://doi.org/10.3390/scipharm90030046
Wetzel EA, Corriero GC, Brown-Ford S, Sem DS, Donaldson WA. Synthesis and Evaluation of (1,4-Disubstituted)-1,2,3-triazoles as Estrogen Receptor Beta Agonists. Scientia Pharmaceutica. 2022; 90(3):46. https://doi.org/10.3390/scipharm90030046
Chicago/Turabian StyleWetzel, Edward A., Grace C. Corriero, Sandra Brown-Ford, Daniel S. Sem, and William A. Donaldson. 2022. "Synthesis and Evaluation of (1,4-Disubstituted)-1,2,3-triazoles as Estrogen Receptor Beta Agonists" Scientia Pharmaceutica 90, no. 3: 46. https://doi.org/10.3390/scipharm90030046
APA StyleWetzel, E. A., Corriero, G. C., Brown-Ford, S., Sem, D. S., & Donaldson, W. A. (2022). Synthesis and Evaluation of (1,4-Disubstituted)-1,2,3-triazoles as Estrogen Receptor Beta Agonists. Scientia Pharmaceutica, 90(3), 46. https://doi.org/10.3390/scipharm90030046







