Design, Synthesis and In Vitro Antimicrobial Activity of 6-(1H-Benzimidazol-2-yl)-3,5-dimethyl-4-oxo-2-thio-3,4-dihydrothieno[2,3-d]pyrimidines
Abstract
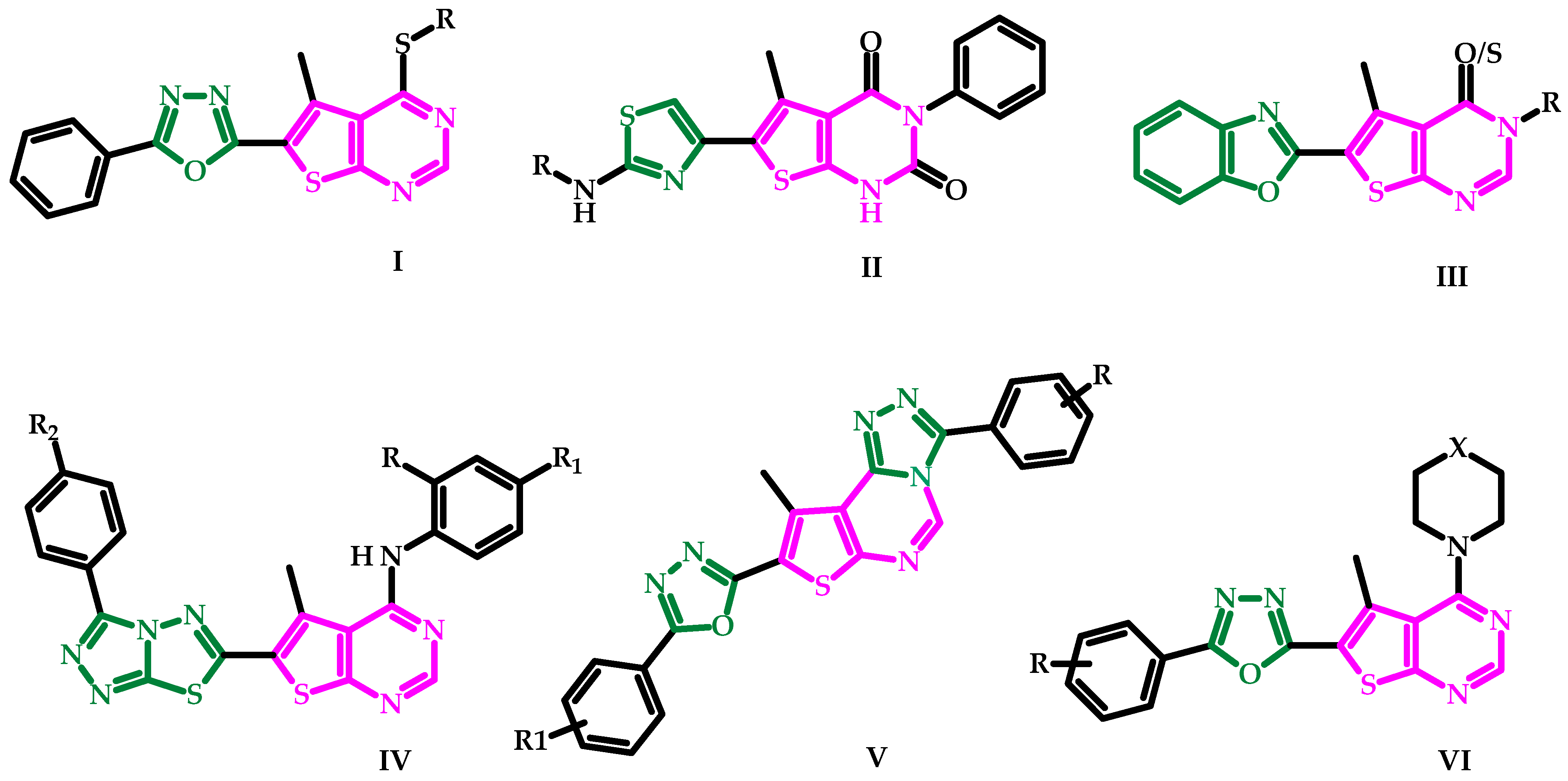
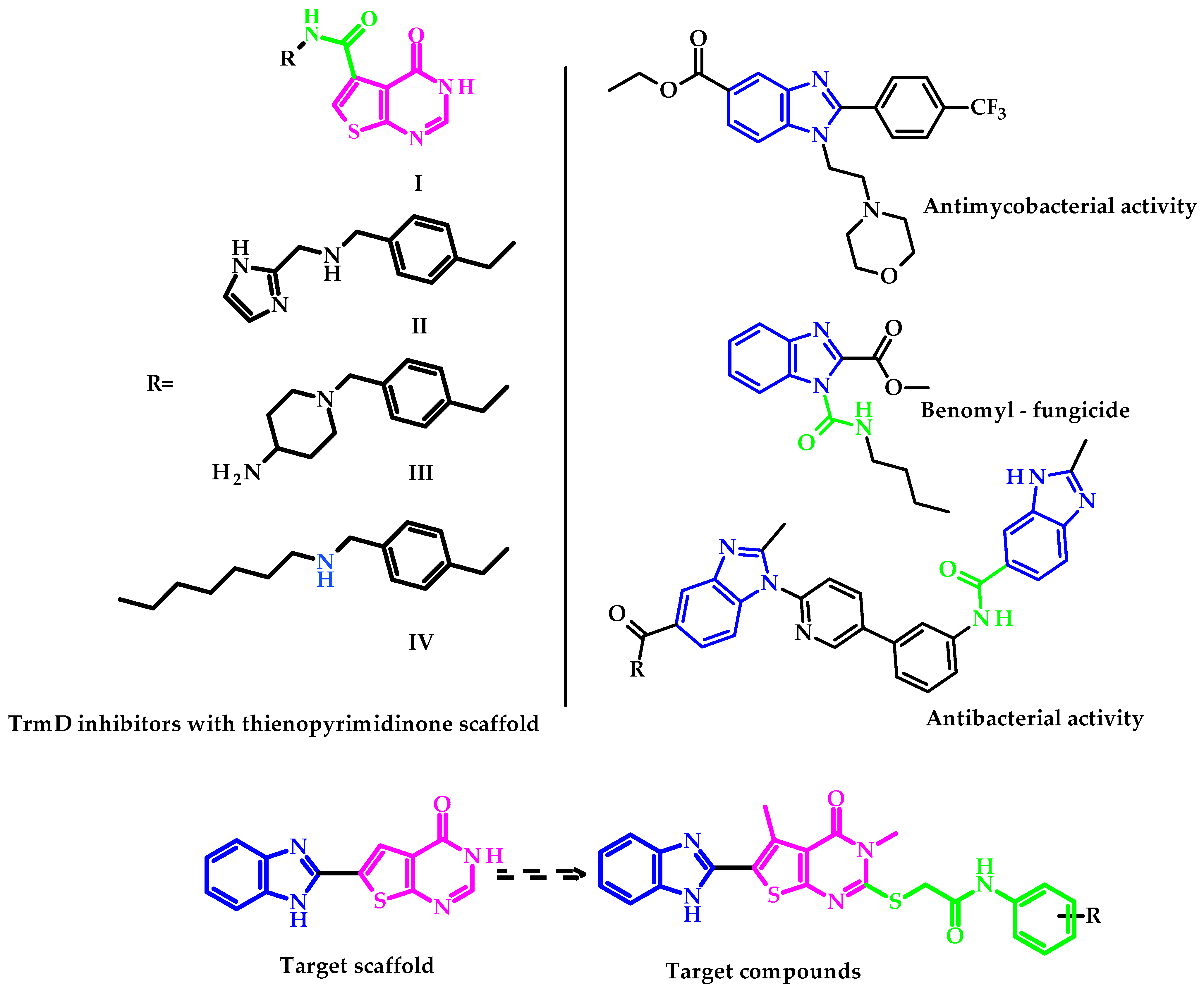
1. Introduction
2. Materials and Methods
2.1. General Information
2.2. Molecular Docking Study
2.3. Synthesis and Characterization of Compounds
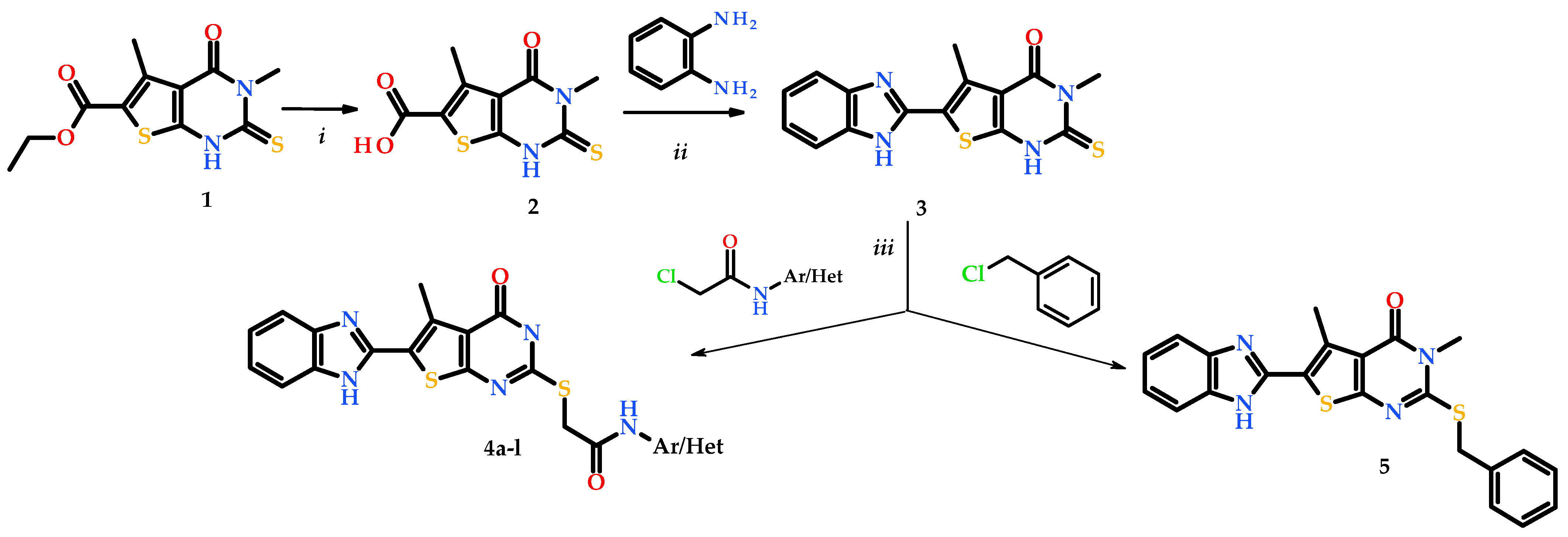
2.3.1. Synthesis of 3,5-Dimethyl-4-oxo-2-thioxo-1,2,3,4-tetrahydrothieno[2,3-d]pyrimidine-6-carboxylic Acid (2)
2.3.2. Synthesis of 6-(1H-Benzimidazol-2-yl)-3,5-dimethyl-2-thioxo-1H-thieno[2,3-d]pyrimidin-4-one (3)
2.3.3. General Method for Synthesis of 2-[6-(1H-Benzimidazol-2-yl)-3,5-dimethyl-4-oxo-thieno[2,3-d]pyrimidin-2-yl]sulfanyl-N-(aryl/heteryl)acetamides (4a-l) and 6-(1H-benzimidazol-2-yl)-2-(benzylsulfanyl)-3,5-dimethylthieno[2,3-d]pyrimidin-4(3H)-one (5)
2.4. Antimicrobial Activity Study
3. Results and Discussion
3.1. Docking Study
3.2. Chemical Synthesis
3.3. Antimicrobial Activity
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kousovista, R.; Athanasiou, C.; Liaskonis, K.; Ivopoulou, O.; Karalis, V. Association of Antibiotic Use with the Resistance Epidemiology of Pseudomonas aeruginosa in a Hospital Setting: A Four-Year Retrospective Time Series Analysis. Sci. Pharm. 2021, 89, 13. [Google Scholar] [CrossRef]
- Sirakanyan, S.N.; Geronikaki, A.; Spinelli, D.; Hakobyan, E.K.; Kartsev, V.G.; Petrou, A.; Hovakimyan, A.A. Synthesis and antimicrobial activity of new amino derivatives of pyrano[4″,3″:4′,5′]pyrido[3′,2′:4,5]thieno[3,2-d]pyrimidine. An. Acad. Bras. Cienc. 2018, 90, 1043–1057. [Google Scholar] [CrossRef]
- Sirakanyan, S.N.; Kartsev, V.G.; Geronikakim, A.; Spinellim, D.; Petrou, A.; Hakobyan, E.K.; Glamoclija, J.; Ivanov, M.; Sokovic, M.; Hovakimyan, A.A. Synthesis and Evaluation of Antimicrobial Activity and Molecular Docking of New N-1,3-thiazol-2-ylacetamides of Condensed Pyrido[3′,2′:4,5] furo(thieno)[3,2-d]pyrimidines. Curr. Top. Med. Chem. 2020, 20, 2192–2209. [Google Scholar] [CrossRef]
- Kumari, M.A.; Triloknadh, S.; Harikrishna, N.; Vijjulatha, M.; Venkata Rao, C. Synthesis, antibacterial activity, and docking studies of 1,2,3-triazole-tagged thieno[2,3-d]pyrimidinone derivatives. J. Heterocycl. Chem. 2017, 54, 3672–3681. [Google Scholar] [CrossRef]
- Khatri, T.T.; Shah, V.H. Effective microwave synthesis of bioactive thieno[2,3-d]pyrimidines. J. Chil. Chem. Soc. 2017, 62, 3354–3358. [Google Scholar] [CrossRef]
- Vlasov, S.V.; Kovalenko, S.M.; Chernykh, V.P.; Krolenko, K.Y. Synthesis of 5-methyl-4-thio-6-(1,3,4-oxadiazol-2-yl)thieno[2,3-d]pyrimidines and their antimicrobial activity study. J. Chem. Pharm. Res. 2014, 6, 22–27. [Google Scholar]
- Vlasov, S.V.; Kovalenko, S.M.; Osolodchenko, T.P.; Vlasov, V.S. The study of the antimicrobial activity of the derivatives of 6-(1,2,4-oxadizol-3-yl)- and 6-(2-aminothiazol-4-yl)thieno[2,3-d]pyrimidin-4-ones by the double dilution method. J. Org. Pharm. Chem. 2019, 17, 26–30. [Google Scholar] [CrossRef]
- Vlasov, S.V.; Kovalenko, S.N.; Osolodchenko, T.P.; Lenitskaya, E.B.; Chernykh, V.P. Synthesis and biological activity of 6-(1,3-benzoxazol-2-yl)-5-methylthieno-[2,3-d]pyrimidines. Pharm. Chem. J. 2018, 52, 510–514. [Google Scholar] [CrossRef]
- Settypalli, T.; Chunduri, V.R.; Kerru, N.; Nallapaneni, H.K.; Chintha, V.R.; Daggupati, T.; Yeguvapalli, S.; Wudayagiri, R. Design, synthesis, neuroprotective and antibacterial activities of 1,2,4-triazolo[3,4-b]1,3,4-thiadiazole linked thieno[2,3-d]pyrimidine derivatives and in silico docking studies. Chem. Select. 2019, 4, 1627–1634. [Google Scholar] [CrossRef]
- Kerru, N.; Settypalli, T.; Nallapaneni, H.; Chunduri, V.R. Novel Thienopyrimidine derivatives containing 1,2,4-triazoles and 1,3,4-oxadiazoles as potent antimicrobial activity. Med. Chem. 2014, 4, 623–629. [Google Scholar] [CrossRef]
- Triloknadh, S.; Rao, C.V.; Nagaraju, K.; Hari Krishna, N.; Ramaiah, C.V.; Rajendra, W.; Trinath, D.; Suneetha, Y. Design, synthesis, neuroprotective, antibacterial activities and docking studies of novel thieno[2,3-d]pyrimidine-alkyne Mannich base and oxadiazole hybrids. Bioorg. Med. Chem. Lett. 2018, 28, 1663–1669. [Google Scholar] [CrossRef]
- Abbas, S.E.; Gawad, N.M.A.; George, R.F.; Akar, Y.A. Synthesis, antitumor and antibacterial activities of some novel tetrahydrobenzo [4, 5] thieno [2, 3-d] pyrimidine derivatives. Eur. J. Med. Chem. 2013, 65, 195–204. [Google Scholar] [CrossRef] [PubMed]
- Mahmoud, M.R.; Abu El-Azm, F.S.M.; Ali, A.T.; Ali, Y.M. Design, synthesis, and antimicrobial evaluation of novel thienopyrimidines and triazolothienopyrimidines. Synth. Commun. 2015, 45, 982–992. [Google Scholar] [CrossRef]
- Chambhare, R.V.; Khadse, B.G.; Bobde, A.S.; Bahekar, R.H. Synthesis and preliminary evaluation of some N-[5-(2-furanyl)-2-methyl-4-oxo-4H-thieno[2,3-d]pyrimidin-3-yl]-carboxamide and 3-substituted-5-(2-furanyl)-2-methyl-3H-thieno[2,3-d]pyrimidin-4-ones as antimicrobial agents. Eur. J. Med. Chem. 2003, 38, 89–100. [Google Scholar] [CrossRef]
- Prabhakar, V.; Kondra, S.B.; Maddula, S.R.; Parandhama, G.; Latha, J. Synthesis, structural elucidation of novel thieno[2,3-d]pyrimidine core unit containing 1,2,4-triazoles and thiophenes as potent antimicrobial activity. Org. Chem. Curr. Res. 2016, 5, 169. [Google Scholar] [CrossRef]
- Saddik, A.A.; Kamal El-Dean, A.M.; El-Said, W.A.; Hassan, K.M.; Abbady, M.S. Synthesis, antimicrobial, and anticancer activities of a new series of thieno[2,3-d]pyrimidine derivatives. J. Heterocyclic. Chem. 2018, 55, 2111–2122. [Google Scholar] [CrossRef]
- Vilchèze, C.; Baughn, A.D.; Tufariello, J.; Leung, L.W.; Kuo, M.; Basler, C.F.; Alland, D.; Sacchettini, J.C.; Freundlich, J.S.; Jacobs, W.R. Novel Inhibitors of InhA Efficiently Kill Mycobacterium tuberculosis under Aerobic and Anaerobic Conditions. Antimicrob. Agents Chemother. 2011, 55, 3889–3898. [Google Scholar] [CrossRef]
- Li, S.-G.; Vilchèze, C.; Chakraborty, S.; Wang, X.; Kim, H.; Anisetti, M.; Ekins, S.; Rhee, K.Y.; Jacobs, W.R., Jr.; Freundlich, J.S. Evolution of a thienopyrimidine antitubercular relying on medicinal chemistry and metabolomics insights. Tetrahedron Lett. 2015, 56, 3246–3250. [Google Scholar] [CrossRef]
- Harrison, G.A.; Mayer Bridwell, A.E.; Singh, M.; Jayaraman, K.; Weiss, L.A.; Kinsella, R.L.; Aneke, J.S.; Flentie, K.; Schene, M.E.; Gaggioli, M.; et al. Identification of 4-Amino-Thieno[2,3-d]Pyrimidines as QcrB Inhibitors in Mycobacterium tuberculosis. mSphere 2019, 4, e00606-19. [Google Scholar] [CrossRef]
- Angula, K.T.; Legoabe, L.J.; Beteck, R.M. Chemical Classes Presenting Novel Antituberculosis Agents Currently in Different Phases of Drug Development: A 2010–2020 Review. Pharmaceuticals 2021, 14, 461. [Google Scholar] [CrossRef]
- Zhong, W.; Koayc, A.; Ngoc, A.; Lic, Y.; Naha, Q.; Wong, Y.H.; Chionh, Y.H.; Ngc, H.Q.; KohStentac, X.; Poulsenc, A.; et al. Targeting the bacterial epitranscriptome: Discovery of novel tRNA-(N1G37) methyltransferase (TrmD) inhibitors with antibiotic activity. ACS Infect. Dis. 2019, 5, 326–335. [Google Scholar] [CrossRef]
- Hill, P.J.; Abibi, A.; Albert, R.; Andrews, B.; Gagnon, M.M.; Gao, N.; Grebe, T.; Hajec, L.I.; Huang, J.; Livchak, S.; et al. Selective inhibitors of bacterial t-RNA-(N(1)G37) methyltransferase (TrmD) that demonstrate novel ordering of the lid domain. J. Med. Chem. 2013, 56, 7278–7288. [Google Scholar] [CrossRef] [PubMed]
- Zhong, W.; Pasunooti, K.K.; Balamkundu, S.; Wong, Y.H.; Nah, Q.; Gadi, V.; Gnanakalai, S.; Chionh, Y.H.; McBee, M.E.; Gopal, P.; et al. Thienopyrimidinone derivatives that inhibit bacterial tRNA (guanine37-N1)-methyltransferase (TrmD) by restructuring the active site with a tyrosine-flipping mechanism. J. Med. Chem. 2019, 62, 7788–7805. [Google Scholar] [CrossRef]
- Alasmary, F.A.S.; Snelling, A.M.; Zain, M.E.; Alafeefy, A.M.; Awaad, A.S.; Karodia, N. Synthesis and evaluation of selected benzimidazole derivatives as potential antimicrobial agents. Molecules 2015, 20, 15206–15223. [Google Scholar] [CrossRef]
- Picconi, P.; Hind, C.; Jamshidi, S.; Nahar, K.; Clifford, M.; Wand, M.E.; Sutton, J.M.; Rahman, K.M. Triaryl benzimidazoles as a new class of antibacterial agents against resistant pathogenic microorganisms. J. Med. Chem. 2017, 60, 6045–6059. [Google Scholar] [CrossRef]
- Antoci, V.; Cucu, D.; Zbancioc, G.; Moldoveanu, C.; Mangalagiu, V.; Amariucai-Mantu, D.; Aricu, A.; Mangalagiu, I. Bis-(imidazole/benzimidazole)-pyridine derivatives: Synthesis, structure and antimycobacterial activity. Future Med. Chem. 2020, 12, 207–222. [Google Scholar] [CrossRef]
- Al–Harthy, T.; Al-Sadi, A.M.; Zoghaib, W.; Moghadam, E.S.; Stoll, R.; Abdel-Jalil, R. Design, Synthesis and Bioactivity of Benzimidazole–2–Carbamates as Soil–Borne Anti–Fungal Agents. Chem. Proc. 2021, 3, 64. [Google Scholar] [CrossRef]
- Pereira, G.A.; Santos, L.H.; Wang, S.C.; Martins, L.C.; Villela, F.S.; Liao, W.; Dessoy, M.A.; Dias, L.C.; Andricopulo, A.D.; Costa, M.A.; et al. Benzimidazole inhibitors of the major cysteine protease of Trypanosoma brucei. Future Med. Chem. 2019, 11, 1537–1551. [Google Scholar] [CrossRef]
- Yoon, Y.K.; Ali, M.A.; Wei, A.C.; Choon, T.S.; Ismail, R. Synthesis and evaluation of antimycobacterial activity of new benzimidazole aminoesters. Eur. J. Med. Chem. 2015, 93, 614–624. [Google Scholar] [CrossRef] [PubMed]
- Miller, S.I. Antibiotic resistance and regulation of the gram-negative bacterial outer membrane barrier by host innate immune molecules. mBio 2016, 7, me01541-16. [Google Scholar] [CrossRef]
- Mookherjee, P.; Green, P.S.; Watson, G. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J. Comput. Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef]
- Protein Data Bank. Available online: http://www.rcsb.org/pdb/home/home.do (accessed on 4 April 2020).
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef]
- Vlasov, S.V.; Chernykh, V.P.; Osolodchenko, T.P. Synthesis and the antimicrobial activity of ethyl 3-alkyl-2-(alkylthio)-5-methyl-4-oxo-3,4-dihydrothieno[2,3-d]pyrimidine-6-carboxylate derivatives. News Pharm. 2015, 3, 3–8. [Google Scholar] [CrossRef][Green Version]
- Clinical and Laboratory Standards Institute (CLSI). Performance Standards for Antimicrobial Susceptibility Testing, 27th ed.; CLSI supplement M100-S26; CLSI: Wayne, PA, USA, 2017; p. 280. [Google Scholar]
- Ministry of Public Health of Ukraine. Bacteriological Control of Culture Media; Newsletter № 05.4.1/1670; Ministry of Public Health of Ukraine: Kiev, Ukraine, 2001; p. 45. [Google Scholar]
- Nekrasova, L.S.; Svita, V.M.; Glushkevich, T.G.; Tomchuk, V.V.; Zherebko, N.M.; Yanovs’ka, V.V. Methodological Guidelines “Determination of the Sensitivity of Microorganisms to Antibiotics”; № MB 9.9.5-143-2007; Ministry of Public Health of Ukraine: Kiev, Ukraine, 2007; p. 24. [Google Scholar]
- Baber, J.C.; Thompson, D.C.; Cross, J.B.; Humblet, C. GARD: A generally applicable replacement for RMSD. J. Chem. Inf. Model. 2009, 49, 1889–1900. [Google Scholar] [CrossRef] [PubMed]
- Vlasov, S.V.; Chernykh, V.P. Synthesis, antiinflammatory and antimicrobial activity of 6-(1H-benzimidazol-2-yl)-5-methyl-4-(alkylthio)thieno[2,3-d]pyrimidines. News Pharm. 2016, 3, 9–16. [Google Scholar] [CrossRef]
- Vianna, J.F.; Bezerra, K.S.; Oliveira, J.I.N.; Albuquerque, E.L.; Fulco, U.L. Binding energies of the drugs capreomycin and streptomycin in complex with tuberculosis bacterial ribosome subunits. Phys. Chem. Chem. Phys. 2019, 21, 19192–19200. [Google Scholar] [CrossRef] [PubMed]
- Germovsek, E.; Barker, C.I.; Sharland, M. What do I need to know about aminoglycoside antibiotics? Arch. Dis. Child. Educ. Pract. Ed. 2017, 102, 89–93. [Google Scholar] [CrossRef]
- Li, R.; Yuan, X.; Wei, J.; Zhang, X.; Cheng, G.; Wang, Z.A.; Du, Y. Synthesis and Evaluation of a Chitosan Oligosaccharide-Streptomycin Conjugate against Pseudomonas aeruginosa Biofilms. Mar. Drugs 2019, 17, 43. [Google Scholar] [CrossRef] [PubMed]
- Peterson, E.; Kaur, P. Antibiotic resistance mechanisms in bacteria: Relationships between resistance determinants of antibiotic producers, environmental bacteria, and clinical pathogens. Fron. Microbiol. 2018, 9, 2928. [Google Scholar] [CrossRef]
- Rodríguez-García, A.; Mares-Alejandre, R.E.; Muñoz-Muñoz, P.L.A.; Ruvalcaba-Ruiz, S.; González-Sánchez, R.A.; Bernáldez-Sarabia, J.; Meléndez-López, S.G.; Licea-Navarro, A.F.; Ramos-Ibarra, M.A. Molecular analysis of Streptomycin resistance genes in clinical strains of Mycobacterium tuberculosis and biocomputational analysis of the MtGidB L101F variant. Antibiotic 2021, 10, 807. [Google Scholar] [CrossRef]
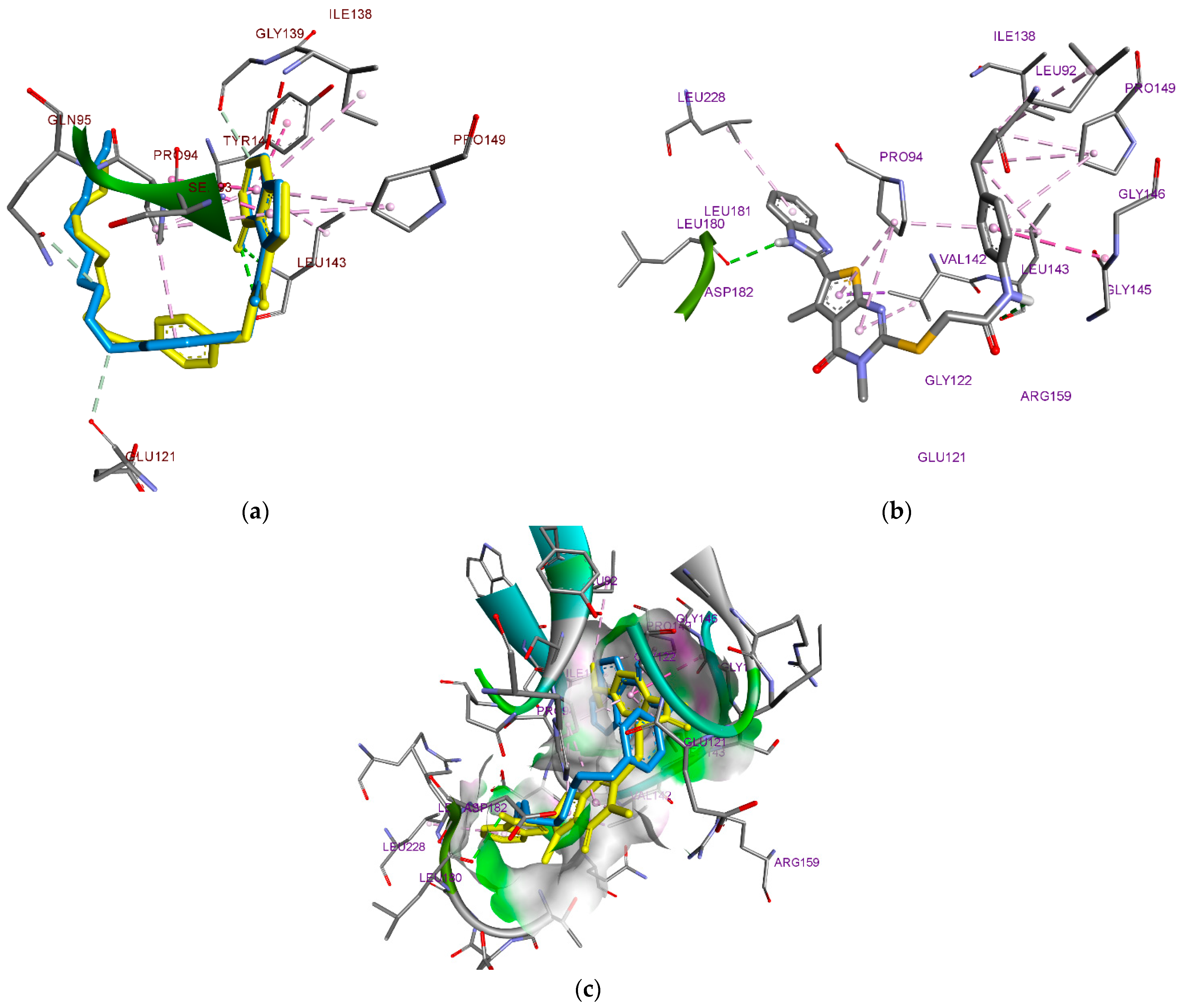
| Ligand | Binding Energy Kcal/Mol | Hydrophobic Interaction | Hydrogen Interaction | Other Interaction |
|---|---|---|---|---|
| Reference ligand | −8.2 | Tyr141, Ser93(2) #, Pro94 (4), Pro149(2), Ile138, Leu143, Gly 45, Gly146 | Leu143, Gln95, Glu121, Gly139, Asp182 | - |
| 3 | −7.9 | Val142(3) *, Glu121, Gly122 *, Pro94(3) | Arg159, Leu143 | - |
| 4a | −9.3 | Glu121(2), Gly122(2) *,Gly145, Gly146, Val142(3) *, Pro94(3), Leu143, Pro149 | Tyt120, Thr177 * | Asp182 (Pi-Anion) |
| 4b | −10.8 | Val142(2) *, Gly145, Gly146, Ile138, Leu143(2), Tyr141, Pro94, Pro149 | Arg159 *, Gly146, Tyr120, Ser144 *, Thr177(2) * | Asp159 * (Pi-Anion) |
| 4c | −10.6 | Glu121(2), Gly122(2) *, Gly145, Gly146, Pro94(3), Ile138, Leu143(2), Tyr141, Val142 *(2), Pro149 | Gln95, Leu92 *, Tyr91 * | Asp159 * (Pi-Cation) |
| 4d | −10.9 | Gly145, Gly146, Leu92 * Ile138, Pro149(2), Leu143(3), Ile138, Leu228 *, Pro94(3), Val142 *, Pro149 | Leu180*, Leu143 | - |
| 4e | −10.6 | Val142(2) *, Gly179, Leu180 *, Ser144, Gly145(2), Gly146, Pro94(3), Pro149, Leu228 | Leu180*, Leu143, Thr177 * | Leu92 *, Gly145 *, Ser93, (Halogen Fluorine) |
| 4f | −10.0 | Val142 *, Gly145(2), Gly146, Leu228 *, Pro94(3), Leu143, Pro149 | Leu180*, Leu143, Leu92 * | Asp182 |
| 4g | −9.9 | Val142(2) *, Gly145(2), Gly146, Leu228 *, Pro94(3), Leu143, Pro149 | Leu180*, Leu143, Leu92 * | - |
| 4h | −9.8 | SER93, Pro94(4), Glu121, Gly122*, Val142(2) *,Leu143, Pro149 | Gln95, Leu143, Gly139 *, Trp136 | Arg159(2) * |
| 4i | −10.4 | Val142(2) *, Gly145(2), Gly146, Pro94(4), Leu143(2), Leu228 *, Pro149 | Leu180 *, Leu143, | - |
| 4j | −10.3 | Val142(2) *, Gly145, Gly146, Pro94(4), Leu143(2), Leu228 *, Pro149 | Leu180 *, Leu143, | Ser93 (Halogen Fluorine) |
| 4k | −10.1 | Val142(3) *, Gly179*, Leu180*, Gly145, Gly146, Leu228 *, Pro94(3), Pro149 | Leu143, Glu121, Ser137* | Asp182 (Pi-Anion) |
| 4l | −9.5 | Val142(2) *, Gly145, Gly146, Pro94(2), Leu228 * | Leu143, Thr177 * | Asp182 (Pi-Anion) |
| 5 | −9.5 | Glu121(2), Gly122(2) *, Gly145, Gly146, Val142(2) *, Pro94(3), Leu143, Pro149 | Tyr120, Ser144 *, Thr177 * | Asp182 (Pi-Anion) |
| Compounds * | Average Diameter (mm) of Growth Inhibition Zone, Number of Test Repetitions n = 3 | |||||
|---|---|---|---|---|---|---|
| Gram-Positive Bacteria | Gram-Negative Bacteria | Fungus | ||||
| S. aureus ATCC 25923 | B. subtilis ATCC 6633 | E. coli ATCC 25922 | P. vulgaris ATCC 4636 | P. aeruginosa ATCC 27853 | C. albicans ATCC 653/885 | |
| 3 | 25, 25, 25 | 25, 26, 25 | 22, 21, 21 | 20, 21, 20 | 22, 21, 21 | 22, 21, 21 |
| 4a | 23, 23, 23 | 27, 27, 26 | 21, 21, 21 | 21, 20, 20 | 21, 21, 21 | 22, 21, 21 |
| 4b | 20, 21, 20 | 26, 26, 26 | 20, 21, 21 | 18, 18, 18 | 19, 20, 20 | 21, 21, 21 |
| 4c | 22, 21, 21 | 27, 26, 26 | 20, 20, 20 | 19, 18, 18 | 21, 21, 21 | 22, 22, 22 |
| 4d | 25, 26, 25 | 28, 28, 28 | 23, 24, 24 | 20, 21, 20 | 22, 21, 22 | 20, 21, 21 |
| 4e | 25, 25, 25 | 26, 25, 25 | 23, 21, 22 | 19, 19, 19 | 22, 21, 21 | 22, 21, 21 |
| 4f | 24, 24, 24 | 26, 25, 25 | 22, 22, 22 | 20, 19, 19 | 22, 21, 21 | 20, 21, 21 |
| 4g | 22, 23, 23 | 26, 26, 26 | 23, 23, 23 | 20, 20, 20 | 20, 21, 21 | 22, 22, 22 |
| 4h | 25, 26, 25 | 25, 26, 26 | 23, 24, 24 | 20, 20, 20 | 21, 21, 21 | 20, 20, 20 |
| 4i | 23, 23, 23 | 25, 25, 25 | 22, 20, 21 | 21, 19, 20 | 21, 21, 21 | 22, 23, 22 |
| 4j | 23, 23, 23 | 25, 26, 26 | 22, 21, 21 | 21, 19, 20 | 20, 20, 20 | 23, 23, 23 |
| 4k | 22, 22, 22 | 27, 27, 27 | 20, 21, 20 | 20, 19, 20 | 21, 20, 20 | 21, 21, 21 |
| 4l | 20, 21, 20 | 23, 24., 23 | 20, 19, 19 | 18, 18, 19 | 20, 21, 21 | 21, 21, 21 |
| 5 | 22, 23, 23 | 26, 26, 26 | 22, 22, 22 | 20, 20, 20 | 20, 21, 20 | 21, 21, 21 |
| Streptomycin | 26, 25, 25 | 27, 28, 27 | 25, 25, 25 | 22, 22, 22 | 22, 21, 21 | – ** |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vlasov, S.V.; Vlasova, O.D.; Severina, H.I.; Krolenko, K.Y.; Borysov, O.V.; Abu Sharkh, A.I.M.; Vlasov, V.S.; Georgiyants, V.A. Design, Synthesis and In Vitro Antimicrobial Activity of 6-(1H-Benzimidazol-2-yl)-3,5-dimethyl-4-oxo-2-thio-3,4-dihydrothieno[2,3-d]pyrimidines. Sci. Pharm. 2021, 89, 49. https://doi.org/10.3390/scipharm89040049
Vlasov SV, Vlasova OD, Severina HI, Krolenko KY, Borysov OV, Abu Sharkh AIM, Vlasov VS, Georgiyants VA. Design, Synthesis and In Vitro Antimicrobial Activity of 6-(1H-Benzimidazol-2-yl)-3,5-dimethyl-4-oxo-2-thio-3,4-dihydrothieno[2,3-d]pyrimidines. Scientia Pharmaceutica. 2021; 89(4):49. https://doi.org/10.3390/scipharm89040049
Chicago/Turabian StyleVlasov, Sergiy V., Olena D. Vlasova, Hanna I. Severina, Konstantin Yu. Krolenko, Oleksandr V. Borysov, Amjad Ibrahim M. Abu Sharkh, Vitaliy S. Vlasov, and Victoriya A. Georgiyants. 2021. "Design, Synthesis and In Vitro Antimicrobial Activity of 6-(1H-Benzimidazol-2-yl)-3,5-dimethyl-4-oxo-2-thio-3,4-dihydrothieno[2,3-d]pyrimidines" Scientia Pharmaceutica 89, no. 4: 49. https://doi.org/10.3390/scipharm89040049
APA StyleVlasov, S. V., Vlasova, O. D., Severina, H. I., Krolenko, K. Y., Borysov, O. V., Abu Sharkh, A. I. M., Vlasov, V. S., & Georgiyants, V. A. (2021). Design, Synthesis and In Vitro Antimicrobial Activity of 6-(1H-Benzimidazol-2-yl)-3,5-dimethyl-4-oxo-2-thio-3,4-dihydrothieno[2,3-d]pyrimidines. Scientia Pharmaceutica, 89(4), 49. https://doi.org/10.3390/scipharm89040049






