UV-Vis Spectrophotometry and UPLC–PDA Combined with Multivariate Calibration for Kappaphycus alvarezii (Doty) Doty ex Silva Standardization Based on Phenolic Compounds
Abstract
:1. Introduction
2. Materials and Methods
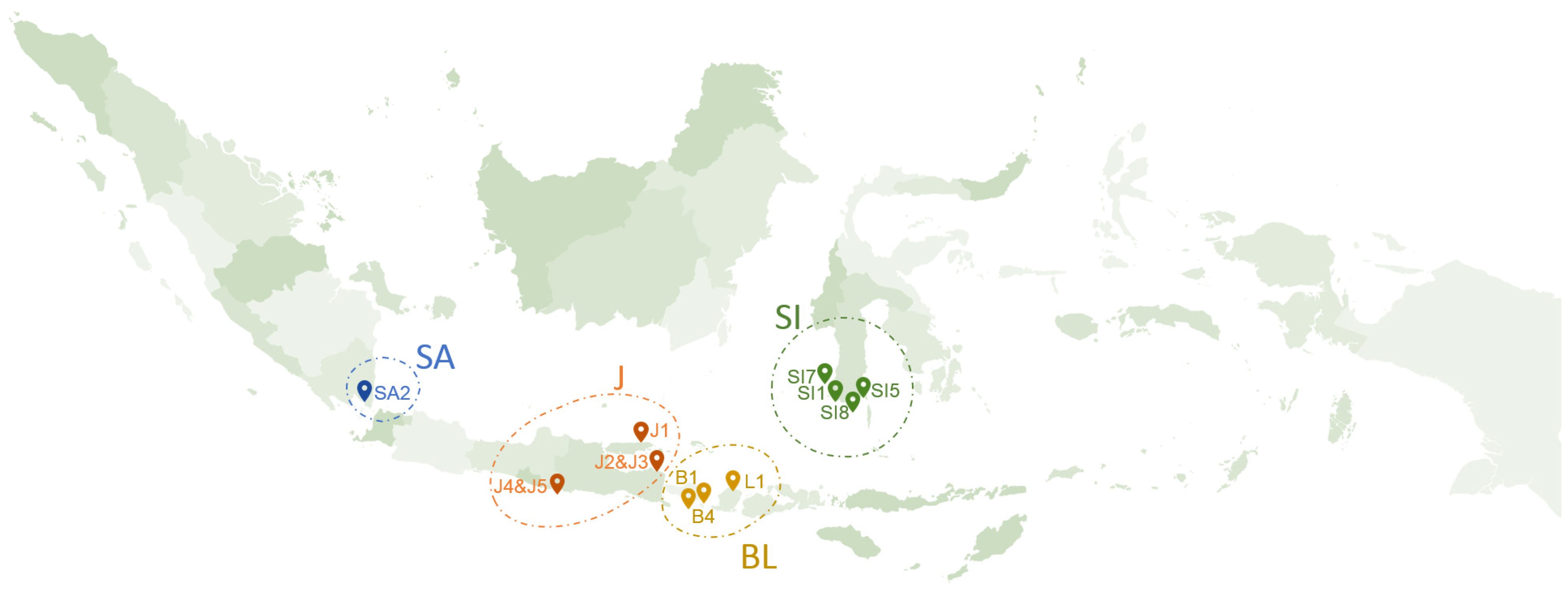
2.1. Sampling and Sample Preparation
2.2. UPLC-PDA
2.3. UV–Vis Spectra Acquisition
2.4. Multivariate Calibration Analysis
2.5. Real Sample Application
3. Result and Discussion
3.1. Identification of Phenolic Compounds in the K. alvarezii Extracts
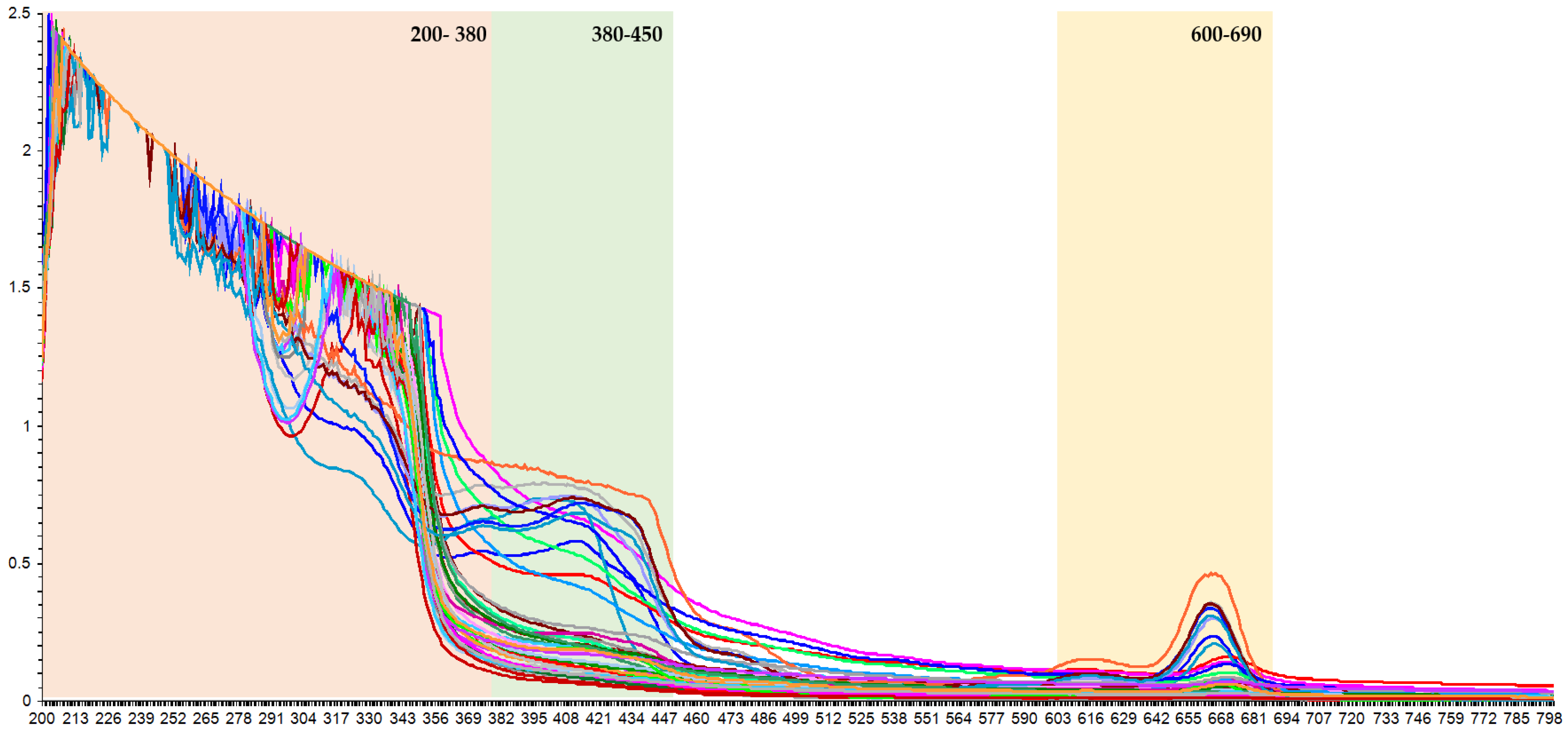
3.2. UV-Vis Spectra of Phenolic Compounds in the K. alvarezii Extracts
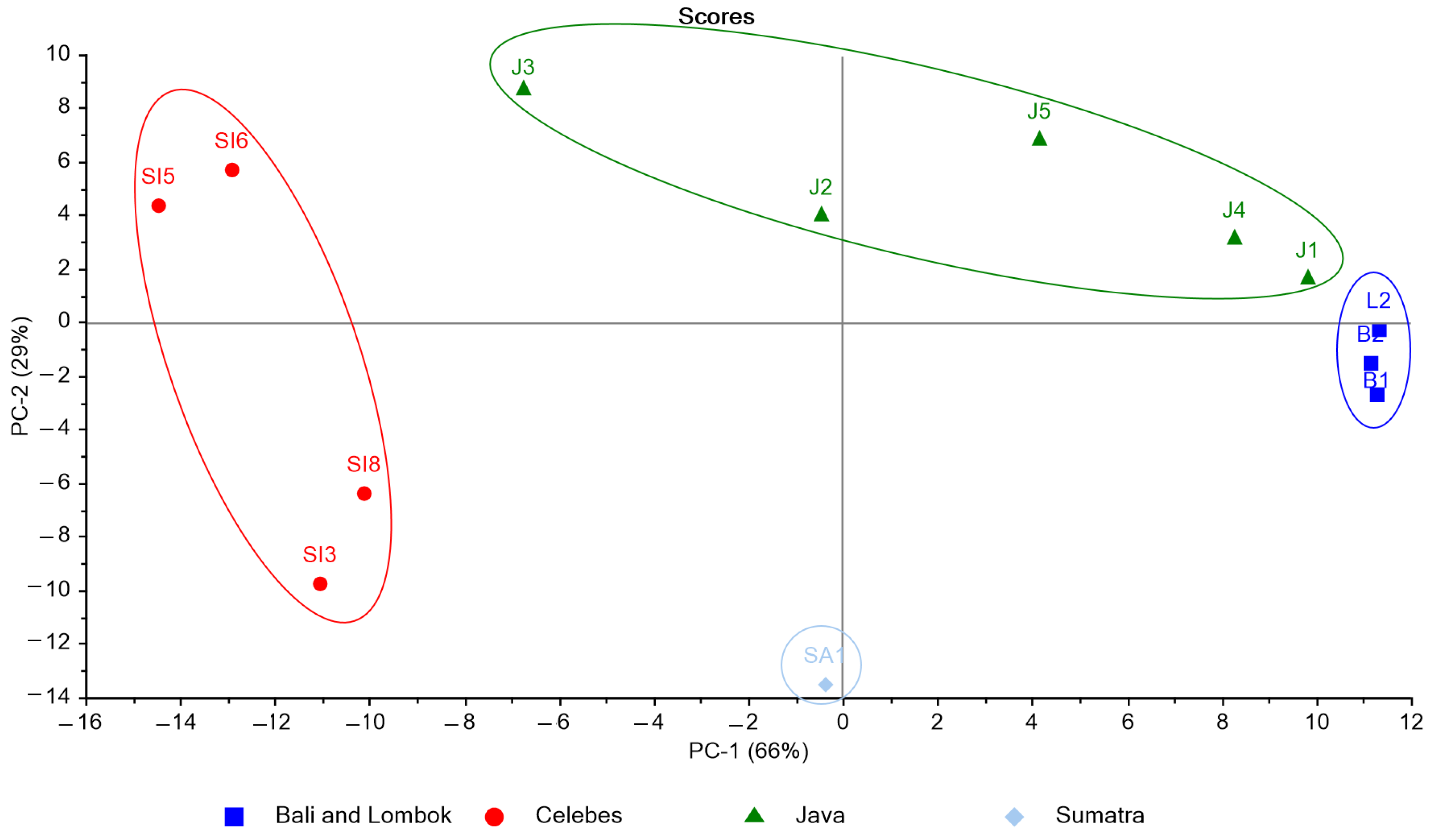
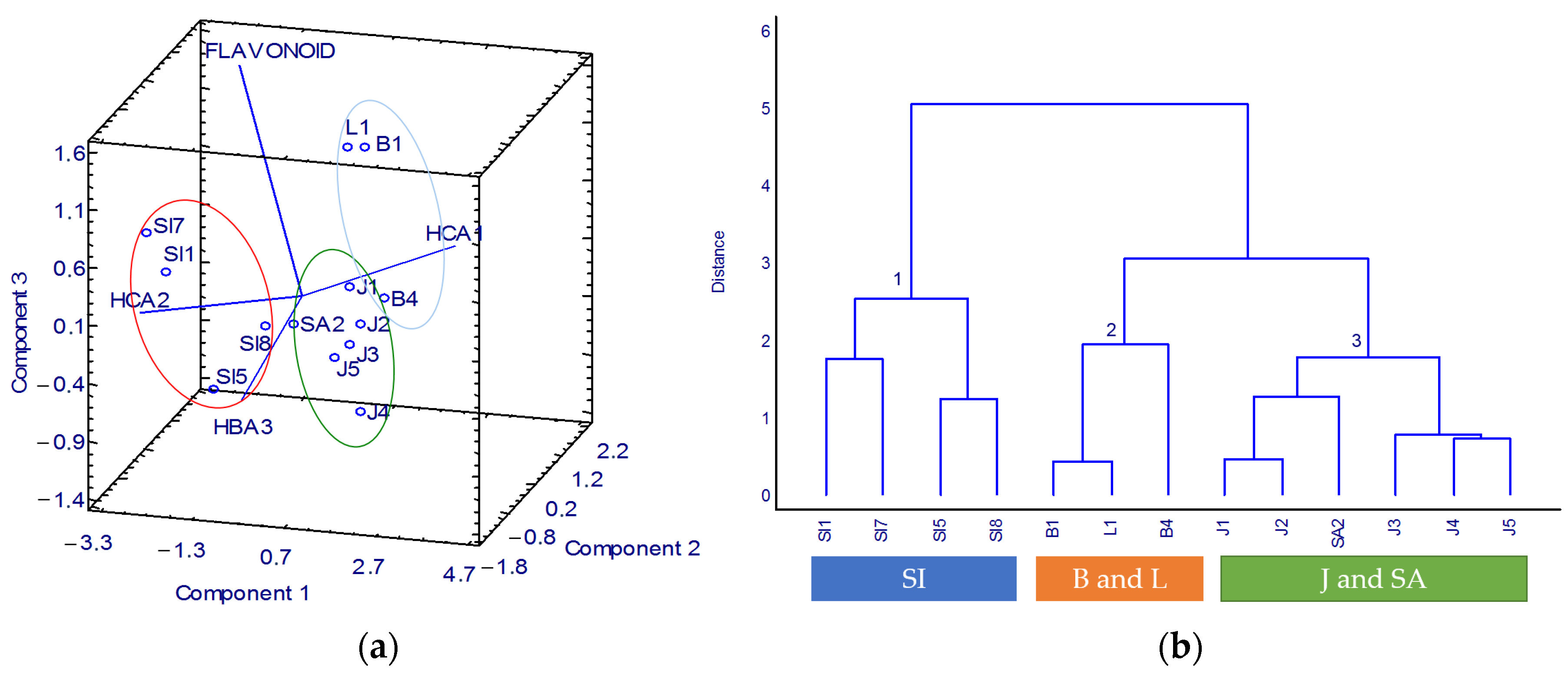
3.3. K. alvarezii Description Based on the Spectroscopic Properties
3.4. Calibration and Validation of PLS Regression
3.5. Phenolic Compounds Measurement in K. alvarezii
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- World Bank Group. Seaweed Aquaculture for Food Security, Income Generation and Environmental Health in Tropical Developing Countries. World Bank USA 2016. [Google Scholar] [CrossRef]
- Hayashi, L.; Hurtado, A.Q.; Msuya, F.E.; Bleicher-Lhonneur, G.; Critchley, A.T. Kappaphycus Farming: Prospects and Constraints; Springer Nature Switzerland: Cham, Switzerland, 2007; Volume 15, ISBN 9789048185696. [Google Scholar]
- Rengasamy, K.R.; Mahomoodally, M.F.; Aumeeruddy, M.Z.; Zengin, G.; Xiao, J.; Kim, D.H. Bioactive compounds in seaweeds: An overview of their biological properties and safety. Food Chem. Toxicol. 2020, 135, 111013. [Google Scholar] [CrossRef]
- Sumayya, S.; Murugan, K. Phytochemical screening, RP-HPLC and FTIR analysis of Kappaphycus alvarezii (Doty) Doty EX P.C Silva: Macro red algae. J. Pharmacogn. Phytochem. 2017, 6, 325–330. [Google Scholar]
- Stengel, D.B.; Connan, S.; Popper, Z.A. Algal chemodiversity and bioactivity: Sources of natural variability and implications for commercial application. Biotechnol. Adv. 2011, 29, 483–501. [Google Scholar] [CrossRef]
- Berrueta, L.A.; Alonso-Salces, R.M.; Héberger, K. Supervised pattern recognition in food analysis. J. Chromatogr. A 2007, 1158, 196–214. [Google Scholar] [CrossRef]
- Bae, I.K.; Ham, H.M.; Jeong, M.H.; Kim, D.H.; Kim, H.J. Simultaneous determination of 15 phenolic compounds and caffeine in teas and mate using RP-HPLC/UV detection: Method development and optimization of extraction process. Food Chem. 2015, 172, 469–475. [Google Scholar] [CrossRef] [PubMed]
- Setyaningsih, W.; Saputro, I.E.; Carrera, C.A.; Palma, M.; García-Barroso, C. Fast Determination of Phenolic Compounds in Rice Grains by Ultraperformance Liquid Chromatography Coupled to Photodiode Array Detection: Method Development and Validation. J. Agric. Food Chem. 2019, 67, 3018–3027. [Google Scholar] [CrossRef] [PubMed]
- Mandhania, S.; Pal, A.; Saharan, V. Simultaneous estimation of twenty eight phenolic compounds by a novel and expeditious method developed on quaternary ultra-performance liquid chromatography system with a photodiode array detector. Biomolecules 2020, 10, 6. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gumustas, M.; Kurbanoglu, S.; Uslu, B.; Ozkan, S.A. UPLC versus HPLC on Drug Analysis: Advantageous, Applications and Their Validation Parameters; Springer Nature Switzerland: Cham, Switzerland, 2013; Volume 76, ISBN 1033701324778. [Google Scholar]
- Palacios-Morillo, A.; Alcázar, Á.; De Pablos, F.; Jurado, J.M. Differentiation of tea varieties using UV-Vis spectra and pattern recognition techniques. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2013, 103, 79–83. [Google Scholar] [CrossRef]
- Riswahyuli, Y.; Rohman, A.; Setyabudi, F.M.C.S.; Raharjo, S. Indonesian wild honey authenticity analysis using attenuated total reflectance-fourier transform infrared (ATR-FTIR) spectroscopy combined with multivariate statistical techniques. Heliyon 2020, 6, e03662. [Google Scholar] [CrossRef]
- Song, X.C.; Canellas, E.; Asensio, E.; Nerín, C. Predicting the antioxidant capacity and total phenolic content of bearberry leaves by data fusion of UV–Vis spectroscopy and UHPLC/Q-TOF-MS. Talanta 2020, 213, 120831. [Google Scholar] [CrossRef]
- Boulet, J.C.; Ducasse, M.A.; Cheynier, V. Ultraviolet spectroscopy study of phenolic substances and other major compounds in red wines: Relationship between astringency and the concentration of phenolic substances. Aust. J. Grape Wine Res. 2017, 23, 193–199. [Google Scholar] [CrossRef]
- Aboulwafa, M.M.; Youssef, F.S.; Gad, H.A.; Sarker, S.D.; Nahar, L.; Al-Azizi, M.M.; Ashour, M.L. Authentication and discrimination of green tea samples using UV–vis, FTIR and HPLC techniques coupled with chemometrics analysis. J. Pharm. Biomed. Anal. 2019, 164, 653–658. [Google Scholar] [CrossRef]
- Shi, H.; Yu, P. Comparison of grating-based near-infrared (NIR) and Fourier transform mid-infrared (ATR-FT/MIR) spectroscopy based on spectral preprocessing and wavelength selection for the determination of crude protein and moisture content in wheat. Food Control 2017, 82, 57–65. [Google Scholar] [CrossRef]
- Rohaeti, E.; Muzayanah, K.; Septaningsih, D.A.; Rafi, M. Fast analytical method for authentication of chili powder from synthetic dyes using uv-vis spectroscopy in combination with chemometrics. Indones. J. Chem. 2019, 19, 668–674. [Google Scholar] [CrossRef]
- Liu, L.; Zuo, Z.-t.; Wang, Y.-z.; Xu, F.-r. A fast multi-source information fusion strategy based on FTIR spectroscopy for geographical authentication of wild Gentiana rigescens. Microchem. J. 2020, 159, 105360. [Google Scholar] [CrossRef]
- Sampaio, P.S.; Soares, A.; Castanho, A.; Almeida, A.S.; Oliveira, J.; Brites, C. Optimization of rice amylose determination by NIR-spectroscopy using PLS chemometrics algorithms. Food Chem. 2018, 242, 196–204. [Google Scholar] [CrossRef] [PubMed]
- Gam, D.H.; Kim, S.Y.; Kim, J.W. Optimization of ultrasound-assisted extraction condition for phenolic compounds, antioxidant activity, and epigallocatechin gallate in lipid-extracted microalgae. Molecules 2020, 25, 454. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rodríguez, E.; Arqués, J.L.; Rodríguez, R.; Nuñez, M.; Medina, M.; Talarico, T.L.; Casas, I.A.; Chung, T.C.; Dobrogosz, W.J.; Axelsson, L.; et al. The Role of UV-Visible Spectroscopy for Phenolic Compounds Quantification in Winemaking. Intech 2011, 32, 137–144. [Google Scholar]
- Widyartini, D.S.; Widodo, P.; Susanto, A.B. Thallus variation of Sargassum polycystum from Central Java, Indonesia. Biodiversitas 2017, 18, 1004–1011. [Google Scholar] [CrossRef]
- Tian, W.; Chen, G.; Gui, Y.; Zhang, G.; Li, Y. Rapid quantification of total phenolics and ferulic acid in whole wheat using UV–Vis spectrophotometry. Food Control 2021, 123, 107691. [Google Scholar] [CrossRef]
- Kim, S.; Kim, H. A new metric of absolute percentage error for intermittent demand forecasts. Int. J. Forecast. 2016, 32, 669–679. [Google Scholar] [CrossRef]
- Maricar, M.A.; Widiadnyana, P.; Arta Wijaya, I.W. Analysis of Data Mining for Forecasting Total Goods Delivery with Moving Average Method. Int. J. Eng. Emerg. Technol. 2017, 2, 7. [Google Scholar] [CrossRef] [Green Version]
- Liu, Y.; Liu, C.; Rubinato, M.; Guo, K.; Zhou, J.; Cui, M. An assessment of soil’s nutrient deficiencies and their influence on the restoration of degraded karst vegetation in Southwest China. Forests 2020, 11, 797. [Google Scholar] [CrossRef]
- Simatupang, N.F.; Pong-Masak, P.R.; Ratnawati, P.; Agusman; Paul, N.A.; Rimmer, M.A. Growth and product quality of the seaweed Kappaphycus alvarezii from different farming locations in Indonesia. Aquac. Rep. 2021, 20, 100685. [Google Scholar] [CrossRef]
| Experiment | Temperature (°C) | Solvent Composition (% Ethanol in Water) | Pulse Duty-Cycle (s−1) | Ultrasound Power (%) | Sample to Solvent Ratio |
|---|---|---|---|---|---|
| 1 | 35 | 90 | 0.6 | 100 | 1:20 |
| 2 | 35 | 70 | 0.6 | 100 | 1:30 |
| 3 | 35 | 70 | 1.0 | 60 | 1:10 |
| 4 | 35 | 70 | 0.6 | 20 | 1:10 |
| 5 | 35 | 50 | 0.6 | 60 | 1:10 |
| 6 | 35 | 70 | 0.2 | 60 | 1:10 |
| 7 | 35 | 50 | 1.0 | 60 | 1:20 |
| 8 | 35 | 70 | 0.6 | 100 | 1:10 |
| 9 | 35 | 70 | 0.6 | 60 | 1:20 |
| 10 | 35 | 90 | 0.2 | 60 | 1:20 |
| 11 | 35 | 70 | 0.2 | 60 | 1:30 |
| 12 | 35 | 50 | 0.2 | 60 | 1:20 |
| 13 | 35 | 70 | 0.2 | 20 | 1:20 |
| 14 | 35 | 70 | 0.2 | 100 | 1:20 |
| 15 | 35 | 90 | 1.0 | 60 | 1:20 |
| 16 | 35 | 70 | 0.6 | 60 | 1:20 |
| 17 | 35 | 70 | 1.0 | 60 | 1:30 |
| 18 | 35 | 50 | 0.6 | 100 | 1:20 |
| 19 | 35 | 90 | 0.6 | 60 | 1:30 |
| 20 | 35 | 70 | 0.6 | 60 | 1:20 |
| 21 | 35 | 70 | 0.6 | 60 | 1:20 |
| 22 | 35 | 50 | 0.6 | 60 | 1:30 |
| 23 | 35 | 70 | 1.0 | 100 | 1:20 |
| 25 | 35 | 90 | 0.6 | 20 | 1:20 |
| 26 | 35 | 70 | 1.0 | 20 | 1:20 |
| 27 | 35 | 50 | 0.6 | 20 | 1:20 |
| 28 | 35 | 90 | 0.6 | 60 | 1:10 |
| 29 | 35 | 70 | 0.6 | 20 | 1:30 |
| 30 | 35 | 70 | 0.6 | 60 | 1:20 |
| 31 | 60 | 50 | 0.6 | 60 | 1:20 |
| 32 | 60 | 70 | 0.6 | 60 | 1:10 |
| 33 | 60 | 70 | 0.6 | 100 | 1:20 |
| 34 | 60 | 70 | 0.6 | 20 | 1:20 |
| 35 | 60 | 90 | 0.6 | 60 | 1:20 |
| 36 | 60 | 70 | 1.0 | 60 | 1:20 |
| 37 | 60 | 70 | 0.2 | 60 | 1:20 |
| 38 | 60 | 70 | 0.6 | 60 | 1:30 |
| 39 | 10 | 70 | 0.6 | 60 | 1:10 |
| 40 | 10 | 90 | 0.6 | 60 | 1:20 |
| 41 | 10 | 70 | 0.2 | 60 | 1:20 |
| 42 | 10 | 50 | 0.6 | 60 | 1:20 |
| 43 | 10 | 70 | 0.6 | 20 | 1:20 |
| 44 | 10 | 70 | 1.0 | 60 | 1:20 |
| 45 | 10 | 70 | 0.6 | 100 | 1:20 |
| 46 | 10 | 70 | 0.6 | 60 | 1:30 |
| Phenolic Compound | Wavelength Range (nm) | Calibration | Cross-Validation | Prediction | |||
|---|---|---|---|---|---|---|---|
| R2C | RMSEC | R2CV | RMSECV | R2P | RMSEP | ||
| HCA1 | 200–800 | 0.9880 | 7.2880 | 0.8769 | 2.2531 | 0.9908 | 6.2261 |
| 200–380 | 0.9434 | 1.4940 | 0.8230 | 3.0557 | 0.9590 | 1.2882 | |
| 200–450 | 0.9044 | 8.4938 | 0.9578 | 1.4726 | 0.9897 | 7.0497 | |
| 200–450, 600–690 | 0.9937 | 5.4024 | 0.9848 | 8.6694 | 0.9954 | 4.6999 | |
| HCA2 | 200–800 | 0.9877 | 8.3960 | 0.9343 | 1.8086 | 0.9879 | 8.3820 |
| 200–380 | 0.8101 | 2.8974 | 0.6689 | 4.4350 | 0.7442 | 3.1775 | |
| 200–450 | 0.9579 | 1.5240 | 0.8882 | 2.2626 | 0.9414 | 1.8079 | |
| 200–450, 600–690 | 0.9927 | 6.3660 | 0.9875 | 1.0113 | 0.9954 | 5.0130 | |
| HBA | 200–800 | 0.8958 | 2.9950 | 0.6172 | 6.0450 | 0.8894 | 2.9120 |
| 200–380 | 0.9805 | 1.2110 | 0.9104 | 2.9620 | 0.9801 | 1.2120 | |
| 200–450 | 0.9641 | 1.7560 | 0.8326 | 4.8300 | 0.9749 | 1.3830 | |
| 200–450, 600–690 | 0.9184 | 2.6490 | 0.7837 | 4.5820 | 0.9056 | 2.6820 | |
| Flavonoid | 200–800 | 0.8516 | 5.4100 | 0.5552 | 1.0947 | 0.7816 | 6.8770 |
| 200–380 | 0.9662 | 3.1950 | 0.9692 | 8.2890 | 0.9625 | 3.1160 | |
| 200–450 | 0.8810 | 5.7590 | 0.6185 | 1.0133 | 0.8501 | 6.4670 | |
| 200–450, 600–690 | 0.8822 | 4.8230 | 0.6286 | 9.3500 | 0.8775 | 4.6640 | |
| Cultivation Site | Code | Sample Appearance | Origin Island | Peak Area (AU × min) | |||
|---|---|---|---|---|---|---|---|
| HCA1 Mean ± SD | HCA2 Mean ± SD | HBA Mean ± SD | Flavonoid Mean ± SD | ||||
| Puntondo | SI1 |  | Sulawesi | 85,704 ± 53 | 60,718 ± 56 | 31,615 ± 21 | 3219 ± 11 |
| Lembongan | B1 |  | Bali Lombok | 274,193 ± 14 | 60,818 ± 40 | 29,010 ± 61 | 4929 ± 14 |
| Sumenep | J1 |  | Java | 1,629,138 ± 15 | 487,439 ± 40 | 115,893 ± 30 | 10,572 ± 9 |
| Banyuwangi Grey | J2 |  | Java | 65,154 ± 57 | 6086 ± 10 | 16,203 ± 65 | 2232 ± 11 |
| Banyuwangi Red | J3 |  | Java | 784,322 ± 68 | 116,678 ± 75 | 144,096 ± 39 | 10,695 ± 15 |
| Pacitan Red | J4 |  | Java | 1,350,759 ± 24 | 331,726 ± 3 | 200,902 ± 67 | 5035 ± 32 |
| Pacitan Green | J5 |  | Java | 569,466 ± 14 | 165,605 ± 24 | 87,646 ± 59 | 4683 ± 17 |
| Lombok | L1 |  | Bali Lombok | 2,744,097 ± 69 | 866,786 ± 77 | 60,167 ± 20 | 7916 ± 49 |
| Bantaeng | SI5 |  | Sulawesi | 107,487 ± 22 | 62,827 ± 43 | 31,389 ± 37 | 1986 ± 36 |
| Tanakeke Island | SI7 |  | Sulawesi | 100,070 ± 31 | 85,310 ± 45 | 25,035 ± 14 | 3325 ± 58 |
| Djene Ponto | SI8 |  | Sulawesi | 50,680 ± 27 | 4146 ± 51 | 22,403 ± 48 | 839 ± 12 |
| Nusa Penida | B4 |  | Bali Lombok | 749,583 ± 27 | 208,965 ± 65 | 85,404 ± 38 | 4434 ± 16 |
| Teluk Pandan | SA2 |  | Sumatera | 2,678,169 ± 77 | 1,133,335 ± 40 | 277,646 ± 43 | 20,782 ± 58 |
| MAPE 1 (%) | 12 | 10 | 9 | 6 | |||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mutiarahma, S.; Putra, V.G.P.; Chaniago, W.; Carrera, C.; Anggrahini, S.; Palma, M.; Setyaningsih, W. UV-Vis Spectrophotometry and UPLC–PDA Combined with Multivariate Calibration for Kappaphycus alvarezii (Doty) Doty ex Silva Standardization Based on Phenolic Compounds. Sci. Pharm. 2021, 89, 47. https://doi.org/10.3390/scipharm89040047
Mutiarahma S, Putra VGP, Chaniago W, Carrera C, Anggrahini S, Palma M, Setyaningsih W. UV-Vis Spectrophotometry and UPLC–PDA Combined with Multivariate Calibration for Kappaphycus alvarezii (Doty) Doty ex Silva Standardization Based on Phenolic Compounds. Scientia Pharmaceutica. 2021; 89(4):47. https://doi.org/10.3390/scipharm89040047
Chicago/Turabian StyleMutiarahma, Selma, Venansius G. P. Putra, Weni Chaniago, Ceferino Carrera, Sri Anggrahini, Miguel Palma, and Widiastuti Setyaningsih. 2021. "UV-Vis Spectrophotometry and UPLC–PDA Combined with Multivariate Calibration for Kappaphycus alvarezii (Doty) Doty ex Silva Standardization Based on Phenolic Compounds" Scientia Pharmaceutica 89, no. 4: 47. https://doi.org/10.3390/scipharm89040047
APA StyleMutiarahma, S., Putra, V. G. P., Chaniago, W., Carrera, C., Anggrahini, S., Palma, M., & Setyaningsih, W. (2021). UV-Vis Spectrophotometry and UPLC–PDA Combined with Multivariate Calibration for Kappaphycus alvarezii (Doty) Doty ex Silva Standardization Based on Phenolic Compounds. Scientia Pharmaceutica, 89(4), 47. https://doi.org/10.3390/scipharm89040047








